Abstract.
Purpose: Radiomic features extracted from medical images acquired in different countries may demonstrate a batch effect. Thus, we investigated the effect of harmonization on a database of radiomic features extracted from dynamic contrast-enhanced magnetic resonance (DCE-MR) breast imaging studies of 3150 benign lesions and cancers collected from international datasets, as well as the potential of harmonization to improve classification of malignancy.
Approach: Eligible features were harmonized by category using the ComBat method. Harmonization effect on features was evaluated using the Davies–Bouldin index for degree of clustering between populations for both benign lesions and cancers. Performance in distinguishing between cancers and benign lesions was evaluated for each dataset using 10-fold cross validation with the area under the receiver operating characteristic curve (AUC) determined on the pre- and postharmonization sets of radiomic features in each dataset and a combined one. Differences in AUCs were evaluated for statistical significance.
Results: The Davies–Bouldin index increased by 27% for benign lesions and by 43% for cancers, indicating that the postharmonization features were more similar. Classification performance using postharmonization features performed better than that using preharmonization features ( for all three).
Conclusion: Harmonization of radiomic features may enable combining databases from different populations for more comprehensive computer-aided diagnosis models of breast cancer.
Keywords: radiomics, computer-aided diagnosis, harmonization, dynamic contrast-enhanced magnetic resonance, breast cancer
1. Introduction
The goal of computer-aided diagnosis of cancer is to support decision making in clinical practice of medicine by providing quantitative information on the state of disease. Quantitative descriptions of lesions extracted from medical images, so-called radiomic features, can contribute to the aims of computer-aided diagnosis.1–4 Many studies investigating the role of radiomic features in computer-aided diagnosis have done so using subjects from a single population. While this offers reduction of variation of image acquisition factors for that particular set of subjects, it is of broader interest to develop decision-making models that incorporate subject images acquired in different countries. Gain, system resolution, and image noise may all vary between imaging protocols that are nominally the same. Additionally, the interval of time between images in dynamic contrast-enhanced magnetic resonance (DCE-MR) series can differ, or a contrast agent may be administered with variation due to factors such as patient kidney health. It can be challenging to combine image databases from multiple centers for use in computer-aided diagnosis without accounting for these differences, which may contribute to false positives or false negatives in classification performance when machine learning models trained on cases imaged in one center are tested on cases imaged in another center. Harmonization may enable the combination of multiple databases. The potential application of harmonization to radiomic features is not unique to image-based computer-aided diagnosis. For example, in the field of genomics, the issue of differences in data collected from samples that are otherwise nominally the same but processed at different centers is referred to as the batch effect.5
The purpose of our study was to (a) assess the impact of harmonization on radiomic features extracted from MR images of breast lesions in two different populations, i.e., patient images acquired in the United States and those acquired in China, and (b) assess the impact of harmonization on the classification performance in the task of distinguishing the lesions as benign or cancerous, within each population and in a combined dataset of both populations.
2. Materials and Methods
2.1. Databases
DCE-MR images of breast lesions were retrospectively collected under Internal Review Board and HIPAA compliance from an institution in the United States (during the period of 2005 to 2017) and in China (during the period of 2015 to 2017) (Table 1). These images constituted the two “populations.” Images collected from the United States were acquired in the axial plane, whereas images collected from China were acquired in the sagittal plane. The time interval between postcontrast images was typically 60 s in the United States database and typically 90 s in the China database. Most images collected in the United States were acquired using Philips scanners (Best, The Netherlands) (1145 out of 1163 cases), whereas all images collected in the China database were acquired using GE Discovery 750 scanners (Waukesha, Wisconsin).
Table 1.
