Abstract
In this study, we evaluated the function and regulation of the long non-coding RNA (lncRNA) FAM83H-AS1 in triple-negative breast cancer (TNBC). Our data show that the FAM83H-AS1 levels are increased in human TNBC cells and tissues. Proliferation, migration, and invasion of TNBC cells are decreased by FAM83H-AS1 suppression, but increased by FAM83H-AS1 overexpression. Bioinformatics analysis revealed that miR-136-5p is a potential target of FAM83H-AS1. MiR-136-5p expression is decreased in TNBC tissues, and its overexpression suppresses TNBC cell proliferation, migration, and invasion. MiR-136-5p suppression reverses the FAM83H-AS1 silencing-mediated inhibition of TNBC cell proliferation, migration, and invasion, suggesting that FAM83H-AS1 exerts its oncogenic effect by inhibiting miR-136-5p. Our data identify metadherin (MTDH) as the target gene of miR-136-5p, and demonstrate that the MTDH expression is increased in human TNBC tissues, which induces proliferation, migration, and invasion of TNBC cells. Importantly, our in vivo data show that FAM83H-AS1 also promotes tumor growth in TNBC mouse xenografts. Together, our results demonstrate that FAM83H-AS1 functions as an oncogenic lncRNA that regulates miR-136-5p and MTDH expression during TNBC progression, and suggest that targeting the FAM83H-AS1/miR-136-5p/MTDH axis may serve as a novel therapeutic target in TNBC.
Keywords: triple-negative breast cancer (TNBC), FAM83H-AS1, miR-136-5p, metadherin (MTDH)
INTRODUCTION
Triple-negative breast cancer (TNBC) cells lack expression of estrogen receptor, progesterone receptor, and human epidermal growth factor receptor 2 (HER2). TNBC accounts for approximately 15% of invasive breast cancers, and is the most aggressive breast cancer subtype with a poor prognosis. While many patients with other breast cancer subtypes have benefited from targeted therapies, due to the lack of a definitive molecular therapeutic target, there have been no specific targeted drugs available for patients with TNBC. Conventional chemotherapy remains the standard of care for patients with advanced TNBC [1, 2]. Thus, it is imperative to identify the underlying mechanisms, and develop diagnostic and prognostic biomarkers for TNBC treatment.
Long noncoding RNAs (lncRNAs) are transcripts longer than 200 nucleotides without a protein-coding potential [3]. LncRNAs exert pivotal roles in physiological and pathological processes, including organ development, cell fate, and stemness maintenance [4–7]. Dysregulation of lncRNAs has been implicated in cancer cell proliferation, invasion, and metastasis [8–10]. LncRNA GClnc1 promotes gastric cancer progression and is associated with a poor prognosis in gastric cancer [10], while Lnc-UCID promotes hepatocellular carcinoma (HCC) tumorigenesis and correlates with HCC progression [11]. Elevated lncRNA TTTY15 promotes prostate cancer progression through targeting miRNA let-7 [12], while lncRNA n384546 promotes thyroid papillary cancer cell proliferation, invasion, and migration by regulating miR-145-5p /AKT3 signaling [13].
Several dysregulated lncRNAs have been implicated also in breast cancer progression. For example, LINC00511 correlates with a poor prognosis in breast cancer, and promotes breast cancer stemness and tumorigenesis via miR-185-3p/E2F1 [14]. LncRNA DANCR promotes breast cancer progression by regulating miR-216a-5p [15]. FOXD2-AS1 facilitates breast cancer cell proliferation, migration, and invasion by regulating miR-150-5p/PFN2 [16]. Moreover, the lncRNAs HCP5 [17], HOTAIR [18], NRAD1 [19], MIR503HG [20], and NAMPT-AS [21] are dysregulated and involved in TNBC progression. However, the function of the lncRNA FAM83H antisense RNA 1, FAM83H-AS1, is still unknown in TNBC.
FAM83H-AS1, also known as onco-lncRNA-3, is located on chromosome 8 (8q24.3), and has 2743 base pairs. FAM83H-AS1 has been shown to function as an oncogene in several human cancers. For example, FAM83H-AS1 is associated with worse survival rates in cervical cancer patients, and its inhibition decreases proliferation, migration, and survival of cervical cancer cells [22]. FAM83H-AS1 promotes radio-resistance and metastasis via targeting HuR protein in ovarian cancer [23]. FAM83H-AS1 also contributes to bladder cancer cell proliferation, migration, and invasion [24], and to glioma progression by epigenetically silencing CDKN1A (p21) [25]. FAM83H-AS1 correlates with poor prognosis in colorectal cancer, and promotes colorectal cancer cell proliferation by targeting the Notch signaling [26]. Moreover, FAM83H-AS1 contributes to lung cancer progression via regulating the MET/EGFR signaling [27]. In breast cancer, Yang et al have found that FAM83H-AS1 is upregulated in the luminal subtype of breast cancer [28]. In addition, the expression of FAM83H-AS1 is increased and correlates with poor survival rates in patients with early-stage breast cancer [29]. However, the function and the underlying mechanisms of FAM83H-AS1 in TNBC progression remain unknown.
In the present study, we investigated the role and the regulation of FAM83H-AS1 during TNBC progression. Our results demonstrate that the expression of FAM83H-AS1 is increased in TNBC cells and tissues, and promotes TNBC cell proliferation, migration, and invasion by regulating miR-136-5p and metadherin (MTDH) expression. These data indicate that the FAM83H-AS1/miR-136-5p/MTDH signaling might represent a potential therapeutic target in managing TNBC.
RESULTS
FAM83H-AS1 is upregulated in human TNBC tissues
First, we analyzed the expression of FAM83H-AS1 in human breast cancer tissues. Online data from the Gene Expression Profiling Interactive Analysis 2 (GEPIA2) (http://gepia2.cancer-pku.cn/#index) database showed that the expression of FAM83H-AS1 is increased in different types of cancer, including breast cancer (Supplementary Figure 1A, 1B, p < 0.05). Moreover, high FAM83H-AS1 levels are associated with a poor overall survival of breast cancer patients (Supplementary Figure 1C, 1D). The data analysis from cBioPortal revealed that 21% of breast cancer samples contain gene amplification of FAM83H-AS1 (Supplementary Figure 1E).
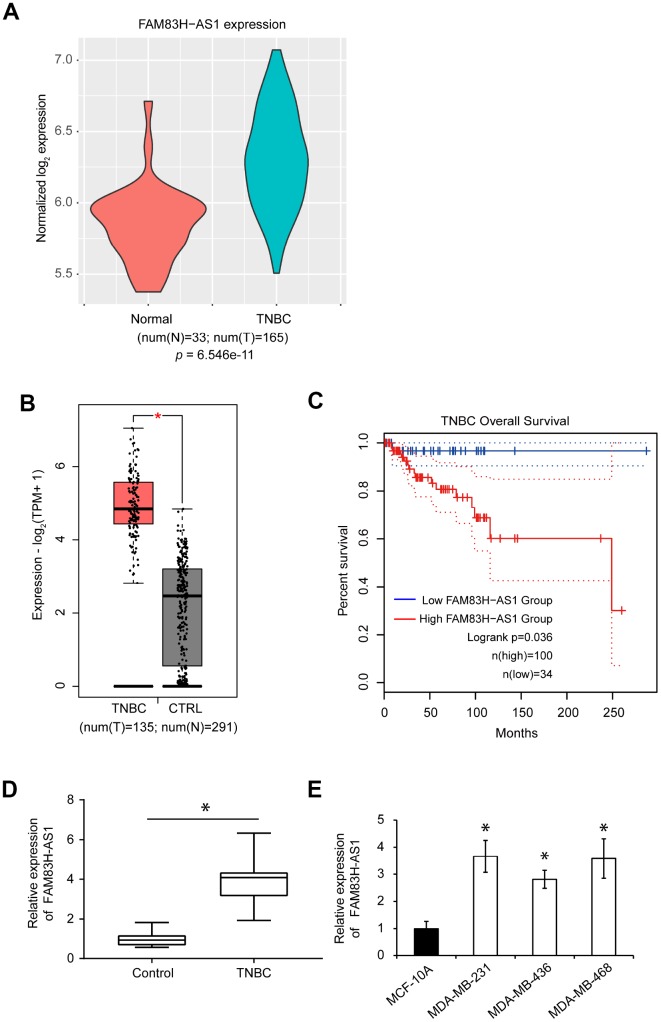
Next, we analyzed the expression of FAM83H-AS1 in human TNBC tissues. The human lncRNA microarray dataset GSE76250 (containing 165 TNBC samples and 33 paired normal breast tissues) was downloaded using the Affymetrix Human Transcriptome Array 2.0 platform to analyze the expression profile of FAM83H-AS1 between TNBC and normal breast tissues. The expression of FAM83H-AS1 was significantly upregulated in TNBC compared to normal tissues (Figure 1A). Analysis of the GEPIA2 database also showed that the expression of FAM83H-AS1 is increased in human TNBC compared to normal breast tissues (Figure 1B, p < 0.05). Moreover, the upregulated expression of FAM83H-AS1 was predictive of a poor overall survival in TNBC patients (Figure 1C, p < 0.05). Furthermore, qRT-PCR analysis confirmed the increased expression of FAM83H-AS1 in TNBC compared to adjacent normal tissues (Figure 1D, p < 0.05). In addition, analysis of FAM83H-AS1 expression in three different TNBC cell lines (MDA-MB-231, MDA-MB-436, and MDA-MB-468) showed that the FAM83H-AS1 levels are increased in TNBC cell lines compared to normal human mammary epithelial cell line MCF-10A (Figure 1E, p < 0.05).
