Abstract
Several artisanal cheeses are elaborated in European countries, being commonly curdled with rennets of animal origin. However, in some Spanish regions some cheeses of type “Torta” are elaborated using Cynara cardunculus L. rennets. Two of these cheeses, “Torta del Casar” and “Torta de Trujillo”, are elaborated in Cáceres province with ewe’s raw milk and matured over at least 60 days without starters. In this work, we identified the lactic acid bacteria present in these cheeses using MALDI-TOF MS and pheS gene analyses, which showed they belong to the species Lactobacillus curvatus, Lactobacillus diolivorans, Lactobacillus paracasei, Lactobacillus plantarum, Lactobacillus rhamnosus, Lactococcus lactis and Leuconostoc mesenteroides. The pheS gene analysis also allowed the identification of the subspecies La. plantarum subsp. plantarum, La. paracasei subsp. paracasei and Le. mesenteroides subsp. jonggajibkimchii. Low similarity values were found in this gene for some currently accepted subspecies of Lc. lactis and for the two subspecies of La. plantarum, and values near to 100% for the subspecies of Le. mesenteroides and La. paracasei. These results, which were confirmed by the calculated ANIb and dDDH values of their whole genomes, showed the need to revise the taxonomic status of these species and their subspecies.
Keywords: lactic acid bacteria, cheese, “Torta” type, MALDI-TOF MS, pheS gene, Spain
1. Introduction
Lactic acid bacteria (LAB) encompass Gram positive cocci and rods distributed in different genera, species and subspecies belonging to different families from the order Lactobacillales [1]. Many of these bacteria are considered probiotics due to their beneficial effects for human health [2] and they are present in fermented foods [3].
Cheeses, including artisanal ones, are commonly curdled with rennet of animal origin, however, the Spanish agronomic writer Columela (4–70 AD) mentioned in his book entitled De Re Rustica that cheese can be curdled with the thistle flowers. This practice is currently maintained in some Spanish regions, where the cheeses of type “Torta” are elaborated using Cynara cardunculus L. rennets. The best known of these cheeses is the “Torta del Casar” elaborated in Cáceres province with ewe’s raw milk and matured over at least 60 days without starters.
The LAB present in “Torta del Casar” cheese were initially identified using phenotypic traits [4], and more recently through the analysis of the 16S rRNA gene sequences [5], which was the methodology also used for the identification of these bacteria in other European artisanal cheeses [6,7,8,9,10,11].
However, the 16S rRNA gene has limitations in differentiating among closely related species and subspecies of LAB needing additional techniques, such as the sequencing of protein-coding genes or MALDI-TOF MS [12]. The latter technique has been used to identify the LAB from a French artisanal cheese, showing the presence of species such as La. plantarum and La. paracasei, which encompass several subspecies [13].
The usefulness of MALDI-TOF MS to differentiate some subspecies of La. paracasei, La. plantarum and Lc. lactis has been shown in some works [14,15,16], but the identification at subspecies level should be assessed by the sequencing of protein-coding genes, which have a higher discriminating power than the 16S rRNA gene among closely related taxa. In the case of LAB, the pheS gene has been used, combined with MALDI-TOF MS, for their identification in some fermented foods [17,18], but, to date, these two techniques have not been used together to identify LAB in cheese samples.
Therefore, the first aim of this work was to identify the LAB isolated from two cheeses of type “Torta” elaborated in two different sites (Casar and Trujillo) in Cáceres province in Spain through MALDI-TOF MS and pheS gene analyses. The second aim was to analyse the results obtained with these two techniques compared to those of whole-genome analysis for the differentiation of the subspecies currently accepted within several species of LAB.
2. Materials and Methods
2.1. Strains Isolation
The strains were isolated from ripened cheeses type “Torta” named “Torta del Casar” (Doña Engracia Torta del Casar, Casar de Cáceres, Spain) and “Torta de Trujillo” (or “Retorta de Trujillo”) (Quesería Finca Pascualete, Trujillo, Spain), both elaborated in Cáceres province. For strains’ isolation, we followed the methodology described by Ordiales et al. [5] using MRS agar (Sigma Co., St. Louis, MO., USA) for strain isolation. The inoculated plates were incubated at 20 °C for 48h.
2.2. MALDI-TOF MS Performing and Data Analysis
The sample preparation and the MALDI-TOF MS analysis were carried out as was previously published [19] using a matrix of saturated solution of α-HCCA (Bruker Daltonics, Bremen, Germany) in 50% acetonitrile and 2.5% trifluoracetic acid. We used amounts of biomass between 5 and 100 mg to obtain the spectra as indicated by the manufacturer. The calibration masses were the Bruker Bacterial Test Standards (BTS), which were as follows (masses as averages): RL36, 4365.3 Da; RS22, 5096.8 Da; RL34, 5381.4 Da; RL33meth, 6255.4 Da; RL29, 7274.5 Da; RS19, 10,300.1 Da; RNase A, 13,683.2 Da and myoglobin, 16,952.3 Da.
The score values proposed by the manufacturer are the following: a score value between 2.3 and 3.00 indicates highly probable species identification; a score value between 2.0 and 2.299 indicates secure genus identification and probable species identification, a score value between 1.7 and 1.999 indicates probable genus identification, and a score value <1.7 indicates no reliable identification.
Cluster analysis was performed based on a comparison of strain-specific main spectra, created as described above. The dendrogram was constructed by the statistical toolbox of Matlab 7.1 (MathWorks Inc., Natick, MA, USA) integrated in the MALDI Biotyper 3.0 software. The parameter settings were: ‘Distance Measure=Euclidean’ and ‘Linkage=Complete’. The linkage function is normalized according to the distance between 0 (perfect match) and 1000 (no match).
2.3. Phylogenetic Analysis of pheS Gene
The amplification and sequencing of pheS gene was carried out as indicated by Doan et al. [17] using the primers pheS-21-F (5’-CAYCCNGCHCGYGAYATGC-3’) and pheS-23-R (5’-GGRTGRACCATVCCNGCHCC-3’). The sequences obtained were compared with those from the GenBank using the BLASTN program [20]. The obtained sequences and those of related bacteria retrieved from GenBank were aligned using the Clustal W program [21]. The phylogenetic distances were calculated according to Kimura´s two-parameter model [22]. The phylogenetic trees were inferred using the neighbour joining model [23] and MEGA 7.09 [24] was used for all the phylogenetic analyses.
2.4. Genome Analysis of the Subspecies from the Species Identified in this Study
The Average nucleotide identity blast (ANIb) and Digital DNA–DNA hybridization (dDDH) was calculated using the JSpecies service [25] (http://imedea.uib-csic.es/jspecies/) and dDDH values were calculated using the genome-to-genome distance calculator website service from DSMZ (GGDC 2.1) [26] (http://ggdc.dsmz.de/ggdc.php/). These values were calculated using the formula two at the GGDC website because it is the only function appropriate to analyse draft genomes [27].
3. Results
3.1. MALDI-TOF MS Analysis
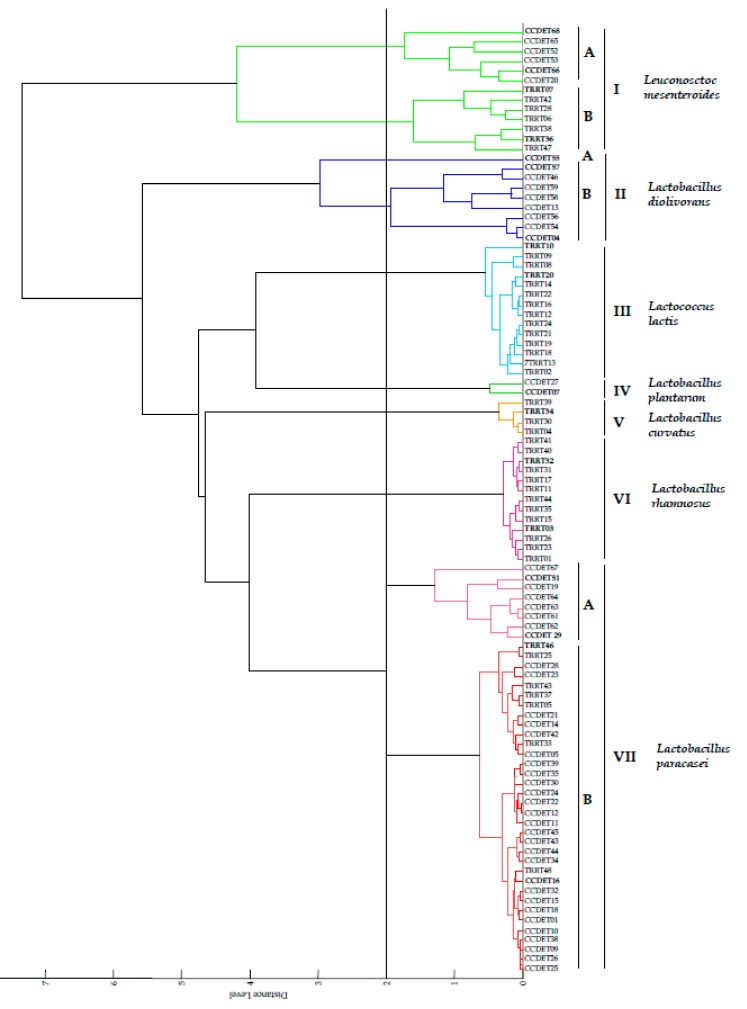
The results of this analysis showed that the isolated strains belong to different genera and species of LAB, namely La. curvatus, La. diolivorans, La. paracasei, La. plantarum, La. rhamnosus, Le. mesenteroides and Lc. lactis. All our strains matched with score values near or higher than 2.0 with strains of these species available in the Biotyper 3.0 database (Table 1). Nevertheless, in most cases, the first matching strain is not the strain type of the identified species, and therefore the identification must be confirmed by gene analysis. In order to select representative strains for this analysis, we grouped the isolated strains through mathematical analysis of their, and the resulting dendrogram is shown in Figure 1.
