Abstract
Background
Iron (Fe) deficiency is a common problem in citrus production. As the second largest superfamily of transcription factors (TFs), the basic/helix-loop-helix (bHLH) proteins have been shown to participate in the regulation of Fe homeostasis and a series of other biological and developmental processes in plants. However, this family of members in citrus and their functions in citrus Fe deficiency are still largely unknown.
Results
In this study, we identified a total of 128 CgbHLHs from pummelo (Citrus grandis) genome that were classified into 18 subfamilies by phylogenetic comparison with Arabidopsis thaliana bHLH proteins. All of these CgbHLHs were randomly distributed on nine known (125 genes) and one unknown (3 genes) chromosomes, and 12 and 47 of them were identified to be tandem and segmental duplicated genes, respectively. Sequence analysis showed detailed characteristics of their intron-exon structures, bHLH domain and conserved motifs. Gene ontology (GO) analysis suggested that most of CgbHLHs were annotated to the nucleus, DNA-binding transcription factor activity, response to abiotic stimulus, reproduction, post-embryonic development, flower development and photosynthesis. In addition, 27 CgbHLH proteins were predicted to have direct or indirect protein-protein interactions. Based on GO annotation, RNA sequencing data in public database and qRT-PCR results, several of CgbHLHs were identified as the key candidates that respond to iron deficiency.
Conclusions
In total, 128 CgbHLH proteins were identified from pummelo, and their detailed sequence and structure characteristics and putative functions were analyzed. This study provides comprehensive information for further functional elucidation of CgbHLH genes in citrus.
Keywords: Citrus, bHLH, Stress, Motif, Gene expression
Background
Citrus is the largest fruit crop in the world, which provides not only necessary nutrition for human but also substantial economic income for farmers. In 2015, the global citrus area was 13.5 million hectares and the yield reached 178.2 million tons (FAO statistics, http://faostat.fao.org/default.aspx). However, citrus production is also continuously influenced by many environmental factors, such as diseases, cold, drought, heat, and nutrient disorders, among which iron (Fe) deficiency is a common problem that can cause severe chlorosis of leaves, impaired tree vigor, and reduction of fruit yield and quality in citrus production [1]. In particular, in calcareous soils, citrus plants are highly sensitive to low Fe availability because bioavailable forms of ferrous Fe (II) are oxidized into insoluble ferric Fe (III) in a high-pH and oxygen-rich environment [2, 3]. To cope with this issue, plants have developed two major adaptive mechanisms for efficient Fe uptake from soils. All dicot and non-graminaceous monocot plants, such as Arabidopsis thaliana (A. thaliana), use a reduction-based strategy (strategy I), while graminaceous plants such as rice employ a chelation-based strategy (strategy II) [3–5]. In both strategy I and strategy II, one class of transcription factors (TFs), the basic/helix-loop-helix genes (bHLHs), is found to play the core regulatory role [3, 4, 6–9].
The first discovery of bHLH was during the study of murine muscle development [10]. Thereafter, the proteins containing this conserved domain have been widely identified in all three eukaryotic kingdoms and constitute one of the largest families of TFs [11–15]. In plants, the bHLH proteins are recognized as the second largest superfamily of TFs [12]. The bHLH domain consists of approximately 60 amino acids with two functionally distinct regions, the basic region and the HLH region [12, 14]. The basic region, containing 13 to 17 amino acids, is located at the N-terminus of the bHLH domain and functions in specific recognition and binding to the DNA motif of the target gene promoter [16]. In contrast, the HLH region is located at the C-terminal end and consists of approximately 40–50 amino acids with two amphipathic α helices that are linked by a loop of variable length. This region often functions in domain dimerization and allows the formation of homodimers or heterodimers to promote protein-protein interactions [12, 16, 17]. Outside of the conserved bHLH domain, the sequence of bHLH proteins is considerably divergent [18].
As an essential superfamily of TFs, bHLH proteins participate in regulating of a series of biological and developmental processes, such as flowering [19], root development [20], seed germination [21], anthocyanin or flavonoid metabolism [22–24], hormonal signaling regulation [25, 26], as well as biotic and abiotic stresses responses [14, 27–29]. Under Fe deficiency stress, bHLHs have been shown to play a predominant regulatory role. The FER-like iron-deficiency-induced transcription factor (FIT), encoding a bHLH29 orthologous to the tomato FER protein, is the first identified TF that regulates Fe homeostasis in A. thaliana [30–32]. Subsequently, bHLH38, bHLH39, bHLH100, and bHLH101, belonging to the Ib subgroup of the bHLH, are found to form heterodimers with FIT and positively regulate the expression of IRT1 (iron-regulated transporter 1) and FRO2 (ferric reduction oxidase 2) under Fe deficiency [9, 33]. POPEYE (PYE), another bHLH protein (bHLH47), acts as a negative regulator to participate in Fe homeostasis [34]. Recently, bHLH34, bHLH104, bHLH105, and bHLH115 have been shown to play essential roles in Fe homeostasis by positively regulating bHLH38, bHLH39, bHLH100, bHLH101 and PYE [8, 35, 36]. Moreover, four IVa subgroups of bHLH members, bHLH18, bHLH19, bHLH20, and bHLH25, were identified as novel interactors of FIT and mediate jasmonic acid-induced FIT protein degradation under Fe deficiency [6].
Although many bHLH genes in response to Fe deficiency have been well documented in model plants, it is still poorly understood in citrus. Pummelo (Citrus grandis) is a major cultivated species of citrus, which also belongs to the progenitor species that contributes to the generation of hybridized mandarins and sweet orange [37]. Both directly cultivated pummelos and the varieties grafted on pummelo rootstocks readily show Fe deficiency. Recently, high quality genome information of pummelo [38] was published, which promoted us to consider that we can use pummelo as material to systematically analyze the bHLH family genes and identify the essential members involved in the response to Fe deficiency in citrus. Therefore, in the present study, we identified 128 bHLH family members from pummelo at the whole genome level. We also carried out a detailed analysis and prediction of their sequence characteristics, phylogeny, gene duplication, chromosomal distribution, gene structure, protein motif, and protein-protein interaction. The bHLH members in response to Fe deficiency were identified by performing gene ontology (GO) annotation and time-course expression analysis under Fe deficiency. Our results provide valuable clues for functional elucidation of the bHLH genes in citrus, especially for revealing the regulatory mechanisms of bHLHs in citrus under Fe deficiency in the future.
Results
Identification and classification of CgbHLH members in pummelo
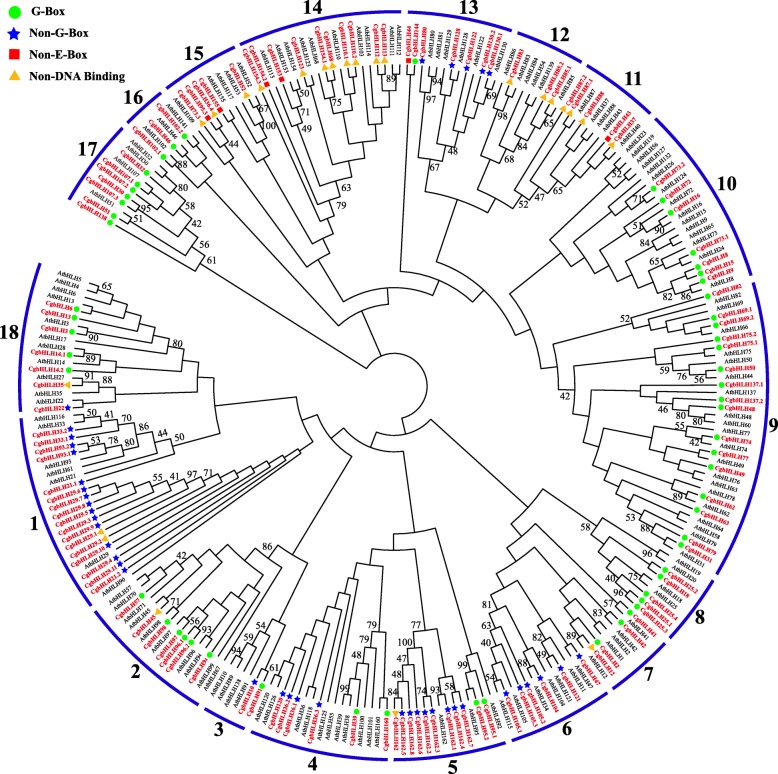
Based on the methods in “Materials and methods,” we finally identified 128 bHLH proteins from pummelo (Table 1 and Additional file 2: Table S1). These bHLHs showed 21.48% to 73.16% sequence identity with bHLHs of A. thaliana (AtbHLHs), and they were named CgbHLHs according to the highest sequence identity of AtbHLH (Table 1). If more than one CgbHLHs corresponded to the same AtbHLH, they were presented with an extra decimal point (e.g. CgbHLH29.1–CgbHLH29.11). The ORF length of the CgbHLHs ranged from 210 bp (CgbHLH29.1) to 4401 bp (CgbHLH73.2), encoding 69–1466 amino acids (Table 1 and Additional file 2: Table S1). Although these CgbHLHs showed large differences in length, 74.2% of them (95/128) were in the range of 700–2000 bp. The predicted MW and pI of CgbHLHs ranged from 7.56 to 163.79 kDa and 4.55 to 10.41, respectively (Additional file 2: Table S1). To explore the evolutionary relationship and the classification of CgbHLHs, a neighbor-joining phylogenetic tree was constructed with conserved sequences of 128 CgbHLHs and 136 AtbHLHs. Based on this phylogenetic analysis and the previous reported classification of the AtbHLHs, we classified 128 CgbHLHs into 18 subfamilies using Arabic numerals 1–18 (Fig. 1). Group 1 was the largest subfamily with 17 CgbHLH members, while the smallest, group 3, contained only 2 members.
Table 1.