Composition of the databases: Number of lesions by status as benign or cancer, as well as maximum linear size and age.
| Number of lesions | Median maximum linear size [95% CI] (mm) | Median age [95% CI] (years) | |
|---|---|---|---|
| United States | 1163 | — | — |
| Benign | 264 (23%) | 12.7 [5.0 to 41.8] | 48 [27 to 74] |
| Cancer | 899 (77%) | 29.0 [8.3 to 106.9] | 55 [33 to 81] |
| China | 1987 | — | — |
| Benign | 481 (24%) | 19.9 [6.3 to 67.5] | 43 [21 to 60] |
| Cancer | 1506 (76%) | 27.2 [11.5 to 87.6] | 47 [30 to 69] |
Note: In the United States database, some ages were given in terms of decade, e.g., “50s.” For the purpose of calculating statistics for age of this database, the age was changed to the middle of the decade, e.g., “55” in the example given. The ages of 6 subjects with benign lesions and 41 subjects with cancers were changed in this manner. Additionally, the ages of 40 subjects with benign lesions and 50 subjects with cancers were unknown.
2.2. Computerized Breast Lesion Segmentation and Radiomic Feature Extraction
Lesions were segmented using a fuzzy c-means method.6 Thirty-two computer-extracted radiomic features were automatically calculated, covering categories of size, shape, morphology, texture, and kinetics of contrast dynamics.7–9 Features that were candidates for harmonization were identified for their dependence upon gray level and potential for intrinsic variation related to gain, image resolution, image noise, and imaging protocol. Features describing lesion geometry (i.e., categories of size and shape) and the kinetic curve assessment feature of volume of most enhancing voxels were deemed to not be candidates for harmonization due to these being characteristics directly of the lesions and not dependent on the image acquisition. Additionally, two other kinetic curve features, washout rate and curve shape index, were excluded from harmonization because they are semi-categorical variables, in which the algorithm sets the feature value to zero if criteria for indication of washout of contrast agent are not met, as is true for many benign lesions.9 Because features were harmonized by feature category, these two features were not included in the harmonization of the other kinetic curve features even when values were not set to zero, as doing so would have resulted in unmatched sets of features of cases after removal of features that were equal to zero. Based upon the consideration of features and their description, features that were candidates for harmonization were determined, whereas the other features did not undergo harmonization (Table 2).
Table 2.
Description of radiomic features.
| Feature abbreviation | Feature name | Feature description |
|---|---|---|
| Radiomic features deemed eligible for harmonization | ||
| M1 | Margin sharpness | Mean of the image gradient at the lesion margin |
| M2 | Variance of margin sharpness | Variance of the image gradient at the lesion margin |
| M3 | Variance of radial gradient histogram | Degree to which the enhancement structure extends in a radial pattern originating from the center of the lesion |
| T1 | Contrast | Location image variations |
| T2 | Correlation | Image linearity |
| T3 | Difference entropy | Randomness of the difference of neighboring voxels’ gray levels |
| T4 | Difference variance | Variations of difference of gray levels between voxel pairs |
| T5 | Energy | Image homogeneity |
| T6 | Entropy | Randomness of the gray levels |
| T7 | Inverse difference moment (homogeneity) | Image homogeneity |
| T8 | Information measure of correlation 1 | Nonlinear gray-level dependence |
| T9 | Information measure of correlation 2 | Nonlinear gray-level dependence |
| T10 | Maximum correlation coefficient | Nonlinear gray-level dependence |
| T11 | Sum average | Overall brightness |
| T12 | Sum entropy | Randomness of the sum of gray levels of neighboring voxels |
| T13 | Sum variance | Spread in the sum of the gray levels of voxel-pairs distribution |
| T14 | Sum of squares (variance) | Spread in the gray-level distribution |
| K1 | Maximum enhancement | Maximum contrast enhancement |