Table 1.
Results obtained using MALDI-TOF MS analysis.
| Torta del Casar | |||
|---|---|---|---|
| Strains | Closest Taxa | Score Values | Groups |
| CCDET 01 | La. paracasei subsp. paracasei DSM 20006 | 2.502 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.194 | ||
| La. paracasei subsp. tolerans DSM 20258T | 1.960 | ||
| CCDET 04 | La. diolivorans DSM 14421T | 2.228 | IIB |
| CCDET 05 | La. paracasei subsp. paracasei DSM 20006 | 2.504 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.193 | ||
| La. paracasei subsp. tolerans DSM 20258T | 2.174 | ||
| CCDET 07 | La. plantarum DSM 2601 | 2.478 | IV |
| La. plantarum subsp. argentoratensis DSM 16365T | 2.322 | ||
| La. plantarum subsp. plantarum DSM 20174T | 2.037 | ||
| CCDET 09 | La. paracasei subsp. paracasei DSM 20006 | 2.511 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.128 | ||
| La. paracasei subsp. tolerans DSM 20258T | 1.476 | ||
| CCDET 10 | La. paracasei subsp. paracasei DSM 20006 | 2.483 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.097 | ||
| La. paracasei subsp. tolerans DSM 20258T | 2.051 | ||
| CCDET 11 | La. paracasei subsp. paracasei DSM 20244 | 2.517 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.063 | ||
| La. paracasei subsp. tolerans DSM 20258T | 1.911 | ||
| CCDET 12 | La. paracasei subsp. paracasei DSM 20006 | 2.433 | VIIB |
| La. paracasei subsp. tolerans DSM 20258T | 2.113 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.018 | ||
| CCDET 13 | La. diolivorans DSM 14421T | 2.218 | IIB |
| CCDET 14 | La. paracasei subsp. paracasei DSM 2649 | 2.43 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.047 | ||
| La. paracasei subsp. tolerans DSM 20258T | 1.773 | ||
| CCDET 15 | La. paracasei subsp. paracasei DSM 20006 | 2.531 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.053 | ||
| La. paracasei subsp. tolerans DSM 20258T | 1.542 | ||
| CCDET 16 | La. paracasei subsp. paracasei DSM 20006 | 2.545 | VIIB |
| La. paracasei subsp. tolerans DSM 20258T | 2.147 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.112 | ||
| CCDET 18 | La. paracasei subsp. paracasei DSM 20006 | 2.463 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.107 | ||
| La. paracasei subsp. tolerans DSM 20258T | 1.911 | ||
| CCDET19 | La. paracasei subsp. paracasei DSM 20244 | 2.500 | VIIA |
| La. paracasei subsp. tolerans DSM 20258T | 2.309 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.103 | ||
| CCDET20 | Le. mesenteroides subsp. dextranicum DSM 20187 | 2.062 | IA |
| Le. mesenteroides subsp. mesenteroides DSM 20343T | 1.683 | ||
| Le. mesenteroides subsp. cremoris DSM 20346T | 1.648 | ||
| CCDET21 | La. paracasei subsp. paracasei DSM 20006 | 2.337 | VIIB |
| La. paracasei subsp. tolerans DSM 20258T | 2.174 | ||
| La. paracasei subsp. paracasei DSM 5622T | 1.952 | ||
| CCDET 22 | La. paracasei subsp. paracasei DSM 20006 | 2.380 | VIIB |
| La. paracasei subsp. paracasei DSM 5622TLa. paracasei subsp. tolerans DSM 20258T | 2.0401.752 | ||
| CCDET 23 | La. paracasei subsp. paracasei DSM 20244 | 2.397 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 1.938 | ||
| La. paracasei subsp. tolerans DSM 20258T | 1.800 | ||
| CCDET 24 | La. paracasei subsp. paracasei DSM 20312 | 2.355 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.038 | ||
| La. paracasei subsp. tolerans DSM 20258T | 1.897 | ||
| CCDET 25 | La. paracasei subsp. paracasei DSM 20006 | 2.544 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.115 | ||
| La. paracasei subsp. tolerans DSM 20258T | 2.092 | ||
| CCDET 26 | La. paracasei subsp. paracasei DSM 20006 | 2.476 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.165 | ||
| La. paracasei subsp. tolerans DSM 20258T | 2.033 | ||
| CCDET 27 | La. plantarum subsp. plantarum DSM 12028 | 2.177 | IV |
| La. plantarum subsp. argentoratensis DSM 16365T | 2.131 | ||
| La. plantarum subsp. plantarum DSM 20174T | 1.963 | ||
| CCDET 28 | La. paracasei subsp. paracasei DSM 20244 | 2.386 | VIIB |
| La. paracasei subsp. tolerans DSM 20258T | 2.097 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.097 | ||
| CCDET 29 | La. paracasei subsp. paracasei DSM 46331 | 2.432 | VIIA |
| La. paracasei subsp. tolerans DSM 20258T | 2.224 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.072 | ||
| CCDET 30 | La. paracasei subsp. paracasei DSM 20006 | 2.492 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.157 | ||
| La. paracasei subsp. tolerans DSM 20258T | 2.003 | ||
| CCDET 32 | La. paracasei subsp. paracasei DSM 20006 | 2.513 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.113 | ||
| La. paracasei subsp. tolerans DSM 20258T | 1.540 | ||
| CCDET 34 | La. paracasei subsp. paracasei DSM 20244 | 2.444 | VIIB |
| La. paracasei subsp. tolerans DSM 20258T | 2.160 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.120 | ||
| CCDET 35 | La. paracasei subsp. paracasei DSM 20006 | 2.475 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.080 | ||
| La. paracasei subsp. tolerans DSM 20258T | 1.962 | ||
| CCDET 38 | La. paracasei subsp. paracasei DSM 20006 | 2.452 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.112 | ||
| La. paracasei subsp. tolerans DSM 20258T | 2.083 | ||
| CCDET 39 | La. paracasei subsp. paracasei DSM 20006 | 2.494 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.059 | ||
| La. paracasei subsp. tolerans DSM 20258T | 1.847 | ||
| CCDET 42 | La. paracasei subsp. paracasei DSM 20006 | 2.223 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 1.871 | ||
| La. paracasei subsp. tolerans DSM 20258T | 1.854 | ||
| CCDET 43 | La. paracasei subsp. paracasei DSM 20244 | 2.348 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.035 | ||
| La. paracasei subsp. tolerans DSM 20258T | 1.990 | ||
| CCDET 44 | La. paracasei subsp. paracasei DSM 20244 | 2.339 | VIIB |
| La. paracasei subsp. tolerans DSM 20258T | 2.106 | ||
| La. paracasei subsp. paracasei DSM 5622T | 1.998 | ||
| CCDET 45 | La. paracasei subsp. paracasei DSM 20244 | 2.353 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.061 | ||
| La. paracasei subsp. tolerans DSM 20258T | 2.053 | ||
| CCDET 46 | La. diolivorans DSM 14421T | 2.235 | IIB |
| CCDET51 | La. paracasei subsp. paracasei DSM 20244 | 2.437 | VIIA |
| La. paracasei subsp. tolerans DSM 20258T | 2.054 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.054 | ||
| CCDET52 | Le. mesenteroides subsp. dextranicum DSM 20187 | 2.000 | IA |
| Le. mesenteroides subsp. cremoris DSM 20346T | 1.690 | ||
| Le. mesenteroides subsp. mesenteroides DSM 20343T | 1.374 | ||
| CCDET53 | Le. mesenteroides subsp. mesenteroides DSM 20241 | 2.120 | IA |
| Le. mesenteroides subsp. mesenteroides DSM 20343T | 2.106 | ||
| Le. mesenteroides subsp. cremoris DSM 20346T | 1.961 | ||
| CCDET 54 | La. diolivorans DSM 14421T | 2.003 | IIB |
| CCDET 55 | La. diolivorans DSM 14421T | 1.911 | IIB |
| CCDET 56 | La. diolivorans DSM 14421T | 2.093 | IIB |
| CCDET 57 | La. diolivorans DSM 14421T | 2.100 | IIB |
| CCDET 58 | La. diolivorans DSM 14421T | 2.149 | IIB |
| CCDET 59 | La. diolivorans DSM 14421T | 2.106 | IIB |
| CCDET 61 | La. paracasei subsp. paracasei DSM 20006 | 2.353 | VIIA |
| La. paracasei subsp. paracasei DSM 5622T | 2.170 | ||
| La. paracasei subsp. tolerans DSM 20258T | 2.101 | ||
| CCDET62 | La. paracasei subsp. paracasei DSM 20244 | 2.536 | VIIA |
| La. paracasei subsp. tolerans DSM 20258T | 2.157 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.148 | ||