Summary of 128 identified CgbHLH members
| No. | Gene Name | ORF Length | Gene ID | A. thaliana Homologs | Abbr. Name | Identity% | No. | Gene Name | ORF Length | Gene ID | A. thaliana Homologs | Abbr. Name | Identity% |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | CgbHLH2 | 1869 | Cg5g002660 | AT1G63650 | EGL3 | 52.09 | 65 | CgbHLH73.3 | 915 | Cg1g019630 | AT5G67110 | ALC | 26.42 |
| 2 | CgbHLH3 | 1512 | Cg5g034370 | AT4G16430 | JAM3 | 58.3 | 66 | CgbHLH74 | 1284 | Cg3g013130 | AT1G10120 | CIB4 | 49.3 |
| 3 | CgbHLH6 | 2064 | CgUng000770 | AT1G32640 | JAI1 | 61.74 | 67 | CgbHLH75.1 | 717 | Cg2g036150 | AT1G25330 | CES | 52.2 |
| 4 | CgbHLH8 | 2229 | Cg5g013200 | AT1G09530 | PIF3 | 44.38 | 68 | CgbHLH75.2 | 1095 | Cg6g019500 | AT1G25330 | CES | 39.9 |
| 5 | CgbHLH9 | 1632 | Cg7g012190 | AT2G43010 | PIF4 | 44.07 | 69 | CgbHLH77 | 1455 | Cg9g014180 | AT3G23690 | CIL2 | 47.21 |
| 6 | CgbHLH10 | 1071 | Cg9g023740 | AT2G31220 | – | 46.5 | 70 | CgbHLH79 | 840 | Cg5g004140 | AT5G62610 | – | 59.71 |
| 7 | CgbHLH12 | 1974 | Cg5g042050 | AT4G00480 | MYC1 | 49.49 | 71 | CgbHLH80 | 765 | Cg3g020720 | AT1G35460 | FBH1 | 63.68 |
| 8 | CgbHLH13 | 1482 | Cg5g040610 | AT1G01260 | JAM2 | 57.11 | 72 | CgbHLH82 | 1563 | Cg4g018170 | AT5G58010 | LRL3 | 43.38 |
| 9 | CgbHLH14.1 | 1551 | Cg5g040200 | AT4G00870 | – | 47.04 | 73 | CgbHLH83 | 888 | Cg6g018550 | AT1G66470 | RHD6 | 48.46 |
| 10 | CgbHLH14.2 | 1563 | Cg5g000450 | AT4G00870 | – | 37.07 | 74 | CgbHLH85.1 | 1089 | Cg8g003130 | AT4G33880 | RSL2 | 51.39 |
| 11 | CgbHLH15 | 1764 | Cg5g032930 | AT2G20180 | PIL5 | 46.61 | 75 | CgbHLH85.2 | 1104 | Cg3g012730 | AT4G33880 | RSL2 | 45.74 |
| 12 | CgbHLH16 | 1386 | Cg5g043840 | AT4G00050 | UNE10 | 59.43 | 76 | CgbHLH87.1 | 1296 | Cg1g009170 | AT3G21330 | – | 52.65 |
| 13 | CgbHLH18 | 783 | Cg5g017710 | AT2G22750 | – | 41.44 | 77 | CgbHLH87.2 | 513 | Cg6g009450 | AT3G21330 | – | 47.87 |
| 14 | CgbHLH21.1 | 1785 | Cg8g001500 | AT2G16910 | AMS | 48.72 | 78 | CgbHLH88 | 822 | Cg3g021250 | AT5G67060 | HEC1 | 50.93 |
| 15 | CgbHLH21.2 | 1419 | Cg7g019250 | AT2G16910 | AMS | 36.78 | 79 | CgbHLH91 | 1428 | Cg5g045050 | AT2G31210 | – | 45.32 |
| 16 | CgbHLH22 | 585 | Cg2g040910 | AT4G21330 | DYT1 | 43.92 | 80 | CgbHLH93.1 | 1065 | Cg1g007400 | AT5G65640 | NFL | 52.13 |
| 17 | CgbHLH25.1 | 570 | Cg8g002160 | AT4G37850 | – | 49.25 | 81 | CgbHLH93.2 | 1155 | Cg3g013530 | AT5G65640 | NFL | 46.11 |
| 18 | CgbHLH25.2 | 1017 | Cg1g007240 | AT4G37850 | – | 44.83 | 82 | CgbHLH94 | 957 | Cg1g008260 | AT1G22490 | – | 46.89 |
| 19 | CgbHLH25.3 | 426 | Cg8g002170 | AT4G37850 | – | 44.56 | 83 | CgbHLH95.1 | 939 | Cg8g004350 | AT1G49770 | RGE1 | 42.18 |
| 20 | CgbHLH25.4 | 1107 | Cg1g007260 | AT4G37850 | – | 44.04 | 84 | CgbHLH95.2 | 804 | Cg3g021160 | AT1G49770 | RGE1 | 41.02 |
| 21 | CgbHLH29.1 | 210 | Cg8g019060 | AT2G28160 | FIT | 64.41 | 85 | CgbHLH95.3 | 729 | Cg2g044520 | AT1G49770 | RGE1 | 24.66 |
| 22 | CgbHLH29.2 | 222 | Cg8g019160 | AT2G28160 | FIT | 61.9 | 86 | CgbHLH96.1 | 993 | Cg3g014510 | AT1G72210 | – | 56.62 |
| 23 | CgbHLH29.3 | 489 | Cg8g018990 | AT2G28160 | FIT | 52 | 87 | CgbHLH96.2 | 906 | Cg2g030740 | AT1G72210 | – | 51.72 |
| 24 | CgbHLH29.4 | 969 | Cg8g019240 | AT2G28160 | FIT | 51.16 | 88 | CgbHLH97 | 1287 | Cg9g012540 | AT3G24140 | FMA | 60.67 |
| 25 | CgbHLH29.5 | 501 | Cg8g019040 | AT2G28160 | FIT | 50.49 | 89 | CgbHLH98 | 966 | Cg9g028240 | AT5G53210 | SPCH | 61.29 |
| 26 | CgbHLH29.6 | 633 | Cg8g019200 | AT2G28160 | FIT | 49.54 | 90 | CgbHLH102.1 | 1029 | Cg7g015780 | AT1G69010 | BIM2 | 60.19 |
| 27 | CgbHLH29.7 | 237 | Cg8g019020 | AT2G28160 | FIT | 49.25 | 91 | CgbHLH102.2 | 1026 | Cg6g020770 | AT1G69010 | BIM2 | 44.37 |
| 28 | CgbHLH29.8 | 660 | Cg8g019140 | AT2G28160 | FIT | 48.15 | 92 | CgbHLH104 | 648 | Cg9g007860 | AT4G14410 | – | 64.62 |
| 29 | CgbHLH29.9 | 651 | Cg8g018940 | AT2G28160 | FIT | 47.27 | 93 | CgbHLH105.1 | 714 | Cg2g023210 | AT5G54680 | ILR3 | 73.16 |
| 30 | CgbHLH29.10 | 1416 | Cg8g019230 | AT2G28160 | FIT | 37.17 | 94 | CgbHLH105.2 | 711 | Cg2g017780 | AT5G54680 | ILR3 | 65.18 |
| 31 | CgbHLH29.11 | 903 | Cg8g018890 | AT2G28160 | FIT | 33.15 | 95 | CgbHLH105.3 | 711 | Cg2g040740 | AT5G54680 | ILR3 | 62.11 |
| 32 | CgbHLH30 | 798 | Cg2g043260 | AT1G68810 | ABS5 | 41.67 | 96 | CgbHLH107.1 | 717 | Cg6g017840 | AT3G56770 | – | 59.26 |
| 33 | CgbHLH31 | 753 | Cg7g020320 | AT1G59640 | ZCW32 | 51.95 | 97 | CgbHLH107.2 | 786 | Cg3g014080 | AT3G56770 | – | 37.04 |
| 34 | CgbHLH32 | 789 | Cg7g014890 | AT3G25710 | TMO5 | 58.65 | 98 | CgbHLH107.3 | 846 | Cg2g008600 | AT3G56770 | – | 35.87 |
| 35 | CgbHLH33.1 | 1659 | Cg7g011080 | AT1G12860 | ICE2 | 57.28 | 99 | CgbHLH110.1 | 1299 | Cg3g023160 | AT1G27660 | – | 51.35 |
| 36 | CgbHLH33.2 | 1443 | Cg2g009820 | AT1G12860 | ICE2 | 55.8 | 100 | CgbHLH110.2 | 1104 | CgUng019530 | AT1G27660 | – | 48.1 |
| 37 | CgbHLH35 | 747 | Cg4g021000 | AT5G57150 | – | 57.2 | 101 | CgbHLH112 | 1479 | Cg2g040950 | AT1G61660 | – | 44.04 |
| 38 | CgbHLH36.1 | 741 | Cg9g025850 | AT5G51780 | – | 53.85 | 102 | CgbHLH113 | 819 | Cg8g003580 | AT3G19500 | – | 45.71 |
| 39 | CgbHLH36.2 | 744 | Cg5g009400 | AT5G51780 | – | 42.77 | 103 | CgbHLH120 | 537 | Cg9g025860 | AT5G51790 | – | 47.46 |
| 40 | CgbHLH36.3 | 726 | Cg9g025870 | AT5G51780 | – | 38.69 | 104 | CgbHLH121 | 1029 | CgUng005820 | AT3G19860 | – | 61.39 |
| 41 | CgbHLH36.4 | 1122 | Cg3g014280 | AT5G51780 | – | 25.81 | 105 | CgbHLH122 | 1296 | Cg2g002540 | AT1G51140 | FBH3 | 45.14 |
| 42 | CgbHLH37 | 807 | Cg1g002590 | AT3G50330 | HEC2 | 53.88 | 106 | CgbHLH123 | 1377 | Cg2g022720 | AT3G20640 | – | 49.59 |
| 43 | CgbHLH39 | 762 | Cg9g016290 | AT3G56980 | ORG3 | 47.5 | 107 | CgbHLH128 | 1116 | Cg5g028960 | AT1G05805 | AKS2 | 55.06 |
| 44 | CgbHLH41 | 1599 | Cg4g008800 | AT5G56960 | – | 29.22 | 108 | CgbHLH130.1 | 2334 | Cg9g022410 | AT2G42280 | FBH4 | 57.14 |
| 45 | CgbHLH42 | 2073 | Cg5g035630 | AT4G09820 | TT8 | 56.51 | 109 | CgbHLH130.2 | 1287 | Cg7g023040 | AT2G42280 | FBH4 | 45.99 |
| 46 | CgbHLH43 | 462 | Cg1g025450 | AT5G09750 | HEC3 | 66.92 | 110 | CgbHLH137.1 | 1116 | Cg7g010980 | AT5G50915 | – | 45.96 |
| 47 | CgbHLH44 | 1200 | Cg1g004400 | AT1G18400 | BEE1 | 21.48 | 111 | CgbHLH137.2 | 837 | Cg7g022310 | AT5G50915 | – | 43.89 |
| 48 | CgbHLH45 | 555 | Cg9g004620 | AT3G06120 | MUTE | 65.03 | 112 | CgbHLH138 | 1542 | Cg1g007950 | AT2G31215 | – | 28 |
