Abstract
Amyotrophic lateral sclerosis (ALS) is a fatal disease, characterized by the selective loss of motor neurons leading to paralysis. Mutations in the gene encoding superoxide dismutase 1 (SOD1) are the second most common cause of familial ALS, and considerable evidence suggests that these mutations result in an increase in toxicity due to protein misfolding. We previously demonstrated in the SOD1G93A rat model that misfolded SOD1 exists as distinct conformers and forms deposits on mitochondrial subpopulations. Here, using SOD1G93A rats and conformation-restricted antibodies specific for misfolded SOD1 (B8H10 and AMF7-63), we identified the interactomes of the mitochondrial pools of misfolded SOD1. This strategy identified binding proteins that uniquely interacted with either AMF7-63 or B8H10-reactive SOD1 conformers as well as a high proportion of interactors common to both conformers. Of this latter set, we identified the E3 ubiquitin ligase TNF receptor–associated factor 6 (TRAF6) as a SOD1 interactor, and we determined that exposure of the SOD1 functional loops facilitates this interaction. Of note, this conformational change was not universally fulfilled by all SOD1 variants and differentiated TRAF6 interacting from TRAF6 noninteracting SOD1 variants. Functionally, TRAF6 stimulated polyubiquitination and aggregation of the interacting SOD1 variants. TRAF6 E3 ubiquitin ligase activity was required for the former but was dispensable for the latter, indicating that TRAF6-mediated polyubiquitination and aggregation of the SOD1 variants are independent events. We propose that the interaction between misfolded SOD1 and TRAF6 may be relevant to the etiology of ALS.
Keywords: amyotrophic lateral sclerosis (ALS) (Lou Gehrig disease), superoxide dismutase (SOD), protein misfolding, mitochondria, TNF receptor–associated factor (TRAF), ubiquitin, protein aggregation
Introduction
Amyotrophic lateral sclerosis (ALS)4 is a progressive paralytic and ultimately fatal disease, characterized by the degeneration of upper and lower motor neurons in the brain, brainstem, and spinal cord. Mutations in the SOD1 gene, encoding superoxide dismutase 1, were the first to be identified as causative of ALS (1). SOD1 mutations are the second most common genetic cause of ALS to date, accounting for about 12% of familial cases (fALS) and 1% of sporadic ALS cases (2). Over 180 different mutations in SOD1 have thus far been reported as pathogenic (RRID:SCR_018075). These mostly comprise missense mutations in coding regions, but rare nonsense mutations, deletions, insertions, and mutations in noncoding regions have also been reported (3). Mutations in the coding region are found in all five exons of SOD1. It is now agreed that SOD1-mediated ALS rises from a toxic gain–of–function due to mutation-induced changes in the folding of the SOD1 protein, collectively referred to as misfolded SOD1.
Misfolded SOD1 exposes certain structural features, normally buried in natively folded SOD1, that have been exploited for the generation of antibodies that detect misfolded SOD1 on a conformation-restricted basis (4, 5). Among those, the mouse monoclonal antibody B8H10 recognizes an epitope within the metal-binding loop (loop IV, within aa. 57–80), the exposure of which is associated with a deficiency in metal co-factor binding (6–8). The rabbit monoclonal AMF7-63 is the high-affinity descendant of the mouse monoclonal DSE2 and was raised against residues in the electrostatic loop (loop VII, aa. 125–142), the exposure of which is associated with increased conformational flexibility in this region (9, 10). Strategies to target and thus eliminate misfolded SOD1 in transgenic rodent models, using either active or passive immunization approaches, show benefit in delaying disease phenotypes (6, 8, 11–15). Whether or not a common target exists for misfolded SOD1 remains unknown. Using B8H10 and AMF7-63, we have previously demonstrated in the SOD1G93A rat model of ALS that different conformers of misfolded SOD1 deposit on the surface of spinal cord mitochondria in an age-dependent manner, supporting the view that distinct misfolded SOD1 conformers exist (10, 16). Interestingly, the accumulation of both conformers positively correlates with mitochondrial damage (10, 16, 17).
Here, we report on the identification of the E3 ubiquitin ligase TNF receptor–associated factor 6 (TRAF6) as a novel binding partner of misfolded SOD1. In cellulo, we show that the interaction with TRAF6 is unique to mutant but not WT SOD1, and we map the interaction to the C terminus of TRAF6. However, not all SOD1 variants interact equally with TRAF6. We show that TRAF6-interacting and noninteracting SOD1 variants are conformationally distinct in terms of their exposure of misfolding-associated epitopes recognized by B8H10 and AMF7-63. We demonstrate that TRAF6 stimulates mutant SOD1 polyubiquitination and aggregation but that these variably depend on TRAF6 E3 ubiquitin ligase activity and are therefore independent events. Specifically, siRNA-mediated reduction of TRAF6 levels reduces mutant SOD1 aggregation in a cellular model. Importantly, we uncover that TRAF6 localization is not restricted to the cytoplasm, with a pool of TRAF6 being constitutively localized to spinal cord mitochondria. Finally, TRAF6 is expressed in the cell types harboring misfolded SOD1 in vivo, establishing relevance to misfolded SOD1-linked pathology in ALS. Taken together, these results suggest that misfolded SOD1 interacts with a mitochondrially-localized pool of TRAF6, and this interaction may underlie misfolded SOD1 accumulation.
Results
Identification of novel binding partners of mitochondria-associated misfolded SOD1
In mutant SOD1 rodent models of ALS, a portion of misfolded SOD1 conformers is localized to the surface of spinal cord mitochondria (10, 16–18). To identify the interactomes of specific misfolded SOD1 conformers at the mitochondria, we performed immunoprecipitation coupled with MS (IP-MS) of spinal cord mitochondria isolated from symptomatic SOD1G93A rats (Fig. 1A). Briefly, misfolded SOD1 was immunoprecipitated with anti-B8H10 or anti-AMF7-63, and eluates were analyzed by MS. As published previously, both B8H10 and AMF7-63 specifically detect misfolded forms of SOD1 but are unreactive for natively-folded SOD1WT (8–10, 16, 17). From all identified prey proteins, only those with a maximum peptide score of ≥40 in three of three biological replicates and not detected in IgG controls were considered as potential binding partners. Identified proteins are listed in Table S1. Human SOD1 was detected in all samples and served as an internal control. Also, known binding partners of SOD1 (e.g. the copper chaperone for SOD1 (CCS) (19)) and misfolded SOD1 (e.g. voltage-gated anion channel 1 (VDAC1) (20)) were detected, which additionally supports the quality and validity of our dataset. Of the 52 proteins identified, three proteins (6%) interacted uniquely with B8H10-reactive SOD1 conformers and seven uniquely interacted with AMF7-63–reactive conformers (13%) (Fig. 1B and Table S2). Thus, the majority of interactors (81%) were common to B8H10- and AMF7-63–reactive conformers of misfolded SOD1.
Figure 1.
TRAF6 is a novel binding partner of B8H10 and AMF7-63-reactive misfolded SOD1 at the mitochondria. A, workflow for immunoprecipitation-based MS proteomics for the identification of binding partners of mitochondria-associated misfolded SOD1 in the SOD1G93A rat model. Misfolded SOD1 was immunoprecipitated with the misfolded SOD1-specific conformation-restricted antibodies B8H10 and AMF7-63 from SOD1G93A rat spinal cord mitochondria and binding partners identified by nanoLC-MS/MS. B, Venn diagram of identified misfolded SOD1-binding partners common to both B8H10 and AMF7-63 conformers or selective for either. C, novel binding-partner TRAF6 (UniProt ID: B5DF45) was identified based on six unique peptides covering 17% of the protein sequence. D and E, Flag-tagged SOD1 (WT or mutants) and Myc-tagged TRAF6 (WT or deletion mutants) were co-expressed in 293FT cells. Co-immunoprecipitations were performed on Flag–SOD1 or reciprocally on Myc–TRAF6, and co-precipitation of either binding partner was analyzed by immunoblotting. Immunoblotting for Flag–SOD1 (bait) is to demonstrate equal IP efficiency across conditions. Mock refers to transfection with an equivalent amount of empty vector. Note, the slower migrating band in TRAF6 lanes is auto-ubiquitinated TRAF6. Whole-cell lysates (WCL) were loaded to demonstrate equal plasmid expression. Actin serves as loading control. Data are representative of at least three independent experiments.
To extract information on the biological significance of our dataset, we used Enrichr to perform functional annotation and pathway enrichment analyses (21, 22). Our dataset revealed a significant enrichment for molecules participating in biological processes, such as response to unfolded protein (GO:006986, q-value = 1.47E−04), removal of superoxide radicals (GO:0019430, q-value = 5.66E−04), succinyl-CoA metabolic process (GO:0006104, q-value = 3.12E−02), intermediate filament bundle assembly (GO:0045110, q-value = 2.84E−02), and cellular response to interleukin-12 (GO: 0071349, q-value = 4.44E−02) (Table S3). On the basis of molecular function, our dataset significantly accumulated molecules that are C3HC4-type RING finger domain binding (GO:0055131, q-value = 3.52E−04) or possess ATPase activity (GO:0016887, q-value = 4.87E−02) (Table S3). Next, KEGG pathway analysis showed a significant enrichment for molecular players associated not only with ALS (q-value = 1.90E−05), but also Huntington disease (q-value = 3.91E−04), Parkinson disease (q-value = 1.24E−02), and Alzheimer disease (q-value = 2.10E−02). In addition, our dataset revealed an accumulation of molecules participating in antigen processing and presentation (q-value = 2.32E−02) and infectious disease pathways (Table S4).
Presence of ubiquitin in AMF7-63 and B8H10 immunoprecipitates
Ubiquitin is highly conserved and encoded by four different genes: UBB, UBC, UBA52, and RPS27A. Although UBB and UBC encode polyubiquitin precursors made up of multiple head–to–tail repeats of ubiquitin, UBA52 and RPS27A are ubiquitin hybrid genes encoding a fusion protein composed of a single N-terminal ubiquitin moiety fused to a C-terminal extension. In reviewing our misfolded SOD1 interactomes, we noticed that ubiquitin-60S ribosomal protein L40 (RL40), which is encoded by UBA52, was a binding partner detected in both B8H10 and AMF7-63 immunoprecipitates (Table S1 and Fig. S1A). To elucidate whether the peptides listed under RL40 correspond to the N-terminal ubiquitin moiety or the C-terminal extension of the protein, we generated a sequence alignment for rat RL40 with ubiquitin (single moiety) (Fig. S1B). All three peptides listed under the identifier RL40 mapped to the N-terminal ubiquitin moiety and not the C-terminal extension of the protein. This indicates that the detected peptides are not unique for RL40, but they generally correspond to ubiquitin and may thus derive from RL40, RS27A, UBB, or UBC proteins. To illustrate this, we also aligned rat RS27A (Fig. S1B). To investigate whether ubiquitin is indeed associated with B8H10- and AMF7-63-reactive misfolded SOD1, we precipitated misfolded SOD1 from SOD1G93A rat spinal mitochondria and immunoblotted for ubiquitin (Fig. S1C). In both B8H10- and AMF7-63-immunoprecipitates, ubiquitin was detected as a high-molecular-weight smear refusing entry into the separating gel, a pattern typically observed with intensely polyubiquitin-modified proteins. Taken together, these data suggest that B8H10 and AMF7-63 conformers are modified with ubiquitin (or polyubiquitin) or associate with stable binding partners that are themselves ubiquitinated.
Identification of the E3 ubiquitin ligase TRAF6 as a novel binding partner of misfolded SOD1
In our dataset, we detected six unique peptides for an E3 ubiquitin ligase: TNF receptor-associated factor 6 (TRAF6). TRAF6 peptides, covering 17% of the protein sequence, were detected in both B8H10 and AMF7-63 immunoprecipitates (Table S1, Fig. 1C, and Fig. S2).
TRAF6 is a cytoplasmic-localized RING-type ubiquitin ligase classically known for its role in the canonical NF-κB–signaling pathway, where it participates in the transduction of signals downstream of a multitude of immunoregulatory receptors (23–26) and functions as a hub protein in the cross-talk of immune-regulatory pathways with the autophagy machinery (27–30). TRAF6 is not strictly localized to the cytoplasm but translocates to mitochondria in certain contexts (31–35). Importantly, TRAF6 has been previously implicated in neurodegenerative diseases, including Parkinson's disease (36–38), Huntington's disease (39), and Alzheimer's disease (40–42), where it stimulates the ubiquitination of disease-associated proteins and exerts a regulatory role on their turnover and aggregation (43). The detection of TRAF6 as a binding partner of misfolded SOD1 was additionally intriguing, because physiological interactions between E3 ubiquitin ligases and their substrates are generally known to be weak, transient, and not readily detected by IP-MS–based techniques (44–46). We speculated that the interaction is unnaturally stable and of possible pathological relevance. Thus, we elected to more fully investigate the interaction between mutant/misfolded SOD1 and TRAF6.
