Abstract
Many primary tumours have low levels of molecular oxygen (hypoxia), and hypoxic tumours respond poorly to therapy. Pan-cancer molecular hallmarks of tumour hypoxia remain poorly understood, with limited comprehension of its associations with specific mutational processes, non-coding driver genes and evolutionary features. Here, as part of the ICGC/TCGA Pan-Cancer Analysis of Whole Genomes (PCAWG) Consortium, which aggregated whole genome sequencing data from 2658 cancers across 38 tumour types, we quantify hypoxia in 1188 tumours spanning 27 cancer types. Elevated hypoxia associates with increased mutational load across cancer types, irrespective of underlying mutational class. The proportion of mutations attributed to several mutational signatures of unknown aetiology directly associates with the level of hypoxia, suggesting underlying mutational processes for these signatures. At the gene level, driver mutations in TP53, MYC and PTEN are enriched in hypoxic tumours, and mutations in PTEN interact with hypoxia to direct tumour evolutionary trajectories. Overall, hypoxia plays a critical role in shaping the genomic and evolutionary landscapes of cancer.
Subject terms: Cancer genomics, Cancer microenvironment, Tumour heterogeneity
Many tumours exhibit hypoxia (low oxygen) and hypoxic tumours often respond poorly to therapy. Here, the authors quantify hypoxia in 1188 tumours from 27 cancer types, showing elevated hypoxia links to increased mutational load, directing evolutionary trajectories.
Introduction
Approximately half of all solid tumours are characterized by low levels of molecular oxygen (hypoxia)1–4. Sub-regions of hypoxia can result from disrupted oxygen supply: irregular and disorganized tumour vasculature can reduce oxygen availability5. Hypoxia can also be caused by changes in oxygen demand: altered tumour metabolism6,7 can increase intra-cellular demand for oxygen, potentially extending hypoxia signalling to liquid tumours. The adaptation of tumour cells to this imbalance in oxygen supply and demand is associated with poor clinical prognosis in several cancer types, attributed at least in part to hypoxia-associated genomic instability and clonal selection8–16.
Previous work has provided insight into the molecular origins and consequences of tumour hypoxia and genomic instability. Dynamic cycling of hypoxia can select for cells with TP53 mutations and for those that are apoptosis-deficient17,18. Indeed mutations in TP53 occur at a higher frequency in hypoxic primary tumours of at least 9 types16. The abundance of proteins involved in homologous recombination (e.g. RAD51) and non-homologous end joining (e.g. Ku70) are reduced under hypoxia, and these changes can persist for 2 days after reoxygenation19–21. Genes central to efficient mismatch repair (e.g. MLH1 and MSH2) are also downregulated under hypoxia22,23. Further, co-presence of tumour hypoxia and high genomic instability14,15, specific cellular morphologies like intraductal and cribriform carcinoma24 or specific mutations like loss of PTEN16, synergistically predict for rapid relapse after definitive local therapy in some tumour types, particularly prostate cancer. These data underscore the relationship between hypoxia and DNA repair defects, and suggest the tumour microenvironment applies a selective pressure leading to the development of specific genomic profiles.
We previously evaluated the exomic and copy-number alteration (CNA) consequences of tumour hypoxia across 19 cancer types16. However, the influence of tumour hypoxia on pan-cancer driver alterations, mutational signatures, and subclonal architectures remains unclear. To fill this gap, we calculated tumour hypoxia scores for 1188 tumours with whole-genome sequencing (WGS) and RNA sequencing, spanning 27 cancer types. Genome sequencing data was aggregated by the Pan-Cancer Analysis of Whole Genomes (PCAWG) consortium and generated by the ICGC and TCGA projects. These sequencing data were re-analyzed with standardized, high-accuracy pipelines to align to the human genome (reference build hs37d5) and identify germline variants and somatic mutations, as described previously25. This sequencing data together with our high-quality hypoxia quantitation represents a powerful hypothesis-generating mechanism to suggest useful back-translational in vitro experiments and better define the hypoxia-associated mutator phenotype across cancers. We associated hypoxia with key driver alterations in coding and non-coding regions of the genome, and find hypoxia is associated with specific mutational signatures of unknown aetiology. We illustrate the joint impact of PTEN and the tumour microenvironment in influencing the evolutionary trajectory of tumours. Overall, these data highlight the genomic changes through which hypoxia drives aggressive cancers.
Results
The pan-cancer landscape of tumour hypoxia
We compiled a cohort of 1188 tumours from 27 cancer types via the PCAWG Consortium. All samples had matched tumour and reference normal WGS and tumour RNA sequencing data generated by the ICGC and TCGA projects. WGS25 and RNA-sequencing26 analyses were systematically carried out by centralized teams with consistent and high-accuracy bioinformatics pipelines. Normal reference samples had a mean WGS coverage of 39 reads per base-pair while coverage for tumour samples had a bimodal distribution with modes at 38 and 60 reads per base-pair25. All samples underwent an extensive and systematic quality assurance process25.
We used linear mixed-effect models to associate hypoxia with features of interest across cancers while adjusting for tumour purity, age, and sex27,28. Cancer type was further incorporated as a random effect in every model, allowing us to consider a different baseline value for the feature of interest for each cancer type. As a measure of effect size we report conditional R2 values, denoted as , which reflect the variance explained by the fixed and random factors in each model29. We also report marginal R2 values, denoted as , which reflect the variance explained only by the fixed factors29.
We scored tumour hypoxia in all 1188 tumours using a trio of mRNA-based hypoxia signatures from Buffa30, Winter31 and Ragnum32 (Fig. 1a, Supplementary Fig. 1a, b, Supplementary Table 1, Supplementary Data 1). Hypoxia scores from each of these independent signatures were strongly correlated (ρ = 0.71–0.88, all p < 2.2 × 10−16, AS89; Supplementary Fig. 1c) and consistently predicted squamous tumours of the lung (Lung-SCC), cervix (Cervix-SCC), and head (Head-SCC) as the most hypoxic (Supplementary Fig. 1d, e). Comparatively, chronic lymphocytic leukaemias (Lymph-CLL) and thyroid adenocarcinomas (Thy-AdenoCA) were the least hypoxic, consistent with previous16 reports (ρ = 0.94, p < 2.2 × 10−16, AS89; Fig. 1b, Supplementary Fig. 1f, h). Remarkably, subsets of patients from 23/27 cancer types have tumours with elevated hypoxia (hypoxia score > 0) and tumours consistently have elevated hypoxia compared to normal tissues (Supplementary Fig. 2a–c).
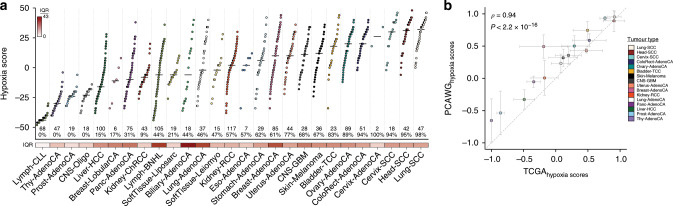
Fig. 1. The pan-cancer landscape of tumour hypoxia.
We quantified tumour hypoxia in 1188 independent tumours spanning 27 different cancer types. a Hypoxia scores for 27 types of cancer, based on the Buffa mRNA abundance signature. Cancer types are sorted by the median hypoxia score (horizontal black line) for each cancer type. Each dot represents one tumour. Sample sizes for each cancer type are listed near the bottom along with the percent of tumours that have elevated hypoxia (hypoxia score > 0). The variability in hypoxia within cancer types was measured by the interquartile range (IQR), shown along the bottom. The IQR was particularly high in biliary adenocarcinoma (IQR = 43.0; Biliary-AdenoCA), lymphoid B-cell non-Hodgkin’s lymphomas (IQR = 36.0; Lymph-BNHL), lung adenocarcinoma (IQR = 34.0; Lung-AdenoCA) and breast adenocarcinoma (IQR = 32; Breast-AdenoCA). By contrast, chronic lymphocytic leukaemia (IQR = 2.0; Lymph-CLL) and thyroid adenocarcinoma (IQR = 11.0; Thy-AdenoCA) showed less variance in hypoxia score. b Analysis of hypoxia between 16 comparable cancer types in PCAWG and TCGA (Spearman’s ρ, AS89). Dots represent the mean of the scaled median hypoxia scores from three different mRNA-based hypoxia signatures. Error bars represent the standard deviation of the scaled median hypoxia scores. Overall, the pan-cancer quantification of hypoxia between the PCAWG and TCGA datasets shows strong agreement.
Considering the strong agreement between the Winter, Buffa and Ragnum hypoxia signatures (Fig. 1a, Supplementary Fig. 1c, d), we used the Buffa signature for subsequent analyses. The Buffa signature has been previously used for pan-cancer analyses and shows results consistent with those from other signatures16. We first assessed the degree of inter-tumoural heterogeneity in hypoxia that lies within individual cancer types rather than between them. Over 42% of the variance in hypoxia scores occurs within individual cancer types, highlighting the microenvironmental diversity between tumours arising in a single tissue. This variability in hypoxia score within cancer types was especially elevated in some tumour types, particularly biliary adenocarcinomas (interquartile range, IQR = 43.0; Biliary-AdenoCA), mature B-cell lymphomas (IQR = 36.0; Lymph-BNHL), lung adenocarcinomas (IQR = 34.0; Lung-AdenoCA) and breast adenocarcinomas (IQR = 32.0; Breast-AdenoCA). This was in contrast to chronic lymphocytic leukaemias (IQR = 2.0; Lymph-CLL) and prostate adenocarcinomas (IQR = 6.0; Prost-AdenoCA) where little inter-tumoural variability in hypoxia was observed. The variability in hypoxia score was not significantly associated with the median hypoxia score within cancer types (ρ = 0.20, p = 0.30, AS89; Supplementary Fig. 2d) or with sample size (ρ = 0.22, p = 0.27, AS89; Supplementary Fig. 2e). Overall, extensive heterogeneity exists in hypoxia levels within and across cancer types.
The genomic correlates of tumour hypoxia
To determine whether genomic instability arising from specific mutational classes is associated with hypoxia, we looked to identify hypoxia-associated pan-cancer mutational density and summary features33. As a positive control, we first considered the percentage of the genome with a copy-number aberration (PGA), an engineered feature that is a surrogate for genomic instability and is associated with hypoxia across several tumour types16 (Supplementary Fig. 2f). Indeed, in this diverse pan-cancer cohort, hypoxic tumours have elevated genomic instability while controlling for cancer type, tumour purity, age and sex27 (p = 2.41 × 10−8, = 0.022, = 0.57, linear mixed-effect model; Fig. 2a).
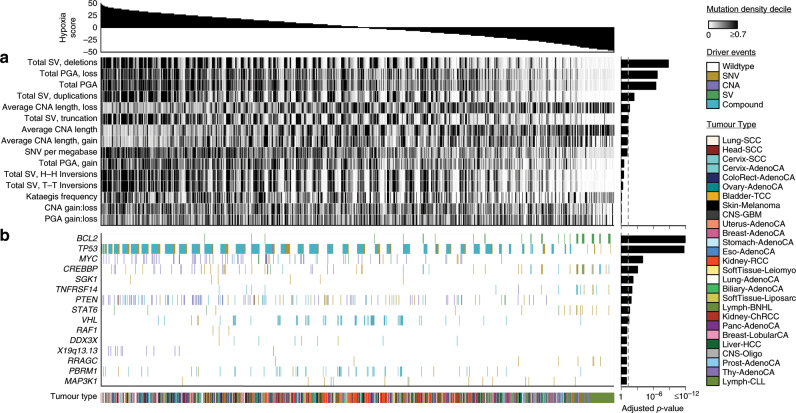
Fig. 2. The genomic correlates of tumour hypoxia.
