Abstract
The task of drug-target interaction prediction holds significant importance in pharmacology and therapeutic drug design. In this paper, we present FRnet-DTI, an auto-encoder based feature manipulation and a convolutional neural network based classifier for drug target interaction prediction. Two convolutional neural networks are proposed: FRnet-Encode and FRnet-Predict. Here, one model is used for feature manipulation and the other one for classification. Using the first method FRnet-Encode, we generate 4096 features for each of the instances in each of the datasets and use the second method, FRnet-Predict, to identify interaction probability employing those features. We have tested our method on four gold standard datasets extensively used by other researchers. Experimental results shows that our method significantly improves over the state-of-the-art method on three out of four drug-target interaction gold standard datasets on both area under curve for Receiver Operating Characteristic (auROC) and area under Precision Recall curve (auPR) metric. We also introduce twenty new potential drug-target pairs for interaction based on high prediction scores. The source codes and implementation details of our methods are available from https://github.com/farshidrayhanuiu/FRnet-DTI/ and also readily available to use as an web application from http://farshidrayhan.pythonanywhere.com/FRnet-DTI/.
Keywords: Bioinformatics, Computer Science, Drug-Target, Class imbalance, Feature engineering, Ensemble classifier, Classification
Bioinformatics; Computer Science; Drug-Target; Class imbalance; Feature engineering; Ensemble classifier; Classification
1. Introduction
The task of drug-target interaction prediction is very important in pharmacology and therapeutic drug design. This problem can be addressed in several ways. Firstly, for an already developed drug compound the task is to find new targets with which the drug might have interactions. Secondly, for a given target protein one might search for potential drugs in the library. Another way to tackle the problem is to find the possibility of interaction given a pair of drug and target protein. In this paper, we are interested in the latter kind. Experimental methods in predicting drug-protein interactions are expensive and time consuming and hence computational methods have been used extensively in the recent years [1], [2].
One of the most successful computational method in drug-target interaction prediction is docking simulations [3]. This method largely depends on the availability of three dimensional native structure of the target protein determined by sophisticated methods like X-Ray Crystallography. However, X-Ray Crystallography is itself a time-consuming and expensive process and thus the native structure of the targets proteins are often unavailable. These have encouraged the researchers to apply machine learning based methods to tackle the prediction problem by formulating it in a supervised learning setting [4], [5].
Success of supervised learning methods largely depend on the training datasets. In a pioneering work on drug-target interaction prediction, Yamanishi et al. [6] proposed gold standard datasets with four sets or target proteins and drugs. A good number of machine learning algorithms have been used in the literature of supervised drug-target interaction prediction that includes: Support Vector Machines (SVM) [7], Boosting [8], [9], [10], Deep Learning [11], [12], etc. One of the major obstacles in drug-target interaction prediction is due to the imbalance in the dataset. Since the known validated interactions among drug target pairs are not large, most of the approaches considers the unknown interactions as negative samples and thus they outnumber positive samples. The representation of the drug-target pair in the supervised learning dataset is another added challenge. Machine learning methods used in prediction of drug-target interaction often use features generated from molecular fingerprints of drugs and sequence or structure based information of proteins [7], [8].
In the recent years, Chemo-genomic methods have received a lot of attention for identifying drug target interaction. They usually include methods like graph theory [13], [14], deep learning [11], machine learning [6], [15] and network analysis methods [16], [17]. In supervised learning setting, K-Nearest Neighbor (KNN) [18], fuzzy logic [19], support vector machines [7], [20] are the most commonly used classification algorithm. In [6], drug target interaction problem was first introduced as a supervised problem and a gold standard dataset was proposed. Later those datasets have been exhaustively used by researchers. The same authors from [6], applied distance based learning to association among pharmacological space of drug target interactions. A non-linear kernel fusion with regularized least square method was proposed by [21].
In [22], Chemical and genomic kernels and Bayesian factorization was applied. Another method, DBSI (drug based similarity interface) [17] proposed two dimensional chemical-structural similarity for drug similarity. Later methods like DASPfind [23], NetCBP [24], SELF-BLM [20] were proposed in order to solve the same problem. Bigram based features as fingerprints extracted from position specific scoring matrix were found very helpful solving the drug target interaction problem [7]. Most of the supervised learning methods do not exploit the structure based features because most protein targets' three dimensional native structure are not available.
In [25], the authors have used extremely randomized trees as classifier. They represented the drugs as molecular fingerprint and proteins as pseudo substitution matrix. These matrix were generated from its amino acid sequence information. Other relevant works include self-organizing theory [26], [27], similarity based methods [28], network and chemoinformatics based tools [29], ensemble methods [30], [31]. A in depth literature review was provided by Chen et al. on computational methods for this particular problem [32].
Wen et al. [33] presented a model which consisted of multiple stacked RBM. The output layer consisted of two neurons each predicting the interaction and non-interaction probability respectively. Chan et al. [11] presented a model that used deep representations for drug target interaction predictions. There were also many other attempts made to use deep learning tools and boosting to make further improvements in DTI [10]. In our recent work, iDTI-ESBoost [8], we exploited evolutionary features along with structural features to predict drug protein interaction. SPIDER3, a successful secondary structural prediction tool [34], [35], was used to generate a novel set of features for supervised learning. The novel set of features includes seven primary set of features. A short description of each feature set is given in Table 1. In that paper, two balancing methods were used to handle the imbalance ratio of the datasets, and Adaboost [36] was used for classification. In [29], authors presented a system of deep auto-encoders to further improve drug target pair interaction prediction.
Table 1.
