Abstract
Amyotrophic lateral sclerosis (ALS) is a neurodegenerative disease for which the pathophysiological mechanisms of motor neuron loss are not precisely clarified. Environmental and epigenetic mechanisms such as microRNAs (miRNAs) could have a role in disease progression. We studied the expression pattern of miRNAs in ALS serum from 60 patients and 29 healthy controls. We also analyzed how deregulated miRNAs found in serum affected cellular pathways such as apoptosis, autophagy and mitochondrial physiology in SH-SY5Y cells. We found that miR-335-5p was downregulated in ALS serum. SH-SY5Y cells were transfected with a specific inhibitor of miR-335-5p and showed abnormal mitochondrial morphology, with an increment of reactive species of oxygen and superoxide dismutase activity. Pro-apoptotic caspases-3 and 7 also showed an increased activity in transfected cells. The downregulation of miR-335-5p, which has an effect on mitophagy, autophagy and apoptosis in SH-SY5Y neuronal cells could have a role in the motor neuron loss observed in ALS.
Subject terms: Neurological disorders, Neurodegenerative diseases, Neuromuscular disease
Introduction
Amyotrophic lateral sclerosis (ALS) is a progressive neurodegenerative disease. It is classically characterized by weakness and atrophy due to loss of lower motor neurons in the brainstem and spinal cord and loss of upper motor neurons in the motor cortex1. Degeneration, however, is now considered to be more widespread and not restricted to the motor system. Up to 50% of patients show evidence of behavioral and cognitive dysfunction, suggesting pathologic changes involving the frontal and temporal cortex, and sometimes fulfilling diagnostic criteria of frontotemporal dementia (FTD). These disorders are now considered to be manifestations within the clinicopathological spectrum of an underlying mechanism of neurodegeneration2,3.
The molecular pathogenic mechanism of ALS onset and progression is not completely known. Environmental factors have been described as potential contributors to neurodegeneration and ALS progression, but none have proven to be causative4,5. Mutations in more than 25 genes have been implicated in familial (fALS) and sporadic ALS (sALS) forms6, suggesting a wide heterogeneity in the molecular bases of the disease. Several ALS-causing genes encode proteins involved in RNA metabolism processes, such as RNA transcription, splicing, mRNA transport and microRNAs (miRNAs) biogenesis7. Dysfunction of these mechanisms is not exclusive and may lead to diverse cellular abnormalities. Epigenetic mechanisms, such as miRNAs, could regulate the expression of genes involved in common cellular pathways that are disrupted in ALS and therefore have an impact on ALS phenotype8. MiRNAs are small, non-coding RNA molecules of approximately 22 nucleotides that regulate gene expression in fundamental cellular processes and, post-transcriptionally, in the translation levels of target mRNA transcripts9. A single miRNA can regulate multiple transcripts and a single transcript can be regulated by multiple miRNAs. MiRNAs mediate silencing of target genes guided by Argonaute proteins, forming the silencing complex which promotes translational repression and degradation of targeted mRNA10. Numerous miRNAs are also temporally and spatially regulated with transcription factors or epigenetic mechanisms11. Deregulation of miRNA function is associated with numerous diseases12, including ALS13,14. Therefore, profiling of the miRNA expression could serve as a potential tool for ALS diagnosis, prognosis, and follow up, or to gain insight into the pathophysiology of the disease13.
Oxidative stress and mitochondrial damage are key features of most neurodegenerative diseases15,16, including ALS17, suggesting that mitochondrial dysfunction might contribute to motor neuron pathology18,19. Mitochondrial oxidative stress is also responsible for mitochondrial DNA damage in spinal motor neurons of sporadic ALS patients20. Mitochondrial dynamics play an important role in the neuronal survival by maintaining ATP biogenesis, Ca2+ homeostasis and regulating oxidative stress. Mitochondria produce ATP through the OXPHOS system, and at the same time, they generate reactive species of oxygen (ROS)21,22. When ROS production is excessive it causes oxidative stress in the cell23. Mitochondrial components may be damaged by ROS and can cause dysfunction of respiratory enzyme activity and mitochondrial depolarization24. Oxidative stress also induces cell death, releasing pro-apoptotic proteins25. A central role for autophagy in ALS is well supported by the pathology of the disease, which commonly includes the accumulation of protein aggregates and swollen mitochondria in motor neurons of affected patients26. In this study, we aimed to analyze the miRNA expression pattern in ALS serum and test the hypothesis that deregulation of some miRNA affects the mitochondrial physiology of neuronal cells.
Results
In the discovery phase thirteen miRNA had deregulated expression in ALS serum compared to controls
Seven ALS patients and six healthy controls were analyzed in the discovery phase. The clinical and demographic characteristics of ALS patients are included in Table 1. We observed significantly different expression levels between ALS patients and controls in 13 out of the 185 mRNAs included in the panel from the discovery phase (Fig. 1). The 13 deregulated miRNAs were: miR-107, miR-142-3p, miR-30c-5p, miR-335-5p, miR-421, miR-423-3p, miR-454-3p, miR-7a-5p, miR-122-5p, miR-125a-5p, miR-30b-5p, miR-30e-5p and miR-2110.
Table 1.