We associated tumour hypoxia with mutational density and summary features, a, and driver mutations, b, across 27 cancer types using linear mixed-effect models. Hypoxia scores for all 1188 tumours are shown along the top. a Elevated tumour hypoxia was strongly associated with more deletions, elevated PGA, smaller CNAs, and a higher number of SNVs per megabase (n = 1188 independent tumours). Bonferroni-adjusted p-values are shown on the right. b We tested if driver mutations (e.g. any of SNV, CNA, SV or a compound event with more than one type of mutation) were associated with hypoxia in 1096 independent tumours with driver mutation data. Tumours with mutations in BCL2 showed lower levels of hypoxia while patients with mutations in TP53 showed remarkably elevated tumour hypoxia. Other driver mutations associated with elevated hypoxia include the oncogene MYC and the tumour suppressor PTEN. FDR-adjusted p-values are shown along the right. SV structural variant; PGA percentage of the genome with a copy-number aberration; CNA copy-number aberration; SNV single nucleotide variant; H-H head-to-head; T-T tail-to-tail. All associations were modelled using linear mixed-effect models while adjusting for cancer type, tumour purity, age and sex.
We then considered the association of hypoxia scores with 14 other metrics of the mutation density of CNAs, structural variants (SVs) and single nucleotide variants (SNVs) using linear mixed-effect models (Fig. 2a, Supplementary Fig. 2f, Supplementary Tables 2, 3). The strongest single correlate of tumour hypoxia was the total number of deletions, where patients with elevated hypoxia had more deletions (p = 1.11 × 10−10, = 0.023, = 0.59, linear mixed-effect model). Elevated numbers of other SVs such as duplications (p = 2.94 × 10−4, = 0.0084, = 0.60, linear mixed-effect model) and truncations (p = 3.29 × 10−3, = 0.0062, = 0.60, linear mixed-effect model) were also associated with high hypoxia, and we confirmed this within individual cancer types (Supplementary Fig. 3a). Other features associated with elevated hypoxia include smaller CNAs (p = 3.51 × 10−3, = 0.0065, = 0.59, linear mixed-effect model) and more SNVs/Mbp (p = 5.55 × 10−3, = 0.0054, = 0.60, linear mixed-effect model). Since mutational density features can be correlated, we wanted to further test if SNVs per megabase were independently associated with hypoxia after adjusting for the total number of deletions. We created a linear mixed-effect model associating hypoxia with the number of SNVs per megabase while adjusting for cancer type, age, sex, tumour purity and the number of deletions. We also created a second model which lacked our feature of interest, SNVs per megabase, and compared the two models using an ANOVA (see the “Methods” section). The p-value for this comparison was 0.011, suggesting that the number of SNVs per megabase are associated with hypoxia independent of the number of deletions (and other potential confounders included in the models). Overall, hypoxia is associated with increased numbers of most types of somatic mutations.
Considering the strong association of hypoxia with mutational density, we next looked to determine if these were only general effects or selectively affected specific genes or chromosome regions. We leveraged a catalogue of 653 driver mutations25, with CNA, SV and SNV data available for 1096 patients. In cases where a patient had multiple mutations in the same gene (e.g. a CNA and an SNV) we denoted these as compound events. We again used linear-mixed effect models to associate hypoxia with each driver feature across cancers (Fig. 2b). Adjusting for cancer type, tumour purity, age and sex, 10 driver events were associated with hypoxia across cancers (FDR < 0.10, linear-mixed effect models; Supplementary Fig. 2f, Supplementary Table 4). Tumours with mutations in BCL2 (FDR = 7.56 × 10−15, = 0.045, = 0.62, linear-mixed effect model) showed lower levels of hypoxia compared to those without. All alterations of BCL2 in this cohort were SVs, so it is important to note that this association could not be identified from previous exome-sequencing data. Similarly, mutations in the tumour suppressor TP53 were associated with elevated hypoxia across cancers (FDR = 1.97 × 10−12, = 0.043, = 0.59, linear-mixed effect model), consistent with previous descriptions of hypoxia-mediated selection of TP53-mutated cells17 and elevated hypoxia in breast cancers with TP53 mutations16. We also confirmed this association within individual cancer types (Supplementary Fig. 3b). Mutations of the oncogene MYC (FDR = 1.07 × 10−4, = 0.016, = 0.60, linear-mixed effect model) and tumour suppressor PTEN (FDR = 1.50 × 10−2, = 0.0098, = 0.59, linear mixed-effect model) were also associated with elevated hypoxia. Alterations in mitochondrial genes34 were not significantly associated with tumour hypoxia (Supplementary Fig. 3c). Thus, hypoxia is associated with both broad elevation of mutation density of most types of somatic variation, along with a consistent signature of alterations in oncogenes and tumour suppressors across cancers.
Hypoxia-associated mutational signatures
Previous work has used nonnegative matrix factorization to identify distinct mutational processes in cancer cells from endogenous and exogenous agents35. To identify hypoxia-associated mutational processes, we tested if hypoxia score was associated with the proportion of mutations attributed to each mutational signature using linear-mixed effect models. Of the 65 single base substitution (SBS) signatures tested, nine showed differential activity in hypoxic tumours compared to non-hypoxic ones, while controlling for cancer type, tumour purity, age and sex (FDR < 0.10, linear mixed-effect models; Fig. 3a, Supplementary Table 5). Of these, six were more active and three less active in tumours with elevated hypoxia. Since previous work has shown that DNA repair is impaired under hypoxia, it was not surprising to observe that a higher proportion of mutations were attributed to SBS3 (related to defective homologous recombination-based repair) in tumours with elevated hypoxia score (FDR = 1.98 × 10−3, = 0.016, = 0.60, linear-mixed effect model). Further, SBS6 (FDR = 1.98 × 10−3, = 0.0086, = 0.61, linear-mixed effect model) and SBS21 (FDR = 4.31 × 10−2, = 0.0051, = 0.61, linear-mixed effect model), both related to defective DNA mismatch repair, had a higher proportion of attributed mutations with increasing hypoxia. A lower proportion of mutations were also attributed to SBS1, previously related to the deamination of 5-methylcytosine, with increasing hypoxia (FDR = 8.33 × 10−8, = 0.033, = 0.61, linear-mixed effect model).
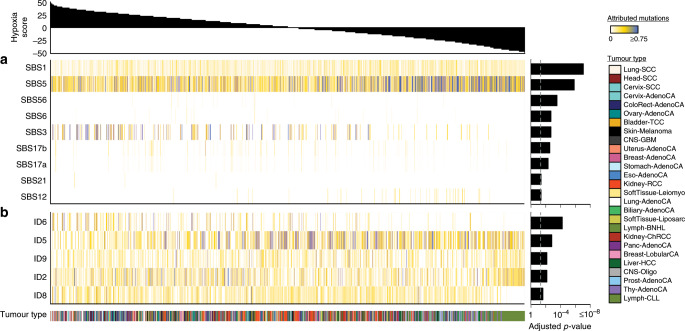
Fig. 3. Hypoxia-associated mutational signatures.
We associated hypoxia with the proportion of mutations attributed to specific mutational signatures using linear-mixed effect models. Hypoxia scores for 1188 independent tumours are shown across the top while FDR-adjusted p-values are shown on the right. a Hypoxia was associated with a series of single base substitution signatures with unknown aetiology including SBS5, SBS17a, SBS17b and SBS12. Some of these mutational signatures may reflect hypoxia-dependent mutational processes. Hypoxia was also associated with a lower proportion of attributed mutations to SBS1, which reflects deamination of 5-methylcytosine, and a higher proportion of attributed mutations to SBS3, which is related to deficiencies in DNA double-strand break repair and homologous recombination. b Several signatures of small insertions and deletions were also associated with hypoxia, including ID6 and ID2, which reflect defective homologous recombination and defective DNA mismatch repair, respectively. ID5, ID9 and ID4, all with unknown aetiology, were significantly associated with hypoxia score. All associations were modelled using linear-mixed effect models while adjusting for cancer type, tumour purity, age and sex.
Intriguingly, hypoxia was also associated with a number of SBS signatures with unknown aetiology (Fig. 3b). The strongest of these was SBS5, where elevated hypoxia was associated with a significantly lower proportion of mutations attributed to the signature (FDR = 1.43 × 10−6, = 0.022, = 0.59, linear-mixed effect model). A significantly lower proportion of mutations were also attributed to SBS12 with increasing hypoxia score (FDR = 4.31 × 10−2, = 0.0066, = 0.60, linear-mixed effect model). In contrast, a higher proportion of mutations were attributed to SBS17a (FDR = 4.80 × 10−3, = 0.0072, = 0.61, linear mixed-effect model) and SBS17b (FDR = 2.83 × 10−3, = 0.079, = 0.61, linear mixed-effect model) with increasing hypoxia.
Analysis of small insertion and deletion (ID) signatures illustrated a similar story. Of the 17 ID signatures analyzed, the activity of 5 were associated with tumour hypoxia scores while controlling for cancer type, tumour purity, age and sex (FDR < 0.10, linear mixed-effect models; Fig. 3b, Supplementary Table 6). Of these, 3 were more active in tumours with elevated hypoxia while 2 were less active in them. The defective homologous recombination signature ID6 (FDR = 5.76 × 10−5, = 0.015, = 0.60, linear mixed-effect model) and defective DNA mismatch repair signature ID2 (FDR = 7.06 × 10−3, = 0.011, = 0.61, linear mixed-effect model) had a higher proportion of attributed mutations as hypoxia score increased. Several signatures with unknown aetiology were also significantly associated with hypoxia score, including ID5 (FDR = 1.54 × 10−3, = 0.016, = 0.60, linear mixed-effect model) and ID9 (FDR = 7.06 × 10−3, = 0.0068, = 0.60, linear mixed-effect model). These data suggest that oxygen levels play a direct or indirect role in the accumulation of specific mutations in cancer cells that are reflected by these signatures.
The subclonal hallmarks of tumour hypoxia
State-of-the-art methods for subclonal reconstruction rely on WGS data36, making the PCAWG dataset ideal for understanding the evolutionary pressures imposed by hypoxia. We and others have shown that some mutations consistently occur early during tumourigenesis while others occur later and that hypoxia is associated with CNAs occurring early in localized prostate cancer16,37,38. To explore if this interaction between the tumour microenvironment and mutational landscape exists more broadly in cancer, we assessed if hypoxia was related to the number of clonal or subclonal mutations across 1188 tumours from 27 cancer types38. Clonal mutations are common to all cells in a tumour while subclonal ones are only present in a subpopulation of cells. We found that elevated hypoxia was significantly associated with an increased number of clonal alterations across cancers (Bonferroni-adjusted p = 4.65 × 10−3, = 0.0074, = 0.60, linear mixed-effect model; Fig. 4a, Supplementary Table 7), particularly clonal SVs (p = 1.17 × 10−5, = 0.013, = 0.60, linear mixed-effect model). In contrast, tumour hypoxia was not significantly associated with the number of subclonal alterations (Bonferroni-adjusted p = 0.28, = 0.0039, = 0.60, linear mixed-effect model; Fig. 4a). Further, consistent with findings in prostate cancer16, hypoxia was not associated with the number of subclones detected (Bonferroni-adjusted p = 0.14, = 0.0051, = 0.60, linear mixed-effect model; Fig. 4a). These data suggest that hypoxia applies a selective pressure on tumours during their early evolution, prior to subclonal diversification.