| 49 | CgbHLH46 | 1761 | Cg5g012000 | AT5G08130 | BIM1 | 53.08 | 113 | CgbHLH144 | 609 | Cg1g004390 | AT1G29950 | SACL3 | 23.28 |
| 50 | CgbHLH47 | 729 | Cg5g003170 | AT3G47640 | PYE | 51.79 | 114 | CgbHLH153 | 435 | Cg5g026370 | AT1G05710 | – | 66.42 |
| 51 | CgbHLH48 | 1227 | Cg9g022350 | AT2G42300 | – | 59.32 | 115 | CgbHLH154.1 | 933 | Cg1g012580 | AT2G31730 | – | 47 |
| 52 | CgbHLH49 | 1680 | Cg7g015600 | AT1G68920 | CIB1 | 54.17 | 116 | CgbHLH154.2 | 735 | Cg7g004740 | AT2G31730 | – | 36.36 |
| 53 | CgbHLH50 | 798 | Cg5g022610 | AT1G73830 | BEE3 | 55.27 | 117 | CgbHLH154.3 | 1107 | Cg5g035790 | AT2G31730 | – | 24.68 |
| 54 | CgbHLH51 | 843 | Cg7g002190 | AT2G40200 | – | 42.34 | 118 | CgbHLH155 | 2250 | Cg5g034200 | AT2G31280 | LHL2 | 47.44 |
| 55 | CgbHLH52 | 939 | Cg2g038660 | AT1G30670 | – | 42.51 | 119 | CgbHLH160 | 999 | Cg3g004340 | AT1G71200 | CITF1 | 40.58 |
| 56 | CgbHLH57 | 1299 | Cg5g039930 | AT4G01460 | – | 51.96 | 120 | CgbHLH162.1 | 570 | Cg2g042210 | AT4G20970 | – | 42.93 |
| 57 | CgbHLH62 | 1665 | Cg6g008950 | AT3G07340 | CIB3 | 53.27 | 121 | CgbHLH162.2 | 345 | Cg2g005390 | AT4G20970 | – | 38.16 |
| 58 | CgbHLH63 | 1215 | Cg8g007040 | AT4G34530 | CIB1 | 46.28 | 122 | CgbHLH162.3 | 279 | Cg2g005350 | AT4G20970 | – | 37.84 |
| 59 | CgbHLH68 | 1023 | Cg1g014350 | AT4G29100 | – | 51.43 | 123 | CgbHLH162.4 | 549 | Cg5g004810 | AT4G20970 | – | 35.23 |
| 60 | CgbHLH69.1 | 633 | Cg3g010300 | AT4G30980 | LRL2 | 55.43 | 124 | CgbHLH162.5 | 498 | Cg2g005320 | AT4G20970 | – | 33.54 |
| 61 | CgbHLH69.2 | 1044 | Cg4g024560 | AT4G30980 | LRL2 | 52.04 | 125 | CgbHLH162.6 | 462 | Cg2g005400 | AT4G20970 | – | 33.56 |
| 62 | CgbHLH72 | 1155 | Cg5g010830 | AT5G61270 | PIF7 | 47.52 | 126 | CgbHLH162.7 | 666 | Cg2g015830 | AT4G20970 | – | 31.28 |
| 63 | CgbHLH73.1 | 1233 | Cg1g005520 | AT5G67110 | ALC | 49.15 | 127 | CgbHLH162.8 | 498 | Cg2g005380 | AT4G20970 | – | 31.06 |
| 64 | CgbHLH73.2 | 4401 | Cg5g039380 | AT5G67110 | ALC | 29.05 | 128 | CgbHLH162.9 | 420 | Cg2g005340 | AT4G20970 | – | 33.33 |
Abbreviation (Abbr.) names are the known gene names in A. thaliana, and their full names are shown in abbreviations list
Fig. 1.
Phylogenetic tree and subfamily classifications of CgbHLH and AtbHLH proteins. The conserved protein sequences of a total of 128 CgbHLHs and 136 AtbHLHs were used to construct neighbor-joining phylogenetic tree. Arabic numerals 1 to 18 represent 18 of subfamilies. All CgbHLH proteins are highlighted with red color. The G-box, non-G-box, non-E-box and non-DNA binding of CgbHLHs are marked with different colored symbols
Multiple sequence alignment and analysis of the gene structure, conserved motif and domain of CgbHLHs
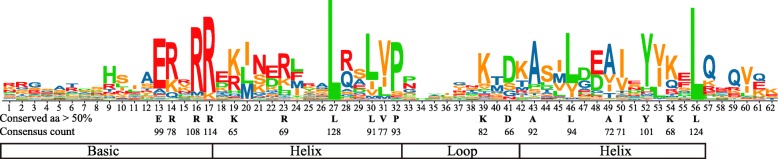
As shown in Additional file 1: Figure S1, multiple sequence alignment of 128 CgbHLHs showed that most of them were highly conserved in their bHLH domains, except that CgbLH37, CgbLH87.1, CgbLH88, CgbLH29.1, CgbLH45 and CgbLH162.9 were absent in the basic region, and CgbLH96.2 and CgbLH25.1 were short of the second helix. There were 19 conserved amino acid residues with a consensus ratio higher than 50%, including the basic region of Glu-13, Arg-14, Arg-16 and Arg-17, the first helix region of Lys-19, Arg-23, Leu-27, Leu-30, Val-31 and Pro-32, the loop region of Lys-39 and Asp-41, and the second helix region of Ala-43, Leu-46, Ala-49, Ile-50, Tyr-52, Lys-54 and Leu-56 (Fig. 2 and Additional file 1: Fig. S1). Among these, Arg-16, Arg-17, Leu-27 and Leu-56 showed extreme conservation, with a consensus ratio higher than 80%. The basic region of CgbHLHs consisted of a maximum of 17 amino acid residues, which determined the DNA binding ability of CgbHLH proteins. Based on the rule developed by Toledo-Ortiz et al. (2003) [6], we identified 104 DNA binding CgbHLHs (more than 5 basic residues existing in their basic region) and 24 non-DNA binding CgbHLHs (less than 6 basic residues existing in their basic region). The DNA binding CgbHLHs were further subdivided into 98 E-box binders (containing Glu-13 and Arg-16) and 6 non-E-box binders (lacking Glu-13 or Arg-16) based on the presence or absence of Glu-13 and Arg-16. Most of non-DNA binding and non-E-box CgbHLHs were distributed in subfamilies 11, 12, 14 and 15 (Fig. 1). Among 98 E-box CgbHLHs, 58 proteins (containing His/Lys-9, Glu-13 and Arg-17) were predicted to bind the G-box motif (CACGTG), while 40 proteins were predicted to recognize other types of E-boxes (CANNTG) and were defined as non-G-box binders (Fig. 1).
Fig. 2.
Conserved amino acids in CgbHLH domains. The height of each amino acid represents its conservation at that position. The amino acids conserved with more than 50% consensus ratio among the 128 CgbHLH domains are indicated in black letters
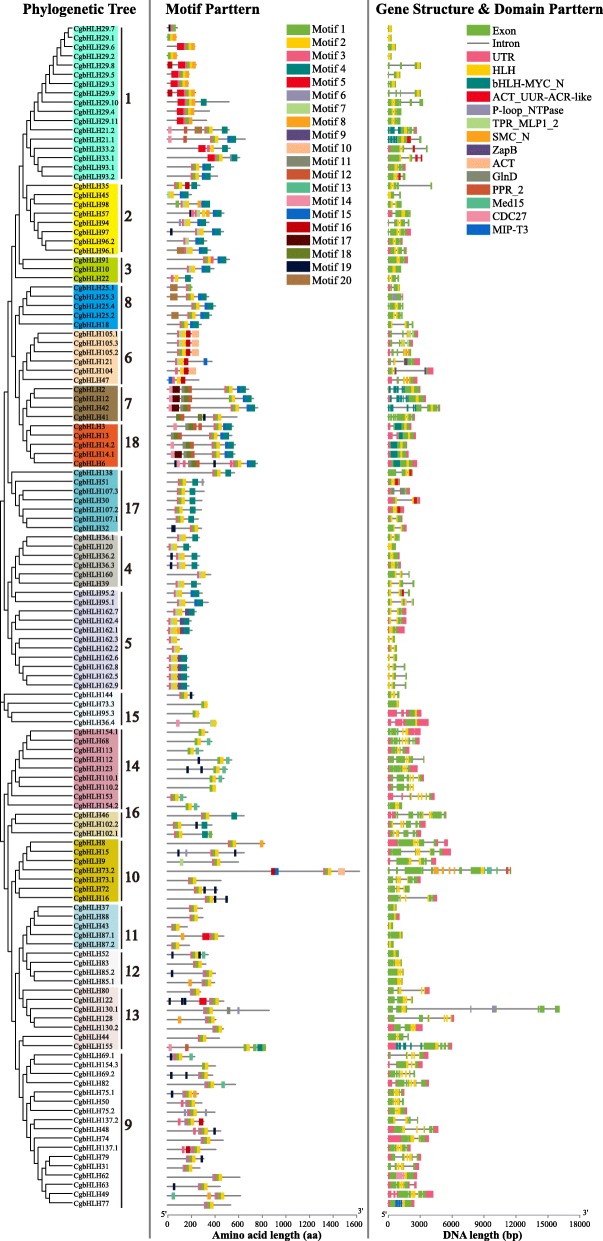
Gene structure analysis showed that 127 CgbHLHs contained 1 to 14 exons, and one CgbHLH (CgbHLH73.2) contained 21 exons (Additional file 2: Table S1). A total of 6 members (CgbHLH37/43/87.2/88 in subfamily 11 and CgbHLH6/14.1 in subfamily 18) were intronless, and 51 members had no untranslated region (UTR). Conserved motif prediction showed the constitutions of motif 1 to motif 20 in each CgbHLH protein (other possible motifs are not shown). Details of the 20 motifs are shown in Additional file 2: Table S1 and Fig. 3. Motifs 1, 2 and 3, located in bHLH domains, were found in almost all CgbHLHs, while the other motifs existed only in certain members. The members that phylogenetically clustered together or in the same subfamily often showed a similar gene structure and motif pattern. For example, the closely clustered pair CgbHLH93.1 and CgbHLH93.2 had identical motifs 1, 2, 3, 4 and 15. Most of the members of subfamily 1 contained motifs 1, 2, 3 and 4, and those of subfamily 2 contained motifs 4, 7 and 15. In addition, we found that motifs 9 and 10 were identified only in subfamily 5 and subfamily 6, respectively. Five types of motifs (motif 1, 3, 6, 18 and 19) were repeated twice or thrice in certain members, such as twice for motif 3 in CgbHLH48/50/57/137.1/137.2, twice for motif 6 in CgbHLH75.2, and thrice for motif 1 in CgbHLH155. As shown in Fig. 3, conserved domain analysis showed that all 128 CgbHLHs had an HLH domain, but there were also 36 CgbHLH genes with other domains, which might be fusion genes.
