Abstract
The engineering of a native-like articular cartilage (AC) is a long-standing objective that could serve the clinical needs of millions of patients suffering from osteoarthritis and cartilage injury. An incomplete understanding of the developmental stages of articular cartilage has contributed to limited success in this endeavor. Using next generation RNA sequencing we have transcriptionally characterized two critical stages of AC development in humans-i.e., immature neonatal and mature adult, as well as tissue-engineered cartilage derived from culture expanded human mesenchymal stem cells (hMSCs). We identified key transcription factors (TFs) and long non-coding RNAs (lncRNAs) as candidate drivers of the distinct phenotypes of these tissues. AGTR2, SCGB3A1, TFCP2L1, RORC, and TBX4 stand out as key transcription factors whose expression may be capable of reprogramming engineered cartilage into a more expandable and neonatal-like cartilage primed for maturation into biomechanically competent cartilage. We also identified that the transcriptional profiles of many annotated but poorly studied lncRNAs were dramatically different between these cartilages, indicating that lncRNAs may be also playing significant roles in cartilage biology. Key neonatal-specific lncRNAs identified include AC092818.1, AC099560.1, and KC877982. Collectively, our results suggest that tissue-engineered cartilage can be optimized for future clinical applications by the specific expression of TFs and lncRNAs.
Keywords: hMSCs, Long non-coding RNAs, Transcription factors, Cartilage, Tissue engineering
INTRODUCTION
Articular cartilage (AC) is the subtype of hyaline cartilage found on the articulating surfaces of joints. AC tissue is cell sparse, comprised mostly of water and highly organized and unique extracellular matrix (ECM), which is composed mostly of collagens and glycosaminoglycans. Its molecularly stratified arrangement allows AC to serve multiple mechanical demands dynamically and operate as a complex biomechanical composite.
AC injury, associated with pain and disability, remains a common clinical challenge that often leads to expensive and invasive total joint replacement surgery which may not necessarily relieve pain and immobility (Woolhead, Donovan, & Dieppe, 2005). Native cartilage demonstrates poor intrinsic regenerative ability due to its avascular, alymphatic, and aneural biology, and so developing clinical efforts have focused on means of producing grafts or supplementing native regenerative capacity. While various treatments can relieve pain and delay or eliminate the need for joint replacement, current therapies suffer several important limitations including ineffectiveness with large defects, long rehabilitation times, limited graft material availability, and donor site morbidity (Makris, Gomoll, Malizos, Hu, & Athanasiou, 2015). Many of these limitations might be ameliorated by improvements to tissue engineered grafts or cell-based therapies, both of which require a source of donor cells to create and maintain the ECM that gives cartilage its unique composite mechanical properties. Sources in use and in study include adult chondrocytes, human mesenchymal stem cells (hMSCs), and induced pluripotent stem cells, yet each source faces limitations and the resultant tissue engineered cartilages have failed to reproduce the structure and function of native AC (Somoza, Welter, Correa, & Caplan, 2014).
hMSCs are derived from perivascular cells found throughout the body and exhibit multipotency in vitro. hMSCs derived from bone marrow have been differentiated into cartilage by first aggregating the cells into a 3D pellet and then introducing a strong chondrogenic signaling molecule such as transforming growth factor β1 or β3 (TGF-β1 or TGF-β3) (Cleary, van Osch, Brama, Hellingman, & Narcisi, 2015; Johnstone, Hering, Caplan, Goldberg, & Yoo, 1998; Worster, Nixon, Brower-Toland, & Williams, 2000). Experimental evidence suggests that hMSCs produce a transient cartilage typical of endochondral processes rather than hyaline AC under current induction protocols. Therefore, the resulting in vitro hMSC-derived cartilage differs significantly from native AC in terms of structure, chemical composition, cell phenotype, and function (Somoza et al., 2014).
Models using this or similar chondrogenic induction protocols have been used to study the generation of cartilage by following the high expression of the classical markers for hyaline and AC, COL2A1 and ACAN. These molecules, especially type II collagen, comprise the bulk of the dry mass of AC but are not sufficient to identify a cartilage as AC (Somoza et al., 2018). The tissue produced by these processes lacks the functional stratification of native cartilage, demonstrates a markedly different transcriptional profile, and is ultimately of limited usefulness to tissue engineering efforts due to inferior biomechanical properties. Despite these limitations, hMSC-derived cartilage is a promising source of graft material for cartilage tissue engineering and is widely studied for this purpose. Thus, the identification of key molecules that can reprogram hMSCs into clinically useful cartilage would have enormous therapeutic uses and benefits (Somoza et al., 2014).
Neonatal AC is functional but is not yet weight-bearing and has not yet stratified into distinct functional zones like mature, adult AC (Brama, TeKoppele, Bank, Barneveld, & van Weeren, 2002; Brommer et al., 2005; Hunziker, Kapfinger, & Geiss, 2007). The cellular density of neonatal AC is much greater than in mature tissue, and it is primed to grow and expand as the body undergoes the rapid weight gain and increasing functional demands of early childhood development (Gannon, Nagel, Bell, Avery, & Kelly, 2015; Jadin, Bae, Schumacher, & Sah, 2007). A key result of this expansion and maturation takes the form of zone dependent collagen fibril organization and stiffening, which ultimately provide the tissue with its complex composite mechanical properties (Gannon et al., 2015). The rapid pace of AC expansion in early development also raises the possibility that it may be the appropriate phenotype to alleviate another long-standing issue in the field of AC grafting – failure of the graft tissue to integrate with the surrounding native tissue. Collectively these properties make neonatal AC an interesting target phenotype for a tissue engineered cartilage, as that tissue could begin as a homogenous, cell-dense cartilage before potentially being expanded and matured in vitro or even in vivo. This strategy for producing a more native-like AC graft than currently available may prove more feasible than efforts to directly produce mature, stratified cartilage.
The structural and histological differences between neonatal, adult, and hMSC-derived cartilage are likely driven by differences in gene expression. Cell and tissue identity is largely determined by the expression of transcription factors (TFs), and recent studies have also demonstrated key role of long non-coding RNAs (lncRNAs) (Glinsky et al., 2018). LncRNAs are transcripts greater than 200 nucleotides that do not encode proteins and have been shown to play numerous roles in many areas of cellular biology, particularly in epigenetic regulation (Jarroux, Morillon, & Pinskaya, 2017), cancer (Forrest & Khalil, 2017), and recently human development (Glinsky et al., 2018). As it becomes increasingly apparent that these molecules are critical elements of human physiology, we chose here to focus on identifying key lncRNAs in AC biology so that their roles may be further investigated. One subset of lncRNAs known as long intergenic non-coding RNAs (lincRNAs) has been shown to have high tissue specificity and evolutionary conservation, as well as developmental specificity, and therefore may play critical roles in cartilage development and homeostasis (Ayupe et al., 2015; Khalil et al., 2009; Morris & Mattick, 2014; Necsulea et al., 2014; Ulitsky, Shkumatava, Jan, Sive, & Bartel, 2011; Washietl, Kellis, & Garber, 2014). We also focus on identification of TFs because as direct regulators of transcription, they are likely to play critical roles in driving AC cellular phenotypes. With a comprehensive characterization of lncRNAs and TFs in native AC and hMSC-derived cartilage, we aim to identify potential candidate molecular drivers of AC that can be introduced into hMSCs with the goal of developing native-like engineered cartilage for future clinical applications.
Here, we utilize next generation RNA sequencing (RNA-seq) to identify the molecular fingerprints of native neonatal and adult AC and compare it to hMSCs undergoing chondrogenic differentiation in 3D pellet culture induced by TGF-β1 in vitro. Our analyses reveal key potential roles of TFs and lncRNAs in both distinguishing these tissues and coordinating the transcriptional changes responsible for their respective biomechanical properties.
METHODS
Human Mesenchymal Stem Cells Isolation and Expansion
hMSCs were isolated as previously described (Lennon & Caplan, 2006). Briefly, bone marrow was collected from the posterior-superior iliac crest of healthy adult donors ages 30–39 (n=3). The marrow was collected using a procedure reviewed and approved by the University Hospitals of Cleveland Institutional Review Board; informed consent was obtained from all de-identified donors. The marrow was centrifuged for 15min at 480g on a preformed Percoll gradient (density 1.03–1.12 g/mL; Sigma. St. Louis, MO). The nucleated cells of the monocyte light-cell fraction were collected, rinsed with serum-containing Dulbecco’s modified Eagle’s medium-low glucose (DMEM-LG; Invitrogen, Grand Island, NY), centrifuged at 450g, re-suspended, counted, and seeded in standard tissue culture vessels at 1.0×105 cells per cm2. Cells were expanded in DMEM-LG supplemented with 10% FBS (selected batch), supplemented with FGF2 (10 ng/ml). Cells were used at passage 2 for chondrogenic differentiation.
hMSC Chondrogenic Differentiation
hMSCs were cultured in cell aggregates in complete chondrogenic medium consisting of high glucose DMEM supplemented with 1% ITS+, 0.1μM dexamethasone, 1mM sodium pyruvate, 120mM ascorbic acid-2 phosphate, 100mM nonessential amino acids, and 10ng/mL TGF-β1 (Lennon, Schluchter, & Caplan, 2012). Chondrogenic pellets were harvested at different time points (3, 7, 10, 14, 21, and 28 days). Day 0 samples were harvested from monolayer cultures on passage 2, just before pellet culture was initiated.
Tissue Dissection
Neonatal femoral condyle AC was carefully dissected from both knees of de-identified 1-month-old cadaveric specimens (n=3) procured from consented guardians of the human tissue donors (Allosource, Centennial, CO). The tissue samples were stored at −80C suspended in RNAlater reagent (Qiagen, Valencia, CA). Snap frozen adult AC was acquired (Articular Engineering, Northbrook, IL) and stored at −80C in RNAlater before processing (Table 1).
Table 1:
Study Specimens
| Sample Name |
Age | Gender | Race | Cause of Death |
|---|---|---|---|---|
| Adult 1 | 35 years | Female | Caucasian | Alprazolam overdose |
| Adult 2 | 20 years | Male | Hispanic | Gunshot wound |
| Adult 3 | 20 years | Male | Hispanic | Acetaminophen overdose |
| Neonate 1 | 1 month | Male | Unknown | Unknown |
| Neonate 2 | 1 month | Male | Unknown | Unknown |
| Neonate 3 | 1 month | Male | Unknown | Unknown |
RNA Isolation from hMSC-derived Pellets and Primary Tissues
Adult and neonatal AC tissues were snap frozen in liquid nitrogen and pulverized by mechanical compression before static rotor homogenization in TRIzol (Life Technologies) at 4C for 5 minutes. TRIzol-chloroform phase separation of nucleic acids preceded column isolation of RNA by Qiagen RNeasy Mini Kit. RNA quality was assessed by high sensitivity Agilent Bioanalyzer (Bioanalyzer RNA 6000 pico) and RIN scores ranged from 5.2 to 7.4. hMSCs and hMSC-derived pellets were processed the same way except were not snap frozen prior to static rotor homogenization, instead being frozen at −20C in TRIzol. RIN scores from hMSCs and pellets ranged from 8.5 to 10.0.
