Abstract
Pathogen-derived lipopolysaccharide (LPS) and cytokine tumor necrosis factor (TNF) activate NFκB with distinct duration dynamics, but how immune response genes decode NFκB duration to produce stimulus-specific expression remains unclear. Here, detailed transcriptomic profiling of combinatorial and temporal control mutants identified 81 genes that depend on stimulus-specific NFκB duration for their stimulus-specificity. Combining quantitative experimentation with mathematical modeling, we found that for some genes a long mRNA half-life allowed effective decoding but for many genes this was insufficient to account for the data; instead, we found that chromatin mechanisms, such as a slow transition rate between inactive and RelA-bound enhancer states, could also decode NFκB dynamics. Chromatin-mediated decoding is favored by genes acting as immune effectors (e.g. tissue remodelers, T-cell recruiters) rather than immune regulators (e.g. signaling proteins, monocyte recruiter). Overall, our results delineate two gene regulatory strategies that decode stimulus-specific NFκB dynamics and determine distinct biological functions.
Graphical Abstract

eTOC Blurb
Sen et al. used combinatorial and temporal coding mutants of NFκB to identify stimulus-specific genes that are dependent on the duration of dynamic NFκB activity. Iterating between quantitative experimentation and mathematical modeling revealed that a long mRNA half-life and a slow chromatin-control step are involved in decoding stimulus-specific NFκB duration for genes with distinct biological functions.
Introduction
The innate immune response is the first line of defense against pathogens. Exposure to a pathogen leads to dramatic changes in the transcriptome of myeloid and fibroblastoid cells (Nau et al., 2002; Novershtern et al., 2011; Ramirez-Carrozzi et al., 2009; Ramsey et al., 2008; Ravasi et al., 2010). These cells coordinate multi-tiered immune responses by inducing cell-intrinsic defenses, recruitment of professional innate immune cells (e.g., neutrophils and macrophages), the initiation of an adaptive immune response (via dendritic cells and T cells), and tissue remodeling for pathogen clearance and subsequent wound healing. Numerous molecular factors have been identified that regulate these pathogen-responsive gene expression programs through complex signal regulatory networks (Amit et al., 2009; Cheng et al., 2011; Gilchrist et al., 2006; Nau et al., 2002; Novershtern et al., 2011; Ramsey et al., 2008; Ravasi et al., 2010).
Previous studies (Cheng et al., 2017; Tong et al., 2016) delineated the pathogen-responsive transcriptome in terms of the combinatorial control of three major transcription factors (TFs), activating protein 1 (AP1), nuclear factor κB (NFκB), and interferon regulatory factors (IRF/ISGF3). Specifically, examining the transcriptomic data quantitatively provided evidence for the p38 pathway functioning combinatorially with NFκB to potentiate LPS-specific expression (Cheng et al. 2017). Further, focusing on highly induced genes and analyzing transcriptomic data at single gene resolution helped to identify five genes that engage the combinatorial functions of IRF3 and NFκB (Tong et al., 2016).
These studies also suggested that combinatorial TF control is not the only regulatory strategy that cells can employ to achieve stimulus-specific gene expression. Several signaling pathways have been found to produce stimulus-specific TF temporal patterns (a.k.a. “dynamics”), leading to the hypothesis that a temporal code specifies stimulus-specific gene expression (Behar and Hoffmann, 2010; Hoffmann and Baltimore, 2006; Purvis and Lahav, 2013; Hoffmann 2016). Indeed, a number of studies have presented evidence that stimulus-specific gene expression depends on the dynamics of transcription factor activity (Batchelor et al., 2014; Hao and O’Shea, 2012; Hoffmann et al., 2002; Purvis and Lahav, 2013; Werner et al., 2005). However, the number of genes controlled by TF dynamics, and the molecular mechanisms they employ to decode such dynamics to achieve stimulus-specific expression has remained unclear.
One confounding aspect of TF dynamics is that they are complex and differ in multiple aspects, for example, in speed of activation, amplitude, oscillatory components, and duration. Thus, focusing on a specific dynamic feature may be key to gain a mechanistic understanding of the gene regulatory mechanisms that decode even complex TF dynamics. Prior work focusing on peak amplitude or fold change of NFκB activation identified an incoherent feedforward loop as a decoding mechanism (Lee et al 2014). In this study, we focused on identifying decoding mechanisms for stimulus-specific duration of NFκB activity.
Based on the gene expression programs induced by a single stimulus, recent studies in other biological systems, showed that mRNA half-life may play a role in decoding the duration of p53 dynamics (Hafner et al., 2017; Porter et al., 2016) and ERK signaling (Uhlitz et al., 2017). This relates to the observation that the temporal ordering of gene expression is governed by mRNA half-life, for example, during the response to H2O2-induced stress in yeast (Shalem et al., 2008), upon IL-2 signaling in murine T cells (Elkon et al., 2010), or upon TNF-induced NFκB signaling in mouse fibroblasts (Hao and Baltimore, 2009). However, it is currently not understood whether stimulus-specific expression of immune response genes is mediated by differential mRNA half-life control, and which genes actually employ this gene regulatory strategy. Indeed, whether this mechanism is sufficient to account for observed stimulus-specific gene expression, or whether other mechanisms of dynamics decoding might also contribute, is an open and important question.
These considerations suggest that dissecting the regulatory strategies of immune response genes requires not only detailed quantitative experimentation, but also quantitative interpretation of the data. That in turn requires a mathematical modeling framework that recapitulates the dynamic processes underlying gene expression. The systems biology approach of combining experimental studies with computational models to achieve a quantitative understanding has been particularly useful in the study of NFκB signaling, uncovering numerous mechanisms that encode NFκB dynamics (Basak et al 2012). A key utility of mathematical modeling has been to determine the sufficiency of a mechanistic explanation to account for the observed phenomena or measurements. However, quantitatively applying mathematical modeling to mammalian gene expression data has proven more challenging. In one study, mathematical models explored the combinatorial control of signaling pathways for LPS-responsive gene clusters (Cheng et al., 2017). In another study, mathematical models were fit to gene expression data produced by a synthetic experimental system for ERK signaling in PC12 cells and MCF7 cells that avoided the combinatorial complexities of endogenous signaling (Uhlitz et al., 2017). However, quantitative modeling at single gene resolution to understand the mechanisms of stimulus-specific gene expression in primary cells has not been reported.
Here, we addressed how TNF- vs. LPS-specific NFκB activation dynamics are decoded by target genes to produce stimulus-specific gene expression responses. We developed a workflow of iterative mathematical modeling and experimentation to evaluate the sufficiency of alternate gene regulatory strategies. We found that a mRNA half-life of just >30 min could be an effective decoding mechanism, but for the majority of genes, stimulus-specific transcription initiation mediated by chromatin-associated control mechanisms contributed substantially to decoding stimulus-specific NFκB dynamics. Using a two-step mathematical model for gene activation could recapitulate the mRNA dynamics of a majority genes and quantify chromatin-associated transcriptional and cytoplasmic post-transcriptional mechanisms to achieve stimulus-specific immune responses.
Results
The duration of NFκB activity controls LPS-specific expression of innate immune genes
The early LPS-responsive transcriptome is largely determined by the activation of the transcription factors NFκB, AP1 and IRF (Fig. 1A, (Cheng et al., 2017)), while the cytokines TNF and IL1 activate only NFκB and AP1. We asked whether LPS-specific (compared to TNF and IL1) gene expression can occur in the absence of the primary IRF family member ISGF3. In Ifnar−/− mice and cells, ISGF3 activation is abrogated, thereby diminishing the encoding of stimulus-specific combinations of transcription factors. However, stimulus-specific dynamics of NFκB activity are maintained in Ifnar−/− murine embryo fibroblasts (MEFs) (Fig. 1B). Thus using Ifnar−/− MEFs allow us to identify genes that may be stimulus-specifically expressed, not by the combinatorial TF code (Cheng et al., 2017), but a temporal TF code (Hoffmann 2016, Behar and Hoffmann, 2010).
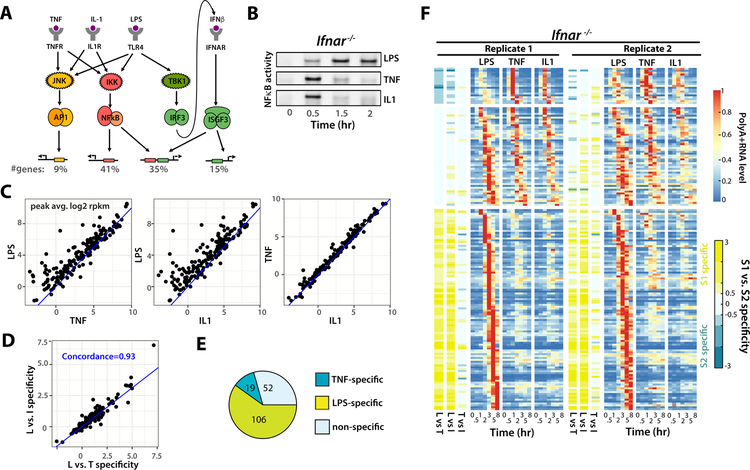
Figure 1. Identifying LPS-specific genes that do not depend on interferon signaling.
(A) Schematic of the signaling pathways for TNF, LPS and IL1 that activate downstream gene expression through transcriptional factors AP1, NFκB and IRF3/ISGF3. The percentage of LPS-induced genes that are targets of each transcription factor, as identified in Cheng et al (Cheng et al., 2017), is indicated. (B) EMSA showing NFκB activity in Ifnar−/− MEFs following treatment with LPS (100 ng/ml), TNF (1 ng/ml), or IL1 (1 ng/ml). (C) Scatter plots of maximum gene expression between LPS and TNF, LPS and IL1, TNF and IL1. (D) Scatter plot showing that LPS vs TNF specificity correlates well to LPS vs IL1 specificity. The pair-wise specificity (L vs. T or L vs. I) for each gene is defined by log2 of the fold ratio between maximum expression in LPS treatment vs. TNF or IL1 treatment respectively. (E) Pie chart showing TNF-specific, LPS-specific or non-specific genes, using a L vs. T specificity threshold value of ≥ 0.5 or ≤−0.5. (F) Heat-map of polyA+ RNA expression profiles (normalized to max) of the 177 induced genes in Ifnar−/− MEFs by LPS (100 ng/ml), TNF (1 ng/ml) or IL1 (1 ng/ml) at each of the indicated time-points. The three groups of genes defined in (E) are shown in distinct expression clusters. Specificity metrics (Lvs.T, Lvs.I, T vs.I) are shown on left. Results from two independent biological replicates are shown (see also Table S1).