Fig. 3.
Gene structures and protein motifs of CgbHLHs. The intron-exon structures, predicted motifs 1 to 20, and conserved domains of all CgbHLHs are shown according to their phylogenic classifications
Chromosomal distribution and gene duplication of CgbHLHs
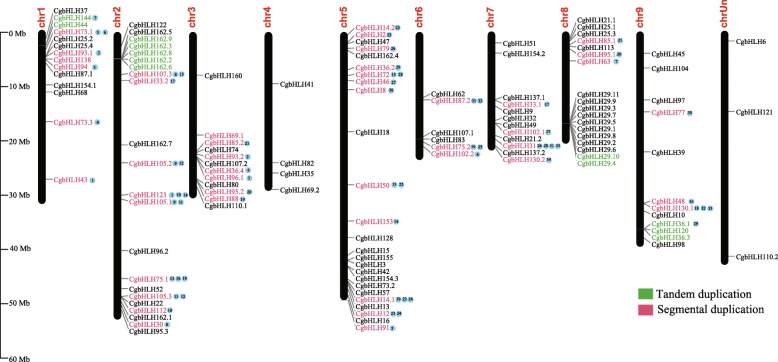
According to the genome annotation information, we found that 125 CgbHLH genes were distributed on nine chromosomes, while the chromosomal location of three CgbHLHs (CgbHLH6/110.2/121) could not be determined (Fig. 4). Chromosome 5 (chr5) contained the most CgbHLHs (25 genes), followed by chr2 (22 genes), chr8 (18 genes) and chr1 (14 genes). Although chr4 was longer than chr7 and chr8, it contained the fewest CgbHLHs (only 4 genes). The gene duplication and collinear correlations analysis showed that a total of 12 CgbHLHs were identified to be tandem duplicated genes, distributed on chr1, chr2, chr8 and chr9 (Fig. 4). In addition, 47 CgbHLHs were predicted to be segmental duplicated genes, accounting for about 37% of all CgbHLH genes, and the identified collinear genes showed great overlapping with these segmental duplicated genes (Fig. 4). These results suggested that the tandem and segmental duplication events that occurred during citrus evolution might have played an essential role in CgbHLH family expansion.
Fig. 4.
Chromosomal localizations and gene duplications of CgbHLH genes. The green and red colors represent tandem duplicated and segmental duplicated genes, respectively. The CgbHLH genes followed by the same number (highlighted with blue circles) are collinear genes
GO annotation and protein interactions of CgbHLHs
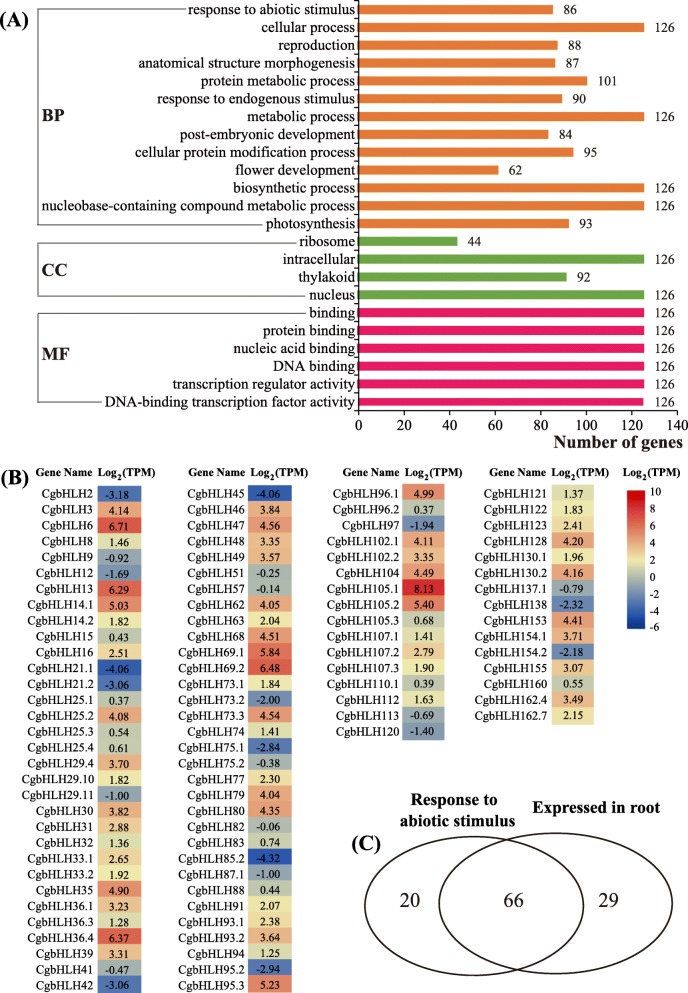
To determine potential functions of each CgbHLH gene, GO annotation was performed for all CgbHLHs. As shown in Fig. 5a, except for two genes that were not annotated a GO term, the other 126 CgbHLHs were annotated in three functional categories: biological process (BP), cellular component (CC) and molecular function (MF). In the BP category, these CgbHLHs were further annotated to respond to abiotic stimulus (86 genes), reproduction (88 genes), post-embryonic development (84 genes), flower development (62 genes) and photosynthesis (93 genes). Under CC and MF categories, we found that all 126 CgbHLHs were annotated to the nucleus, protein or DNA binding, and transcription factor activity, which agreed well with TF property of these CgbHLHs.
Fig. 5.
Functional annotation and tissue expression of CgbHLH genes. a All annotated GO terms including biological process (BP), cellular component (CC) and molecular function (MF) of 126 CgbHLHs (two other CgbHLHs were not annotated a GO term). b Expression heatmap of CgbHLHs. The TPM values were generated from the RNAseq data of pummelo roots in a public database. c Venn diagram shows the number of the genes that respond to abiotic stimulus and expresssed in roots
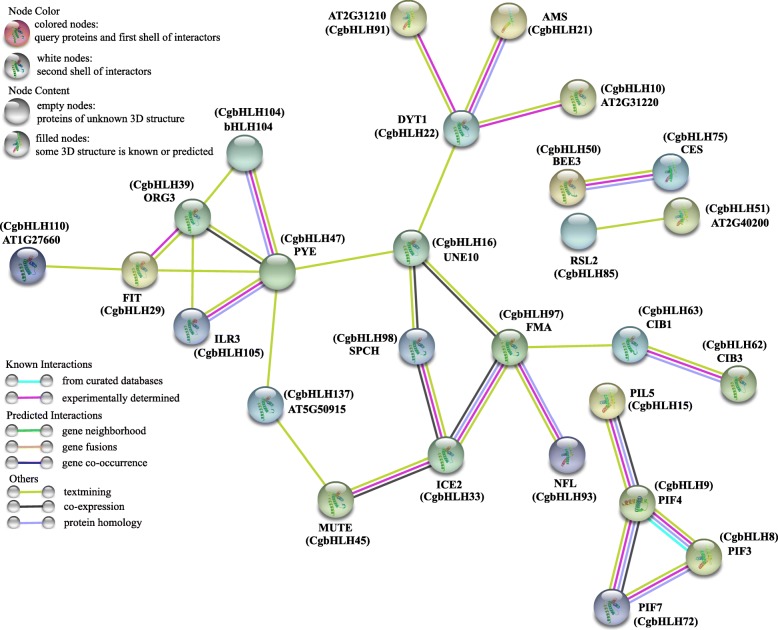
Proteins in the bHLH family often interact with each other by forming homodimers or heterodimers, which are essential for binding and regulating downstream target gene expression. Thus, prediction of protein interactions of CgbHLHs was performed by using orthologous bHLHs of A. thaliana. The result showed that a total of 27 CgbHLH proteins were predicted to have protein interaction relationships (Fig. 6). For example, CgbHLH29 (FIT ortholog), CgbHLH39, CgbHLH47 (PYE ortholog), CgbHLH104, CgbHLH105 (ILR3 ortholog) and CgbHLH110 were predicted to have direct or indirect interactions with each other. CgbHLH33 (ICE2 ortholog) was predicted to interact with CgbHLH45 (MUTE ortholog), CgbHLH97 (FMA ortholog) and CgbHLH98 (SPCH) directly. Moreover, interactions among several phytochrome interacting factor (PIF) orthologous proteins (CgbHLH8, CgbHLH9, CgbHLH15 and CgbHLH72) were predicted. Overall, the result provides an important reference for identifying true interactions of CgbHLHs with biochemical experiments.
Fig. 6.
Predicted protein-protein interactions of CgbHLHs according to their orthologs in A. thaliana. In the network, only the pairs with higher than 40% sequence identity between CgbHLHs and AtbHLHs and with an interaction score > 0.7 are shown. Line and node colors indicate the different kinds and degrees of interactions, respectively. The filled or empty nodes represent known or unknown 3D structures, respectively. The abbreviated names are the genes that have been reported in A. thaliana
Candidate CgbHLH members in response to Fe deficiency
As described in the “Introduction,” bHLH genes play an essential regulatory role in plant Fe homeostasis. To identify candidate CgbHLH members in response to Fe deficiency in pummelo, their expression levels in the root were evaluated by analyzing the previous RNAseq data and performing qRT-PCR confirmation. Based on analysis of previous RNAseq data published by Guo et al. (2017) [39], a total of 95 CgbHLH genes were determined to be transcribed in the root of normal cultured pummelo (Fig. 5b). Their transcription levels (expressed as the TPM value) ranged from 0.05 (CgbHLH85.2) to 281.08 (CgbHLH105.1). These root-expressed genes are preliminarily considered to respond to Fe deficiency, as the root is directly responsible for ion uptake. To further narrow the range of candidates, we searched for root-expressed genes among the GO annotated abiotic stimulus responsive genes; as a result, 66 CgbHLHs were found to overlap (Fig. 5c). Due to lower sensitivity of qRT-PCR than RNAseq in determining gene expression, we finally selected 39 CgbHLHs with TPM values higher than 3.00 from these 66 genes for qRT-PCR confirmation. In addition, three genes (CgbHLH104, CgbHLH105.1 and CgbHLH105.2) that are orthologous to known Fe-related bHLHs but that were not included in the 39 CgbHLHs were also determined by qRT-PCR in this study.