TRAF6 interacts with mutant but not WT SOD1
To characterize the interaction between mutant SOD1 and TRAF6, we established an in cellulo system based on transient co-expression of Flag-tagged SOD1 and Myc-tagged TRAF6 in human 293FT cells. To elucidate whether this system can recapitulate our in vivo results, we performed reciprocal co-immunoprecipitations. SOD1G93A and SOD1G85R, but not SOD1WT, co-precipitated TRAF6WT (Fig. 1D, lanes 5–7). Reversely, TRAF6WT co-precipitated mutant SOD1, but not SOD1WT (Fig. 1E, lanes 5–7), thus confirming our IP-MS results. Note, the slower-migrating TRAF6 band is the auto-ubiquitinated form, consistent with previous work (47). In addition, these data indicate that TRAF6 selectively interacts with mutant but not WT SOD1, indicating that the interaction with TRAF6 is unique to nonnative SOD1 conformers.
Mutant SOD1 binds the TRAF6 C terminus
TRAF6 is a multidomain protein and is highly conserved across mammalian species (48, 49). The N terminus features a really interesting new gene domain (RING) with a series of zinc finger motifs, which are connected via an α-helical coiled-coil motif to a C-terminal meprin and TRAF homology (MATH) domain (Fig. 2, A and B). The RING domain and adjacent zinc finger motifs are involved in E2 ubiquitin conjugase binding and are required for TRAF6 function as an E3 ubiquitin ligase (50). The MATH domain is an eight-stranded anti-parallel β-sandwich with which TRAF6 engages in a multitude of protein–protein interactions (32, 51–53). To determine the domain of TRAF6 that interacts with mutant SOD1, two TRAF6 deletion constructs were generated: TRAF6ΔC (aa. 1–288) comprising the N-terminal RING domain and zinc finger motifs, and TRAF6ΔN (aa. 289–522) comprising the central coiled-coil motif and C-terminal MATH domain (Fig. 2, A and B). SOD1G93A and SOD1G85R efficiently co-precipitated TRAF6ΔN but not TRAF6ΔC (Fig. 2C, compare lanes 14 and 15 to lanes 11 and 12). SOD1WT did not interact with TRAF6WT nor the two TRAF6 deletion mutants (Fig. 2C, lanes 7, 10, and 13). We conclude that the C terminus of TRAF6 is necessary and sufficient for mutant SOD1 binding.
Figure 2.
C terminus of TRAF6 is necessary and sufficient for mutant SOD1 binding. A, schematic representation of generated Myc–TRAF6 deletion constructs. TRAF6ΔC (aa. 1–288) comprises the N-terminal RING domain and zinc finger motifs. TRAF6ΔN (aa. 289–522) comprises the central coiled-coil motif (CC) and the C-terminal MATH domain. B, protein structure models for TRAF6WT, TRAF6ΔC, and TRAF6ΔN. C, Flag-tagged SOD1 (WT or mutants) and Myc-tagged TRAF6 (WT or deletion mutants) were co-expressed in 293FT cells. Co-immunoprecipitations were performed on Flag–SOD1, and Myc–TRAF6 co-precipitation was assessed by immunoblotting. Immunoblotting for Flag–SOD1 (bait) is to demonstrate equal IP efficiency across conditions. Mock refers to transfection with an equivalent amount of empty vector. WCL were loaded to demonstrate equal plasmid expression. Actin serves as loading control. Data are representative of at least three independent experiments.
SOD1 mutation-dependent variation in TRAF6 binding
To evaluate whether the interaction of mutant SOD1 with TRAF6 is a general property of SOD1 variants, we generated Flag-tagged expression plasmids for six additional fALS-associated SOD1 missense mutations: SOD1A4V, SOD1G37R, SOD1H46R, SOD1E100G, SOD1V148G, and SOD1V148I. We also cloned SOD1G127X, a fALS-associated SOD1 truncation mutant lacking the C terminus. Again, we co-expressed these mutants with Myc–TRAF6 (WT or ΔN) in 293FT cells, immunoprecipitated Flag–SOD1 from cell lysates, and assessed Myc–TRAF6 co-precipitation by immunoblot (Fig. 3, A and B). As demonstrated previously, SOD1G93A and SOD1G85R co-precipitated TRAF6WT to equivalent amounts (Fig. 1, D and E). A similar level of interaction was also observed between TRAF6WT and SOD1E100G and SOD1G127X, SOD1V148G, and SOD1G37R, whereas SOD1A4V showed the most robust interaction (Fig. 3A, lanes 7–13). In contrast, SOD1H46R very weakly co-precipitated TRAF6WT (Fig. 3A, lane 6). Interestingly, SOD1V148I did not co-precipitate with TRAF6WT (Fig. 3A, lane 5) nor TRAF6ΔN (Fig. 3C, lane 11). Taken together, there is variation in TRAF6 binding by SOD1 mutant proteins.
Figure 3.
SOD1 mutation-dependent variation in the interaction with TRAF6 in cellulo. A–C, Flag-tagged SOD1 (WT or mutants) and Myc-tagged TRAF6 (WT or ΔN) were co-expressed in 293FT cells. Co-immunoprecipitations were performed on Flag–SOD1, and Myc–TRAF6 co-precipitation was analyzed by immunoblot. Immunoprecipitates were immunoblotted for Flag–SOD1 to demonstrate equal IP efficiency across conditions. Mock refers to transfection with an equivalent amount of empty vector. WCLs were loaded to demonstrate equal plasmid expression. Actin serves as loading control. B, densitometric analysis of the co-immunoprecipitated amount of Myc–TRAF6 relative to the amount of immunoprecipitated Flag–SOD1. Plotted values are the mean ± S.D. of four independent experiments. *, p < 0.05; **, p < 0.01; ***, p < 0.001; and ****, p < 0.0001.
Consistent with the view that more than one misfolded SOD1 conformer exists (10, 54), these data suggest that many SOD1 mutant proteins adopt a misfolded conformation(s) that is (are) favorable for TRAF6 binding, but that these are not conformationally identical, which may account for the observed variable interaction with TRAF6. Interestingly, one ALS-causing mutant, SOD1V148I, did not interact with TRAF6, consistent with previous data indicating that this mutant is WT-like in terms of stability and structure (55–57). Strikingly, the substitution at Val-148 with glycine is sufficient for mutant SOD1 to adopt a TRAF6-interacting conformation.
Finally, the variation in mutant SOD1 binding was observed for TRAF6WT but not for TRAF6ΔN. SOD1A4V and SOD1V148G co-precipitated TRAF6WT to variable amounts but TRAF6ΔN to similar amounts (Fig. 3C, compare lanes 6 and 7 to lanes 9 and 10). Although the TRAF6 C terminus is required for mutant SOD1 binding, these data suggest that the N terminus of TRAF6 modulates the interaction with mutant SOD1.
SOD1 misfolding–associated functional loop exposure is required for TRAF6 interaction
Next, we sought to investigate the relevance of SOD1 misfolding-associated exposure of the metal-binding loop (loop IV) and electrostatic loop (loop VII) to TRAF6 binding. For this, Flag–SOD1 variants expressed in 293FT cells were evaluated for reactivity to the conformation-restricted misfolded SOD1 antibodies B8H10 (epitope in loop IV) or AMF7-63 (epitope in loop VII) by immunoprecipitation (Fig. 4, A and B). We incorporated the C-terminal truncation mutant SOD1G127X as a negative control for AMF7-63 immunoprecipitations, as this mutant lacks the electrostatic loop (loop VII). Although most SOD1 variants immunoprecipitated with both B8H10 and AMF7-63, SOD1V148I was not immunoprecipitated with either antibody (Fig. 4B, compare lanes 3–9 to lane 10). SOD1G127X was not immunoprecipitated with AMF7-63 (as expected), but was with B8H10 (Fig. 4B, lane 11). As expected, neither antibody immunoprecipitated SOD1WT (Fig. 4B, lane 2). To summarize, all TRAF6-interacting SOD1 variants were reactive for either B8H10 and/or AMF7-63. In contrast, SOD1WT and SOD1V148I, which both do not interact with TRAF6, were unreactive for both antibodies suggesting that SOD1 misfolding-associated exposure of the functional loops or monomerization/unfolding of SOD1 are conformational requirements for TRAF6 interaction. Finally, data obtained with SOD1G127X indicate that exposure of the metal-binding loop is sufficient for TRAF6 binding.
Figure 4.
Exposure of epitopes associated with SOD1 misfolding discerns TRAF6-interacting from noninteracting mutants. A, schematic representation of Flag–SOD1 constructs, with the location of mutation sites relative to B8H10 (loop IV) and AMF7-63 (loop VII) antibody epitopes. SOD1G127X is a truncation mutant with a nonnative C terminus and inherently lacks the AMF7-63 epitope. B, Flag-tagged SOD1 (WT or mutants) was expressed in 293FT cells. Misfolded SOD1 was immunoprecipitated with the B8H10 or AMF7-63 antibody, and precipitates were analyzed by Western blotting. Mock refers to transfection with an equivalent amount of empty vector. WCLs were loaded to demonstrate equal plasmid expression. Actin serves as loading control. Data are representative of three independent experiments.
TRAF6 stimulates mutant SOD1 ubiquitination
TRAF6 is reported to stimulate the ubiquitination and aggregation of mutant/misfolded proteins in the context of other neurodegenerative diseases (36, 39, 42). To determine whether mutant SOD1 is also a substrate of TRAF6, we performed in cellulo ubiquitination assays (Fig. 5A). For this, we co-expressed Flag–SOD1 and Myc–TRAF6WT or E3 ubiquitin ligase-inactive mutant TRAF6C70A (58) with HA-ubiquitin, immunoprecipitated HA-ubiquitin from whole-cell lysates, and immunoblotted for Flag–SOD1. In this assay, we selected a SOD1 variant that demonstrates robust interaction with TRAF6 and is dually reactive for B8H10 and AMF7-63 (SOD1A4V), a SOD1 variant that exhibits a slightly less robust interaction with TRAF6 and is also reactive for B8H10 and AMF7-63) (SOD1V148G), and a variant that does not interact with TRAF6 and does not display B8H10- or AMF7-63–reactive SOD1 epitopes (SOD1V148I). TRAF6 strongly stimulated the polyubiquitination of SOD1A4V and SOD1V148G, but not SOD1V148I (Fig. 5A, compare lanes 5–7). The polyubiquitin signature on SOD1A4V and SOD1V148G was absent in cells co-expressing TRAF6C70A (Fig. 5A, compare lanes 5 and 6 to 8 and 9). These data indicate that TRAF6 can stimulate the polyubiquitination of mutant SOD1 in an E3 ubiquitin ligase activity-dependent manner. Moreover, this effect is selective to TRAF6-interacting SOD1 variants, and the level of detected polyubiquitin-modified mutant SOD1 positively correlates with the observed differences in the TRAF6–mutant SOD1 interaction.
Figure 5.
TRAF6-mediated polyubiquitination of mutant SOD1 depends on its ubiquitin ligase activity and is selective to TRAF6-interacting SOD1 variants. A, in cellulo ubiquitination assays with mutant SOD1 as substrate and TRAF6 as catalyzing E3 ubiquitin ligase. Flag–SOD1A4V and SOD1V148G (TRAF6-interacting) or Flag–SOD1V148I (TRAF6-noninteracting) was co-expressed with Myc–TRAF6WT or ubiquitin ligase-inactive TRAF6C70A in the presence of HA-ubiquitinWT in 293FT cells. Immunoprecipitations were performed on HA-ubiquitin, and immunoprecipitates were immunoblotted for Flag–SOD1 and HA-ubiquitin to demonstrate equal IP efficiency across conditions. B, in cellulo ubiquitin-linkage assays were performed similar to A, but in the presence of HA-ubiquitinWT or single-lysine mutants. TRAF6 mediates mutant SOD1 polyubiquitination with primarily Lys-6, Lys-27, and Lys-29 linkages. Note that no polyubiquitinated mutant SOD1 was recovered in the 4% stacking gel (A) and therefore was removed prior to transfer in subsequent experiments. Mock refers to transfection with an equivalent amount of empty vector. WCLs were loaded to demonstrate plasmid expression. Actin serves as loading control. Data are representative of three independent experiments.
Next, to determine the fate of mutant SOD1 with respect to TRAF6-mediated polyubiquitination, we performed ubiquitination assays with single-lysine ubiquitin mutants to elucidate the polyubiquitin chain linkages with which mutant SOD1 is modified (Fig. 5B). Single-lysine ubiquitin mutants have only one of seven lysines available (all other lysines are mutated to arginine) and thus can only be used to catalyze polyubiquitin chains linked through one particular lysine residue. We performed these assays with Flag–SOD1A4V as we determined that this mutant had the strongest interaction with TRAF6 and was the most ubiquitinated. Our data suggest that TRAF6 promotes the polyubiquitination of mutant SOD1, with Lys-6, Lys-27, and Lys-29 linkages prevailing (Fig. 5B, lanes 3, 4, and 6).