A short summary of structural and evolutionary features used for fingerprint features and protein targets for drugs. Each Group column shows a different feature group, used in this experiment and they are discussed in a later sections.
| Feature group | Number of features | Reference |
|---|---|---|
| Molecular finger print | 881 | [7] |
| PSSM bigram | 400 | [7] |
| Secondary Structure Composition | 3 | [8] |
| Accessible Surface Area Composition | 1 | [8] |
| Torsional Angles Composition | 8 | [8] |
| Torsional Angles Auto-Covariance | 80 | [8] |
| Structural Probabilities Auto-Covariance | 30 | [8] |
| Torsional Angles bigram | 64 | [8] |
| Structural Probabilities bigram | 9 | [8] |
| Total | 1476 | |
In this paper, we propose two deep convolutional methods for feature manipulation and predicting drug target interaction: FRnet-Encode and FRnet-Predict. FRnet-Encode is used to generate 4096 features or deep representation of each dataset and FRnet-Predict is used for classification using the extracted features. We use the latest version of 4 gold standard datasets with 1476 features to test our method. In the experimental results combining both methods, we have observed improved auROC and auPR metric scores and therefore we strongly claim that our method is an excellent alternative for most other proposed methods for Drug-Target-Interaction (DTI).
2. Methodology
This section provides a description of the methodology used in this paper: algorithmic details, datasets and performance evaluation methods.
2.1. Convolutional models
In this paper, we propose two novel deep learning architectures for drug target interaction, each network having its own purpose and goal. Throughout the rest of the paper these two models are referred as FRnet-Encode and FRnet-Predict. FRnet-Encode is used as a auto encoder that extracts 4096 features from the given feature sets which is than fed as input to FRnet-Predict for classification.
The proposed models FRnet-Encode and FRnet-Predict both have over millions of hyper-parameter to tune. While exhaustively tuning each of the parameter would provide the best result it is seldom done as it requires tremendous computational cost and time and risks over-fit. Four of the most effective hyper parameters are chosen [37] to tune and from a given range. Aggressive regularization is also employed to remove over-fitting to maximum extend.
2.2. Rational for FRnet-DTI
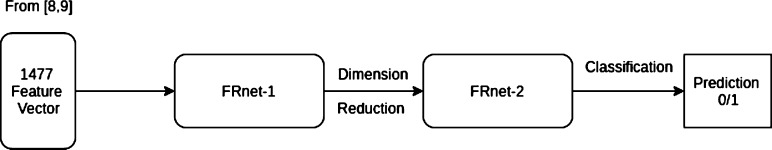
Recently, Deep learning methods have been receiving a lot of attention for biological applications. In [38], the authors used a model called Wide-and-Deep. Wen et al. [33] showed impressive improvement in prediction capability using deep learning. The authors of [39] used stacked auto-encoders to further improve the problem of DTI. We closely follow that article and employ an auto encoder to extract features and a classifier for final classification. We follow the architectural design of GoogleNet [40] which is regarded as one of the most successful classification network [41], [42]. They use a module called the inception module (see Fig. 3) where convolution operation with different filters sizes are done in parallel with a pooling operation. Each of the output are then merged together as the final output of the module. This process reliefs the burden of choosing of proper filter size or operation type between convolution and pooling. Following this intuition, we design our two models. Top view of the proposed system is shown in Fig. 5 where the 1477 feature are generated using [8], [9].
Figure 3.
Figure 5.
Top view of the proposed system.
2.2.1. FRnet-Encode
In order to perform a convolution operation, a 4D tensor with the shape is required [43] where are the 3d representation of the features and X is the input batch size. Molecular fingerprint based drug features and structure and evolutionary features for proteins extracted for the gold standard datasets showed very optimistic results in several recent works [7], [8], [44]. For convenience, a value 0 was added which extended the feature length to 1476 so that the input can be reshaped into . Here 211 and 7 are unique numbers and interchanging them has no effect and 1 represents that the input has only 1 channel. Basically the dataset is represented as a gray scale sized image.
This model consists of several convolutional layers, max-Pooling layer and fully-connected layer. Fig. 1 shows the visual representation of the complete network. Input layer takes the input in the shape of and passes to a Convolutional layer with 32 filters and 2 strides which outputs a tensor with the shape . Here convolution means the size of the filter were . ‘Relu’ activation, ‘SAME’ padding and ‘L2’ regularizer were used in each layer. After convolution, a Max-Pool operation is done with a kernel and stride value of 2 to reduce the tensor shape to . This network output is then fed to four parallel processes. They are depicted in Fig. 1 from left to right. The first process is a convolution with 8 filters followed by a convolution with 64 filter. Next is a convolution with 8 filters followed by a convolution with 64 filters. Then, there is a convolution with 8 filters followed by a convolution with 32 filters and lastly, a Max-pool operation with kernel and stride 1. In the next stage, a merge operator combines the 4 network outputs on the 3rd axis of the resulting tensor with the shape . Detailed description is provided in Table 2. The merged network is then fed to fully connected layers with 4096 and 2048 neurons followed by a dropout operation with value at 0.5. Finally the output layer consists of 1476 neurons each neuron representing a feature value of an instance in the dataset. The model was trained with a learning rate 0.001, ‘’ as optimizer [45] and as loss function. The fully connected layer with 4096 neurons is used as features to predict interaction probability using the FRnet-Predict method. The model achieved 85% accuracy just after 3 iterations and reached over 90% after 20 iterations. Accuracy curves with respect to each iteration are described in Fig. 2.
Figure 1.
Architecture of FRnet-Encode.
Table 2.