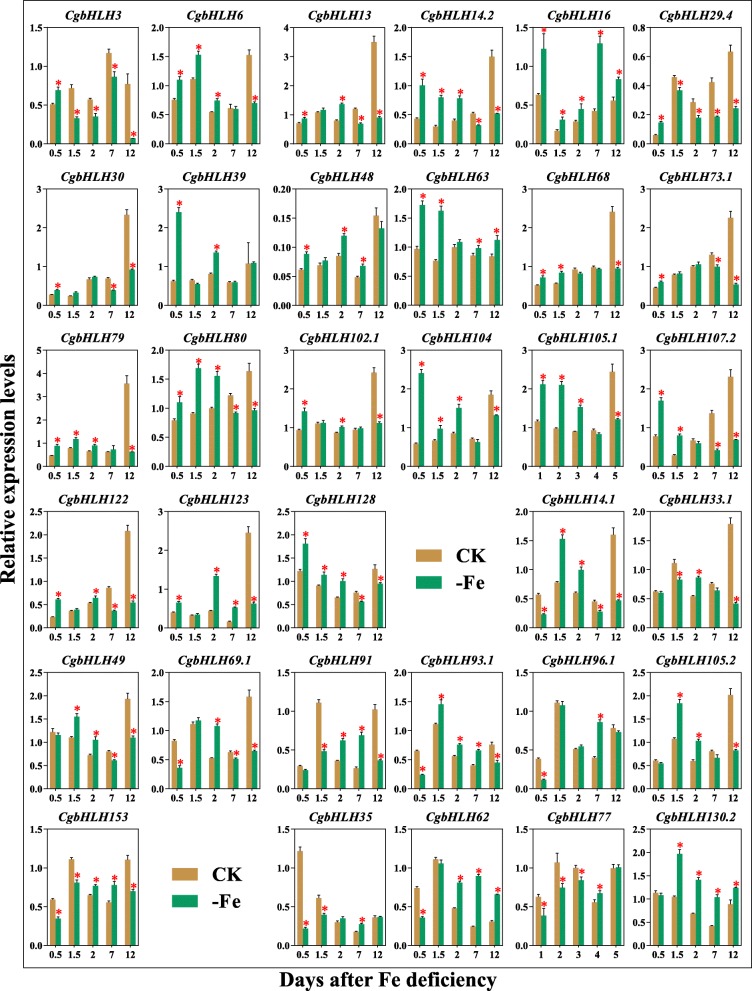
Except for eight genes were still undetectable, the other 34 genes were determined to be up- or down-regulated in roots of pummelo after 0.5 d, 1.5 d, 2 d, 7 d, and 12 d of Fe-deficient (−Fe) treatments when comparing with those at corresponding time points of CK (Fig. 7). In general, the tested genes showed three types of expression patterns. The first type, such as CgbHLH3, CgbHLH6, CgbHLH13, CgbHLH14.2, CgbHLH16, CgbHLH29.4, CgbHLH30, CgbHLH39, CgbHLH48, CgbHLH63, CgbHLH68, CgbHLH73.1, CgbHLH79, CgbHLH80, CgbHLH102.1, CgbHLH104, CgbHLH105.1, CgbHLH107.2, CgbHLH122, CgbHLH123 and CgbHLH128, was up-regulated from an early period (0.5 d), which indicated an early response of these genes to Fe deficiency. The second type, such as CgbHLH14.1, CgbHLH33.1, CgbHLH49, CgbHLH69.1, CgbHLH91, CgbHLH93.1, CgbHLH96.1, CgbHLH105.2 and CgbHLH153, was up-regulated at medium periods but was down-regulated during early and late periods. The third type was only up-regulated during late periods, such as CgbHLH35, CgbHLH62, CgbHLH77 and CgbHLH130.2. Statistical analysis showed that most of the tested genes were significantly differentially expressed at multiple time points. In particular, CgbHLH6, CgbHLH14.2, CgbHLH16, CgbHLH48, CgbHLH63, CgbHLH79, CgbHLH80, CgbHLH104, CgbHLH105.1, CgbHLH123, CgbHLH128, CgbHLH93.1, CgbHLH62 and CgbHLH130.2 were significantly up-regulated while CgbHLH3, CgbHLH29.4, CgbHLH14.1, CgbHLH69.1, CgbHLH153 and CgbHLH77 were significantly down-regulated at least three time points. Moreover, CgbHLH16 and CgbHLH63 showed continuous up-regulation at all tested times. We speculate that these significantly differentially expressed CgbHLH genes are possibly the key candidates that respond to Fe deficiency in pummelo.
Fig. 7.
qRT-PCR analysis of 34 CgbHLH genes in pummelo roots. Relative expression levels of 34 CgbHLHs were determined in 0.5 d, 1.5 d, 2 d, 7 d and 12 d of Fe-deficient (−Fe) roots and normal (CK) roots sampled at corresponding time points. Data are means ± SE of three biological replicates. Asterisks on the error bars indicate significant differences (t-test, P < 0.05) between -Fe and CK
Discussion
The genomes of several citrus varieties have been released, including the sweet orange (Citrus sinensis) genome published by Xu et al. (2011) [40], the clementine mandarin genome published by Wu et al. (2014) [41], and the pummelo, papeda (Citrus ichangensis) and citron (Citrus medica) genomes published by Wang et al. (2017) [38]. After the completion of these genomes, systematic identification and analysis of important gene families have been widely reported for citrus [1, 42–44]. These studies have provided us a comprehensive understanding of potential gene functions and have promoted related research progress. However, no such study has been done on the citrus bHLH family, except that Geng and Liu (2018) [45] identified 56 putative bHLH genes from sweet orange. In this study, we used a genome-wide approach to identify 128 CgbHLHs from pummelo. This number possibly indicates the real quantity of bHLH genes in most citrus varieties because the ratio of CgbHLHs in the pummelo genome (0.42%) is very similar to that of tomato (0.46%) [46], rice (0.44%) [13], poplar (0.40%) [11], apple (0.42%) [27], and grape (0.40%) [47]. With respect to the bHLHs of these plants, a high proportion of tandem duplications and segmental duplications has been determined, which indicates that bHLH expansion was possibly derived from gene duplication during evolution [48]. In citrus, an ancient whole-genome duplication event (WGD) occurred [40], and it could have led to this chromosome segmental duplication.
In the CgbHLH family, 19 amino acid residues are highly conserved, with a consensus ratio higher than 50%. These residues are also highly conserved in bHLHs of A. thaliana [15], maize [49] and grape [47], indicating conservation of bHLH families among different plants. Evidence has shown that at least five basic residues in the basic region of the bHLH domain determine DNA binding activity of bHLHs; Glu-13 is critical in specific recognition of the E-box DNA binding motif, while Arg-16 functions in fixing and stabilizing the position of Glu-13; moreover, His/Lys-9 and Arg-17 confer specificity to G-box (CACGTG) recognition [15]. Based on these findings, 128 CgbHLHs can be classified into four types: G-Box, non-G-Box, non-E-Box and non-DNA binding (Fig. 1). This classification indicates the basic functional property of each CgbHLH. We also noticed that Leu-27 was conserved in all CgbHLH proteins, and a similar result was found in AtbHLHs [15]. A previous study showed that the Leu residue at position 27 is necessary for dimer formation [50]. Thus, Leu-27 of CgbHLHs suggests their dimerization capacity.
To date, only very limited citrus bHLH genes have been functionally characterized, including PtrbHLH, an ortholog of AtbHLH33 (ICE2), that was isolated from trifoliate orange (Poncirus trifoliata) and proved to confer cold tolerance by regulating both POD- and CAT-mediated reactive oxygen species (ROS) scavenging [29, 45]. CsbHLH18 of sweet orange was also proved to modulate cold tolerance and ROS homeostasis [51]. Moreover, a CubHLH1 of Satsuma mandarin (Citrus unshiu Marc.) was overexpressed in transgenic tomato fruit, resulting in modulation of carotenoid metabolism [52]. Generally, the potential functions of the unknown genes could be predicted through their orthologous genes. Based on phylogenetic and orthologous analyses, the CgbHLH family was classified into 18 subfamilies (Fig. 1). The closely clustered CgbHLHs and AtbHLHs in the same subfamily/group may have similar functions. In group 1, AtbHLH29 (FIT), AtbHLH33 (ICE2), AtbHLH116 (ICE1), AtbHLH93 (NFL), AtbHLH61, and AtbHLH21 (AMS) have been shown to be involved in Fe deficiency, cold tolerance and flower development [32, 53–55], suggesting that there are similar functions for the CgbHLHs of this group. In the other groups, many AtbHLH genes such as AtbHLH70 (MYC70) and AtbHLH45 (MUTE) in group 2, AtbHLH10, AtbHLH89 and AtbHLH91 in group 3, AtbHLH38, AtbHLH39, AtbHLH100 and AtbHLH101 in group 4, etc. have also been functionally characterized [9, 33, 56–58]. Based on these previous studies, the potential functions of most CgbHLH genes could be predicted. Furthermore, GO annotation and protein-protein interaction analysis of the CgbHLHs could help us to understand their possible functions.
Fe deficiency-regulated bHLHs are the main concerns in this study. qRT-PCR showed that most of the tested CgbHLH genes were significantly differentially expressed at two or more time points under Fe deficiency. Among them, CgbHLH29.4, CgbHLH39, CgbHLH104, CgbHLH105.1 and CgbHLH105.2 are orthologs of known Fe deficiency-responsive AtbHLHs [32, 33, 36], and their expression patterns further supported the putative function on Fe deficiency. CgbHLH6, CgbHLH13, CgbHLH14.1 and CgbHLH14.2 are orthologs of AtHLH6 (MYC2), AtHLH13 (JAM2) and AtHLH14, respectively, and these three AtHLHs of A. thaliana have been shown to participate in jasmonic acid (JA) signaling [6, 59]. Because JA is a negative regulator of Fe uptake, it suggests that these four CgbHLH genes may also be involved in the response to Fe deficiency. Interestingly, several of the significantly expressed CgbHLHs such as CgbHLH105.1, CgbHLH104, CgbHLH13 and CgbHLH130.2 also showed a high expression level in roots based on RNAseq data, which further indicates their possible functions in Fe deficiency. Moreover, we found that CgbHLH104 and CgbHLH105.1 belong to the same subfamily (Group 6) and have a similar gene structure, pI and exon number. Their orthologous bHLHs of A. thaliana, AtbHLH104 and AtbHLH105, as well as all the other members in group 6 (AtbHLH47 and AtbHLH121) have been functionally identified in Fe deficiency [7, 8, 34–36]. Herein, CgbHLH104 and CgbHLH105.1 also showed significant up-regulation under Fe deficiency. Thus, we speculate that these two CgbHLHs most likely have similar functions with AtbHLH104 and AtbHLH105 under Fe deficiency. Apart from these known Fe deficiency-responsive bHLHs, there were also significantly differentially expressed CgbHLHs, but direct evidence could not be found regarding their orthologs that have a function in plant Fe deficiency, such as CgbHLH16, CgbHLH48, CgbHLH62, CgbHLH63, CgbHLH79, CgbHLH80, CgbHLH123, CgbHLH128 and CgbHLH130.2. New and essential bHLHs that regulate Fe homeostasis are expected to be identified from these candidates in future. In particular, CgbHLH16 was continuously up-regulated at all tested times of –Fe treatment, and it was also predicted to interact with an important Fe deficiency-responsive TF, PYE (bHLH47), based on protein-protein interaction prediction (Fig. 6). In addition, CgbHLH62 and CgbHLH63 were predicted to interact with each other while their expression levels showed the opposite tendency at the same time points, which indicates that there may be negative regulation relationships between them under Fe deficiency. Overall, these results provide us abundant information for understanding of the citrus bHLH genes, but more work including transgenic and biochemical experiments needs to be done to further confirm our speculations and illustrate their functions.