RNA-Sequencing
Low input total RNA (5ng-1000ng) was used for 150bp paired-end stranded RNA-seq at Novogene. Strand specific cDNA (insert size: 250–300bp) libraries were constructed using the NEBNext Ultra Kit for following poly-T oligo-attached magnetic bead purification. Quality assessed by insert size (Agilent 2100).
Data Analysis
TPM quantification was performed by Salmon (v0.9.1) quasimapping against ENSEMBL GRCh38 (v93) cDNA and ncRNA indexes. Differential expression analysis was performed by DESeq2 after Salmon quantifications were condensed from transcript level to gene level. 2,556 transcription factors were defined by UNIPROT designation. Raw FASTQ and processed data files are deposited in GEO under accession number GSE128554. PCA was performed by prcomp of the stats R package (v 3.5.1) on all the top 2,859 transcripts by average TPM expression. Transcripts for which no replicate expressed at ≥ 1 TPM were excluded. Heat maps were built by GeneCluster (v3.0) and TreeViewer (v1.1.6r4) using differentially expressed genes with an adjusted p-value ≤ 0.05. Hierarchical clustering was determined by uncentered Pearson correlation with average linkage. PANTHER (version 14.1, released 2019.03.12) GO pathway analysis (PANTHER Overrepresentation Test, released 2019.05.17) conducted using differentially expressed genes with adjusted p-values less than 0.05 with default settings (Fisher’s exact test with calculated false discovery rate correction against the Homo sapiens all genes reference).
RESULTS
Neonatal and Adult Articular Cartilage Are Highly Distinct
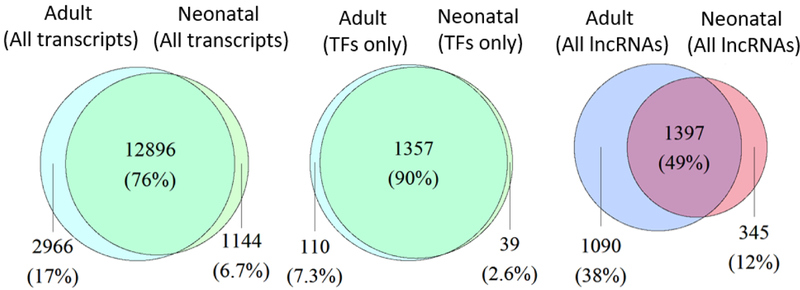
Due to advances in gene expression analysis, it has become clear that AC at different sites in the human body and during human development is likely to have distinct, site-specific gene expression profiles. However, these differences are yet to be examined. To that end, we began our studies by examining gene expression differences between neonatal and adult AC. We isolated and sequenced RNA from AC tissues of de-identified cadaverous human donors (Table 1). Transcripts were considered consistently expressed if each replicate quantified to at least 1 transcript per million (TPM); using this criteria, developmentally immature (neonatal) and mature (adult) AC tissues share the consistent expression of 12,896 transcripts (76%) and demonstrate 1,144 and 2,966 uniquely and consistently expressed transcripts respectively for a total of 4,110 developmentally specific transcripts in these tissues (Fig 1A). Interestingly, mature and immature AC share 90% of TF expression and demonstrate only 149 developmentally specific TFs, yet are much more divergent in expression of long noncoding RNAs as these tissues share only 49% of lncRNAs and express 1,435 developmentally specific lncRNAs (Fig 1A). Differential expression analysis distinguishes 2,792 transcripts, including 247 TFs and 200 lincRNAs, as significant differentially expressed genes (DEGs) between neonatal and adult AC (Fig 1B) (p<0.05).
Figure 1. Gene expression comparison of human adult vs. neonatal cartilage.
A. Venn diagrams comparing all transcripts (left), TFs (middle), and lincRNAs (right) consistently expressed (all replicates TPM ≥ 1) in adult and neonatal AC.
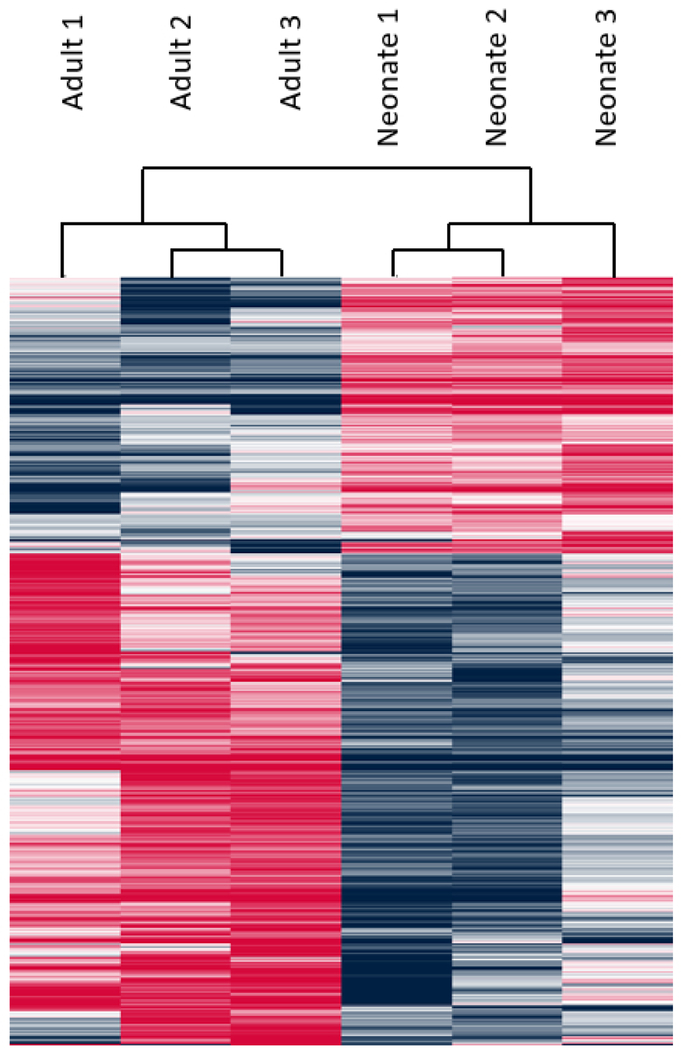
B. Heat map of all differentially expressed genes between adult and neonatal AC samples with hierarchical clustering of biological replicate groupings.
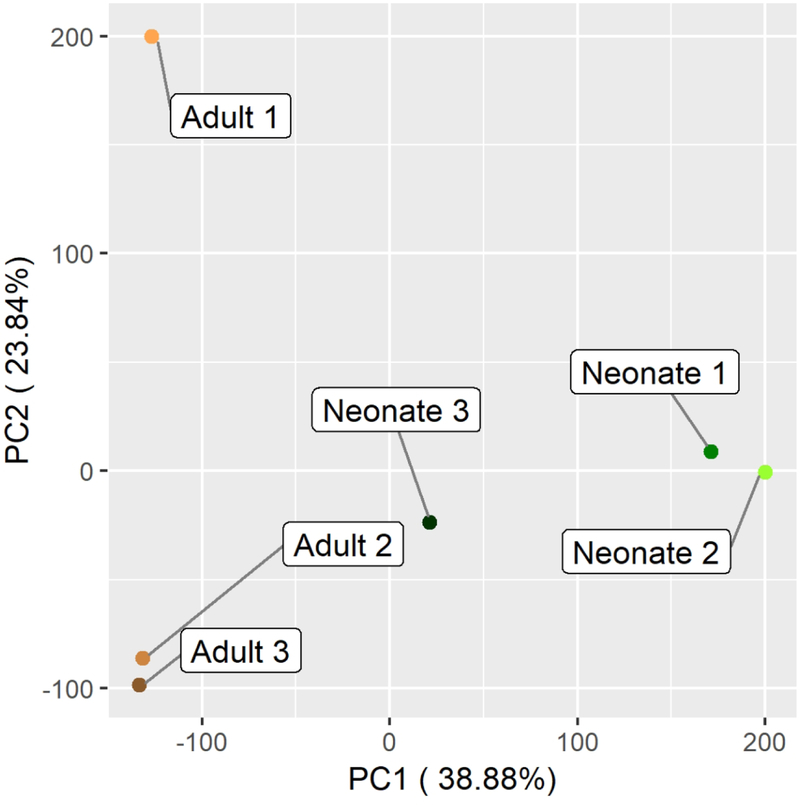
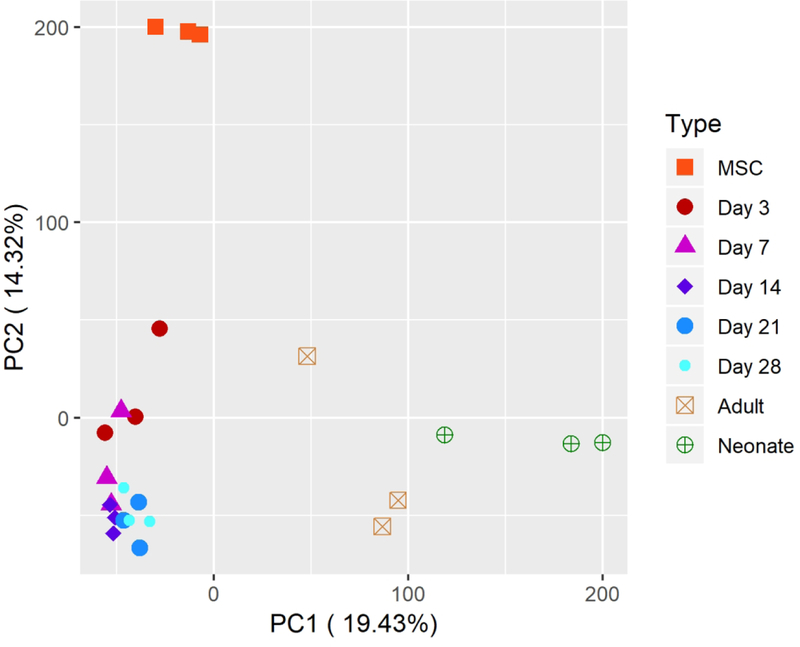
C. Two dimensional principal component analysis comparing likeness of adult and neonatal AC by transcriptional profile of transcripts with at least one sample expressing at ≥ TPM of 1.