TRAF6 is a modifier of mutant SOD1 aggregation
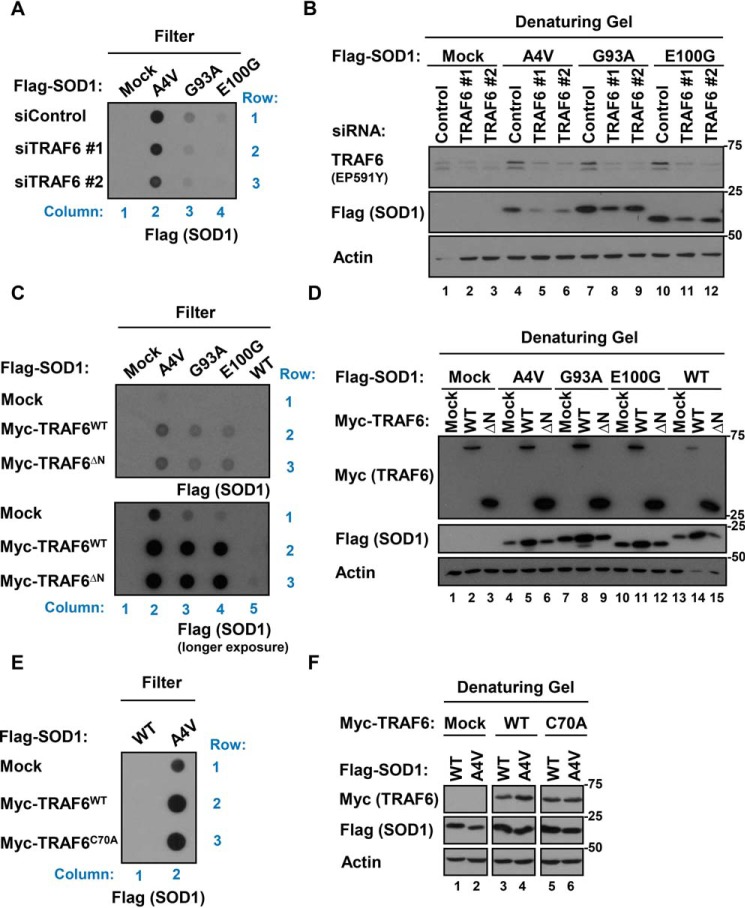
To address whether mutant SOD1 aggregation could be causally linked to the interaction with TRAF6, we performed filter retardation assays using 293FT cells expressing Flag–SOD1 (WT or mutants), either following siRNA-mediated knockdown of TRAF6 (Fig. 6A) or co-expression of Myc–TRAF6 (WT, ΔN, or C70A) (Fig. 6, C and E). Cell lysates were filtered through cellulose acetate membrane filters, which trap protein aggregates with a diameter of ≥200 nm, and immunoblotted for Flag–SOD1. Western blottings of cell lysates were included to verify the efficiency of TRAF6 knockdown, to demonstrate equal plasmid expression across conditions, and to provide an indirect loading control for the filter membranes (Fig. 6, B, D, and F). Note, cell viability was observed to be equivalent across conditions. Flag–SOD1 variants tested in the assays were as follows: SOD1A4V (high-aggregation propensity and strong TRAF6 interaction); SOD1G93A and SOD1E100G (high-aggregation propensity and less strong TRAF6 interaction). In experiments with TRAF6 knockdown, all SOD1 variants aggregated less in cells treated with two independent siRNAs for TRAF6 compared with siControl-treated cells (Fig. 6A, compare rows 2 and 3 to row 1). TRAF6 protein levels were effectively reduced with both siRNAs (Fig. 6B). In contrast, co-expression of TRAF6WT or TRAF6ΔN stimulated the aggregation of mutant SOD1 (Fig. 6C, columns 2–4 compare row 1 to rows 2 and 3), but not SOD1WT (Fig. 6C, column 5). Taken together, these data indicate that lowering TRAF6 levels alleviates mutant SOD1 aggregation, whereas the interaction of mutant SOD1 with the TRAF6 C terminus is sufficient to exacerbate mutant SOD1 aggregation. This is consistent with the nondegradative ubiquitin linkage types observed.
Figure 6.
TRAF6 modifies mutant SOD1 aggregation but independently of its ubiquitin ligase activity. A, C, and E, filter retardation assays in which protein aggregates ≥200 nm in diameter are captured on cellulose acetate membrane filters. Flag–SOD1 (WT or mutants) was either co-expressed with Myc–TRAF6WT, TRAF6ΔN, or ubiquitin ligase–inactive TRAF6C70A, or Flag–SOD1 variants were expressed in 293FT cells that received siTRAF6 or siControl. Filter membranes were immunoblotted for Flag–SOD1. B, D, and F, Western blotting of cell lysates to demonstrate equal plasmid expression and to verify TRAF6 knockdown across conditions. Immunoblots also serve as loading controls for the filter membranes. Mock refers to transfection with an equivalent amount of empty vector. Actin serves as loading control. All data are representative of three independent experiments.
Our results with the TRAF6 N terminus harboring the RING domain, which is required for ubiquitin ligase activity, suggested that the E3 ligase activity may not be implicated in SOD1 aggregation. To directly test this, we co-expressed mutant SOD1 with ubiquitin ligase–inactive TRAF6C70A. TRAF6C70A stimulated SOD1A4V aggregation comparable with TRAF6WT (Fig. 6E, columns 2–4 compare row 2 to row 3), confirming that TRAF6 ubiquitin ligase activity is indeed not required for TRAF6-mediated mutant SOD1 aggregation. A similar result was obtained for SOD1G37R (data not shown). Finally, we examined whether TRAF6C70A-stimulated aggregation of mutant SOD1 correlates with a bimolecular interaction between TRAF6 and mutant SOD1. We observed that mutant SOD1 interacted comparably with TRAF6C70A and TRAF6WT (Fig. S3, compare lanes 7 and 8 to 10 and 11). Taken together, these data indicate that TRAF6 ubiquitin ligase activity is not required for TRAF6 interaction with mutant SOD1 nor TRAF6's capacity to stimulate mutant SOD1 aggregation. We conclude that the interaction of mutant SOD1 with TRAF6 is sufficient to modulate mutant SOD1 aggregation.
Mitochondrial recruitment of TRAF6 in SOD1G93A rats
A pool of mutant SOD1 exists in an aggregated state on the surface of spinal cord mitochondria in SOD1G85R mice and SOD1G93A rats (18). In addition, we have demonstrated that B8H10 and AMF7-63–reactive misfolded SOD1 accumulate in mitochondrial fractions isolated from SOD1G93A rat spinal cords and correlate with disease progression (10). We speculated that these observations could be linked to misfolded SOD1 interacting with TRAF6. TRAF6 is known as a cytoplasmic-localized protein, but its capacity to translocate to mitochondria in certain contexts has been previously reported (31, 33, 34). Because we detected TRAF6 as a binding partner of misfolded SOD1 at the mitochondria, we reasoned that TRAF6 must be recruited. To experimentally address this, we examined TRAF6 levels in spinal cord lysates and purified floated mitochondria from SOD1G93A rats (Fig. S4). We collected samples at two different disease stages: 10 weeks (pre-symptomatic) and 14 weeks (pre-symptomatic/prior to disease onset). These two age groups were chosen for comparison because misfolded SOD1 is not readily detectable in spinal cord mitochondria in 10-week-old SOD1G93A rats but is at 14 weeks (10). Age-matched nontransgenic and SOD1WT-expressing rats were included as controls. Collectively, no differences in TRAF6 protein levels were observed in SOD1G93A rat spinal cord lysates compared with controls and between-age groups (Fig. S4A) nor was there a difference observed in the level of TRAF6 in mitochondrial fractions compared with controls and between-age groups (Fig. S4B). Surprisingly, however, we observed TRAF6 in mitochondrial fractions from nontransgenic animals, indicating that a pool of TRAF6 is constitutively localized to spinal cord mitochondria. These data suggest that misfolded SOD1 interacts with a mitochondrial-localized pool of TRAF6 rather than requiring TRAF6 to be recruited.
Finally, we noticed that the TRAF6 (EP591Y) antibody we used to detect TRAF6 at 60 kDa, corresponding to its reported size, additionally detected two faster-migrating bands at ∼48 and ∼35 kDa (both unreported previously) enriched in spinal cord mitochondrial fractions compared with lysates (Figs. S4 and S5C). Using a range of tools and approaches, we validated that all three detected bands are specific to this antibody (Fig. S5).
Effect of mutant SOD1 on TRAF6-stimulated NF-κB activation
Because the constitutive localization of TRAF6 at the mitochondria is unreported, the function of TRAF6 as a resident mitochondrial protein is unclear. Thus, we further investigated the functional repercussions of mutant/misfolded SOD1 binding to TRAF6 using a proof–of–principle approach. TRAF6 plays a central role in cytoplasmic signal transduction from a multitude of immune receptors in response to stimuli (induced NF-κB signaling) but is also crucial for cellular homeostasis through maintaining constitutive NF-κB activity (basal NF-κB signaling) (59). Constitutive NF-κB activity in the mature CNS is restricted to neurons and is required for neuronal survival (60, 61). Glial cells have very low levels of constitutive NF-κB activity, but activity is strongly induced in inflammatory contexts (62). An up-regulation of NF-κB activity is seen at late stages of disease in mutant SOD1 transgenic mice and primarily in glial cells, which is associated with neuroinflammatory processes (63, 64). Thus, we reasoned that mutant SOD1 binding to TRAF6 may have repercussions on TRAF6's capacity to transduce signals in the NF-κB pathway. To experimentally address this, we performed dual NF-κB luciferase reporter assays in cells expressing Myc–TRAF6 and an escalating dose of Flag–SOD1WT or Flag–SOD1A4V, together with a firefly luciferase NF-κB reporter and a Renilla luciferase internal control (Fig. S6). The amount of transfected Myc–TRAF6 plasmid was previously determined by dose titration and was chosen to yield ∼50% NF-κB activation relative to maximal TRAF6-stimulated activation, thus allowing for the detection of decreased or increased NF-κB activity. SOD1A4V was chosen because it most strongly interacted with TRAF6 in other experiments, and SOD1WT served as TRAF6-noninteracting control. Flag–p65/RelA was incorporated as positive control. The sole expression of Myc–TRAF6 increased NF-κB activity by 7-fold relative to basal level (as expected), but mutant SOD1 did not have a significant effect on this across the entire titration range. This suggests that in this cellular context, mutant SOD1 binding does not adversely affect TRAF6 in activating NF-κB and that the signaling transduction cascade remained intact.
TRAF6 is expressed in the cell types affected in ALS
We and others have previously demonstrated in mutant SOD1 transgenic rodent models that misfolded SOD1 is primarily detected in motor neurons at disease end-stage and rarely in glial cells (8, 10, 16, 65–67). To assess whether TRAF6 is expressed in these cell types, we attempted to detect TRAF6 expression by immunohistochemistry in SOD1G93A rat spinal cords. Unfortunately, none of the four commercial antibodies we tried gave specific labeling in rats, mice, or in cultured cells. As an alternative, we immunoblotted whole-cell lysates prepared from human iPSC–derived motor neurons (Fig. 7A and Fig. S7) and mouse primary cortical neurons and astrocytes (Fig. 7B). TRAF6 was detected in all cell types, but we observed a difference in the ratio of the 60-kDa TRAF6 and the 48-kDa band in mouse cortical astrocytes compared with neurons. Densitometry revealed that 60-kDa TRAF6 was 3-fold more abundant in neurons than in astrocytes, whereas the 48-kDa band was 0.7-fold less abundant in neurons than in astrocytes (Fig. 7C). Moreover, in neurons the 60-kDa TRAF6 was 6.5-fold more abundant than the 48-kDa band and 1.4-fold in astrocytes. These data indicate that TRAF6 is expressed in the cell types that primarily harbor misfolded SOD1 in vivo. Thus, we explored the interaction between misfolded SOD1 and TRAF6 in post-mortem tissues from SOD1 mutation-carriers affected by ALS. Although misfolded SOD1 was detected in the motor cortex of three independent SOD1A4V cases (data not shown), we were unable to detect TRAF6 in misfolded SOD1 immunoprecipitates, possibly because of the low overall abundance of TRAF6 in CNS tissues (68).
Figure 7.
TRAF6 is expressed in ALS disease–relevant cell types. A, Western blotting of whole-cell lysates from human iPSC-derived motor neurons differentiated for 14 days. Data are representative of two independent experiments. B and C, Western blotting of whole-cell lysates from mouse primary cortical astrocytes and neurons and quantification of the abundance of TRAF6 relative to total protein stained with Ponceau S. Data represent the mean ± S.D. from two independent cell preparations. TRAF6 is detected in all cell types as a 60-kDa immunoreactive band (corresponding to the reported size of TRAF6) and a faster-migrating band at about 48 kDa (unreported).