Demographic and clinical characteristics of patients included in the miRNAs study.
| Id | Gender | Age | Age at onset | Age at death | Mutated gene | Mutation | Phenotype | Onset | Familial |
|---|---|---|---|---|---|---|---|---|---|
| 1 | F | 69,8 | 67,6 | 71,1 | — | — | ALS | espinal | No |
| 2 | M | 64,6 | 62,2 | 67,7 | — | — | ALS | espinal | No |
| 3 | F | 70,3 | 68,6 | Alive | — | — | ALS | espinal | No |
| 4 | M | 68,4 | 60,9 | Alive | — | — | ALS | espinal | No |
| 5 | F | 49,4 | 46,0 | Alive | SOD1 | p.Asp77Val | fALS | espinal | Yes |
| 6 | M | 39,9 | 34,3 | Alive | TBK1 | p.Gly121Asp | ALS | espinal | No |
| 7 | M | 60,9 | 60,5 | Alive | — | — | ALS | espinal | No |
| 8 | M | 57,0 | 55,5 | Alive | — | — | ALS | espinal | No |
| 9 | M | 78,4 | 77,8 | 78,9 | — | — | ALS | espinal | No |
| 10 | M | 72,2 | 64,0 | 72,7 | — | — | ALS | espinal | No |
| 11 | F | 71,0 | 70,1 | 72,5 | — | — | ALS | bulbar | No |
| 12 | M | 75,0 | 74,4 | 75,2 | — | — | ALS | espinal | No |
| 13 | M | 61,9 | 53,9 | 64,2 | — | — | ALS | espinal | No |
| 14 | M | 52,7 | 52,0 | Alive | — | — | ALS | espinal | No |
| 15 | M | 71,5 | 69,0 | Alive | — | — | ALS | bulbar | No |
| 16 | F | 78,3 | 73,9 | Alive | — | — | ALS-FTD | bulbar | No |
| 17 | M | 64,2 | 35,3 | 66,6 | SOD1 | p.GLy38Arg | fALS | espinal | Yes |
| 18 | F | 71,9 | 68,5 | Alive | — | — | ALS | espinal | No |
| 19 | F | 64,4 | 62,8 | 65,5 | — | — | ALS | bulbar | No |
| 20 | F | 56,6 | 54,3 | 57,3 | — | — | ALS | espinal | No |
| 21 | F | 32,1 | 25,7 | Alive | — | — | ALS | espinal | No |
| 22 | F | 55,9 | 42,0 | 57,6 | SOD1 | p.Ile113Met | fALS | espinal | Yes |
| 23 | F | 85,7 | 84,6 | 86,2 | — | — | ALS | bulbar | No |
| 24 | M | 68,0 | 67,4 | 71,6 | — | — | ALS | espinal | Yes |
| 25 | M | 55,6 | 54,0 | 57,8 | — | — | ALS-FTD | bulbar | Yes |
| 26 | M | 46,4 | 45,1 | 47,5 | FUS | p.Arg521His | fALS | espinal | Yes |
| 27 | M | 65,1 | 62,2 | 65,5 | TARDBP | p.Arg90Val | ALS-FTD | bulbar | Yes |
| 28 | F | 86,3 | 84,2 | 86,3 | — | — | ALS | bulbar | No |
| 29 | M | 82,9 | 75,3 | Alive | — | — | ALS | espinal | No |
| 30 | F | 69,2 | 65,2 | 70,2 | — | — | ALS | espinal | No |
| 31 | F | 66,6 | 63,4 | 67,1 | — | — | ALS | espinal | No |
| 32 | F | 53,5 | 38,5 | Alive | SOD1 | p.Gly37Arg | fALS | espinal | Yes |
| 33 | F | 62,2 | 57,3 | 64,9 | — | — | ALS | espinal | No |
| 34 | M | 64,5 | 61,9 | 64,7 | — | — | ALS | espinal | No |
| 35 | F | 74,1 | 64,7 | Alive | — | — | ALS | espinal | No |
| 36 | F | 63,3 | 60,7 | Alive | — | — | ALS | espinal | No |
| 37 | F | 73,1 | 70,4 | 74,0 | — | — | ALS-FTD | bulbar | No |
| 38 | F | 82,9 | 81,9 | 83,2 | — | — | ALS | bulbar | No |
| 39 | M | 79,6 | 71,6 | 80,5 | — | — | ALS | espinal | No |
| 40 | M | 73,7 | 73,3 | 73,8 | — | — | ALS | bulbar | No |
| 41 | F | 69,3 | 68,0 | 70,9 | — | — | ALS | espinal | No |
| 42 | F | 61,8 | 59,5 | 60,5 | — | — | ALS | espinal | No |
| 43 | F | 72,5 | 70,3 | 73,4 | — | — | ALS | bulbar | No |
| 44 | M | 46,0 | 42,6 | 47,7 | — | — | ALS | bulbar | No |
| 45 | F | 71,1 | 70,7 | 71,2 | — | — | ALS | espinal | No |
| 46 | F | 68,9 | 68,4 | 71,2 | — | — | ALS | bulbar | No |
| 47 | F | 30,8 | 29,8 | Alive | FUS | p.Ser57del | fALS | espinal | Yes |
| 48 | F | 56,4 | 54,6 | Alive | — | — | ALS | espinal | No |
| 49 | F | 66,4 | 66,4 | 67,1 | — | — | ALS | espinal | No |
| 50 | M | 70,1 | 68,0 | 70,6 | — | — | ALS-FTD | bullbar | No |
| 51 | M | 50,9 | 50,1 | 52,4 | — | — | ALS | bulbar | No |
| 52 | M | 75,0 | 74,0 | Alive | — | — | ALS | espinal | No |
| 53 | F | 63,1 | 62,3 | 53,8 | — | — | ALS-FTD | espinal | No |
| 54 | F | 63,8 | 62,8 | 65,5 | — | — | ALS | bulbar | No |
| 55 | M | 76,4 | 75,5 | 77,2 | — | — | ALS | bulbar | No |
| 56 | M | 60,1 | 58,7 | 62,3 | — | — | ALS | bulbar | No |
| 57 | M | 74,0 | 67,5 | Alive | — | — | ALS | espinal | No |
| 58 | F | 79,0 | 78,0 | 79,0 | — | — | ALS | espinal | No |
| 59 | H | 62,9 | 61,9 | 64,7 | — | — | ALS | espinal | No |
| 60 | F | 69,7 | 67,6 | 71,1 | — | — | ALS | espinal | No |
Sera from patients 1 to 7 (in bold) were used in the discovery phase to study miRNAs profile. Sera from patients 8 to 60 were analyzed in the validation phase.
Figure 1.