| CCDET 63 | La. paracasei subsp. paracasei DSM 8741 | 2.383 | VIIA |
| La. paracasei subsp. paracasei DSM 5622T | 2.100 | ||
| La. paracasei subsp. tolerans DSM 20258T | 2.082 | ||
| CCDET64 | La. paracasei subsp. paracasei DSM 20244 | 2.493 | VIIA |
| La. paracasei subsp. tolerans DSM 20258T | 2.149 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.092 | ||
| CCDET65 | Le. mesenteroides subsp. dextranicum DSM 20187 | 2.072 | IA |
| Le. mesenteroides subsp. cremoris DSM 20346T | 1.692 | ||
| Le. mesenteroides subsp. mesenteroides DSM 20343T | 1.454 | ||
| CCDET66 | Le. mesenteroides subsp. dextranicum DSM 20187 | 2.071 | IA |
| Le. mesenteroides subsp. cremoris DSM 20346T | 1.633 | ||
| Le. mesenteroides subsp. mesenteroides DSM 20343T | 1.355 | ||
| CCDET67 | La. paracasei subsp. paracasei DSM 20244 | 2.468 | VIIA |
| La. paracasei subsp. tolerans DSM 20258T | 2.233 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.047 | ||
| CCDET68 | Le. mesenteroides subsp. mesenteroides DSM 20343T | 2.204 | IA |
| Le. mesenteroides subsp. dextranicum DSM 20187 | 2.089 | ||
| Le. mesenteroides subsp. cremoris DSM 20346T | 1.953 | ||
| Torta de Trujillo | |||
| Strains | Closest taxa | Score values | Groups |
| TRRT01 | La. rhamnosus CIP A157T | 2.362 | VI |
| TRRT02 | Lc. lactis subsp. lactis DSM 20661 | 2.433 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.149 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 1.913 | ||
| TRRT03 | La. rhamnosus CIP A157T | 2.389 | VI |
| TRRT04 | La. curvatus DSM 20499 | 2.430 | V |
| La. curvatus DSM 20019T | 2.007 | ||
| TRRT05 | La. paracasei subsp. paracasei DSM 20006 | 2.393 | VIIB |
| La. paracasei subsp. tolerans DSM 20258T | 2.193 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.109 | ||
| TRRT06 | Le. mesenteroides subsp. mesenteroides DSM 20343T | 2.368 | IB |
| Le. mesenteroides subsp. dextranicum DSM 20187 | 2.097 | ||
| Le. mesenteroides subsp. cremoris DSM 20346T | 1.963 | ||
| TRRT07 | Le. mesenteroides subsp. mesenteroides DSM 20343T | 2.380 | IB |
| Le. mesenteroides subsp. cremoris DSM 20346T | 2.221 | ||
| Le. mesenteroides subsp. dextranicum DSM 20187 | 2.026 | ||
| TRRT08 | Lc. lactis subsp. lactis DSM 20661 | 2.373 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.209 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 1.848 | ||
| TRRT09 | Lc. lactis subsp. lactis DSM 20661 | 2.283 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.214 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 1.896 | ||
| TRRT10 | Lc. lactis subsp. lactis DSM 20661 | 2.236 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.198 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 1.771 | ||
| TRRT11 | La. rhamnosus CIP A157T | 2.366 | VI |
| TRRT12 | Lc. lactis subsp. lactis DSM 20661 | 2.310 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.150 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 1.983 | ||
| TRRT13 | Lc. lactis subsp. lactis DSM 20661 | 2.392 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.255 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 1.988 | ||
| TRRT14 | Lc. lactis subsp. lactis DSM 20661 | 2.371 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.223 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 1.901 | ||
| TRRT15 | La. rhamnosus CIP A157T | 2.324 | VI |
| TRRT16 | Lc. lactis subsp. lactis DSM 20661 | 2.456 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.228 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 1.868 | ||
| TRRT17 | La. rhamnosus CIP A157T | 2.360 | VI |
| TRRT18 | Lc. lactis subsp. lactis DSM 20661 | 2.521 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.215 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 1.998 | ||
| TRRT19 | Lc. lactis subsp. lactis DSM 20661 | 2.514 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.196 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 1.927 | ||
| TRRT20 | Lc. lactis subsp. lactis DSM 20661 | 2.461 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.157 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 1.992 | ||
| TRRT21 | Lc. lactis subsp. lactis DSM 20661 | 2.538 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.226 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 2.045 | ||
| TRRT22 | Lc. lactis subsp. lactis DSM 20661 | 2.345 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.286 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 1.955 | ||
| TRRT23 | La. rhamnosus CIP A157T | 2.426 | VI |
| TRRT24 | Lc. lactis subsp. lactis DSM 20661 | 2.468 | III |
| Lc. lactis subsp. lactis DSM 20481T | 2.243 | ||
| Lc. lactis subsp. cremoris DSM 20069T | 2.036 | ||
| TRRT25 | La. paracasei subsp. paracasei DSM 20006 | 2.425 | VIIB |
| La. paracasei subsp. tolerans DSM 20258T | 2.181 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.144 | ||
| TRRT26 | La. rhamnosus CIP A157T | 2.357 | VI |
| TRRT28 | Le. mesenteroides subsp. mesenteroides DSM 20343T | 2.358 | IB |
| Le. mesenteroides subsp. dextranicum DSM 20187 | 2.090 | ||
| Le. mesenteroides subsp. cremoris DSM 20346T | 2.044 | ||
| TRRT30 | La. curvatus DSM 20499 | 2.340 | V |
| La. curvatus DSM 20019T | 2.116 | ||
| TRRT31 | La. rhamnosus CIP A157T | 2.433 | VI |
| TRRT32 | La. rhamnosus CIP A157T | 2.350 | VI |
| TRRT33 | La. paracasei subsp. paracasei DSM 20006 | 2.435 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.136 | ||
| La. paracasei subsp. tolerans DSM 20258T | 2.103 | ||
| TRRT34 | La. curvatus DSM 20499 | 2.405 | V |
| La. curvatus DSM 20019T | 2.166 | ||
| TRRT35 | La. rhamnosus CIP A157T | 2.367 | VI |
| TRRT36 | Le. mesenteroides subsp. mesenteroides DSM 20343T | 2.389 | IB |
| Le. mesenteroides subsp. dextranicum DSM 20187 | 2.035 | ||
| Le. mesenteroides subsp. cremoris DSM 20346T | 1.985 | ||
| TRRT37 | La. paracasei subsp. paracasei DSM 20006 | 2.448 | VIIB |
| La. paracasei subsp. tolerans DSM 20258T | 2.114 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.000 | ||
| TRRT38 | Le. mesenteroides subsp. mesenteroides DSM 20343T | 2.359 | IB |
| Le. mesenteroides subsp. dextranicum DSM 20187 | 2.131 | ||
| Le. mesenteroides subsp. cremoris DSM 20346T | 1.908 | ||
| TRRT39 | La. curvatus DSM 20499 | 2.234 | V |
| La. curvatus DSM 20019T | 2.118 | ||
| TRRT40 | La. rhamnosus CIP A157T | 2.361 | VI |
| TRRT41 | La. rhamnosus CIP A157T | 2.354 | VI |
| TRRT42 | Le. mesenteroides subsp. mesenteroides DSM 20343T | 2.308 | IB |
| Le. mesenteroides subsp. dextranicum DSM 20187 | 2.004 | ||
| Le. mesenteroides subsp. cremoris DSM 20346T | 1.982 | ||
| TRRT43 | La. paracasei subsp. paracasei DSM 20006 | 2.362 | VIIB |
| La. paracasei subsp. paracasei DSM 5622T | 2.234 | ||
| La. paracasei subsp. tolerans DSM 20258T | 2.182 | ||
| TRRT44 | La. rhamnosus CIP A157T | 2.405 | VI |
| TRRT46 | La. paracasei subsp. paracasei DSM 20006 | 2.459 | VIIB |
| La. paracasei subsp. tolerans DSM 20258T | 2.221 | ||
| La. paracasei subsp. paracasei DSM 5622T | 2.000 | ||
| TRRT47 | Le. mesenteroides subsp. mesenteroides DSM 20343T | 2.322 | IB |
| Le. mesenteroides subsp. dextranicum DSM 20187 | 2.041 | ||
| Le. mesenteroides subsp. cremoris DSM 20346T | 1.864 | ||
| TRRT48 | La. paracasei subsp. paracasei DSM 20006 | 2.242 | VIIB |
| La. paracasei subsp. tolerans DSM 20258T | 1.983 | ||
| La. paracasei subsp. paracasei DSM 5622T | 1.658 | ||
Figure 1.