Figure 1.
FAM83H-AS1 is upregulated in TNBC tissues and predicts worse overall survival. (A) Expression profiles of FAM83H-AS1 in TNBC and normal breast tissues using the human lncRNA microarray dataset GSE76250. The p value was calculated by Wilcoxon rank-sum test. (B) Expression profiles of FAM83H-AS1 in TNBC and normal breast tissues using the GEPIA 2 dataset. (C) Overall survival rates in low and high FAM83H-AS1 expression groups in TNBC patients using the GEPIA2 dataset. (D) qRT-PCR of FAM83H-AS1 expression in human TNBC and adjacent control tissues. (E) qRT-PCR of FAM83H-AS1 mRNA in MDA-MB-231, MDA-MB-436, MDA-MB-468, and MCF-10A cells.
FAM83H-AS1 promotes proliferation, migration, and invasion in TNBC cells
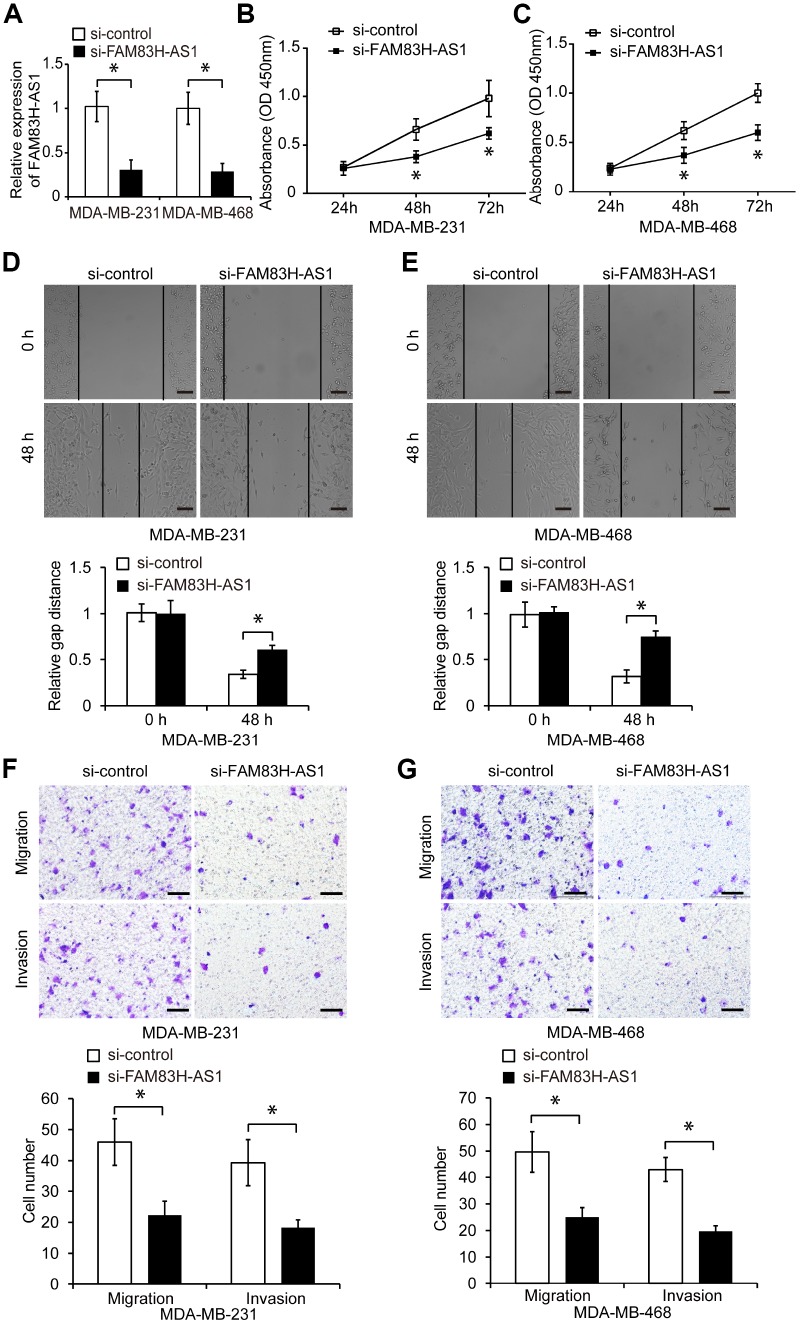
To investigate the function of FAM83H-AS1 in TNBC cells, we first suppressed the FAM83H-AS1 expression by specific siRNA in MDA-MB-231 and MDA-MB-468 cells (Figure 2A, p < 0.05). As shown in Figure 2B, 2C, FAM83H-AS1 suppression significantly reduced proliferation of TNBC cells measured by the CCK8 assay (p < 0.05). In addition, wound healing and transwell assays demonstrated that FAM83H-AS1 suppression markedly inhibited migration and invasion of TNBC cells compared to cells transfected with control siRNA (Figure 2D–2G, p < 0.05).
Figure 2.
FAM83H-AS1 suppression inhibits TNBC cell proliferation, migration, and invasion. (A) qRT-PCR of FAM83H-AS1 expression in TNBC cells transfected with si-control or si-FAM83H-AS1 RNA. (B, C) Proliferation of TNBC cells transfected with si-control or si-FAM83H-AS1 RNA, analyzed by CCK8 assay. (D, E) Wound healing assay of the migration capacity of MDA-MB-231 and MDA-MB-468 cells transfected with si-control or si-FAM83H-AS1. (F, G) Migration and invasion of MDA-MB-231 and MDA-MB-468 cells transfected with si-control or si-FAM83H-AS1. Scale bars, 100 μm. * p < 0.05 compared to controls.
Next, we overexpressed FAM83H-AS1 in TNBC cells using the pcDNA-FAM83H-AS1 or empty vector pcDNA-control plasmids (Supplementary Figure 2A, p < 0.05). Overexpression of FAM83H-AS1 promoted proliferation of TNBC cells (Supplementary Figure 2B, 2C, p < 0.05), and increased their migration and invasion (Supplementary Figure 2D–2G, p < 0.05). These results indicate that FAM83H-AS1 promotes proliferation, migration, and invasion of TNBC cells in vitro.
FAM83H-AS1 functions as a sponge for miR-136-5p in TNBC cells
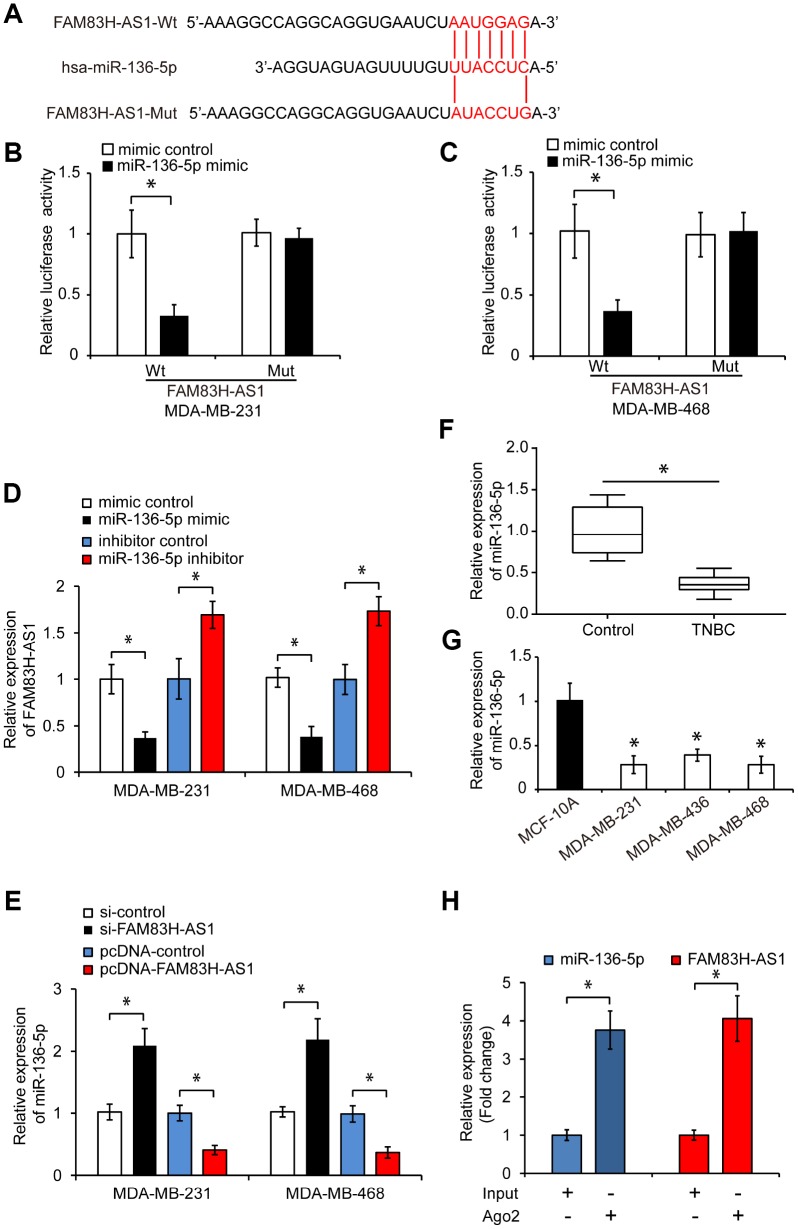
To explore the mechanisms underlying the pro-oncogenic effects of FAM83H-AS1 in TNBC cells, we searched for potential FAM83H-AS1-targeted miRNAs using the LncBase Predicted v.2 bioinformatics database. We found that miR-136-5p was a potential target of FAM83H-AS1 (Figure 3A). To confirm this prediction, we constructed a luciferase reporter containing wild-type (Wt) or mutated (Mut) binding sites for miR-136-5p in FAM83H-AS1. As shown in Figure 3B and 3C, miR-136-5p overexpression significantly inhibited the luciferase activity in TNBC cells transfected with FAM83H-AS1-Wt, but not in FAM83H-AS1-Mut transfected cells (p < 0.05). In addition, overexpression of miR-136-5p significantly inhibited the FAM83H-AS1 expression, while miR-136-5p knockdown promoted the FAM83H-AS1 expression in TNBC cells (Figure 3D, p < 0.05). Moreover, FAM83H-AS1 knockdown increased the miR-136-5p expression, while FAM83H-AS1 overexpression decreased the miR-136-5p expression in TNBC cells (Figure 3E, p < 0.05). Analysis of miR-136-5p levels in human TNBC cells and tissues by qRT-PCR revealed that the miR-136-5p expression is reduced in TNBC tissues and cell lines (Figure 3F–3G, p < 0.05). In addition, RNA immunoprecipitation demonstrated that FAM83H-AS1 and miR-136-5p were highly enriched in Ago2-containing beads compared to controls (Figure 3H, p < 0.05), indicating that FAM83H-AS1 directly binds to miR-136-5p. Together, these data suggest that FAM83H-AS1 functions as a sponge for miR-136-5p in TNBC cells.