Shapes of tensor after each convolution operation leading up to the merge operation. Also know as Inception Operation.
| Index | Input shape | Output shape after first convolution | Output shape after second convolution/Max-pool |
|---|---|---|---|
| a | X, 53, 2, 32 | X, 53, 2, 16 | X, 53, 2, 64 |
| b | X, 53, 2, 32 | X, 53, 2, 16 | X, 53, 2, 64 |
| c | X, 53, 2, 32 | X, 53, 2, 16 | X, 53, 2, 32 |
| d | X, 53, 2, 32 | – | X, 53, 2, 32 |
| Final Tensor shape | (X, 53, 2, 192) |
Figure 2.
Accuracy curves of FRnet-Encodeusing Binary crossentropy as loss function on four datasets: (a) enzymes (b) ion channels (ic) (c) GPCRs (d) nuclear receptors (nr).
The merge operation is used to retire the burden of choosing filter size from , and . In stead, the model exploits all of them and chooses the better set of features by itself. This concept was inspired form the model [40] which was later used to build the various versions of imageNet by the same authors [41], [42]. However, this model also increases computational complexity as different sized filters are used at the same time. In order to reduce some computational cost, FRnet-Encode employs convolution before and after each convolution operation with different filter size to reduce computational complexity of the model. This concept was first introduced in 2013 in the article by [46]. They showed that convolutional operations can be used as tool to reduce channel size of a tensor. The hypothesis of [46] states that converting a tensor form shape to will cost much more than converting to using a convolution using 16 filters than converting to and have the same effect on the network. This same methodology is later incorporated in FRnet-Predict also.
2.2.2. FRnet-Predict
FRnet-Predict serves for the purpose of classifying interaction probability between a given drug-target pair. Similar to FRnet-Encode this model employs the inception module (details figure of the model is given in Fig. 3). It uses the 4096 features generated by FRnet-Encode as a shaped instance. In this model, the first convolution and Max-Pool operation is kept the same as the previous method. Following those operations, the tensors are parallelly fed to 2 inception modules, one with stride size of 1 (left module of Fig. 4) and another with stride size 2.
Figure 4.
Architecture of FRnet-Predict.
The model merges those outputs in order to take befit of both stride size and put it through a final inception layer before connecting in to fully connected layers of 2048, 512 and finally 1 neuron for prediction. Similar to FRnet-Encode this model also uses L2 regularization, ‘’ as optimizer with as loss function with learning rate set to 0.001 and value set to 0.5 in the dropout layer.
2.3. Datasets
The benchmark datasets used in this article were first introduced by [6] in 2008 using DrugBank [47], KEGG [48], BRENDA [49] and SuperTarget [50] to extract information about drug-target interactions. These datasets are regarded as and have been exhaustively been used by researchers throughout the years [7], [8], [11], [32]. These datasets are publicly available at: http://web.kuicr.kyoto-u.ac.jp/supp/yoshi/drugtarget/.
In this paper, an extended version of those datasets is used which consists of structural and evolutionary features. This version of dataset was first introduced in 2016 by [7] and later further extended by [8], [9]. The exacted dataset from [8] were used in this project for experimentation. A short description of each dataset is given in Table 3.
Table 3.
Description of the gold standard datasets with structural and evolutionary features [8].
| Dataset | Drugs | Proteins | Positive interactions | Imbalance ratio |
|---|---|---|---|---|
| Enzyme | 445 | 664 | 2926 | 99.98 |
| Ion Chanel | 210 | 204 | 1476 | 28.02 |
| GPCR | 223 | 95 | 635 | 32.36 |
| Nuclear Receptor | 54 | 26 | 90 | 14.6 |
2.4. Performance evaluation
A wide variety of performance metrics are available to show case and compare performance of classification models. Even though the metric is sufficient enough to show the accuracy percentage of a model, in highly imbalanced dataset, such as gold standard datasets used in this experiment, that value holds little to no significance. In imbalance binary datasets, one class highly outnumbers samples of other class thus a measure of accuracy in this case makes little sense.
There are two metrics, who are independent from the imbalance ratio of the datasets, called area under curve for Receiver Operating Characteristic (auROC) and area under Precision Recall curve (auPR). Due to their ignorance towards the imbalance ratio of datasets, they have been widely used [8], [9], [11], [24], [51] as standard metric for comparison. Both metrics value range from 0 to 1 where a random classifier should have a score of 0.5 and a perfect classification model will have a auPR and auROC score of 1. In both cases the higher the value the better.
Another important factor is the balance of bias and variance trade off [52]. k-fold cross validation and jack knife tests are mostly used as an attempt to solve the bias-variance problem. In our experiment, we used 5-fold shuffled cross validation on each dataset. Each time the dataset is shuffled and spitted into 5 equal parts. Then 4 of them are used for training and the rest for testing.
3. Results and discussion
In the experiments reported in this paper, Python v3.6, Library and Sci-kit learn [53] were used for the implementation. Each experiments were executed 10 times and the average result was considered. Each dataset were split into two sets, train set and test set using 5 fold cross validation, thus for each fold we used 80% of each dataset for training and used the 20% for testing.