Conclusions
In summary, 128 CgbHLH proteins were identified from pummelo (Citrus grandis) genome and were classified into 18 subfamilies based on the phylogenetic relationship with AtbHLH proteins. The detailed analysis of Mw, pI, sequence alignment, phylogeny, gene duplication, chromosomal distribution, gene structure and protein motif suggested basic properties of each CgbHLH. GO annotation and protein-protein interaction analysis further revealed the potential functions of CgbHLHs. qRT-PCR showed that 34 of the tested CgbHLH genes were up- or down-regulated under Fe deficiency, and some of them such as CgbHLH6, CgbHLH14.2, CgbHLH16, CgbHLH48, CgbHLH63, CgbHLH79, CgbHLH80, CgbHLH104, CgbHLH105.1, CgbHLH123, CgbHLH128, CgbHLH93.1, CgbHLH62, CgbHLH130.2, CgbHLH3, CgbHLH29.4, CgbHLH14.1, CgbHLH69.1, CgbHLH153 and CgbHLH77 were considered the key candidates that respond to Fe deficiency. This study lays the foundation for further functional elucidation of CgbHLH genes in citrus.
Methods
Plant material and treatment
Seeds of pummelo (Citrus grandis) were collected from the Citrus Research Institute, Southwest University, Chongqing, China. After surface disinfection with 2% sodium hypochlorite, seed coats were removed, and seeds were then germinated on wet filter papers that filled 96-cell plastic boxes with lids. After 2 weeks germination, uniform seedlings were transferred into modified Hoagland’s solution for hydroponics as described by Fu et al. (2017) [1]. When the seedlings had grown four leaves, half of them were renewed with Fe deficient (0.5 μM Fe-EDTA) solution (−Fe), while the others received normal (50 μM Fe-EDTA) solution (CK). The culture conditions were a temperature of 28 °C and a relative humidity of 70% under a 16-h photoperiod (50 μmol m− 2 s− 1). Both –Fe and CK groups had three biological replicates, and each replicate contained 14 seedlings. After 0.5 d, 1.5 d, 2 d, 7 d, and 12 d of treatment, roots of –Fe and CK were sampled and then frozen in liquid nitrogen immediately.
Identification of bHLH members in pummelo
The genome data of pummelo were downloaded from the Orange (Citrus sinensis) Genome Annotation Project (http://citrus.hzau.edu.cn/orange/download/index.php). Then, we used two methods to search for all bHLHs of pummelo. Firstly, the hidden Markov model (HMM) file of the HLH domain (PF00010) obtained from the Pfam database (http://pfam.xfam.org/) was used as the query to scan the pummelo genome with HMMER software (http://hmmer.org/). Secondly, the bHLH domain sequences of A. thaliana (AtbHLHs) were used as queries to blast against the pummelo genome. After removal of redundant sequences, all putative bHLHs obtained from the two methods were submitted to the NCBI (National Center of Biotechnology Information) conserved domain database (CDD, https://www.ncbi.nlm.nih.gov/Structure/bwrpsb/bwrpsb.cgi) to further verify the existence of the conserved bHLH domain. All the identified bHLH proteins of pummelo were named as CgbHLHs.
Multiple sequence alignment and phylogenetic tree construction
Molecular weights (MWs) and isoelectric points (pIs) of CgbHLH proteins were analyzed with ExPASy (http://web.expasy.org/compute_pi/). Conserved domain sequences of CgbHLHs were submitted to ClustalW to perform multiple sequence alignment, and the BioEdit was used to edit the aligned sequences and shade the conserved sites. To visualize the conserved motifs, the sequences were analyzed with WEBLOGO programs (http://weblogo.berkeley.edu). The neighbor-joining phylogenetic tree was constructed by using MEGA7 and 1000 replicates in bootstrap analysis. In addition, the full-length protein sequences of all CgbHLHs and known AtbHLHs were also aligned in Clustal Omega (https://www.ebi.ac.uk/Tools/msa/clustalo/) to obtain their sequence identity (%) data, and the CgbHLH-AtbHLH pairs with highest percent identity were considered orthologs.
Gene structure, conserved motifs, chromosomal location and gene duplication analysis
Gene structure information of each CgbHLH was obtained from the generic feature format (GFF) file downloaded from the pummelo genome database, and TBtools [60] was used to display their intron-exon structures. Conserved motifs of each CgbHLH protein were predicted in the MEME [61] web tool with the maximum number of motifs set at 20, and the other parameters were the default values. The chromosomal locations of the CgbHLH genes were provided by the pummelo genome database, and TBtools software was used to draw the location images. To predict the gene duplication, the BLASTP program was first used to blast against the pummelo genome using all peptide sequences of pummelo as the queries; then, the top matches (e value ≤1e− 05) and their GFF files were inputted into MCScanX software [62] to identify tandem duplications, segmental duplications, and collinear correlations.
Gene ontology (GO) annotation and RNA-sequencing (RNAseq) data analysis
GO annotation of all the CgbHLH proteins was performed with the Blast2GO program, and A. thaliana was chosen as the reference. To evaluate the expression levels of CgbHLH genes in pummelo, the published RNAseq raw data (SRR4050288/SRR4050289) of pummelo [39] were downloaded from the NCBI database (https://www.ncbi.nlm.nih.gov/sra/). Then, the raw data were analyzed with the linux system of the fastqc and trimommatic programs to remove adaptors and low quality sequences. The kallisto program was used to map sequence data to the pummelo genome, and the featurecount program was used to calculate the values of the transcripts per million reads (TPM).
Prediction of the protein-protein interaction network
The interaction network of CgbHLHs was predicted in STRING (version 11.0, http://string-db.org) website that contains known and predicted protein-protein interactions of different organisms [63]. In detail, all the CgbHLH protein sequences were first submitted to the STRING website as queries, and A. thaliana was chosen as the reference organism for blasting because pummulo has no protein-protein interaction database and is not included in STRING. After blasting, the matched homologs of A. thaliana with the highest scores (Bitscore) and higher than 40% identity were used to construct the network. To ensure reliability, only the predicted interactions with interaction scores > 0.7 (high confidence level was set in STRING) were shown in the network.
RNA extraction and quantitative real-time PCR (qRT-PCR) analysis
Total RNA was extracted from the –Fe and CK roots of pummelo by using the RNAprep pure plant kit (Cat#DP432, Tiangen Biotech Co., Ltd., Beijing, China), and the RNA concentration and quality were determined with a Nanodrop 2000 spectrophotometer (Thermo Scientific, Waltham, MA, USA). Then, 1 μg of high-quality RNA was used for cDNA synthesis with HiScript II Q RT Super Mix (Cat#R222–01, Vazyme Biotech Co., Ltd., Nanjing, China) according to the manufacturer’s instructions. qRT-PCR was performed on the Bio-Rad CFX Connect RealTime system by using ChamQ™ Universal SYBR® qPCR Master Mix (Cat#Q711, Vazyme Biotech Co., Ltd., Nanjing, China). Each PCR reaction contained 5.0 μL SYBR mix, 0.2 μM primers, and 1.0 μL diluted cDNA in a final volume of 10 μL. The 2-ΔΔCT method was used to normalize and calculate the expression level of each tested gene relative to an internal reference gene, actin (Cs1g05000.1). All the primers of CgbHLH genes are listed in Additional file 3: Table S2. Three biological replicates and three technical replicates were performed for each treatment. The presented data herein are means ± standard errors (SE) of three biological replicates. Student’s t-test was used to analyze the statistical differences between -Fe and corresponding CK samples, taking P < 0.05 as significantly different.
Supplementary information
Additional file 1: Figure S1. Sequence alignment of 128 CgbHLH proteins.
Additional file 2: Table S1. Predicted MW, pI, exon number, UTR and motifs of 128 CgbHLH proteins.
Additional file 3: Table S2. The primers of CgbHLH genes used for qRT-PCR.
Acknowledgments
We thank LetPub (http://www.letpub.com) for linguistic assistance during the preparation of this manuscript.
Abbreviations
- ABS5
Abnormal shoot 5
- AKS2
ABA-responsive kinase substrate 2
- ALC
ALCATRAZ
- AMS
Aborted microspores
- BEE1
BR enhanced expression 1
- bHLH
Basic/helix-loop-helix
- BIM1
BES1-interacting myc-like 1
- BP
Biological process
- CC
Cellular component
- CDD
Conserved domain database
- CES
CESTA
- CIB1
Cryptochrome-interacting basic/helix-loop-helix 1
- CIL2
CIB1 like protein 2
- CITF1
Cu-deficiency induced transcription factor 1
- DYT1
Dysfunctional tapetum 1
- EGL3
Enhancer of glabra 3
- FBH1
Flowering bHLH 1
- Fe
Iron
- FIT
FER-like iron-deficiency-induced transcription factor
- FRO2
Ferric reduction oxidase 2
- FRU
Fer-like iron deficiency induced transcription factor 3
- GO
Gene ontology
- HEC
Hecate
- HMM
Hidden markov model
- ICE2
Inducer of CBF expression 2
- ILR3
IAA-leucine resistant 3
- IRT1
Iron-regulated transporter 1
- JA
Jasmonic acid
- JAI1
Jasmonate insensitive 1
- JAM
Jasmonate associated MYC2 like
- LHL2
Lonesome highway like 2
- LRL2
LJRHL1-like 2
- MF
Molecular function
- MW
Molecular weight
- NCBI
National center of biotechnology information
- NFL
No flowering in short day
- ORG3
OBP3-responsive gene 3
- pI
Isoelectric point
- PIF
Phytochrome interacting factor
- PIL5
Phytochrome interacting factor 3-like 5
- PYE
POPEYE
- qRT-PCR
Quantitative real-time PCR
- RGE1
Retarded growth of embryo 1
- RHD6
Root hair defective 6
- ROS
Reactive oxygen species
- RSL2
Root hair defective 6-like 2
- SPCH
Speechless
- SPT
SPATULA
- TFs
Transcription factors
- TMO5
Target of monopteros 5
- TPM
Transcripts per million reads
- TT8
Transparent testa 8
- UNE10
Unfertilized embryo sac 10
- UTR
Untranslated region
- WGD
Whole-genome duplication event
Authors’ contributions
XZF and LZP conceived and designed the study. XYZ collected the public data and performed bioinformatics analysis. JYQ and XYZ performed qRT-PCR. QLH, YYX and YZH prepared experimental materials and performed the Fe-deficient treatment. XYZ and XZF prepared all the figures and tables and wrote the paper. All authors read and approved the final manuscript.