Deregulated miRNA expression in the discovery phase. Normalized relative expression levels of serum miRNAs in ALS patients compared to healthy controls. Data are presented as mean ± SD. *p ≤ 0.05 (Mann-Whitney U test). Graphics of miRNA expression were created using GraphPad Prism, version 5.03 (https://www.graphpad.com/scientific-software/prism/).
miR-335-5p is differentially expressed between ALS serum and controls in the validation phase
In the validation phase we assessed the expression of the 13 miRNA deregulated in the discovery phase in sera from 53 ALS patients and 23 healthy controls. After Gene-Globe analysis (Qiagen, Hilden, Germany), the expression of miR-335-5p in serum showed a significant reduction (−1.67 fold decrease, p = 0.023) in ALS patients compared to controls. The Mann-Whitney U test of each miRNA expression between ALS and controls showed statistically significant differences in miR-335-5p (Fig. 2) No significant differences were observed in the other miRNAs analyzed in the validation phase (Fig. 2).
Figure 2.
MiRNA expression in validation phase. Only miR-335-5p expression showed statistically significant differences between ALS patients and healthy controls. Normalized relative expression levels of serum miRNAs in ALS patients compared to healthy controls. Data are presented as mean ± SD. *p ≤ 0.05 (Mann-Whitney U test). Graphics of miRNA expression were created using GraphPad Prism, version 5.03 (https://www.graphpad.com/scientific-software/prism/).
Inhibition of miR-335-5p altered mitochondrial dynamics
To assess the involvement of decreased miR-335-5p in mitochondrial dynamics, SH-SY5Y cells were transfected with an antisense miR-335 vector, a negative control (NC) together with mKeima-Red-Mito7. The fluorescent protein Keima has an excitation spectrum that changes according to pH. In a neutral environment the protein is excited at a short wavelength and displays a green signal, whereas in an acidic environment it is excited at a long wavelength and displays a red signal27. The miR-335 inhibitor forms a duplex with the miRNA guide strand and prevents miRNA from binding to its intended target. At 24 h after miR-335 inhibitor transfection, miR-335-5p levels were drastically reduced (Ct ≥ 40) while the NC showed a Ct of 32.2 ± 0.5.
After 24 h of transfection, mitochondria displayed no morphological changes. However, 48 h and 72 h after transfection, the number of mitochondria displaying a red signal increased significantly, indicating that they were located in acidic environment, probably due to mitophagy mediated by lysosomes (Fig. 3A). Furthermore, at 72 hours some cells presented round-shaped mitochondria which differed from those in NC. Excitation at 561 nm showed red mitochondria located in an acidic environment whereas excitation at 458 nm showed green mitochondria under a neutral environment. The excitation ratio 561/458 nm thus shows the ratio between mitophagic and healthy mitochondria. This ratio showed statistically significant differences at 48 h (**p < 0.01) and 72 h (*p < 0.05) between the miR-335 inhibitor and NC transfected cells (Fig. 3B).
Figure 3.
Co-transfection of miR-335 inhibitor or NC together with Keima Red. (A) Confocal images showed an increased pool of mitochondria 48 and 72 h after transfection with the miR-335 inhibitor.0 (B) 561/458 nm excitation ratio showed an increase number of red mitochondria (excited at 561), indicating there localization in an acidic environment. The graphic representation was created using GraphPad Prism, version 5.03, (https://www.graphpad.com/scientific-software/prism/).
At 72 h after transfection, some of the miR-335 inhibitor transfected cells presented swollen and rounded mitochondria, and they co-localized with p62, an autophagosome marker. In miR-335 inhibitor transfected cells, p62 remained accumulated on perinuclear clustered-mitochondria, while in NC no p62 positive vacuoles were observed (Fig. 4A). The Western-Blot (WB) of p62, showed an increased expression (*p < 0.05) at 72 h (Fig. 4B,C), indicating a potentially abnormal autophagy process in SH-SY5Y cells after miR-335 inhibitor transfection.
Figure 4.
Mitochondria co-localize with p62 after miR-335 inhibitor transfection. (A) Co-localization of mitochondria (red) with p62 autophagic marker (green), 72 h after transfection with miR-335 inhibitor. (B) Representative WB of p62 in cells after 24, 48 or 72 hours after transfection. Bands corresponding to p62 and β-actin were cropped from the same gel. (C) WB quantification showed a decrease in p62 expression at 48 h, and an increase at 72 h after transfection, indicating potential impairment of autophagy in transfected cells. The graphic representation was created using GraphPad Prism, version 5.03, (https://www.graphpad.com/scientific-software/prism/).
Oxidative stress is increased in miR-335 inhibitor transfected cells
The generation of ROS in cells has a deleterious effect on mitochondria normal function. For this reason, we studied the production of ROS in SH-SY5Y cells transfected with the miR-355 inhibitor. Cells transfected with this inhibitor showed green fluorescence, indicating that the fluorogenic probe of the assay was oxidized by ROS. This was not observed in NC (Fig. 5A). We also observed a significant increase in SOD activity 24 h after miR-335 inhibitor transfection (t-test **p < 0.01) compared to NC (Fig. 5B). No differences were observed at 48 and 72 h after transfection (Data not shown).
Figure 5.
Increment of ROS and SOD activity after miR-335 inhibitor transfection. (A) ROS detection in SH-SY5Y transfected cells 24 h after transfection correlated with an increment of SOD activity as a measure to regulate ROS levels (B). The graphic representation was created using GraphPad Prism, version 5.03, (https://www.graphpad.com/scientific-software/prism/).
Inhibition of miR-335-5p activates caspase 3/7 apoptotic pathway
To determine whether apoptosis could be influenced by miR-335-5p downregulation, we analyzed the expression of caspase-3 and caspase-7 as final effectors of the classical apoptotic pathway. Furthermore, CASP-7 is directly targeted by miR-335-5p (http://www.targetscan.org). After cell transfection with the miR-335 inhibitor, we observed a 1.63 fold-increase in caspase-7 and a 1.52 fold-increase of expression in caspase-3 at 72 h. However, these differences did not reach statistical significance (Fig. 6A,B).