FRnet-Encode method is multilayer deep auto-encoder that uses convolution, max-pool and fully connected layers to regenerate the input as output in the final fully connected layer. For each of the datasets, the model was trained to achieve accuracy over of 95%. Due to the use of aggressive regularization, value of 0.5 in dropout and L2 regularization in each layer using learning rate of 0.001 to avoid over fitting, the models were unable to achieve accuracy higher than 97% for any of the datasets. The first fully connected layer in the network has 4096 neurons in the output and those were used to extract 4096 features from each dataset. For a fair sake of comparison, FRnet-Predict was tested with several states of the art machine learning algorithms like, Decision Tree [54], SVM [55], MEBoost [44] and CUSBoost [56]. Each of these classifiers were fed the 4096 features generated by FRnet-Encode. Results in terms of auPR and auROC are given Table 4. Table 4 shows that FRnet-Predict is able to produce results with better auROC for all the datasets. However, in terms of auPR the results in three datasets are better than the competitor algorithms. However, for the ‘enzymes’ dataset, the performance of FRnet-Predict is very close to the best performing CUSBoost. Note that other classifiers also achieved very impressive auROC and auPR score which shows the effectiveness of the features generated by FRnet-Encode. Therefore even though FRnet-Encode is designed for feature manipulation of the four datasets mentioned in this article, it can be used as a strong feature manipulation tool on other domains as well [8], [9].
Table 4.
A comparison of performances among FRnet-Predict and other classifiers on the gold standard datasets in terms of auROC and auPR using 4096 features generated by FRnet-Encode.
| Dataset | Classifier | auPR | auROC |
|---|---|---|---|
| enzymes | Decision Tree | 0.28 | 0.9376 |
| SVM | 0.53 | 0.9010 | |
| MEBoost | 0.41 | 0.9404 | |
| CUSBoost | 0.71 | 0.9345 | |
| FRnet-Predict | 0.70 | 0.9754 | |
| GPCR | Decision Tree | 0.31 | 0.9038 |
| SVM | 0.44 | 0.8859 | |
| MEBoost | 0.46 | 0.9075 | |
| CUSBoost | 0.65 | 0.8989 | |
| FRnet-Predict | 0.69 | 0.9512 | |
| Ion Channel | Decision Tree | 0.29 | 0.933 |
| SVM | 0.40 | 0.8904 | |
| MEBoost | 0.39 | 0.928 | |
| CUSBoost | 0.45 | 0.8851 | |
| FRnet-Predict | 0.49 | 0.9478 | |
| NR | Decision Tree | 0.46 | 0.8147 |
| SVM | 0.41 | 0.7605 | |
| MEBoost | 0.23 | 0.9165 | |
| CUSBoost | 0.71 | 0.8989 | |
| FRnet-Predict | 0.73 | 0.9241 | |
We have compared our method with other state of the art classifiers mentioned in recent literatures such as SVM, AdaBoost and random forest. FRnet-Predict shows superior performance in both metric on all the datasets except for NR which holds only 1048 instances and is the smallest dataset among the others. Comparison with other classification models are shown in Table 5. Results for the other methods were taken from the experiments reported in the literature [7], [8], [9]. Note that, for each of the datasets except the nuclear receptor (NR) dataset, performance of FRnet-Predict is superior to the other methods both in terms of auPR and auROC. For the NR dataset, the performance of FRnet-Predict is almost similar to the best performing boosting classifier. The auPR value is second best and probably because of the fact that this dataset is highly clustered and clustered sampling techniques for balancing used in [8] makes it perform better in this particular case.
Table 5.
A performance comparison among FRnet-Predict with AdaBoost, Support Vector Machine and Random Forest classifiers on the gold standard datasets auROC and auPR curve.
| Dataset | Reference | Classifier | auPR | auROC |
|---|---|---|---|---|
| enzymes | [8] | AdaBoost | 0.68 | 0.9689 |
| [8] | Random Forest | 0.43 | 0.9457 | |
| [7] | SVM | 0.54 | 0.9194 | |
| FRnet-Predict | 0.70 | 0.9754 | ||
| GPCR | [8] | AdaBoost | 0.31 | 0.9128 |
| [8] | Random Forest | 0.30 | 0.9168 | |
| [7] | SVM | 0.28 | 0.8720 | |
| FRnet-Predict | 0.69 | 0.9512 | ||
| Ion Channel | [8] | AdaBoost | 0.48 | 0.9369 |
| [8] | Random Forest | 0.40 | 0.9234 | |
| [7] | SVM | 0.39 | 0.8890 | |
| FRnet-Predict | 0.49 | 0.9512 | ||
| NR | [8] | AdaBoost | 0.79 | 0.9285 |
| [8] | Random Forest | 0.29 | 0.7723 | |
| [7] | SVM | 0.41 | 0.8690 | |
| FRnet-Predict | 0.73 | 0.9241 | ||
We have also compared our results against methods which used unsupervised and semi-supervised methods reported in the literature [6], [13], [17], [22], [24], [57]. Table 6 shows comparisons of auROC scores of other methods including supervised methods [7], [8]. Note that, our proposed method achieves significantly higher auROC for three datasets among four and for the NR dataset, the performance is only second best and very close to the best performing one. Although the performance of our model is not better than the models of [5] it is superior to other recent models based on the given result which makes it a competitive substitute.
Table 6.
Performance of FRnet-Predict on the four benchmark gold datasets in terms of auROC with comparison to other state-of-the-art methods.'N/A' denotes where that particular dataset was not used in the article.