Genome-wide expression measurements by mRNA-seq in replicate at six time-points (0.5, 1, 2, 3, 5, 8 hr) in response to pulse stimulation with a 80% saturating dose of LPS, TNF or IL1 identified 177 differentially induced genes with fold change of > 4-fold and False Discovery Rate (FDR) <0.01 at any timepoint. Binary comparisons of the maximum expression quantified by Reads Per Kilobase per Million (RPKM) revealed that many genes showed higher expression in response to LPS than TNF or IL1, but that IL1 and TNF produced roughly equivalent levels of expression for any gene (Fig. 1C, Table S1). Pairwise stimulus-specificity for each gene was determined by calculating the log2 ratio of maximum gene expression (RPKM) in any two stimulation conditions. Using this specificity metric, we found that LPS vs. IL1 specificity strongly correlated with LPS vs. TNF specificity (Fig. 1D, Table S1), allowing us to focus on the latter as being representative for pathogen vs cytokine-responsive gene expression.
To further categorize the 177 inducible genes, we used a threshold value of 0.5 (i.e. 20.5=1.4 fold difference) for specificity; by this measure, 106 genes were LPS-specific (LPS vs. TNF specificity > 0.5), 19 were TNF-specific (LPS vs. TNF specificity < −0.5), and the remaining 52 genes fell below the stimulus-specificity threshold (Fig. 1E, Table S1). Expression profiles from two biological replicates showed excellent reproducibility in mRNA abundance measurements (Fig. S1A, Table S1), in the evolution of their time-courses, and in specificity metrics (Fig. 1F, Table S1). Our results demonstrate that even in the absence of combinatorial TF coding by the IFNAR-ISGF3 axis, there is a high degree of LPS-specific gene expression. Two mechanisms may underlie this phenomenon: either combinatorial coding via another LPS-specific signaling pathway, such as for example the TBK1-IRF3, MAPKp38-TTP pathways (Cheng et al., 2017), or temporal coding by stimulus-specific dynamic control of NFκB (Werner et al., 2005).
To address the role of stimulus-specific NFκB dynamics in mediating stimulus-specific gene expression programs, we quantitated time-course NFκB activity in these TNF and LPS stimulation conditions, and found LPS-specific long-lasting activity (Fig. 2A). To diminish the stimulus-specificity in NFκB dynamics, we generated MEFs that are deficient in the key NFκB negative feedback regulator IκBα (encoded by Nfkbia) in the genetic background (Ifnar−/−) of combinatorial control deficiency (hereafter referred to as “control”, as temporal coding is tested), resulting in a temporal control “mutant” (Ifnar−/−Nfkbia−/−). We found that in the mutant MEFs, NFκB activity was extended in response to TNF, and largely unaltered in response to LPS. Particularly, at 2 hrs, nuclear NFκB activity was still present in mutant cells stimulated with TNF whereas in control cells, it had returned to basal levels (Fig. 2A & Fig. S2A). Thus, IκBα deficiency extended TNF-induced NFκB activity by about 1 hr causing the duration of TNF-induced and LPS-induced NFκB activity to be similar.
Figure 2. Determining the NFκB duration dependency of LPS-specific gene expression.
(A) Dynamics of nuclear NFκB activity measured by EMSA in control (Ifnar−/−) and mutant (Ifnar−/−Nfκbia−/−) in response to 30 min pulse of 100 ng/ml LPS or 1ng/ml TNF. Error bars are standard deviations (SD) from 3–5 independent experiments. p value was calculated by multiple t-test using Graph-pad Prizm. P value >0.05 considered significant (B) Pie charts showing TNF-specific, LPS-specific, or nonspecific genes using the specificity threshold of 0.5 in control vs mutant (same definition as in Figure 1D). (C) 177 differentially induced genes are categorized into 3 groups based on the loss of specificity in the mutant. Group I. not LPS-specific: if Lvs.T in control as well as mutant is less than 0.5; group II. NFκB dynamics independent and LPS-specific: if Lvs.T in control is higher than 0.5 and L vs. T in mutant is not reduced by 2^0.5 fold (i.e. above line y=0.71x); group III. NFκB dynamics dependent and LPS-specific: if Lvs.T in control is higher than 0.5 and Lvs.T in mutant is reduced by 2^0.5 fold (i.e. below line y=0.71x). The expression trajectories for example genes in each category are shown on the right. (D) Heat-map of polyA+ RNA-seq gene expression profiles (normalized to max) for control and mutant cells for indicated time-points (See also Table S2). (E) Known TF binding motifs (κB, AP1 and IRE) and (F) Gene Ontology (GO) terms enrichment level is shown as the p-value in -log10, using genes in each category and remaining genes as background. If p-value >0.05, it is shown as ‘–‘.)
To test how diminished stimulus-specificity in NFκB duration affects the stimulus-specificity of gene expression, we compared the transcriptomes of mutant and control cells stimulated with LPS or TNF (Fig. S1B, Table S2). Of the 177 inducible genes, 104 genes were found to fall below the threshold of stimulus-specificity in the mutant (compared to 52 in control), 55 genes were still LPS-specific (compared to 106 in control) and 18 genes (compared to 19) were found to be TNF-specific (Fig. 2B, Table S2). Thus, by reducing the difference in the duration of NFκB activity in the LPS and TNF stimulation scenarios, we diminished the specificity of gene expression responses.
In order to determine in a more quantitative and gene-specific manner which genes showed diminished stimulus-specificity in the mutant, we plotted the specificity of LPS vs. TNF for each gene for the mutant (y-axis) against that for the control (x-axis) as a scatterplot (Fig. 2C, left panel). One example of a stimulus-non-specific gene (group I) is Fos, whose expression was equivalent for LPS and TNF in both control and mutant cells (Fig. 2C, right panel), which is consistent with its transcriptional control being regulated by the MAPK/JNK axis rather than by NFκB. Focusing on the 106 genes that were categorized as LPS-specific in control cells, we found that many genes showed diminished specificity in the mutant, but others were unaffected. We categorized 18 genes as LPS-specific but independent of NFκB duration (group II) as their specificity score was maintained in the mutant; a representative gene is Il1a (Fig. 2C, right panel) as its substantial LPS specificity was maintained in the mutant, and whose expression is known to be subject to combinatorial control by the MAPK p38 axis (Cheng et al., 2017). However, the majority of genes (88 genes), LPS-specific expression was either fully or partially dependent on the duration of NFκB dynamics (group III), as their specificity score was reduced by at least 1/3 (dashed blue line in Fig. 2C, left panel). Ccl5 serves as an example of this category as its expression is highly LPS-specific in the control but this specificity was almost entirely lost in the mutant (Fig. 2C, right panel). In fact, visual inspection of line graphs (Fig. S2B) proved useful as a quality control measure for our analysis. We identified 7 genes as being incorrectly categorized in group III due to elevated basal levels in the mutant (Fig. S2B). The recategorization thus resulted in a total of 25 genes in group II, and 81 genes in group III. In subsequent analyses we focused on the 81 genes whose LPS-specific expression was at least partially dependent on the duration of NFκB dynamics.
Based on these groupings, we plotted the time-course RNA-seq data (normalized to max) from control and mutant cells as a heat-map, ordering genes within each group by the time of peak expression (Fig. 2, Table S2). As described, focusing on the 106 genes whose expression was LPS-specific in control cells, group II contained 25 genes whose LPS-specific expression in control was largely maintained in the mutant, and group III contained 81 genes whose LPS-specific expression in control showed substantially diminished specificity in the mutant (Table S2). We examined the relative enrichment for the motifs (κB, AP1 and IRE) of relevant transcription factors in the promoter proximal regions (−500bp to +200bp) of genes in each group against the remaining LPS-induced genes (Fig. 2E). These results revealed that whereas non-specific group I genes had a preponderance of both κB and AP1 motifs, and LPS-specific group II genes had both κB and IRE motifs, LPS-specific, NFκB-dynamics-dependent group III genes showed only a preponderance of κB motifs. Further, gene ontology (GO) enrichment analysis revealed that non-specific group I genes code for a variety of transcriptional regulators, and group II genes showed an enrichment for regulators of interferon signaling, while group III genes were enriched in the GO term “innate immune response”, suggesting that NFκB temporal coding is functionally important for mounting an effective immune response (Fig. 2F).
Long-mRNA half-lives are correlated with LPS specificity
We next set out to characterize the gene regulatory strategies that allow group III LPS-specific genes to decode the duration of stimulus-specific NFκB dynamics.
Mathematical modelling of RNA synthesis and degradation suggests (Yang et al., 2003) and experimental data confirm (Cheng et al., 2017; Elkon et al., 2010; Hao and Baltimore, 2009; Nagashima et al., 2015; Porter et al., 2016; Shalem et al., 2008) that short-lived transcripts achieve their half-maximal induction more rapidly than long-lived transcripts. This suggests that long-lived transcripts may only be fully induced by persistent TF activities.