Figure 6.
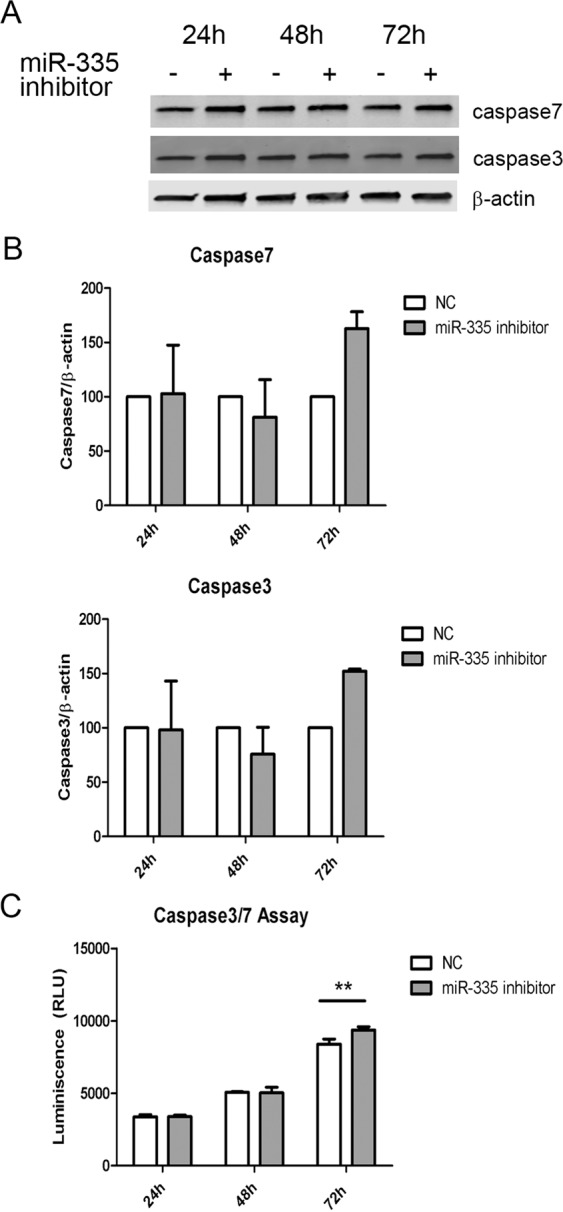
Abnormal activity of apoptosis effector caspases after miR-335 inhibitor transfection. (A) Representative WB of caspase-3 and 7 expressions after miR-335 transfection. Bands corresponding to caspase-7, caspase-3 and β-actin were cropped from the same gel. We can quantify the bands corresponding to caspase-7 and caspase-3 (B), and although no significant differences were observed, when we measured the activity of these caspases, we observed a significant increase in caspase activity (**p < 0.01) (C). The graphic representations were created using GraphPad Prism, version 5.03, (https://www.graphpad.com/scientific-software/prism/).
To study whether the increased expression observed by WB would correlate with an increment of caspase 3/7 activity, we measured this by luminescence ((RLU = Relative Light Units)). No differences were observed at 24 h or 48 h after transfection. However, a significant increase of caspase 3/7 activity was observed at 72 h (2way ANOVA, p < 0.01)(Fig. 6C).
Discussion
Our screening of 185 miRNAs in serum from ALS patients and controls revealed a downregulation of miR-335-5p in ALS patients, and this was replicated in an independent validation cohort. MiR-335-5p has high potential activity since it targets 2544 genes28. Changes in the expression of this miRNA could therefore have an impact on multiple cell functions.
In other motor neuron diseases, such as spinal muscle atrophy, miR-335-5p was highly reduced in neural stem cells29,30. Our results in ALS patients add evidence to a possible role of this specific non-coding RNA in the pathogenic process of motor neuron degeneration. It has been reported that miR335-5p downregulation is necessary to stabilize plasticity and memory in mice31 and may play a role in the appearance of cognitive symptoms in ALS patients with concomitant FTD, thus theoretically, contributing to the whole clinical spectrum of the disease.
Previous studies have described that miR-206, miR-143-3p and miR-374b-5p are deregulated in ALS patients’ serum32. Our panel did not include miR-206 and we could not therefore test its expression in our samples. We did not observe significant differences in miR-143 expression in our panel. Although miR-374b-5p was represented in our panel and showed a tendency to be downregulated in ALS patients, no significant differences were reached between groups.
Low levels of miR-30b-5p and high expression of miR-2110 have been reported to correlate with more rapid progression of the disease33. Although we observed that levels of miR-30b-5p and miR-2110 were low in the serum of our ALS cohort, again our results did not reach statistical significance.
Our results suggest that downregulation of miR-335-5p affects mitochondrial dynamics, autophagy and apoptosis in SH-SY5Y neuronal cells. Mitophagy is the selective degradation of damaged mitochondria by autophagy, contributing to maintenance of a healthy population of mitochondria. Since damaged mitochondria lead to a collapse of cell homeostasis, mitophagy is believed to protect against diseases related to mitochondrial dysfunction, such as neurodegenerative disorders34. In our study, we observed a co-localization of mitochondria with p62. P62 can act as an autophagy receptor that connects autophagic substrates with autophagosomes35, and it is increased following induction of mitophagy36. These findings, together with the observation of mitochondria in an acidic environment, probably inside autophagic vacuoles, indicate that downregulation of miR-335-5p could induce mitophagy in SH-SY5Y cells. Oxidative stress and mitochondrial alterations are also observed in the muscle of SOD1mutant mice37,38. Under oxidative stress conditions, ROS production is dramatically increased, resulting in subsequent alteration of membrane lipids, proteins and nucleic acids39 which could induce cell death. We observed an increase of ROS in miR-335 inhibitor transfected cells and a proportional increase of SOD activity, suggesting an attempt to regulate ROS levels after miR-335-5p downregulation. Mitochondrial SOD2 and SOD1 are usually sufficient to remove superoxide efficiently to avoid propagation of damage in the other compartments of the cell20. Mitochondria regulate apoptosis in the nervous system both in healthy and pathological conditions18. Caspase-7 together with caspase-3 are executioner caspases that lead to apoptosis by cleaving specific sets of substrates40. Mature caspase-3 and 7 activation results in nuclear condensation and genomic DNA break up, among other cellular events. Since CASP-7 is a direct target of miR-335-p, our results suggest that the downregulation of miR-335-p increases caspase-7 expression which in turn activates apoptosis and leads to cell death of SH-SY5Y neuronal cells. In fact, it has been reported that increased caspase-7 expression correlates with excessive neuronal cell death in some neurodegenerative disorders41. Therefore, caspase-7 activation constitutes a potential therapeutic target that could be beneficial in diseases such as ALS.