Discussion
In this study, we conducted IP-MS proteomics using the SOD1G93A rat model to identify the interactomes of misfolded SOD1 conformers at the mitochondria. Using antibodies reported to recognize distinct misfolded SOD1 conformers, we found a high degree of overlap for interactors (81%). Gene ontology analyses revealed a convergence on cellular responses related to unfolded proteins and protein ubiquitination. It is noteworthy that a number of interactors identified here are chaperone family members, which is in agreement with studies performed by others in different SOD1 models (69–72). In addition, we confirmed the previously reported interaction between human misfolded SOD1 and VDAC1 in rat spinal cord mitochondria (20). We also report that both misfolded SOD1 conformers interact with CCS, which may be expected given the known role of CCS in SOD1 maturation (73). However, the CCS–SOD1 heterodimer is an obligatorily transient complex (74). Thus, the detection of CCS in complex with misfolded SOD1 proteins here could reflect a sustained interaction that disrupts the available pool of CCS, thus impeding SOD1 maturation, as suggested by others (74). This could effectively contribute to a feed-forward loop of SOD1 misfolding. However, CCS has been previously ruled out as a modifier of the development of ALS-related phenotypes in transgenic SOD1 mice (75). Perhaps this concept should be revisited, given the recent data in monozygotic siblings discordant for ALS where CCS is found to be down-regulated in the affected twin (76).
Interestingly, a few binding partners on our proteomics list were also selectively detected in B8H10 but not AMF7-63 IPs and vice versa (Table S2). Although the significance of these conformer-selective binding partners remains to be investigated, these data support the possibility that different misfolded conformers of SOD1 may have one or more distinct interactome-dependent pathomechanisms. One binding partner that was uniquely identified in B8H10 immunoprecipitates was 2-oxoglutarate dehydrogenase (OGDH), which has been described as an endogenous protein processed for mitochondrial antigen presentation in certain conditions (77, 78). This raises the question whether misfolded SOD1 is itself presented on mitochondrial-derived vesicles, similar to OGDH, or whether misfolded SOD1 may interfere with OGDH antigen presentation.
Another binding partner uniquely identified in B8H10 immunoprecipitates was ATP synthase subunit β (ATP5B), which has been previously reported as an interactor of apoSOD1 (metal co-factor deficient) in rat spinal cords (70). Consistent with this, B8H10 immunoprecipitates the metal-deficient mutants H46R, G85R, and G127X comparably in our in cellulo experiments, and we have previously shown that B8H10 preferentially recognizes apoSOD1 (10).
Interestingly, another ATP synthase subunit, ATP synthase F0 complex subunit β1 (ATP5F1), was uniquely recovered in AMF7-63 immunoprecipitates. Thus, the misfolded SOD1 conformers examined here each bound a different portion of the F1F0-ATP synthetase complex that drives mitochondrial ATP production. Defective mitochondrial ATP production has been previously reported in mutant SOD1-mediated disease and could positively contribute to compromised mitochondrial axonal trafficking (79).
We identified the E3 ubiquitin ligase TRAF6 as a novel binding partner of mitochondria-associated misfolded SOD1. We showed that SOD1 misfolding-associated exposure of the metal-binding loop and electrostatic loop in mutant SOD1 is a contributing factor for interaction with TRAF6. To this end, Garg et al.5 have performed in silico molecular dynamics simulations to further investigate the biophysical nature of the intermolecular interaction between mutant/misfolded SOD1 and TRAF6. It is our view, which is supported by this in silico work and our experimental evidence with SOD1G127X, that the electrostatic loop is not required for TRAF6 binding, because it is absent in this truncation mutant. This leads us to conclude that the interface for SOD1 binding to TRAF6 is formed primarily by residues located in the metal-binding loop, with a secondary contribution of residues in the electrostatic loop, as well as a potential co-option of the native SOD1 dimer interface. Interestingly, the conformational criterion for TRAF6 interaction is not universally fulfilled by all SOD1 variants. SOD1V148I, which is WT-like with respect to stability and structure (55–57), does not adopt a conformation that is characterized by the exposure of the B8H10 or AMF7-63 epitopes and does not interact with TRAF6 in cellulo. These data are reminiscent of the absence of an interaction of SOD1V148I with Derlin-1, a functional component of endoplasmic reticulum–associated degradation (80). Indeed, all TRAF6-interacting SOD1 variants also interact with Derlin-1, suggesting that these mutants may collectively adopt a misfolded conformation that exposes a similar or possibly identical binding site for both interacting proteins. These findings certainly strengthen the notion that different SOD1 mutations may lead to motor neuron toxicity via distinct mechanisms. This possibility could be a contributor to the heterogeneity in disease onset observed for genetic subtypes of SOD1–ALS (81, 82).
As shown previously in the context of other neurodegenerative diseases, TRAF6 stimulates the ubiquitination and aggregation of mutant/misfolded proteins (36, 39, 42, 43). Here, TRAF6 stimulates the polyubiquitination of mutant SOD1 in an E3 ubiquitin ligase activity-dependent manner in cellulo. The polyubiquitin chain linkages we detected on mutant SOD1 were primarily Lys-6-, Lys-27-, and Lys-29–linked, which is reminiscent of the type of chain linkages that have been found on other mutant/misfolded proteins when TRAF6 is co-expressed in cultured cells (36, 39). TRAF6 also stimulated the aggregation of mutant SOD1, but surprisingly, the TRAF6 N terminus and E3 ubiquitin ligase activity were dispensable for this. Therefore, we conclude that these two events are independent, contrary to the currently proposed concept for the modification and autophagic degradation of mutant/misfolded protein aggregates (43). Moreover, polyubiquitinated mutant SOD1 was readily detectable in our assays in the absence of proteasome or autophagy inhibitors. This indicates that TRAF6-stimulated polyubiquitination and/or aggregation of mutant SOD1 does not prompt mutant SOD1 turnover but instead leads to its cellular accumulation. In fact, we observed in multiple experiments (e.g. Figs. 2C (2nd blot from bottom, compare lanes 1 and 7), 3A (2nd blot from bottom, compare lanes 1 and 4, 2 and 5), and 6D (2nd blot from bottom, compare lanes 13–15)) that the co-expression of TRAF6 resulted in increased mutant SOD1 levels in whole-cell lysates. This was observed with TRAF6WT and TRAF6C70A, but not TRAF6ΔN or TRAF6ΔC, and was observed for TRAF6-interacting SOD1 variants as well as for TRAF6 noninteracting SOD1WT and SOD1V148I. Collectively, this suggests that TRAF6 negatively affects mutant SOD1 turnover but that this effect is not dependent on an intermolecular interaction between the two proteins and does not require TRAF6 E3 ligase activity. In line with this conclusion, we observed a reduction in the total level of mutant SOD1 in TRAF6-depleted cells (Fig. 6B), suggesting that lowering TRAF6 levels alleviates the aggregation of mutant SOD1 and prompts mutant SOD1 turnover. Taken together, this alludes to a scenario in which TRAF6 may act as a molecular scaffold serving a mechanical function in the context of mutant/misfolded SOD1 accumulation/aggregation at the mitochondria. That TRAF6 has E3 ubiquitin ligase-independent roles as a scaffolding protein has been previously demonstrated in the context of IL-1 signaling (83). However, given the role of TRAF6 in NF-κB signaling, which is intricately connected with regulating autophagy flux, the possibility remains that mutant SOD1 aggregation is indirectly and dynamically modulated in response to effects of TRAF6 on the autophagy machinery (28, 29, 84).
In addition to being localized to the cytoplasm, we found that a pool of TRAF6 is constitutively localized to mitochondria in the spinal cord. We propose that this mitochondrial TRAF6 interacts with and scaffolds the accumulation of misfolded SOD1. The TRAF6 (EP591Y) antibody we used for immunoblotting specifically detected TRAF6 at 60 kDa in spinal cord mitochondrial fractions and two other bands at 48 and 35 kDa, which were enriched in mitochondrial fractions compared with whole-tissue lysates. This raises the possibility that there is more than one form of TRAF6 and that these forms may be differentially targeted to the cytoplasmic or mitochondrial compartments. How such short forms of TRAF6 are generated and what their function is at the mitochondria remains to be investigated.
TRAF6 is a multifaceted protein; it is a central player at the interface of inflammatory signaling and the autophagy machinery (27–30) and plays a role in mitophagy (34, 35) and RNA metabolism (85, 86), all of which are recurrent pathomechanistic themes in ALS, independent of genetic etiology. As such, we speculate that TRAF6 dysfunction due to mutant/misfolded SOD1 binding could affect cellular integrity at various levels and in different cellular compartments. In fact, TRAF6 could participate in a variety of pathomechanisms underlying a broad spectrum of genetic subtypes of ALS, given that TRAF6 is reported to interact with many proteins already implicated in ALS (e.g. heterogeneous nuclear ribonucleoprotein A1 (86), Matrin-3 (86), Profilin-1 (86), Sqstm1/p62 (87), and TBK1 (88)). Our initial examination of the impact of mutant SOD1/TRAF6 on NF-κB signaling did not yield any differences. However, it is possible that the portion of TRAF6 bound to mutant SOD1 may be small compared with the total TRAF6 pool, which may explain our experimental outcome. Further investigations into the role of TRAF6 in ALS could yield important and unexpected insights, which may help explain the heterogeneity and apparent complexity of the disease.
Experimental procedures
Animals
Transgenic rats expressing human SOD1WT and SOD1G93A have been previously described (89, 90). SOD1G93A rats were bred and monitored biweekly as previously published (16). SOD1G93A rats followed the expected disease course and reached end-stage at around 110 days. SOD1G93A rats at 10 and 14 weeks were pre-symptomatic, defined as no loss of body weight and intact hindlimb reflex. Animals described as symptomatic displayed bilateral hindlimb paralysis with no phenotypic involvement of the forelimbs. SOD1WT-expressing and nontransgenic control animals were age-matched. Animals of both sexes were used. Breeding, housing, and all manipulations were carried out in strict accordance with approved protocols from the Centre de Recherche du Centre Hospitalier de Université de Montréal (CRCHUM) Institutional Committee for the Protection of Animals (CIPA) and the Canadian Council on Animal Care (CCAC).
Antibodies and siRNAs
Rabbit anti-AMF7-63 has been previously published (9, 10). Commercially available primary antibodies used were as follows: mouse anti-actin (MP Biomedicals, 69100); mouse anti-B8H10 (Medimabs, MM-0070); mouse anti-Flag (Sigma, F1804); rabbit anti-GAPDH (Cell Signaling, 5174); chicken anti-GFAP (Abcam, ab4674); rat anti-HA (Chromotek, 7c9); rabbit anti-Myc (Sigma, C3956); rabbit anti-NeuN (Novus Biologicals, NBP1-92693); rabbit anti-SOD1 (Enzo Life Sciences, ADI–SOD-101); rabbit anti-TRAF6 (EP591Y) (Abcam, ab33915, with ab183540 blocking peptide); and mouse anti-VDAC1 (Millipore, MABNS04). Secondary horseradish peroxidase (HRP)-conjugated secondary antibodies were from Jackson ImmunoResearch: anti-chicken HRP(703-035-155); anti-mouse HRP(715-035-151); anti-rabbit HRP(711-035-152); and anti-rat HRP(712-035-150). Stealth siRNAs were from Thermo Fisher Scientific: siTRAF6 HSS110968 (referred to as siTRAF6 #1); siTRAF6 #HSS110970 (referred to as siTRAF6 #2); and siRNA negative control #12935200 (referred to as siControl).
Preparation of mitochondria and lysates from rat spinal cords
Rat spinal cords were homogenized in mitochondria isolation buffer I (10 mm Tris, pH 7.5, 1 mm EDTA, 210 mm d-mannitol, 70 mm d-sucrose) with protease inhibitors (10 μg/ml leupeptin, 10 μg/ml pepstatin A, 10 μg/ml chymotrypsin). Mitochondria were isolated by sedimentation according to previously published protocols (91). For Fig. S4, spinal cords were homogenized in mitochondria isolation buffer II (20 mm HEPES-NaOH, pH 7.4, 1 mm EDTA, 250 mm d-sucrose), and mitochondria were isolated by buoyant density centrifugation on OptiPrep (Sigma, D1556) gradients as published previously (17). For tissue lysate preparations, an aliquot of homogenate was retained, adjusted to 1% v/v SDS and 1% v/v Nonidet P-40, and incubated for 10 min on ice and 10 min at ambient temperature. Extracts were mildly sonicated three times for 30 s with 1 min icing between pulses with a Branson 2510R-DTH bath sonicator and cleared by centrifugation at 13,500 × g for 10 min at 4 °C.