To test this hypothesis, we developed a simple kinetic model (model v1) of TF-dependent mRNA production (Bintu et al., 2005) (Fig. 3A), and we asked how NFκB activities in response to LPS or TNF may affect the abundances of mRNAs of different half-lives. We interpolated the quantitated NFκB activities measured by EMSA for both control and mutant cells (Fig. 2A) to use as input to the ODE model (Fig. 3A). The mRNA abundances (color scale, z-axis) were then simulated over time (x-axis) in control (Fig. 3A) as a function of mRNA half-lives ranging from 1 min to 1000 mins (y-axis). The log2 peak expression ratio (P.R., representing LPS vs. TNF specificity for mRNA) was calculated and indicated in a yellow color bar on the right side of the heat-map. As expected, we found that LPS-specific expression is more pronounced in mRNAs with long half-lives. These model simulations suggest that for mRNAs whose half-lives are >30 mins, TNF-induced NFκB activity is too transient to produce target gene mRNA of half the maximum response amplitude. The simulated mRNA trajectories for 5 min, 1 hr, and 8 hr half-lifes also documented this phenomenon (shown on right of panel 3A).
Figure 3. Long-mRNA half-lives are correlated with LPS specificity.
(A-B) Heat-map of simulated gene expression profiles with different mRNA half-lives (from 1 min to 1000 mins) using a simple ordinary differential equation (ODE) model. NFκB activity profiles in response to LPS (left) or TNF (right) are used as input, in control (A) or in mutant (B). ODE is shown in the top panel: mRNA abundance is determined by NFκB-dependent synthesis using a Hill-equation (with a basal synthesis) and a first-order degradation term. The yellow color-bar on the right sight of the heat-map shows the peak expression ratio (P.R.) between LPS and TNF input given the same mRNA half-life. Example trajectories of different half-lives are shown on the right for LPS input (red line) and TNF input (green). Grey dashed lines indicate a half-life of 30 mins and 100 mins. (C) Line-graph showing that the predicted Lvs.T specificity from simulation results in (A) and (B) is correlated with mRNA half-life. (D) Gene expression profiles (log2 ratio with respect to unstimulated timepoint) for the 177 inducible genes after Actinomycin D treatment (ActD-seq) in control cells at indicated time points for two independent biological replicates are shown. The right-side annotation color-bars indicate the derived mRNA half-life using an adapted linear regression method on log2 expression of actinomycin-D time-course data (See also Table S3). (E) Four example genes are shown on the right. Dots are the values from ActD-seq data with two different colors to indicate replicates. The dots with an open circle are the data points selected for linear regression. If the initial reads are not enough (less than 32), the half-life is not determined (e.g. Mmp1b). (F) Scatter of derived half-life between the two replicates with density distribution shown on the top and right. (G) Box-plots showing that the measured Lvs.T specificity from the RNA-seq in Fig. 2D is correlated with the derived mRNA half-life from Act-D-seq.
We next asked whether the model would predict that mutant cells produce diminished stimulus-specific gene expression. Using the measured NFκB time-courses associated with mutant cells for analogous model simulations (Fig. 3B), we found that the specificity metric associated with mRNAs with half-lives between 30 and 100 minutes were substantially diminished, though mRNAs with half-lives of >100 mins largely retained it. Taken together, these simulation results suggested that mRNAs with half-lives of greater than 30 mins might decode the duration of stimulus-specific NFκB dynamics and that the IκBα mutant may render mRNA of 30 to 100 min half-lives incapable of stimulus-specific gene expression. These findings are summarized in a graph of Lvs.T specificity of NFκB target genes as a function of their mRNA half-life (Fig. 3C): whereas in control cells, NFκB target genes with > 30 min half-life may show LPS-specificity (greater than 0.5 threshold), in mutant cells, LPS specific genes must have a half-life of > 100 min.
To address this model prediction experimentally, we measured mRNA half-lives using actinomycin D (actD)-mediated transcriptional inhibition in two biological replicates (Fig. 3D, Table S3). Though this approach is used for mRNA half-life determinations, it is not always reliable (Lugowski et al., 2018). To focus on likely reliable data, we developed an algorithm and software (Material & Methods, Table S3) for deriving half-life estimates by linear regression in log2 scale only when specific conditions were met. We show four example genes, for three of which the data allowed for half-life derivations (Fig. 3E). Overall, for the genes for which we are able to derive half-lives from both datasets, the values were reasonably reproducible (Fig. 3F) and matched with the values that were reported by a previous study (Hao and Baltimore, 2009).
Using these measured mRNA half-lives, we plotted the measured Lvs.T specificity of control and mutant against the median measured mRNA half-life values, binned into five ranges as indicated (Fig. 3G). We found that Lvs.T specificity indeed correlated with mRNA half-life up to about a 5 hr of half-life. Furthermore, we found that the genes that had lost their Lvs.T specificity in the mutant were in the (30–60) and (60–180) mins bins, though very long-lived mRNAs retained some Lvs.T specificity. These results showed a remarkable resemblance to the model simulations (Fig. 3C). In sum, these results suggest that an intermediate mRNA half-life of 30–180 minutes is correlated with LPS-specific NFκB target gene expression in control cells, but not in IκBα-mutant cells, which show diminished stimulus-specific NFκB dynamics.
A long mRNA half-life may be sufficient for decoding stimulus-specific NFκB duration
To extend the correlative finding, we aimed to test whether the LPS specificity of NFκB dynamics-dependent genes can indeed be accounted by their mRNA-half-lives. To this end we embedded the mathematical model v1 used for hypothesis generation (Fig. 3A) into a gene-by-gene parameterization workflow (Fig. 4A). This allowed us to determine whether for a given gene a set of value for the model parameters (k0, kf, Kd) could be found that satisfy the available data, namely the quantified NFκB activity, mRNA half-life estimates, and mRNA-seq time-course data in control and mutant cells stimulated with LPS and TNF. Thereby, we aimed to determine whether mRNA half-life was itself a sufficient decoding mechanism to explain the stimulus-specific expression of group III duration-dependent NFκB target genes. We performed the fitting procedure for all 81 genes in group III and plotted both the experimental data and the simulation data as a heat-map (Fig. 4B). We found that for 31 genes, the model simulations recapitulated the RNA-seq data well, showing a fit with a normalized RMSD (nRMSD) of <0.13 (Table S4). As expected, many of these genes were induced rather late indicating a long mRNA half-life. Line graphs of mRNA time-courses for some genes deemed to be well-fit or poorly fit clarified that an nRMSD≤0.13 provided a stringent cut-off. Whereas Ccl2, Gsap, Rab15 and Mmp3 showed a good fit between simulation and data, Rel, Nfkb1 and Ccl1 showed a less good fit (Fig. 4C) and in most of these cases the TNF simulations in the mutant typically fell short of what the data indicated.
Figure 4. Model-aided analysis to determine sufficiency of mRNA half-life as the decoding mechanism.
(A) Diagram of model v1 showing NFκB activity and mRNA half-life (range) as inputs to output mRNA expression profiles. By comparing the predicted trajectory with the experimental measurement from RNA-seq, we can test for each gene in category II in Figure 2D, whether a parameter set can be found that allows the model to fit the data. (B) Comparing gene expression profiles between data and best-fit model simulation in response to indicated stimulus. The expression levels are normalized to the maximum in control or mutant individually. The yellow and black color bar indicates whether the best fit model is acceptable (normalized RMSD <0.13) or not (See also Table S4) (C) Line graphs for seven genes are shown to represent how well the best fit model matches the experimental data. (D) Testing whether NFκB dynamics-dependent LPS-specificity is dependent on the mRNA half-life for the fitted genes. Predicted Lvs.T specificity in best-fit model for control (black closed circle) and mutant (black open circle) for the genes with nRMSD < 0.13 in 4B is shown. The length of the line connecting control and mutant indicates the degree to which specificity is dependent on NFκB dynamics. The blue lines and dots are the in-silico perturbation results obtained by only changing the mRNA half-life in the best-fit model to 15 mins. (E) Expression trajectories for six genes with estimated half-life and fixed short half-life (15 mins) are shown. (F) Graph showing the comparison of Lvs.T specificity of model fitted (yellow) and non-fitted (black dots) genes in control vs mutant. Lvs.T specificity of model fitted (yellow) genes is significantly higher compared to non-fitted (black dots) genes in mutant, but not in control. P-values are generated by one-tailed Student’s T-test.
To test whether NFκB duration-dependent Lvs.T specificity was indeed dependent on the mRNA half-life for the fitted genes, we plotted the Lvs.T specificity of each of the 31 best-fit models for control cells (black closed circle) and mutant cells (black open circle) (Fig. 4D). The length of the line connecting Lvs.T specificity of control and mutant indicates the dependence of specificity on NFκB duration as probed with IκBα-deficiency. We then considered how this Lvs.T specificity metric would be affected if these genes had a short mRNA half-life. With mRNA half-life set to 15 mins, we simulated Lvs.T specificity of control and mutant in each of the 31 genes, plotted in blue (Fig. 4D). This analysis revealed that the Lvs.T specificity was universally dependent on a long mRNA half-life. Plotting the simulation time-course data directly for a few genes confirmed these results (Fig. 4E). When the estimated mRNA half-lives were used, the four sample genes showed high Lvs.T specificity in control conditions, but diminished Lvs.T specificity in the mutant. However, when the mRNA half-life was set to 15 min, Lvs.T specificity was either eliminated (Rab15) or significantly diminished (Ccl2, Gsap, Mmp3) in control and thus the mutant showed little effect.
While our simple modeling approach provides a sufficient explanation for 31of the 81 genes that show NFκB-duration-dependent Lvs.T specificity, we wondered whether remaining 50 genes that did not show an adequate fit to the simple model were in some manner distinct. We graphed Lvs.T specificity for control and mutant for the 31 best-fit genes and for the remaining 50 genes (Fig 4F). Remarkably, we found that while the good-fit genes showed only partial NFκB-duration-dependence for Lvs.T specificity, the remaining 50 genes showed a higher degree of NFκB-duration-dependence (p-value < 0.002). These results suggest that mRNA half-life may not be solely responsible for controlling Lvs.T specificity of these genes but that there may be additional regulatory layers.