| K2 | Time to peak (s) | Time at which the maximum enhancement occurs |
| K3 | Uptake rate (1/s) | Uptake speed of the contrast enhancement |
| K6 | Enhancement at first postcontrast time point | Enhancement at first postcontrast time point |
| K7 | Signal enhancement ratio | Ratio of initial enhancement to overall enhancement |
| Radiomic features deemed not eligible for harmonization | ||
| S1 | Volume () | Volume of lesion |
| S2 | Effective diameter (mm) | Greatest dimension of a sphere with the same volume as the lesion |
| S3 | Surface area () | Lesion surface area |
| S4 | Maximum linear size (mm) | Maximum distance between any 2 voxels in the lesion |
| G1 | Sphericity | Similarity of the lesion shape to a sphere |
| G2 | Irregularity | Deviation of the lesion surface from the surface of a sphere |
| G3 | Surface area/volume (1/mm) | Ratio of surface area to volume |
| K4 | Washout rate (1/s) | Washout speed of the contrast enhancement |
| K5 | Curve shape index | Difference between late and early enhancement |
| K8 | Volume of most enhancing voxels () | Volume of the most enhancing voxels |
2.3. Radiomic Feature Harmonization
Harmonization was conducted using the parametric version of the ComBat method, which is based on additive and multiplicative batch effects using empirical Bayes estimates, with which to transform the features.5 In our study, the two populations (i.e., patients imaged in the United States and patients imaged in China) served as the two “batches,” yielding the source of differences the ComBat algorithm seeks to reduce. The harmonization was applied separately within categories features, i.e., morphology, texture, and kinetics, as described above. The ComBat harmonization method can accommodate covariates, i.e., confounding variables, which for our study was malignancy status of the lesions. The cancer prevalence was similar between the two populations (see Table 1), an important consideration for the implementation of ComBat harmonization for classification of lesions as benign or malignant.10
2.4. Effect of Harmonization on Radiomic Features
The effect of harmonization on feature value distributions in each population in benign lesions and in cancers was evaluated for each eligible feature using the Kolmogorov–Smirnov (K-S) test to compare the distributions.11,12 The K-S test statistic was used to characterize the change in feature value distribution due to harmonization.
To visualize the impact of harmonization methods on clustering of features by population, t-distributed stochastic neighbor embedding (t-SNE) methods were used to reduce the dimensionality of the feature sets from 32 to two.13 The Davies–Bouldin index14 was used after k-means clustering of the t-SNE values to assess inter- and intra-cluster agreement across the two populations, separately for benign lesions and for cancers.
2.5. Effect of Harmonization on CAD Performance
Classification of lesions as benign or malignant was performed using two sets of 32 radiomic features: (a) a feature set with all features in their original form (called as “preharmonization” in this work) and (b) a feature set comprised of features not eligible for harmonization combined with those that had undergone harmonization in their respective feature groups (called as “postharmonization” in this work).
The classification performance evaluations were performed separately for lesions imaged in the United States, for lesions imaged in China, and the combined dataset. For each classification evaluation on each set, 10-fold cross validation was performed using a random forest classifier with 100 decision trees, with the posterior probability of malignancy used as the classifier output for receiver operating characteristic (ROC) curve analysis. No feature selection was conducted prior to the use of the random forest classifier. The area under the ROC curve (AUC), determined using the proper binormal model,15 served as figure of merit, with its value and 95% CI determined using ROCkit software.16 The difference in AUC for each set of cases was deemed to be statistically significantly different if .