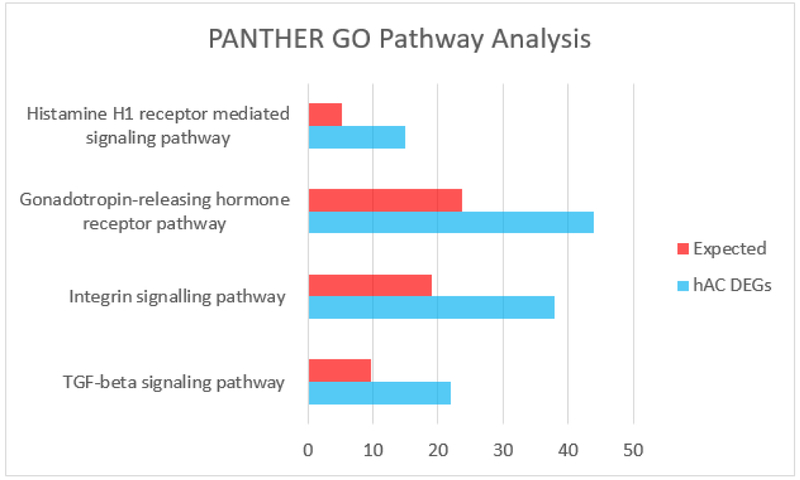
D. PANTHER GO pathway analysis demonstrating the pathways overrepresented by differentially expressed genes.
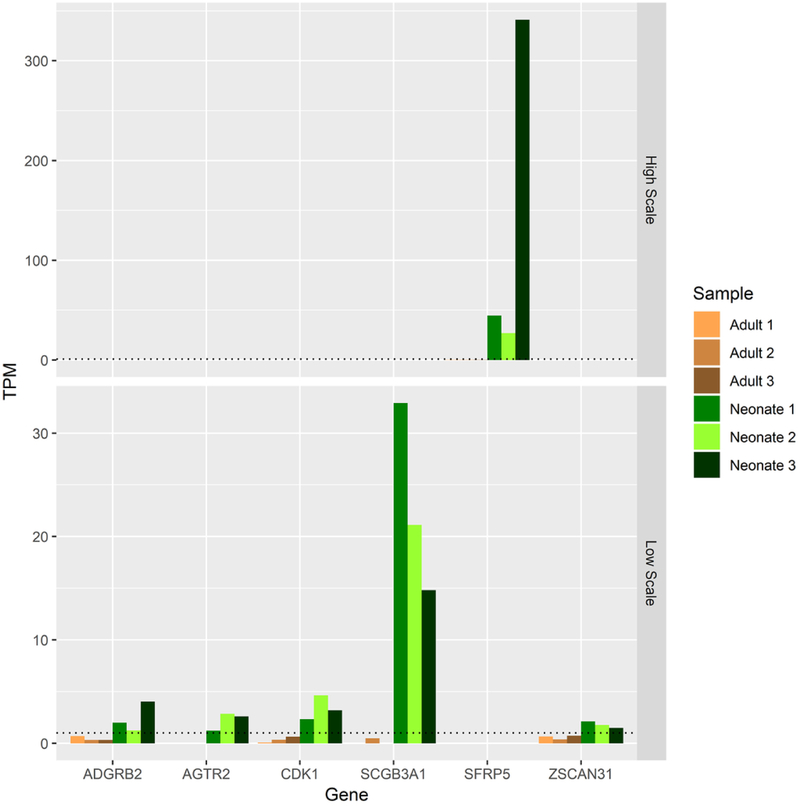
E. Expression patterns of potential molecular driver TFs for neonatal AC as defined by a minimum expression of 1 TPM in each neonatal replicate and ≤ TPM of 1 in all adult replicates.
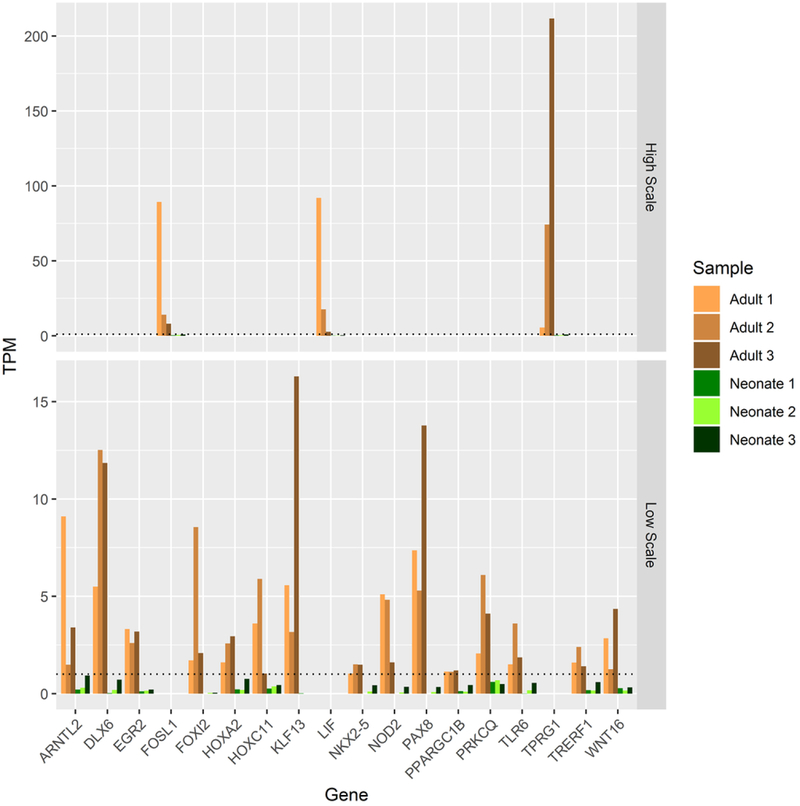
F. Expression patterns of potential molecular driver TFs for adult AC as defined by a minimum expression of 1 TPM in each adult replicate and ≤ TPM of 1 in all neonatal replicates.
Principal component analysis (PCA) confirms the similarity of these biological replicates within their respective groups (Fig 1C). PANTHER pathway overrepresentation testing revealed several signaling pathways enriched in neonatal upregulated genes including TGF-β signaling – a known pathway critical to chondrogenic development (Fig 1D) (Mi, Muruganujan, Ebert, Huang, & Thomas, 2019; Mi & Thomas, 2009). Another identified pathway, integrin signaling, is crucial to chondrocyte development and maintenance, as collagen, fibronectin, and laminin-specific integrin heterodimers are expressed in chondrocytes and provide cell-ECM interactions (Song & Park, 2014).
To identify molecular drivers potentially responsible for the differences between these tissues, we initially focused on TFs consistently expressed or not expressed in each biological replicate of neonatal and adult AC, respectively. We identified six TFs that followed this pattern of conserved neonatal-specific expression: SFRP5, CDK1, ADGRB2, ZSCAN31, SCGB3A1, and AGTR2 (Fig 1E). The inverse situation, TFs consistently expressed in adult but not expressed in neonatal biological replicates, revealed 20 TFs (Fig 1F). Additionally, we identified eight lncRNAs with dramatic expression differences between neonatal and adult AC using stringent criteria to highlight only RNAs most likely to be functional (positive lncRNA expression ≥ 5 TPM in each replicate, negative lncRNA expression ≤ 1 TPM) (Fig 5A).
Figure 5. Uniquely expressed lncRNAs in either neonatal or adult cartilage and example co-expression of lncRNAs with genomic co-occupant cartilage-related protein-coding gene.
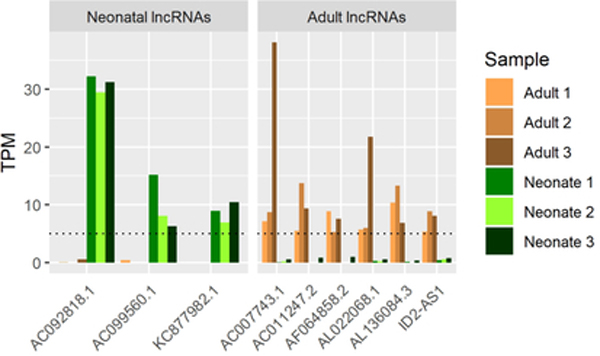
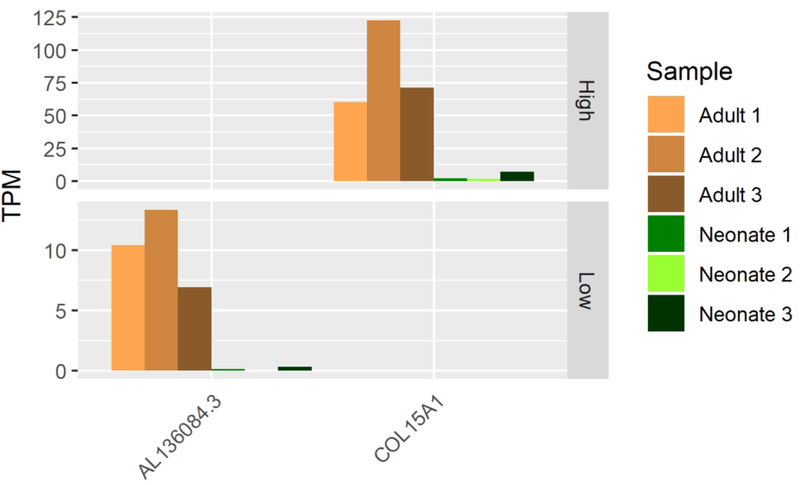
A. lncRNAs consistently expressed under stringent criteria (TPM≥5) in each primary AC but not in the other (TPM <1).
B. Co-expression of COL15A1 with AL136084.3.
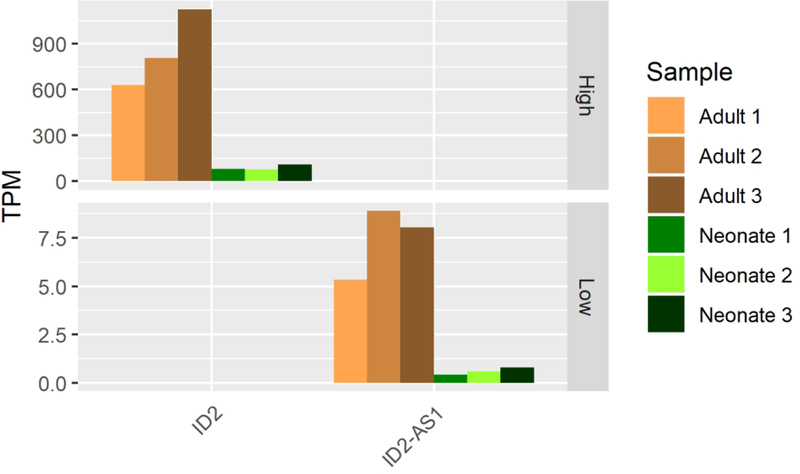
C. Co-expression of ID2 with ID2-AS1.
Chondrogenic Differentiation of hMSCs Demonstrates Dramatic Early Changes and Large Differences in lncRNA Profiles
3D pellet culture of hMSCs in a defined medium containing TGF-β1 creates a form of cartilage that highly expresses classical markers of hyaline cartilage but eventually expresses markers of hypertrophy, a characteristic of transient cartilage that has limited usefulness in AC repair clinical application (Johnstone et al., 1998; Pelttari et al., 2006). Thus, we analyzed the kinetics of this process to ultimately help determine how it might be modified to improve tissue engineered cartilage outcomes. Specifically, we analyzed hMSCs at day 0, and at specific days post induction including days 3, 7, 14, 21 and 28. These time points were chosen because they represent key phases of the pellet culture differentiation process, wherein early changes are rapid and dramatic before transcriptional changes slow and pellet maturation is reached by day 21 and hypertrophy by day 28 (Somoza et al., 2018; Sorrell, Somoza, & Caplan, 2018).