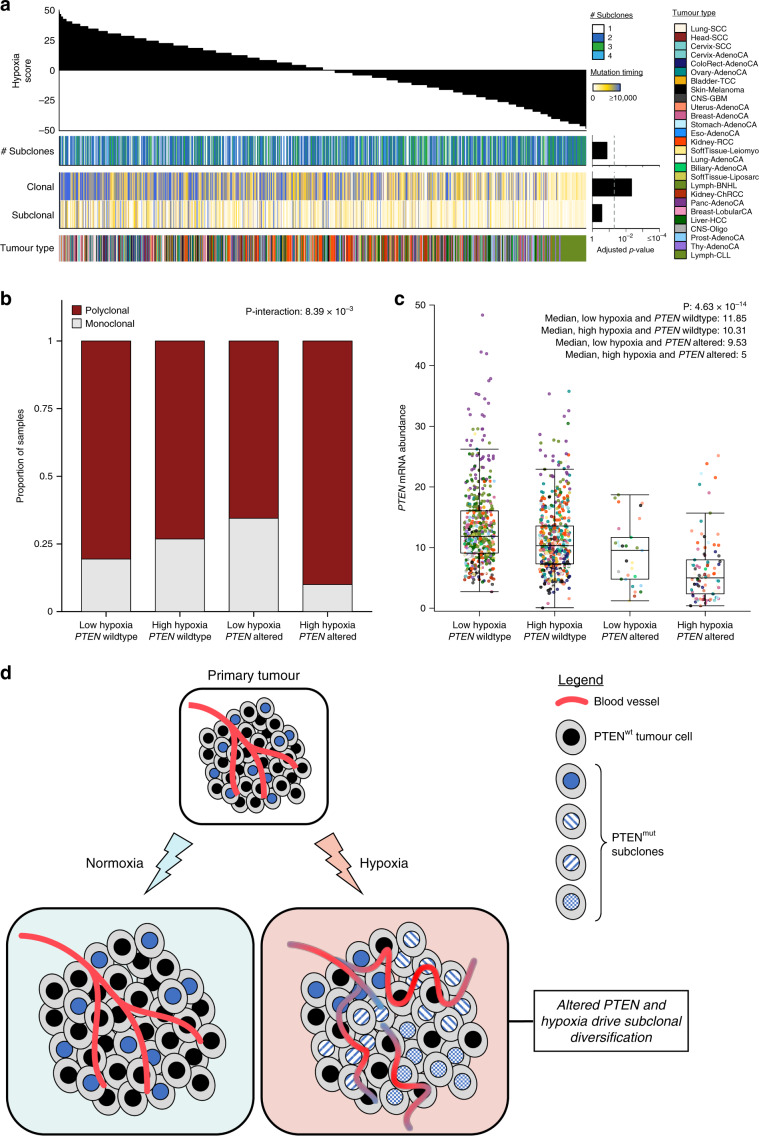
Fig. 4. The subclonal hallmarks of tumour hypoxia.
We associated tumour hypoxia with features related to the subclonal architecture of 1188 independent tumours from 27 cancer types using linear mixed-effect models. a Hypoxia scores are shown along the top while Bonferroni-adjusted p-values are shown on the right. Hypoxia was not associated with the number of subclones in the tumour but elevated hypoxia was associated with a higher number of clonal mutations. b We also observed a significant interaction between hypoxia and altered PTEN where tumours with both of these features were particularly likely to be polyclonal. c The mRNA abundance of PTEN is modulated by both PTEN mutational status and tumour hypoxia. Tumours with altered PTEN and elevated hypoxia have the lowest abundance of PTEN mRNA. mRNA abundance is reported as FPKM with upper-quartile normalization. A Tukey box plot is shown. Box plots represent the median (centre line) and upper and lower quartiles (box limits), and whiskers extend to the minimum and maximum values within 1.5× the interquartile range. All associations were modelled using linear mixed-effect models while adjusting for cancer type, tumour purity, age and sex. d Altered PTEN and hypoxia may drive subclonal diversification. Many primary tumours have elevated hypoxia due to increased demand or decreased supply of oxygen. Tumours with elevated hypoxia tend to have altered PTEN. Elevated hypoxia and altered PTEN may drive subclonal diversification and poor outcomes.
Next, we assessed if the mutational background of a tumour together with its oxygenation level was linked to its evolutionary trajectory. We previously demonstrated that patients with hypoxic polyclonal prostate tumours with loss of the tumour suppressor PTEN tend to have a poor prognosis16. Indeed, here we observed a significant interaction between tumour hypoxia and loss of PTEN in predicting subclonal architecture (pinteraction = 8.39 × 10−3, = 0.60, = 0.87, linear mixed-effect model; Fig. 4b). Specifically, tumours with both of these features tend to have a polyclonal architecture across cancers. The downstream impact of this interaction between the genome and the tumour microenvironment was observed in RNA data: tumours with both altered PTEN and elevated hypoxia had the lowest abundance of PTEN mRNA (p = 4.63 × 10−14, = 0.054, = 0.47, linear mixed-effect model; Fig. 4c). Thus, the evolutionary trajectory of a tumour may be driven by the presence of a mutation in a specific microenvironmental context (Fig. 4d).
Discussion
Hypoxia is a feature of many solid and liquid tumours and is associated with aggressive disease. We calculated hypoxia scores for 1188 tumours from 27 cancer types and showed the vast heterogeneity that exists in this microenvironmental feature within and across cancer types. This reinforces previous pushes for careful patient selection in prospective trials of hypoxia-targeting agents16.
For the first time, we characterized the pan-cancer whole-genome correlates of tumour hypoxia. We show the broad influence of the hypoxia-associated mutator phenotype: elevated hypoxia is associated with increased mutational load across all mutational classes (i.e. CNAs, SVs and SNVs). This supports previous in vitro work that demonstrated the contextual synthetic lethality of PARP inhibition in cells with defective DNA repair due to hypoxia39. Regarding this co-occurrence of genomic instability and hypoxia, our group16 and others40 have previously described this metabolic reprogramming as a series of distinct genomic alterations. This is supported by our finding that alterations in TP53, MYC and PTEN are more common in tumours with elevated hypoxia across cancers. Supporting these findings, previous in vitro work has shown that heterogeneous populations of cells where a small subpopulation have mutant TP53 can rapidly expand under cycling hypoxia to become the major subpopulation due to deficient apoptosis and selection17. We have also previously shown that tumours with TP53 mutations have elevated hypoxia within individual breast cancer subtypes, confirming that this association is not simply reflecting previously described molecular subtypes16.
Our study cannot conclusively say whether hypoxia exerts a selective pressure that enriches for specific genomic alterations or if these genomic changes directly result in hypoxia. Experimental studies of single genes support that both effects may contribute to the associations we describe17,22,41–43. While we have not specifically included in vitro experimental validation data in this report, we and others have previously validated associations first revealed by analysis of mRNA-based hypoxia signatures. For example, our group previously described microRNA-133a-3p as a hypoxia-associated miRNA in prostate cancer based on mRNA signature-based associations across multiple independent datasets16. We went on to validate that microRNA-133a-3p was indeed induced under hypoxia in multiple prostate cancer cell lines and confirmed its capacity to modulate cell proliferation and invasion. Similarly, Ye et al. applied hypoxia signatures to 10 independent datasets of cell lines and primary tumour fragments under hypoxia and normoxia44. These 10 datasets represented seven cancer types and within each dataset samples under hypoxia showed higher hypoxia scores compared to normoxic samples. Further, they generated predictions of drugs that would be more or less potent under hypoxia and validated four drug–hypoxia interactions in vitro. These data illustrate that hypoxia signatures applied to large cohorts of primary tumours can generate reliable hypotheses, many of which have been validated in controlled systems. However, some aerobic cancer cells may also mimic the biological state of hypoxia (i.e., pseudohypoxia) and this may affect signature-derived hypoxia estimates. Further, while pimonidazole (which was used to develop the Ragnum hypoxia signature32) reflects oxygen tensions below 10 mmHg (1.3% O2), it is difficult to directly relate hypoxia signature scores with oxygen tension45. Overall, hypoxia signalling can be distinct from microenvironmental hypoxia and this remains a critical caveat of this study.
Diving into the mutational processes related to hypoxia, we confirmed that several SBS and small indel signatures related to impaired DNA repair were associated with hypoxia. This raises the potential confounder that because hypoxic tumours have more mutations, we have more power to detect related mutational signatures. However, we demonstrated that hypoxia is indeed strongly associated with many mutational signatures with unknown aetiology, particularly SBS5, which is found in nearly all cancer types. Modelling these associations in vitro is particularly difficult and these data provide a high confidence measure of the mutational signatures that may be directly or indirectly driven by tumour oxygen levels. It is difficult to disentangle the timing of these events: whether a specific driver mutation gives rise to a specific mutational signature or if these are separate processes. Better mapping of the evolutionary timing of hypoxia will be particularly important in addressing this question and the advent of hypoxia signatures may facilitate future studies in this area.
We observed a significant association between elevated hypoxia and the number of clonal mutations. This supports the idea that hypoxia is an early event in cancer, as we have suggested previously16, and other models that link hypoxia to genomic instability and downstream clonal selection20,42. Previous work has also demonstrated that patients with allelic loss of PTEN and elevated hypoxia rapidly relapse after definitive treatment for localized prostate cancer16. Here, we showed that tumours with alterations in PTEN and elevated hypoxia are enriched for a polyclonal tumour architecture. This illustrates the joint influence of the tumour mutational landscape and microenvironment in guiding evolutionary trajectories across cancers. Further, these data suggest that increased subclonal diversification may be a novel route via which PTEN drives aggressive tumour phenotypes, in concert with tumour hypoxia, and this can be better defined with future back-translational in vitro experiments. The PCAWG dataset is the largest publicly available pan-cancer dataset to date and this limits our ability to validate our discoveries in independent datasets. Ultimately, it will be necessary to validate our findings in large, independent cohorts. Hypotheses generated here, particularly those around hypoxia and tumour evolution, will require long term, systematic in vitro modelling and will be the subject of future studies. Overall, this work shows that a hypoxic tumour microenvironment is associated with specific mutational processes and distinct somatic mutational profiles, and may direct the subclonal architecture of cancers.
Methods
Pan-cancer hypoxia scoring
Hypoxia scores were calculated for all 1188 tumours with mRNA abundance data (FPKM with upper-quartile normalization) using mRNA-abundance-based signatures of tumour hypoxia developed previously by Winter et al.31, Buffa et al. 30 and Ragnum et al. 32, as described previously14,16 (Supplementary Data 1). Briefly, patients with the top 50% of mRNA abundance values for each gene in a signature were given a score of +1. Patients with the bottom 50% of mRNA abundance values for that gene were given a score of −1. This was repeated for every gene in the signature to generate a hypoxia score for each patient, and this process was repeated for each of the three signatures used in the study. High scores suggest that the tumour was hypoxic and low scores are indicative of normoxia.
Hypoxia score comparison
To compare hypoxia scores generated by the different signatures, the median hypoxia score was calculated for each of the PCAWG cancer types based on each signature. The median hypoxia scores from each signature were then scaled from +1 to −1 using the plotrix package (v3.7). Scaled median hypoxia values for the PCAWG cancer types were also compared to scaled median hypoxia values from previously published16 TCGA data between comparable cancer groups. The groups compared are as follows (PCAWG cancer type and TCGA cancer type): Bladder-TCC and BLCA; Breast-AdenoCA and BRCA; Cervix-SCC and CESC; CNS-GBM and GBM; ColoRect-AdenoCA and COADREAD; Head-SCC and HNSC; Kidney-RCC and KIRC; Liver-HCC and LIHC; Lung-AdenoCA and LUAD; Lung-SCC and LUSC; Ovary-AdenoCA and OV; Panc-AdenoCA and PAAD; Prost-AdenoCA and PRAD; Skin-Melanoma and SKCM; Thy-AdenoCA and THCA; Uterus-Adeno and UCEC. Of the 27 cancer types in PCAWG, 11 (Cervix-AdenoCA, Stomach-AdenoCA, Eso-AdenoCA, Breast-LobularCA, SoftTissue-Leiomyo, Lymph-BNHL, SoftTissue-Liposarc, Biliary-AdenoCA, Kidney-ChRCC, CNS-Oligo and Lymph-CLL) did not have hypoxia data from comparable cancers in TCGA and were not used for the comparison16. For Spearman’s correlations, p-values were calculated using algorithm AS89.