| Method | Dataset |
|||
|---|---|---|---|---|
| Enzyme | GPCR | ion channels | nuclear receptor | |
| Yamanishi et al. [6] | 0.904 | 0.8510 | 0.8990 | 0.8430 |
| Yamanishi et al. [57] | 0.8920 | 0.8120 | 0.8270 | 0.8350 |
| DBSI [17] | 0.8075 | 0.8029 | 0.8022 | 0.7578 |
| KBMF2K [22] | 0.8320 | 0.7990 | 0.8570 | 0.8240 |
| NetCBP [24] | 0.8251 | .8034 | 0.8235 | 0.8394 |
| Wang et al. [13] | 0.8860 | 0.8930 | 0.8730 | 0.8240 |
| Mutowo et al. [58] | 0.9480 | 0.8990 | 0.8720 | 0.8690 |
| iDTI-ESBoost [8] | 0.9689 | 0.9369 | 0.9222 | 0.9285 |
| Wang et al. [29] | 0.9425 | 0.8743 | 0.9107 | 0.8176 |
| CFSBoost [9] | 0.9563 | 0.9377 | 0.9278 | 0.8147 |
| Our Method | 0.9754 | 0.9478 | 0.9512 | 0.9241 |
These two methods works well together because we use the first network to represent the 1476 features as a grey scale image and reduce the dimension of the feature vector to a smaller scale. This eases the difficulty of the problem. Then we use another deep network to use those feature vector for prediction. Further improvements can be made by making the networks more deeper but at the cost of more computational expense. We also have considered the fact that making the network more deeper may cause it to over-fit the data-set, therefore a dataset of larger scale will also be required.
In the literature of imbalanced classification problems, it has been often argued that between area under Precision Recall curve (auPR) and area under Receiver Operating Curve (auROC), auPR should be considered more significant. However, only [7] and [8] reported auPR scores in their paper. A comparison in terms of auPR score are given in Table 7. Here too, its interesting to note the superior performance of our proposed model on all the datasets.
Table 7.
Comparison of the performance of FRnet-Predict on the four benchmark gold datasets from [8] in terms of auPR with other the state-of-the-art methods.
We have also tested our method with input shape instead of and found similar results which concludes that changing the input shape has little to no effect on the performance on the models. Results using input shape is provided in Table 8.
Table 8.
Performance comparison of FRnet-Predict and other classifiers on the gold standard datasets in terms of auROC and auPR using 4096 features generated by FRnet-Encode with input shape (X, 7, 211, 1).
| Dataset | Classifier | auPR | auROC |
|---|---|---|---|
| enzymes | Decision Tree | 0.27 | 0.9299 |
| SVM | 0.54 | 0.9035 | |
| FRnet-Predict | 0.70 | 0.9713 | |
| GPCR | Decision Tree | 0.32 | 0.9038 |
| SVM | 0.48 | 0.8859 | |
| FRnet-Predict | 0.70 | 0.9255 | |
| Ion Channel | Decision Tree | 0.60 | 0.9235 |
| SVM | 0.52 | 0.8894 | |
| FRnet-Predict | 0.50 | 0.9507 | |
| NR | Decision Tree | 0.43 | 0.8207 |
| SVM | 0.42 | 0.7588 | |
| FRnet-Predict | 0.62 | 0.9134 | |
Since the prediction scores with high confidence are interesting in practical applications, A list of top five the false positive interactions based on FRnet-Predict's prediction score is given in Table 9. These are the interactions that are known as not interacting pair but the model highly suggests other wise.
Table 9.
New prediction made by FRnet-Predict for four gold standard datasets used in this paper.
| Dataset | Protein Id | Drug Id | Drug name | Score |
|---|---|---|---|---|
| enzymes | hsa:10825 | D00041 | Threonine (USP) | 0.8351 |
| hsa:4759 | D00041 | Threonine (USP) | 0.8321 | |
| hsa:129807 | D00041 | Threonine (USP) | 0.8255 | |
| hsa:4953 | D00041 | Threonine (USP) | 0.8048 | |
| hsa:1845 | D00041 | Threonine (USP) | 0.7923 | |
| ion channels | hsa:285242 | D00294 | Diazoxide (JAN/USP/INN) | 0.9823 |
| hsa:779 | D00294 | Diazoxide (JAN/USP/INN) | 0.9712 | |
| hsa:2561 | D00294 | Diazoxide (JAN/USP/INN) | 0.9723 | |
| hsa:785 | D00294 | Diazoxide (JAN/USP/INN) | 0.9634 | |
| hsa:11254 | D00294 | Diazoxide (JAN/USP/INN) | 0.9565 | |
| GPCRs | hsas:9052 | D04625 | Isoetharine (USP) | 0.9013 |
| hsa:9052 | D00632 | Dobutamine hydrochloride (JP17/USP) | 0.9013 | |
| hsa:9052 | D03880 | Dobutamine lactobionate (USAN) | 0.8912 | |
| hsa:9052 | D03881 | Dobutamine tartrate (USP) | 0.8904 | |
| hsa:1909 | D03621 | Cyclizine (INN) | 0.8898 | |
| nuclear receptors | hsa:2099 | D01132 | Tazarotene (JAN/USAN/INN) | 0.9912 |
| hsa:2101 | D00956 | Nandrolone phenpropionate (USP) | 0.9876 | |
| hsa:2101 | D00443 | Spironolactone (JP17/USP/INN) | 0.9885 | |
| hsa:2099 | D00316 | Etretinate (JAN/USAN/INN) | 0.9472 | |
| hsa:9971 | D00316 | Etretinate (JAN/USAN/INN) | 0.9102 | |
4. Conclusion
In This paper, we propose two novel deep neural net architectures, FRnet-Encode and FRnet-Predict where FRnet-Encode aims to extract convolutional features and FRnet-Predict tries to identify drug target interaction using the extracted features. From [8], we exploit our algorithm with datasets consisting with both structural and evolutionary features and with the help of FRnet-Encode we try to generate 4096 informative features. These datasets are regarded as gold standard datasets and are exhaustively used by researchers. We have conducted extensive experiments and produced the results in term of auROC and auPR scores. In many previous literatures like [7], [8], it was argued that in case of drug target interaction, it is more appropriate to use auPR metric over auROC as the gold standard datasets are highly imbalanced with very few interaction samples. For this reason, FRnet-Predict was focused on getting a superior auPR score even by sacrificing auROC score. We also proposed 5 new possible interaction pair for each of the 4 datasets based on prediction score. Up to this moment, our proposed method outperforms other state of that art methods in 3 of the 4 benchmark datasets in auPR and auROC metric. We believe the excellent performance our method will motivate other practitioners and researchers to exploit both methods for not only drug target interaction but also in other domains.