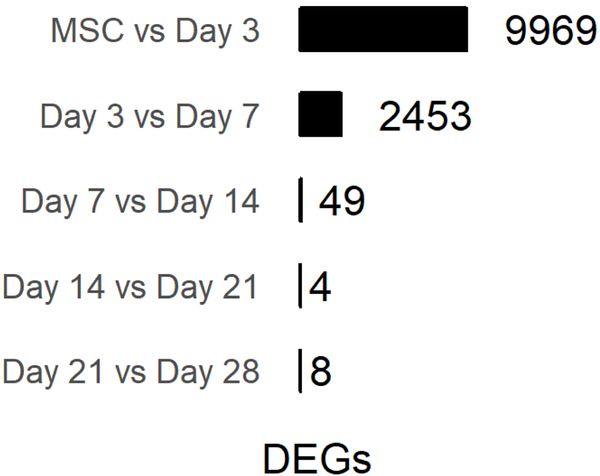
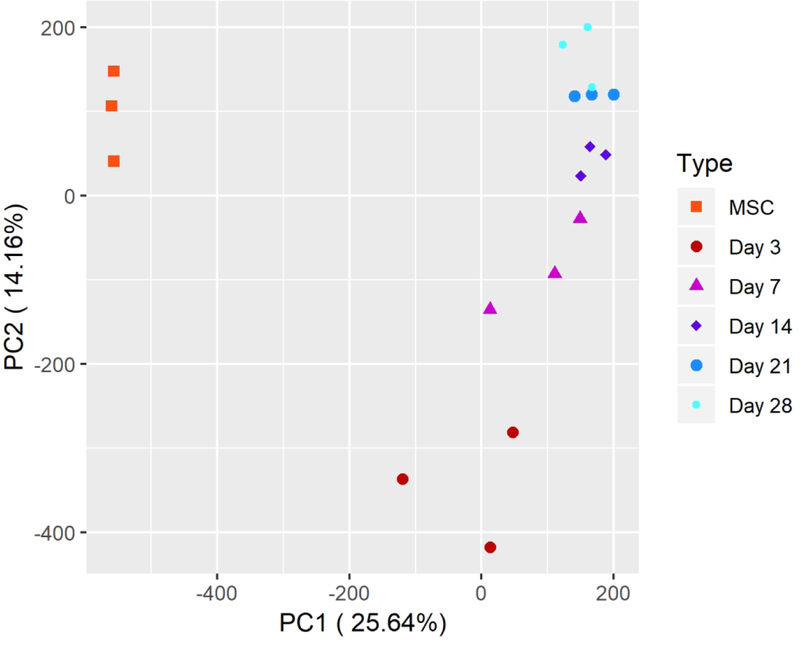
The transcriptional profiles of hMSCs on passage two is distinct from those undergoing pellet culture differentiation (Fig 2A) and the samples we profiled clustered neatly into their respective groupings under principal component analysis (Fig 2C). Differences in transcriptomes drop dramatically throughout the differentiation process as 9,969 DEGs were identified between hMSCs and day 3, while only 2,453, 49, 4, and 8 DEGs were identified between days 3–7, 7–14, 14–21, and 21–28, respectively (Fig 2B). Of the eight genes that are differentially expressed between day 21 (pellet maturity) and day 28 (hypertrophy), four are upregulated in day 28 pellets. The highest upregulation occurs in DYSF, a skeletal muscle involved in muscle contraction, and a non-coding transcript AL137782.1 with no known function. Expression of these transcripts most closely tracks with the onset of hypertrophy in pellet culture chondrogenesis.
Figure 2. Dynamic changes in gene expression are observed during hMSC differentiation in AC in vitro.
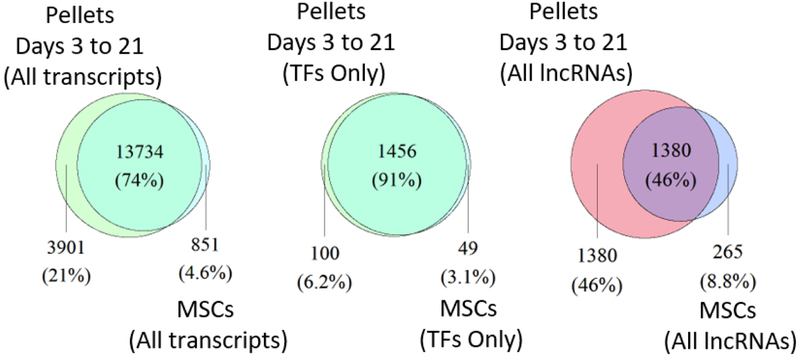
A. Venn diagram of all transcripts, TFs, and lncRNAs for hMSCs versus all transcripts, TFs, and lncRNAs expressed at any point between pellet culture day 3 and day 21.
B. Count of DEGs between consecutive days throughout pellet culture (DEGs defined here as any replicate TPM ≥ 1 and adjusted p-value ≤ 0.05).
C. PCA comparing hMSCs and chondrogenic pellets at various time points throughout differentiation by transcriptional profile of transcripts with at least one sample expressing at ≥ TPM of 1.
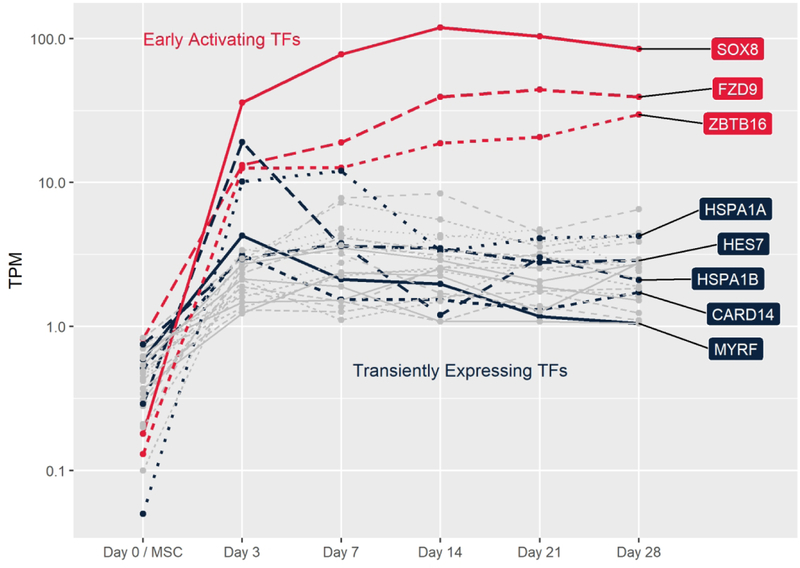
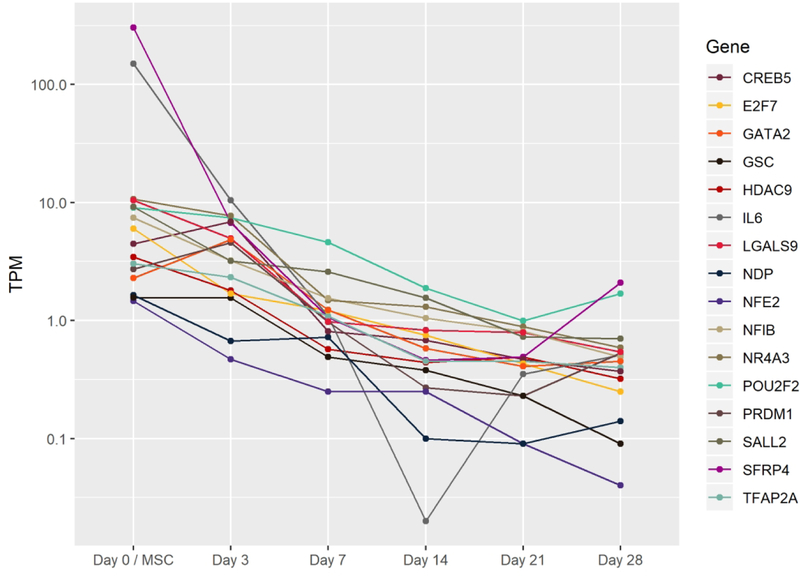
D. Highlight of rapidly and highly expressed TFs (red) and transiently expressed TFs (blue). Also shows all other TFs consistently (mean TPM≥1) expressed throughout chondrogenesis not highlighted in (E) or (F) (gray).