Tumour vs. normal hypoxia comparison
Previously calculated pan-cancer tumour hypoxia scores were gathered for 7791 independent tumours from 19 cancer types based on the Buffa hypoxia signature16. Hypoxia scores were then calculated for all normal tissue samples related to the 19 cancer types with hypoxia scores (n = 640 independent normal tissue samples). Tumour hypoxia scores were compared to normal tissue hypoxia scores for each tissue type where at least 15 normal tissue samples were available (Mann–Whitney U-test). A total of 5649 independent tumours and 625 independent normal tissue samples were evaluated in the comparisons.
Linear mixed-effect models
We used linear mixed-effect models to associate hypoxia with features of interest (e.g., PGA, TP53 mutational status, etc.) across cancers using the lme4 package (v1.1-17). For each feature of interest, we compared a full model (i.e., a model with the feature of interest) to a null model (i.e. a model without the feature of interest) using an ANOVA to determine if hypoxia was significantly associated with the feature of interest across cancers. A generic example of this is shown below with Eqs. (1) and (2):
| 1 |
| 2 |
All models were adjusted for tumour purity, patient age and sex27,28. Cancer type was incorporated as a random effect in every model. This allowed us to consider a different baseline value for the feature of interest for each cancer type. For each model a conditional R2 value is reported () which reflects the variance explained by the fixed and random factors29. We also report marginal R2 values for each model () which reflect the variance explained by the fixed factors only29. and values were calculated as described previously29.
All model diagnostics were done using the DHARMa package (0.2.0) which uses a simulation-based approach to create standardized residuals46. For each model, scaled residuals were generated using the simulateResiduals function. The full model was used as the input for fittedModel parameter and 1000 simulations were run. For correctly specified models, the scaled residuals were expected to be uniformly distributed and this was verified for each full model. We also compared the standardized residuals to the rank transformed predicted values to assess deviations from uniformity for each full model.
Mutational density analysis
Previously published data for 15 mutational density and summary features were downloaded for 1188 tumours33. We used linear mixed-effect models to associate each feature with hypoxia score across cancers and compared each full model with a null model. Cancer type was incorporated as a random effect in each model while tumour purity, age and sex were incorporated as fixed effects. Tumours belonging to cancer types with fewer than 15 samples were excluded from the analysis. A Bonferroni p-value adjustment was applied to the p-values from linear mixed-effect modelling since fewer than 20 tests were conducted. All models were adjusted for tumour purity based on previously published purity data33. The full model for evaluating PGA is shown below as an example as follows:
| 3 |
To assess if SNVs per megabase were independently associated with hypoxia after adjusting for the total number of deletions we created two linear mixed-effect models. The full model associated hypoxia with SNVs per megabase while adjusting for cancer type, age, sex, tumour purity and the number of deletions (Eq. (4)). For comparison, a null model was created without our feature of interest, SNVs per megabase (Eq. (5)). The two models were compared using an ANOVA.
| 4 |
| 5 |
Driver mutations analysis
Data for driver mutations was first summarized at the gene level for 1096 tumours with previously published driver mutation data25. For each of the 653 driver features, we summarized if a patient had an SNV, CNA or SV. Some tumours had more than one type of event in a gene (e.g. a CNA and an SNV) and these events were classified as compound events. We then used linear mixed-effect models to associate the mutational status of each gene with hypoxia score and compared each full model with a null model. Cancer type was incorporated as a random effect in each model while tumour purity, age and sex were incorporated as fixed effects. The driver mutation analysis did not specifically consider the type of mutation in the gene and only considered if the gene had a mutation or was wildtype. Tumours belonging to cancer types with fewer than 15 samples were excluded from the analysis. An FDR adjustment was applied to the p-values from linear mixed-effect modelling. The full model for evaluating PTEN is shown below as an example as follows:
| 6 |
Mutational signature analysis
Previously published data for mutations attributed to various specific signatures was downloaded for 1188 tumours35. For each tumour, we calculated the proportion of total mutations attributed to each mutational signature. The proportion of mutations attributed to each signature were calculated by dividing the number of mutations attributed to each signature by the total number of mutations in the tumour. We used linear mixed-effect models to associate the proportion of mutations attributed to each signature with hypoxia score and compared each full model with a null model. Cancer type was incorporated as a random effect in each model while purity, age and sex were incorporated as fixed effects. Tumours belonging to cancer types with fewer than 15 samples were excluded from the analysis. An FDR adjustment was applied to the p-values from linear mixed-effect modelling. The full model for SBS1 is shown below as an example as follows:
| 7 |
Subclonality analysis
Previously reported38 subclonal reconstruction data was used to summarize the number of clonal and subclonal mutations in all 1188 tumours. We used linear mixed-effect models to associate the number of these timed mutations with hypoxia score and compared each full model with a null model. Cancer type was incorporated as a random effect in each model while purity, age and sex were incorporated as fixed effects. Tumours belonging to cancer types with fewer than 15 samples were excluded from the analysis. A Bonferroni adjustment was applied to the p-values from linear mixed-effect modelling since fewer than 20 tests were conducted.
The number of subclones was calculated for all 1188 tumours based on the number of clusters of cells identified in each sample. A linear mixed-effects model was used to associate the number of subclones with hypoxia score and this model was compared to a null model. Cancer type was incorporated as a random effect while purity, age and sex were incorporated as fixed effects. Tumours belonging to cancer types with fewer than 15 samples were excluded from the analysis. The full model for associating the number of subclones with hypoxia score is shown below as follows:
| 8 |
Patients with only one identified cluster of cells were defined as monoclonal and patients with more than one identified cluster of cells were defined as polyclonal37. Hypoxia scores were median dichotomized to classify patients as hypoxic or normoxic. To test for an interaction between tumour hypoxia and PTEN mutational status in selecting for a particular subclonal architecture, we used linear mixed-effect models together with an ANOVA. An interaction model was first created where the relationship between the hypoxia scores and PTEN mutational status was modelled as an interaction (Eq. (9)). An additive model was also created where the relationship between hypoxia scores and PTEN mutational status was modelled in an additive manner (Eq. (10)):
| 9 |
| 10 |
The two models were compared using an ANOVA to test if hypoxia scores significantly interact with PTEN mutational status. Tumours belonging to cancer types with fewer than 15 samples were excluded from the analysis. The full model diagnostics were carried out using the DHARMa package, as described above.
All data analysis was performed in the R statistical environment (v3.4.3). Data visualization was performed using the BPG package47 (v5.9.1). Figures were compiled using Inkscape (v0.91).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Supplementary information
Description of Additional Supplementary Files
Acknowledgements
The authors gratefully thank Dr. Marianne Koritzinsky for insightful suggestions and thank Jenna E. van Leeuwen for visualization support. The authors also thank all members of the Boutros and Bristow labs for helpful suggestions. We acknowledge the contributions of the many clinical networks across ICGC and TCGA who provided samples and data to the PCAWG Consortium, and the contributions of the Technical Working Group and the Germline Working Group of the PCAWG Consortium for collation, realignment and harmonised variant calling of the cancer genomes used in this study. We thank the patients and their families for their participation in the individual ICGC and TCGA projects. This study was conducted with the support of Movember funds through Prostate Cancer Canada, and with the additional support of the Ontario Institute for Cancer Research, funded by the Government of Ontario. This work was supported by Prostate Cancer Canada and is proudly funded by the Movember Foundation Team Grant T2013 and #RS2014-01. Paul C. Boutros was supported by a Terry Fox Research Institute New Investigator Award, a Prostate Cancer Canada Rising Star Fellowship, a CIHR New Investigator Award, a CIHR Project Grant, the Government of Canada through Genome Canada and the Ontario Genomics Institute (OGI-125), Canadian Cancer Society (grant #705649) and by the University of California. This work has been funded by a fellowship from the Canadian Institutes of Health Research to Vinayak Bhandari. Constance Li was supported by an award from the Government of Ontario. The authors gratefully thank the Princess Margaret Cancer Centre Foundation and Radiation Medicine Programme Academic Enrichment Fund for support (to Robert G. Bristow). This work was supported by a Terry Fox Research Institute Programme Project Grant. Robert G. Bristow is a recipient of a Canadian Cancer Society Research Scientist Award. Laboratory work for R.G.B is supported by the CRUK Manchester Institute through Cancer Research UK. This work was supported by the NIH/NCI through award number P30CA016042.
Author contributions
Bioinformatics and statistical analysis: V.B. Data processing: V.B., C.H.L. Data visualization: V.B. Supervised research: R.G.B., P.C.B. Initiated the project: V.B., P.C.B. Wrote the first draft of the manuscript: V.B. Approved the manuscript: all authors.
Data availability
Aligned sequencing data, somatic and germline variant calls, single nucleotide variants, indels, copy-number alterations, structural variants, mutational signatures, subclonal reconstructions, transcript abundances and other core data generated by the PCAWG Consortium are described here25 and available for download at dcc.icgc.org/releases/PCAWG. Additional information on accessing the data, including raw read files, can be found at docs.icgc.org/pcawg/data. In accordance with the data access policies of the ICGC and TCGA projects, most molecular, clinical and specimen data are in an open tier which does not require approval for access.
To access potentially identifiable information, such as germline alleles and underlying sequencing data, researchers will need to apply to the TCGA Data Access Committee (DAC) via dbGaP (dbgap.ncbi.nlm.nih.gov/aa/wga.cgi?page = login) for access to the TCGA portion of the dataset, and to the ICGC Data Access Compliance Office (DACO; icgc.org/daco) for the ICGC portion of the dataset. To access somatic single nucleotide variant data derived from TCGA donors, researchers will also need to obtain dbGaP authorization.
The analyses in this paper used several PCAWG datasets that were derived from the raw sequencing data and variant calls. The individual datasets are available at Synapse (synapse.org) and are denoted with accession numbers starting with syn. All of these datasets are also mirrored at dcc.icgc.org. The datasets encompass: clinical data for each patient, including information about age and sex (syn10389158, dcc.icgc.org/releases/PCAWG/clinical_and_histology); histopathology data for each tumour (syn10389164, dcc.icgc.org/releases/PCAWG/clinical_and_histology); mRNA abundance data for each tumour (syn5553991, dcc.icgc.org/releases/PCAWG/transcriptome/gene_expression); driver mutations for each tumour spanning all classes of variants (syn11639581, dcc.icgc.org/releases/PCAWG/driver_mutations); single base substitution and small indel signature exposures for each tumour (syn8366024, dcc.icgc.org/releases/PCAWG/mutational_signatures); subclonal architecture, tumour evolution and tumour purity data for each tumour (syn8272483, dcc.icgc.org/releases/PCAWG/subclonal_reconstruction).
Code availability
The core computational pipelines used by the PCAWG Consortium for alignment, quality control and variant calling are available to the public at dockstore.org/search?search = pcawg under the GNU General Public License v3.0, which allows for reuse and distribution. Code specific to this manuscript is available upon request.