Declarations
Author contribution statement
Author Contribution Statement F. Rayhan: Conceived and designed the experiments; Performed the experiments; Analyzed and interpreted the data; Contributed reagents, materials, analysis tools or data; Wrote the paper.
S. Ahmed, Z. Mousavian, D. M. Farid, S. Shatabda: Analyzed and interpreted the data; Contributed reagents, materials, analysis tools or data; Wrote the paper.
Funding statement
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Competing interest statement
The authors declare no conflict of interest.
Additional information
No additional information is available for this paper.
References
- 1.Haggarty S.J., Koeller K.M., Wong J.C., Butcher R.A., Schreiber S.L. Multidimensional chemical genetic analysis of diversity-oriented synthesis-derived deacetylase inhibitors using cell-based assays. Chem. Biol. 2003;10(5):383–396. doi: 10.1016/s1074-5521(03)00095-4. [DOI] [PubMed] [Google Scholar]
- 2.Kuruvilla F.G., Shamji A.F., Sternson S.M., Hergenrother P.J., Schreiber S.L. Dissecting glucose signalling with diversity-oriented synthesis and small-molecule microarrays. Nature. 2002;416(6881):653–657. doi: 10.1038/416653a. [DOI] [PubMed] [Google Scholar]
- 3.Kitchen D.B., Decornez H., Furr J.R., Bajorath J. Docking and scoring in virtual screening for drug discovery: methods and applications. Nat. Rev. Drug Discov. 2004;3(11):935–949. doi: 10.1038/nrd1549. [DOI] [PubMed] [Google Scholar]
- 4.Mousavian Z., Masoudi-Nejad A. Drug–target interaction prediction via chemogenomic space: learning-based methods. Expert Opinion on Drug Metabolism & Toxicology. 2014;10(9):1273–1287. doi: 10.1517/17425255.2014.950222. [DOI] [PubMed] [Google Scholar]
- 5.Olayan R.S., Ashoor H., Bajic V.B. Ddr: efficient computational method to predict drug–target interactions using graph mining and machine learning approaches. Bioinformatics. 2017;34(7):1164–1173. doi: 10.1093/bioinformatics/btx731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Yamanishi Y., Araki M., Gutteridge A., Honda W., Kanehisa M. Prediction of drug–target interaction networks from the integration of chemical and genomic spaces. Bioinformatics. 2008;24(13) doi: 10.1093/bioinformatics/btn162. i232–i240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Mousavian Z., Khakabimamaghani S., Kavousi K., Masoudi-Nejad A. Drug–target interaction prediction from pssm based evolutionary information. J. Pharmacol. Toxicol. Methods. 2016;78:42–51. doi: 10.1016/j.vascn.2015.11.002. [DOI] [PubMed] [Google Scholar]
- 8.Rayhan F., Ahmed S., Shatabda S., Farid D.M., Mousavian Z., Dehzangi A., Rahman M.S. idti-esboost: identification of drug target interaction using evolutionary and structural features with boosting. Sci. Rep. 2017;7(1) doi: 10.1038/s41598-017-18025-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Rayhan F., Ahmed S., Farid D.M., Dehzangi A., Shatabda S. Cfsboost: cumulative feature subspace boosting for drug-target interaction prediction. J. Theor. Biol. 2019;464:1–8. doi: 10.1016/j.jtbi.2018.12.024. [DOI] [PubMed] [Google Scholar]
- 10.Tian K., Shao M., Wang Y., Guan J., Zhou S. Boosting compound-protein interaction prediction by deep learning. Methods. 2016;110:64–72. doi: 10.1016/j.ymeth.2016.06.024. [DOI] [PubMed] [Google Scholar]
- 11.Chan K.C., You Z.-H. Large-scale prediction of drug-target interactions from deep representations. 2016 International Joint Conference on Neural Networks; IJCNN, IEEE; IEEE; 2016. pp. 1236–1243. [Google Scholar]
- 12.Rayhan F. Fr-mrinet: a deep convolutional encoder-decoder for brain tumor segmentation with relu-RGB and sliding-window. Int. J. Comput. Appl. 2019;975:8887. [Google Scholar]
- 13.Wang W., Yang S., Li J. Drug target predictions based on heterogeneous graph inference. Pacific Symposium on Biocomputing. Pacific Symposium on Biocomputing; NIH Public Access; NIH Public Access; 2013. p. 53. [PMC free article] [PubMed] [Google Scholar]
- 14.Chen X., Liu M.-X., Yan G.-Y. Drug–target interaction prediction by random walk on the heterogeneous network. Mol. BioSyst. 2012;8(7):1970–1978. doi: 10.1039/c2mb00002d. [DOI] [PubMed] [Google Scholar]
- 15.Bleakley K., Yamanishi Y. Supervised prediction of drug–target interactions using bipartite local models. Bioinformatics. 2009;25(18):2397–2403. doi: 10.1093/bioinformatics/btp433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Alaimo S., Pulvirenti A., Giugno R., Ferro A. Drug–target interaction prediction through domain-tuned network-based inference. Bioinformatics. 2013;29(16):2004–2008. doi: 10.1093/bioinformatics/btt307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Cheng F., Liu C., Jiang J., Lu W., Li W., Liu G., Zhou W., Huang J., Tang Y. Prediction of drug-target interactions and drug repositioning via network-based inference. PLoS Comput. Biol. 2012;8(5) doi: 10.1371/journal.pcbi.1002503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.He Z., Zhang J., Shi X.-H., Hu L.-L., Kong X., Cai Y.-D., Chou K.-C. Predicting drug-target interaction networks based on functional groups and biological features. PLoS ONE. 2010;5(3) doi: 10.1371/journal.pone.0009603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Xiao X., Min J.-L., Wang P., Chou K.-C. icdi-psefpt: identify the channel–drug interaction in cellular networking with pseaac and molecular fingerprints. J. Theor. Biol. 2013;337:71–79. doi: 10.1016/j.jtbi.2013.08.013. [DOI] [PubMed] [Google Scholar]