Stimulus-specific NFκB duration may also be decoded by transcriptional mechanisms
In order to examine whether an additional regulatory layer for decoding stimulus-specific NFκB-duration may occur at the levl of transcriptional intiiation, we measured nascent transcript levels in control and mutant cells by isolating total chromatin-associated RNA (caRNA) followed by ribosomal RNA depletion and Next Gen Sequencing at six time-points in response to TNF and LPS stimulation (Fig. 5A). Of the 81 LPS-specific dynamics-dependent genes, 39 were found to be substantially LPS-specific at the level of caRNA in control cells (Fig. 5B). Among the 39 LPS-specific nascent transcripts, in the mutant 16 genes lost specificity to below the threshold. These data suggested that for some genes the mechanisms controlling transcriptional initiation may indeed be capable of decoding the duration of NFκB activity.
Figure 5. Stimulus-specific NFκB dynamics may be decoded at the level of transcriptional initiation.
(A) Cartoon showing the synthesis of nascent, chromatin-associated RNA (caRNA) in the nucleus and transport of mature RNA (mRNA) to cytoplasm after post-transcriptional processing. (B) Pie charts showing LPS specific or nonspecific genes using a threshold of 20.5 in control vs mutant at the caRNA level. (C) Heat-maps comparing gene expression (normalized to max) at PolyA+ RNA (mature-mRNA) and caRNA (pre-mRNA) levels for control and mutant cells for indicated time-points (See also Table S5) (D) Relative expression levels are shown by line graphs. (E) Heatmap of RelA ChIP-seq in control and mutant cells for genes in cluster 3 (top panel). Grayed out rows were genes for which no peaks were mapped to it. Boxplots show maximum peak intensity for each gene in the heatmap (normalized individually for control and mutant cells) (bottom panel) with p values indicated (two-sided t-test).
To relate caRNA-seq and mature polyA+ RNA time-courses, we plotted both datasets side by side for the 81 genes in a heatmap (Fig. 5C), showing them in the three clusters based on specificity. The first cluster contains 42 genes that did not show substantial LPS-specific caRNA levels and presumably achieves stimulus-specific polyA+ RNA through post-transcriptional mechanisms, including mRNA half-life. The second cluster contains 23 genes that showed LPS-specific caRNA in control cells and retained that specificity in the mutant; indeed, their stimulus-specificity at the polyA+ RNA level was only partially diminished in the mutant. These genes might be targets of other LPS-specific transcription factor(s) such as IRF3, or they may be sensitive to durations of NFκB longer than 2 hrs. The third cluster contains 16 genes that showed LPS-specific caRNA in control cells but lost that specificity in the mutant; in other words, their caRNA-seq data mirrored the polyA+ RNA-seq data, indicating that NFκB-duration-dependent LPS-specific gene expression for this cluster is produced primarily by chromatin-associated mechanisms that control transcriptional initiation.
Line graphs of a few example genes (Fig. 5D) confirmed these conclusions. For example, Ccl5 is LPS-specific at both the caRNA and polyA+-RNA level in control cells, with the Lvs.T specificity score > 0.5 for both data types. In mutant cells, this gene still showed some, though diminished LPS-specificity in caRNA but specificity at the polyA+-RNA level was almost entirely lost. Cgn is LPS-specific in caRNA as well as in polyA+-RNA in control but not in mutant cells indicating a critical role for the transcriptional initiation mechanism in decoding NFκB duration. Fpr1 showed LPS-specificity in control at caRNA as well as in polyA+ RNA level. This specificity was lost at the caRNA level in the mutant but only partially at the level of polyA+ RNA indicating the presence of LPS-specific post-transcriptional mechanism underlying LPS-specific gene expression.
NFκB-duration-dependent transcriptional initiation may in principle be mediated either by molecular mechanisms that control chromatin accessibility to NFκB or by some duration-sensitive downstream step in co-activator recruitment, pre-initiation complex assembly or activation. For the former, NFκB with longer nuclear residence time may gain better access to its binding site. We examined this possibility by performing RelA ChIP-seq analysis in both control and mutant cells. Focusing on RelA binding events in the proximity of genes in the third cluster, we found that their LPS-specificity was weakly correlated not just with the duration of RelA binding but peak ChIP read counts (Fig. 5E) indicating that RelA may not achieve equivalent access to its binding motif when it is transiently activated. In contrast, in mutant cells peak read-counts were similar in response to LPS and TNF (Fig. 5E, lower panel). This result indicates that the duration of NFκB activation determines whether RelA is able to gain access to its binding site and suggests that there is a chromatin-associated mechanism that decodes the duration of NFκB signaling.
Model-aided quantification of transcriptional and post-transcriptional contributions
In order to more quantitatively relate and interpret the caRNA and polyA+-RNA datasets, we constructed a mathematical model v2 in which we considered transcriptional initiation by a chromatin-associated mechanism that is sensitive to the duration of NFκB activity. Based on the observations from caRNA-seq and RelA-ChIP-seq data, model v2 describes three states of chromatin – closed, open, and active – with NFκB not only being able to activate the gene, but also acting to transition it from a closed to open state (Fig. 6A). This may reflect a conformational or chromatin-state transition that prior studies reported NFκB to be capable of either via SWI/SNF recruitment (Ramirez-Carrozzi et al., 2009) or de novo enhancer formation (Kaikkonen et al., 2013; Ostuni et al., 2013), and has been used previously to describe the control of the IL-4 gene (Mariani et al., 2010). A detailed description of the model equations is included in Methods.
Figure 6. Model-aided analysis to quantify chromatin-associated decoding of stimulus-specific NFκB dynamics.
(A) Model diagram and ODEs of two-step model v2. NFκB (red dimer) serves to both open chromatin and activate transcription. Chromatin transitions between closed, open, and active states. Transcription can only occur from the active state. Parameters in parentheses correspond to the parameters in the model. The details of the model can be found in the Method section. (B) Experimental data and model simulation of three example genes with nRMSD for each gene shown. (C) Simulation heatmap for caRNA and polyA+ RNA in control and mutant MEFs stimulated with LPS or TNF. (nRMSD <0.13 defined as good fit for v2, marked with Yellow; For v1, same as Fig. 4B, see also Table S6) (See also Fig.S3 for side-by-side comparison of data and simulation.)
To quantitatively interpret the available data, we followed the established parameterization workflow. Specifically, we first determined whether there is a set of values for the free parameters (k−1, k−2, Kd1, Kd2) that satisfy the available data for a given gene, namely, the quantified NFκB activity, measured mRNA half-life, and caRNA-seq and polyA+-RNA-seq time-course data in control and mutant cells stimulated with LPS and TNF. When the normalized root-mean square difference (nRMSD) between model simulations and data time-points was ≤0.13, we deemed the model to provide an appropriate fit to the data (Table S6). Line graphs showing the best fit model simulation results for caRNA and mRNA dynamics (Fig. 6B) for 3 genes (Ccl5, Cgn and Fpr1) provide a visual confirmation that these models recapitulate the data reasonably well. By this measure, we found that model v2 accounts for 37 additional genes that were not accounted for in model v1, indicative of a role of a chromatin-associated mechanism in decoding NFκB duration. Meanwhile, 1 gene (Slc6a12) whose mRNA data was accounted for by Model v1 fell outside the fit threshold with v2, due to poor concordance of the caRNA simulations and caRNA-seq data. Overall, model v2 accounted for 67 of 81 (~83%) stimulus-specific genes that are dependent on NFκB dynamics (Fig. 6C, Fig. S3). The remaining 14 genes whose expression dynamics could not be explained by v2 suggest more complex regulation, potentially involving other stimulus-induced transcription factors or signaling pathways.
Gene regulatory strategies correlate with gene functions in immune responses
The described studies revealed that a slow chromatin-opening mechanism and slow mRNA decay may combine to decode the differential duration of LPS- vs TNF-induced NFκB dynamics more effectively (Fig. 7A). Using the mathematical model, we parameterized for each of the 67 genes for which a satisfactory fit was produced, we could now quantify the relative contribution of these two regulatory strategies for decoding the duration of NFκB dynamics. Starting with caRNA data recapitulated by the model (Fig. 6C), we obtained the duration-dependent specificity observable at the chromatin level. Taking the difference of mutant vs control specificity using the simulated mRNA data further describes the total dynamics-dependent specificity, a portion of which was the chromatin-associated specificity. Remaining Lvs.T specificity in the mutant is a result of other unknown mechanism(s). In this manner, for 67 genes whose expression profiles could be accounted for by model v2, we calculated the contribution of each mechanism and plotted the quantification result as a stacked bar graph that also indicates the residual as yet unaccounted specificity for each gene (Fig. 7B). Using this analysis, we found that the vast majority of genes involve multiple mechanisms to decode the duration of NFκB dynamics. Although most genes employ multiple mechanisms for decoding, among the 67 genes, 11 genes employed predominantly slow mRNA decay and 30 genes slow chromatin transition. In support of this, small molecule histone deacetylase (HDAC) inhibitor, Trichostatin A, showed more hyper-expression effect on chromatin regulated genes (e.g. Lcn2, Fpr1) after TNF stimulation than genes that are not predominantly chromatin regulated (e.g. Rab20, Ccl7) (Fig. S4).
Figure 7. Distinct regulatory strategies that decode NFκB duration-dependent gene expression.
(A) Diagram illustrating two broad regulatory mechanisms that may decode stimulus-specific duration of NFκB dynamics to contribute to LPS vs.TNF-specific gene expression. (B) Stacked bar graphs showing the relative contribution of mRNA half-life (T1/2) and chromatin-regulation to the decoding of stimulus-specific gene expression. The grey portion of each bar denotes stimulus-specificity that is not mitigated by the NFκB dynamic mutant. Genes are ordered by whether they are controlled by a predominantly chromatin-associated mechanism, mixed mechanisms, or a predominantly mRNA T1/2 mechanism. (C) Scatterplot of chromatin contribution versus half-life contribution for all genes. Four selected genes from Fig. 7B that are controlled by either predominantly a chromatin-associated mechanism or an mRNA half-life mechanism are indicated. (D) Example ATAC-seq track view of a predominantly chromatin-controlled gene. (E) Boxplots of read counts for ATAC-seq data indicates that chromatin-regulated genes are more likely to have closed chromatin at promoter regions (peaks found within −1000 to 100 basepairs of the TSS). Genes are grouped by the categories shown in Fig. 7B. Two-sided Wilcoxon-Mann-Whitney U test p values are shown. (F) Boxplots of the difference in read counts of Control and Mutant cells after 2hrs treated with TNF pulse, compared to the 0 hr basal counts. Genes are grouped by the categories shown in Fig. 7B. Shown are one-sided Wilcoxon-Mann-Whitney U test p values that chromatin-controlled genes shown greater differences from baseline. Lines inside boxplots represent the 25th, 50th, and 75th quantiles. Whiskers extend up to 1.5 the interquartile range, with genes outside this represented as outlier points.