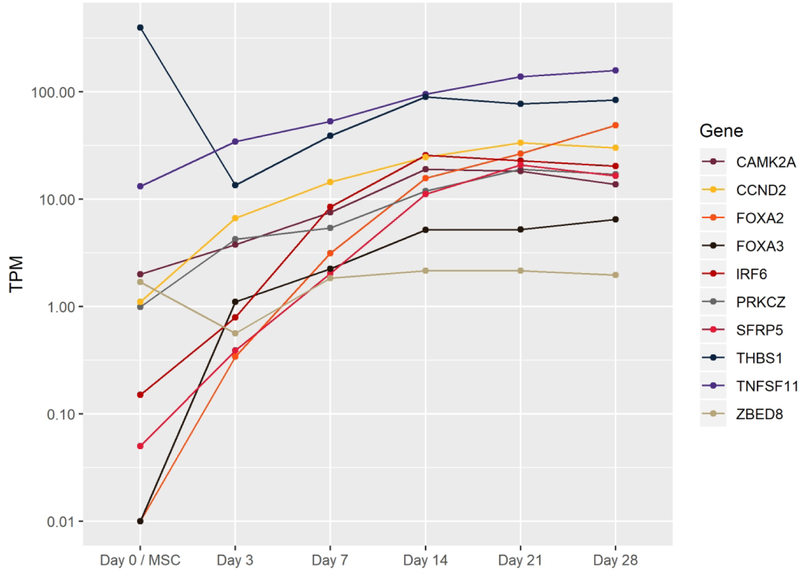
E. TFs increasing by four-fold between day 3 and day 21 of pellet culture.
F. TFs decreasing by four-fold between day 3 and day 21 of pellet culture..
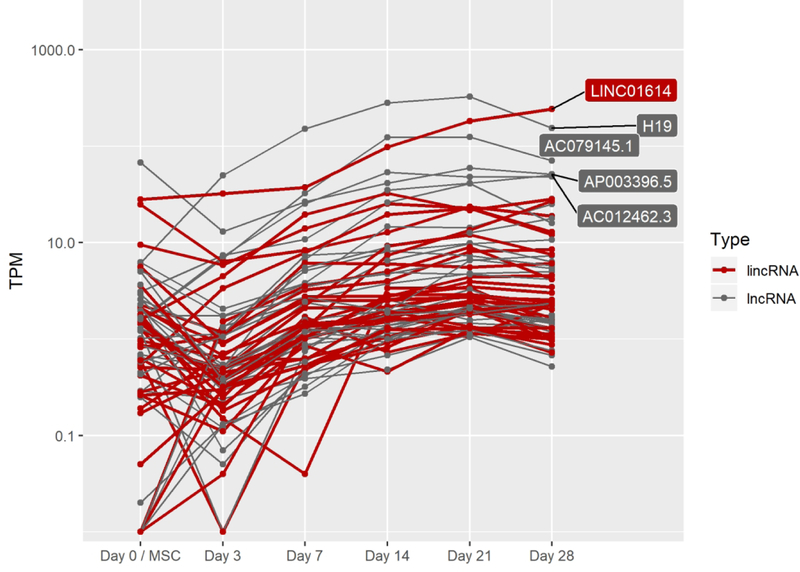
G. lncRNAs increasing by four-fold between day 3 and day 21 of pellet culture.
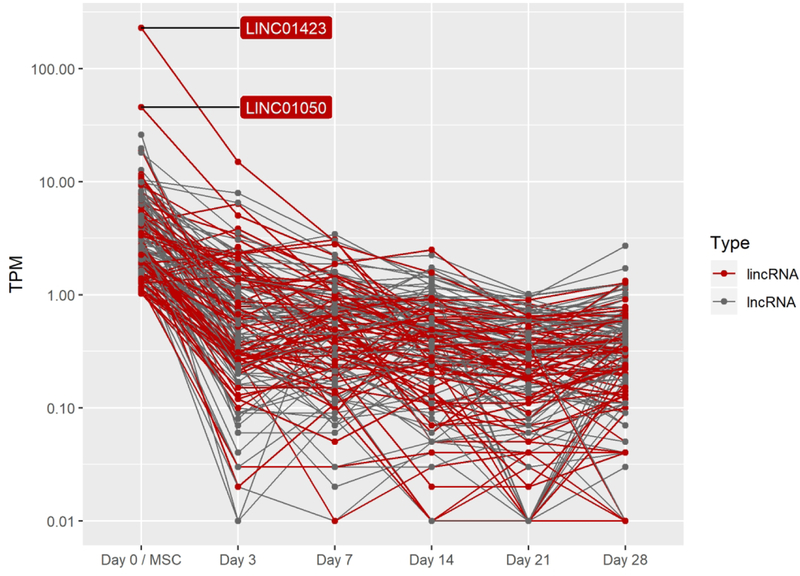
H. lncRNAs decreasing by four-fold between day 3 and day 21 of pellet culture.
With 91% of TFs shared between chondrogenic pellets and hMSCs (Fig 2A), relatively few TFs may play a controlling role in chondrogenic differentiation in vitro. We identified some such critical TFs by defining those increasing at least four fold between days 3 and 21 as well as those decreasing at least four fold from a minimum TPM of 1 at the hMSC stage. Fewer TFs met our criteria for increasing as compared to decreasing throughout the pellet culture process (Figs 2E, 2F), while a few highly expressed potential pioneer factors were rapidly transcribed upon chondrogenic induction (Early Activating TFs, Fig 2D). Five TFs also demonstrated a strong transient expression pattern at some point in differentiation by dropping expression levels by 50% or more between consecutive time points, while 18 others were expressed consistently throughout differentiation (Fig 2D).
Contrary to the relatively small portion of TFs whose expression was turned on or off (9.3%) during chondrogenic differentiation, many lncRNAs were turned on or off during this process (56%) (Fig 2A). Expression of the chondrogenic lincRNA H19, for example, dramatically increases (67-fold) from hMSC stage (day 0) to day 21 of differentiation, while several other less-studied lncRNAs are also highly expressed (day 21 > 50 TPM) as in vitro chondrogenesis progresses (Fig 2G). 146 lncRNAs, of which 64 are classified by ENSEMBL as lincRNAs, were expressed in hMSCs but fell below our 1 TPM expression threshold while also dropping at least four fold by day 21 (Fig 2G).
hMSC-derived Cartilage is Markedly Distinct from Primary Articular Cartilages
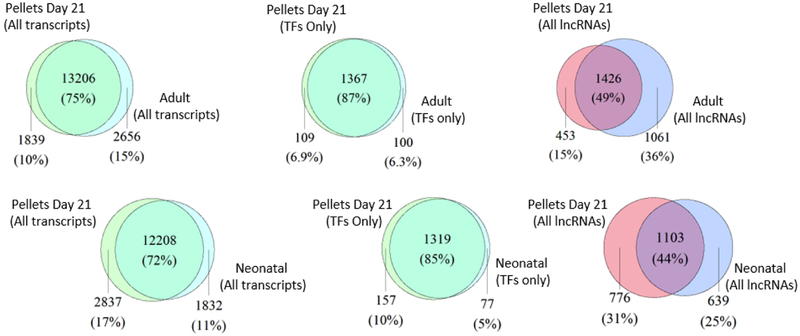
To comprehensively characterize the differences between chondrogenic pellets and both mature and immature human AC, we performed gene expression analyses between day 21 pellets and these tissues. As distinct as adult and neonatal AC are, they are both more similar in transcriptional profile to each other than they are to hMSC-derived cartilage (chondrogenic pellet at day 21). Primary tissues express 4,110 developmentally specific, uniquely expressed transcripts, 149 of which are TFs (Fig 1A). Comparison of hMSC-derived cartilage to adult tissue transcriptional profiles, however, demonstrate 4,495 (+9.6%) transcripts and 209 TFs (+28.7%) that are uniquely expressed in one tissue or the other. The same comparison between hMSC-derived cartilage and neonatal AC identifies 4,669 transcripts (+13.6%) and 234 TFs (+57.0%) (Figs 1A, 3A). The degree of variability in lncRNA transcriptional profiles when comparing each of these three tissues is not markedly different. It is worth noting that each category of hMSC-derived cartilage transcripts had more in common with adult than with neonatal AC and so it appears this cultured cartilage is more similar to adult tissue than to neonatal (Fig 3A). Additionally, PCA demonstrates greater similarity to hMSC-derived cartilage for adult samples than for neonatal samples (Fig 3B). Conversely, we find in differential expression analysis that many genes qualify as differentially expressed in neonatal AC versus day 21 pellets (3,840) and even more (5,377) qualify in adult versus day 21 pellets (Fig 3C), yet chondrogenic pellets become more similar to each native tissue’s transcriptional profile as the in vitro differentiation process progresses (Figs S2A, S2B). These metrics indicate that cultured cartilage is markedly distinct from primary tissues, though appears a closer approximation of adult tissue than of immature neonatal tissue.
Figure 3. Human native cartilage show substantial gene expression differences with hMSC-derived cartilage in vitro.
A. Venn diagram of day 21 pellets vs adult and neonatal AC (All, TFs, and lncRNAs).
B. PCA comparing pellets and primary ACs by transcriptional profile of transcripts with at least one sample expressing at ≥ TPM of 1.
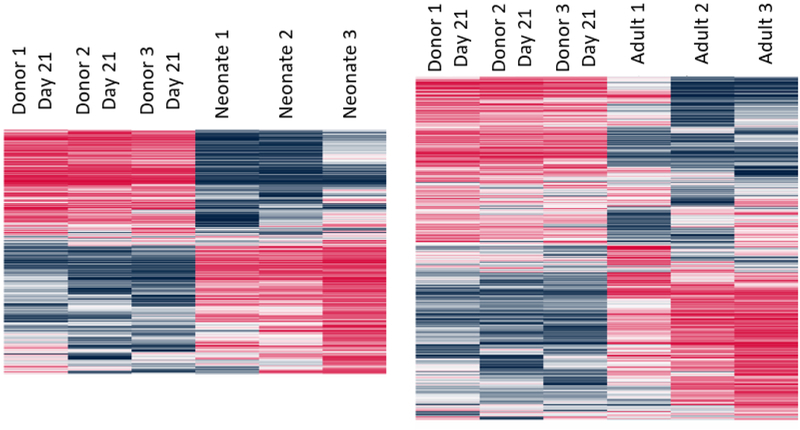
C. Heat map of DEGs for day 21 pellets vs neonatal and adult AC.
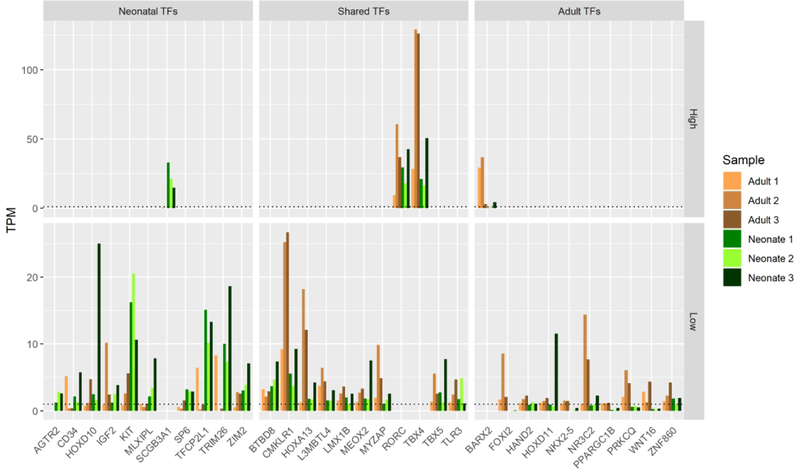
Successful reprogramming of hMSC-derived pellet cartilage into phenotypes resembling native AC may be achieved by modulation of critical chondrogenic TFs. To identify candidate chondrogenic TFs, we identified factors that were consistently expressed (TPM ≥ 1) in each replicate of primary tissue but never expressed throughout pellet culture (TPM ≤ 1 days 3–21). 21 TFs were found to meet these criteria in neonatal AC and 22 were found in adult AC with 11 factors shared between both sets (Fig 4A). Two TFs highly expressed in both tissues include RORC (range 18–61 TPM) and TBX4 (range 16–129 TPM). KIT, SCGB3A1, and TFCP2L1 each expressed consistently over 10 TPM in neonatal AC with relatively low expression for each in adult AC and little to no expression in hMSC-derived cartilage, indicating these as potentially key factors unique to the neonatal AC phenotype missing in TGF-β induced chondrogenesis.
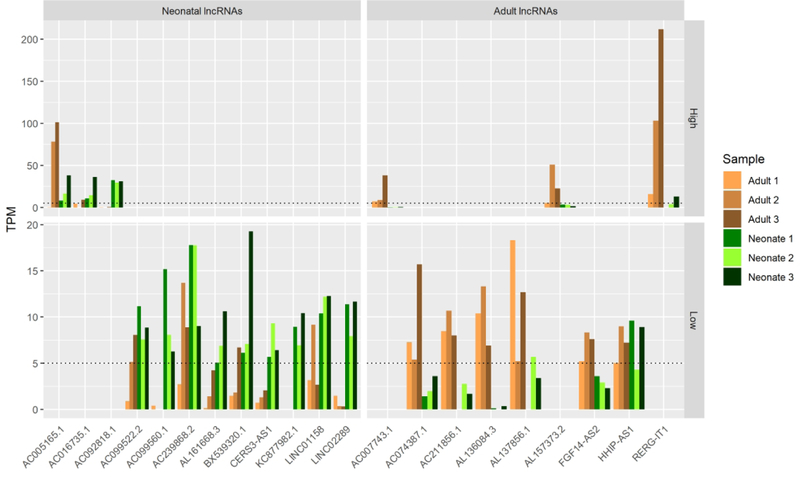
Figure 4. Key Transcription factors and long non-coding RNAs that are either unique or shared between neonatal vs. adult cartilage and absent in in vitro chondrogenesis.