Funding
This work was financially supported by the National Key Research and Development Program of China (2018YFD1000300, 2017YFD0202000), the National Natural Science Foundation of China (31772280), the National Citrus Engineering Research Center (NCERC2019001), and the Technological Innovation and Application Program of Chongqing (cstc2018jscx-mszdX0002).
Availability of data and materials
All data generated during this study are included in this published article and its supplementary information files. In addition, the protein and CDS sequences of CgbHLHs analysed in this study were retrieved from the Orange Genome Annotation Project, http://citrus.hzau.edu.cn/orange/download/index.php. The protein sequences of AtbHLHs were retrieved from the TAIR database, https://www.arabidopsis.org/browse/genefamily/bHLH.jsp.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information accompanies this paper at 10.1186/s12864-020-6644-7.
References
- 1.Fu XZ, Zhou X, Xing F, Ling LL, Chun CP, Cao L, et al. Genome-wide identification, cloning and functional analysis of the zinc/iron-regulated transporter-like protein (ZIP) gene family in trifoliate orange (Poncirus trifoliata L. Raf.) Front Plant Sci. 2017;8:588. doi: 10.3389/fpls.2017.00588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Fu LN, Zhu QQ, Sun YY, Du W, Pan ZY, Peng SA. Physiological and transcriptional changes of three citrus rootstock seedlings under iron deficiency. Front Plant Sci. 2017;8:1104. doi: 10.3389/fpls.2017.01104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Brumbarova T, Bauer P, Ivanov R. Molecular mechanisms governing Arabidopsis iron uptake. Trends Plant Sci. 2015;20(2):124–133. doi: 10.1016/j.tplants.2014.11.004. [DOI] [PubMed] [Google Scholar]
- 4.Jeong J, Guerinot ML. Homing in on iron homeostasis in plants. Trends Plant Sci. 2009;14(5):280–285. doi: 10.1016/j.tplants.2009.02.006. [DOI] [PubMed] [Google Scholar]
- 5.Kobayashi T, Nishizawa NK. Iron uptake, translocation, and regulation in higher plants. Annu Rev Plant Biol. 2012;63:131–152. doi: 10.1146/annurev-arplant-042811-105522. [DOI] [PubMed] [Google Scholar]
- 6.Cui Y, Chen CL, Cui M, Zhou WJ, Wu HL, Ling HQ. Four IVa bHLH transcription factors are novel interactors of FIT and mediate JA inhibition of iron uptake in Arabidopsis. Mol Plant. 2018;11(9):1166–1183. doi: 10.1016/j.molp.2018.06.005. [DOI] [PubMed] [Google Scholar]
- 7.Gao Fei, Robe Kevin, Bettembourg Mathilde, Navarro Nathalia, Rofidal Valérie, Santoni Véronique, Gaymard Frédéric, Vignols Florence, Roschzttardtz Hannetz, Izquierdo Esther, Dubos Christian. The Transcription Factor bHLH121 Interacts with bHLH105 (ILR3) and Its Closest Homologs to Regulate Iron Homeostasis in Arabidopsis. The Plant Cell. 2019;32(2):508–524. doi: 10.1105/tpc.19.00541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Li XL, Zhang HM, Ai Q, Liang G, Yu DQ. Two bHLH transcription factors, bHLH34 and bHLH104, regulate Iron homeostasis in Arabidopsis thaliana. Plant Physiol. 2016;170(4):2478–2493. doi: 10.1104/pp.15.01827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wang N, Cui Y, Liu Y, Fan H, Du J, Huang Z, et al. Requirement and functional redundancy of ib subgroup bHLH proteins for iron deficiency responses and uptake in Arabidopsis thaliana. Mol Plant. 2013;6(2):503–513. doi: 10.1093/mp/sss089. [DOI] [PubMed] [Google Scholar]
- 10.Murre C, Mccaw PS, Baltimore D. A new DNA-binding and dimerization motif in immunoglobulin enhancer binding, daughterless, myod, and myc proteins. Cell. 1989;56(5):777–783. doi: 10.1016/0092-8674(89)90682-X. [DOI] [PubMed] [Google Scholar]
- 11.Carretero-Paulet L, Galstyan A, Roig-Villanova I, Martinez-Garcia JF, Bilbao-Castro JR, Robertson DL. Genome-wide classification and evolutionary analysis of the bHLH family of transcription factors in Arabidopsis, poplar, Rice, Moss, and algae. Plant Physiol. 2010;153(3):1398–1412. doi: 10.1104/pp.110.153593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Feller A, Machemer K, Braun EL, Grotewold E. Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. Plant J. 2011;66(1):94–116. doi: 10.1111/j.1365-313X.2010.04459.x. [DOI] [PubMed] [Google Scholar]
- 13.Li XX, Duan XP, Jiang HX, Sun YJ, Tang YP, Yuan Z, et al. Genome-wide analysis of basic/helix-loop-helix transcription factor family in rice and Arabidopsis. Plant Physiol. 2006;141(4):1167–1184. doi: 10.1104/pp.106.080580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Sun X, Wang Y, Sui N. Transcriptional regulation of bHLH during plant response to stress. Biochem Bioph Res Co. 2018;503(2):397–401. doi: 10.1016/j.bbrc.2018.07.123. [DOI] [PubMed] [Google Scholar]
- 15.Toledo-Ortiz G, Huq E, Quail PH. The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell. 2003;15(8):1749–1770. doi: 10.1105/tpc.013839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Massari ME, Murre C. Helix-loop-helix proteins: regulators of transcription in eucaryotic organisms. Mol Cell Biol. 2000;20(2):429–440. doi: 10.1128/MCB.20.2.429-440.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Buck MJ, Atchley WR. Phylogenetic analysis of plant basic helix-loop-helix proteins. J Mol Evol. 2003;56(6):742–750. doi: 10.1007/s00239-002-2449-3. [DOI] [PubMed] [Google Scholar]
- 18.Atchley WR, Terhalle W, Dress A. Positional dependence, cliques, and predictive motifs in the bHLH protein domain. J Mol Evol. 1999;48(5):501–516. doi: 10.1007/PL00006494. [DOI] [PubMed] [Google Scholar]
- 19.Wang HP, Li Y, Pan JJ, Lou DJ, Hu YR, Yu DQ. The bHLH transcription factors MYC2, MYC3, and MYC4 are required for jasmonate-mediated inhibition of flowering in Arabidopsis. Mol Plant. 2017;10(11):1461–1464. doi: 10.1016/j.molp.2017.08.007. [DOI] [PubMed] [Google Scholar]
- 20.Ding WN, Yu ZM, Tong YL, Huang W, Chen HM, Wu P. A transcription factor with a bHLH domain regulates root hair development in rice. Cell Res. 2009;19(11):1309–1311. doi: 10.1038/cr.2009.109. [DOI] [PubMed] [Google Scholar]
- 21.Penfield S, Josse EM, Kannangara R, Gilday AD, Halliday KJ, Graham IA. Cold and light control seed germination through the bHLH transcription factor SPATULA. Curr Biol. 2005;15(22):1998–2006. doi: 10.1016/j.cub.2005.11.010. [DOI] [PubMed] [Google Scholar]
- 22.Wang LH, Tang W, Hu YW, Zhang YB, Sun JQ, Guo XH, et al. A MYB/bHLH complex regulates tissue-specific anthocyanin biosynthesis in the inner pericarp of red-centered kiwifruit Actinidia chinensis cv. Hongyang. Plant J. 2019;99(2):359–378. doi: 10.1111/tpj.12894. [DOI] [PubMed] [Google Scholar]
- 23.Hu DG, Sun CH, Zhang QY, An JP, You CX, Hao YJ. Glucose sensor MdHXK1 phosphorylates and stabilizes MdbHLH3 to promote anthocyanin aiosynthesis in apple. PLoS Genet. 2016;12(8):e1006273. doi: 10.1371/journal.pgen.1006273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Xu WJ, Dubos C, Lepiniec L. Transcriptional control of flavonoid biosynthesis by MYB-bHLH-WDR complexes. Trends Plant Sci. 2015;20(3):176–185. doi: 10.1016/j.tplants.2014.12.001. [DOI] [PubMed] [Google Scholar]
- 25.An JP, Li HH, Song LQ, Su L, Liu X, You CX, et al. The molecular cloning and functional characterization of MdMYC2, a bHLH transcription factor in apple. Plant Physiol Biochem. 2016;108:24–31. doi: 10.1016/j.plaphy.2016.06.032. [DOI] [PubMed] [Google Scholar]
- 26.Nakata M, Mitsuda N, Herde M, Koo AJK, Moreno JE, Suzuki K, et al. A bHLH-type transcription factor, ABA-inducible bHLH-type transcription factor/JA-associated MYC2-like1, acts as a repressor to negatively regulate jasmonate signaling in Arabidopsis. Plant Cell. 2013;25(5):1641–1656. doi: 10.1105/tpc.113.111112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Mao K, Dong QL, Li C, Liu CH, Ma FW. Genome wide identification and characterization of apple bHLH transcription factors and expression analysis in response to drought and salt stress. Front Plant Sci. 2017;8:480. doi: 10.3389/fpls.2017.00480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Chen YY, Li MY, Wu XJ, Huang Y, Ma J, Xiong AS. Genome-wide analysis of basic helix-loop-helix family transcription factors and their role in responses to abiotic stress in carrot. Mol Breed. 2015;35(5):125. doi: 10.1007/s11032-015-0319-0. [DOI] [Google Scholar]
- 29.Huang XS, Wang W, Zhang Q, Liu JH. A basic helix-loop-helix transcription factor, PtrbHLH, of Poncirus trifoliata confers cold tolerance and modulates peroxidase-mediated scavenging of hydrogen peroxide. Plant Physiol. 2013;162(2):1178–1194. doi: 10.1104/pp.112.210740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Ling HQ, Bauer P, Bereczky Z, Keller B, Ganal M. The tomato fer gene encoding a bHLH protein controls iron-uptake responses in roots. Proc Natl Acad Sci U S A. 2002;99(21):13938–13943. doi: 10.1073/pnas.212448699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Jakoby M, Wang H-Y, Reidt W, Weisshaar B, Bauer P. FRU (BHLH029) is required for induction of iron mobilization genes in Arabidopsis thaliana. FEBS Lett. 2004;577(3):528–534. doi: 10.1016/j.febslet.2004.10.062. [DOI] [PubMed] [Google Scholar]