3. Results
3.1. Effect of Harmonization on Radiomic Features
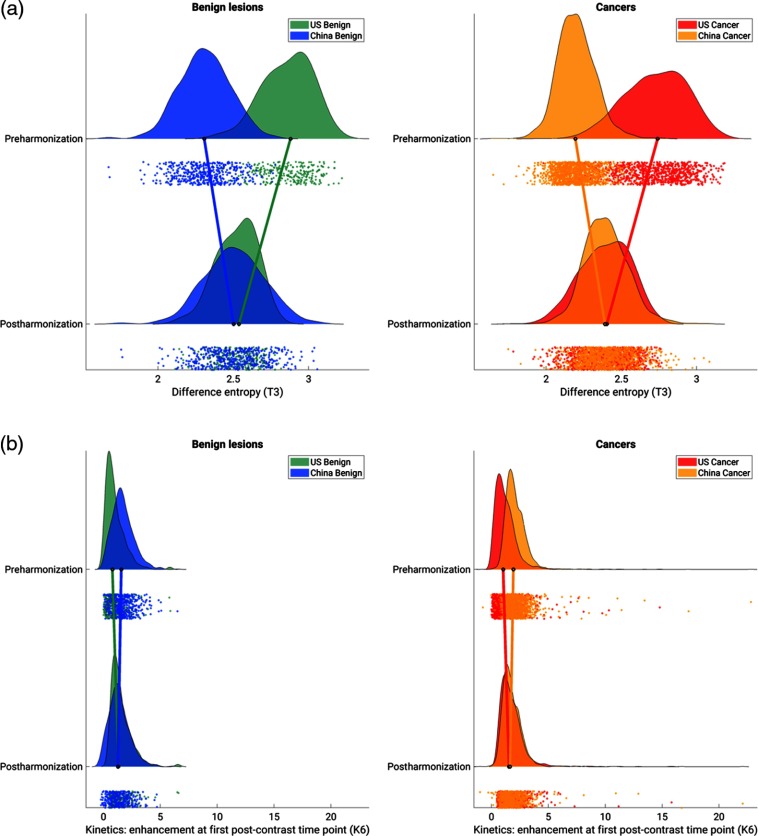
The harmonization method changed the distribution of values of the features that had been determined to be eligible for harmonization (Fig. 1). In general, for each radiomic feature eligible for harmonization, the difference in median feature value for benign lesions and for cancers was reduced after harmonization, and the distributions became more similar. The K-S test statistic for comparing feature value distributions pre- and postharmonization showed that in benign lesions, larger changes in most feature value distributions were seen in the lesions imaged in the United States. For cancers, some feature value distributions demonstrated more change in feature values in lesions imaged in China (features T2, T4, T9, T10, T12, and T13), and others demonstrated more change in feature values in lesions imaged in the United States (mostly, morphology and kinetic curve features, as well as the remaining texture features, Fig. 2). T-SNE space visualized pre- and postharmonization in benign lesions and in cancers (Fig. 3) show that the degree of clustering between populations was reduced after harmonization was applied to eligible features. The Davies–Bouldin index for degree of interclustering and intraclustering of t-SNE values increased after harmonization was applied to eligible features, demonstrating that the features were more similar between populations postharmonization (Fig. 4). The Davies–Bouldin index for benign lesions increased by 27%, whereas it increased for cancers by 43%.
Fig. 1.
Example raincloud plots for the distribution of the values of (a) the texture feature of difference entropy and (b) the kinetic curve feature of enhancement at first postcontrast time point, before and after harmonization for benign lesions and cancers within each population. The black circles at the base of each distribution indicate the median of the distribution, and the connecting lines between the medians of the distributions pre- and postharmonization demonstrate the change of the median with harmonization of the feature. The dots below each distribution show individual data points.
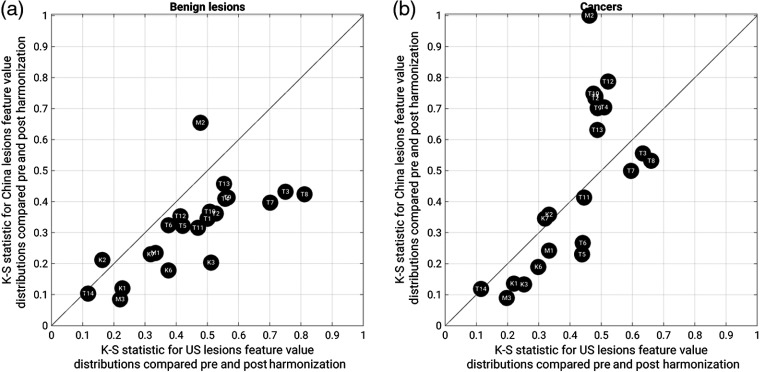
Fig. 2.