Our results provide evidence that downregulation of miR-335-5p may enhance mitophagy and apoptosis and support the notion that dysregulation of miRNAs may be involved in the pathogenic process of neuron degeneration in ALS.
Patients and Methods
Subjects, sample collection and isolation of miRNAs in serum samples
In this study, we analyzed samples from 89 subjects (60 ALS patients and 29 sex and age-matched healthy controls). The study was designed in two steps: a discovery phase (7 ALS patients and 6 healthy controls) and a validation phase (53 ALS patients and 23 healthy controls) (Table 1). ALS patients were prospectively recruited from the Motor Neuron Disease Clinic at Hospital de la Santa Creu i Sant Pau. They all fulfilled El Escorial revised criteria for probable, probable laboratory-supported, or definite ALS42 and underwent comprehensive cognitive and behavioral screening at the Sant Pau Memory Unit. Patients were categorized as ALS-FTD according to current diagnostic criteria for the behavioral variant of frontotemporal dementia43,44. Healthy controls were recruited from non-genetically related relatives and serum samples were taken after signed informed consent was obtained. All patients were able to sign an informed consent form by themselves. The study was approved by the local ethics committee (Ethic Committee of Investigation with Drugs from Fundación de Gestió Sanitària de l’Hospital de la Santa Creu i Sant Pau) in compliance with the 1964 Helsinki Declaration and its later amendments.
Analysis of miRNA expression in serum
After whole blood samples were collected and left for 30 minutes to clot. The tubes were then centrifuged at 1800 rpm for 10 minutes. miRNA extraction was performed as previously described45.miRNAs were isolated from serum using the miRCURY RNA Isolation Kit – Biofluids (Qiagen). 200 µl of serum were lysed with lysis solution following manufacturer’s instructions. MiRNAs were isolated after isopropanol precipitation and centrifugation in spin columns. MS2 RNA (Roche, Basel, Switzerland) was added to the samples as a carrier to improve the efficiency of RNA isolation. 2 μl of isolated miRNAs were then retrotranscribed into a cDNA with the miRCURY LNA Universal RT microRNA PCR (Qiagen). A 1/60 cDNA dilution was used to perform each assay included in Serum/Plasma Focus microRNA PCR 384 wells Panels, (V4.M) (Qiagen) and to perform individual assays of the validation phase.
To ensure the quality of the results the samples used in the panels were selected according to two criteria: the efficiency of miRNA isolation and the level of haemolysis. We analyzed miR-451a and miR-23a-3p as haemolysis markers.
The study was designed in two steps: a discovery phase and a validation phase.
In the first step or discovery phase, we analyzed expression levels of 185 miRNA included in a PCR panel in 7 ALS patients (Table 1, in bold) and 6 age-matched healthy controls, to identify differentially expressed candidate miRNAs.
In the validation phase, the significantly deregulated miRNAs were analyzed in a larger cohort of patients and controls (53 ALS patients and 23 healthy controls).
The results of miRNAs expression were evaluated using Applied Biosystems SDS 2.4 software and the results were transformed into an Excel file. Mean and SD were calculated and statistical analysis was performed using Gene-Globe Data Analysis Center, an online platform provided by Qiagen. The p values were calculated based on a Student’s t-test of the replicate 2^(-ΔCt) values for each gene in the control group and in the ALS group. Samples were normalized using the global Ct mean of expressed miRNAs. Whether an miRNA was expressed or not was determined by the lower limit of detection that was set up to Ct 37.
Discovery phase
The miRNAs profile was studied by qPCR using Serum/Plasma Focus microRNA PCR 384 wells Panels, (V4.M) (Qiagen). This tool includes 185 different miRNA oligonucleotides, all of which were analyzed in each patient (n = 7) and healthy control (n = 6).
The deregulated miRNAs between ALS patients and controls found in the discovery phase were assessed in a second step or validation phase, which included 53 ALS patients and 23 healthy controls (Table 1).
The results were analyzed using software developed by Anaxomics (Barcelona, Spain).
Validation phase
For the validation phase, the 13 deregulated miRNAs identified in the discovery phase were evaluated in serum samples from 23 healthy controls and 53 ALS patients. miRCURY LNA miRNAs PCR Assays (Qiagen) were used for each of the miRNAs, and amplified fragments were detected using ExiLENT SYBR Green master mix (Qiagen). In total, we studied 13 candidate miRNAs (miR-107, miR-142-3p, miR-30c-5p, miR-335-5p, miR-421, miR-423-3p, miR-454-3p, miR-7a-5p, miR-122-5p, miR-125a-5p, miR-30b-5p, miR-30e-5p, miR-2110), 2 haemolysis controls (miR-451a and miR-23a-3p), and 4 exogenous controls (spike-ins UniSp2, UniSp4, UniSp5 and UniSp6), as described above.
SH-SY5Y cell culture
Human neuroblastoma cell line SH-SY5Y was obtained from the general collection of the European Collection of Authenticated Cell Culture (ECACC n°9403034) and purchased at Sigma-Aldrich (St. Louis, MI, USA). SH-SY5Y cells were seeded on culture well plates which were coated with 0.15% gelatin (Sigma-Aldrich) for 30 minutes. SH-SY5Y were grown in proliferation media (EMEM (Lonza Group Ltd, Basel, Switzerland):F12 medium (Sigma-Aldrich) (1:1) supplemented with 15% FBS (Lonza), 2 mM glutamine (Lonza), 1 mM sodium pyruvate (Lonza) and penicillin-streptomycin (Lonza).