Protein extraction, quantification, and Western blotting
Unless otherwise stated, whole-cell lysates were prepared by extraction in RIPA buffer (50 mm Tris, pH 7.4, 150 mm NaCl, 1% v/v Triton X-100, 0.1% v/v SDS, 1% v/v sodium deoxycholate) with protease inhibitors (10 μg/ml leupeptin, 10 μg/ml pepstatin A, 10 μg/ml chymotrypsin) for 10 min on ice and 10 min at ambient temperature. Supernatants were recovered by centrifugation at 13,500 × g for 5 min at 4 °C. Protein quantifications were performed with a BCA protein assay (Thermo Fisher Scientific, 23227). Samples were boiled in 1× Laemmli buffer (60 mm Tris-HCl, pH 6.8, 2% w/v SDS, 10% v/v glycerol, 0.025% w/v bromphenol blue, 100 mm DTT) at 98 °C for 5 min and separated on 12.5% Tris-glycine polyacrylamide gels. Gels were run at 100 V in running buffer (25 mm Tris, 192 mm glycine, 0.1% w/v SDS) and wet-transferred at 60 V for 1 h at 4 °C in transfer buffer (25 mm Tris, 190 mm glycine, 20% v/v methanol) onto 0.45-μm nitrocellulose membranes (Bio-Rad, 1620155). Membranes were blocked in 5% milk/PBS-T (5% w/v Carnation® instant skim milk powder in PBS, pH 7.4, 0.1% v/v Tween 20) for at least 30 min at ambient temperature. Primary antibodies were immunoblotted in 5% milk/PBS-T for 2 h at ambient temperature or overnight at 4 °C. For antibody pre-absorption with blocking peptide (Fig. S5), anti-TRAF6 (EP591Y) was diluted at 0.412 μg/ml in 5% milk/PBS-T, and blocking peptide was added at 2-fold excess of the antibody, and the solution was rotated end–over–end at ambient temperature for 1 h. HRP-coupled secondary antibodies were bound for 2 h at ambient temperature. After primary and secondary antibody incubation, the membranes were washed three times with PBS-T. For all experiments, signals were captured on CL-XPosureTM radiography films (Thermo Fisher Scientific, 34090) using ECL Western blotting substrate (Pierce, 32106), with the exception of Fig. 3A, which was captured on the Bio-Rad ChemiDoc system.
Misfolded SOD1 co-immunoprecipitation from mitochondria and MS
Rabbit anti-AMF7-63 (concentration 1.54 μg/μl) was coupled to protein A Dynabeads (Invitrogen, 10002D) at full saturation according to bead-binding capacity given by the manufacturer (6.15 μl per 40 μl of beads). Mouse anti-B8H10 (concentration not provided by the manufacturer) was coupled to protein G Dynabeads (Invitrogen, 10004D) at an equivalent volume to AMF7-63. Beads coupled to isotype-matched ChromPure whole molecule IgGs (Jackson ImmunoResearch, 015-000-003 or 011-000-003) at full saturation were used as control. B8H10 and AMF7-63-coupled beads were cross-linked with BS3 (Thermo Fisher Scientific, 21580), according to the manufacturer's instructions. Beads were then tumbled with 150 μg of spinal cord mitochondria isolated from symptomatic SOD1G93A rats at 1 μg/μl final protein concentration in co-immunoprecipitation (IP) buffer (50 mm Tris-HCl, pH 7.4, 150 mm NaCl, 1 mm EDTA, 0.5% v/v Nonidet P-40) overnight at 4 °C. The antigen-bound beads were washed three times with co-IP buffer and snap-frozen in liquid nitrogen. For MS analysis, all samples were reduced on-bead with 5 mm DTT for 10 min at 95 °C, alkylated with 5 mm iodoacetamide for 1 h at ambient temperature, and trypsin-digested on-column for 18 h at 37 °C. Tryptic peptides were subjected to nanoLC-MS/MS analysis using a NanoAcquity UPLC (Waters) coupled to ESI-LTQ-XL-ETD mass spectrometer (Thermo Fisher Scientific), as published previously (92). The .raw files were used to generate .mgf files using ProteoWizard (version 3.0.18250) and submitted to the Matrix Science Mascot search engine (version 2.6.2) to search against a rat Uniprot database (2019-02 release, number of entries 36,064), with human SOD1 sequence added (UniProt ID: P00441), and a decoy database (same database but randomized). Searches were performed with a specified trypsin enzymatic cleavage with one missed cleavage allowed and a 0.6-Da mass tolerance for precursor and fragment ions. Allowed modifications included carbamidomethyl at cysteines (+57 Da) as fixed and oxidation at methionines (+16 Da) as variable. Peptides showing Mascot scores ≥40 show an overall false discovery rate less than 2.5% (decoy/target hits) and were determined as positive hits. The MS proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD01527 (93).
Cell culture
HAP1 cells edited by CRISPR/Cas9 containing a 10-bp deletion in the first coding exon of TRAF6 (referred to as TRAF6-KO) and HAP1 parental cells (referred to as TRAF6-WT) (Horizon Discovery, HZGHC003466c011) were cultured in Iscove's modified Dulbecco's medium (Thermo Fisher Scientific, 12440) supplemented with 10% fetal bovine serum (Wisent Bioproducts, 080150). 293FT cells (human embryonic kidney) were cultured in Dulbecco's high-glucose modified Eagle's medium (GE Healthcare, SH30081.01) supplemented with 10% fetal bovine serum and 2 mm l-glutamine (Sigma, G7513). Primary mouse cortical neurons and astrocytes cultures were performed as described previously (94, 95). For iPSC-derived motor neurons, motor neuron progenitors were generated from reprogrammed human fibroblast lines SOD1A4V (Coriell, ND35671*C), SOD1V148G (Coriell, ND35670*D), and healthy control line NCRM-1 (National Institutes of Health) using a monolayer approach and a mixture of small molecules optimized from published protocols (96). Motor neuron progenitors were then differentiated into motor neurons (50% for Dulbecco's modified Eagle's medium/F-12 and 50% Neurobasal medium, supplemented with N2, B27, l-Glutamax, antibiotic–antimycotic, 100 μm ascorbic acid, 0.5 μm retinoic acid, 0.1 μm purmorphamine, 0.1 μm compound E, 10 ng/ml brain-derived neurotrophic factor, 10 ng/ml ciliary neurotrophic factor, 10 ng/ml IGF-1) and cultured for 14 days. All cells were maintained at 37 °C with a humidified atmosphere of 5% CO2. Studies with patient-derived cells abided by the Declaration of Helsinki principles and were conducted in accordance with approved protocols from the CRCHUM Institutional Review Board.
Plasmids and cloning
pCMV–Myc–TRAF6WT was a gift from Dr. Douglas Leaman (Wright State University, OH) and has been described previously (97). pCMV–Myc–TRAF6WT was used as a template to generate the deletion constructs TRAF6ΔC (comprising aa. 1–288) and TRAF6ΔN (comprising aa. 289–522) by classic restriction-based cloning into EcoRI and HindIII sites. pCMV–Myc–TRAF6C70A was generated from pCMV–Myc–TRAF6WT by site-directed mutagenesis with the QuikChange II kit (Agilent, 200524) according to the manufacturer's instructions. pCMV empty vector was used as a control. pcDNA3–Flag–SOD1WT, SOD1G85R, and SOD1G93A were gifts from Dr. Jean-Pierre Julien (Université Laval, Quebec, Canada). pcDNA3–Flag–SOD1WT was used as a template to generate pcDNA3–Flag–SOD1A4V, SOD1G37R, SOD1H46R, SOD1E100G, SOD1V148G, and SOD1V148I by site-directed mutagenesis. pcDNA3–Flag–SOD1G127X was subcloned from pCI-neo-SOD1G127X (Gly-127insTGGG), a gift from Dr. Don Cleveland (University of California San Diego). pCI–Flag empty vector was used as control. pRK5–HA–ubiquitinWT and single-lysine mutants were gifts from Dr. Edward Fon (McGill University, Quebec, Canada) and are commercially available (Addgene, UbWT catalog numbers 17608, UbK6 22900; UbK11 22901; UbK27 22902; UbK29 22903; UbK33 17607; UbK48 17605; and UbK63 17606). pCMV–Flag–p65, pRL–null, and pGL3–P2(2×)TK for NF-κB luciferase assays have been published previously (98). All newly-cloned plasmids were verified by Sanger sequencing (Genome Quebec). Cloning primers are listed in Table S5.
Transfections
400 ng per expression plasmid or respective empty vector (referred to as mock) were transfected into 293FT cells with Lipofectamine LTX with Plus reagent (Invitrogen, 15338030) according to the manufacturer's instructions in Opti-MEM Reduced Serum Medium (Thermo Fisher Scientific, 22600050). Normal growth medium was replaced after 3 h, and cells were collected after 24 h. For siRNA delivery (150 pmol), 293FT cells were transfected with Lipofectamine 2000 (Invitrogen, 11668027) according to the manufacturer's instructions. Normal growth medium was replaced after 5 h, and cells were collected after 72 h. If cDNA expression plasmids were introduced into siRNA-treated cells, this was 24 h prior to cell collection. For NF-κB luciferase assays, 293FT cells were co-transfected with pGL3–P2(2X)TK (25 ng, firefly luciferase NF-κB reporter), pRL-null (50 ng, Renilla luciferase internal control), p65–Flag (0.5 ng, positive control) or Myc–TRAF6 (20 ng), and an escalating dose of Flag–SOD1WT or Flag–SOD1A4V (5–80 ng) with Lipofectamine LTX. Transfections were balanced with pCI-neo-Flag empty vector.
In cellulo protein–protein interaction assays
293FT cells were transfected with the indicated plasmid combinations. Cell pellets were resuspended in PBS (137 mm NaCl, 2.7 mm KCl, 8 mm Na2HPO4, 1.5 mm KH2PO4, pH7.4) with protease inhibitors, and lysates were prepared by freeze–thaw extraction (three cycles of freezing on dry ice and snap thawing in a 37 °C water bath). The supernatants were recovered after centrifugation at 13,500 × g for 5 min at 4 °C. Lysates were adjusted with 2× IP-buffer (100 mm Tris-HCl, pH 7.4, 300 mm NaCl, 2 mm EDTA, 1% v/v Nonidet P-40, with protease inhibitors) to 1.5× final concentration and tumbled overnight at 4 °C with protein G Dynabeads coupled to mouse anti-Flag or rabbit anti-Myc. Antigen-bound beads were washed three times with 1× IP-buffer and eluted by boiling at 98 °C for 5 min in 2.5× Laemmli buffer. The eluates were analyzed by Western blotting as described.
In cellulo ubiquitination assays
293FT cells were transfected with the indicated plasmid combinations. Whole-cell lysates were prepared by extraction in RIPA buffer for 15 min on ice and were sonicated three times for 30 s with 1 min icing between pulses with a Branson 2510R-DTH bath sonicator. Extracts were cleared by centrifugation at 13,500 × g for 10 min at 4 °C. Protein G Dynabeads were coupled to rat anti-HA, and 400 μg of cell lysate were immunoprecipitated as described.
Filter retardation assays
293FT cells were transfected with the indicated plasmid combinations as described. Filter retardation assays were performed along previously published methods with slight modifications (99). Cells were lysed by freeze–thaw extraction in PBS with protease inhibitors. The DNA condensate was manually removed with a pipette tip, and lysates were additionally digested with 0.35 Kunitz units of DNase I (Roche Applied Science, 11284932001) per μl of extraction buffer for 30 min at 37 °C. Extracts were cleared by low-speed centrifugation at 800 × g for 10 min at 4 °C. Supernatants were recovered and quantified, and samples of 60 μg in a total volume of 150 μl of PBS were loaded onto cellulose acetate membrane filters with 0.2-μm pore size (Whatman, 10404131) using the Bio-Dot® microfiltration apparatus (Bio-Rad, 1706545). After 1 h of passive filtration by gravity flow, the wells were vacated and washed three times with 200 μl of PBS by gentle vacuum application. Filters were removed from the apparatus, blocked in 5% milk/PBS-T for 1 h, and immunoblotted as described.
NF-κB luciferase assays
NF-κB luciferase assays were performed with the Dual-Luciferase® reporter assay system (Promega, E1960) according to the manufacturer's instructions and as published previously (100). Briefly, 293FT cells were transfected with the indicated plasmid combinations in quadruplicate conditions. Luciferase luminescence was measured with a BioTek Synergy HT multimode microplate reader equipped with an automated substrate injection system. Data are expressed as the ratio of mean luminescence ± S.D. of firefly luciferase/Renilla luciferase. For SDS-PAGE, quadruplicate samples were pooled, and 30 μl were loaded.
qRT-PCR
Human iPSC-derived motor neurons were differentiated for 14 days, and total RNA was extracted with the miRNeasy mini kit (Qiagen, 217004) according to the manufacturer's instructions. Reverse transcriptions were performed on 400 ng of total RNA extract in a 4-μl total volume, using random primers (Invitrogen, 48190011), and Moloney murine leukemia virus reverse transcriptase (Invitrogen, 28025013), according to the manufacturer's instructions. Reactions were conducted in singleplex, in a 10-μl total volume containing 1× TaqMan Fast Advanced Master Mix (Applied Biosystems, 4444556), 1× TaqMan primers/probe set (Thermo Fisher Scientific), and 1 μl of diluted cDNA. qRT-PCRs were performed on a QuantStudio 3 real-time PCR system (Thermo Fisher Scientific). Probes were from Applied Biosystems: ACTB (Hs01060665_g1); CHAT (Hs00758143_m1); GAPDH (Hs02786624_g1); HB9 (Hs00907365_m1); and SOX1 (Hs01057642_s1). The data were analyzed using the ΔCT method (101).