Figure 3.
FAM83H-AS1 functions as a sponge for miR-136-5p. (A) Predicted binding site of miR-136-5p in the FAM83H-AS1 sequence and mutated nucleotides. (B, C) Overexpression of miR-136-5p repressed the luciferase activity in TNBC cells transfected with FAM83H-AS1-Wt assessed by luciferase reporter assays. (D) Relative FAM83H-AS1 expression in TNBC cells transfected with mimic control, miR-136-5p mimic, inhibitor control, or miR-136-5p inhibitor. (E) Relative miR-136-5p expression in TNBC cells transfected with si-control, si-FAM83H-AS1, pcDNA-control, or pcDNA-FAM83H-AS1. (F) qRT-PCR analysis of miR-136-5p expression in human TNBC and adjacent control tissues. (G) qRT-PCR analysis of miR-136-5p levels in TNBC cell lines MDA-MB-231, MDA-MB-436, and MDA-MB-468, and a control breast epithelial cell line MCF-10A. (H) RIP assay demonstrating the enrichment of FAM83H-AS1 and miR-136-5p. * p < 0.05 compared to controls.
MiR-136-5p inhibits proliferation, migration, and invasion in TNBC cells
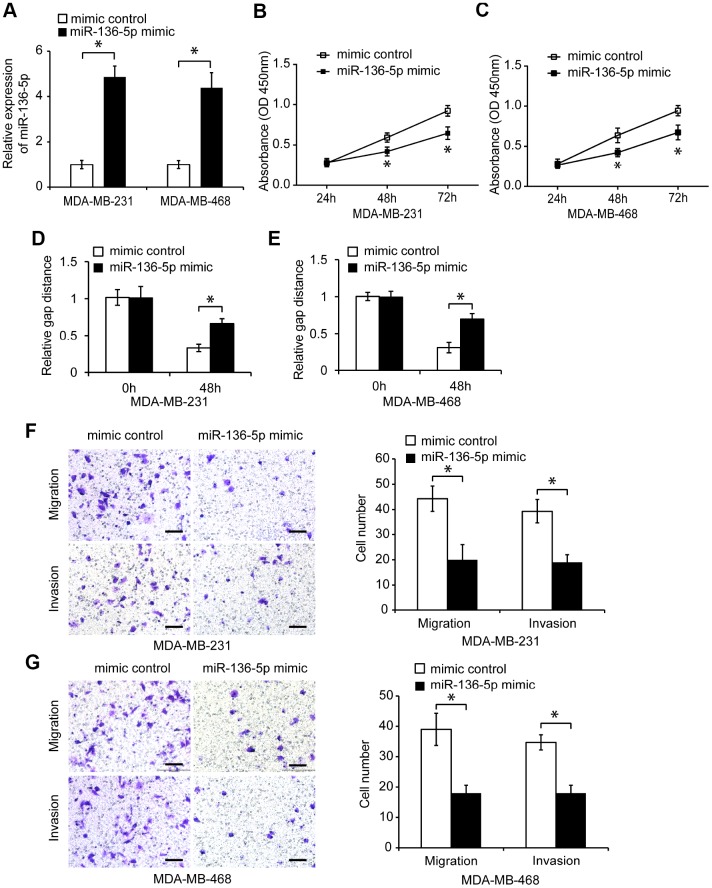
To investigate the function of miR-136-5p in TNBC cells, we first overexpressed miR-136-5p in MDA-MB-231 and MDA-MB-468 cells. MiR-136-5p mimic transfection significantly increased the endogenous miR-136-5p expression in TNBC cells (Figure 4A, p < 0.05), and reduced their proliferation (Figure 4B, 4C, p < 0.05). In addition, miR-136-5p overexpression significantly decreased migration and invasion of TNBC cells (Figure 4D–4G, p < 0.05). Conversely, transfection of miR-136-5p inhibitor decreased the expression of miR-136-5p in TNBC cells (Supplementary Figure 3A, p < 0.05), and promoted TNBC cell proliferation (Supplementary Figure 3B, 3C, p < 0.05). Moreover, the reduced miR-136-5p expression increased TNBC cell migration and invasion (Supplementary Figure 3D–3G, p < 0.05), indicating that miR-136-5p acts as a tumor suppressor in TNBC cells.
Figure 4.
Overexpression of miR-136-5p reduces TNBC cell proliferation, migration, and invasion. (A) Relative miR-136-5p expression in TNBC cells transfected with mimic control or miR-136-5p mimic. (B, C) Proliferation of TNBC cells transfected with mimic control or miR-136-5p mimic, analyzed by CCK8 assay. (D, E) Wound healing assay of the migration capacity of TNBC cells transfected with mimic control or miR-136-5p mimic. (F, G) Migration and invasion of TNBC cells transfected with mimic control or miR-136-5p mimic, analyzed by transwell assays. Scale bars, 100 μm. * p < 0.05 compared to controls.
FAM83H-AS1 exerts its pro-oncogenic function in TNBC cells by inhibiting miR-136-5p
To determine whether FAM83H-AS1 exerts its pro-oncogenic function in TNBC cells via inhibiting miR-136-5p, we first knocked down miR-136-5p expression in FAM83H-AS1-silenced TNBC cells (Figure 5A, p < 0.05), and then analyzed their proliferation, migration and invasion. As shown in Figure 5B–5G, miR-136-5p suppression blocked the suppressive effect of FAM83H-AS1 knockdown on TNBC cell proliferation, migration, and invasion (p < 0.05). These results show that FAM83H-AS1 promotes cell proliferation, migration, and invasion through inhibiting miR-136-5p in TNBC cells.
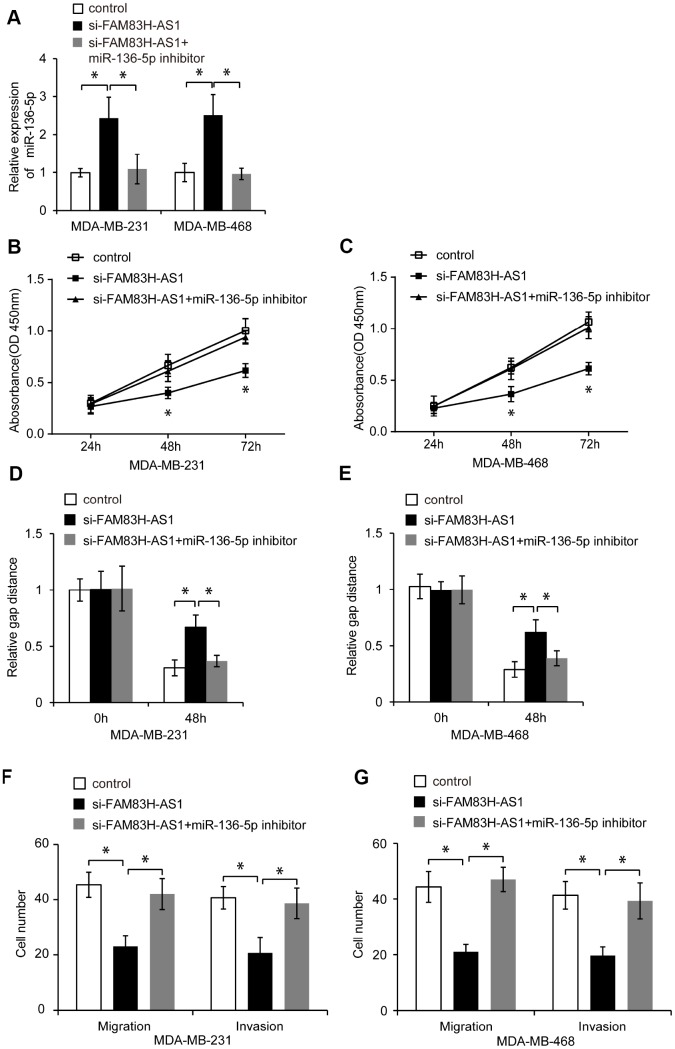
Figure 5.