SH-SY5Y transient transfection
For transfection experiments, SH-SY5Y cells were seeded at 20,000 cells/cm2.
We transfected 10 nM of IDT miRNA Inhibitor (Integrated DNA Technologies (IDT), Newark, NJ, USA), specific for miR-335-5p (miR-335 inhibitor). This oligonucleotide is perfectly complementary to the mature miRNA target. As a negative control (NC) we used a validated sequence (5′-ucguuaaucggcuauaauacgc-3′) developed by IDT.
To test mitophagy, we co-transfected the miRNA-335 inhibitor together with mKeima-Red-Mito-7 (a gift from Michael Davidson (Addgene plasmid # 56018; http://n2t.net/addgene:56018; RRID:Addgene_56018)). As a negative control, we co-transfected NC vector with mKeima-Red-Mito-7. MKeima-Red-Mito-7 contains a mitochondrial targeting sequence corresponding to the subunit VIII of human cytochrome C oxidase. The transfection reagent used was Fugene HD (Promega, Madison, WI, USA). The fluorescence was measured with Zeiss Zen software (Zeiss, Oberkochen, Germany), and the ratio 561/458 nm excited Keima fluorescence was calculated to reflect the underlying level of mitophagy. Images were acquired with a Leica Confocal microscope (Leica, Wetxlar, Germany).
Mitochondrial staining and Immunohistochemistry
To study mitochondrial morphology we stained transfected cells with MitoTracker Red CMXRos (Thermo Fisher Scientific), a red-fluorescent dye that stains mitochondria in living cells.
After mitotracker incubation, transfected cells were permeabilized with blocking goat solution 5% together with Triton X-100 0.1% (Sigma-Aldrich) during 30 minutes. Cells were incubated with p62 antibody (1/200 dilution) (Enzo LifeSciences, NY, USA, Cat# BML-PW9860, RRID:AB_2196009) and goat-anti-rabbit Alexa 488 (1/500 dilution) (Molecular Probes, Eugene, OR, USA, Cat# A-11008, RRID:AB_143165) was used as secondary antibody. Images were acquired with a Leica Confocal microscope (Leica).
Oxidative stress detection
To measure oxidative stress in transfected cells we used CellROX Green Reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s instructions. The cell-permeant dye is weakly fluorescent while in a reduced state but exhibits bright green photostable fluorescence upon oxidation by reactive oxygen species (ROS).
Superoxide dismutase (SOD) activity
We measured SOD activity in transfected cell samples (NC1 and miR-335 inhibitor) at 24, 48 and 72 h, using the SOD colorimetric activity kit (Invitrogen) following the manufacturer’s instructions. We used curve-fitting software to generate standard curve because a four parameter algorithm provides the best standard curve fit (https://www.myassays.com).
Caspase 3/7 assay
The caspase activity assay (Caspase-Glo 3/7 assay, Promega) was performed at 24, 48 and 72 h after transfection of SH-SY5Y cells with miR-335 inhibitor and following the manufacturer’s instructions.
Western-Blot
WB was performed as previously described46. Protein extraction was performed with RIPA buffer (Sigma-Aldrich). 20 μg of protein was loaded in each well. Gels were transferred to nitrocellulose transfer membranes. Unspecific binding sites on the blots were blocked by incubation for 1 hour in Odyssey Blocking Buffer (LI-COR Biosciences, Lincoln, NE, USA) diluted 1:1 in phosphate buffer saline (PBS). The blots were incubated for 1 hour with the primary antibodies rabbit anti-caspase-3 (1/250 dilution) (Genetex, CA, USA, Cat#GTX110543, RRID:AB_10722709), mouse monoclonal anti-caspase-7 (1/250 dilution) (Thermo Fisher Scientific, Cat#MA1-91896, RRID:AB_2068163) and rabbit anti-p62 (1/250 dilution)(Enzo LifeSciences, Cat# BML-PW9860, RRID:AB_2196009). To normalize the results, mouse anti-β-actin (1/1000 dilution) (Sigma-Aldrich, Cat# A5441, RRID:AB_476744) was added simultaneously with the primary antibodies. The secondary antibodies used were goat anti-rabbit labeled with IR-Dye 680 (1/5000 dilution) (LI-COR Biosciences Cat# 926-32221, RRID:AB_621841) and goat anti-mouse labeled with IR-Dye 800 (1/5000 dilution) (LI-COR Biosciences Cat# 926-32210, RRID:AB_621842). After extensive washing, the immunoreactive bands were visualized and quantified using the Odyssey Infrared Imaging System (LI-COR).
Statistical analysis
Experimental data were processed using SPSS Statistics 21.0 software (IBM Corp. Armonk, NY, USA). Normality and homogeneity of variance tests were performed. The data conforming to normal distribution and homogeneity of variance were expressed as mean ± standard deviation and analyzed with t-test45.
P values of <0.05 are indicated with one asterisk (*) and P values of less or equal than 0.01 are indicated with two asterisks (**).
Acknowledgements
The authors thank the patients and their relatives for their support for this study. The study was supported by grants from the Fondo de Investigación Sanitaria (FIS), Instituto de Salud Carlos III, Spanish Ministry of Health (PI15/1618 and PI19/01543 to R.R.), and partly jointly funded by Fondo Europeo de Desarrollo Regional (FEDER), Unión Europea, Una manera de hacer Europa. This work was also supported by Fundació La Marató de TV3 (201437.10 to R.R.). I. I-G is supported by the Rio Hortega grant (CM17/00074) from “Acción Estratégica en Salud 2013-2016” and the European Social Fund and the Global Brain Health Institute (Atlantic fellow for equity and brain health at GBHI; https://www.gbhi.org/).