We observed that genes that have higher stimulus-specificity tend to rely on chromatin-mediated mechanisms rather than post-transcriptional mechanisms to decode duration. Indeed, genes that use only mRNA half-life as a decoding mechanism have a specificity index of below 1.5. In contrast, genes that employ chromatin associated mechanisms to decode duration tend to have high specificity indices, with more than half genes having a specificity greater than 1.5 (Fig. 7B). This suggests that chromatin-mediated mechanisms can in general generate more pronounced stimulus-specificity in gene expression from stimulus-specific NFκB duration.
After sorting the genes by their gene regulatory strategy (i.e. predominantly chromatin-associated, mRNA half-life, or both, Fig. 7B), we examined their cellular and physiological functions. We found that several functionally important genes decoded predominantly by mRNA half-life were cell-intrinsic regulators of innate immune function, such as transcription factors and their regulators (Nfkb2, Nfkbie) and genes involved in cytokine signal transduction pathways (Pim1, Il1rl1) (Fig. 7C). In addition, the monocyte chemoattractant Ccl2 was in this category, indicating that recruitment of immune surveillance cells that can appropriately regulate immune responses was LPS-specific via the mRNA decay gene regulatory strategy. Examining the genes whose stimulus-specificity employed a chromatin-associated gene regulatory strategy, we identified the phagocytosis receptor, Fpr1, and the tissue remodeling protease, Mmp3, that are key effectors of immune responses, which may then synergize with the function of Ccl5, a key chemoattractant of cytotoxic T-cells (Fig. 7C). Thus, our analysis suggests that the stimulus-specific expression of immune effector genes may be more dependent on cell type, microenvironmental context, and history than that of immune regulatory genes due to their employment of a chromatin-associated rather than mRNA decay-mediated gene regulatory strategy to decode stimulus-specific NFκB dynamics.
To examine whether molecular correlates of a chromatin-associated mechanism for decoding NFκB signaling duration could be identified, we turned to ATAC-seq and grouped genes by their predominant duration decoding strategy. We made two observations – that chromatin-regulated genes are more likely to have closed chromatin at promoter regions (Fig. 7E) at the basal state, and that these genes showed increased accessibility compared to the basal state after TNF stimulation in the dynamic control mutant, as seen in the gene Il34 (Fig. 7D, 7F). This experimental evidence further supports a slow chromatin opening step for a subset of genes as a gene regulatory strategy for decoding stimulus-specific signaling duration.
Discussion
In this study, we report progress in addressing the longstanding question of how immune response genes decode differential temporal profiles of NFκB. This question was first raised by the discovery that NFκB activation showed complex dynamic control (Hoffmann et al., 2002) that was indeed found to be stimulus-specific (Covert et al., 2005; Werner et al., 2005). Further, the duration of NFκB activity was found to be correlated with the expression of some NFκB target genes (Hoffmann et al., 2002; Werner et al., 2005), even when NFκB duration distributed into several discrete pulses (Ashall et al., 2009). However, the mechanisms by which genes are able to achieve stimulus-specific expression by distinguishing differential durations of NFκB activity remained unknown. The present study addressed this question in a quantitative manner by employing mathematical models to interpret quantitative experimental data produced at high temporal resolution. Here, we have demonstrated that not only mRNA half-life, previously shown to play a critical role in duration decoding of p53 and ERK (Hafner et al., 2017; Uhlitz et al., 2017), but also chromatin regulated mechanism(s) are important for duration decoding of NFκB target genes.
Our analysis revealed that the genes most strongly regulated by duration employ both the transcriptional and post-transcriptional decoding mechanisms. Interestingly, immune effectors and immune regulators were found predominately regulated by the former and latter, respectively. How may the chromatin mechanism work? As no binding sites other than NFκB were identified among LPS-specific dynamics-dependent genes (Fig. 2E), we found no evidence for coherent feedforward loops as the prevalent regulatory mechanism, but instead suggest that slow steps in chromatin opening or PIC assembly are required for decoding. A functional consequence of this purely kinetic explanation for why genes may employ one mechanism over another may be that chromatin may be a more versatile regulatory node; it may receive inputs from diverse micro-environmental contexts or exposure histories. For example, for a gene with high LPS specificity based on chromatin control, memory of a prior direct pathogen exposure may allow the gene to be expressed later with less stimulus-specificity, also in response to cytokines released from pathogen-infected neighbors. In addition, chromatin-based mechanisms are strongly cell-type-specific, thus rendering stimulus-specificity a function of cell type.
The present study provides a general approach that may be used to elucidate the regulatory strategies underlying TF control of gene expression at single gene resolution. Iterative mechanistic modeling and experimentation has been successful in elucidating the mechanisms that encode dynamic transcription factor activities (Basak et al., 2012), but for understanding the regulatory strategies of gene expression, this approach encounters particular challenges. While Next Generation Sequencing methods provide highly quantitative genome-wide data, errors are associated with low abundance mRNAs, and variabilities in library preparation methodologies and data normalization methods. This is compounded for caRNA-seq data where the read depth tends to be lower, and the benefits of polyA+ selection do not apply. Previous studies that fit transcriptomic data to models either did not include TF activities as inputs to elucidate the regulatory logic (Rabani et al., 2011) or did not attempt to provide quantitative model fits at single gene resolution (Cheng et al., 2017). Indeed, the transcriptional control mechanisms are highly complex, involving numerous factors and potentially multiple regulatory steps; these must be abstracted without losing critical regulatory behavior.
A key principle of mathematical biology holds true here also: when a model fits the data it does not mean that the model is correct, but it represents a starting point for further iterative testing. This is illustrated in our study: when we iterated with additional data in the form of caRNA-seq, our initial conclusions about the key role of mRNA half-life were confirmed for only some genes (11 out of 67), while we found substantial contributions by a chromatin mechanism for the remaining 56 genes, 30 of which were predominantly controlled by the chromatin mechanism. This in turn suggests that further iterations may lead to additional insights. For example, other gene regulatory strategies or mechanisms such as pre-mRNA processing, splicing or mRNA transport might play regulatory roles in decoding NFκB duration.
Theoretical considerations (Behar and Hoffmann, 2010) suggest a diverse set of potential decoding mechanisms. First, any slow kinetic process that controls either gene activation (e.g. mRNA synthesis, slow chromatin opening, slow pre-initiation complex (PIC) assembly) or slow mRNA decay (e.g. long mRNA half-life) can in principle decode the duration of TF activity. Second, coherent feedforward logic where two TFs present coincidently (AND gate) might also decode duration of TF activity. The inclusion of additional transcription factors (e.g. AP1 or IRF) in subsequent model versions may also be key to accounting for the remaining specificity not accounted for by model v2, or the 14 of 81 dynamics-controlled genes not satisfactorily recapitulated.
STAR Methods
The Materials Availability Statement (CONTACT FOR REAGENT AND RESOURCE SHARING)
This study did not generate new materials. Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contact, Alexander Hoffmann (ahoffmann@ucla.edu).
EXPERIMENTAL MODEL AND SUBJECT DETAILS
Cell Culture:
Mouse embryonic fibroblasts (MEFs) were prepared from male or female E12-E14 embryos from C57BL/6 mice that were Ifnar−/− and Ifnar−/−Nfkbia−/− and cultured in DMEM containing 10% BCS for 5–6 passages before being stimulated (Werner et al., 2005).
Stimulation Conditions:
MEFs were stimulated with 100ng/ml LPS (Sigma, B5:055), 1 ng/ml murine TNF (R&D Systems) or 1 ng/ml IL1 (Pepro Tech, USA) for 30 minutes and washed with warm PBS for 3 times. Cells were cultured with conditioned media (without stimulus) until harvested at indicated time points for further manipulations. For TSA experiment, Ifnar−/− MEFs were treated for 24 hrs. with 50 nM TSA (T1952, Sigma-Aldrich) and then stimulated with 1 ng/ml murine TNF for 30 minutes and washed with warm PBS for 3 times and then cultured until harvested for RNA isolation.
Animal Use:
The use of mice and isolation of MEFs have been approved by the Animal Care and Use Committee of University of California, Los Angeles.
METHOD DETAILS
In all Figures, the data presented are representative of at least 2 or more independent experiments. Data were not divided into training and test datasets for any of the analyses performed in this paper. Blinding of the experimenter to the sample genotypes was not performed at any stage of the study.
Electrophoretic mobility shift assays (EMSAs):
EMSAs were conducted with standard methods as described previously for fibroblasts (Basak et al., 2007; Hoffmann et al., 2003). Whole cell lysates were made with RIPA buffer; nuclear extracts by hypotonic cell lysis and high salt extraction of nuclear proteins.
Cell Fractionation and RNA isolation:
After stimulation, cells were harvested at desired time points. For PolyA+ RNA, cells were harvested in TRIzol reagent (Life Technologies, Carlsbad, CA). Then, DNA-free RNA was extracted from cell using DIRECTzol kit (Zymo Research, Irvine, CA) according to manufacturer’s instructions. For chromatin RNA, subcellular fractions were prepared as described (Pandya-Jones and Black, 2009), with minor changes. The cell lysis buffer contained 0.15% NP-40, and the sucrose cushion did not contain detergent. For chromatin RNA isolation, chromatin fraction was suspended in TRIzol reagent. Followed by chloroform extraction, aqueous phase containing RNA was used for purification with Direct-zol RNA Miniprep Kit (Zymo Research). Chromatin fraction purity was confirmed by immunoblot analysis with anti-B-Tubulin (Sigma, T5201), anti-SNRP70 (Sigma, AV40275) and anti-Histone H3 (abcam, ab1791) antibodies.
cDNA synthesis and qPCR:
RNA was reverse transcribed with iScript Reverse Transcription Supermix for RT-PCR (Bio-Rad). qPCR was performed using SsoAdvanced Universal SYBR Green Supermix (Bio-Rad).