Cluster analysis of MALDI-TOF MS spectra of strains isolated in this study. Distance is displayed in relative units. Representative strains of each group selected for pheS gene analysis are marked in bold.
The strains were distributed into seven groups with similarity values lower than 2, which correspond to the different species identified in this study (Figure 1). Group I encompasses strains that matched with score values higher than 2.0 with Le. mesenteroides strains and was divided into two subgroups. The strains from the subgroup IA matched with the type strains of Le. mesenteroides subsp. mesenteroides DSM 20343T and Le. mesenteroides subsp. cremoris DSM 20346T and with the non-type strain of Le. mesenteroides subsp. dextranicum DSM 20187 with score values lower than 2.3, whereas those from the subgroup IB matched with the type strain of Le. mesenteroides subsp. mesenteroides DSM 20343T with score values higher than 2.3 (Table 1).
Group II encompasses strains that matched with the type strain of La. diolivorans DSM 14421T and comprised the independent branch IIA and the subgroup IIB (Figure 1). The strain CCDET 55 formed an independent branch and matched with the type strain of La. diolivorans DSM 14421T with a score value lower than 2.0, whereas the strains from subgroup IIB matched with score values higher than 2.0 and lower than 2.3 with the same type strain (Table 1).
Group III encompasses strains that matched with score values higher than 2.0 with Lc. lactis strains (Figure 1). All strains isolated in this study matched with the non-type strain Lc. lactis subsp. lactis DSM 20661, with score values near to or higher than 2.3 with Lc. lactis subsp. lactis DSM 20481T with score values lower than 2.3 and with Lc. lactis subsp. cremoris DSM 20069T with score values lower than 2.0 in most of cases (Table 1).
Group IV encompasses two strains that matched with score values higher than 2.0 with La. plantarum strains (Figure 1). The strain CCDET07 matched with score values higher than 2.3 with the non type strain La. plantarum DSM 2601 and with the type strain of La. plantarum subsp. argentoratensis DSM 16365T, whereas these values were lower than 2.3 with respect to the type strain of La. plantarum subsp. plantarum DSM 20174T. The strain CCDET27 matched with score values higher than 2.0 with respect to the non-type strain La. plantarum DSM 12028 and with the type strain of La. plantarum subsp. argentoratensis DSM 16365T, whereas these values were lower than 2.0 with respect to the type strain of La. plantarum subsp. plantarum DSM 20174T (Table 1).
Group V encompasses strains matching with score values higher than 2.0 with La. curvatus strains (Figure 1). The higher score values, near or higher than 2.3, were found with respect to the non-type strain DSM 20499, whereas these values were lower than 2.3 with respect to the type strain of La. curvatus DSM 20499T (Table 1).
Group VI encompasses strains that matched with the type strain of La. rhamnosus CIP A157T with score values higher than 2.3 in all cases (Figure 1, Table 1).
Finally, group VII encompasses strains that matched with score values higher than 2.0 with La. paracasei strains (Figure 1). This group was divided into two subgroups whose strains mostly matched with score values higher than 2.3 with different non-type strains of La. paracasei subsp. paracasei (DSM 20006, DSM 20244, DSM 2649, DSM 20312 or DSM 8741). Only the strain CCDET19 matched with values higher than 2.3 with respect to the type strain of La. paracasei subsp. tolerans DSM 20258T and the remaining strains matched with La. paracasei subsp. paracasei DSM 5622T and/or La. paracasei subsp. tolerans DSM 20258T with score values lower, near or higher than 2.0, but in all cases lower than 2.3 (Table 1).
Since the type strains of several subspecies identified in this study are included in the Biotyper 3.0 database, we calculated the score values between the subspecies from the same species (Table 2). Score values higher than 2.3, typically found in strains from the same species, were presented by the type strains of the subspecies plantarum and argentoratensis of La. plantarum (2.424) and by those of the subspecies mesenteroides and cremoris of Le. mesenteroides (2.456). However, score values lower than 2.3, which can be found in strains of different species, were found between by the type strains of the subspecies lactis and cremoris of Lc. lactis (2.174) and by those of the subspecies paracasei and tolerans of La. paracasei (1.846). These results show the need to carry out genetic analyses to verify the taxonomic status of these subspecies.
Table 2.
Results of the comparison of the type strains of subspecies from different species of LAB identified in this study obtained with different methodologies.
| Strains | Closest Species | Score Values MALDI-TOF | pheS Gene Similarity (%) | ANIb (%) | dDDH (%) |
|---|---|---|---|---|---|
| La. plantarum subsp plantarum ATCC 14917T (DSM 20174T) | La. plantarum subsp. argentoratensis DSM 16365T | 2.424 | 90.5% | 94.9 | 62.9 |
| La. paracasei subsp paracasei DSM 5622T | La. paracasei subsp. tolerans DSM 20258T | 1.846 | 99.5 | 97.9 | 84.9 |
| Le. mesenteroides subsp mesenteroides ATCC 8293T (DSM 20343T) | Le. mesenteroides subsp. cremoris ATCC 19254T (DSM 20346T) | 2.456 | 99.5 | 98.1 | 90.9 |
| Le. mesenteroides subsp mesenteroides ATCC 8293T (DSM 20343T) | Le. mesenteroides subsp. dextranicum DSM 20484T | nd | 99.2 | 98.2 | 91.9 |
| Le. mesenteroides subsp mesenteroides ATCC 8293T (DSM 20343T) | Le. mesenteroides subsp. jonggajibkimchii DRC1506T | nd | 99.7 | 98.4 | 90.1 |
| Le. mesenteroides subsp. cremoris ATCC 19254T (DSM 20346T) | Le. mesenteroides subsp. dextranicum DSM 20484T | nd | 99.7 | 98.5 | 91.5 |
| Le. mesenteroides subsp. cremoris ATCC 19254T (DSM 20346T) | Le. mesenteroides subsp. jonggajibkimchii DRC1506T | nd | 99.7 | 98.1 | 88.5 |
| Le. mesenteroides subsp. dextranicum DSM 20484T | Le. mesenteroides subsp. jonggajibkimchii DRC1506T | nd | 99.5 | 98.4 | 90.1 |
| Lc.lactis subsp lactis ATCC 19435T (DSM 20481T) | Lc. lactis subsp. cremoris NBRC 100676T (DSM 20069T) | 2.174 | 92.2 | 86.7 | 32.7 |
| Lc.lactis subsp lactis ATCC 19435T (DSM 20481T) | Lc. lactis subsp. hordniae CCUG 32210T | nd | 99.2 | 96.7 | 79.9 |
| Lc.lactis subsp lactis ATCC 19435T (DSM 20481T) | Lc. lactis subsp. tructae DSM 21502T | nd | 92.5 | 86.1 | 31.7 |
| Lc. lactis subsp. cremoris NBRC 100676T (DSM 20069T) | Lc. lactis subsp. hordniae CCUG 32210T | nd | 91.5 | 86.0 | 31.4 |
| Lc. lactis subsp. cremoris NBRC 100676T (DSM 20069T) | Lc. lactis subsp. tructae DSM 21502T | nd | 98.5 | 97.5 | 83.6 |
| Lc. lactis subsp. hordniae CCUG 32210T | Lc. lactis subsp. tructae DSM 21502T | nd | 91.8 | 85.9 | 31.6 |
nd: no data because the type strains of some subspecies are not included in the Biotyper 3.0 database.