A. TFs expressed (TPM ≥ 1) in primary tissues but never in hMSC chondrogenic differentiation (TPM < 1 days 3–21).
B. lncRNAs expressed (TPM ≥ 5) in primary tissues but never in hMSC chondrogenic differentiation (TPM < 1 days 3–21).
The characterization of lncRNA profiles in mature and immature AC may prove necessary for successful developmental recapitulation in vitro as many lncRNAs demonstrate dramatic differences in transcription between these primary tissues and hMSC-derived cartilage. To increase selectivity in identifying key lncRNAs absent during hMSC-derived cartilage differentiation, we increased our TPM threshold to 5 (while maintaining ≤1 TPM days 3–21 throughout pellet culture). These criteria highlighted 12 neonatal lncRNAs and 9 adult lncRNAs uniquely expressed throughout their respective groups (Fig 4B). Notably, several of these seem to be expressed highly across 5 of our 6 primary AC samples but absent or below threshold in another, indicating that they may be common to both immature and mature AC. Identification of these highly expressed lncRNAs within AC tissues is a critical first step towards understanding their roles in AC biology.
Mitigating Uncertainty in Human Tissue Data
Due to uncontrolled genetic, medical, and environmental differences, human data exhibits uncertainty that many model organisms do not. Yet, it is imperative that we study these data to most directly serve translational efforts. Rarely, healthy human AC is collected from the selfless donations of donor families. The human adult samples in this study were purchased and originated from both males and females of multiple racial backgrounds with an age range of 20–35 years. The causes of death for these donors were varied as well, but no cause gives any reason to suspect damage to cartilage tissue. The neonatal samples in this study, graciously donated by AlloSource, were all males approximately one month old with unknown racial backgrounds and causes of death. There are many unknowns in the backgrounds of these donors, notably any relevant medical conditions, weight, and physical activity level. The harvested AC was undamaged in each sample, however, and so we can at least be certain that osteoarthritis was not a factor. To minimize false positives hits for candidate genes, we chose to ensure that all positive hits were expressed over listed thresholds in each sample and not merely on average. We believe this strategy will mitigate some of the confounding effects of age, gender, race, medical history, and environmental effects.
DISCUSSION
With the growing need for better AC defect treatment options in mind, we aim to empower further studies exploring the potential of neonatal-like engineered cartilage in meeting such needs. With its capacity for expansion and self-organization, neonatal AC may be an ideal phenotypic target of engineered AC. Our results here highlight TFs and lncRNAs that we propose as candidate molecular drivers of this phenotype while distinguishing it from mature AC and hMSC-derived cartilage.
Relatively Few TFs Express Consistently and Uniquely in Neonatal AC
Six TFs and three lncRNAs met our criteria of expression (TPM≥1 for TFs, TPM≥5 for lncRNAs in all replicates) in neonatal AC and negative expression (TPM<1 in all replicates) in adult AC. Among these, the highest levels of expression were in SFRP5 and SCGB3A1. SFRP5 is a WNT signaling inhibitor of the secreted frizzled-related protein family and has been shown to be expressed at low levels in the deep zone of neonatal mouse cartilage by E15.5 (Lui et al., 2015; Witte, Dokas, Neuendorf, Mundlos, & Stricker, 2009) as well as in equine neonatal tissue AC (Duesterdieck-Zellmer, Semevolos, Kinsley, & Riddick, 2015). Given the importance of WNT signaling in limb bud and cartilage development and the dramatic differences in expression between adult and neonatal AC, SFRP5 stands out as a key candidate molecular driver of human neonatal AC phenotypes. Another interesting candidate is SCGB3A1, a growth inhibitory putative cytokine that is sensitive to oxygen levels in neonatal murine lung tissue (Krop et al., 2001; Raffay et al., 2013). Since AC is avascular and accordingly relatively hypoxic, oxygen-sensitive differentially expressed TFs are of particular interest to the tissue’s phenotype. The presence of CDK1 in this set reinforces the hypothesis that it is an essential regulator of skeletal development that acts as a switch from proliferative to hypertrophic states in chondrocytes (Saito et al., 2016). Little or nothing is known about roles for ADGRB2, AGTR2, or ZSCAN31 or for the neonatal-specific lncRNAs AC092818.1, AC099560.1, and KC877982.1 in cartilage biology to date. These nine factors appear to be distinctive within neonatal AC and their expression may be required to recapitulate the tissue in vitro.
Neonatal AC Lacks Consistent Expression of Many Factors Found in Mature AC
Mature AC expresses twenty TFs and five lncRNAs which neonatal AC lacks. Expression of two of the TFs, EGR2 and FOSL1, may be artifacts of the tissue harvesting procedure as they are indicators of stress response and are known to express in cartilage tissue allografts stored at 4C° for extended periods (Lin et al., 2016). Others are more likely to be functional drivers of cartilage biology for various reasons including disease association (Table 2), ontological database association (CAPN15, GO database v1.2, ECM degradation and collagen remodeling), or direct evidence of a known role in cartilage biology (Table 3).Of the five lncRNAs that met our stringent expression profile criteria, two overlap genes important to cartilage biology (AL136084.3 and COL15A1, ID2-AS1 and ID2). Both these lncRNAs may upregulate their respective neighboring protein coding genes as indicated by their co-expression (Figs 5A, 5B, 5C). While COL15A1 is expressed in each of our cartilage tissues, it is expressed dramatically more in adult than in neonatal AC (log2-fold-change 4.2, p-value = 7.6 × 10−12, Supp. File 1). ID2 is similarly differentially expressed in adults when compared to neonatal AC (log2-fold-change 2.2, p-value = 7.2 × 10−7, Supp. File 1) and is known to play roles in cartilage development (Lorda-Diez, Torre-Perez, Garcia-Porrero, Hurle, & Montero, 2009; Sakata-Goto et al., 2012). These data suggest that these uniquely expressed lncRNAs are playing important roles in AC biology and may serve as useful biomarkers when surveying the maturation of engineered cartilage. Together with the aforementioned TFs found only in adult AC, these factors may prove critical to both driving and monitoring the maturation of an immature, neonatal-like engineered AC into an adult-like post-expansion, tissue maintenance state.
Table 2:
Phenotypes of Diseases Associated with Highlighted Transcription Factors in Human Articular Cartilage
| Gene | Disease | Relevant Phenotype |
|---|---|---|
| DLX6 | Isolated split hand-split foot malformation | Skeletal defects |
| FOXI2 | Noonan syndrome | Short stature, skeletal defects |
| GLI1 | Ellis-van Creveld syndrome | Short stature, skeletal defects |
| HOXD11 | Congenital vertical talus | Primary dislocation of the talonavicular joint, foot deformity |
| NOD2 | Blau syndrome | Arthritis, synovitis |
| PPARGC1B | Kashin-Beck Disease | Dwarfism, severe articular deformities |
| WNT16 | Osteoarthritis | Polymorphisms associated with hip OA phenotypes (hypertrophic vs atrophic) (García-Ibarbia et al., 2017) |
Table 3:
Expression levels and Known Roles of Genes in Human Articular/Engineered Cartilage
| Gene | TPM in hMSC- derived cartilage |
TPM in Adult AC |
TPM in Neonatal AC |
Known role in cartilage biology |
|---|---|---|---|---|
| Hoxa2 | 1.58 | 2.38 | 0.39 | Inhibits bone morphogenetic protein signaling during osteogenic differentiation of the palatal mesenchyme (Iyyanar & Nazarali, 2017) |
| HOXC11 | 1.75 | 3.51 | 0.36 | Required for synovial joint formation (Koyama et al., 2010; Wellik & Capecchi, 2003) |
| HOXD11 | 0.05 | 1.51 | 4.37 | Required for limb bud development, SHH induction (Johnson & Tabin, 1997) |
| KLF13 | 6.89 | 11.90 | 36.65 | Cross-regulates BMP2 (Pabona, Zeng, Simmen, & Simmen, 2010) |
| LIF | 6.00 | 37.42 | 0.14 | Stimulates proteoglycan release and inhibit proteoglycan synthesis in cultured pig articular cartilage explants (Hui, Bell, Carroll, & Layton, 1998) |
| Nkx2.5 | 0.12 | 1.34 | 0.18 | Required for proper craniofacial development in zebrafish (Iklé et al., 2017) |
| Pax8 | 20.81 | 8.81 | 0.14 | Impaired linear growth, delayed endochondral ossification, reduced cortical bone, reduced bone mineralization in knockout mice (Mansouri, Chowdhury, & Gruss, 1998) |
| PRKCQ | 0.18 | 4.09 | 0.60 | Mediates BMP-4-induced osteoblastic differentiation (Park et al., 2010) |
| TLR6 | 0.67 | 2.32 | 0.24 | Treatment of chondrocytes with TLR6 ligands resulted in collagen resorption and MMP1 upregulation (Zhang et al., 2008) |
| WNT16 | 0.39 | 2.82 | 0.25 | Antagonizes excessive canonical WNT activation and protects cartilage in osteoarthritis (Nalesso et al., 2017); Synovial overexpression of Wnt16 led to canonical Wnt signaling in cartilage which increased protease activity and induced cartilage damage (van den Bosch et al., 2015) |
hMSC Chondrogenic Differentiation Transcriptional Profiles Suggest High Degree of lncRNA Activity in Chondrogenesis
The functional roles of lncRNAs in human biology are rapidly emerging, and the dramatic shifts in lncRNA expression demonstrated throughout in vitro chondrogenesis suggest numerous important roles for this class of molecules in cartilage biology. The most highly upregulated lncRNAs in TGF-β1/MSC chondrogenesis include LINC01614, AP003396.5, AC079145.1, and AC012462.3. While LINC01614 has been shown to activate FOXP1, which may be involved in osteoclast biology (A.-N. Liu, Qu, Yu, & Sun, 2018), AP003396.5 and AC079145.1 co-occupy loci with genes with interesting backgrounds. MATN3 has known roles in both development and homeostasis of cartilage and co-expresses with AC079145.1 (Fig S1A) while THY1 encodes a cell adhesion glycoprotein with no known role in cartilage biology, yet is expressed in all primary AC replicates and co-expresses with AP003396.5 during hMSC chondrogenic differentiation (Fig S1B). Most other upregulated lncRNAs in chondrogenic differentiation remain unstudied and demand attention in order to further our understanding of this process.
Several lncRNAs express highly in primary AC but are missing in hMSC chondrogenic differentiation. Of particular note are the highest expressing lncRNAs including AC092818.1 in neonatal AC and RERG-IT1 in adult AC. RERG-IT1 is a sense intronic transcript that co-expresses with RERG in adult and neonatal AC but not in differentiating hMSC cartilage pellets (Fig S1C), which may explain the higher expression levels of the cell proliferation and angiogenesis inhibiting RERG in primary AC tissues relative to chondrogenic pellets (Finlin et al., 2001; Zhao et al., 2017). AC092818.1, on the other hand, is an uncharacterized lincRNA located on chromosome 8 whose expression profile suggests a functional role unique to neonatal AC biology. Expression of these and other lncRNAs may or may not be required for engineering native-like AC, but may prove useful as easily identified biomarkers of successful recapitulation of primary AC phenotypes in vitro.