Author contributions
N.d.L. and R.R. were responsible for the conception and design of the study. Patients were visited by J.T.-S., E.C.-V., A.L.L., I.I.-G. and R.R. Data acquisition was performed by N.d.L., O.D.-I. and A.C.-R. Data analysis was done by N.d.L., J.T.-S., J.C., A.L.L. and R.R., N.d.L., E.G., I.I. and R.R. drafted the manuscript and figures.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Noemi De Luna, Email: nluna@santpau.cat.
Ricardo Rojas-García, Email: rrojas@santpau.cat.
References
- 1.Brown, R. H. & Al-Chalabi, A. Amyotrophic Lateral Sclerosis. N Engl J Med. 377(2), 162–72 (2017 Jul 13). [DOI] [PubMed]
- 2.Geser, F. et al. Evidence of multisystem disorder in whole-brain map of pathological TDP-43 in amyotrophic lateral sclerosis. Arch Neurol. 65(5), 636–41 (2008 May). [DOI] [PubMed]
- 3.Wilson, C. M., Grace, G. M., Munoz, D. G., He, B. P. & Strong, M. J. Cognitive impairment in sporadic ALS: a pathologic continuum underlying a multisystem disorder. Neurology. 57(4), 651–7 (2001 Aug 28). [DOI] [PubMed]
- 4.Gunnarsson, L. G. & Bodin, L. Amyotrophic Lateral Sclerosis and Occupational Exposures: A Systematic Literature Review and Meta-Analyses. Int J Environ Res Public Health. 15(11) (2018 Oct 26). [DOI] [PMC free article] [PubMed]
- 5.Oskarsson, B., Gendron, T. F. & Staff, N. P. Amyotrophic Lateral Sclerosis: An Update for 2018. Mayo Clin Proc. 93(11), 1617–28 (2018 Nov). [DOI] [PubMed]
- 6.Al-Chalabi, A. et al. The genetics and neuropathology of amyotrophic lateral sclerosis. Acta Neuropathol. 124(3), 339–52 (2012 Sep). [DOI] [PubMed]
- 7.Butti Z, Patten SA. RNA Dysregulation in Amyotrophic Lateral Sclerosis. Front. Genet. 2018;9:712. doi: 10.3389/fgene.2018.00712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Dardiotis, E. et al. The Role of MicroRNAs in Patients with Amyotrophic Lateral Sclerosis. J Mol Neurosci. 66(9), 617–28 (2018 Nov 10). [DOI] [PubMed]
- 9.Chen, P. Y. & Meister, G. microRNA-guided posttranscriptional gene regulation. Biol Chem. 386(12), 1205–18 (2005 Dec). [DOI] [PubMed]
- 10.Gebert, L. F. R. & MacRae, I. J. Regulation of microRNA function in animals. Nat Rev Mol Cell Biol. 20(1), 21–37 (2018 Aug 14). [DOI] [PMC free article] [PubMed]
- 11.Goodall, E. F., Heath, P. R., Bandmann, O., Kirby, J. & Shaw, P. J. Neuronal dark matter: the emerging role of microRNAs in neurodegeneration. Front Cell Neurosci. 7, 178 (2013 Oct 10). [DOI] [PMC free article] [PubMed]
- 12.Esteller, M. Non-coding RNAs in human disease. Nat Rev Genet. 12(12), 861–74 (2011 Nov 18). [DOI] [PubMed]
- 13.Kye MJ, Goncalves Ido C. The role of miRNA in motor neuron disease. Front. Cell Neurosci. 2014;8:15. doi: 10.3389/fncel.2014.00015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Volonte C, Apolloni S, Parisi C. MicroRNAs: newcomers into the ALS picture. CNS Neurol. Disord. Drug. Targets. 2015;14(2):194–207. doi: 10.2174/1871527314666150116125506. [DOI] [PubMed] [Google Scholar]
- 15.Gandhi, S. et al. PINK1-associated Parkinson’s disease is caused by neuronal vulnerability to calcium-induced cell death. Mol Cell. 33(5), 627–38 (2009 Mar 13). [DOI] [PMC free article] [PubMed]
- 16.Kim SM, et al. Intermittent hypoxia can aggravate motor neuronal loss and cognitive dysfunction in ALS mice. PLoS One. 2013;8(11):e81808. doi: 10.1371/journal.pone.0081808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Manfredi, G. & Xu, Z. Mitochondrial dysfunction and its role in motor neuron degeneration in ALS. Mitochondrion. 5(2), 77–87 (2005 Apr). [DOI] [PubMed]
- 18.Kroemer, G. & Reed, J. C. Mitochondrial control of cell death. Nat Med. 6(5), 513–9 (2000 May). [DOI] [PubMed]
- 19.Panchal, K. & Tiwari, A. K. Mitochondrial dynamics, a key executioner in neurodegenerative diseases. Mitochondrion. 47, 151-73 (2018 Nov 5). [DOI] [PubMed]
- 20.Cozzolino, M. & Carri, M. T. Mitochondrial dysfunction in ALS. Prog Neurobiol. 97(2), 54–66 (2012 May). [DOI] [PubMed]
- 21.Federico, A. et al. Mitochondria, oxidative stress and neurodegeneration. J Neurol Sci. 322(1–2), 254–62 (2012 Nov 15). [DOI] [PubMed]
- 22.Zorov, D. B., Juhaszova, M. & Sollott, S. J. Mitochondrial reactive oxygen species (ROS) and ROS-induced ROS release. Physiol Rev. 94(3), 909–50 (2014 Jul). [DOI] [PMC free article] [PubMed]
- 23.Burton, G. J. & Jauniaux, E. Oxidative stress. Best Pract Res Clin Obstet Gynaecol. 25(3), 287–99 (2011 Jun). [DOI] [PMC free article] [PubMed]