Competing interests
The authors declare no competing interests.
Footnotes
Peer review information Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
PCAWG Consortium members and their affiliations appears online
Change history
12/8/2022
A Correction to this paper has been published: 10.1038/s41467-022-32339-4
Contributor Information
Robert G. Bristow, Email: robert.bristow@manchester.ac.uk
Paul C. Boutros, Email: pboutros@mednet.ucla.edu
PCAWG Consortium:
Lauri A. Aaltonen, Federico Abascal, Adam Abeshouse, Hiroyuki Aburatani, David J. Adams, Nishant Agrawal, Keun Soo Ahn, Sung-Min Ahn, Hiroshi Aikata, Rehan Akbani, Kadir C. Akdemir, Hikmat Al-Ahmadie, Sultan T. Al-Sedairy, Fatima Al-Shahrour, Malik Alawi, Monique Albert, Kenneth Aldape, Ludmil B. Alexandrov, Adrian Ally, Kathryn Alsop, Eva G. Alvarez, Fernanda Amary, Samirkumar B. Amin, Brice Aminou, Ole Ammerpohl, Matthew J. Anderson, Yeng Ang, Davide Antonello, Pavana Anur, Samuel Aparicio, Elizabeth L. Appelbaum, Yasuhito Arai, Axel Aretz, Koji Arihiro, Shun-ichi Ariizumi, Joshua Armenia, Laurent Arnould, Sylvia Asa, Yassen Assenov, Gurnit Atwal, Sietse Aukema, J. Todd Auman, Miriam R. R. Aure, Philip Awadalla, Marta Aymerich, Gary D. Bader, Adrian Baez-Ortega, Matthew H. Bailey, Peter J. Bailey, Miruna Balasundaram, Saianand Balu, Pratiti Bandopadhayay, Rosamonde E. Banks, Stefano Barbi, Andrew P. Barbour, Jonathan Barenboim, Jill Barnholtz-Sloan, Hugh Barr, Elisabet Barrera, John Bartlett, Javier Bartolome, Claudio Bassi, Oliver F. Bathe, Daniel Baumhoer, Prashant Bavi, Stephen B. Baylin, Wojciech Bazant, Duncan Beardsmore, Timothy A. Beck, Sam Behjati, Andreas Behren, Beifang Niu, Cindy Bell, Sergi Beltran, Christopher Benz, Andrew Berchuck, Anke K. Bergmann, Erik N. Bergstrom, Benjamin P. Berman, Daniel M. Berney, Stephan H. Bernhart, Rameen Beroukhim, Mario Berrios, Samantha Bersani, Johanna Bertl, Miguel Betancourt, Vinayak Bhandari, Shriram G. Bhosle, Andrew V. Biankin, Matthias Bieg, Darell Bigner, Hans Binder, Ewan Birney, Michael Birrer, Nidhan K. Biswas, Bodil Bjerkehagen, Tom Bodenheimer, Lori Boice, Giada Bonizzato, Johann S. De Bono, Arnoud Boot, Moiz S. Bootwalla, Ake Borg, Arndt Borkhardt, Keith A. Boroevich, Ivan Borozan, Christoph Borst, Marcus Bosenberg, Mattia Bosio, Jacqueline Boultwood, Guillaume Bourque, Paul C. Boutros, G. Steven Bova, David T. Bowen, Reanne Bowlby, David D. L. Bowtell, Sandrine Boyault, Rich Boyce, Jeffrey Boyd, Alvis Brazma, Paul Brennan, Daniel S. Brewer, Arie B. Brinkman, Robert G. Bristow, Russell R. Broaddus, Jane E. Brock, Malcolm Brock, Annegien Broeks, Angela N. Brooks, Denise Brooks, Benedikt Brors, Søren Brunak, Timothy J. C. Bruxner, Alicia L. Bruzos, Alex Buchanan, Ivo Buchhalter, Christiane Buchholz, Susan Bullman, Hazel Burke, Birgit Burkhardt, Kathleen H. Burns, John Busanovich, Carlos D. Bustamante, Adam P. Butler, Atul J. Butte, Niall J. Byrne, Anne-Lise Børresen-Dale, Samantha J. Caesar-Johnson, Andy Cafferkey, Declan Cahill, Claudia Calabrese, Carlos Caldas, Fabien Calvo, Niedzica Camacho, Peter J. Campbell, Elias Campo, Cinzia Cantù, Shaolong Cao, Thomas E. Carey, Joana Carlevaro-Fita, Rebecca Carlsen, Ivana Cataldo, Mario Cazzola, Jonathan Cebon, Robert Cerfolio, Dianne E. Chadwick, Dimple Chakravarty, Don Chalmers, Calvin Wing Yiu Chan, Kin Chan, Michelle Chan-Seng-Yue, Vishal S. Chandan, David K. Chang, Stephen J. Chanock, Lorraine A. Chantrill, Aurélien Chateigner, Nilanjan Chatterjee, Kazuaki Chayama, Hsiao-Wei Chen, Jieming Chen, Ken Chen, Yiwen Chen, Zhaohong Chen, Andrew D. Cherniack, Jeremy Chien, Yoke-Eng Chiew, Suet-Feung Chin, Juok Cho, Sunghoon Cho, Jung Kyoon Choi, Wan Choi, Christine Chomienne, Zechen Chong, Su Pin Choo, Angela Chou, Angelika N. Christ, Elizabeth L. Christie, Eric Chuah, Carrie Cibulskis, Kristian Cibulskis, Sara Cingarlini, Peter Clapham, Alexander Claviez, Sean Cleary, Nicole Cloonan, Marek Cmero, Colin C. Collins, Ashton A. Connor, Susanna L. Cooke, Colin S. Cooper, Leslie Cope, Vincenzo Corbo, Matthew G. Cordes, Stephen M. Cordner, Isidro Cortés-Ciriano, Kyle Covington, Prue A. Cowin, Brian Craft, David Craft, Chad J. Creighton, Yupeng Cun, Erin Curley, Ioana Cutcutache, Karolina Czajka, Bogdan Czerniak, Rebecca A. Dagg, Ludmila Danilova, Maria Vittoria Davi, Natalie R. Davidson, Helen Davies, Ian J. Davis, Brandi N. Davis-Dusenbery, Kevin J. Dawson, Francisco M. De La Vega, Ricardo De Paoli-Iseppi, Timothy Defreitas, Angelo P. Dei Tos, Olivier Delaneau, John A. Demchok, Jonas Demeulemeester, German M. Demidov, Deniz Demircioğlu, Nening M. Dennis, Robert E. Denroche, Stefan C. Dentro, Nikita Desai, Vikram Deshpande, Amit G. Deshwar, Christine Desmedt, Jordi Deu-Pons, Noreen Dhalla, Neesha C. Dhani, Priyanka Dhingra, Rajiv Dhir, Anthony DiBiase, Klev Diamanti, Li Ding, Shuai Ding, Huy Q. Dinh, Luc Dirix, HarshaVardhan Doddapaneni, Nilgun Donmez, Michelle T. Dow, Ronny Drapkin, Oliver Drechsel, Ruben M. Drews, Serge Serge, Tim Dudderidge, Ana Dueso-Barroso, Andrew J. Dunford, Michael Dunn, Lewis Jonathan Dursi, Fraser R. Duthie, Ken Dutton-Regester, Jenna Eagles, Douglas F. Easton, Stuart Edmonds, Paul A. Edwards, Sandra E. Edwards, Rosalind A. Eeles, Anna Ehinger, Juergen Eils, Roland Eils, Adel El-Naggar, Matthew Eldridge, Kyle Ellrott, Serap Erkek, Georgia Escaramis, Shadrielle M. G. Espiritu, Xavier Estivill, Dariush Etemadmoghadam, Jorunn E. Eyfjord, Bishoy M. Faltas, Daiming Fan, Yu Fan, William C. Faquin, Claudiu Farcas, Matteo Fassan, Aquila Fatima, Francesco Favero, Nodirjon Fayzullaev, Ina Felau, Sian Fereday, Martin L. Ferguson, Vincent Ferretti, Lars Feuerbach, Matthew A. Field, J. Lynn Fink, Gaetano Finocchiaro, Cyril Fisher, Matthew W. Fittall, Anna Fitzgerald, Rebecca C. Fitzgerald, Adrienne M. Flanagan, Neil E. Fleshner, Paul Flicek, John A. Foekens, Kwun M. Fong, Nuno A. Fonseca, Christopher S. Foster, Natalie S. Fox, Michael Fraser, Scott Frazer, Milana Frenkel-Morgenstern, William Friedman, Joan Frigola, Catrina C. Fronick, Akihiro Fujimoto, Masashi Fujita, Masashi Fukayama, Lucinda A. Fulton, Robert S. Fulton, Mayuko Furuta, P. Andrew Futreal, Anja Füllgrabe, Stacey B. Gabriel, Steven Gallinger, Carlo Gambacorti-Passerini, Jianjiong Gao, Shengjie Gao, Levi Garraway, Øystein Garred, Erik Garrison, Dale W. Garsed, Nils Gehlenborg, Josep L. L. Gelpi, Joshy George, Daniela S. Gerhard, Clarissa Gerhauser, Jeffrey E. Gershenwald, Mark Gerstein, Moritz Gerstung, Gad Getz, Mohammed Ghori, Ronald Ghossein, Nasra H. Giama, Richard A. Gibbs, Bob Gibson, Anthony J. Gill, Pelvender Gill, Dilip D. Giri, Dominik Glodzik, Vincent J. Gnanapragasam, Maria Elisabeth Goebler, Mary J. Goldman, Carmen Gomez, Santiago Gonzalez, Abel Gonzalez-Perez, Dmitry A. Gordenin, James Gossage, Kunihito Gotoh, Ramaswamy Govindan, Dorthe Grabau, Janet S. Graham, Robert C. Grant, Anthony R. Green, Eric Green, Liliana Greger, Nicola Grehan, Sonia Grimaldi, Sean M. Grimmond, Robert L. Grossman, Adam Grundhoff, Gunes Gundem, Qianyun Guo, Manaswi Gupta, Shailja Gupta, Ivo G. Gut, Marta Gut, Jonathan Göke, Gavin Ha, Andrea Haake, David Haan, Siegfried Haas, Kerstin Haase, James E. Haber, Nina Habermann, Faraz Hach, Syed Haider, Natsuko Hama, Freddie C. Hamdy, Anne Hamilton, Mark P. Hamilton, Leng Han, George B. Hanna, Martin Hansmann, Nicholas J. Haradhvala, Olivier Harismendy, Ivon Harliwong, Arif O. Harmanci, Eoghan Harrington, Takanori Hasegawa, David Haussler, Steve Hawkins, Shinya Hayami, Shuto Hayashi, D. Neil Hayes, Stephen J. Hayes, Nicholas