Software and statistics
Enrichr was used to perform gene ontology (BP_2018, MF_2018, and CC_2018) and pathway enrichment (KEGG_2019_Human) analyses (21, 22). Protein sequence alignments were generated with MultAlin (102). Three-dimensional protein structure models were generated using iTasser (Iterative Threading Assembly Refinement) (103). The full-length model was constrained with Protein Data Bank code 3HCS.A, without alignment. No constraints were used for the deletion models. The highest C score models were chosen for display. Densitometry was performed in Adobe Photoshop CS4, and graphs were generated with GraphPad Prism 8. Statistics were performed in GraphPad Prism 8 (one-way analysis of variance with Dunnett's multiple comparisons or t test, where appropriate).
Author contributions
S. S., S. R. P., E. M. M., and C. V. V. conceptualization; S. S. and C. V. V. data curation; S. S., P. G., A. N. B., J.-F. T., A. S. H., S. S. P., and C. V. V. formal analysis; S. S. and C. V. V. funding acquisition; S. S., M. G., S. R. P., C. B., E. H.-K., L. D., Y. K., M. C., and E. C. investigation; S. S., P. G., A. T. S., A. S. H., N. G., and S. S. P. methodology; S. S. and C. V. V. writing-original draft; S. S., M. G., P. G., S. R. P., C. B., E. H.-K., L. D., Y. K., M. C., E. C., A. N. B., J.-F. T., N. R. C., A. T. S., A. S. H., T. M. D., E. M. M., J. R., N. G., S. S. P., H. M. M., and C. V. V. writing-review and editing; N. R. C., T. M. D., and J. R. resources; S. S. P. software; H. M. M. and C. V. V. supervision; C. V. V. project administration.
Supplementary Material
Acknowledgments
We thank Don Cleveland (University of California San Diego), Heather Durham (McGill University), Edward Fon (McGill University), Jean-Pierre Julien (Laval University), Douglas Leaman (Wright State University), Giovanni Manfredi (Cornell University), Diana Matheoud (University of Montreal), and Timothy Miller (Washington University) for contributing reagents and/or for helpful discussions throughout the project.
This work was supported by grants from the ALS Society of Canada (to C. V. V.), the Muscular Dystrophy Association (to C. V. V.), the Frick Foundation for ALS Research (to C. V. V.), and the Rare Disease Foundation (to S. S. and C. V. V.). The authors declare that they have no conflicts of interest with the contents of this article.
The MS proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier accession no. PXD015279.
This article contains Figs. S1–S7 and Tables S1–S5.
P. Garg, S. Semmler, C. Vande Velde, and S. S. Plotkin, manuscript in preparation.
- ALS
- amyotrophic lateral sclerosis
- aa.
- amino acids
- CCS
- copper chaperone for SOD1
- fALS
- familial ALS
- GO
- gene ontology
- IP
- immunoprecipitation
- IP-MS
- mass spectrometry-based immunoprecipitation proteomics
- MATH
- meprin and TRAF homology domain
- KEGG
- Kyoto Encyclopedia of Genes and Genomes
- NF-κB
- nuclear factor-κB
- RING
- really interesting new gene domain
- RPS27A
- ubiquitin-40S ribosomal protein S27A
- SOD1
- superoxide dismutase 1
- TRAF6
- TNF receptor–associated factor 6
- RL40
- ubiquitin-60S ribosomal protein L40
- UBB
- polyubiquitin-B
- UBC
- polyubiquitin-C
- VDAC1
- voltage-gated anion channel 1
- WCL
- whole-cell lysate
- Ub
- ubiquitin
- iPSC
- induced pluripotent stem cell
- HRP
- horseradish peroxidase
- GAPDH
- glyceraldehyde-3-phosphate dehydrogenase
- OGDH
- 2-oxoglutarate dehydrogenase
- CNS
- central nervous system
- qRT
- quantitative RT.
References
- 1. Rosen D. R., Siddique T., Patterson D., Figlewicz D. A., Sapp P., Hentati A., Donaldson D., Goto J., O'Regan J. P., and Deng H. X. (1993) Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 362, 59–62 10.1038/362059a0 [DOI] [PubMed] [Google Scholar]
- 2. Renton A. E., Chiò A., and Traynor B. J. (2014) State of play in amyotrophic lateral sclerosis genetics. Nat. Neurosci. 17, 17–23 10.1038/nn.3584 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Gaudette M., Hirano M., and Siddique T. (2000) Current status of SOD1 mutations in familial amyotrophic lateral sclerosis. Amyotroph. Lateral Scler. Other Motor Neuron Disord. 1, 83–89 10.1080/14660820050515377 [DOI] [PubMed] [Google Scholar]
- 4. Pickles S., and Vande Velde C. (2012) Misfolded SOD1 and ALS: zeroing in on mitochondria. Amyotroph. Lateral Scler. 13, 333–340 10.3109/17482968.2012.648645 [DOI] [PubMed] [Google Scholar]
- 5. Peng X., Cashman N. R., and Plotkin S. S. (2018) Prediction of misfolding-specific epitopes in SOD1 using collective coordinates. J. Phys. Chem. B 122, 11662–11676 10.1021/acs.jpcb.8b07680 [DOI] [PubMed] [Google Scholar]
- 6. Patel P., Kriz J., Gravel M., Soucy G., Bareil C., Gravel C., and Julien J. P. (2014) Adeno-associated virus-mediated delivery of a recombinant single-chain antibody against misfolded superoxide dismutase for treatment of amyotrophic lateral sclerosis. Mol. Ther. 22, 498–510 10.1038/mt.2013.239 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Roberts B. R., Lim N. K., McAllum E. J., Donnelly P. S., Hare D. J., Doble P. A., Turner B. J., Price K. A., Lim S. C., Paterson B. M., Hickey J. L., Rhoads T. W., Williams J. R., Kanninen K. M., Hung L. W., et al. (2014) Oral treatment with Cu(II)(atsm) increases mutant SOD1 in vivo but protects motor neurons and improves the phenotype of a transgenic mouse model of amyotrophic lateral sclerosis. J. Neurosci. 34, 8021–8031 10.1523/JNEUROSCI.4196-13.2014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Gros-Louis F., Soucy G., Larivière R., and Julien J. P. (2010) Intracerebroventricular infusion of monoclonal antibody or its derived Fab fragment against misfolded forms of SOD1 mutant delays mortality in a mouse model of ALS. J. Neurochem. 113, 1188–1199 10.1111/j.1471-4159.2010.06683.x [DOI] [PubMed] [Google Scholar]
- 9. Atlasi R. S., Malik R., Corrales C. I., Tzeplaeff L., Whitelegge J. P., Cashman N. R., and Bitan G. (2018) Investigation of anti-SOD1 antibodies yields new structural insight into SOD1 misfolding and surprising behavior of the antibodies themselves. ACS Chem. Biol. 13, 2794–2807 10.1021/acschembio.8b00729 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Pickles S., Semmler S., Broom H. R., Destroismaisons L., Legroux L., Arbour N., Meiering E., Cashman N. R., and Vande Velde C. (2016) ALS-linked misfolded SOD1 species have divergent impacts on mitochondria. Acta Neuropathol. Commun. 4, 43 10.1186/s40478-016-0313-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Liu H. N., Tjostheim S., Dasilva K., Taylor D., Zhao B., Rakhit R., Brown M., Chakrabartty A., McLaurin J., and Robertson J. (2012) Targeting of monomer/misfolded SOD1 as a therapeutic strategy for amyotrophic lateral sclerosis. J Neurosci. 32, 8791–8799 10.1523/JNEUROSCI.5053-11.2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Takeuchi S., Fujiwara N., Ido A., Oono M., Takeuchi Y., Tateno M., Suzuki K., Takahashi R., Tooyama I., Taniguchi N., Julien J. P., and Urushitani M. (2010) Induction of protective immunity by vaccination with wild-type apo superoxide dismutase 1 in mutant SOD1 transgenic mice. J. Neuropathol. Exp. Neurol. 69, 1044–1056 10.1097/NEN.0b013e3181f4a90a [DOI] [PubMed] [Google Scholar]
- 13. Urushitani M., Ezzi S. A., and Julien J. P. (2007) Therapeutic effects of immunization with mutant superoxide dismutase in mice models of amyotrophic lateral sclerosis. Proc. Natl. Acad. Sci. U.S.A. 104, 2495–2500 10.1073/pnas.0606201104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Zhao B., Marciniuk K., Gibbs E., Yousefi M., Napper S., and Cashman N. R. (2019) Therapeutic vaccines for amyotrophic lateral sclerosis directed against disease specific epitopes of superoxide dismutase 1. Vaccine 37, 4920–4927 10.1016/j.vaccine.2019.07.044 [DOI] [PubMed] [Google Scholar]
- 15. Maier M., Welt T., Wirth F., Montrasio F., Preisig D., McAfoose J., Vieira F. G., Kulic L., Späni C., Stehle T., Perrin S., Weber M., Hock C., Nitsch R. M., and Grimm J. (2018) A human-derived antibody targets misfolded SOD1 and ameliorates motor symptoms in mouse models of amyotrophic lateral sclerosis. Sci. Transl. Med. 10, eaah3924 10.1126/scitranslmed.aah3924 [DOI] [PubMed] [Google Scholar]
- 16. Pickles S., Destroismaisons L., Peyrard S. L., Cadot S., Rouleau G. A., Brown R. H. Jr., Julien J. P., Arbour N., and Vande Velde C. (2013) Mitochondrial damage revealed by immunoselection for ALS-linked misfolded SOD1. Hum. Mol. Genet. 22, 3947–3959 10.1093/hmg/ddt249 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Vande Velde C., Miller T. M., Cashman N. R., and Cleveland D. W. (2008) Selective association of misfolded ALS-linked mutant SOD1 with the cytoplasmic face of mitochondria. Proc. Natl. Acad. Sci. U.S.A. 105, 4022–4027 10.1073/pnas.0712209105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Liu J., Lillo C., Jonsson P. A., Vande Velde C., Ward C. M., Miller T. M., Subramaniam J. R., Rothstein J. D., Marklund S., Andersen P. M., Brännstrom T., Gredal O., Wong P. C., Williams D. S., and Cleveland D. W. (2004) Toxicity of familial ALS-linked SOD1 mutants from selective recruitment to spinal mitochondria. Neuron 43, 5–17 10.1016/j.neuron.2004.06.016 [DOI] [PubMed] [Google Scholar]
- 19. Casareno R. L., Waggoner D., and Gitlin J. D. (1998) The copper chaperone CCS directly interacts with copper/zinc superoxide dismutase. J. Biol. Chem. 273, 23625–23628 10.1074/jbc.273.37.23625 [DOI] [PubMed] [Google Scholar]
- 20. Israelson A., Arbel N., Da Cruz S., Ilieva H., Yamanaka K., Shoshan-Barmatz V., and Cleveland D. W. (2010) Misfolded mutant SOD1 directly inhibits VDAC1 conductance in a mouse model of inherited ALS. Neuron 67, 575–587 10.1016/j.neuron.2010.07.019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Kuleshov M. V., Jones M. R., Rouillard A. D., Fernandez N. F., Duan Q., Wang Z., Koplev S., Jenkins S. L., Jagodnik K. M., Lachmann A., McDermott M. G., Monteiro C. D., Gundersen G. W., and Ma'ayan A. (2016) Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 10.1093/nar/gkw377 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Chen E. Y., Tan C. M., Kou Y., Duan Q., Wang Z., Meirelles G. V., Clark N. R., and Ma'ayan A. (2013) Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 10.1186/1471-2105-14-128 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Walsh M. C., Lee J., and Choi Y. (2015) Tumor necrosis factor receptor-associated factor 6 (TRAF6) regulation of development, function, and homeostasis of the immune system. Immunol. Rev. 266, 72–92 10.1111/imr.12302 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Inoue Ji., Ishida T., Tsukamoto N., Kobayashi N., Naito A., Azuma S., and Yamamoto T. (2000) Tumor necrosis factor receptor-associated factor (TRAF) family: adapter proteins that mediate cytokine signaling. Exp. Cell Res. 254, 14–24 10.1006/excr.1999.4733 [DOI] [PubMed] [Google Scholar]
- 25. Nakhaei P., Genin P., Civas A., and Hiscott J. (2009) RIG-I-like receptors: sensing and responding to RNA virus infection. Semin. Immunol. 21, 215–222 10.1016/j.smim.2009.05.001 [DOI] [PubMed] [Google Scholar]
- 26. Bradley J. R., and Pober J. S. (2001) Tumor necrosis factor receptor-associated factors (TRAFs). Oncogene 20, 6482–6491 10.1038/sj.onc.1204788 [DOI] [PubMed] [Google Scholar]