FAM83H-AS1 promotes TNBC cell proliferation, migration, and invasion through inhibiting miR-136-5p. (A) Relative miR-136-5p expression in TNBC cells transfected with control, si- FAM83H-AS1, and si-FAM83H-AS1+miR-136-5p inhibitor evaluated by qRT-PCR. (B, C) Proliferation of TNBC cells transfected with control, si-FAM83H-AS1, and si-FAM83H-AS1+miR-136-5p inhibitor, analyzed by CCK8 assay. (D, E) Wound healing assay of the migration of MDA-MB-231 and MDA-MB-468 cells transfected with control, si-FAM83H-AS1, and si-FAM83H-AS1+miR-136-5p inhibitor. (F, G) Migration and invasion of TNBC cells transfected with control, si-FAM83H-AS1, and si-FAM83H-AS1+miR-136-5p inhibitor, assessed by transwell assays. * p < 0.05 compared to controls.
MiR-136-5p suppresses metadherin expression in TNBC cells
We searched for the potential target genes of miR-136-5p through bioinformatics analysis using miRPathDB and miRTarBase. We found that two genes, BCL2 and metadherin (MTDH), were the potential targets of miR-136-5p (Supplementary Figure 4A). However, since transfection with miR-136-5p mimic did not have any significant effect on BCL2 expression in TNBC cells (Supplementary Figure 4B), the BCL2 gene was excluded from further analysis. Using bioinformatics analysis, we found that there is a putative binding site for miR-136-5p in the 3’-UTR of MTDH mRNA (Figure 6A). Online data analysis using cBioPortal and GEPIA 2 showed an increased MTDH expression in breast cancer tissues, including the TNBC subtype (Supplementary Figure 5A–5C, Figure 6B, p < 0.05). The increased expression of MTDH in breast cancer tissues was validated by gene mutation analysis using the cBioPortal data (Supplementary Figure 5D, 5E). The elevated MTDH expression was associated with poor overall survival rates in breast cancer patients, based on the cBioPortal and GEPIA 2 data analyses (Supplementary Figure 5F, 5G).
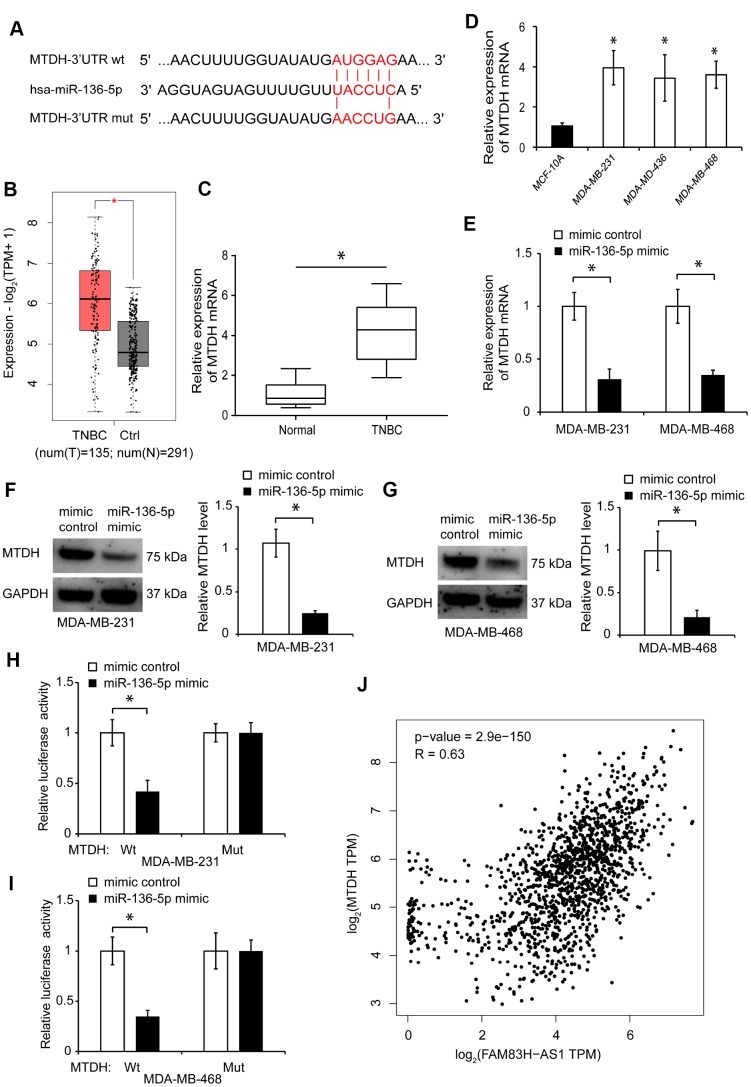
Figure 6.
MiR-136-5p suppresses MTDH expression in TNBC cells. (A) Predicted binding site of miR-136-5p in MTDH sequence and mutated nucleotides. (B) MTDH expression in TNBC tissues analyzed using GEPIA 2 dataset. (C) qRT-PCR analysis of MTDH expression in human TNBC tissues. (D) qRT-PCR analysis of MTDH expression in MDA-MB-231, MDA-MB-436, MDA-MB-468, and MCF-10A cells. (E) MTDH expression in TNBC cells transfected with mimic control or miR-136-5p mimic. (F, G) Western blot analysis of MTDH protein levels in TNBC cells transfected with mimic control or miR-136-5p mimic. (H, I) Overexpression of miR-136-5p represses the luciferase activity in TNBC cells transfected with MTDH-Wt, evaluated by luciferase reporter assays. (J) Correlation between FAM83H-AS1 and MTDH expression in breast cancer tissues analyzed by Spearman’s rank test using the GEPIA 2 dataset. * p < 0.05 compared to controls.
In addition, using qRT-PCR, we found that the MTDH gene expression is increased in TNBC tissues and cell lines (Figure 6C, 6D, p < 0.05). Furthermore, we found that miR-136-5p overexpression inhibits the MTDH expression in TNBC cells (Figure 6E–6G, p < 0.05). To determine whether MTDH binds to miR-136-5p, we constructed luciferase reporters containing Wt or Mut miR-136-5p binding sites. Overexpression of miR-136-5p significantly inhibited the luciferase activity in TNBC cells transfected with MTDH-Wt, but not MTDH-Mut (Figure 6H, 6I, p < 0.05). Moreover, analysis of the GEPIA 2 database indicated that the expression of MTDH positively correlates with the level of FAM83H-AS1 in breast cancer tissues (Figure 6J). These findings indicate that miR-136-5p suppresses the MTDH expression in TNBC cells.
MiR-136-5p exerts its tumor-suppressive function in TNBC cells by suppressing MTDH
We investigated whether miR-136-5p exerts its tumor-suppressive function in TNBC cells by suppressing the MTDH expression. To examine the function of MTDH in TNBC cells, we suppressed the endogenous MTDH expression by using MTDH specific si-RNA (Supplementary Figure 6A, p < 0.05). MTDH knockdown markedly suppressed proliferation (Supplementary Figure 6B, 6C, p < 0.05), migration, and invasion of TNBC cells compared to control siRNA (Supplementary Figure 6D–6G, p < 0.05).
Next, we overexpressed MTDH in miR-136-5p mimic-transfected TNBC cells via pcDNA-MTDH (oeMTDH) transfection. The transfection efficiency in control, miR-136-5p mimic, and miR-136-5p mimic plus oeMTDH groups was analyzed by qRT-PCR and western blotting (Figure 7A–7C, p < 0.05). Elevated MTDH expression abolished the inhibitory effect of miR-136-5p overexpression on TNBC cell proliferation, migration, and invasion (Figure 7D–7I, p < 0.05), suggesting that miR-136-5p exerts its tumor-suppressive function in TNBC cells by suppressing the MTDH expression.