3.2. pheS Gene Analysis
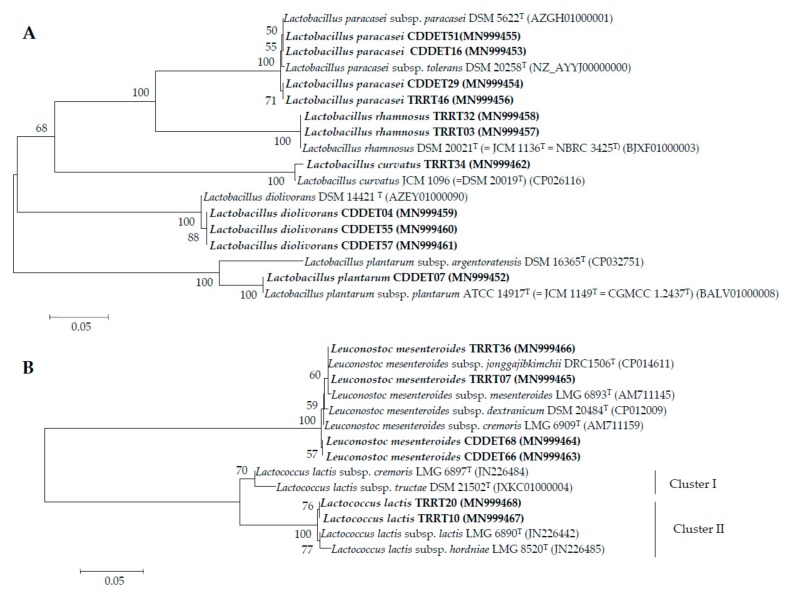
The analysis of partial sequences of pheS gene of representative strains of different MALDI-TOF MS groups are shown in Figure 2 and Table 2 and Table 3. The results of this analysis confirmed the identification obtained after MALDI-TOF MS analysis at genus and species levels for all strains isolated in this study.
Figure 2.
(A) Neighbour-joining phylogenetic unrooted tree based on pheS gene partial sequences (400 nt) showing the taxonomic location of representative strains from different groups of MALDI-TOF MS within the genus Lactobacillus. (B) Neighbour-joining phylogenetic unrooted tree based on pheS gene partial sequences (400 nt) showing the taxonomic location of representative strains from different groups of MALDI-TOF MS within the genera Lactobacillus and Leuconostoc. Bootstrap values calculated for 1000 replications are indicated. Bar, 5 nt substitution per 1000 nt. Accession numbers from Genbank are given in brackets.
Table 3.
Results obtained using MALDI-TOF MS and pheS gene analyses.
| MALDI-TOF MS Group | Number of Strains | Selected Strains | Closest Taxa | Score Values | pheS Gene Similarity (%) |
|---|---|---|---|---|---|
| Group IA | 6 from “Torta del Casar” | CCDET66, | Le. mesenteroides subsp. mesenteroides DSM 20343T | 1.3–2.2 | 99.2 |
| 1 from “Torta de Trujillo” | CCDET68 | Le. mesenteroides subsp. cremoris DSM 20346T | 1.6–2.0 | 99.7 | |
| Group IB * | 6 from “Torta de Trujillo” | TRRT07, | Le. mesenteroides subsp. mesenteroides DSM 20343T | 2.3–2.4 | 99.7 |
| TRRT36 | Le. mesenteroides subsp. cremoris DSM 20346T | 1.8–2.2 | 99.5 | ||
| Branch IIA | 1 from “Torta del Casar” | CCDET55 | La. diolivorans DSM 14421T | 1.9 | 99.3 |
| Group IIB | 9 from “Torta del Casar” | CCDET04, CCDET57 | La. diolivorans DSM 14421T | 1.9–2.2 | 99.3 |
| Group III | 13 from “Torta de Trujillo” | TRRT10, | Lc. lactis subsp. lactis DSM 20481T | 1.9–2.3 | 99.5 |
| TRRT20 | Lc. lactis subsp. cremoris DSM 20069T | 1.7–2.1 | 99.0 | ||
| Group IV | 2 from “Torta del Casar” | CCDET07 | La. plantarum subsp. plantarum DSM 20174T | 2.1–2.3 | 100 |
| La. plantarum subsp. argentoratensis DSM 16365T | 1.9–2.0 | 90.8 | |||
| Group V | 4 from “Torta de Trujillo” | TRRT34 | La. curvatus DSM 20019T | 2.0–2.2 | 99.2 |
| Group VI | 13 from “Torta de Trujillo” | TRRT03, TRRT32 | La. rhamnosus CIP A157T | 2.3–2.4 | 100 |
| VIIA | 8 from “Torta del Casar” | CCDET29 | La. paracasei subsp. paracasei DSM 5622T | 2.0–2.1 | 99.5 |
| La. paracasei subsp. tolerans DSM 20258T | 2.0–2.3 | 99.5 | |||
| VIIA | 8 from “Torta del Casar” | CCDET51 | La. paracasei subsp. paracasei DSM 5622T | 2.0–2.1 | 100 |
| La. paracasei subsp. tolerans DSM 20258T | 2.0–2.3 | 99.7 | |||
| VIIB | 22 from “Torta del Casar” | CCDET16 | La. paracasei subsp. paracasei DSM 5622T | 1.8–2.3 | 100 |
| 7 from “Torta de Trujillo” | |||||
| La. paracasei subsp. tolerans DSM 20258T | 1.9–2.2 | 99.7 | |||
| VIIB | 22 from “Torta del Casar” | TRRT46 | La. paracasei subsp. paracasei DSM 5622T | 1.8-2.3 | 99.5 |
| 7 from “Torta de Trujillo” | La. paracasei subsp. tolerans DSM 20258T | 1.9-2.2 | 99.5 |
* These strains presented 100% similarity with respect to L. mesenteroides subsp. jonggajibkimchii which is not included in Biotyper 3.0.
According to the results of the pheS gene analysis, several strains were identified with high similarity values as Lactobacillus species that, to date, do not encompasses subspecies (Figure 2A, Table 3). The strains TRRT03, TRRT32 representative of group VI, were identified as La. rhamnosus with 100% similarity. The strain CCDET55, representative of subgroup IIA, and the strains CCDET04, CCDET57, representative of subgroup IIB, were identified as La. diolivorans with 99.3% similarity. The strain TRRT34, representative of group V, was identified as La. curvatus with 99.2% similarity.
After the pheS gene analysis, the remaining strains were identified with high similarity values with LABs of species that contain two or more subspecies (Figure 2A, Table 3). This happened in the case of the strain CCDET07, representative of group IV, which was identified as La. plantarum which currently encompasses two subspecies, L. plantarum subsp. plantarum and L. plantarum subsp. argentoratensis, whose type strains showed 90.5% similarity in their pheS gene sequences (Table 2). The strain CCDET07, representative of group II, can be assigned to the subspecies plantarum, since it presented 100% similarity with respect to the type strain of this subspecies and 90.5% similarity with respect to the type strain of the subspecies argentoratensis (Figure 2B, Table 3).
The representative strains from group I were identified with pheS gene similarity values higher than 99.2% with the species Le. mesenteroides, whose subspecies mesenteroides, cremoris, dextranicum and jonggajibkimchii showed values ranging from 99.2% to 99.7% (Figure 2B, Table 2). The strains CCDET66, CCDET68 representative of subgroup IA were slightly more closesly related to the type strain of Le. mesenteroides subsp. cremoris, with 99.7% similarity, than to the type strains of the remaining subspecies, with similarity values ranging from 99.2% to 99.5%. The strains TRRT07, TRRT36, representative of subgroup IB, presented 100% similarity with respect to the type strain of Le. mesenteroides subsp. jonggajibkimchii, and values ranging from 99.5% to 99.7% with respect to the type strains of the other three subspecies. Therefore, the strains from the group IB can be assigned to the subspecies Le. mesenteroides subsp. jonggajibkimchii, whereas it is difficult to assign those of group IA to any of the subspecies from Le. mesenteroides (Figure 2B, Table 3).
The representative strains from group III were identified with pheS gene similarity values higher than 99.0% as Lc. lactis, whose subspecies formed two clearly separated clusters with less than 93% similarity (Figure 2B, Table 2). Cluster I contains the subspecies lactis and hordniae showing 99.2% similarity and cluster II the subspecies cremoris and tructae, showing 98.5% similarity (Table 2). The strains TRRT10, TRRT20, representative of group III, belong to cluster II and, since they presented 99.0% and 99.5% similarity, respectively, to the subspecies lactis and hordniae, it is difficult to assign the strains of group III to any of these two subspecies (Figure 2B, Table 3).