Improving the Utility of hMSC-derived Cartilage
Despite very high expression and production of both the master chondrogenic regulator SOX9 and classical markers of hyaline cartilage such as COL2A1 and ACAN (Fig S3), hMSC-derived cartilage expresses a molecular profile reliably distinct from both neonatal and adult AC in humans. Given the degree to which the cartilage derived from hMSCs differs in transcriptional profile from primary tissues, caution must be exercised when using it as a model. The clinical utility of engineered cartilages using current differentiation protocols is limited, but may be improved by inducing a more native-like transcriptional profile. The properties of neonatal AC make it a compelling target phenotype for such tissue engineering efforts and the factors highlighted here may prove useful in reprogramming hMSC chondrogenic differentiation to that end.
Given that the developmental state of neonatal AC is one of rapid expansion while adult AC is in a more stable state of tissue maintenance, it is reasonable to suggest that the factors unique to neonatal AC are potentially more transformative than those found in adult AC. With this in mind, the TFs AGTR2 and SCGB3A1 are the only two TFs that are both unique to neonatal AC compared to adult AC and also never expressed in hMSCs undergoing chondrogenic differentiation. These two factors may be among the most critically lacking in current tissue engineering efforts to create an engineered cartilage capable of expanding to cover a defect and of maturing into a stratified cartilage with native-like biomechanical properties. Other factors that may be of particular importance to creating a more articular engineered cartilage include those present in primary AC tissues but absent in hMSC-derived cartilage, listed in Figure 4A. Especially noteworthy are the most highly expressed TFs among the neonatal replicates, TFCP2L1, RORC, and TBX4. TFCP2L1 has been shown to be critical in maintaining stemness in human and mouse embryonic stem cells through Wnt/β-catenin signaling (K. Liu, Zhang, Liu, Ying, & Ye, 2017; Qiu et al., 2015; Sun et al., 2018) and may be acting to preserve chondrogenic progenitors in neonatal AC. TBX4 is a known factor in hindlimb development (Sheeba & Logan, 2017) and mutations in it cause Ischiocoxopodopatellar Syndrome, a rare genetic condition in which the patella and other hindlimb skeletal features fail to develop properly (Bongers et al., 2001). RORC, which has not availed itself as a key player in cartilage biology to date, is among the most consistently and highly expressed TFs in our dataset and warrants further study. These TFs are among the most promising candidates for reprogramming hMSC-derived cartilage into neonatal-like engineered AC. The successful engineering of such a cartilage may help to alleviate the unmet clinical need for a robust graft capable of resurfacing joint surfaces ravaged by osteoarthritis or of successfully integrating into large AC defects.
Supplementary Material
Supplementary Figure 1. Co-expression of lncRNAs upregulated during in vitro chondrogenesis with their genomic co-occupants.
A. Co-expression of MATN3 with AC079145.1.
B. Co-expression of THY1 with AP003396.5.
C. Co-expression of RERG with RERG-IT1.
Supplementary Figure 2
A. DEGs of all types with expression ≥ 1 TPM in at least one sample and adjusted p-value ≤ 0.05 throughout in vitro chondrogenesis compared to adult AC.
B. DEGs of all types with expression ≥ 1 TPM in at least one sample and adjusted p-value ≤ 0.05 throughout in vitro chondrogenesis compared to neonatal AC.
Supplementary Figure 3. Expression of common genes of interest in cartilage development, biology, and tissue engineering (average TPM, n=3 per group).
Acknowledgements:
The research was supported by grants from NIH/NIBIB (P41 EB021911), and the L. David and E. Virginia Baldwin Fund. We wish to sincerely thank the families of the tissue donors that made this study possible, and also AlloSource for their support via tissue donation.
Footnotes
Conflict of interest:
The authors declare that there is no conflict of interest to be declared.
References
- Ayupe AC, Tahira AC, Camargo L, Beckedorff FC, Verjovski-Almeida S, & Reis EM (2015). Global analysis of biogenesis, stability and sub-cellular localization of lncRNAs mapping to intragenic regions of the human genome. RNA Biology, 12(8), 877–892. 10.1080/15476286.2015.1062960 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bongers EM, Van Bokhoven H, Van Thienen MN, Kooyman MA, Van Beersum SE, Boetes C, … Hamel BC (2001). The small patella syndrome: description of five cases from three families and examination of possible allelism with familial patella aplasia-hypoplasia and nail-patella syndrome. Journal of Medical Genetics, 38(3), 209–214. Retrieved from http://www.ncbi.nlm.nih.gov/pubmed/11303519 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brama PAJ, TeKoppele JM, Bank RA, Barneveld A, & van Weeren PR (2002). Development of biochemical heterogeneity of articular cartilage: influences of age and exercise. Equine Veterinary Journal, 34(3), 265–269. Retrieved from http://www.ncbi.nlm.nih.gov/pubmed/12108744 [DOI] [PubMed] [Google Scholar]
- Brommer H, Brama PAJ, Laasanen MS, Helminen HJ, van Weeren PR, & Jurvelin JS (2005). Functional adaptation of articular cartilage from birth to maturity under the influence of loading: a biomechanical analysis. Equine Veterinary Journal, 37(2), 148–154. Retrieved from http://www.ncbi.nlm.nih.gov/pubmed/15779628 [DOI] [PubMed] [Google Scholar]
- Cleary MA, van Osch GJVM, Brama PA, Hellingman CA, & Narcisi R (2015). FGF, TGF β and Wnt crosstalk: embryonic to in vitro cartilage development from mesenchymal stem cells. Journal of Tissue Engineering and Regenerative Medicine, 9(4), 332–342. 10.1002/term.1744 [DOI] [PubMed] [Google Scholar]
- Duesterdieck-Zellmer K, Semevolos S, Kinsley M, & Riddick T (2015). Age-related differential gene and protein expression in postnatal cartilage canal and osteochondral junction chondrocytes. Gene Expression Patterns, 17(1), 1–10. 10.1016/J.GEP.2014.11.002 [DOI] [PubMed] [Google Scholar]
- Finlin BS, Gau C-L, Murphy GA, Shao H, Kimel T, Seitz RS, … Perou CM (2001). RERG Is a Novel ras-related, Estrogen-regulated and Growth-inhibitory Gene in Breast Cancer. Journal of Biological Chemistry, 276(45), 42259–42267. 10.1074/jbc.M105888200 [DOI] [PubMed] [Google Scholar]
- Forrest ME, & Khalil AM (2017). Review: Regulation of the cancer epigenome by long non-coding RNAs. Cancer Letters, 407, 106–112. 10.1016/J.CANLET.2017.03.040 [DOI] [PubMed] [Google Scholar]
- Gannon AR, Nagel T, Bell AP, Avery NC, & Kelly DJ (2015). Postnatal changes to the mechanical properties of articular cartilage are driven by the evolution of its collagen network. European Cells & Materials, 29, 105–121; discussion 121–3. Retrieved from http://www.ncbi.nlm.nih.gov/pubmed/25633309 [DOI] [PubMed] [Google Scholar]
- García-Ibarbia C, Neila S, Garcés C, Alonso MA, Zarrabeitia MT, Valero C, … Riancho JA (2017). Non-synonymous WNT16 polymorphisms alleles are associated with different osteoarthritis phenotypes. Rheumatology International, 37(10), 1667–1672. 10.1007/s00296-017-3783-5 [DOI] [PubMed] [Google Scholar]
- Glinsky G, Durruthy-Durruthy J, Wossidlo M, Grow EJ, Weirather JL, Au KF, … Sebastiano V (2018). Single cell expression analysis of primate-specific retroviruses-derived HPAT lincRNAs in viable human blastocysts identifies embryonic cells co-expressing genetic markers of multiple lineages. Heliyon, 4(6), e00667. 10.1016/j.heliyon.2018.e00667 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hui W, Bell MC, Carroll GJ, & Layton MJ (1998). MODULATION OF CARTILAGE PROTEOGLYCAN METABOLISM BY LIF BINDING PROTEIN. Cytokine, 10(3), 220–226. 10.1006/CYTO.1997.0279 [DOI] [PubMed] [Google Scholar]
- Hunziker EB, Kapfinger E, & Geiss J (2007). The structural architecture of adult mammalian articular cartilage evolves by a synchronized process of tissue resorption and neoformation during postnatal development. Osteoarthritis and Cartilage, 15(4), 403–413. 10.1016/j.joca.2006.09.010 [DOI] [PubMed] [Google Scholar]
- Iklé JM, Tavares ALP, King M, Ding H, Colombo S, Firulli BA, … Clouthier DE (2017). Nkx2.5 regulates endothelin converting enzyme-1 during pharyngeal arch patterning. Genesis (New York, N.Y. : 2000), 55(3). 10.1002/dvg.23021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iyyanar PPR, & Nazarali AJ (2017). Hoxa2 Inhibits Bone Morphogenetic Protein Signaling during Osteogenic Differentiation of the Palatal Mesenchyme. Frontiers in Physiology, 8, 929 10.3389/fphys.2017.00929 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jadin KD, Bae WC, Schumacher BL, & Sah RL (2007). Three-dimensional (3-D) imaging of chondrocytes in articular cartilage: Growth-associated changes in cell organization. Biomaterials, 28(2), 230–239. 10.1016/j.biomaterials.2006.08.053 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jarroux J, Morillon A, & Pinskaya M (2017). History, Discovery, and Classification of lncRNAs. In Advances in experimental medicine and biology (Vol. 1008, pp. 1–46). 10.1007/978-981-10-5203-3_1 [DOI] [PubMed] [Google Scholar]
- Johnson RL, & Tabin CJ (1997). Molecular models for vertebrate limb development. Cell, 90(6), 979–990. Retrieved from http://www.ncbi.nlm.nih.gov/pubmed/9323126 [DOI] [PubMed] [Google Scholar]
- Johnstone B, Hering TM, Caplan AI, Goldberg VM, & Yoo JU (1998). In VitroChondrogenesis of Bone Marrow-Derived Mesenchymal Progenitor Cells. Experimental Cell Research, 238(1), 265–272. 10.1006/excr.1997.3858 [DOI] [PubMed] [Google Scholar]
- Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D., … Rinn JL (2009). Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proceedings of the National Academy of Sciences, 106(28), 11667–11672. 10.1073/pnas.0904715106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koyama E, Yasuda T, Minugh-Purvis N, Kinumatsu T, Yallowitz AR, Wellik DM, & Pacifici M (2010). Hox11 genes establish synovial joint organization and phylogenetic characteristics in developing mouse zeugopod skeletal elements. Development, 137(22), 3795–3800. 10.1242/dev.053447 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krop IE, Sgroi D, Porter DA, Lunetta KL, LeVangie R, Seth P, … Polyak K (2001). HIN-1, a putative cytokine highly expressed in normal but not cancerous mammary epithelial cells. Proceedings of the National Academy of Sciences of the United States of America, 98(17), 9796–9801. 10.1073/pnas.171138398 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lennon DP, & Caplan AI (2006). Isolation of human marrow-derived mesenchymal stem cells. Experimental Hematology, 34(11), 1604–1605. 10.1016/j.exphem.2006.07.014 [DOI] [PubMed] [Google Scholar]