The K-S test statistic for feature value distributions compared pre- and postharmonization, for ( axis) lesions imaged in the United States and ( axis) lesions imaged in China [(a) benign lesions and (b) cancers]. Data points below the diagonal indicate that there is a larger difference between the feature value distributions compared pre- and postharmonization in lesions imaged in the United States, and data points above the diagonal indicate that there is a larger difference between the feature value distributions compared pre- and postharmonization in lesions imaged in China. Feature abbreviations are the same as in Table 2.
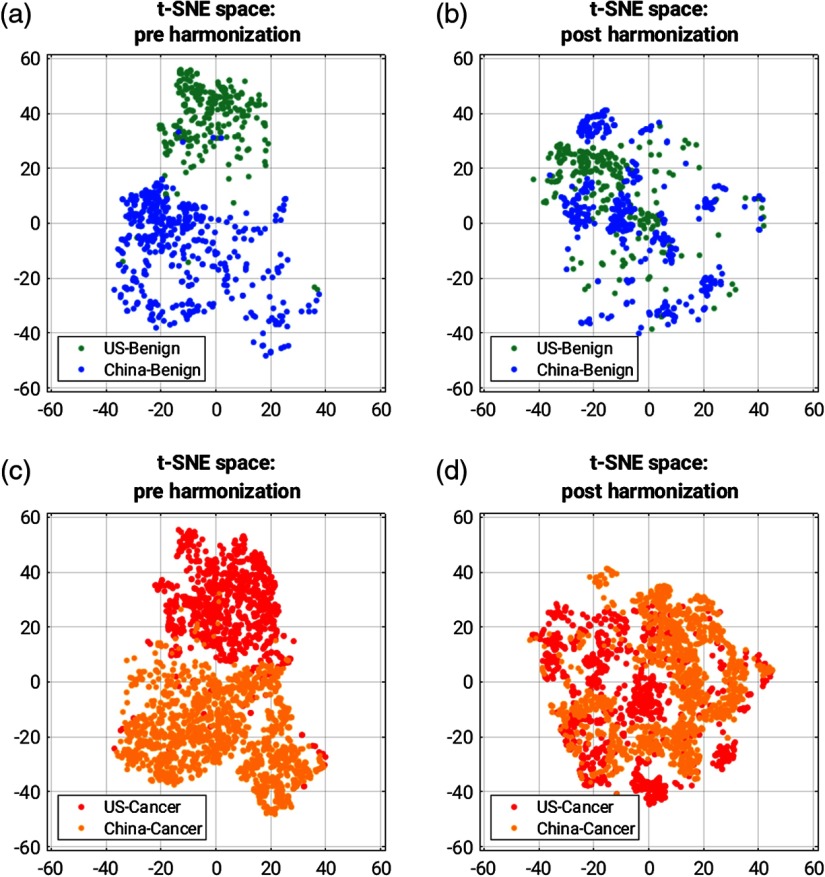
Fig. 3.
t-SNE space for the 32 features reduced to 2 features for (a), (c) preharmonization state and (b), (d) postharmonization. (a), (b) t-SNE space for benign features and (c), (d) t-SNE space for cancers.
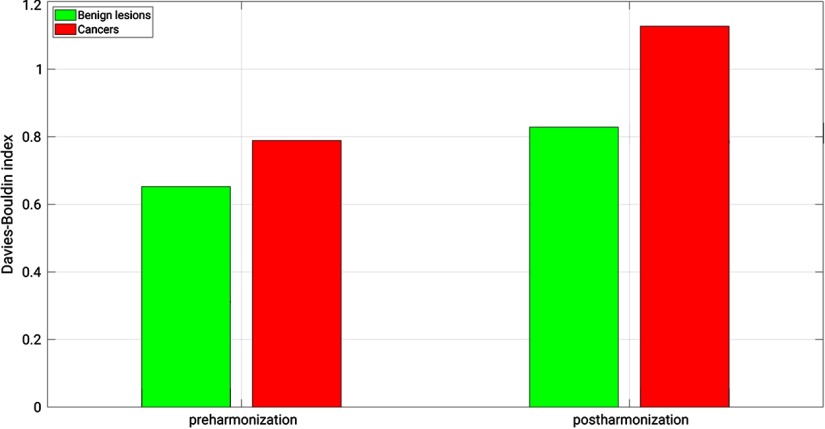
Fig. 4.