The representative strains from group VII with pheS gene similarity values higher than 99.5% were identified as La. paracasei, which contains two subspecies, paracasei and tolerans, showing 99.5% similarity between their type strains (Figure 2A, Table 2). The representative strains for both subgroups VIIA and VIIB were divided into two subclusters with 99.5% similarity, each one containing strains of these both subgroups (Figure 2A). The strains CCDET51 and CCDET16 can be assigned to the subspecies paracasei since they showed 100% similarity with the type strain of this subspecies, however, the strains CCDET29 and CCDET46 cannot be assigned to these subspecies because they showed 99.5% similarity with respect to their type strains (Figure 2A, Table 3).
Therefore, the identification at species level obtained by MALDI-TOF MS was confirmed by pheS gene sequencing. Moreover, the pheS gene analysis supports the identification at subspecies level for some strains isolated in this work, but it is remarkable that several others cannot be assigned to any subspecies because they formed subclusters whose similarity values are similar to those found among the currently accepted subspecies of LAB identified in this study.
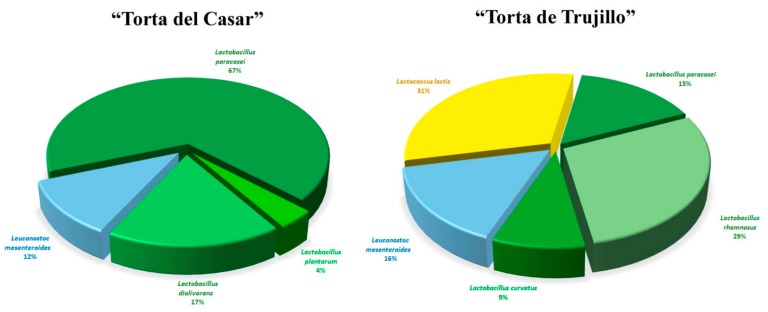
Collectively, the data from MALDI-TOF MS and pheS gene analyses showed that most of the strains isolated from “Torta del Casar” belong to the species La. paracasei, which was also present in “Torta de Trujillo” and that the species Le. mesenteroides was present in both cheeses in similar proportions. However, other species only were found in one of the two cheeses, La. diolivorans and La. plantarum in “Torta del Casar”, and La. curvatus, La. rhamnosus and Lc. lactis in “Torta de Trujillo” (Figure 3, Table 2).
Figure 3.
Pie charts showing the distribution of the different species of LAB in the two cheeses type “Torta” analysed in this study.
3.3. Taxonomic Status of the Subspecies from the Species Identified in this Study
The pheS gene analysis showed that similarity values ranging from 98.5% to 99.7% are presented by the type strains of the subspecies of La. paracasei and Le. mesenteroides, whereas values lower than 93% were found between the type strains of the subspecies of La. plantarum and those of some subspecies of Lc. lactis (Table 2). These results should be compared with those obtained after whole genome analysis, taking into account the threshold values of ANIb and dDDH for bacterial species differentiation (95%~96% and 70%, respectively) [28] and the dDDH cut-off values for bacterial subspecies differentiation (79%~80%) [29].
The whole genomes of the type strains of all subspecies found in this study are available in Genbank and we calculated the ANIb and dDDH values for all of them, whether or not they are present in the Biotyper 3.0 database (Table 2). In agreement with the results of both pheS gene and MALDI-TOF MS analyses, the type strains of the subspecies mesenteroides and cremoris of Le. mesenteroides showed ANIb and dDDH values typical of the same species, 98.1% and 90.9%, respectively (Table 2). Concerning the subspecies dextranicum and jonggajibkimchii, whose type strains are not in Biotyper 3.0 database, in agreement with the results of pheS gene analysis, their ANIb and dDDH values, between them and with respect to the remaining two subspecies, were higher than those proposed for bacterial species differentiation (Table 2).
In agreement with the results of both pheS gene and MALDI-TOF MS analyses, the type strains of the subspecies lactis and cremoris of Lc. lactis showed ANIb and dDDH values typical of different species, 86.7% and 33.1 %, respectively (Table 2). These results confirmed that the type strains of the subspecies cremoris and lactis belong to different species, making it necessary to reclassify the subspecies cremoris into a different, novel species. Nevertheless, it is also necessary to analyse the two subspecies of Lc. lactis that are not present in the Biotyper 3.0 database, as the pheS gene analysis showed that they belong to two divergent clusters, one of them containing the type strains of the subspecies lactis and hordniae and the other containing the type strains of the subspecies cremoris and tructae. Taking into account the ANIb and dDDH values found among the type strains of these subspecies, the subspecies hordniae should be maintained within the species Lc. lactis, and the subspecies tructae, together with the subspecies cremoris, should be transferred to a novel species (Table 2).
In agreement with the results of the pheS gene analysis, but not with those of MALDI-TOF MS analysis, the type strains of the subspecies plantarum and argentoratensis of La. plantarum, which showed ANIb and dDDH values typical of different species, 94.9% and 62.9%, respectively, should be considered different species, making it necessary to reclassify the subspecies argentoratensis in a novel species. In agreement with the results of the pheS gene analysis, but not with those of MALDI-TOF MS analysis, the type strains of the subspecies paracasei and tolerans of La. paracasei showed ANIb and dDDH values typical of the same species, 97.9% and 84.9%, respectively (Table 2).
4. Discussion
There is a growing interest in the identification of LAB present in artisanal cheeses elaborated with raw milk in Europe, with those elaborated with cow or/and goat raw milks being more analysed [6,7,8,10,11,13] compared to those elaborated with ewe’s raw milk [5,9].
In Spain, one of the most appreciated cheeses is the named type “Torta”, elaborated with ewe’s raw milk in Caceres province, and therefore it is also interesting to know the species of LAB present in these cheeses. In a work published in the last century, the lactic bacteria present in the “Torta del Casar” cheese were identified on the basis of phenotypic traits [4]. More recently, by 16S rRNA gene analysis, the species Lactobacillus sakei, Lactobacillus casei, Lactobacillus helveticus and Lc. lactis subsp. cremoris have been identified in this cheese [5]. These two works were only carried out in the “Torta del Casar” cheese and using techniques that have limitations for species and particularly for subspecies differentiation. For this reason, in this study we compared the results obtained in two cheeses type “Torta” elaborated in the same region by using more recent methodologies. From the species previously identified in the “Torta del Casar” cheese [5], in the present work only L. lactis has been identified in the “Torta de Trujillo” cheese. Nevertheless, the species identified in this study have been found in some of the European artisanal cheeses elaborated with raw milk [7,8,9,10,11,12,13].
From the mentioned cheeses, only the LAB present in the French cheese Maroilles were identified by MALDI-TOF MS [13]. The authors showed that this methodology is very useful to identify LAB belonging to different genera and species, but they do not demonstrate its usefulness in differentiating among subspecies. Considering that many species of LAB contain several subspecies, this is an essential issue to be discussed by comparison with other molecular techniques, particularly genomic ones.
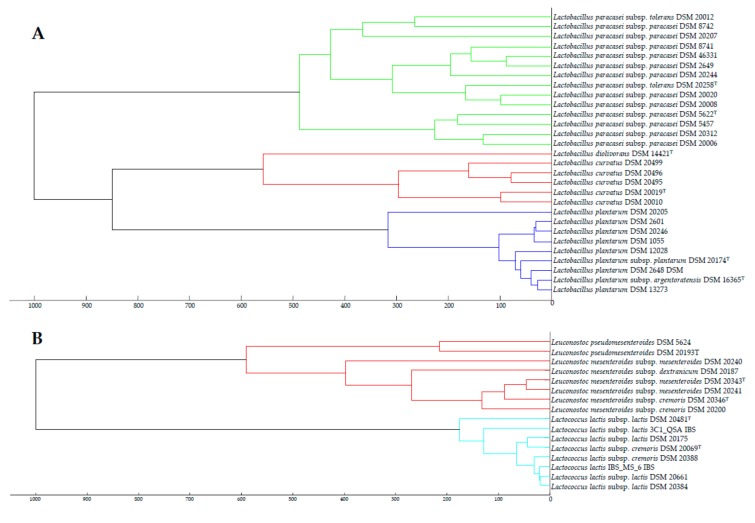
In this study, we identified four species which contain several subspecies, La. plantarum, La. paracasei, Le. mesenteroides and Lc. lactis (http://www.bacterio.net/) of which the most common inhabitants in milk-related sources are present in the database Biotyper 3.0. In addition to the type strain, several strains are included in this database for Le. mesenteroides subsp. mesenteroides, Lc. lactis subsp. lactis, Lc. lactis subsp. cremoris, La. paracasei subsp. paracasei, La. paracasei subsp. tolerans and La. plantarum subsp. plantarum. The presence of most than one strain for a taxon in a database is a priori positive, but this can be an important disadvantage if some strains are not correctly assigned to a taxon, as seems to occur for several strains from the subspecies of Le. mesenteroides, Lc. lactis, La. paracasei and La. Plantarum, present in the Biotyper 3.0 database. For example, the non-type strain of La. plantarum subsp. plantarum DSM 20205 is more distant from the type strain of this subspecies than the type strain of La. plantarum subsp. argentoratensis (Figure 4A). In the case of La. paracasei, there is a greater distance among strains of the same subspecies than among strains of different species (Figure 4A). In the case of Le. mesenteroides, the non-type strain Le. mesenteroides subsp. mesenteroides DSM 2040 is more distant from the type strain of this subspecies than the strain Le. mesenteroides subsp. dextranicum DSM 20187 (Figure 4B). Several strains assigned to Lc. lactis subsp. lactis are more distant from the type strain of this subspecies than to that of Lc. lactis subsp. cremoris (Figure 4B). These results indicate that several non-type strains held in DSMZ culture collection which are included in the Biotyper 3.0 database are not correctly classified at species or subspecies levels, but, as no gene sequences are available for these strains, we cannot know their correct taxonomic name and this could lead to errors in the identification of any tested strain. For this reason, we always referred to a type strain in the identification of our strains, although the score values were lower than those found for non-type strains (Table 1).