- Lennon DP, Schluchter MD, & Caplan AI (2012). The Effect of Extended First Passage Culture on the Proliferation and Differentiation of Human Marrow-Derived Mesenchymal Stem Cells. STEM CELLS Translational Medicine, 1(4), 279–288. 10.5966/sctm.2011-0011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin Y, Lewallen EA, Camilleri ET, Bonin CA, Jones DL, Dudakovic A, … Krych AJ (2016). RNA-seq analysis of clinical-grade osteochondral allografts reveals activation of early response genes. Journal of Orthopaedic Research : Official Publication of the Orthopaedic Research Society, 34(11), 1950–1959. 10.1002/jor.23209 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu A-N, Qu H-J, Yu C-Y, & Sun P (2018). Knockdown of LINC01614 inhibits lung adenocarcinoma cell progression by up-regulating miR-217 and down-regulating FOXP1. Journal of Cellular and Molecular Medicine, 22(9), 4034–4044. 10.1111/jcmm.13483 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu K, Zhang Y, Liu D, Ying Q-L, & Ye S (2017). TFCP2L1 represses multiple lineage commitment of mouse embryonic stem cells through MTA1 and LEF1. Journal of Cell Science, 130(22), 3809–3817. 10.1242/jcs.206532 [DOI] [PubMed] [Google Scholar]
- Lorda-Diez CI, Torre-Perez N, Garcia-Porrero JA, Hurle JM, & Montero JA (2009). Expression of Id2 in the developing limb is associated with zones of active BMP signaling and marks the regions of growth and differentiation of the developing digits. The International Journal of Developmental Biology, 53(8–9–10), 1495–1502. 10.1387/ijdb.072415cl [DOI] [PubMed] [Google Scholar]
- Lui JC, Chau M, Chen W, Cheung CSF, Hanson J, Rodriguez-Canales J, … Baron J (2015). Spatial regulation of gene expression during growth of articular cartilage in juvenile mice. Pediatric Research, 77(3), 406–415. 10.1038/pr.2014.208 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makris EA, Gomoll AH, Malizos KN, Hu JC, & Athanasiou KA (2015). Repair and tissue engineering techniques for articular cartilage. Nature Reviews. Rheumatology, 11(1), 21–34. 10.1038/nrrheum.2014.157 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mansouri A, Chowdhury K, & Gruss P (1998). Follicular cells of the thyroid gland require Pax8 gene function. Nature Genetics, 19(1), 87–90. 10.1038/ng0598-87 [DOI] [PubMed] [Google Scholar]
- Mi H, Muruganujan A, Ebert D, Huang X, & Thomas PD (2019). PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Research, 47(D1), D419–D426. 10.1093/nar/gky1038 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mi H, & Thomas P (2009). PANTHER Pathway: An Ontology-Based Pathway Database Coupled with Data Analysis Tools (pp. 123–140). 10.1007/978-1-60761-175-2_7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morris KV, & Mattick JS (2014). The rise of regulatory RNA. Nature Reviews Genetics, 15(6), 423–437. 10.1038/nrg3722 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nalesso G, Thomas BL, Sherwood JC, Yu J, Addimanda O, Eldridge SE, … Dell’Accio F (2017). WNT16 antagonises excessive canonical WNT activation and protects cartilage in osteoarthritis. Annals of the Rheumatic Diseases, 76(1), 218–226. 10.1136/annrheumdis-2015-208577 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Necsulea A, Soumillon M, Warnefors M, Liechti A, Daish T, Zeller U, … Kaessmann H (2014). The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature, 505(7485), 635–640. 10.1038/nature12943 [DOI] [PubMed] [Google Scholar]
- Pabona JMP, Zeng Z, Simmen FA, & Simmen RCM (2010). Functional Differentiation of Uterine Stromal Cells Involves Cross-Regulation between Bone Morphogenetic Protein 2 and Krüppel-Like Factor (KLF) Family Members KLF9 and KLF13. Endocrinology, 151(7), 3396–3406. 10.1210/en.2009-1370 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park KH, Han DI, Rhee Y-H, Jeong S-J, Kim S-H, & Park YG (2010). Protein kinase C βII and δ/θ play critical roles in bone morphogenic protein-4-stimulated osteoblastic differentiation of MC3T3-E1 cells. Biochemical and Biophysical Research Communications, 403(1), 7–12. 10.1016/j.bbrc.2010.10.074 [DOI] [PubMed] [Google Scholar]
- Pelttari K, Winter A, Steck E, Goetzke K, Hennig T, Ochs BG, … Richter W (2006). Premature induction of hypertrophy during in vitro chondrogenesis of human mesenchymal stem cells correlates with calcification and vascular invasion after ectopic transplantation in SCID mice. Arthritis & Rheumatism, 54(10), 3254–3266. 10.1002/art.22136 [DOI] [PubMed] [Google Scholar]
- Qiu D, Ye S, Ruiz B, Zhou X, Liu D, Zhang Q, & Ying Q-L (2015). Klf2 and Tfcp2l1, Two Wnt/β-Catenin Targets, Act Synergistically to Induce and Maintain Naive Pluripotency. Stem Cell Reports, 5(3), 314–322. 10.1016/j.stemcr.2015.07.014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raffay TM, Locy ML, Hill CL, Jindal NS, Rogers LK, Welty SE, & Tipple TE (2013). Neonatal hyperoxic exposure persistently alters lung secretoglobins and annexin A1. BioMed Research International, 2013, 408485. 10.1155/2013/408485 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saito M, Mulati M, Talib SZA, Kaldis P, Takeda S, Okawa A, & Inose H (2016). The Indispensable Role of Cyclin-Dependent Kinase 1 in Skeletal Development. Scientific Reports, 6, 20622. 10.1038/srep20622 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakata-Goto T, Takahashi K, Kiso H, Huang B, Tsukamoto H, Takemoto M, … Bessho K (2012). Id2 controls chondrogenesis acting downstream of BMP signaling during maxillary morphogenesis. Bone, 50(1), 69–78. 10.1016/j.bone.2011.09.049 [DOI] [PubMed] [Google Scholar]
- Sheeba CJ, & Logan MPO (2017). The Roles of T-Box Genes in Vertebrate Limb Development. In Current topics in developmental biology (Vol. 122, pp. 355–381). 10.1016/bs.ctdb.2016.08.009 [DOI] [PubMed] [Google Scholar]
- Somoza RA, Correa D, Labat I, Sternberg H, Forrest ME, Khalil AM, … Caplan AI (2018). Transcriptome-Wide Analyses of Human Neonatal Articular Cartilage and Human Mesenchymal Stem Cell-Derived Cartilage Provide a New Molecular Target for Evaluating Engineered Cartilage. Tissue Engineering. Part A, 24(3–4), 335–350. 10.1089/ten.TEA.2016.0559 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Somoza RA, Welter JF, Correa D, & Caplan AI (2014). Chondrogenic differentiation of mesenchymal stem cells: challenges and unfulfilled expectations. Tissue Engineering. Part B, Reviews, 20(6), 596–608. 10.1089/ten.TEB.2013.0771 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song EK, & Park TJ (2014). Integrin signaling in cartilage development. Animal Cells and Systems, 18(6), 365–371. 10.1080/19768354.2014.987319 [DOI] [Google Scholar]
- Sorrell JM, Somoza RA, & Caplan AI (2018). Human mesenchymal stem cells induced to differentiate as chondrocytes follow a biphasic pattern of extracellular matrix production. Journal of Orthopaedic Research®, 36(6), 1757–1766. 10.1002/jor.23820 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun H, You Y, Guo M, Wang X, Zhang Y, & Ye S (2018). Tfcp2l1 safeguards the maintenance of human embryonic stem cell self-renewal. Journal of Cellular Physiology, 233(9), 6944–6951. 10.1002/jcp.26483 [DOI] [PubMed] [Google Scholar]
- Ulitsky I, Shkumatava A, Jan CH, Sive H, & Bartel DP (2011). Conserved Function of lincRNAs in Vertebrate Embryonic Development despite Rapid Sequence Evolution. Cell, 147(7), 1537–1550. 10.1016/j.cell.2011.11.055 [DOI] [PMC free article] [PubMed] [Google Scholar]
- van den Bosch MH, Blom AB, Sloetjes AW, Koenders MI, van de Loo FA, van den Berg WB, … van der Kraan PM (2015). Induction of Canonical Wnt Signaling by Synovial Overexpression of Selected Wnts Leads to Protease Activity and Early Osteoarthritis-Like Cartilage Damage. The American Journal of Pathology, 185(7), 1970–1980. 10.1016/j.ajpath.2015.03.013 [DOI] [PubMed] [Google Scholar]
- Washietl S, Kellis M, & Garber M (2014). Evolutionary dynamics and tissue specificity of human long noncoding RNAs in six mammals. Genome Research, 24(4), 616–628. 10.1101/gr.165035.113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wellik DM, & Capecchi MR (2003). Hox10 and Hox11 Genes Are Required to Globally Pattern the Mammalian Skeleton. Science, 301(5631), 363–367. 10.1126/science.1085672 [DOI] [PubMed] [Google Scholar]
- Witte F, Dokas J, Neuendorf F, Mundlos S, & Stricker S (2009). Comprehensive expression analysis of all Wnt genes and their major secreted antagonists during mouse limb development and cartilage differentiation. Gene Expression Patterns, 9(4), 215–223. 10.1016/J.GEP.2008.12.009 [DOI] [PubMed] [Google Scholar]
- Woolhead GM, Donovan JL, & Dieppe PA (2005). Outcomes of total knee replacement: a qualitative study. Rheumatology, 44(8), 1032–1037. 10.1093/rheumatology/keh674 [DOI] [PubMed] [Google Scholar]
- Worster AA, Nixon AJ, Brower-Toland BD, & Williams J (2000). Effect of transforming growth factor beta1 on chondrogenic differentiation of cultured equine mesenchymal stem cells. American Journal of Veterinary Research, 61(9), 1003–1010. Retrieved from http://www.ncbi.nlm.nih.gov/pubmed/10976727 [DOI] [PubMed] [Google Scholar]
- Zhang Q, Hui W, Litherland GJ, Barter MJ, Davidson R, Darrah C, … Young DA (2008). Differential Toll-like receptor-dependent collagenase expression in chondrocytes. Annals of the Rheumatic Diseases, 67(11), 1633–1641. 10.1136/ard.2007.079574 [DOI] [PubMed] [Google Scholar]
- Zhao W, Ma N, Wang S, Mo Y, Zhang Z, Huang G, … Takeuchi K (2017). RERG suppresses cell proliferation, migration and angiogenesis through ERK/NF-κB signaling pathway in nasopharyngeal carcinoma. Journal of Experimental & Clinical Cancer Research, 36(1), 88 10.1186/s13046-017-0554-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Figure 1. Co-expression of lncRNAs upregulated during in vitro chondrogenesis with their genomic co-occupants.
A. Co-expression of MATN3 with AC079145.1.
B. Co-expression of THY1 with AP003396.5.
C. Co-expression of RERG with RERG-IT1.
Supplementary Figure 2
A. DEGs of all types with expression ≥ 1 TPM in at least one sample and adjusted p-value ≤ 0.05 throughout in vitro chondrogenesis compared to adult AC.
B. DEGs of all types with expression ≥ 1 TPM in at least one sample and adjusted p-value ≤ 0.05 throughout in vitro chondrogenesis compared to neonatal AC.
Supplementary Figure 3. Expression of common genes of interest in cartilage development, biology, and tissue engineering (average TPM, n=3 per group).