The Davies–Bouldin index, a measure of inter- and intra-clustering, for the t-SNE values (32 features reduced to 2) for benign lesions and cancers. A higher Davies–Bouldin index indicates improved harmonization across the features between populations, as is also seen in Fig. 3.
3.2. Effect of Harmonization on CAD Performance
In the task of classification of lesions as benign or malignant, classification performances, as measured by AUC, demonstrated statistically significant increase when postharmonization features were used compared with preharmonization features (Fig. 5, Table 3)
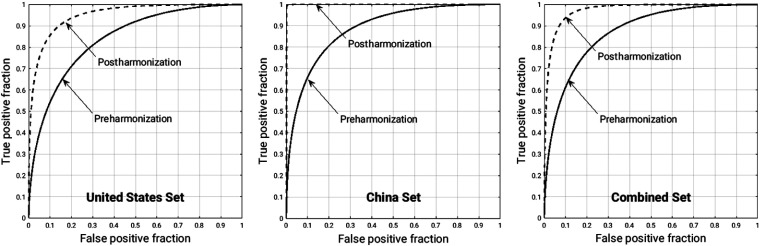
Fig. 5.
ROC curves in the task of classification of lesions as benign or malignant, for the three datasets pre- and postharmonization of selected radiomic features. Within each dataset, ROC analysis using the posterior probability of malignancy was performed after 10-fold cross validation with random forest classifier.
Table 3.
The AUC in the task of classification of lesions as benign or malignant, for classification using preharmonization features and for classification using postharmonization features. Within each dataset, the AUC was determined using classification with random forest classifier via 10-fold cross validation, with the posterior probability of malignancy used as the classifier output for ROC curve analysis.
| Dataset | [95% CI] | [95% CI] | [95% CI] | value |
|---|---|---|---|---|
| US set | 0.839 [0.810 to 0.864] | 0.951 [0.937 to 0.964] | 0.122 [0.095 to 0.131] | |
| China set | 0.886 [0.869 to 0.902] | 0.999 [0.995 to 1.000] | 0.113 [0.097 to 0.131] | |
| Combined set (US and China) | 0.872 [0.857 to 0.886] | 0.974 [0.968 to 0.980] | 0.102 [0.090 to 0.115] |
4. Discussion and Conclusion
Preliminary efforts toward harmonization of image-based features of cancer have been previously described for radiomic features extracted from postreconstruction positron-emission tomography images of breast cancer17 and from full-field digital mammography images of breast lesions.18 Our findings extend the application of this harmonization method to radiomic features extracted from DCE-MR images of breast lesions in two populations and demonstrate the importance of applying the method to features in terms of categories as well as carefully identifying which features can actually be submitted for harmonization. It is important to note that the scope of this work is limited to harmonization of radiomic features and does not consider harmonization in the context of feature selection or classifier training, which also could affect performance.
The larger changes in feature value distributions observed for most features in patients imaged in the United States may be due to the longer time period over which the database was collected in the United States, during which more variation in MR scanners and imaging protocol could have occurred, or the differences in field strength acquisition between the two groups. For example, the field strength of acquisition can have an inherent effect on features that depend upon field strength, as well as coincidental effect, due to changes in image acquisition parameters such as spatial resolution.19 Understanding these differences will be the subject of future work, particularly in the context of field strength of acquisition.
One limitation of our work is that there are additional biological covariates within our data, such as the variety of molecular subtypes within the database of the cancerous lesions. However, the classification of lesions as benign or cancerous is a clinically relevant task in its own right, and our future studies will investigate the application of harmonization in the context of molecular subtype and field strength of image acquisition, as noted earlier.