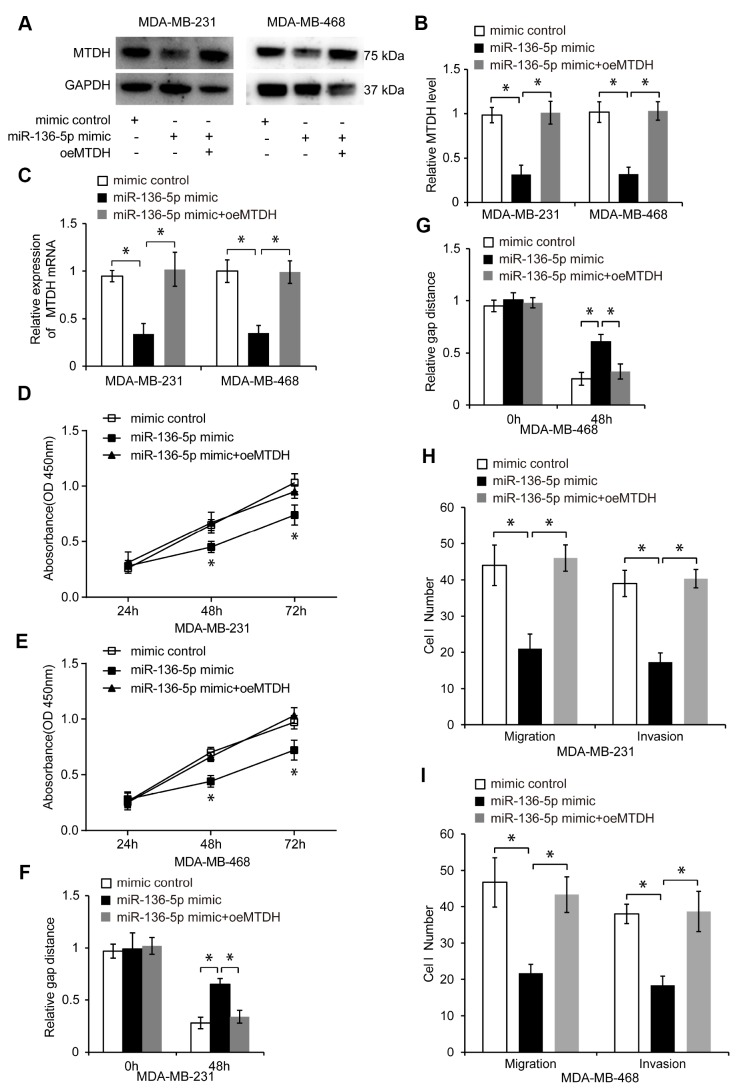
Figure 7.
MiR-136-5p inhibits proliferation, migration, and invasion of TNBC cells through suppressing MTDH. (A, B) Western blot analyses in TNBC cells transfected with mimic control, miR-136-5p mimic, or miR-136-5p mimic plus oeMTDH. (C) Expression of MTDH in TNBC cells transfected with mimic control, miR-136-5p mimic, or miR-136-5p mimic plus oeMTDH. (D, E) CCK8 assay utilized to evaluate cell proliferation of TNBC cells transfected with mimic control, miR-136-5p mimic, or miR-136-5p mimic plus oeMTDH. (F, G) Wound healing assay used to determine migration of TNBC cells transfected with mimic control, miR-136-5p mimic, or miR-136-5p mimic plus oeMTDH. (H, I) Migration and invasion of TNBC cells transfected with mimic control, miR-136-5p mimic, or miR-136-5p mimic plus oeMTDH, analyzed by transwell assays. * p < 0.05 compared to controls.
FAM83H-AS1 promotes tumor growth in TNBC xenograft mouse model
To investigate the function of FAM83H-AS1 in regulating TNBC progression in vivo, we used a mouse TNBC xenograft model. TNBC cells were transfected with sh-control, sh-FAM83H-AS1, LV-control, or LV-FAM83H-AS1, and subcutaneously injected into female nude mice. As shown in Figure 8A–8C, mice injected with sh-FAM83H-AS1 had reduced tumor volumes and weight compared to mice injected with sh-control (p < 0.05), suggesting that FAM83H-AS1 suppression inhibits the TNBC tumor growth in vivo. Moreover, the tumor volume and weight were higher in LV-FAM83H-AS1 transfection group than in LV-control group (Figure 8D–8F, p < 0.05), suggesting that FAM83H-AS1 overexpression promotes the TNBC tumor growth in vivo. These data indicate that FAM83H-AS1 promotes the TNBC progression in vivo.
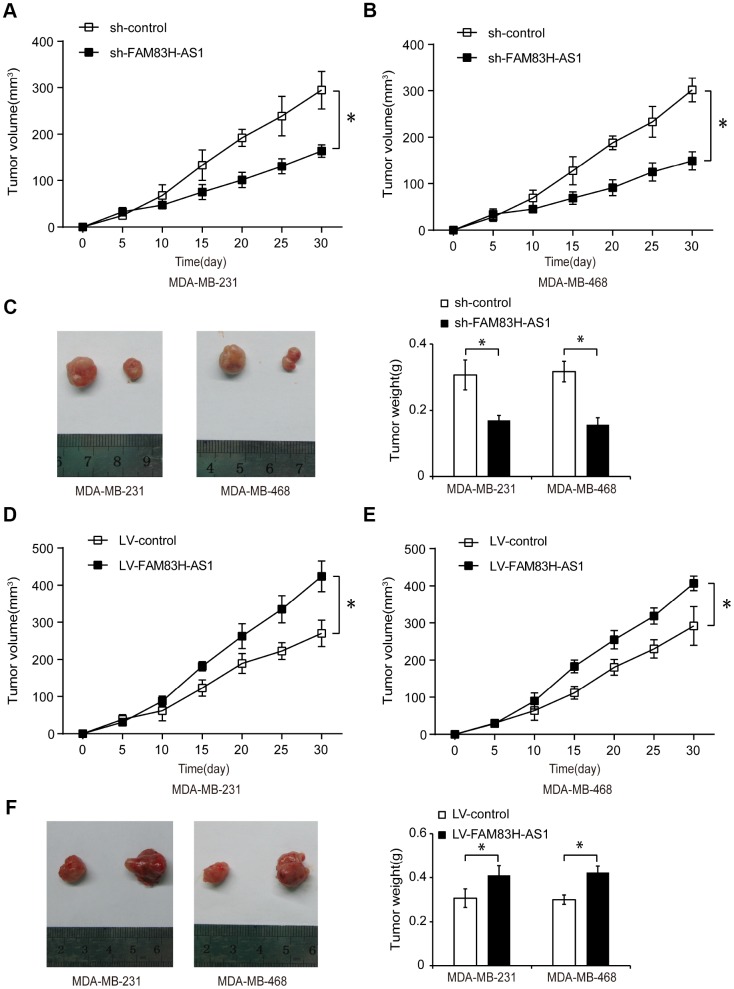
Figure 8.
FAM83H-AS1 promotes tumor growth in TNBC xenograft mouse model. (A, B) Tumor volume measured every 5 days in mice injected with TNBC cells transfected with sh-control or sh-FAM83H-AS1. (C) Tumor weight in mice injected with TNBC cells transfected with sh-control or sh-FAM83H-AS1. (D, E) Tumor size measured in mice injected with TNBC cells transfected with LV-control or LV-FAM83H-AS1. (F) Tumor weight in mice injected with TNBC cells transfected with LV-control or LV-FAM83H-AS1. * p < 0.05 compared to controls.
DISCUSSION
Dysregulation of lncRNAs plays a critical role in the development, progression, and prognosis in a variety of human cancers, including breast cancer [19, 21, 30]. However, the function and regulation of the lncRNA FAM83H-AS1 in TNBC remain largely unknown. In our present study, we have found that FAM83H-AS1 promotes TNBC cell proliferation, migration, and invasion in vitro, and induces TNBC tumor growth in vivo. MiR-136-5p is the downstream target of FAM83H-AS1. MiR-136-5p functions as a tumor suppressor in TNBC cells, and inhibits their proliferation, migration, and invasion. MiR-136-5p exerts its tumor suppressive effect by targeting the protein metadherin (MTDH). Our data show that FAM83H-AS1 promotes the TNBC progression via targeting the miR-136-5p/MTDH axis.
Previous studies have indicated that FAM83H-AS1 plays an important role in cancer progression. For instance, high expression of FAM83H-AS1 correlates with advanced tumor grade and FIGO stage, and predicts radio-resistance, metastasis risk, and poor overall survival in ovarian cancer patients [23, 31]. Upregulated FAM83H-AS1 expression is also associated with worse survival rates in human cervical cancer [22], bladder cancer [24] and lung cancer [27]. FAM83H-AS1 overexpression predicts short survival times, and promotes tumor cell proliferation via targeting the Notch signaling in colorectal cancer [26, 32]. Moreover, increased FAM83H-AS1 expression is associated with poor survival rates in patients with breast cancer [28, 29]. Our results demonstrate that FAM83H-AS1 promotes TNBC cell proliferation in vitro, and induces TNBC tumor growth in an in vivo xenograft model.
LncRNAs may exert their functions by targeting miRNAs as miRNA sponges [30, 33]. To explore the molecular mechanism by which FAM83H-AS1 promotes TNBC progression, we searched for the potential miRNA targets of FAM83H-AS1 in TNBC cells by using bioinformatics prediction. Interestingly, miR-136-5p was identified as a potential target for FAM83H-AS1; this was further confirmed by qRT-PCR, luciferase activity, and the RIP assays. MiR-136-5p was found to inhibit cell proliferation, migration and invasion in renal cell carcinoma [34], liver cancer - by regulating IRX5 [35], and in colon cancer, through targeting LRH-1/Wnt signaling [36]. MiR-136 also inhibits cell survival, proliferation, cancer stem cell spheroid formation, and tumor angiogenesis in paclitaxel-resistant ovarian cancer cells by targeting Notch3 [37]. A few studies have demonstrated that miR-136 plays an essential role in breast cancer progression. Yan et al have found that miR-136 inhibits migration and invasion of TNBC cells through targeting RASAL2 [38]. Additionally, miR-136 inhibits breast cancer progression via regulating the Wnt/β-catenin signaling [39]. Consistent with the above studies, we have found that miR-136-5p overexpression inhibits TNBC cell proliferation, migration, and invasion, while MiR-136-5p knockdown increases proliferation, migration, and invasion of TNBC cells. Our data indicate that FAM83H-AS1 promotes TNBC progression through inhibiting miR-136-5p.