K. Hayward, Steven Hazell, Yao He, Allison P. Heath, Simon C. Heath, David Hedley, Apurva M. Hegde, David I. Heiman, Michael C. Heinold, Zachary Heins, Lawrence E. Heisler, Eva Hellstrom-Lindberg, Mohamed Helmy, Seong Gu Heo, Austin J. Hepperla, José María Heredia-Genestar, Carl Herrmann, Peter Hersey, Julian M. Hess, Holmfridur Hilmarsdottir, Jonathan Hinton, Satoshi Hirano, Nobuyoshi Hiraoka, Katherine A. Hoadley, Asger Hobolth, Ermin Hodzic, Jessica I. Hoell, Steve Hoffmann, Oliver Hofmann, Andrea Holbrook, Aliaksei Z. Holik, Michael A. Hollingsworth, Oliver Holmes, Robert A. Holt, Chen Hong, Eun Pyo Hong, Jongwhi H. Hong, Gerrit K. Hooijer, Henrik Hornshøj, Fumie Hosoda, Yong Hou, Volker Hovestadt, William Howat, Alan P. Hoyle, Ralph H. Hruban, Jianhong Hu, Taobo Hu, Xing Hua, Kuan-lin Huang, Mei Huang, Mi Ni Huang, Vincent Huang, Yi Huang, Wolfgang Huber, Thomas J. Hudson, Michael Hummel, Jillian A. Hung, David Huntsman, Ted R. Hupp, Jason Huse, Matthew R. Huska, Barbara Hutter, Carolyn M. Hutter, Daniel Hübschmann, Christine A. Iacobuzio-Donahue, Charles David Imbusch, Marcin Imielinski, Seiya Imoto, William B. Isaacs, Keren Isaev, Shumpei Ishikawa, Murat Iskar, S. M. Ashiqul Islam, Michael Ittmann, Sinisa Ivkovic, Jose M. G. Izarzugaza, Jocelyne Jacquemier, Valerie Jakrot, Nigel B. Jamieson, Gun Ho Jang, Se Jin Jang, Joy C. Jayaseelan, Reyka Jayasinghe, Stuart R. Jefferys, Karine Jegalian, Jennifer L. Jennings, Seung-Hyup Jeon, Lara Jerman, Yuan Ji, Wei Jiao, Peter A. Johansson, Amber L. Johns, Jeremy Johns, Rory Johnson, Todd A. Johnson, Clemency Jolly, Yann Joly, Jon G. Jonasson, Corbin D. Jones, David R. Jones, David T. W. Jones, Nic Jones, Steven J. M. Jones, Jos Jonkers, Young Seok Ju, Hartmut Juhl, Jongsun Jung, Malene Juul, Randi Istrup Juul, Sissel Juul, Natalie Jäger, Rolf Kabbe, Andre Kahles, Abdullah Kahraman, Vera B. Kaiser, Hojabr Kakavand, Sangeetha Kalimuthu, Christof von Kalle, Koo Jeong Kang, Katalin Karaszi, Beth Karlan, Rosa Karlić, Dennis Karsch, Katayoon Kasaian, Karin S. Kassahn, Hitoshi Katai, Mamoru Kato, Hiroto Katoh, Yoshiiku Kawakami, Jonathan D. Kay, Stephen H. Kazakoff, Marat D. Kazanov, Maria Keays, Electron Kebebew, Richard F. Kefford, Manolis Kellis, James G. Kench, Catherine J. Kennedy, Jules N. A. Kerssemakers, David Khoo, Vincent Khoo, Narong Khuntikeo, Ekta Khurana, Helena Kilpinen, Hark Kyun Kim, Hyung-Lae Kim, Hyung-Yong Kim, Hyunghwan Kim, Jaegil Kim, Jihoon Kim, Jong K. Kim, Youngwook Kim, Tari A. King, Wolfram Klapper, Kortine Kleinheinz, Leszek J. Klimczak, Stian Knappskog, Michael Kneba, Bartha M. Knoppers, Youngil Koh, Jan Komorowski, Daisuke Komura, Mitsuhiro Komura, Gu Kong, Marcel Kool, Jan O. Korbel, Viktoriya Korchina, Andrey Korshunov, Michael Koscher, Roelof Koster, Zsofia Kote-Jarai, Antonios Koures, Milena Kovacevic, Barbara Kremeyer, Helene Kretzmer, Markus Kreuz, Savitri Krishnamurthy, Dieter Kube, Kiran Kumar, Pardeep Kumar, Sushant Kumar, Yogesh Kumar, Ritika Kundra, Kirsten Kübler, Ralf Küppers, Jesper Lagergren, Phillip H. Lai, Peter W. Laird, Sunil R. Lakhani, Christopher M. Lalansingh, Emilie Lalonde, Fabien C. Lamaze, Adam Lambert, Eric Lander, Pablo Landgraf, Luca Landoni, Anita Langerød, Andrés Lanzós, Denis Larsimont, Erik Larsson, Mark Lathrop, Loretta M. S. Lau, Chris Lawerenz, Rita T. Lawlor, Michael S. Lawrence, Alexander J. Lazar, Ana Mijalkovic Lazic, Xuan Le, Darlene Lee, Donghoon Lee, Eunjung Alice Lee, Hee Jin Lee, Jake June-Koo Lee, Jeong-Yeon Lee, Juhee Lee, Ming Ta Michael Lee, Henry Lee-Six, Kjong-Van Lehmann, Hans Lehrach, Dido Lenze, Conrad R. Leonard, Daniel A. Leongamornlert, Ignaty Leshchiner, Louis Letourneau, Ivica Letunic, Douglas A. Levine, Lora Lewis, Tim Ley, Chang Li, Constance H. Li, Haiyan Irene Li, Jun Li, Lin Li, Shantao Li, Siliang Li, Xiaobo Li, Xiaotong Li, Xinyue Li, Yilong Li, Han Liang, Sheng-Ben Liang, Peter Lichter, Pei Lin, Ziao Lin, W. M. Linehan, Ole Christian Lingjærde, Dongbing Liu, Eric Minwei Liu, Fei-Fei Fei Liu, Fenglin Liu, Jia Liu, Xingmin Liu, Julie Livingstone, Dimitri Livitz, Naomi Livni, Lucas Lochovsky, Markus Loeffler, Georgina V. Long, Armando Lopez-Guillermo, Shaoke Lou, David N. Louis, Laurence B. Lovat, Yiling Lu, Yong-Jie Lu, Youyong Lu, Claudio Luchini, Ilinca Lungu, Xuemei Luo, Hayley J. Luxton, Andy G. Lynch, Lisa Lype, Cristina López, Carlos López-Otín, Eric Z. Ma, Yussanne Ma, Gaetan MacGrogan, Shona MacRae, Geoff Macintyre, Tobias Madsen, Kazuhiro Maejima, Andrea Mafficini, Dennis T. Maglinte, Arindam Maitra, Partha P. Majumder, Luca Malcovati, Salem Malikic, Giuseppe Malleo, Graham J. Mann, Luisa Mantovani-Löffler, Kathleen Marchal, Giovanni Marchegiani, Elaine R. Mardis, Adam A. Margolin, Maximillian G. Marin, Florian Markowetz, Julia Markowski, Jeffrey Marks, Tomas Marques-Bonet, Marco A. Marra, Luke Marsden, John W. M. Martens, Sancha Martin, Jose I. Martin-Subero, Iñigo Martincorena, Alexander Martinez-Fundichely, Yosef E. Maruvka, R. Jay Mashl, Charlie E. Massie, Thomas J. Matthew, Lucy Matthews, Erik Mayer, Simon Mayes, Michael Mayo, Faridah Mbabaali, Karen McCune, Ultan McDermott, Patrick D. McGillivray, Michael D. McLellan, John D. McPherson, John R. McPherson, Treasa A. McPherson, Samuel R. Meier, Alice Meng, Shaowu Meng, Andrew Menzies, Neil D. Merrett, Sue Merson, Matthew Meyerson, William Meyerson, Piotr A. Mieczkowski, George L. Mihaiescu, Sanja Mijalkovic, Tom Mikkelsen, Michele Milella, Linda Mileshkin, Christopher A. Miller, David K. Miller, Jessica K. Miller, Gordon B. Mills, Ana Milovanovic, Sarah Minner, Marco Miotto, Gisela Mir Arnau, Lisa Mirabello, Chris Mitchell, Thomas J. Mitchell, Satoru Miyano, Naoki Miyoshi, Shinichi Mizuno, Fruzsina Molnár-Gábor, Malcolm J. Moore, Richard A. Moore, Sandro Morganella, Quaid D. Morris, Carl Morrison, Lisle E. Mose, Catherine D. Moser, Ferran Muiños, Loris Mularoni, Andrew J. Mungall, Karen Mungall, Elizabeth A. Musgrove, Ville Mustonen, David Mutch, Francesc Muyas, Donna M. Muzny, Alfonso Muñoz, Jerome Myers, Ola Myklebost, Peter Möller, Genta Nagae, Adnan M. Nagrial, Hardeep K. Nahal-Bose, Hitoshi Nakagama, Hidewaki Nakagawa, Hiromi Nakamura, Toru Nakamura, Kaoru Nakano, Tannistha Nandi, Jyoti Nangalia, Mia Nastic, Arcadi Navarro, Fabio C. P. Navarro, David E. Neal, Gerd Nettekoven, Felicity Newell, Steven J. Newhouse, Yulia Newton, Alvin Wei Tian Ng, Anthony Ng, Jonathan Nicholson, David Nicol, Yongzhan Nie, G. Petur Nielsen, Morten Muhlig Nielsen, Serena Nik-Zainal, Michael S. Noble, Katia Nones, Paul A. Northcott, Faiyaz Notta, Brian D. O’Connor, Peter O’Donnell, Maria O’Donovan, Sarah O’Meara, Brian Patrick O’Neill, J. Robert O’Neill, David Ocana, Angelica Ochoa, Layla Oesper, Christopher Ogden, Hideki Ohdan, Kazuhiro Ohi, Lucila Ohno-Machado, Karin A. Oien, Akinyemi I. Ojesina, Hidenori Ojima, Takuji Okusaka, Larsson Omberg, Choon Kiat Ong, Stephan Ossowski, German Ott, B. F. Francis Ouellette, Christine P’ng, Marta Paczkowska, Salvatore Paiella, Chawalit Pairojkul, Marina Pajic, Qiang Pan-Hammarström, Elli Papaemmanuil, Irene Papatheodorou, Nagarajan Paramasivam, Ji Wan Park, Joong-Won Park, Keunchil Park, Kiejung Park, Peter J. Park, Joel S. Parker, Simon L. Parsons, Harvey Pass, Danielle Pasternack, Alessandro Pastore, Ann-Marie Patch, Iris Pauporté, Antonio Pea, John V. Pearson, Chandra Sekhar Pedamallu, Jakob Skou Pedersen, Paolo Pederzoli, Martin Peifer, Nathan A. Pennell, Charles M. Perou, Marc D. Perry, Gloria M. Petersen, Myron Peto, Nicholas Petrelli, Robert Petryszak, Stefan