- 20.Keum J., Nam Self-blm H. Prediction of drug-target interactions via self-training svm. PLoS ONE. 2017;12(2) doi: 10.1371/journal.pone.0171839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Hao M., Wang Y., Bryant S.H. Improved prediction of drug-target interactions using regularized least squares integrating with kernel fusion technique. Anal. Chim. Acta. 2016;909:41–50. doi: 10.1016/j.aca.2016.01.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Gönen M. Predicting drug–target interactions from chemical and genomic kernels using Bayesian matrix factorization. Bioinformatics. 2012;28(18):2304–2310. doi: 10.1093/bioinformatics/bts360. [DOI] [PubMed] [Google Scholar]
- 23.Ba-Alawi W., Soufan O., Essack M., Kalnis P., Bajic V.B. Daspfind: new efficient method to predict drug–target interactions. J. Cheminform. 2016;8(1):15. doi: 10.1186/s13321-016-0128-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Chen H., Zhang Z. A semi-supervised method for drug-target interaction prediction with consistency in networks. PLoS ONE. 2013;8(5) doi: 10.1371/journal.pone.0062975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Huang Y.-A., You Z.-H., Chen X. A systematic prediction of drug-target interactions using molecular fingerprints and protein sequences. Curr. Protein Pept. Sci. 2018;19(5):468–478. doi: 10.2174/1389203718666161122103057. [DOI] [PubMed] [Google Scholar]
- 26.Daminelli S., Thomas J.M., Durán C., Cannistraci C.V. Common neighbours and the local-community-paradigm for topological link prediction in bipartite networks. New J. Phys. 2015;17(11) [Google Scholar]
- 27.Durán C., Daminelli S., Thomas J.M., Haupt V.J., Schroeder M., Cannistraci C.V. Pioneering topological methods for network-based drug–target prediction by exploiting a brain-network self-organization theory. Brief. Bioinform. 2018;19(6):1183–1202. doi: 10.1093/bib/bbx041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yuan Q., Gao J., Wu D., Zhang S., Mamitsuka H., Zhu S. Druge-rank: improving drug–target interaction prediction of new candidate drugs or targets by ensemble learning to rank. Bioinformatics. 2016;32(12) doi: 10.1093/bioinformatics/btw244. i18–i27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Wang L., You Z.-H., Chen X., Xia S.-X., Liu F., Yan X., Zhou Y., Song K.-J. A computational-based method for predicting drug–target interactions by using stacked autoencoder deep neural network. J. Comput. Biol. 2018;25(3):361–373. doi: 10.1089/cmb.2017.0135. [DOI] [PubMed] [Google Scholar]
- 30.Ezzat A., Wu M., Li X.-L., Kwoh C.-K. Drug-target interaction prediction via class imbalance-aware ensemble learning. BMC Bioinform. 2016;17(19):509. doi: 10.1186/s12859-016-1377-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ezzat A., Wu M., Li X.-L., Kwoh C.-K. Drug-target interaction prediction using ensemble learning and dimensionality reduction. Methods. 2017;129:81–88. doi: 10.1016/j.ymeth.2017.05.016. [DOI] [PubMed] [Google Scholar]
- 32.Chen X., Yan C.C., Zhang X., Zhang X., Dai F., Yin J., Zhang Y. Drug–target interaction prediction: databases, web servers and computational models. Brief. Bioinform. 2015;17(4):696–712. doi: 10.1093/bib/bbv066. [DOI] [PubMed] [Google Scholar]
- 33.Wen M., Zhang Z., Niu S., Sha H., Yang R., Yun Y., Lu H. Deep-learning-based drug–target interaction prediction. J. Proteome Res. 2017;16(4):1401–1409. doi: 10.1021/acs.jproteome.6b00618. [DOI] [PubMed] [Google Scholar]
- 34.López Y., Dehzangi A., Lal S.P., Taherzadeh G., Michaelson J., Sattar A., Tsunoda T., Sharma A. Sucstruct: prediction of succinylated lysine residues by using structural properties of amino acids. Anal. Biochem. 2017;527:24–32. doi: 10.1016/j.ab.2017.03.021. [DOI] [PubMed] [Google Scholar]
- 35.Taherzadeh G., Zhou Y., Liew A.W.-C., Yang Y. Structure-based prediction of protein-peptide binding regions using random forest. Bioinformatics. 2018;34(3):477–484. doi: 10.1093/bioinformatics/btx614. [DOI] [PubMed] [Google Scholar]
- 36.Freund Y., Schapire R.E. A desicion-theoretic generalization of on-line learning and an application to boosting. European Conference on Computational Learning Theory; Springer; Springer; 1995. pp. 23–37. [Google Scholar]
- 37.Goodfellow I., Bengio Y., Courville A., Bengio Y. MIT Press; Cambridge: 2016. Deep Learning, Vol. 1. [Google Scholar]