In addition, our study shows that miR-136-5p exerts its suppressive function in TNBC by inhibiting the MTDH expression. MTDH, also known as AEG-1 (Astrocyte Elevated Gene 1) and Lyric, has been implicated in the development and progression of a variety of human cancers [40], including hepatic cancer [41], lung cancer [42], esophageal squamous cell carcinoma [43], glioblastoma multiforme [44], and gastric cancer [45]. Moreover, the MTDH gene is frequently amplified in breast cancer, and the increased MTDH expression is associated with increased aggressiveness [46, 47], paclitaxel resistance [48], and trastuzumab resistance [49] in breast cancer patients. Our study demonstrates that MTDH is the downstream target of miR-136-5p, and its expression is upregulated in human TNBC tissues and cells. Overexpression of MTDH is able to abrogate the inhibitory effect of miR-136-5p on TNBC cells. Thus, these data show that miR-136-5p inhibits TNBC cell proliferation, migration, and invasion through suppressing the MTDH expression.
In summary, our data demonstrate that FAM83H-AS1 functions as an oncogenic lncRNA during TNBC progression. FAM83H-AS1 promotes proliferation, migration, and invasion of TNBC cells by inhibiting the miR-136-5p levels, resulting in the increased MTDH expression. The FAM83H-AS1/miR-136-5p/MTDH axis may thus serve as a novel therapeutic target for TNBC patients.
MATERIALS AND METHODS
Patient tissues
Human primary TNBC tissues and the corresponding adjacent non-tumorous tissues were obtained from ten newly diagnosed TNBC patients (Stage I–IIA) between Jan 2018 and Jun 2018 at Tianjin Medical University Cancer Institute and Hospital (Tianjin, China). All specimens were snap-frozen and stored in liquid nitrogen for mRNA and protein extraction. None of these patients received any pre-operative chemotherapy, hormonotherapy, or radiotherapy. All patients signed the informed consent. The study protocol was approved by the Ethics Committee of Tianjin Medical University Cancer Institute and Hospital (P.R. China).
Bioinformatics analysis
The online available transcriptome microarray gene expression data (GSE76250) were downloaded from the Gene Expression Omnibus database (GEO, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE76250). The cohort (GSE76250) provided the microarray data of lncRNAs, contained 165 TNBC samples and 33 paired normal breast tissues. The microarray data were based on [HTA-2_0] Affymetrix Human Transcriptome Array 2.0. In addition, we analyzed online data using the Gene Expression Profiling Interactive Analysis 2 (GEPIA2) (http://gepia2.cancer-pku.cn/#index) [50] and cBioPortal (https://www.cbioportal.org/) databases.
Cell culture and transfection
Human TNBC cell lines (MDA-MB-231, MDA-MB-436, and MDA-MB-468) and normal breast cell line (MCF-10A) were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM; Gibco, Grand Island, NY, USA) supplemented with 10% fetal bovine serum (FBS; Gibco, Carlsbad, CA, USA), 100 U/ml penicillin and 100 U/ml streptomycin (Gibco). Cells were incubated at 37°C with 5% CO2 in a humidified atmosphere. siRNAs (si-control and si-FAM83H-AS1), miR-136-5p mimic, mimic control, miR-136-5p inhibitor and inhibitor control were obtained from Ribobio Co., Ltd. (Guangzhou, P.R. China). The pcDNA-FAM83H-AS1 and control pcDNA plasmids were constructed by GenePharma (Shanghai, China). Cell transfection was performed using Lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s instructions.
CCK8 assay
TNBC cells were incubated for 24, 48, or 72 hours at 37°C, and cell proliferation was measured by the CCK-8 assay (CCK-8; Sigma-Aldrich, St. Louis, MO, USA) according to the manufacturer’s instructions.
Transwell migration and invasion assays
Transfected TNBC cells were harvested for transwell invasion and migration assays. For migration assay, TNBC cells (1 × 105 cells) were seeded into the upper side of chambers (Corning, Corning, NY, USA). For invasion assay, TNBC cells were seeded into the upper side of chambers pre-coated with Matrigel (BD Biosciences, San Jose, CA, USA). DMEM medium (500 μl) containing 10% FBS was added into the lower chambers. After incubation for 12 hours (migration assays) or 24 hours (invasion assays), cells in the upper chamber were wiped off, and cells in the lower chamber were fixed with 4% formaldehyde, and stained with crystal violet. After washing with PBS, the number of cells that migrated or invaded were counted and the photographs were taken under a light microscope.
Wound healing assay
Wound healing assay was used to evaluate the cell migration. After transfection, cells were seeded into six well plates at 5×105 cells/well until to about 90% confluence. Linear scratches were made on the cell layer using a 200 uL pipette tip. Then, cells were maintained in serum-free media for 48 hours. The wound healing process was observed under an inverted microscope (Olympus, Japan). The photographs of wounded areas were taken at 100×magnifcation.
Quantitative RT-PCR (qRT-PCR)
Total RNA was extracted using TRIzol reagent (Invitrogen), and reverse transcribed into cDNAs by using a Reverse Transcription Kit with the M-MLV reverse transcriptase (Promega, Madison, WI, USA). Quantitative RT-PCR was utilized to determine the levels of FAM83H-AS1, miR-136-5p and MTDH with the SYBR Green detection system and the 7500 Real Time PCR System (Applied Biosystems). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) or U6 were used as internal controls for mRNA or miRNA, respectively. The relative expression was measured using 2-ΔΔCt method. All experiments were conducted in triplicates.
Luciferase reporter assay
Dual-Luciferase® Reporter Assay System (Promega, Madison, WI, USA) was used to determine the luciferase activity 48 hours after transfection according to the manufacturer’s protocol. TNBC cells were co-transfected with a wild-type or mutant FAM83H-AS1 reporter plasmid, and either a miR-136-5p mimic or a miRNA mimic control using Lipofectamine 2000 (Invitrogen, Carlsbad, CA) in accordance with the manufacturer’s protocol.
RNA immunoprecipitation (RIP)
Magna RNA immunoprecipitation (RIP) kit (Millipore, Billerica, USA) was used to conduct RIP assays in accordance with the manufacturer’s protocol. All cells were lysed using RIP lysis buffer and incubated with RIP immunoprecipitation buffer containing magnetic beads conjugated to human anti-Ago2 antibody (Abcam, Cambridge, MA) or control anti-IgG antibody. Co-precipitated RNAs were analyzed by qRT-PCR for the expression of FAM83H-AS1 and miR-136-5p.
Western blot analysis
Total protein was extracted using RIPA lysis buffer (Beyotime Institute of Biotechnology, Beijing, China), and protein concentration was measured by BCA Protein Assay Kit (Beyotime Institute of Biotechnology, Beijing, China). Equal amounts of proteins (30 μg) were separated by 10% SDS-PAGE and transferred onto PVDF membranes (Millipore, Boston, MA, USA). The membranes were blocked with 5% (w/v) nonfat milk in Tris-buffered saline containing 0.1% Tween 20 (TBST) at room temperature, and incubated with primary antibody against MTDH (Cell Signaling Technology, Danvers, MA, USA) at 4°C overnight. Following washing with TBST, the membranes were incubated with horseradish peroxidase-conjugated secondary antibody (Santa Cruz Biotechnology, Dallas, TX, USA), and visualized using Enhanced Chemiluminescence Kit (GE Healthcare, Chicago, IL, USA).
Lentivirus transduction and in vivo mouse model
The full-length sequence of FAM83H-AS1 was subcloned into pCDH-CMV-MCS-EF1-Puro lentiviral vector. The shRNA sequence specific to FAM83H-AS1 was subcloned into pLKO.1 lentiviral vector. The lentiviral vectors were co-transfected with the packaging vectors pAX8 and pCMV-VSVG into 293FT cells by Lipofectamine 2000. Two days later, viral supernatants were collected and used to infect cells. Forty-eight hours following infections, cells were selected with 1.5 μg/mL puromycin. TNBC cell lines were infected with either lentivirus-pLKO.1-sh-FAM83H-AS1 (sh-FAM83H-AS1), control lentivirus-shRNA (sh-control), lentivirus-FAM83H-AS1 (LV-FAM83H-AS1) or lentivirus-control virus (LV-control).
Nude mice (4~6 weeks old, female) were maintained under pathogen free conditions. All animal procedures were approved by the Animal Care Committee of Tianjin Medical University. For the tumor xenograft experiments, stable TNBC cells (2×106) transfected with either sh-control, sh-FAM83H-AS1, LV-control or LV-FAM83H-AS1 were subcutaneously injected into mice (n = 3 per group). The tumor volume was measured every 5 days, and calculated by the formula: length × width2 × 0.5. At 30 days post-injection, the mice were euthanized and tumors were surgically isolated and photographed.
Statistical analysis
Student’s unpaired t-test was used for comparisons between groups. All data are presented as mean ± standard deviation (SD). A value of p < 0.05 was regarded statistically significant. Statistical analysis was performed using the IBM SPSS 22.0 (IBM Corp., Armonk, NY, USA) and GraphPad Prism 8.0 (San Diego, CA, USA).