- 27. Shi C. S., and Kehrl J. H. (2010) TRAF6 and A20 regulate lysine 63-linked ubiquitination of Beclin-1 to control TLR4-induced autophagy. Sci. Signal. 3, ra42 10.1126/scisignal.2000751 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Linares J. F., Duran A., Yajima T., Pasparakis M., Moscat J., and Diaz-Meco M. T. (2013) K63 polyubiquitination and activation of mTOR by the p62–TRAF6 complex in nutrient-activated cells. Mol. Cell 51, 283–296 10.1016/j.molcel.2013.06.020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Nazio F., Strappazzon F., Antonioli M., Bielli P., Cianfanelli V., Bordi M., Gretzmeier C., Dengjel J., Piacentini M., Fimia G. M., and Cecconi F. (2013) mTOR inhibits autophagy by controlling ULK1 ubiquitylation, self-association and function through AMBRA1 and TRAF6. Nat. Cell Biol. 15, 406–416 10.1038/ncb2708 [DOI] [PubMed] [Google Scholar]
- 30. Paul P. K., Bhatnagar S., Mishra V., Srivastava S., Darnay B. G., Choi Y., and Kumar A. (2012) The E3 ubiquitin ligase TRAF6 intercedes in starvation-induced skeletal muscle atrophy through multiple mechanisms. Mol. Cell. Biol. 32, 1248–1259 10.1128/MCB.06351-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. West A. P., Brodsky I. E., Rahner C., Woo D. K., Erdjument-Bromage H., Tempst P., Walsh M. C., Choi Y., Shadel G. S., and Ghosh S. (2011) TLR signalling augments macrophage bactericidal activity through mitochondrial ROS. Nature 472, 476–480 10.1038/nature09973 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Min Y., Wi S. M., Shin D., Chun E., and Lee K. Y. (2017) Peroxiredoxin-6 negatively regulates bactericidal activity and NF-κB activity by interrupting TRAF6-ECSIT complex. Front. Cell. Infect. Microbiol. 7, 94 10.3389/fcimb.2017.00094 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Lee N. R., Ban J., Lee N. J., Yi C. M., Choi J. Y., Kim H., Lee J. K., Seong J., Cho N. H., Jung J. U., and Inn K. S. (2018) Activation of RIG-I-mediated antiviral signaling triggers autophagy through the MAVS–TRAF6–Beclin-1 signaling axis. Front. Immunol. 9, 2096 10.3389/fimmu.2018.02096 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Murata H., Sakaguchi M., Kataoka K., and Huh N. H. (2013) SARM1 and TRAF6 bind to and stabilize PINK1 on depolarized mitochondria. Mol. Biol. Cell 24, 2772–2784 10.1091/mbc.e13-01-0016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Lim G. G., Chua D. S., Basil A. H., Chan H. Y., Chai C., Arumugam T., and Lim K. L. (2015) Cytosolic PTEN-induced putative kinase 1 is stabilized by the NF-κB pathway and promotes non-selective mitophagy. J. Biol. Chem. 290, 16882–16893 10.1074/jbc.M114.622399 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Zucchelli S., Codrich M., Marcuzzi F., Pinto M., Vilotti S., Biagioli M., Ferrer I., and Gustincich S. (2010) TRAF6 promotes atypical ubiquitination of mutant DJ-1 and α-synuclein and is localized to Lewy bodies in sporadic Parkinson's disease brains. Hum. Mol. Genet. 19, 3759–3770 10.1093/hmg/ddq290 [DOI] [PubMed] [Google Scholar]
- 37. Chung J. Y., Park H. R., Lee S. J., Lee S. H., Kim J. S., Jung Y. S., Hwang S. H., Ha N. C., Seol W. G., Lee J., and Park B. J. (2013) Elevated TRAF2/6 expression in Parkinson's disease is caused by the loss of Parkin E3 ligase activity. Lab. Invest. 93, 663–676 10.1038/labinvest.2013.60 [DOI] [PubMed] [Google Scholar]
- 38. Vilotti S., Codrich M., Dal Ferro M., Pinto M., Ferrer I., Collavin L., Gustincich S., and Zucchelli S. (2012) Parkinson's disease DJ-1 L166P alters rRNA biogenesis by exclusion of TTRAP from the nucleolus and sequestration into cytoplasmic aggregates via TRAF6. PLoS One 7, e35051 10.1371/journal.pone.0035051 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Zucchelli S., Marcuzzi F., Codrich M., Agostoni E., Vilotti S., Biagioli M., Pinto M., Carnemolla A., Santoro C., Gustincich S., and Persichetti F. (2011) Tumor necrosis factor receptor–associated factor 6 (TRAF6) associates with huntingtin protein and promotes its atypical ubiquitination to enhance aggregate formation. J. Biol. Chem. 286, 25108–25117 10.1074/jbc.M110.187591 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Yan R., Farrelly S., and McCarthy J. V. (2013) Presenilins are novel substrates for TRAF6-mediated ubiquitination. Cell. Signal. 25, 1769–1779 10.1016/j.cellsig.2013.05.015 [DOI] [PubMed] [Google Scholar]
- 41. Broering T. J., Wang H., Boatright N. K., Wang Y., Baptista K., Shayan G., Garrity K. A., Kayatekin C., Bosco D. A., Matthews C. R., Ambrosino D. M., Xu Z., and Babcock G. J. (2013) Identification of human monoclonal antibodies specific for human SOD1 recognizing distinct epitopes and forms of SOD1. PLoS One 8, e61210 10.1371/journal.pone.0061210 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Babu J. R., Geetha T., and Wooten M. W. (2005) Sequestosome 1/p62 shuttles polyubiquitinated tau for proteasomal degradation. J. Neurochem. 94, 192–203 10.1111/j.1471-4159.2005.03181.x [DOI] [PubMed] [Google Scholar]
- 43. Popovic D., Vucic D., and Dikic I. (2014) Ubiquitination in disease pathogenesis and treatment. Nat. Med. 20, 1242–1253 10.1038/nm.3739 [DOI] [PubMed] [Google Scholar]
- 44. Iconomou M., and Saunders D. N. (2016) Systematic approaches to identify E3 ligase substrates. Biochem. J. 473, 4083–4101 10.1042/BCJ20160719 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Pierce N. W., Kleiger G., Shan S. O., and Deshaies R. J. (2009) Detection of sequential polyubiquitylation on a millisecond timescale. Nature 462, 615–619 10.1038/nature08595 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Coyaud E., Mis M., Laurent E. M., Dunham W. H., Couzens A. L., Robitaille M., Gingras A. C., Angers S., and Raught B. (2015) BioID-based identification of Skp cullin F-box (SCF)β-TrCP1/2 E3 ligase substrates. Mol. Cell. Proteomics 14, 1781–1795 10.1074/mcp.M114.045658 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Wang K. Z., Galson D. L., and Auron P. E. (2010) TRAF6 is autoinhibited by an intramolecular interaction which is counteracted by trans-ubiquitination. J. Cell. Biochem. 110, 763–771 10.1002/jcb.22589 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Ishida T., Mizushima Si., Azuma S., Kobayashi N., Tojo T., Suzuki K., Aizawa S., Watanabe T., Mosialos G., Kieff E., Yamamoto T., and Inoue J. (1996) Identification of TRAF6, a novel tumor necrosis factor receptor-associated factor protein that mediates signaling from an amino-terminal domain of the CD40 cytoplasmic region. J. Biol. Chem. 271, 28745–28748 10.1074/jbc.271.46.28745 [DOI] [PubMed] [Google Scholar]
- 49. Cao Z., Xiong J., Takeuchi M., Kurama T., and Goeddel D. V. (1996) TRAF6 is a signal transducer for interleukin-1. Nature 383, 443–446 10.1038/383443a0 [DOI] [PubMed] [Google Scholar]
- 50. Lamothe B., Campos A. D., Webster W. K., Gopinathan A., Hur L., and Darnay B. G. (2008) The RING domain and first zinc finger of TRAF6 coordinate signaling by interleukin-1, lipopolysaccharide, and RANKL. J. Biol. Chem. 283, 24871–24880 10.1074/jbc.M802749200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Shi Z., Zhang Z., Zhang Z., Wang Y., Li C., Wang X., He F., Sun L., Jiao S., Shi W., and Zhou Z. (2015) Structural insights into mitochondrial antiviral signaling protein (MAVS)-tumor necrosis factor receptor-associated factor 6 (TRAF6) signaling. J. Biol. Chem. 290, 26811–26820 10.1074/jbc.M115.666578 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Geetha T., and Wooten M. W. (2002) Structure and functional properties of the ubiquitin binding protein p62. FEBS Lett. 512, 19–24 10.1016/S0014-5793(02)02286-X [DOI] [PubMed] [Google Scholar]
- 53. Schimmack G., Schorpp K., Kutzner K., Gehring T., Brenke J. K., Hadian K., and Krappmann D. (2017) YOD1/TRAF6 association balances p62-dependent IL-1 signaling to NF-κB. Elife 6, e22416 10.7554/eLife.22416 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Bergh J., Zetterström P., Andersen P. M., Brännström T., Graffmo K. S., Jonsson P. A., Lang L., Danielsson J., Oliveberg M., and Marklund S. L. (2015) Structural and kinetic analysis of protein-aggregate strains in vivo using binary epitope mapping. Proc. Natl. Acad. Sci. U.S.A. 112, 4489–4494 10.1073/pnas.1419228112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Broom H. R., Rumfeldt J. A., Vassall K. A., and Meiering E. M. (2015) Destabilization of the dimer interface is a common consequence of diverse ALS-associated mutations in metal free SOD1. Protein Sci. 24, 2081–2089 10.1002/pro.2803 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Broom H. R., Vassall K. A., Rumfeldt J. A., Doyle C. M., Tong M. S., Bonner J. M., and Meiering E. M. (2016) Combined isothermal titration and differential scanning calorimetry define three-state thermodynamics of fALS-associated mutant Apo SOD1 dimers and an increased population of folded monomer. Biochemistry 55, 519–533 10.1021/acs.biochem.5b01187 [DOI] [PubMed] [Google Scholar]
- 57. Doyle C. M., Rumfeldt J. A., Broom H. R., Sekhar A., Kay L. E., and Meiering E. M. (2016) Concurrent increases and decreases in local stability and conformational heterogeneity in Cu,Zn superoxide dismutase variants revealed by temperature-dependence of amide chemical shifts. Biochemistry 55, 1346–1361 10.1021/acs.biochem.5b01133 [DOI] [PubMed] [Google Scholar]
- 58. Lamothe B., Besse A., Campos A. D., Webster W. K., Wu H., and Darnay B. G. (2007) Site-specific Lys-63–linked tumor necrosis factor receptor-associated factor 6 auto-ubiquitination is a critical determinant of IκB kinase activation. J. Biol. Chem. 282, 4102–4112 10.1074/jbc.M609503200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Fang J., Muto T., Kleppe M., Bolanos L. C., Hueneman K. M., Walker C. S., Sampson L., Wellendorf A. M., Chetal K., Choi K., Salomonis N., Choi Y., Zheng Y., Cancelas J. A., Levine R. L., and Starczynowski D. T. (2018) TRAF6 mediates basal activation of NF-κB necessary for hematopoietic stem cell homeostasis. Cell Rep. 22, 1250–1262 10.1016/j.celrep.2018.01.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Bhakar A. L., Tannis L. L., Zeindler C., Russo M. P., Jobin C., Park D. S., MacPherson S., and Barker P. A. (2002) Constitutive nuclear factor-κB activity is required for central neuron survival. J. Neurosci. 22, 8466–8475 10.1523/JNEUROSCI.22-19-08466.2002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Schmidt-Ullrich R., Mémet S., Lilienbaum A., Feuillard J., Raphaël M., and Israel A. (1996) NF-κB activity in transgenic mice: developmental regulation and tissue specificity. Development 122, 2117–2128 [DOI] [PubMed] [Google Scholar]
- 62. Kaltschmidt B., and Kaltschmidt C. (2009) NF-κB in the nervous system. Cold Spring Harb. Perspect. Biol. 1, a001271 10.1101/cshperspect.a001271 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Frakes A. E., Ferraiuolo L., Haidet-Phillips A. M., Schmelzer L., Braun L., Miranda C. J., Ladner K. J., Bevan A. K., Foust K. D., Godbout J. P., Popovich P. G., Guttridge D. C., and Kaspar B. K. (2014) Microglia induce motor neuron death via the classical NF-κB pathway in amyotrophic lateral sclerosis. Neuron 81, 1009–1023 10.1016/j.neuron.2014.01.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Ouali Alami N., Schurr C., Olde Heuvel F., Tang L., Li Q., Tasdogan A., Kimbara A., Nettekoven M., Ottaviani G., Raposo C., Röver S., Rogers-Evans M., Rothenhäusler B., Ullmer C., Fingerle J., et al. (2018) NF-κB activation in astrocytes drives a stage-specific beneficial neuroimmunological response in ALS. EMBO J. 37, e98697 10.15252/embj.201798697 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Vande Velde C., McDonald K. K., Boukhedimi Y., McAlonis-Downes M., Lobsiger C. S., Bel Hadj S., Zandona A., Julien J. P., Shah S. B., and Cleveland D. W. (2011) Misfolded SOD1 associated with motor neuron mitochondria alters mitochondrial shape and distribution prior to clinical onset. PLoS One 6, e22031 10.1371/journal.pone.0022031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Saxena S., Roselli F., Singh K., Leptien K., Julien J. P., Gros-Louis F., and Caroni P. (2013) Neuroprotection through excitability and mTOR required in ALS motoneurons to delay disease and extend survival. Neuron 80, 80–96 10.1016/j.neuron.2013.07.027 [DOI] [PubMed] [Google Scholar]