M. Pfister, Mark Phillips, Oriol Pich, Hilda A. Pickett, Todd D. Pihl, Nischalan Pillay, Sarah Pinder, Mark Pinese, Andreia V. Pinho, Esa Pitkänen, Xavier Pivot, Elena Piñeiro-Yáñez, Laura Planko, Christoph Plass, Paz Polak, Tirso Pons, Irinel Popescu, Olga Potapova, Aparna Prasad, Shaun R. Preston, Manuel Prinz, Antonia L. Pritchard, Stephenie D. Prokopec, Elena Provenzano, Xose S. Puente, Sonia Puig, Montserrat Puiggròs, Sergio Pulido-Tamayo, Gulietta M. Pupo, Colin A. Purdie, Michael C. Quinn, Raquel Rabionet, Janet S. Rader, Bernhard Radlwimmer, Petar Radovic, Benjamin Raeder, Keiran M. Raine, Manasa Ramakrishna, Kamna Ramakrishnan, Suresh Ramalingam, Benjamin J. Raphael, W. Kimryn Rathmell, Tobias Rausch, Guido Reifenberger, Jüri Reimand, Jorge Reis-Filho, Victor Reuter, Iker Reyes-Salazar, Matthew A. Reyna, Sheila M. Reynolds, Esther Rheinbay, Yasser Riazalhosseini, Andrea L. Richardson, Julia Richter, Matthew Ringel, Markus Ringnér, Yasushi Rino, Karsten Rippe, Jeffrey Roach, Lewis R. Roberts, Nicola D. Roberts, Steven A. Roberts, A. Gordon Robertson, Alan J. Robertson, Javier Bartolomé Rodriguez, Bernardo Rodriguez-Martin, F. Germán Rodríguez-González, Michael H. A. Roehrl, Marius Rohde, Hirofumi Rokutan, Gilles Romieu, Ilse Rooman, Tom Roques, Daniel Rosebrock, Mara Rosenberg, Philip C. Rosenstiel, Andreas Rosenwald, Edward W. Rowe, Romina Royo, Steven G. Rozen, Yulia Rubanova, Mark A. Rubin, Carlota Rubio-Perez, Vasilisa A. Rudneva, Borislav C. Rusev, Andrea Ruzzenente, Gunnar Rätsch, Radhakrishnan Sabarinathan, Veronica Y. Sabelnykova, Sara Sadeghi, S. Cenk Sahinalp, Natalie Saini, Mihoko Saito-Adachi, Gordon Saksena, Adriana Salcedo, Roberto Salgado, Leonidas Salichos, Richard Sallari, Charles Saller, Roberto Salvia, Michelle Sam, Jaswinder S. Samra, Francisco Sanchez-Vega, Chris Sander, Grant Sanders, Rajiv Sarin, Iman Sarrafi, Aya Sasaki-Oku, Torill Sauer, Guido Sauter, Robyn P. M. Saw, Maria Scardoni, Christopher J. Scarlett, Aldo Scarpa, Ghislaine Scelo, Dirk Schadendorf, Jacqueline E. Schein, Markus B. Schilhabel, Matthias Schlesner, Thorsten Schlomm, Heather K. Schmidt, Sarah-Jane Schramm, Stefan Schreiber, Nikolaus Schultz, Steven E. Schumacher, Roland F. Schwarz, Richard A. Scolyer, David Scott, Ralph Scully, Raja Seethala, Ayellet V. Segre, Iris Selander, Colin A. Semple, Yasin Senbabaoglu, Subhajit Sengupta, Elisabetta Sereni, Stefano Serra, Dennis C. Sgroi, Mark Shackleton, Nimish C. Shah, Sagedeh Shahabi, Catherine A. Shang, Ping Shang, Ofer Shapira, Troy Shelton, Ciyue Shen, Hui Shen, Rebecca Shepherd, Ruian Shi, Yan Shi, Yu-Jia Shiah, Tatsuhiro Shibata, Juliann Shih, Eigo Shimizu, Kiyo Shimizu, Seung Jun Shin, Yuichi Shiraishi, Tal Shmaya, Ilya Shmulevich, Solomon I. Shorser, Charles Short, Raunak Shrestha, Suyash S. Shringarpure, Craig Shriver, Shimin Shuai, Nikos Sidiropoulos, Reiner Siebert, Anieta M. Sieuwerts, Lina Sieverling, Sabina Signoretti, Katarzyna O. Sikora, Michele Simbolo, Ronald Simon, Janae V. Simons, Jared T. Simpson, Peter T. Simpson, Samuel Singer, Nasa Sinnott-Armstrong, Payal Sipahimalani, Tara J. Skelly, Marcel Smid, Jaclyn Smith, Karen Smith-McCune, Nicholas D. Socci, Heidi J. Sofia, Matthew G. Soloway, Lei Song, Anil K. Sood, Sharmila Sothi, Christos Sotiriou, Cameron M. Soulette, Paul N. Span, Paul T. Spellman, Nicola Sperandio, Andrew J. Spillane, Oliver Spiro, Jonathan Spring, Johan Staaf, Peter F. Stadler, Peter Staib, Stefan G. Stark, Lucy Stebbings, Ólafur Andri Stefánsson, Oliver Stegle, Lincoln D. Stein, Alasdair Stenhouse, Chip Stewart, Stephan Stilgenbauer, Miranda D. Stobbe, Michael R. Stratton, Jonathan R. Stretch, Adam J. Struck, Joshua M. Stuart, Henk G. Stunnenberg, Hong Su, Xiaoping Su, Ren X. Sun, Stephanie Sungalee, Hana Susak, Akihiro Suzuki, Fred Sweep, Monika Szczepanowski, Holger Sültmann, Takashi Yugawa, Angela Tam, David Tamborero, Benita Kiat Tee Tan, Donghui Tan, Patrick Tan, Hiroko Tanaka, Hirokazu Taniguchi, Tomas J. Tanskanen, Maxime Tarabichi, Roy Tarnuzzer, Patrick Tarpey, Morgan L. Taschuk, Kenji Tatsuno, Simon Tavaré, Darrin F. Taylor, Amaro Taylor-Weiner, Jon W. Teague, Bin Tean Teh, Varsha Tembe, Javier Temes, Kevin Thai, Sarah P. Thayer, Nina Thiessen, Gilles Thomas, Sarah Thomas, Alan Thompson, Alastair M. Thompson, John F. F. Thompson, R. Houston Thompson, Heather Thorne, Leigh B. Thorne, Adrian Thorogood, Grace Tiao, Nebojsa Tijanic, Lee E. Timms, Roberto Tirabosco, Marta Tojo, Stefania Tommasi, Christopher W. Toon, Umut H. Toprak, David Torrents, Giampaolo Tortora, Jörg Tost, Yasushi Totoki, David Townend, Nadia Traficante, Isabelle Treilleux, Jean-Rémi Trotta, Lorenz H. P. Trümper, Ming Tsao, Tatsuhiko Tsunoda, Jose M. C. Tubio, Olga Tucker, Richard Turkington, Daniel J. Turner, Andrew Tutt, Masaki Ueno, Naoto T. Ueno, Christopher Umbricht, Husen M. Umer, Timothy J. Underwood, Lara Urban, Tomoko Urushidate, Tetsuo Ushiku, Liis Uusküla-Reimand, Alfonso Valencia, David J. Van Den Berg, Steven Van Laere, Peter Van Loo, Erwin G. Van Meir, Gert G. Van den Eynden, Theodorus Van der Kwast, Naveen Vasudev, Miguel Vazquez, Ravikiran Vedururu, Umadevi Veluvolu, Shankar Vembu, Lieven P. C. Verbeke, Peter Vermeulen, Clare Verrill, Alain Viari, David Vicente, Caterina Vicentini, K. VijayRaghavan, Juris Viksna, Ricardo E. Vilain, Izar Villasante, Anne Vincent-Salomon, Tapio Visakorpi, Douglas Voet, Paresh Vyas, Ignacio Vázquez-García, Nick M. Waddell, Nicola Waddell, Claes Wadelius, Lina Wadi, Rabea Wagener, Jeremiah A. Wala, Jian Wang, Jiayin Wang, Linghua Wang, Qi Wang, Wenyi Wang, Yumeng Wang, Zhining Wang, Paul M. Waring, Hans-Jörg Warnatz, Jonathan Warrell, Anne Y. Warren, Sebastian M. Waszak, David C. Wedge, Dieter Weichenhan, Paul Weinberger, John N. Weinstein, Joachim Weischenfeldt, Daniel J. Weisenberger, Ian Welch, Michael C. Wendl, Johannes Werner, Justin P. Whalley, David A. Wheeler, Hayley C. Whitaker, Dennis Wigle, Matthew D. Wilkerson, Ashley Williams, James S. Wilmott, Gavin W. Wilson, Julie M. Wilson, Richard K. Wilson, Boris Winterhoff, Jeffrey A. Wintersinger, Maciej Wiznerowicz, Stephan Wolf, Bernice H. Wong, Tina Wong, Winghing Wong, Youngchoon Woo, Scott Wood, Bradly G. Wouters, Adam J. Wright, Derek W. Wright, Mark H. Wright, Chin-Lee Wu, Dai-Ying Wu, Guanming Wu, Jianmin Wu, Kui Wu, Yang Wu, Zhenggang Wu, Liu Xi, Tian Xia, Qian Xiang, Xiao Xiao, Rui Xing, Heng Xiong, Qinying Xu, Yanxun Xu, Hong Xue, Shinichi Yachida, Sergei Yakneen, Rui Yamaguchi, Takafumi N. Yamaguchi, Masakazu Yamamoto, Shogo Yamamoto, Hiroki Yamaue, Fan Yang, Huanming Yang, Jean Y. Yang, Liming Yang, Lixing Yang, Shanlin Yang, Tsun-Po Yang, Yang Yang, Xiaotong Yao, Marie-Laure Yaspo, Lucy Yates, Christina Yau, Chen Ye, Kai Ye, Venkata D. Yellapantula, Christopher J. Yoon, Sung-Soo Yoon, Fouad Yousif, Jun Yu, Kaixian Yu, Willie Yu, Yingyan Yu, Ke Yuan, Yuan Yuan, Denis Yuen, Christina K. Yung, Olga Zaikova, Jorge Zamora, Marc Zapatka, Jean C. Zenklusen, Thorsten Zenz, Nikolajs Zeps, Cheng-Zhong Zhang, Fan Zhang, Hailei Zhang, Hongwei Zhang, Hongxin Zhang, Jiashan Zhang, Jing Zhang, Junjun Zhang, Xiuqing Zhang, Xuanping Zhang, Yan Zhang, Zemin Zhang, Zhongming Zhao, Liangtao Zheng, Xiuqing Zheng, Wanding Zhou, Yong Zhou, Bin Zhu, Hongtu Zhu, Jingchun Zhu, Shida Zhu, Lihua Zou, Xueqing Zou, Anna deFazio, Nicholas van As, Carolien H. M. van Deurzen, Marc J. van de Vijver, L. van’t Veer, and Christian von Mering
Supplementary information
Supplementary information is available for this paper at 10.1038/s41467-019-14052-x.