Supplementary Material
ACKNOWLEDGMENTS
We thank the National Natural Science Foundation of China and Tianjin Medical University Cancer Institute and Hospital for supporting this study.
Abbreviations
- TNBC
triple-negative breast cancer
- lncRNAs
long noncoding RNAs
- miR
microRNA
- FAM83H-AS1
FAM83H antisense RNA 1
- MTDH
Metadherin
- qRT-PCR
quantitative real-time PCR
- Wt
wild-type
- Mut
mutant binding sites
- CCK8
Cell Counting Kit-8
- GEPIA2
Gene Expression Profiling Interactive Analysis 2
- siRNA
small interfering RNA
Footnotes
AUTHOR CONTRIBUTIONS: C. Han, J. Yin and Q. Li designed the experiments; C. Han, N. Zeng and Y, Fu performed the experiments; C. Han, J. Yin and Q. Li analyzed data; and C. Han, J. Yin and Q. Li wrote the paper.
CONFLICTS OF INTEREST: The authors have declared that no conflicts of interest exists.
FUNDING: The work was supported by the National Natural Science Foundation of China [No. 81602341 and 81702637]; Tianjin Medical University Cancer Institute and Hospital Innovative and Excellent Young Talents Program (2018-1-40); Tianjin "The Belt and Road" Technological Innovation and Cooperation Grant (No. 18PTZWHZ00050); Tianjin Health and Care Committee Science Fund (2014KZ019).
REFERENCES
- 1.Waks AG, Winer EP. Breast Cancer Treatment: A Review. JAMA. 2019; 321:288–300. 10.1001/jama.2018.19323 [DOI] [PubMed] [Google Scholar]
- 2.Denkert C, Liedtke C, Tutt A, von Minckwitz G. Molecular alterations in triple-negative breast cancer-the road to new treatment strategies. Lancet. 2017; 389:2430–42. 10.1016/S0140-6736(16)32454-0 [DOI] [PubMed] [Google Scholar]
- 3.Kopp F, Mendell JT. Functional Classification and Experimental Dissection of Long Noncoding RNAs. Cell. 2018; 172:393–407. 10.1016/j.cell.2018.01.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Batista PJ, Chang HY. Long noncoding RNAs: cellular address codes in development and disease. Cell. 2013; 152:1298–307. 10.1016/j.cell.2013.02.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Fatica A, Bozzoni I. Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet. 2014; 15:7–21. 10.1038/nrg3606 [DOI] [PubMed] [Google Scholar]
- 6.Häfner SJ, Talvard TG, Lund AH. Long noncoding RNAs in normal and pathological pluripotency. Semin Cell Dev Biol. 2017; 65:1–10. 10.1016/j.semcdb.2016.07.011 [DOI] [PubMed] [Google Scholar]
- 7.Zhu P, Wu J, Wang Y, Zhu X, Lu T, Liu B, He L, Ye B, Wang S, Meng S, Fan D, Wang J, Yang L, et al. LncGata6 maintains stemness of intestinal stem cells and promotes intestinal tumorigenesis. Nat Cell Biol. 2018; 20:1134–44. 10.1038/s41556-018-0194-0 [DOI] [PubMed] [Google Scholar]
- 8.Mendell JT. Targeting a Long Noncoding RNA in Breast Cancer. N Engl J Med. 2016; 374:2287–89. 10.1056/NEJMcibr1603785 [DOI] [PubMed] [Google Scholar]
- 9.Wang Z, Yang B, Zhang M, Guo W, Wu Z, Wang Y, Jia L, Li S; Cancer Genome Atlas Research Network, Xie W, Yang D. lncRNA Epigenetic Landscape Analysis Identifies EPIC1 as an Oncogenic lncRNA that Interacts with MYC and Promotes Cell-Cycle Progression in Cancer. Cancer Cell. 2018; 33:706–720.e9. 10.1016/j.ccell.2018.03.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Sun TT, He J, Liang Q, Ren LL, Yan TT, Yu TC, Tang JY, Bao YJ, Hu Y, Lin Y, Sun D, Chen YX, Hong J, et al. LncRNA GClnc1 Promotes Gastric Carcinogenesis and May Act as a Modular Scaffold of WDR5 and KAT2A Complexes to Specify the Histone Modification Pattern. Cancer Discov. 2016; 6:784–801. 10.1158/2159-8290.CD-15-0921 [DOI] [PubMed] [Google Scholar]
- 11.Wang YL, Liu JY, Yang JE, Yu XM, Chen ZL, Chen YJ, Kuang M, Zhu Y, Zhuang SM. Lnc-UCID Promotes G1/S Transition and Hepatoma Growth by Preventing DHX9-Mediated CDK6 Down-regulation. Hepatology. 2019; 70:259–75. 10.1002/hep.30613 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Xiao G, Yao J, Kong D, Ye C, Chen R, Li L, Zeng T, Wang L, Zhang W, Shi X, Zhou T, Li J, Wang Y, et al. The Long Noncoding RNA TTTY15, Which Is Located on the Y Chromosome, Promotes Prostate Cancer Progression by Sponging let-7. Eur Urol. 2019; 76:315–26. 10.1016/j.eururo.2018.11.012 [DOI] [PubMed] [Google Scholar]
- 13.Feng J, Zhou Q, Yi H, Ma S, Li D, Xu Y, Wang J, Yin S. A novel lncRNA n384546 promotes thyroid papillary cancer progression and metastasis by acting as a competing endogenous RNA of miR-145-5p to regulate AKT3. Cell Death Dis. 2019; 10:433. 10.1038/s41419-019-1637-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lu G, Li Y, Ma Y, Lu J, Chen Y, Jiang Q, Qin Q, Zhao L, Huang Q, Luo Z, Huang S, Wei Z. Long noncoding RNA LINC00511 contributes to breast cancer tumourigenesis and stemness by inducing the miR-185-3p/E2F1/Nanog axis. J Exp Clin Cancer Res. 2018; 37:289. 10.1186/s13046-018-0945-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Tao W, Wang C, Zhu B, Zhang G, Pang D. LncRNA DANCR contributes to tumor progression via targetting miR-216a-5p in breast cancer: lncRNA DANCR contributes to tumor progression. Biosci Rep. 2019; 39:BSR20181618. 10.1042/BSR20181618 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Jiang M, Qiu N, Xia H, Liang H, Li H, Ao X. Long non-coding RNA FOXD2-AS1/miR-150-5p/PFN2 axis regulates breast cancer malignancy and tumorigenesis. Int J Oncol. 2019; 54:1043–52. 10.3892/ijo.2019.4671 [DOI] [PubMed] [Google Scholar]
- 17.Wang L, Luan T, Zhou S, Lin J, Yang Y, Liu W, Tong X, Jiang W. LncRNA HCP5 promotes triple negative breast cancer progression as a ceRNA to regulate BIRC3 by sponging miR-219a-5p. Cancer Med. 2019; 8:4389–403. 10.1002/cam4.2335 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Collina F, Aquino G, Brogna M, Cipolletta S, Buonfanti G, De Laurentiis M, Di Bonito M, Cantile M, Botti G. LncRNA HOTAIR up-regulation is strongly related with lymph nodes metastasis and LAR subtype of Triple Negative Breast Cancer. J Cancer. 2019; 10:2018–24. 10.7150/jca.29670 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Vidovic D, Huynh TT, Konda P, Dean C, Cruickshank BM, Sultan M, Coyle KM, Gujar S, Marcato P. ALDH1A3-regulated long non-coding RNA NRAD1 is a potential novel target for triple-negative breast tumors and cancer stem cells. Cell Death Differ. 2020; 27:363–78. 10.1038/s41418-019-0362-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Fu J, Dong G, Shi H, Zhang J, Ning Z, Bao X, Liu C, Hu J, Liu M, Xiong B. LncRNA MIR503HG inhibits cell migration and invasion via miR-103/OLFM4 axis in triple negative breast cancer. J Cell Mol Med. 2019; 23:4738–45. 10.1111/jcmm.14344 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zhang H, Zhang N, Liu Y, Su P, Liang Y, Li Y, Wang X, Chen T, Song X, Sang Y, Duan Y, Zhang J, Wang L, et al. Epigenetic Regulation of NAMPT by NAMPT-AS Drives Metastatic Progression in Triple-Negative Breast Cancer. Cancer Res. 2019; 79:3347–59. 10.1158/0008-5472.CAN-18-3418 [DOI] [PubMed] [Google Scholar]
- 22.Barr JA, Hayes KE, Brownmiller T, Harold AD, Jagannathan R, Lockman PR, Khan S, Martinez I. Long non-coding RNA FAM83H-AS1 is regulated by human papillomavirus 16 E6 independently of p53 in cervical cancer cells. Sci Rep. 2019; 9:3662. 10.1038/s41598-019-40094-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Dou Q, Xu Y, Zhu Y, Hu Y, Yan Y, Yan H. LncRNA FAM83H-AS1 contributes to the radioresistance, proliferation, and metastasis in ovarian cancer through stabilizing HuR protein. Eur J Pharmacol. 2019; 852:134–41. 10.1016/j.ejphar.2019.03.002 [DOI] [PubMed] [Google Scholar]
- 24.Shan H, Yang Y, Zhu X, Han X, Zhang P, Zhang X. FAM83H-AS1 is associated with clinical progression and modulates cell proliferation, migration, and invasion in bladder cancer. J Cell Biochem. 2019; 120:4687–93. 10.1002/jcb.27758 [DOI] [PubMed] [Google Scholar]