- 24.Kalogeris T, Bao Y, Korthuis RJ. Mitochondrial reactive oxygen species: a double edged sword in ischemia/reperfusion vs preconditioning. Redox Biol. 2014;2:702–14. doi: 10.1016/j.redox.2014.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Jacobson, J. & Duchen, M. R. Mitochondrial oxidative stress and cell death in astrocytes–requirement for stored Ca2+ and sustained opening of the permeability transition pore. J Cell Sci. 115(Pt 6), 1175–88 (2002 Mar 15). [DOI] [PubMed]
- 26.Taylor, J. P., Brown, R. H. Jr. & Cleveland, D. W. Decoding ALS: from genes to mechanism. Nature. 539(7628), 197–206 (2016 Nov 10). [DOI] [PMC free article] [PubMed]
- 27.Kogure, T., Kawano, H., Abe, Y. & Miyawaki, A. Fluorescence imaging using a fluorescent protein with a large Stokes shift. Methods. 45(3), 223–6 (2008 Jul). [DOI] [PubMed]
- 28.Hamzeiy H, et al. Visualization and Analysis of miRNAs Implicated in Amyotrophic Lateral Sclerosis Within Gene Regulatory Pathways. Stud. Health Technol. Inform. 2018;253:183–7. [PubMed] [Google Scholar]
- 29.Luchetti, A. et al. A Perturbed MicroRNA Expression Pattern Characterizes Embryonic Neural Stem Cells Derived from a Severe Mouse Model of Spinal Muscular Atrophy (SMA). Int J Mol Sci. 16(8), 18312–27 (2015 Aug 6). [DOI] [PMC free article] [PubMed]
- 30.Murdocca, M. et al. SMA Human iPSC-Derived Motor Neurons Show Perturbed Differentiation and Reduced miR-335-5p Expression. Int J Mol Sci. 17(8), e1231 (2016 Jul 30). [DOI] [PMC free article] [PubMed]
- 31.Capitano, F. et al. MicroRNA-335-5p modulates spatial memory and hippocampal synaptic plasticity. Neurobiol Learn Mem. 139, 63–8 (2017 Mar). [DOI] [PubMed]
- 32.Waller, R. et al. Serum miRNAs miR-206, 143-3p and 374b-5p as potential biomarkers for amyotrophic lateral sclerosis (ALS). Neurobiol Aging. 55:123–31 (2017 Jul). [DOI] [PMC free article] [PubMed]
- 33.Raheja, R. et al. Correlating serum micrornas and clinical parameters in amyotrophic lateral sclerosis. Muscle Nerve. 58(2), 261–9 (2018 Aug). [DOI] [PMC free article] [PubMed]
- 34.Golpich, M. et al. Mitochondrial Dysfunction and Biogenesis in Neurodegenerative diseases: Pathogenesis and Treatment. CNS Neurosci Ther. 23(1), 5–22 (2017 Jan). [DOI] [PMC free article] [PubMed]
- 35.Katsuragi, Y., Ichimura, Y. & Komatsu, M. p62/SQSTM1 functions as a signaling hub and an autophagy adaptor. FEBS J. 282(24), 4672–8 (2015 Dec). [DOI] [PubMed]
- 36.Ivankovic, D., Chau, K. Y., Schapira, A. H. & Gegg, M. E. Mitochondrial and lysosomal biogenesis are activated following PINK1/parkin-mediated mitophagy. J Neurochem. 136(2), 388–402 (2016 Jan). [DOI] [PMC free article] [PubMed]
- 37.Halter, B. et al. Oxidative stress in skeletal muscle stimulates early expression of Rad in a mouse model of amyotrophic lateral sclerosis. Free Radic Biol Med. 48(7), 915–23 (2010 Apr 1). [DOI] [PubMed]
- 38.Jokic, N. et al. Early activation of antioxidant mechanisms in muscle of mutant Cu/Zn-superoxide dismutase-linked amyotrophic lateral sclerosis mice. Ann N Y Acad Sci. 1010, 552–6 (2003 Dec). [DOI] [PubMed]
- 39.Hoye, A. T., Davoren, J. E., Wipf, P., Fink, M. P. & Kagan, V. E. Targeting mitochondria. Acc Chem Res. 41(1), 87–97 (2008 Jan). [DOI] [PubMed]
- 40.Lamkanfi, M. & Kanneganti, T. D. Caspase-7: a protease involved in apoptosis and inflammation. Int J Biochem Cell Biol. 42(1), 21–4 (2010 Jan). [DOI] [PMC free article] [PubMed]
- 41.Berning BA, Walker AK. The Pathobiology of TDP-43 C-Terminal Fragments in ALS and FTLD. Front. Neurosci. 2019;13:335. doi: 10.3389/fnins.2019.00335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Brooks, B. R., Miller, R. G., Swash, M. & Munsat, T. L. El Escorial revisited: revised criteria for the diagnosis of amyotrophic lateral sclerosis. Amyotroph Lateral Scler Other Motor Neuron Disord. 1(5), 293–9 (2000 Dec). [DOI] [PubMed]
- 43.Illan-Gala, I. et al. CSF sAPPbeta, YKL-40, and NfL along the ALS-FTD spectrum. Neurology. 91(17), e1619–e28 (2018 Oct 23). [DOI] [PubMed]
- 44.Rascovsky, K. et al. Sensitivity of revised diagnostic criteria for the behavioural variant of frontotemporal dementia. Brain. 134(Pt 9), 2456–77 (2011 Sep). [DOI] [PMC free article] [PubMed]
- 45.Carrasco-Rozas, A. et al. Identification of serum microRNAs as potential biomarkers in Pompe disease. Ann Clin Transl Neurol. 6(7), 1214–24 (2019 Jul). [DOI] [PMC free article] [PubMed]
- 46.De Luna, N et al. Effect of MAPK Inhibition on the Differentiation of a Rhabdomyosarcoma Cell Line Combined With CRISPR/Cas9 Technology: An In Vitro Model of Human Muscle Diseases. J Neuropathol Exp Neurol. 77(10), 964–72 (2018 Oct 1). [DOI] [PubMed]