References
- 1.Weber CE, Kuo PC. The tumor microenvironment. Surg. Oncol. 2012;21:172–177. doi: 10.1016/j.suronc.2011.09.001. [DOI] [PubMed] [Google Scholar]
- 2.Blagosklonny MV. Antiangiogenic therapy and tumor progression. Cancer Cell. 2004;5:13–17. doi: 10.1016/S1535-6108(03)00336-2. [DOI] [PubMed] [Google Scholar]
- 3.Vaupel P, Thews O, Hoeckel M. Treatment resistance of solid tumors. Med. Oncol. 2001;18:243–260. doi: 10.1385/MO:18:4:243. [DOI] [PubMed] [Google Scholar]
- 4.Dhani N, Fyles A, Hedley D, Milosevic M. The clinical significance of hypoxia in human cancers. Semin. Nucl. Med. 2015;45:110–121. doi: 10.1053/j.semnuclmed.2014.11.002. [DOI] [PubMed] [Google Scholar]
- 5.Brown JM, Wilson WR. Exploiting tumour hypoxia in cancer treatment. Nat. Rev. Cancer. 2004;4:437–447. doi: 10.1038/nrc1367. [DOI] [PubMed] [Google Scholar]
- 6.Zannella VE, et al. Reprogramming metabolism with metformin improves tumor oxygenation and radiotherapy response. Clin. Cancer Res. 2013;19:6741–6750. doi: 10.1158/1078-0432.CCR-13-1787. [DOI] [PubMed] [Google Scholar]
- 7.Mucaj V, Shay JES, Simon MC. Effects of hypoxia and HIFs on cancer metabolism. Int. J. Hematol. 2012;95:464–470. doi: 10.1007/s12185-012-1070-5. [DOI] [PubMed] [Google Scholar]
- 8.Rouschop KMA, et al. The unfolded protein response protects human tumor cells during hypoxia through regulation of the autophagy genes MAP1LC3B and ATG5. J. Clin. Invest. 2010;120:127–141. doi: 10.1172/JCI40027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lassen P, et al. HPV-associated p16-expression and response to hypoxic modification of radiotherapy in head and neck cancer. Radiother. Oncol. 2010;94:30–35. doi: 10.1016/j.radonc.2009.10.008. [DOI] [PubMed] [Google Scholar]
- 10.Hoskin PJ, Rojas AM, Bentzen SM, Saunders MI. Radiotherapy with concurrent carbogen and nicotinamide in bladder carcinoma. J. Clin. Oncol. 2010;28:4912–4918. doi: 10.1200/JCO.2010.28.4950. [DOI] [PubMed] [Google Scholar]
- 11.Milosevic M, et al. Tumor hypoxia predicts biochemical failure following radiotherapy for clinically localized prostate cancer. Clin. Cancer Res. 2012;18:2108–2114. doi: 10.1158/1078-0432.CCR-11-2711. [DOI] [PubMed] [Google Scholar]
- 12.Kim B, et al. Prognostic assessment of hypoxia and metabolic markers in cervical cancer using automated digital image analysis of immunohistochemistry. J. Transl. Med. 2013;11:185. doi: 10.1186/1479-5876-11-185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Yang L, et al. Development and validation of a 28-gene hypoxia-related prognostic signature for localized prostate cancer. EBioMedicine. 2018;31:182–189. doi: 10.1016/j.ebiom.2018.04.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lalonde E, et al. Tumour genomic and microenvironmental heterogeneity for integrated prediction of 5-year biochemical recurrence of prostate cancer: a retrospective cohort study. Lancet Oncol. 2014;15:1521–1532. doi: 10.1016/S1470-2045(14)71021-6. [DOI] [PubMed] [Google Scholar]
- 15.Lalonde E, et al. Translating a prognostic DNA Genomic Classifier into the clinic: retrospective validation in 563 localized prostate tumors. Eur. Urol. 2017;72:22–31. doi: 10.1016/j.eururo.2016.10.013. [DOI] [PubMed] [Google Scholar]
- 16.Bhandari V, et al. Molecular landmarks of tumor hypoxia across cancer types. Nat. Genet. 2019;51:308–318. doi: 10.1038/s41588-018-0318-2. [DOI] [PubMed] [Google Scholar]
- 17.Graeber TG, et al. Hypoxia-mediated selection of cells with diminished apoptotic potential in solid tumours. Nature. 1996;379:88–91. doi: 10.1038/379088a0. [DOI] [PubMed] [Google Scholar]
- 18.Greijer AE, van der Wall E. The role of hypoxia inducible factor 1 (HIF-1) in hypoxia induced apoptosis. J. Clin. Pathol. 2004;57:1009–1014. doi: 10.1136/jcp.2003.015032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Bindra RS, et al. Down-regulation of Rad51 and decreased homologous recombination in hypoxic cancer cells. Mol. Cell. Biol. 2004;24:8504–8518. doi: 10.1128/MCB.24.19.8504-8518.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bristow RG, Hill RP. Hypoxia and metabolism: hypoxia, DNA repair and genetic instability. Nat. Rev. Cancer. 2008;8:180–192. doi: 10.1038/nrc2344. [DOI] [PubMed] [Google Scholar]
- 21.Meng AX, et al. Hypoxia down-regulates DNA double strand break repair gene expression in prostate cancer cells. Radiother. Oncol. 2005;76:168–176. doi: 10.1016/j.radonc.2005.06.025. [DOI] [PubMed] [Google Scholar]
- 22.Mihaylova VT, et al. Decreased expression of the DNA mismatch repair gene Mlh1 under hypoxic stress in mammalian cells. Mol. Cell. Biol. 2003;23:3265–3273. doi: 10.1128/MCB.23.9.3265-3273.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Koshiji M, et al. HIF-1α induces genetic instability by transcriptionally downregulating MutSα expression. Mol. Cell. 2005;17:793–803. doi: 10.1016/j.molcel.2005.02.015. [DOI] [PubMed] [Google Scholar]
- 24.Chua MLK, et al. A prostate cancer “ Nimbosus ”: genomic instability and SChLAP1 dysregulation underpin aggression of intraductal and cribriform subpathologies. Eur. Urol. 2017;71:183–192. doi: 10.1016/j.urology.2017.04.035. [DOI] [PubMed] [Google Scholar]
- 25.The ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature10.1038/s41586-020-1969-6 (2020).
- 26.PCAWG Transcriptome Core Group. et al. Genomic basis for RNA alterations in cancer. Nature10.1038/s41586-020-1970-0 (2020). [DOI] [PMC free article] [PubMed]
- 27.Li CH, Haider S, Shiah Y-J, Thai K, Boutros PC. Sex differences in cancer driver genes and biomarkers. Cancer Res. 2018;78:5527–5537. doi: 10.1158/0008-5472.CAN-18-0362. [DOI] [PubMed] [Google Scholar]
- 28.Li, C. H. et al. Sex differences in oncogenic mutational processes. Nat. Commun. In Press (2020). [DOI] [PMC free article] [PubMed]
- 29.Nakagawa S, Schielzeth H. A general and simple method for obtaining R2 from generalized linear mixed-effects models. Methods Ecol. Evol. 2013;4:133–142. doi: 10.1111/j.2041-210x.2012.00261.x. [DOI] [Google Scholar]
- 30.Buffa FM, Harris AL, West CM, Miller CJ. Large meta-analysis of multiple cancers reveals a common, compact and highly prognostic hypoxia metagene. Br. J. Cancer. 2010;102:428–435. doi: 10.1038/sj.bjc.6605450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Winter SC, et al. Relation of a hypoxia metagene derived from head and neck cancer to prognosis of multiple cancers. Cancer Res. 2007;67:3441–3449. doi: 10.1158/0008-5472.CAN-06-3322. [DOI] [PubMed] [Google Scholar]
- 32.Ragnum HB, et al. The tumour hypoxia marker pimonidazole reflects a transcriptional programme associated with aggressive prostate cancer. Br. J. Cancer. 2015;112:382–390. doi: 10.1038/bjc.2014.604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Yousif, F. et al. The origins and consequences of localized and global somatic hypermutation. Preprint at https://www.biorxiv.org/content/10.1101/287839v2 (2018).
- 34.Yuan, Y. et al. Comprehensive molecular characterization of mitochondrial genomes in human cancers. Nat. Genet. 10.1038/s41588-019-0557-x (2020). [DOI] [PMC free article] [PubMed]
- 35.Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature10.1038/s41586-020-1943-3 (2020). [DOI] [PMC free article] [PubMed]
- 36.Deshwar AG, et al. PhyloWGS: reconstructing subclonal composition and evolution from whole-genome sequencing of tumors. Genome Biol. 2015;16:35. doi: 10.1186/s13059-015-0602-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Espiritu SMG, et al. The evolutionary landscape of localized prostate cancers drives clinical aggression. Cell. 2018;173:1003–1013. doi: 10.1016/j.cell.2018.03.029. [DOI] [PubMed] [Google Scholar]
- 38.Gerstung, M. et al. The evolutionary history of 2,658 cancers. Nature10.1038/s41586-019-1907-7 (2020). [DOI] [PMC free article] [PubMed]
- 39.Chan N, et al. Contextual synthetic lethality of cancer cell kill based on the tumor microenvironment. Cancer Res. 2010;70:8045–8054. doi: 10.1158/0008-5472.CAN-10-2352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Haider S, et al. Genomic alterations underlie a pan-cancer metabolic shift associated with tumour hypoxia. Genome Biol. 2016;17:140. doi: 10.1186/s13059-016-0999-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Nguyen AD, McDonald JG, Bruick RK, DeBose-Boyd RA. Hypoxia stimulates degradation of 3-hydroxy-3-methylglutaryl-coenzyme A reductase through accumulation of lanosterol and hypoxia-inducible factor-mediated induction of insigs. J. Biol. Chem. 2007;282:27436–27446. doi: 10.1074/jbc.M704976200. [DOI] [PubMed] [Google Scholar]
- 42.Luoto KR, Kumareswaran R, Bristow RG. Tumor hypoxia as a driving force in genetic instability. Genome Integr. 2013;4:5. doi: 10.1186/2041-9414-4-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Chen Y, Cairns R, Papandreou I, Koong A, Denko NC. Oxygen consumption can regulate the growth of tumors, a new perspective on the Warburg effect. PLoS One. 2009;4:e7033. doi: 10.1371/journal.pone.0007033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Ye Y, et al. Characterization of hypoxia-associated molecular features to aid hypoxia-targeted therapy. Nat. Metab. 2019;1:431–444. doi: 10.1038/s42255-019-0045-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Rosenberger C, Rosen S, Paliege A, Heyman SN. Pimonidazole adduct immunohistochemistry in the rat kidney: detection of tissue hypoxia. Methods Mol. Biol. 2009;466:161–174. doi: 10.1007/978-1-59745-352-3_12. [DOI] [PubMed] [Google Scholar]
- 46.Dunn PK, Smyth GK. Randomized quantile residuals. J. Comput. Graph. Stat. 1996;5:236. [Google Scholar]
- 47.P’ng C, et al. BPG: seamless, automated and interactive visualization of scientific data. BMC Bioinforma. 2019;20:42. doi: 10.1186/s12859-019-2610-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Description of Additional Supplementary Files
Data Availability Statement
Aligned sequencing data, somatic and germline variant calls, single nucleotide variants, indels, copy-number alterations, structural variants, mutational signatures, subclonal reconstructions, transcript abundances and other core data generated by the PCAWG Consortium are described here25 and available for download at dcc.icgc.org/releases/PCAWG. Additional information on accessing the data, including raw read files, can be found at docs.icgc.org/pcawg/data. In accordance with the data access policies of the ICGC and TCGA projects, most molecular, clinical and specimen data are in an open tier which does not require approval for access.
To access potentially identifiable information, such as germline alleles and underlying sequencing data, researchers will need to apply to the TCGA Data Access Committee (DAC) via dbGaP (dbgap.ncbi.nlm.nih.gov/aa/wga.cgi?page = login) for access to the TCGA portion of the dataset, and to the ICGC Data Access Compliance Office (DACO; icgc.org/daco) for the ICGC portion of the dataset. To access somatic single nucleotide variant data derived from TCGA donors, researchers will also need to obtain dbGaP authorization.
The analyses in this paper used several PCAWG datasets that were derived from the raw sequencing data and variant calls. The individual datasets are available at Synapse (synapse.org) and are denoted with accession numbers starting with syn. All of these datasets are also mirrored at dcc.icgc.org. The datasets encompass: clinical data for each patient, including information about age and sex (syn10389158, dcc.icgc.org/releases/PCAWG/clinical_and_histology); histopathology data for each tumour (syn10389164, dcc.icgc.org/releases/PCAWG/clinical_and_histology); mRNA abundance data for each tumour (syn5553991, dcc.icgc.org/releases/PCAWG/transcriptome/gene_expression); driver mutations for each tumour spanning all classes of variants (syn11639581, dcc.icgc.org/releases/PCAWG/driver_mutations); single base substitution and small indel signature exposures for each tumour (syn8366024, dcc.icgc.org/releases/PCAWG/mutational_signatures); subclonal architecture, tumour evolution and tumour purity data for each tumour (syn8272483, dcc.icgc.org/releases/PCAWG/subclonal_reconstruction).
The core computational pipelines used by the PCAWG Consortium for alignment, quality control and variant calling are available to the public at dockstore.org/search?search = pcawg under the GNU General Public License v3.0, which allows for reuse and distribution. Code specific to this manuscript is available upon request.