Application of harmonization methods to eligible radiomic features extracted from DCE-MR images of breast lesions resulted in increased similarity of features by population group. It also demonstrated statistically significant improvement in classification performance in each population separately and in the combined database, compared with classification performance using all radiomic features in their original preharmonization form. These findings may contribute to the advancement of machine learning models for computer-aided diagnosis that are developed using cases collected from multiple imaging centers, reducing the batch effect on classification performance.
Acknowledgments
This work was funded by NIH National Cancer Institute (NCI) U01 CA195564, NIH NCI R15 CA227948, the National Natural Science Foundation of China (81801781), and The University of Chicago Comprehensive Cancer Center Dancing with Chicago Celebrities Fund. The authors are grateful to Alexandra Edwards and John Papaioannou for their contributions to this work and would like to thank the NVIDIA Corporation for donating the GeForce GTX 1060 used in this article.
Biographies
Heather M. Whitney is an associate professor of physics at Wheaton College, Wheaton, Illinois, USA, and a visiting scholar in the Department of Radiology, The University of Chicago, Chicago, Illinois, USA. Her experience in quantitative medical imaging has ranged from polymer gel dosimetry to radiation damping in nuclear magnetic resonance to now focusing on computer-aided diagnosis (CADx) of breast cancer imaging. She is interested in investigating the effects of the physical basis of imaging on CADx, as well as the repeatability and robustness of CADx.
Hui Li is a research associate professor of radiology at The University of Chicago, Chicago, Illinois, USA, and has been involved in quantitative imaging analysis on medical images for over a decade. His research interests include breast cancer risk assessment, diagnosis, prognosis, response to therapy, understanding the relationship between radiomics and genomics, and their future roles in precision medicine with both conventional and deep learning approaches.
Yu Ji was a visiting scholar in the Department of Radiology, The University of Chicago, Chicago, Illinois, USA. He is currently an attending physician in the Department of Breast Imaging, Tianjin Medical University Cancer Institute and Hospital, Tianjin, China. He has been working for over five years on mammography, ultrasound, and magnetic resonance imaging (MRI). His current research interests include quantitative image analysis in breast cancer diagnosis, prognosis, and response to therapy.
Peifang Liu is currently the director of the Department of Breast Imaging, Tianjin Medical University Cancer Institute and Hospital, Tianjin, China. She has been working, for several decades, in mammography, ultrasound, and MRI. Her research interests include breast cancer diagnosis and management.
Maryellen L. Giger (fellow, IEEE) is the A. N. Professor of Radiology/Medical Physics at The University of Chicago, Chicago, Illinois, USA, and has been working, for multiple decades, in CADx, computer vision, machine learning, and deep learning in cancer diagnosis and management. Her research interests include understanding the role of quantitative radiomics and machine learning in personalized medicine.
Disclosures
M. L. Giger is a stockholder in R2 Technology/Hologic and a cofounder and equity holder in Quantitative Insights (now Qlarity Imaging). M. L. Giger receives royalties from Hologic, GE Medical Systems, MEDIAN Technologies, Riverain Medical, Mitsubishi, and Toshiba. It is the University of Chicago Conflict of Interest Policy that investigators disclose publicly actual or potential significant financial interest that would reasonably appear to be directly and significantly affected by the research activities.
Contributor Information
Heather M. Whitney, Email: heather.whitney@wheaton.edu.
Hui Li, Email: huili@uchicago.edu.
Yu Ji, Email: 364446831@qq.com.
Peifang Liu, Email: cjr.liupeifang@vip.163.com.
Maryellen L. Giger, Email: m-giger@uchicago.edu.