- 38.Du Y., Wang J., Wang X., Chen J., Chang H. Proceedings of the 2018 6th International Conference on Bioinformatics and Computational Biology. ACM; 2018. Predicting drug-target interaction via wide and deep learning; pp. 128–132. [Google Scholar]
- 39.Wang L., You Z.-H., Chen X., Xia S.-X., Liu F., Yan X., Zhou Y. International Symposium on Bioinformatics Research and Applications. Springer; 2017. Computational methods for the prediction of drug-target interactions from drug fingerprints and protein sequences by stacked auto-encoder deep neural network; pp. 46–58. [Google Scholar]
- 40.Szegedy C., Liu W., Jia Y., Sermanet P., Reed S., Anguelov D., Erhan D., Vanhoucke V., Rabinovich A. Going deeper with convolutions. 2014. arXiv:1409.4842
- 41.Szegedy C., Vanhoucke V., Ioffe S., Shlens J., Wojna Z. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. 2016. Rethinking the inception architecture for computer vision; pp. 2818–2826. [Google Scholar]
- 42.Szegedy C., Ioffe S., Vanhoucke V., Alemi A.A. vol. 4. 2017. Inception-v4, inception-resnet and the impact of residual connections on learning; p. 12. (AAAI). [Google Scholar]
- 43.Abadi M., Barham P., Chen J., Chen Z., Davis A., Dean J., Devin M., Ghemawat S., Irving G., Isard M. OSDI. vol. 16. 2016. Tensorflow: a system for large-scale machine learning; pp. 265–283. [Google Scholar]
- 44.Rayhan F., Ahmed S., Mahbub A., Jani M., Shatabda S., Farid D.M. Cusboost: cluster-based under-sampling with boosting for imbalanced classification. arXiv:1712.04356 arXiv preprint.
- 45.Kingma D.P., Ba Adam J. A method for stochastic optimization. arXiv:1412.6980 arXiv preprint.
- 46.Lin M., Chen Q., Yan S. Network in network. arXiv:1312.4400 arXiv preprint.
- 47.Wishart D.S., Knox C., Guo A.C., Cheng D., Shrivastava S., Tzur D., Gautam B., Hassanali M. Drugbank: a knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res. 2008;36(suppl 1):D901–D906. doi: 10.1093/nar/gkm958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Kanehisa M., Araki M., Goto S., Hattori M., Hirakawa M., Itoh M., Katayama T., Kawashima S., Okuda S., Tokimatsu T. Kegg for linking genomes to life and the environment. Nucleic Acids Res. 2008;36(suppl 1):D480–D484. doi: 10.1093/nar/gkm882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Schomburg I., Chang A., Ebeling C., Gremse M., Heldt C., Huhn G., Schomburg D. Brenda, the enzyme database: updates and major new developments. Nucleic Acids Res. 2004;32(suppl 1):D431–D433. doi: 10.1093/nar/gkh081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Günther S., Kuhn M., Dunkel M., Campillos M., Senger C., Petsalaki E., Ahmed J., Urdiales E.G., Gewiess A., Jensen L.J. Supertarget and matador: resources for exploring drug-target relationships. Nucleic Acids Res. 2008;36(suppl 1):D919–D922. doi: 10.1093/nar/gkm862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Cao D.-S., Liu S., Xu Q.-S., Lu H.-M., Huang J.-H., Hu Q.-N., Liang Y.-Z. Large-scale prediction of drug–target interactions using protein sequences and drug topological structures. Anal. Chim. Acta. 2012;752:1–10. doi: 10.1016/j.aca.2012.09.021. [DOI] [PubMed] [Google Scholar]
- 52.Friedman J.H. On bias, variance, 0/1—loss, and the curse-of-dimensionality. Data Min. Knowl. Discov. 1997;1(1):55–77. [Google Scholar]
- 53.Pedregosa F., Varoquaux G., Gramfort A., Michel V., Thirion B., Grisel O., Blondel M., Prettenhofer P., Weiss R., Dubourg V. Scikit-learn: machine learning in python. J. Mach. Learn. Res. 2011;12(Oct):2825–2830. [Google Scholar]
- 54.Safavian S.R., Landgrebe D. A survey of decision tree classifier methodology. IEEE Trans. Syst. Man Cybern. 1991;21(3):660–674. [Google Scholar]
- 55.Joachims T. Universität Dortmund; 1998. Making large-scale svm learning practical. Tech. Rep., Technical report, SFB 475: Komplexitätsreduktion in Multivariaten Datenstrukturen. [Google Scholar]
- 56.Rayhan F., Ahmed S., Mahbub A., Jani M., Shatabda S., Farid D.M., Rahman C.M. Meboost: mixing estimators with boosting for imbalanced data classification. arXiv:1712.06658 arXiv preprint.
- 57.Yamanishi Y., Kotera M., Kanehisa M., Goto S. Drug-target interaction prediction from chemical, genomic and pharmacological data in an integrated framework. Bioinformatics. 2010;26(12) doi: 10.1093/bioinformatics/btq176. i246–i254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Mutowo P., Bento A.P., Dedman N., Gaulton A., Hersey A., Lomax J., Overington J.P. A drug target slim: using gene ontology and gene ontology annotations to navigate protein-ligand target space in chembl. J. Biomed. Semant. 2016;7(1):59. doi: 10.1186/s13326-016-0102-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Ezzat A., Wu M., Li X.-L., Kwoh C.-K. Computational prediction of drug-target interactions using chemogenomic approaches: an empirical survey. Brief. Bioinform. 2019;20(4):1337–1357. doi: 10.1093/bib/bby002. [DOI] [PubMed] [Google Scholar]