Library Preparation and RNA sequencing:
After RNA extraction, libraries for polyA+ RNA were prepared using KAPA Stranded RNA-Seq Kit for Illumina Platforms (KAPA Biosystems, Wilmington, MA) according to the manufacturer’s instructions. caRNA sequencing libraries were prepared using the KAPA Stranded RNA-Seq Kit with Ribo Erase for Illumina Platforms (KAPA Biosystems). Resulting cDNA libraries were single-end sequenced with a length of 50bp on an Illumina HiSeq 2000 (Illumina, San Diego, CA).
RelA ChIP-seq:
After stimulation, cells were harvested at desired time points. ChIP-seq libraries were prepared using the NEB Next Ultra DNA Library Prep Kit for Illumina (New England Biolabs). ChIP-seq was performed as described (Barish et al., 2010) using anti-RelA (Santa Cruz Biotechnology, sc-372) antibody.
ATAC-seq:
For ATAC-seq libraries, cells were dissociated with Accutase (Thermo Fisher Scientific, Waltham, MA), and 50,000 cells were used to prepare nuclei. Cell membrane was lysed using cold lysis buffer (10mM Tris-HCl pH7.5, 3mM MgCl2, 10mM NaCl and 0.1% IGEPAL CA-630). Nuclei were pelleted by centrifugation for 10 minutes at 500 x g, and suspended in the transposase reaction mixture (25 µl of 2X TD Buffer (Illumina), 2.5 µ l of TD Enzyme 1 (Illumina), and 22.5 µ l of nuclease-free water). The transposase reaction was performed for 30 minutes at 37C in a thermomixer shaker. Then, fragmented DNA in the reaction was purified using MinElute PCR purification kit (QIAGEN, Hilden, Germany). The purified DNA fragments were amplified by PCR to obtain ATAC-seq libraries with Illumina Nextera sequencing primers. The libraries were purified using MinElute PCR purification kit (QIAGEN) and quantified using KAPA Library Quantification Kit (KAPA Biosystems). The libraries were single-end sequenced with a length of 50bp on an Illumina HiSeq 4000.
STATISTICAL ANALYSIS
Sequence Mapping and Analysis of polyA+ RNA-seq and caRNA-seq:
After adapter trimming, single-end reads were mapped to reference mouse genome (mm10) using STAR (Dobin et al., 2013) with default parameters. Only primary mapped reads with alignment score (MAPQ)>30 were selected by ‘samtools view -F 2820 -q 30’. Transcript abundance was quantified based on GENECODE M6 (GRCm38.p4) annotation using featureCounts (Liao et al., 2014) using option ‘-t exon -g gene_id. The raw counts of PolyA+ RNA-seq were input to DESeq2 to determine LPS-induced genes by comparing each sample of LPS stimulation time points against unstimulated samples with threshold log2Fold change >2 and Benjamini-Hochberg False discovery rate (FDR) <0.01. The expression levels of genes in each sample were normalized by means of Reads Per Kilobase per Million (RPKM) mapped reads in the downstream analysis.
Specificity score calculation:
The stimulus-specificity score was calculated for each pair of stimuli gene-wise. The stimuli A vs B specificity score for gene g is calculated as follow:
i.e. the log2 ratio of maximum gene expression (mean RPKM of replicates) for all the measured timepoints between stimulation condition A and B.
Transcription factor motif and Gene Ontology (GO term) analysis:
Known transcription factor motif enrichment analysis and GO term enrichment analysis was performed using ‘findMotif.pl’ from HOMER (Heinz et al., 2010) on promoter regions (−500bp to +200bp TSS) of genes in each category in Fig. 2C and the remaining genes in the other two categories as control. Only results for κB, AP-1 and ISRE motifs and the GO:0016070, GO:30035458 and GO:0045087 are displayed in Fig.2E, F.
Determination of mRNA half-life:
Half-lives of mRNAs (Fig. 3D, E, F) were determined from unstimulated Ifnar−/− MEFs treated with 10 mg/ml Actinomycin D (A9415, Sigma-Aldrich) for 0.5, 1, 2, 3, 4, and 6 hr in replicate. 2µ L of 1:100 diluted RNA Spike-In control mix (Ambion ERCC RNA Spike-In Mix Part no 4456740) were added with RNA during library preparation. These libraries were sequenced and processed to get the counts of transcripts per gene using the same process pipeline as described earlier. The gene counts were normalized by size factor per library using the median counts of all the spike-ins that have at least 32 counts. Reads counts normalized, then log2 transformed after adding pesudocount of 1. Then half-life was derived at the log2 normalized count scale based on the procedure below.
Due to confounding drug-induced stress responses, mRNA levels of some genes were increased transiently following Actinomycin treatment, and some mRNAs displayed two-phase reduction: a fast decay followed by a slow decay. Due to these observations, we designed the following strategy to derive the mRNA half-life (Table S3): First, select one of the first three time points (0, 0.5hr, and 1hr), which has the highest level of expression as the starting point. To ensure the reliability of the data, we only considered the mRNAs with ≥32 count. Second, from the selected time point, select the next two or more consecutive time points to perform linear regression at log2 scale (indicated as startID and endID in Table S3). Third, select the linear regression with highest adjusted R2 score to derive the slope. Fourth, calculate the 95% confidence interval (CI) based on CI of potential slopes.
Mapping of RelA ChIP-seq and analysis:
RelA ChIP seq reads were mapped to mm10 using bowtie2 with option `--non-deterministic – very-sensitivè. Peaks were called using macs2 and peaks across all time points and conditions were merged to obtain a consensus peak set. Reads were normalized based on the total number of reads mapped per sample. HOMER was first used to annotate peaks, resulting in a many-to-one peak to gene mapping. Because peaks can be associated with genes from a long distance, to map functional peaks to genes, we utilized the caRNA data and calculated Spearman’s correlation of peaks to caRNA-seq measurements for peaks within −100000/+50000 of the TSS. The single peak with the highest correlation was chosen as representing a functional binding event for that gene. Counts were further scaled to the maximum count for each genotype.
ATAC-seq analysis:
Raw fastq files were processed by taking the reads and trimming, filtering, and aligning to mm10. Peaks were called using MACS2 for each individual sample. Reads that mapped to mitochondrial genes or blacklisted regions were removed. Peak files were merged using bedtools merge to obtain a consensus set of peaks across samples. The number of reads that fell into the peaks was obtained using bedtools multicov, with the parameter –q 30 to exclude reads with poor mapping quality. Reads were then normalized based on the total number of reads mapped per sample. HOMER was used to associate peaks to genes based on the nearest TSS. Because peaks can be associated with genes from a long distance, we further utilized the caRNA data and calculated Spearman’s correlation of peaks to caRNAseq measurements for all peaks, selecting peaks to the nearest gene. Genes were grouped as predominantly chromatin-controlled, half-life controlled, if either chromatin of half-life accounted for greater than 30% of the known specificity, respectively, and the rest were categorized as ‘mixed’. Normalized counts were then used to find the difference in counts for the relevant peaks by subtracting the counts in the basal timepoint, and then plotting these values as boxplots.
Mathematical Modeling:
Model v1 (Fig.3 and 4):
For the simulations shown in Fig.3, the activity of transcription factor (TF(t)) was interpolated from quantified EMSA measurements (in controls: TNF0:1, TNF30: 42.1, TNF60: 25.2, TNF90: 4.6, TNF120: 1, TNF240: 1, TNF360: 1, TNF480:1; LPS0: 1, LPS30:22.5, LPS60: 37.8, LPS90, 42.6, LPS120: 40.6, LPS240: 30.8, LPS360: 20.6, LPS480: 18.6; and in mutant: TNF0:1, TNF30: 36.6, TNF60: 28.9, TNF90: 20.1, TNF120: 18.5, TNF240: 1.8, TNF360: 1.2, TNF480:1.3; LPS0: 1, LPS30:13.1, LPS60: 42.2, LPS90, 34.0, LPS120: 27.8, LPS240: 24.5, LPS360: 15.4, LPS480: 10.1) by using the `pchipfun` from R package `pracmà. These were then inputted into the TF(t) term of the ODE, which was numerically solved by òdè from R package `deSolvè with time step 0.1 min for the sweep of different mRNA half-lives (50 evenly spaced in log10 scale ranging from 1 min to 1000 mins). Other parameters in the simulation are: k0 = 0.001, kf = 0.5, Kd=0.5, n=6.
To fit the ODE model to each LPS induced gene in Fig.4, we first introduced a delay parameter tau in TF(t-tau) to account for the time gap between TF nuclear localization and the appearance of polyA+ mRNA in the cytoplasm. The model is numerically solved by ode function from `deSolvè package for all input functions to predict this gene’s expression dynamics. The parameters are fitted for gene j by minimizing the difference between simulation’s result and the experimental RNA-seq data (Table S2) using normalized RMSD (by range of data):
using `nlminb` function from R in log10 scale. Except for n, which is discrete from 1 to 6, kj 0, kj f and Kj d are bound by 10−3 to 103, kj deg is bound by the experimental half-life determinations (Table S3), and tau is bound by 1e-5 to 120 mins. The results of the model fitting are listed in Table S4.
Model v2 (Fig.6)
The version 2 model includes three states of the promoter: closed (C), open (O), and activated in transcription (A) and the transitions among these states. Only state A promoter can produce mRNA with rate constant kp. We assume that the sum of the fractions of these promoter states is constant, that the transitions from C to O to A are regulated by NFκB (modeled by Michaelis-Menten kinetics), and that they are reversible. The process and degradation of mature mRNA is modeled by a first-order equation. The model equations are shown in Fig. 6A.