References
- 1.Gillies R. J., Kinahan P. E., Hricak H., “Radiomics: images are more than pictures, they are data,” Radiology 278(2), 563–577 (2016). 10.1148/radiol.2015151169 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Saha A., Harowicz M. R., Mazurowski M. A., “Breast cancer MRI radiomics: an overview of algorithmic features and impact of inter-reader variability in annotating tumors,” Med. Phys. 45(7), 3076–3085 (2018). 10.1002/mp.2018.45.issue-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Reig B., et al. , “Machine learning in breast MRI,” J. Magn. Reson. Imaging (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Net J. M., et al. , “Relationships between human-extracted MRI tumor phenotypes of breast cancer and clinical prognostic indicators including receptor status and molecular subtype,” Curr. Probl. Diagn. Radiol. 48(5), 467–472 (2019). 10.1067/j.cpradiol.2018.08.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Johnson W. E., Li C., Rabinovic A., “Adjusting batch effects in microarray expression data using empirical Bayes methods,” Biostatistics 8(1), 118–127 (2007). 10.1093/biostatistics/kxj037 [DOI] [PubMed] [Google Scholar]
- 6.Chen W., Giger M. L., Bick U., “A fuzzy C-means (FCM)-based approach for computerized segmentation of breast lesions in dynamic contrast-enhanced MR images,” Acad. Radiol. 13(1), 63–72 (2006). 10.1016/j.acra.2005.08.035 [DOI] [PubMed] [Google Scholar]
- 7.Gilhuijs K. G., Giger M. L., Bick U., “Computerized analysis of breast lesions in three dimensions using dynamic magnetic-resonance imaging,” Med. Phys. 25(9), 1647–1654 (1998). 10.1118/1.598345 [DOI] [PubMed] [Google Scholar]
- 8.Chen W., et al. , “Volumetric texture analysis of breast lesions on contrast-enhanced magnetic resonance images,” Magn. Reson. Med. 58(3), 562–571 (2007). 10.1002/(ISSN)1522-2594 [DOI] [PubMed] [Google Scholar]
- 9.Chen W., et al. , “Automatic identification and classification of characteristic kinetic curves of breast lesions on DCE-MRI,” Med. Phys. 33(8), 2878–2887 (2006). 10.1118/1.2210568 [DOI] [PubMed] [Google Scholar]
- 10.Nygaard V., Rødland E. A., Hovig E., “Methods that remove batch effects while retaining group differences may lead to exaggerated confidence in downstream analyses,” Biostatistics 17(1), 29–39 (2016). 10.1093/biostatistics/kxv027 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kolmogorov A., “Sulla Determinazione Empirica di una Legge di Distributione,” G. dell’Istituto Ital. degli Attuari 4, 1–11 (1933). [Google Scholar]
- 12.Smirnov N., “Table for estimating the goodness of fit of empirical distributions,” Ann. Math. Stat. 19, 279–281 (1948). 10.1214/aoms/1177730256 [DOI] [Google Scholar]
- 13.van der Maaten L., Hinton G. E., “Visualizing data using t-SNE,” J. Mach. Learn. Resour. 9, 2579–2605 (2008). [Google Scholar]
- 14.Davies D. L., Bouldin D. W., “A cluster separation measure,” IEEE Trans. Pattern Anal. Mach. Intell. PAMI-1(2), 224–227 (1979). 10.1109/TPAMI.1979.4766909 [DOI] [PubMed] [Google Scholar]
- 15.Metz C. E., Pan X., “‘Proper’ binormal ROC curves: theory and maximum-likelihood estimation,” J. Math. Psychol. 43(1), 1–33 (1999). 10.1006/jmps.1998.1218 [DOI] [PubMed] [Google Scholar]
- 16.Metz C. E., “ROCkit,” http://metz-roc.uchicago.edu/ (1998).
- 17.Orlhac F., et al. , “A postreconstruction harmonization method for multicenter radiomic studies in PET,” J. Nucl. Med. 59(8), 1321–1328 (2018). 10.2967/jnumed.117.199935 [DOI] [PubMed] [Google Scholar]
- 18.Robinson K., et al. , “Radiomics robustness assessment and classification evaluation: a two-stage method demonstrated on multi-vendor FFDM,” Med. Phys. 46(5), 2145–2156 (2019). 10.1002/mp.2019.46.issue-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Whitney H. M., et al. , “Robustness of radiomic breast features of benign lesions and luminal A cancers across MR magnet strengths,” Proc. SPIE 10575, 105750A (2018). 10.1117/12.2293764 [DOI] [Google Scholar]