- 25.Bi YY, Shen G, Quan Y, Jiang W, Xu F. Long noncoding RNA FAM83H-AS1 exerts an oncogenic role in glioma through epigenetically silencing CDKN1A (p21). J Cell Physiol. 2018; 233:8896–907. 10.1002/jcp.26813 [DOI] [PubMed] [Google Scholar]
- 26.Lu S, Dong W, Zhao P, Liu Z. lncRNA FAM83H-AS1 is associated with the prognosis of colorectal carcinoma and promotes cell proliferation by targeting the Notch signaling pathway. Oncol Lett. 2018; 15:1861–68. 10.3892/ol.2017.7520 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhang J, Feng S, Su W, Bai S, Xiao L, Wang L, Thomas DG, Lin J, Reddy RM, Carrott PW, Lynch WR, Chang AC, Beer DG, et al. Overexpression of FAM83H-AS1 indicates poor patient survival and knockdown impairs cell proliferation and invasion via MET/EGFR signaling in lung cancer. Sci Rep. 2017; 7:42819. 10.1038/srep42819 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yang F, Lv SX, Lv L, Liu YH, Dong SY, Yao ZH, Dai XX, Zhang XH, Wang OC. Identification of lncRNA FAM83H-AS1 as a novel prognostic marker in luminal subtype breast cancer. OncoTargets Ther. 2016; 9:7039–45. 10.2147/OTT.S110055 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Deva Magendhra Rao AK, Patel K, Korivi Jyothiraj S, Meenakumari B, Sundersingh S, Sridevi V, Rajkumar T, Pandey A, Chatterjee A, Gowda H, Mani S. Identification of lncRNAs associated with early-stage breast cancer and their prognostic implications. Mol Oncol. 2019; 13:1342–55. 10.1002/1878-0261.12489 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wang M, Mao C, Ouyang L, Liu Y, Lai W, Liu N, Shi Y, Chen L, Xiao D, Yu F, Wang X, Zhou H, Cao Y, Liu S, Yan Q, Tao Y, Zhang B. Long noncoding RNA LINC00336 inhibits ferroptosis in lung cancer by functioning as a competing endogenous RNA. Cell Death Differ. 2019; 26:2329–43. 10.1038/s41418-019-0304-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Gong YB, Zou YF. Clinical significance of lncRNA FAM83H-AS1 in ovarian cancer. Eur Rev Med Pharmacol Sci. 2019; 23:4656–62. 10.26355/eurrev_201906_18045 [DOI] [PubMed] [Google Scholar]
- 32.Yang L, Xu L, Wang Q, Wang M, An G. Dysregulation of long non-coding RNA profiles in human colorectal cancer and its association with overall survival. Oncol Lett. 2016; 12:4068–74. 10.3892/ol.2016.5138 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Furió-Tarí P, Tarazona S, Gabaldón T, Enright AJ, Conesa A. spongeScan: A web for detecting microRNA binding elements in lncRNA sequences. Nucleic Acids Res. 2016; 44:W176–80. 10.1093/nar/gkw443 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Chen P, Zhao L, Pan X, Jin L, Lin C, Xu W, Xu J, Guan X, Wu X, Wang Y, Yang S, Wang T, Lai Y. Tumor suppressor microRNA-136-5p regulates the cellular function of renal cell carcinoma. Oncol Lett. 2018; 15:5995–6002. 10.3892/ol.2018.8081 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Zhu L, Liu Y, Chen Q, Yu G, Chen J, Chen K, Yang N, Zeng T, Yan S, Huang A, Tang H. Long-Noncoding RNA Colorectal Neoplasia Differentially Expressed Gene as a Potential Target to Upregulate the Expression of IRX5 by miR-136-5P to Promote Oncogenic Properties in Hepatocellular Carcinoma. Cell Physiol Biochem. 2018; 50:2229–48. 10.1159/000495084 [DOI] [PubMed] [Google Scholar]
- 36.Yuan Q, Cao G, Li J, Zhang Y, Yang W. MicroRNA-136 inhibits colon cancer cell proliferation and invasion through targeting liver receptor homolog-1/Wnt signaling. Gene. 2017; 628:48–55. 10.1016/j.gene.2017.07.031 [DOI] [PubMed] [Google Scholar]
- 37.Jeong JY, Kang H, Kim TH, Kim G, Heo JH, Kwon AY, Kim S, Jung SG, An HJ. MicroRNA-136 inhibits cancer stem cell activity and enhances the anti-tumor effect of paclitaxel against chemoresistant ovarian cancer cells by targeting Notch3. Cancer Lett. 2017; 386:168–78. 10.1016/j.canlet.2016.11.017 [DOI] [PubMed] [Google Scholar]
- 38.Yan M, Li X, Tong D, Han C, Zhao R, He Y, Jin X. miR-136 suppresses tumor invasion and metastasis by targeting RASAL2 in triple-negative breast cancer. Oncol Rep. 2016; 36:65–71. 10.3892/or.2016.4767 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Huan J, Xing L, Lin Q, Xui H, Qin X. Long noncoding RNA CRNDE activates Wnt/β-catenin signaling pathway through acting as a molecular sponge of microRNA-136 in human breast cancer. Am J Transl Res. 2017; 9:1977–89. [PMC free article] [PubMed] [Google Scholar]
- 40.Dhiman G, Srivastava N, Goyal M, Rakha E, Lothion-Roy J, Mongan NP, Miftakhova RR, Khaiboullina SF, Rizvanov AA, Baranwal M. Metadherin: A Therapeutic Target in Multiple Cancers. Front Oncol. 2019; 9:349. 10.3389/fonc.2019.00349 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Srivastava J, Siddiq A, Gredler R, Shen XN, Rajasekaran D, Robertson CL, Subler MA, Windle JJ, Dumur CI, Mukhopadhyay ND, Garcia D, Lai Z, Chen Y, et al. Astrocyte elevated gene-1 and c-Myc cooperate to promote hepatocarcinogenesis in mice. Hepatology. 2015; 61:915–29. 10.1002/hep.27339 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Lu Q, Shan S, Li Y, Zhu D, Jin W, Ren T. Long noncoding RNA SNHG1 promotes non-small cell lung cancer progression by up-regulating MTDH via sponging miR-145-5p. FASEB J. 2018; 32:3957–67. 10.1096/fj.201701237RR [DOI] [PubMed] [Google Scholar]
- 43.Yang C, Zheng S, Liu T, Liu Q, Dai F, Zhou J, Chen Y, Sheyhidin I, Lu X. Down-regulated miR-26a promotes proliferation, migration, and invasion via negative regulation of MTDH in esophageal squamous cell carcinoma. FASEB J. 2017; 31:2114–22. 10.1096/fj.201601237 [DOI] [PubMed] [Google Scholar]
- 44.Hu B, Emdad L, Bacolod MD, Kegelman TP, Shen XN, Alzubi MA, Das SK, Sarkar D, Fisher PB. Astrocyte elevated gene-1 interacts with Akt isoform 2 to control glioma growth, survival, and pathogenesis. Cancer Res. 2014; 74:7321–32. 10.1158/0008-5472.CAN-13-2978 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Li G, Wang Z, Ye J, Zhang X, Wu H, Peng J, Song W, Chen C, Cai S, He Y, Xu J. Uncontrolled inflammation induced by AEG-1 promotes gastric cancer and poor prognosis. Cancer Res. 2014; 74:5541–52. 10.1158/0008-5472.CAN-14-0968 [DOI] [PubMed] [Google Scholar]
- 46.Tokunaga E, Nakashima Y, Yamashita N, Hisamatsu Y, Okada S, Akiyoshi S, Aishima S, Kitao H, Morita M, Maehara Y. Overexpression of metadherin/MTDH is associated with an aggressive phenotype and a poor prognosis in invasive breast cancer. Breast Cancer. 2014; 21:341–49. 10.1007/s12282-012-0398-2 [DOI] [PubMed] [Google Scholar]
- 47.Liu P, Tang H, Chen B, He Z, Deng M, Wu M, Liu X, Yang L, Ye F, Xie X. miR-26a suppresses tumour proliferation and metastasis by targeting metadherin in triple negative breast cancer. Cancer Lett. 2015; 357:384–92. 10.1016/j.canlet.2014.11.050 [DOI] [PubMed] [Google Scholar]
- 48.Yang L, Tian Y, Leong WS, Song H, Yang W, Wang M, Wang X, Kong J, Shan B, Song Z. Efficient and tumor-specific knockdown of MTDH gene attenuates paclitaxel resistance of breast cancer cells both in vivo and in vitro. Breast Cancer Res. 2018; 20:113. 10.1186/s13058-018-1042-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Du C, Yi X, Liu W, Han T, Liu Z, Ding Z, Zheng Z, Piao Y, Yuan J, Han Y, Xie M, Xie X. MTDH mediates trastuzumab resistance in HER2 positive breast cancer by decreasing PTEN expression through an NFκB-dependent pathway. BMC Cancer. 2014; 14:869. 10.1186/1471-2407-14-869 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Tang Z, Kang B, Li C, Chen T, Zhang Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019; 47:W556–60. 10.1093/nar/gkz430 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.