Forward reactions, which are regulated by NFκB and parameters Kd1 and Kd2, contain parameters k1 and k2. To simplify parameter fitting, we assume k1 = k2 = 1. mRNA production, which is regulated by fractional state A, contains parameter kp. To simplify parameter fitting, we assume kp = 1. Other parameters are fitted to experimental measurements of caRNAseq and polyA+RNAseq data (Table S2 and Table S5) using the same algorithm as model v1. The results of the model fitting are listed in Table S6.
Calculation of contribution of different regulatory strategies:
To calculate the contributions of the chromatin and mRNA half-life associated strategies to the total specificity, we used the simulated data from model v2. For each gene, we first calculated the difference in specificity for control versus mutant from the caRNA data to assess the amount of specificity at the caRNA level that is reduced by the dynamics mutant (ΔspecificitycaRNA). These are the dynamics-generated specificities that are controlled at the chromatin level, without any influence from mRNA half-life. The chromatin-contribution is therefore ΔspecificitycaRNA. We next made the same calculation of difference in specificities for control versus mutant from the mRNA data (ΔspecificitymRNA). At the mRNA level, the specificity in the control represents the total specificity, and thus the difference in specificity between mutant and control at the mRNA level represents the contributions of both chromatin and mRNA half-life–associated mechanisms. It follows from this that by subtracting the two deltas (Δ specificitymRNA - Δ specificitycaRNA), we can obtain the mRNA half-life contribution to the specificity. The ‘remaining’ mechanisms that are not accounted for by either chromatin or mRNA half-life associated mechanisms are represented by the amount specificity still remaining in the mutant at the mRNA level.
DATA AND CODE AVAILABILITY
All sequencing data were deposited to Sequence Read Archive (SRA, NCBI) under BioProject IDs PRJNA453806 (polyA+ RNA-seq), PRJNA454496 (ActD RNA-seq), PRJNA454896 (ca-RNA seq), PRJNA517534 (RelA-ChIPseq) and SAMN12325160, SAMN12325163, SAMN12325167, SAMN12325170 (ATAC-seq). All the code necessary for reproducing the figures, model fitting and simulation is available on the GitHub site mentioned in the STAR methods.
Supplementary Material
Table S1: RPKM values for the time points in ifnar−/−; nominal and FDR corrected p values of TNF, LPS, IL1-induced genes; and scaled expression values of RNA-seq in ifnar−/−. Related to Fig. 1.
Table S2: RPKM of RNAseq in ifnar−/− (control) and ifnar−/−nfkbia−/− (mutant); specificity scores and p values for control and mutant replicates (one-sided t- test); and scaled expression values of RNA-seq in ifnar−/− (control) and ifnar−/−nfkbia−/− (mutant) used for heatmap and math modeling. Related to Fig. 2.
Table S3: Log2 fold change for actD RNA-seq and derivated mRNA half-life; normalized counts for actD RNA-seq; and mRNA degradation rates regression for stimulus-specific genes. Related to Fig. 3.
Table S4: Values of parameters for Model v1. Related to Fig. 4.
Table S5: RPKM and scaled expression values of caRNA for the time points in control and mutant. Related to Fig. 5.
Table S6: Values of parameters for Model v2. Related to Fig. 6.
KEY RESOURCES TABLE
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| anti-RelA | Santa Cruz Biotechnology | sc-372 |
| anti-Acetyl Histone H4 | Millipore-Sigma | 06-866 |
| anti-H3K27Ac | abcam | ab4729 |
| anti-Histone H3 | abcam | ab1791 |
| Biological Samples | ||
| Chemicals, Peptides, and Recombinant Proteins | ||
| LPS | Sigma | B5:055 |
| TNF | R&D Systems | 410-MT |
| IL1 | Pepro Tech, USA | 211-11B |
| Actinomycin D | Sigma-Aldrich | A9415 |
| Trichostatin A | Sigma-Aldrich | T1952 |
| iScript Reverse Transcription Supermix | Biorad | 1708841 |
| SsoAdvanced Universal SYBR Green Supermix | Biorad | 172-5270 |
| ERCC RNA Spike-In Mix | Ambion | Part no 4456740 |
| Critical Commercial Assays | ||
| KAPA Stranded RNA-Seq Kit | KAPA Biosystems, Wilmington, MA | KR0934 |
| KAPA Stranded RNA-Seq Kit with Ribo Erase for Illumina Platforms | KAPA Biosystems, Wilmington, MA | KR1151 |
| NEB Next Ultra DNA Library Prep Kit for Illumina | New England Biolabs | E7645 |
| Nextera tagmentation master mix | Illumina | FC-121-1030 |
| KAPA Library Quantification Kit | KAPA Biosystems, Wilmington, MA | KR0405 |
| MinElute PCR purification kit | Qiagen | 28004 |
| DIRECTzol kit | Zymo Research, Irvine, CA | R2070 |
| Deposited Data | ||
| PRJNA453806 (polyA+ RNA-seq) | This paper | PRJNA453806 |
| PRJNA454496 (ActD RNA-seq) | This paper | PRJNA454496 |
| PRJNA454896 (ca-RNA seq) | This paper | PRJNA454896 |
| PRJNA517534 (RelA ChIP-seq) | This paper | PRJNA517534 |
| SAMN12325160 (Control-0 hr ATAC-seq) | This paper | SAMN12325160 |
| SAMN12325163 (Control-TNF 2 hr-ATAC-seq) | This paper | SAMN12325163 |
| SAMN12325167(Mutant-0 hr ATAC-seq) | This paper | SAMN12325167 |
| SAMN12325170 (Mutant-TNF 2 hr-ATAC-seq) | This paper | SAMN12325170 |
| Experimental Models: Cell Lines | ||
| Experimental Models: Organisms/Strains | ||
| Ifnar−/− C57BL/6 mice | This paper | NA |
| Ifnar−/−Nfkbia−/− C57BL/6 mice | This paper | NA |
| Oligonucleotides | ||
| 5’ttctgtgcctgctgctcata3’ | Ccl7 F | |
| 5’ttgacatagcagcatgtggat3’ | Ccl7 R | |
| 5’tgttttccctggtccaagtc3’ | Fpr1 F | |
| 5’atcgtaccctggatttgtgc3’ | Fpr1 R | |
| 5’atttcccagagtgaactggc3’ | Lcn2 F | |
| 5’aatgtcacctccatcctggt3’ | Lcn2 R | |
| 5’tgctctaatcaggacccattg3’ | Sod2 F | |
| 5’gtagtaagcgtgctcccacac3’ | Sod2 R | |
| Software and Algorithms | ||
| STAR | (Dobin et al., 2013) | https://github.com/alexdobin/STAR |
| featureCounts | (Liao et al., 2014) | http://bioinf.wehi.edu.au/featureCounts/ |
| DESeq2 | (Love et al., 2014) | https://bioconductor.org/packages/release/bioc/html/DESeq2.html |
| HOMER | (Heinz et al., 2010) | http://homer.ucsd.edu/homer/ |
| MACS2 | (Zhang et al., 2008) | https://github.com/taoliu/MACS/ |
| GenSA | https://github.com/cran/GenSA/blob/master/R/GenSA.R | |
| deSolve | https://cran.r-project.org/web/packages/deSolve/index.html | |
| Pracma | https://cran.r-project.org/web/packages/pracma/index.html | |
| Other | ||
| Code for reproducing figures, model fitting and simulation | https://github.com/biomystery/duration_decode_manuscript | |
Highlights.
We identify 81 genes whose pathogen-specific expression depends on NFκB duration
Iterative math modeling delineates gene-specific decoding of NFκB duration
A mRNA half-life of >30 min enables pathogen specificity of many immune regulators
Many immune effectors decode NFκB duration via a slow chromatin regulatory step
Acknowledgements
We thank lab members Diane Lefaudeux and Quen Cheng for helpful discussions. The work was supported by NIH grants GM117134, AI127864, and AI132835 (to AH). KMS is supported by the UCLA Medical Scientist Training Program (NIH NIGMS T32 GM008042). Sequencing was performed at the UCLA Broad Stem Cell Center Sequencing Core.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Conflict of Interest
The authors have no conflict of interest.