Figure 4.
Cluster analysis of MALDI-TOF MS spectra of the strains belonging to the species identified in this study which are included in the biotyper 3.0 database within genera Lactobacillus (A) and Leuconostoc and Lactococcus (B). Distance is displayed in relative units.
Moreover, we found some surprising score values for the type strains of the subspecies from La. paracasei and Lc. lactis because the subspecies are infraspecific taxa and score values higher than 2.3 among these subspecies are expected after MALDI-TOF MS analysis (Table 2). In the case of the type strains of the subspecies paracasei and tolerans of La. paracasei, the score value was clearly lower than 2.0, indicating that these strains do not belong to the same species (Table 2). This contrasts with the high similarity value of the pheS gene sequences and the high ANIb and dDDH values calculated from their genomes (Table 2). These two values, which are clearly higher than those proposed for species differentiation [28], confirmed that the type strains of paracasei and tolerans belong to the same species, therefore the type strains of these subspecies held in DSMZ culture collection and in the Biotyper 3.0 database should be revised. In addition, they showed that the dDDH values are higher than those proposed for subspecies differentiation [29], therefore the taxonomic status of these subspecies should be revised.
In the case of the type strains of the subspecies lactis and cremoris of Lc. lactis, the low values found in the pheS gene analysis agree with the calculated ANIb and dDDH values, which were lower than those proposed for species differentiation, confirming that they belong to different species (Table 2). Concerning to the other two subspecies, hordniae and tructae, not included in Biotyper database, the pheS gene analysis showed that their type strains are phylogenetically related to the subspecies lactis and cremoris, respectively (Table 2). The calculated ANIb and dDDH values confirmed that Lc. lactis really contains two different species with two subspecies each, although the dDDH values were near to or slightly lower than those proposed for subspecies differentiation in both cases (Table 2). These results clearly indicate that the taxonomic status of the subspecies currently included within Lc. lactis should be revised in order to separate the subspecies cremoris as a novel species and to evaluate whether the subspecies hordniae and tructae can maintain their current taxonomic status.
Conversely, the score value found between the type strains of the subspecies plantarum and argentoratensis of La. plantarum was surprisingly high (2.424) considering the low similarity value found between their pheS genes (90.5%), and that the calculated ANIb and dDDH values were lower than those proposed for bacterial species differentiation (Table 2). These results indicate that the type strain of the subspecies argentoratensis held in DSMZ culture collection and in the Biotyper 3.0 database should be revised and that it should be reclassified as a novel species.
In the case of the subspecies mesenteroides and cremoris of Le. mesenteroides score, values higher than 2.3 were expected, as this corresponds to strains from the same species (Table 2). In agreement, they showed high similarity in their pheS genes and the calculated ANIb and dDDH values were higher than those proposed for bacterial species differentiation (Table 2). In the case of the subspecies dextranicum and jonggajibkimchii, absent in the Biotyper 3.0 database, the pheS gene analysis also showed high similarity values in agreement with those of the ANIb and dDDH, which were also higher than those proposed for bacterial species’ differentiation. Moreover, the dDDH values among the type strains of all these subspecies considerably exceed the upper limit proposed for subspecies differentiation, and therefore their taxonomic status should be revised.
Therefore, the application of the currently accepted ANIb and dDDH cut-off values for species differentiation [28] will lead to the promotion of some subspecies to the taxonomic status of species. Conversely, the application of the dDDH threshold value for subspecies differentiation [29] will lead to the loss of taxonomic status for some subspecies, as recently occurred with the subspecies sakuensis of Serratia marcescens [30]. Although Chun et al. [28] considered that currently there is not enough information to establish general guidelines for species differentiation on the basis of genome data, we face the dilemma of whether to increase the dDDH threshold value for subspecies differentiation, maintain the existing ones, or reject many of the existing subspecies in several species of LAB. Before trying to solve this, we should take into account that the increase necessary to maintain the current subspecies would cause a dramatic increase in the number of these taxa, since the pheS gene similarity cut-off values would be above 99% and only in this study several strains cannot be assigned to any of the described subspecies because they fall within these limits and could be considered as novel subspecies. We should also consider that applying the dDDH thresholds values proposed by Meier-Kolthoff et al. [29] would mean that none of the subspecies from the species identified in this study can maintain their taxonomic status, except perhaps the subspecies hordniae of Lc. lactis. This second option better agrees with the results of the MALDI-TOF MS analysis that clearly allows the identification of the strains isolated at species level, compared to identification at subspecies level. In any case, this situation should be clarified, since it affects several LABs from different genera and species, and currently there is an increasing interest in the identification of these bacteria, particularly in fermented foods.
5. Conclusions
The LAB present in the two cheeses of type “Torta” analysed in this study were identified as La. curvatus, La. diolivorans, La. paracasei, La. plantarum, La. rhamnosus, Lc. lactis and Le. mesenteroides through MALDI-TOF MS and pheS gene analyses. These results confirmed that MALDI-TOF MS is a reliable method for the identification of LAB comparable to pheS gene sequence analysis and presents important advantages over gene sequencing in terms of rapidity and cost per sample. The analysis of pheS gene showed low similarity values for some subspecies of Lc. lactis and for the two subspecies of La. plantarum and values near to 100% for the subspecies of Le. mesenteroides and La. paracasei. These results were confirmed by the calculated ANIb and dDDH values of their whole genomes, showing the need for a revision of the taxonomic status of these species and their subspecies, which should be based on additional criteria.
Acknowledgments
The authors thank the Strategic Research Programs for Units of Excellence from Junta de Castilla y León (CLU-2O18-04). The pheS genes were sequenced in the Sequencing DNA service (NUCLEUS) from Salamanca University (Spain).
Abbreviations
| LAB | Lactic acid bacteria |
| La. | Lactobacillus |
| Le. | Leuconostoc |
| Lc. | Lactococcus |
Author Contributions
Conceptualization, J.D.F.-F., E.V. and F.S.-J.; methodology, F.S.-J E.V. and J.D.F.-F.; software, J.D.F.-F. and F.S.-J.; validation, J.D.F.-F. and J.M.G.-B.; formal analysis, V.T.-M., E.V., F.S.J. and J.D.F.-F.; investigation, J.D.F.-F., V.T.-M., E.V., F.S.-J. and J.M.G.-B.; data curation, E.V. and F.S.-J.; writing—original draft preparation, F.S.-J., J.D.F.-F. and E.V.; writing—review and editing, F.S.-J., J.D.F.-F., V.T.-M., J.M.G.-B. and E.V.; visualization, F.S.-J., J.D.F.-F., V.T.-M., J.M.G.-B. and E.V.; supervision, F.S.-J. and J.D.F.-F. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding
Conflicts of Interest
The authors declare no conflict of interest.