- 67. Rakhit R., Robertson J., Vande Velde C., Horne P., Ruth D. M., Griffin J., Cleveland D. W., Cashman N. R., and Chakrabartty A. (2007) An immunological epitope selective for pathological monomer-misfolded SOD1 in ALS. Nat. Med. 13, 754–759 10.1038/nm1559 [DOI] [PubMed] [Google Scholar]
- 68. Carithers L. J., Ardlie K., Barcus M., Branton P. A., Britton A., Buia S. A., Compton C. C., DeLuca D. S., Peter-Demchok J., Gelfand E. T., Guan P., Korzeniewski G. E., Lockhart N. C., Rabiner C. A., Rao A. K., et al. (2015) A novel approach to high-quality postmortem tissue procurement: the GTEx Project. Biopreserv. Biobank. 13, 311–319 10.1089/bio.2015.0032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Ruegsegger C., Maharjan N., Goswami A., Filézac de L'Etang A., Weis J., Troost D., Heller M., Gut H., and Saxena S. (2016) Aberrant association of misfolded SOD1 with Na+/K+ATPase-α3 impairs its activity and contributes to motor neuron vulnerability in ALS. Acta Neuropathol. 131, 427–451 10.1007/s00401-015-1510-4 [DOI] [PubMed] [Google Scholar]
- 70. Zetterström P., Graffmo K. S., Andersen P. M., Brännström T., and Marklund S. L. (2011) Proteins that bind to misfolded mutant superoxide dismutase-1 in spinal cords from transgenic amyotrophic lateral sclerosis (ALS) model mice. J. Biol. Chem. 286, 20130–20136 10.1074/jbc.M111.218842 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Watanabe Y., Morita E., Fukada Y., Doi K., Yasui K., Kitayama M., Nakano T., and Nakashima K. (2008) Adherent monomer-misfolded SOD1. PLoS One 3, e3497 10.1371/journal.pone.0003497 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Jain M. R., Ge W. W., Elkabes S., and Li H. (2008) Amyotrophic lateral sclerosis: protein chaperone dysfunction revealed by proteomic studies of animal models. Proteomics Clin. Appl. 2, 670–684 10.1002/prca.200780023 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Furukawa Y., Torres A. S., and O'Halloran T. V. (2004) Oxygen-induced maturation of SOD1: a key role for disulfide formation by the copper chaperone CCS. EMBO J. 23, 2872–2881 10.1038/sj.emboj.7600276 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Wright G. S., Antonyuk S. V., and Hasnain S. S. (2016) A faulty interaction between SOD1 and hCCS in neurodegenerative disease. Sci. Rep. 6, 27691 10.1038/srep27691 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Subramaniam J. R., Lyons W. E., Liu J., Bartnikas T. B., Rothstein J., Price D. L., Cleveland D. W., Gitlin J. D., and Wong P. C. (2002) Mutant SOD1 causes motor neuron disease independent of copper chaperone-mediated copper loading. Nat. Neurosci. 5, 301–307 10.1038/nn823 [DOI] [PubMed] [Google Scholar]
- 76. Tarr I. S., McCann E. P., Benyamin B., Peters T. J., Twine N. A., Zhang K. Y., Zhao Q., Zhang Z. H., Rowe D. B., Nicholson G. A., Bauer D., Clark S. J., Blair I. P., and Williams K. L. (2019) Monozygotic twins and triplets discordant for amyotrophic lateral sclerosis display differential methylation and gene expression. Sci. Rep. 9, 8254 10.1038/s41598-019-44765-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Matheoud D., Sugiura A., Bellemare-Pelletier A., Laplante A., Rondeau C., Chemali M., Fazel A., Bergeron J. J., Trudeau L. E., Burelle Y., Gagnon E., McBride H. M., and Desjardins M. (2016) Parkinson's disease-related proteins PINK1 and Parkin repress mitochondrial antigen presentation. Cell 166, 314–327 10.1016/j.cell.2016.05.039 [DOI] [PubMed] [Google Scholar]
- 78. Matheoud D., Cannon T., Voisin A., Penttinen A. M., Ramet L., Fahmy A. M., Ducrot C., Laplante A., Bourque M. J., Zhu L., Cayrol R., Le Campion A., McBride H. M., Gruenheid S., Trudeau L. E., and Desjardins M. (2019) Intestinal infection triggers Parkinson's disease-like symptoms in Pink1(−/−) mice. Nature 571, 565–569 10.1038/s41586-019-1405-y [DOI] [PubMed] [Google Scholar]
- 79. Granatiero V., and Manfredi G. (2019) Mitochondrial transport and turnover in the pathogenesis of amyotrophic lateral sclerosis. Biology 8, E36 10.3390/biology8020036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Fujisawa T., Homma K., Yamaguchi N., Kadowaki H., Tsuburaya N., Naguro I., Matsuzawa A., Takeda K., Takahashi Y., Goto J., Tsuji S., Nishitoh H., and Ichijo H. (2012) A novel monoclonal antibody reveals a conformational alteration shared by amyotrophic lateral sclerosis-linked SOD1 mutants. Ann. Neurol. 72, 739–749 10.1002/ana.23668 [DOI] [PubMed] [Google Scholar]
- 81. Andersen P. M., Nilsson P., Keränen M. L., Forsgren L., Hägglund J., Karlsborg M., Ronnevi L. O., Gredal O., and Marklund S. L. (1997) Phenotypic heterogeneity in motor neuron disease patients with CuZn-superoxide dismutase mutations in Scandinavia. Brain 120, 1723–1737 10.1093/brain/120.10.1723 [DOI] [PubMed] [Google Scholar]
- 82. Wang Q., Johnson J. L., Agar N. Y., and Agar J. N. (2008) Protein aggregation and protein instability govern familial amyotrophic lateral sclerosis patient survival. PLoS Biol. 6, e170 10.1371/journal.pbio.0060170 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Strickson S., Emmerich C. H., Goh E. T. H., Zhang J., Kelsall I. R., Macartney T., Hastie C. J., Knebel A., Peggie M., Marchesi F., Arthur J. S. C., and Cohen P. (2017) Roles of the TRAF6 and Pellino E3 ligases in MyD88 and RANKL signaling. Proc. Natl. Acad. Sci. U.S.A. 114, E3481–E3489 10.1073/pnas.1702367114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Tang H. W., Liao H. M., Peng W. H., Lin H. R., Chen C. H., and Chen G. C. (2013) Atg9 interacts with dTRAF2/TRAF6 to regulate oxidative stress-induced JNK activation and autophagy induction. Dev. Cell 27, 489–503 10.1016/j.devcel.2013.10.017 [DOI] [PubMed] [Google Scholar]
- 85. Tenekeci U., Poppe M., Beuerlein K., Buro C., Müller H., Weiser H., Kettner-Buhrow D., Porada K., Newel D., Xu M., Chen Z. J., Busch J., Schmitz M. L., and Kracht M. (2016) K63-ubiquitylation and TRAF6 pathways regulate mammalian P-body formation and mRNA decapping. Mol. Cell 62, 943–957 10.1016/j.molcel.2016.05.017 [DOI] [PubMed] [Google Scholar]
- 86. Fang J., Bolanos L. C., Choi K., Liu X., Christie S., Akunuru S., Kumar R., Wang D., Chen X., Greis K. D., Stoilov P., Filippi M. D., Maciejewski J. P., Garcia-Manero G., Weirauch M. T., et al. (2017) Ubiquitination of hnRNPA1 by TRAF6 links chronic innate immune signaling with myelodysplasia. Nat. Immunol. 18, 236–245 10.1038/ni.3654 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87. Sanz L., Diaz-Meco M. T., Nakano H., and Moscat J. (2000) The atypical PKC-interacting protein p62 channels NF-κB activation by the IL-1-TRAF6 pathway. EMBO J. 19, 1576–1586 10.1093/emboj/19.7.1576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88. Konno H., Yamamoto T., Yamazaki K., Gohda J., Akiyama T., Semba K., Goto H., Kato A., Yujiri T., Imai T., Kawaguchi Y., Su B., Takeuchi O., Akira S., Tsunetsugu-Yokota Y., and Inoue J. (2009) TRAF6 establishes innate immune responses by activating NF-κB and IRF7 upon sensing cytosolic viral RNA and DNA. PLoS One 4, e5674 10.1371/journal.pone.0005674 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89. Howland D. S., Liu J., She Y., Goad B., Maragakis N. J., Kim B., Erickson J., Kulik J., DeVito L., Psaltis G., DeGennaro L. J., Cleveland D. W., and Rothstein J. D. (2002) Focal loss of the glutamate transporter EAAT2 in a transgenic rat model of SOD1 mutant-mediated amyotrophic lateral sclerosis (ALS). Proc. Natl. Acad. Sci. U.S.A. 99, 1604–1609 10.1073/pnas.032539299 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90. Chan P. H., Kawase M., Murakami K., Chen S. F., Li Y., Calagui B., Reola L., Carlson E., and Epstein C. J. (1998) Overexpression of SOD1 in transgenic rats protects vulnerable neurons against ischemic damage after global cerebral ischemia and reperfusion. J. Neurosci. 18, 8292–8299 10.1523/JNEUROSCI.18-20-08292.1998 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91. Pickles S., Arbour N., and Vande Velde C. (2014) Immunodetection of outer membrane proteins by flow cytometry of isolated mitochondria. J. Vis. Exp. 2014, 51887 10.3791/51887 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92. Haqqani A. S., Delaney C. E., Tremblay T. L., Sodja C., Sandhu J. K., and Stanimirovic D. B. (2013) Method for isolation and molecular characterization of extracellular microvesicles released from brain endothelial cells. Fluids Barriers CNS 10, 4 10.1186/2045-8118-10-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93. Vizcaíno J. A., Côté R. G., Csordas A., Dianes J. A., Fabregat A., Foster J. M., Griss J., Alpi E., Birim M., Contell J., O'Kelly G., Schoenegger A., Ovelleiro D., Pérez-Riverol Y., Reisinger F., et al. (2013) The PRoteomics IDEntifications (PRIDE) database and associated tools: status in 2013. Nucleic Acids Res. 41, D1063–D1069 10.1093/nar/gks1262 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94. Khalfallah Y., Kuta R., Grasmuck C., Prat A., Durham H. D., and Vande Velde C. (2018) TDP-43 regulation of stress granule dynamics in neurodegenerative disease-relevant cell types. Sci. Rep. 8, 7551 10.1038/s41598-018-25767-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95. Goslin K., Asmussen H., and Banker G. (1998) in Culturing Nerve Cells (Banker G., and Goslin K. eds) 2nd Ed, pp. 339–370, MIT Press, Cambridge, MA [Google Scholar]
- 96. Du Z. W., Chen H., Liu H., Lu J., Qian K., Huang C. L., Zhong X., Fan F., and Zhang S. C. (2015) Generation and expansion of highly pure motor neuron progenitors from human pluripotent stem cells. Nat. Commun. 6, 6626 10.1038/ncomms7626 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Korchnak A. C., Zhan Y., Aguilar M. T., and Chadee D. N. (2009) Cytokine-induced activation of mixed lineage kinase 3 requires TRAF2 and TRAF6. Cell. Signal. 21, 1620–1625 10.1016/j.cellsig.2009.06.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98. Fink K., Duval A., Martel A., Soucy-Faulkner A., and Grandvaux N. (2008) Dual role of NOX2 in respiratory syncytial virus- and Sendai virus-induced activation of NF-κB in airway epithelial cells. J. Immunol. 180, 6911–6922 10.4049/jimmunol.180.10.6911 [DOI] [PubMed] [Google Scholar]
- 99. Gal J., Chen J., Barnett K. R., Yang L., Brumley E., and Zhu H. (2013) HDAC6 regulates mutant SOD1 aggregation through two SMIR motifs and tubulin acetylation. J. Biol. Chem. 288, 15035–15045 10.1074/jbc.M112.431957 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100. Rengachari S., Groiss S., Devos J. M., Caron E., Grandvaux N., and Panne D. (2018) Structural basis of STAT2 recognition by IRF9 reveals molecular insights into ISGF3 function. Proc. Natl. Acad. Sci. U.S.A. 115, E601–E609 10.1073/pnas.1718426115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101. Schmittgen T. D., and Livak K. J. (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 3, 1101–1108 10.1038/nprot.2008.73 [DOI] [PubMed] [Google Scholar]
- 102. Corpet F. (1988) Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 16, 10881–10890 10.1093/nar/16.22.10881 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103. Yang J., Yan R., Roy A., Xu D., Poisson J., and Zhang Y. (2015) The I-TASSER suite: protein structure and function prediction. Nat. Methods 12, 7–8 10.1038/nmeth.3213 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.