References
- Amit I, Garber M, Chevrier N, Leite AP, Donner Y, Eisenhaure T, Guttman M, Grenier JK, Li W, Zuk O, et al. (2009). Unbiased Reconstruction of a Mammalian Transcriptional Network Mediating Pathogen Responses. Science 326, 257–263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ashall L, Horton CA, Nelson DE, Paszek P, Harper CV, Sillitoe K, Ryan S, Spiller DG, Unitt JF, Broomhead DS, et al. (2009). Pulsatile stimulation determines timing and specificity of NF-kappaB-dependent transcription. Science 324, 242–246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barish GD, Yu RT, Karunasiri M, Ocampo CB, Dixon J, Benner C, Dent AL, Tangirala RK, and Evans RM (2010). Bcl-6 and NF- B cistromes mediate opposing regulation of the innate immune response. Genes & Development 24, 2760–2765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Basak S, Behar M, and Hoffmann A (2012). Lessons from mathematically modeling the NF-κB pathway: Mathematical modeling the NF-κB pathway. Immunological Reviews 246, 221–238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Batchelor E, Loewer A, Mock C, and Lahav G (2014). Stimulus-dependent dynamics of p53 in single cells. Molecular Systems Biology 7, 488–488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Behar M, and Hoffmann A (2010). Understanding the temporal codes of intra-cellular signals. Current Opinion in Genetics & Development 20, 684–693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhaumik SR (2011). Distinct regulatory mechanisms of eukaryotic transcriptional activation by SAGA and TFIID. Biochim Biophys Acta 1809, 97–108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bintu L, Buchler NE, Garcia HG, Gerland U, Hwa T, Kondev J, and Phillips R (2005). Transcriptional regulation by the numbers: models. Current Opinion in Genetics & Development 15, 116–124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng CS, Feldman KE, Lee J, Verma S, Huang D-B, Huynh K, Chang M, Ponomarenko JV, Sun S-C, Benedict CA, et al. (2011). The Specificity of Innate Immune Responses Is Enforced by Repression of Interferon Response Elements by NF- B p50. Science Signaling 4, ra11–ra11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng CS, Behar MS, Suryawanshi GW, Feldman KE, Spreafico R, and Hoffmann A (2017). Iterative Modeling Reveals Evidence of Sequential Transcriptional Control Mechanisms. Cell Systems 4, 330–343.e5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Covert MW, Leung TH, Gaston JE, and Baltimore D (2005). Achieving stability of lipopolysaccharide-induced NF-kappaB activation. Science 309, 1854–1857. [DOI] [PubMed] [Google Scholar]
- Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, and Gingeras TR (2013). STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elkon R, Zlotorynski E, Zeller KI, and Agami R (2010). Major role for mRNA stability in shaping the kinetics of gene induction. BMC Genomics 11, 259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilchrist M, Thorsson V, Li B, Rust AG, Korb M, Roach JC, Kennedy K, Hai T, Bolouri H, and Aderem A (2006). Systems biology approaches identify ATF3 as a negative regulator of Toll-like receptor 4. Nature 441, 173–178. [DOI] [PubMed] [Google Scholar]
- Hafner A, Stewart-Ornstein J, Purvis JE, Forrester WC, Bulyk ML, and Lahav G (2017). p53 pulses lead to distinct patterns of gene expression albeit similar DNA-binding dynamics. Nat Struct Mol Biol [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hao N, and O’Shea EK (2012). Signal-dependent dynamics of transcription factor translocation controls gene expression. Nat Struct Mol Biol 19, 31–39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hao S, and Baltimore D (2009). The stability of mRNA influences the temporal order of the induction of genes encoding inflammatory molecules. Nat Immunol 10, 281–288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, Cheng JX, Murre C, Singh H, and Glass CK (2010). Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell 38, 576–589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffmann A (2016). Immune Response Signaling: Combinatorial and Dynamic Control. Trends in Immunology 37, 570–572. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffmann A, and Baltimore D (2006). Circuitry of nuclear factor kappaB signaling. Immunological Reviews 210, 171–186. [DOI] [PubMed] [Google Scholar]
- Hoffmann A, Levchenko A, Scott ML, and Baltimore D (2002). The IkappaB-NF-kappaB signaling module: temporal control and selective gene activation. Science 298, 1241–1245. [DOI] [PubMed] [Google Scholar]
- Kaikkonen MU, Spann NJ, Heinz S, Romanoski CE, Allison KA, Stender JD, Chun HB, Tough DF, Prinjha RK, Benner C, et al. (2013). Remodeling of the enhancer landscape during macrophage activation is coupled to enhancer transcription. Mol Cell 51, 310–325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee RE, Walker SR, Savery K, Frank DA, and Gaudet S (2014). Fold change of nuclear NF-kappaB determines TNF-induced transcription in single cells. Mol Cell 53, 867–879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liao Y, Smyth GK, and Shi W (2014). featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930. [DOI] [PubMed] [Google Scholar]
- Litvak V, Ramsey SA, Rust AG, Zak DE, Kennedy KA, Lampano AE, Nykter M, Shmulevich I, and Aderem A (2009). Function of C/EBPdelta in a regulatory circuit that discriminates between transient and persistent TLR4-induced signals. Nat Immunol 10, 437–443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Love MI, Huber W, and Anders S (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology 15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lugowski A, Nicholson B, and Rissland OS (2018). Determining mRNA half-lives on a transcriptome-wide scale. Methods 137, 90–98. [DOI] [PubMed] [Google Scholar]
- Mariani L; Schulz EG; Lexberg MH; Helmstetter C; Radbruch A; Lohning M; and Hofer T (2010). Short term memory in gene induction reveals the regulatory principle behind stochastic IL-4 expression. Mol Syst Biol, 13, 1–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagashima T, Inoue N, Yumoto N, Saeki Y, Magi S, Volinsky N, Sorkin A, Kholodenko BN, and Okada-Hatakeyama M (2015). Feedforward regulation of mRNA stability by prolonged extracellular signal-regulated kinase activity. Febs J 282, 613–629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nau GJ, Richmond JFL, Schlesinger A, Jennings EG, Lander ES, and Young RA (2002). Human macrophage activation programs induced by bacterial pathogens. Proceedings of the National Academy of Sciences 99, 1503–1508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nelson DE, Ihekwaba AE, Elliott M, Johnson JR, Gibney CA, Foreman BE, Nelson G, See V, Horton CA, Spiller DG, et al. (2004). Oscillations in NF-kappaB signaling control the dynamics of gene expression. Science 306, 704–708. [DOI] [PubMed] [Google Scholar]
- Novershtern N, Subramanian A, Lawton LN, Mak RH, Haining WN, McConkey ME, Habib N, Yosef N, Chang CY, Shay T, et al. (2011). Densely interconnected transcriptional circuits control cell states in human hematopoiesis. Cell 144, 296–309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ostuni R, Piccolo V, Barozzi I, Polletti S, Termanini A, Bonifacio S, Curina A, Prosperini E, Ghisletti S, and Natoli G (2013). Latent Enhancers Activated by Stimulation in Differentiated Cells. Cell 152, 157–171. [DOI] [PubMed] [Google Scholar]
- Porter JR, Fisher BE, and Batchelor E (2016). p53 Pulses Diversify Target Gene Expression Dynamics in an mRNA Half-Life-Dependent Manner and Delineate Co-regulated Target Gene Subnetworks. Cell Syst 2, 272–282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Purvis JE, and Lahav G (2013). Encoding and Decoding Cellular Information through Signaling Dynamics. Cell 152, 945–956. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rabani M, Levin JZ, Fan L, Adiconis X, Raychowdhury R, Garber M, Gnirke A, Nusbaum C, Hacohen N, Friedman N, et al. (2011). Metabolic labeling of RNA uncovers principles of RNA production and degradation dynamics in mammalian cells. Nat Biotechnol 29, 436–442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramirez-Carrozzi VR, Braas D, Bhatt DM, Cheng CS, Hong C, Doty KR, Black JC, Hoffmann A, Carey M, and Smale ST (2009). A Unifying Model for the Selective Regulation of Inducible Transcription by CpG Islands and Nucleosome Remodeling. Cell 138, 114–128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramsey SA, Klemm SL, Zak DE, Kennedy KA, Thorsson V, Li B, Gilchrist M, Gold ES, Johnson CD, Litvak V, et al. (2008). Uncovering a macrophage transcriptional program by integrating evidence from motif scanning and expression dynamics. Plos Comput Biol 4, e1000021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ravasi T, Suzuki H, Cannistraci CV, Katayama S, Bajic VB, Tan K, Akalin A, Schmeier S, Kanamori-Katayama M, Bertin N, et al. (2010). An atlas of combinatorial transcriptional regulation in mouse and man. Cell 140, 744–752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shalem O, Dahan O, Levo M, Martinez MR, Furman I, Segal E, and Pilpel Y (2008). Transient transcriptional responses to stress are generated by opposing effects of mRNA production and degradation. Mol Syst Biol 4, 223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tong AJ, Liu X, Thomas BJ, Lissner MM, Baker MR, Senagolage MD, Allred AL, Barish GD, and Smale ST (2016). A Stringent Systems Approach Uncovers Gene-Specific Mechanisms Regulating Inflammation. Cell 165, 165–179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uhlitz F, Sieber A, Wyler E, Fritsche-Guenther R, Meisig J, Landthaler M, Klinger B, and Bluthgen N (2017). An immediate-late gene expression module decodes ERK signal duration. Mol Syst Biol 13, 928. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Werner SL, Barken D, and Hoffmann A (2005). Stimulus specificity of gene expression programs determined by temporal control of IKK activity. Science 309, 1857–1861. [DOI] [PubMed] [Google Scholar]
- Yang E, van Nimwegen E, Zavolan M, Rajewsky N, Schroeder M, Magnasco M, and Darnell JE (2003). Decay rates of human mRNAs: correlation with functional characteristics and sequence attributes. Genome Res 13, 1863–1872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, Nussbaum C, Myers RM, Brown M, Li W, et al. (2008). Model-based Analysis of ChIP-Seq (MACS). Genome Biology 9, R137. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Table S1: RPKM values for the time points in ifnar−/−; nominal and FDR corrected p values of TNF, LPS, IL1-induced genes; and scaled expression values of RNA-seq in ifnar−/−. Related to Fig. 1.
Table S2: RPKM of RNAseq in ifnar−/− (control) and ifnar−/−nfkbia−/− (mutant); specificity scores and p values for control and mutant replicates (one-sided t- test); and scaled expression values of RNA-seq in ifnar−/− (control) and ifnar−/−nfkbia−/− (mutant) used for heatmap and math modeling. Related to Fig. 2.
Table S3: Log2 fold change for actD RNA-seq and derivated mRNA half-life; normalized counts for actD RNA-seq; and mRNA degradation rates regression for stimulus-specific genes. Related to Fig. 3.
Table S4: Values of parameters for Model v1. Related to Fig. 4.
Table S5: RPKM and scaled expression values of caRNA for the time points in control and mutant. Related to Fig. 5.
Table S6: Values of parameters for Model v2. Related to Fig. 6.
Data Availability Statement
All sequencing data were deposited to Sequence Read Archive (SRA, NCBI) under BioProject IDs PRJNA453806 (polyA+ RNA-seq), PRJNA454496 (ActD RNA-seq), PRJNA454896 (ca-RNA seq), PRJNA517534 (RelA-ChIPseq) and SAMN12325160, SAMN12325163, SAMN12325167, SAMN12325170 (ATAC-seq). All the code necessary for reproducing the figures, model fitting and simulation is available on the GitHub site mentioned in the STAR methods.