References
- 1.Ludwig W., Schleifer K.H., Whitman W.B. Lactobacillales ord. nov. In: Whitman W.B., Rainey F., Kämpfer P., Trujillo M., Chun J., DeVos P., Hedlund B., Dedysh S., editors. Bergey’s Manual of Systematics of Archaea and Bacteria. Wiley Online Library; Hoboken, NJ, USA: 2015. [DOI] [Google Scholar]
- 2.Zielińska D., Kolożyn-Krajewska D. Food-origin lactic acid bacteria may exhibit probiotic properties: Review. Biomed Res. Int. 2018;2018:5063185. doi: 10.1155/2018/5063185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ruiz-Rodríguez L., Bleckwedel J., Ortiz E., Pescuma M., Mozzi F. Lactic Acid Bacteria. In: Wittmann C., Liao J.C., editors. Industrial Biotechnology. Wiley-VCH; Weinheim, Germany: 2016. pp. 395–451. [Google Scholar]
- 4.Poullet B., Huertas M., Sánchez A., Cáceres P., Larriba G. Main lactic acid bacteria isolated during ripening of Casar de Caceres cheese. J. Dairy Res. 1993;60:123–127. doi: 10.1017/S0022029900027412. [DOI] [Google Scholar]
- 5.Ordiales J., Benito M.J., Martín A., Casquete R., Serradilla M.J., de Guía Córdoba M. Bacterial communities of the traditional raw ewe’s milk cheese “Torta del Casar” made without the addition of a starter. Food Control. 2013;33:448–454. doi: 10.1016/j.foodcont.2013.03.027. [DOI] [Google Scholar]
- 6.Morandi S., Brasca M., Lodi R. Technological, phenotypic and genotypic characterisation of wild lactic acid bacteria involved in the production of Bitto PDO Italian cheese. Dairy Sci. Technol. 2011;91:341–359. doi: 10.1007/s13594-011-0016-7. [DOI] [Google Scholar]
- 7.Colombo E., Franzetti L., Frusca M., Scarpellini M. Phenotypic and genotypic characterization of lactic acid bacteria isolated from Artisanal Italian goat cheese. J. Food Prot. 2010;73:657–662. doi: 10.4315/0362-028X-73.4.657. [DOI] [PubMed] [Google Scholar]
- 8.Pogačić T., Mancini A., Santarelli M., Bottari B., Lazzi C., Neviani E., Gatti M. Diversity and dynamic of lactic acid bacteria strains during aging of a long ripened hard cheese produced from raw milk and undefined natural starter. Food Microbiol. 2013;36:207–215. doi: 10.1016/j.fm.2013.05.009. [DOI] [PubMed] [Google Scholar]
- 9.Pangallo D., Saková N., Koreňová J., Puškárová A., Kraková L., Valík L., Kuchta T. Microbial diversity and dynamics during the production of May bryndza cheese. Int. J. Food Microbiol. 2014;170:38–43. doi: 10.1016/j.ijfoodmicro.2013.10.015. [DOI] [PubMed] [Google Scholar]
- 10.Franciosi E., Carafa I., Nardin T., Schiavon S., Poznanski E., Cavazza A., Larcher R., Tuohy K.M. Biodiversity and γ-aminobutyric acid production by lactic acid bacteria isolated from traditional alpine raw cow’s milk cheeses. Biomed Res. Int. 2015;2015:625740. doi: 10.1155/2015/625740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Domingos-Lopes M.F.P., Stanton C., Ross P.R., Dapkevicius M.L.E., Silva C.C.G. Genetic diversity, safety and technological characterization of lactic acid bacteria isolated from artisanal Pico cheese. Food Microbiol. 2017;63:178–190. doi: 10.1016/j.fm.2016.11.014. [DOI] [PubMed] [Google Scholar]
- 12.Foschi C., Laghi L., Parolin C., Giordani B., Compri M., Cevenini R., Marangoni A., Vitali B. Novel approaches for the taxonomic and metabolic characterization of lactobacilli: Integration of 16S rRNA gene sequencing with MALDI-TOF MS and 1H-NMR. PLoS ONE. 2017;12:e0172483. doi: 10.1371/journal.pone.0172483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Nacef M., Chevalier M., Chollet S., Drider D., Flahaut C. MALDI-TOF mass spectrometry for the identification of lactic acid bacteria isolated from a French cheese: The Maroilles. Int. J. Food Microbiol. 2017;247:2–8. doi: 10.1016/j.ijfoodmicro.2016.07.005. [DOI] [PubMed] [Google Scholar]
- 14.Sato H., Torimura M., Kitahara M., Ohkuma M., Hotta Y., Tamura H. Characterization of the Lactobacillus casei group based on the profiling of ribosomal proteins coded in S10-spc-alpha operons as observed by MALDI-TOF MS. Syst. Appl. Microbiol. 2012;35:447–454. doi: 10.1016/j.syapm.2012.08.008. [DOI] [PubMed] [Google Scholar]
- 15.Soro-Yao A.A., Schumann P., Thonart P., Djè K.M., Pukall R. The use of MALDI-TOF Mass Spectrometry, ribotyping and phenotypic tests to identify lactic acid bacteria from fermented cereal foods in Abidjan (Côte d’Ivoire) Open Microbiol. J. 2014;8:78–86. doi: 10.2174/1874285801408010078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Tanigawa K., Kawabata H., Watanabe K. Identification and typing of Lactococcus lactis by matrix-assisted laser desorption ionization-time of flight mass spectrometry. Appl. Environ. Microbiol. 2010;76:4055–4062. doi: 10.1128/AEM.02698-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Doan N.T., Van Hoorde K., Cnockaert M., De Brandt E., Aerts M., Le Thanh B., Vandamme P. Validation of MALDI-TOF MS for rapid classification and identification of lactic acid bacteria, with a focus on isolates from traditional fermented foods in Northern Vietnam. Lett. Appl. Microbiol. 2012;55:265–273. doi: 10.1111/j.1472-765X.2012.03287.x. [DOI] [PubMed] [Google Scholar]
- 18.Nguyen D.T., Van Hoorde K., Cnockaert M., De Brandt E., Aerts M., Binh Thanh L., Vandamme P. A description of the lactic acid bacteria microbiota associated with the production of traditional fermented vegetables in Vietnam. Int. J. Food Microbiol. 2013;163:19–27. doi: 10.1016/j.ijfoodmicro.2013.01.024. [DOI] [PubMed] [Google Scholar]
- 19.Ferreira L., Sánchez-Juanes F., García-Fraile P., Rivas R., Mateos P.F., Martínez-Molina E., González-Buitrago J.M., Velázquez E. MALDI-TOF mass spectrometry is a fast and reliable platform for identification and ecological studies of species from family Rhizobiaceae. PLoS ONE. 2011;6:e20223. doi: 10.1371/journal.pone.0020223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Altschul S.F., Gish W., Miller W., Myers E.W., Lipman D.J. Basic local alignment search tool. J. Mol. Biol. 1990;215:403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- 21.Thompson J.D., Gibson T.J., Plewniak F., Jeanmougin F., Higgins D.G. The clustalX windows interface: Flexible strategies for multiple sequence alignement aided by quality analysis tools. Nucleic Acids Res. 1997;24:4876–4882. doi: 10.1093/nar/25.24.4876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980;16:111–120. doi: 10.1007/BF01731581. [DOI] [PubMed] [Google Scholar]
- 23.Saitou N., Nei M. A neighbour-joining method: A new method for reconstructing phylogenetics trees. Mol. Biol. Evol. 1987;44:406–425. doi: 10.1093/oxfordjournals.molbev.a040454. [DOI] [PubMed] [Google Scholar]
- 24.Kumar S., Stecher G., Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016;3:1870–1874. doi: 10.1093/molbev/msw054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Richter M., Rosselló-Mora R., Glöckner F.O., Peplies J. JSpeciesWS: A web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics. 2016;32:929–931. doi: 10.1093/bioinformatics/btv681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Meier-Kolthoff J.P., Auch A.F., Klenk H.P., Göker M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform. 2013;14:60. doi: 10.1186/1471-2105-14-60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Auch A.F., von Jan M., Klenk H.P., Göker M. Digital DNA–DNA hybridization for microbial species delineation by means of genome-to-genome sequence comparison. Stand. Genom. Sci. 2010;2:117–134. doi: 10.4056/sigs.531120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Chun J., Oren A., Ventosa A., Christensen H., Arahal D.R., da Costa M.S., Rooney A.P., Yi H., Xu X.W., De Meyer S., et al. Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int. J. Syst. Evol. Microbiol. 2018;68:461–466. doi: 10.1099/ijsem.0.002516. [DOI] [PubMed] [Google Scholar]
- 29.Meier-Kolthoff J.P., Hahnke R.L., Petersen J., Scheuner C., Michael V., Fiebig A., Rohde C., Rohde M., Fartmann B., Goodwin L.A., et al. Complete genome sequence of DSM 30083(T), the type strain (U5/41(T) of Escherichia coli, and a proposal for delineating subspecies in microbial taxonomy. Stand. Genomic. Sci. 2014;9:2. doi: 10.1186/1944-3277-9-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Doijad S., Chakraborty T. Genome-based analyses indicate that Serratia marcescens subsp. marcescens and Serratia marcescens subsp. sakuensis do not merit separation to subspecies status. Int. J. Syst. Evol. Microbiol. 2019;69:3924–3926. doi: 10.1099/ijsem.0.003706. [DOI] [PubMed] [Google Scholar]