- 32.Yuan YX, Zhang J, Wang DW, Ling HQ. AtbHLH29 of Arabidopsis thaliana is a functional ortholog of tomato FER involved in controlling iron acquisition in strategy I plants. Cell Res. 2005;15(8):613–621. doi: 10.1038/sj.cr.7290331. [DOI] [PubMed] [Google Scholar]
- 33.Yuan YX, Wu HL, Wang N, Li J, Zhao WN, Du J, et al. FIT interacts with AtbHLH38 and AtbHLH39 in regulating iron uptake gene expression for iron homeostasis in Arabidopsis. Cell Res. 2008;18(3):385–397. doi: 10.1038/cr.2008.26. [DOI] [PubMed] [Google Scholar]
- 34.Long TA, Tsukagoshi H, Busch W, Lahner B, Salt DE, Benfey PN. The bHLH transcription factor POPEYE regulates response to iron deficiency in Arabidopsis roots. Plant Cell. 2010;22(7):2219–2236. doi: 10.1105/tpc.110.074096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Liang G, Zhang HM, Li XL, Ai Q, Yu DQ. bHLH transcription factor bHLH115 regulates iron homeostasis in Arabidopsis thaliana. J Exp Bot. 2017;68(7):1743–1755. doi: 10.1093/jxb/erx043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zhang J, Liu B, Li MS, Feng DR, Jin HL, Wang P, et al. The bHLH transcription factor bHLH104 interacts with IAA-LEUCINE RESISTANT3 and modulates iron homeostasis in Arabidopsis. Plant Cell. 2015;27(3):787–805. doi: 10.1105/tpc.114.132704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wu GA, Terol J, Ibanez V, Lopez-Garcia A, Perez-Roman E, Borreda C, et al. Genomics of the origin and evolution of Citrus. Nature. 2018;554(7692):311–316. doi: 10.1038/nature25447. [DOI] [PubMed] [Google Scholar]
- 38.Wang X, Xu YT, Zhang SQ, Cao L, Huang Y, Cheng JF, et al. Genomic analyses of primitive, wild and cultivated citrus provide insights into asexual reproduction. Nat Genet. 2017;49(5):765–722. doi: 10.1038/ng.3839. [DOI] [PubMed] [Google Scholar]
- 39.Guo P, Qi YP, Yang LT, Lai NW, Ye X, Yang Y, et al. Root adaptive responses to aluminum-treatment revealed by RNA-seq in two citrus species with different aluminum-tolerance. Front Plant Sci. 2017;8:330. doi: 10.3389/fpls.2017.00330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Xu Q, Chen LL, Ruan X, Chen D, Zhu A, Chen C, et al. The draft genome of sweet orange (Citrus sinensis) Nat Genet. 2013;45(1):59–66. doi: 10.1038/ng.2472. [DOI] [PubMed] [Google Scholar]
- 41.Wu GA, Prochnik S, Jenkins J, Salse J, Hellsten U, Murat F, et al. Sequencing of diverse mandarin, pummelo and orange genomes reveals complex history of admixture during citrus domestication. Nat Biotechnol. 2014;32(7):656–662. doi: 10.1038/nbt.2906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Li SB, OuYang WZ, Hou XJ, Xie LL, Hu CG, Zhang JZ. Genome-wide identification, isolation and expression analysis of auxin response factor (ARF) gene family in sweet orange (Citrus sinensis) Front Plant Sci. 2015;6:119. doi: 10.3389/fpls.2015.00119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Magalhaes DM, Scholte LLS, Silva NV, Oliveira GC, Zipfel C, Takita MA, et al. LRR-RLK family from two Citrus species: genome-wide identification and evolutionary aspects. BMC Genomics. 2016;17:623. doi: 10.1186/s12864-016-2930-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Xie RJ, Li YJ, He SL, Zheng YQ, Yi SL, Lv Q, et al. Genome-wide analysis of citrus R2R3MYB genes and their spatiotemporal expression under stresses and hormone treatments. PLoS One. 2014;9(12):e113971. doi: 10.1371/journal.pone.0113971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Geng J, Wei T, Wang Y, Huang X, Liu J-H. Overexpression of PtrbHLH, a basic helix-loop-helix transcription factor from Poncirus trifoliata, confers enhanced cold tolerance in pummelo (Citrus grandis) by modulation of H2O2 level via regulating a CAT gene. Tree Physiol. 2019. 10.1093/treephys/tpz081. [DOI] [PubMed]
- 46.Sun H, Fan HJ, Ling HQ. Genome-wide identification and characterization of the bHLH gene family in tomato. BMC Genomics. 2015;16:9. doi: 10.1186/s12864-014-1209-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Gao M, Zhu YX, Yang JH, Zhang HJ, Cheng CX, Zhang YC, et al. Identification of the grape basic helix-loop-helix transcription factor family and characterization of expression patterns in response to different stresses. Plant Growth Regul. 2019;88(1):19–39. doi: 10.1007/s10725-019-00485-3. [DOI] [Google Scholar]
- 48.Cannon SB, Mitra A, Baumgarten A, Young ND, May G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004;4(1):10. doi: 10.1186/1471-2229-4-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Zhang TT, Lv W, Zhang HS, Ma L, Li PH, Ge L, et al. Genome-wide analysis of the basic Helix-loop-Helix (bHLH) transcription factor family in maize. BMC Plant Biol. 2018;18:235. doi: 10.1186/s12870-018-1441-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Brownlie P, Ceska TA, Lamers M, Romier C, Stier G, Teo H, et al. The crystal structure of an intact human max-DNA complex: new insights into mechanisms of transcriptional control. Structure. 1997;5(4):509–520. doi: 10.1016/S0969-2126(97)00207-4. [DOI] [PubMed] [Google Scholar]
- 51.Geng JJ, Liu JH. The transcription factor CsbHLH18 of sweet orange functions in modulation of cold tolerance and homeostasis of reactive oxygen species by regulating the antioxidant gene. J Exp Bot. 2018;69(10):2677–2692. doi: 10.1093/jxb/ery065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Endo T, Fujii H, Sugiyama A, Nakano M, Nakajima N, Ikoma Y, et al. Overexpression of a citrus basic helix-loop-helix transcription factor (CubHLH1), which is homologous to Arabidopsis activation-tagged bri1 suppressor 1 interacting factor genes, modulates carotenoid metabolism in transgenic tomato. Plant Sci. 2016;243:35–48. doi: 10.1016/j.plantsci.2015.11.005. [DOI] [PubMed] [Google Scholar]
- 53.Kurbidaeva A, Ezhova T, Novokreshchenova M. Arabidopsis thaliana ICE2 gene: phylogeny, structural evolution and functional diversification from ICE1. Plant Sci. 2014;229:10–22. doi: 10.1016/j.plantsci.2014.08.011. [DOI] [PubMed] [Google Scholar]
- 54.Poirier BC, Feldman MJ, Lange BM. bHLH093/NFL and bHLH061 are required for apical meristem function in Arabidopsis thaliana. Plant Signal Behav. 2018;13(7):e1486146. doi: 10.1080/15592324.2018.1486146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Xu J, Ding ZW, Vizcay-Barrena G, Shi JX, Liang WQ, Yuan Z, et al. ABORTED MICROSPORES acts as a master regulator of pollen wall formation in Arabidopsis. Plant Cell. 2014;26(4):1544–1556. doi: 10.1105/tpc.114.122986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Ohta M, Sato A, Renhu N, Yamamoto T, Oka N, Zhu JK, et al. MYC-type transcription factors, MYC67 and MYC70, interact with ICE1 and negatively regulate cold tolerance in Arabidopsis. Sci Rep-Uk. 2018;8:11622. doi: 10.1038/s41598-018-29722-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Raissig MT, Matos JL, Gil MXA, Kornfeld A, Bettadapur A, Abrash E, et al. Mobile MUTE specifies subsidiary cells to build physiologically improved grass stomata. Science. 2017;355(6330):1215–1218. doi: 10.1126/science.aal3254. [DOI] [PubMed] [Google Scholar]
- 58.Zhu EG, You CJ, Wang SS, Cui J, Niu BX, Wang YX, et al. The DYT1-interacting proteins bHLH010, bHLH089 and bHLH091 are redundantly required for Arabidopsis anther development and transcriptome. Plant J. 2015;83(6):976–990. doi: 10.1111/tpj.12942. [DOI] [PubMed] [Google Scholar]
- 59.Song SS, Qi TC, Fan M, Zhang X, Gao H, Huang H, et al. The bHLH subgroup IIId factors negatively regulate jasmonate-mediated plant defense and development. PLoS Genet. 2013;9(7):e1003653. doi: 10.1371/journal.pgen.1003653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Chen C, Chen H, He Y, Xia R. TBtools, a Toolkit for Biologists integrating various HTS-data handling tools with a user-friendly interface. bioRxiv. 2018:289660. 10.1101/289660.
- 61.Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 2009;37:W202–W208. doi: 10.1093/nar/gkp335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Wang YP, Tang HB, DeBarry JD, Tan X, Li JP, Wang XY, et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012;40(7):e49. doi: 10.1093/nar/gkr1293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D607–D613. doi: 10.1093/nar/gky1131. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: Figure S1. Sequence alignment of 128 CgbHLH proteins.
Additional file 2: Table S1. Predicted MW, pI, exon number, UTR and motifs of 128 CgbHLH proteins.
Additional file 3: Table S2. The primers of CgbHLH genes used for qRT-PCR.
Data Availability Statement
All data generated during this study are included in this published article and its supplementary information files. In addition, the protein and CDS sequences of CgbHLHs analysed in this study were retrieved from the Orange Genome Annotation Project, http://citrus.hzau.edu.cn/orange/download/index.php. The protein sequences of AtbHLHs were retrieved from the TAIR database, https://www.arabidopsis.org/browse/genefamily/bHLH.jsp.