Abstract
Human enterovirus 71 (EV71) is the major pathogen that causes hand, foot and mouth disease that particularly affects young children. Growing hand, foot and mouth disease outbreaks were observed worldwide in recent years and caused devastating losses both economically and politically. However, vaccines or effective drugs are unavailable to date. The genome of EV71 consists of a positive sense, single-stranded RNA of ∼ 7400 bp, encoding a large precursor polyprotein that requires proteolytic processing to generate mature viral proteins. The proteolytic processing mainly depends on EV71 3C protease (3Cpro) that possesses both proteolysis and RNA binding activities, which enable the protease to perform multiple tasks in viral replication and pathogen–host interactions. The central roles played by EV71 3Cpro make it an appealing target for antiviral drug development. We determined the first crystal structure of EV71 3Cpro and analyzed its enzymatic activity. The crystal structure shows that EV71 3Cpro has a typical chymotrypsin-like fold that is common in picornaviral 3Cpro. Strikingly, we found an important surface loop, also denoted as β-ribbon, which adopts a novel open conformation in EV71 3Cpro. We identified two important residues located at the base of the β-ribbon, Gly123 and His133, which form hinges that govern the intrinsic flexibility of the ribbon. Structure-guided mutagenesis studies revealed that the hinge residues are important to EV71 3Cpro proteolytic activities. In summary, our work provides the first structural insight into EV71 3Cpro, including a mobile β-ribbon, which is relevant to the proteolytic mechanism. Our data also provides a framework for structure-guided inhibitor design against EV71 3Cpro.
Abbreviations: EV71, human enterovirus 71; HFMD, hand, foot and mouth disease; 3Cpro, 3C protease; FMDV, foot-and-mouth disease virus; HAV, hepatitis A virus; PV, poliovirus; HRV, human rhinovirus; CVB, coxsackievirus B; ASU, asymmetric unit; SARS-CoV, severe acute respiratory syndrome-coronavirus; WT, wild-type; PDB, Protein Data Bank; SLS, Swiss Light Source
Keywords: chymotrypsin-like fold, β-ribbon, picornaviral 3C, HFMD, crystallography
Graphical Abstract
Research Highlights
► EV71 3Cpro adopts the previously unobserved open β-ribbon conformation. ► The flexibility of the β-ribbon is governed by the hinge residues. ► The mobility of the β-ribbon is relevant to the protease activity of EV71 3Cpro.
Introduction
Human enterovirus 71 (EV71) is the primary pathogen of herpangina or exanthema, also known as hand, foot and mouth disease (HFMD). Children and infants are especially vulnerable due to their underdeveloped immunocompetence.1 Generally, HFMD starts with fever, headache, or vomiting. Lesions and rashes may become evident on patient's hand, foot and mouth in the following days, which give rise to the name of the disease. In severe cases, EV71 infections may lead to neurological damages, including aseptic meningitis, encephalitis, acute flaccid paralysis, myocarditis, pulmonary edema and hemorrhage2 with high incidence of fatality. In recent years, the increasing outbreaks of HFMD have posed the global public health threats. HFMD epidemics were observed in China, Malaysia, Singapore, Australia, the United States and Germany.3, 4, 5, 6, 7, 8 In China, a total number of 192,300 cases were reported from January to April, 2010, an increase of almost 40% up compared to the same period in 2009. From the total number of cases, 2119 cases were diagnosed as severe and 94 cases were fatal.9 An unusual outbreak of HFMD happened in Fuyang city (mainland China) in 2008, one of the most densely populated regions, causing the panic and endangering the social stability in the region. The initial fatality rate of the Fuyang outbreak was 2.9%, and all fatal cases died of serious neurogenic pulmonary edema were due to EV71 infections.10 Unfortunately, effective drug or vaccine against EV71 is currently unavailable.11
EV71 belongs to the genus Enterovirus of the Picornaviridae family. The genome of EV71 consists of a positive sense, single-stranded RNA of ∼ 7400 bp in length,12 encoding a large precursor polyprotein that requires proteolytic processing to produce viral structural and replication proteins. In picornaviruses, except the cleavage between VP1 and 2A (catalyzed by the 2A protease13) and the RNA-dependent cleavage between VP2 and VP4,14 the 3C protease (3Cpro) is absolutely required for all other polyprotein processing during the virus replication. Weng et al. discovered that EV71 3Cpro can interfere with the polyadenylation of host cellular RNA by digesting the host protein CstF-64, a critical factor for 3′ pre-mRNA processing.15 The finding proposed a novel mechanism by which the picornavirus utilizes 3Cpro to impair host cellular functions.
Most picornaviral 3C proteases preferentially cleave polypeptide with Gln–Gly junctions, with the exception of foot-and-mouth disease virus (FMDV) 3Cpro.16 This unusual member of the picornaviral 3Cpro cleaves peptide junctions of P1–Gln or P1–Glu followed by a large apolar amino acid, such as leucine.17, 18, 19 Structural studies on hepatitis A virus (HAV) 3Cpro, human rhinovirus (HRV) 3Cpro, poliovirus (PV) 3Cpro, FMDV 3Cpro and coxsackievirus B (CVB) 3Cpro have shown that this family of cysteine protease shares an overall fold with many serine proteases, such as chymotrypsin, α-lytic protease and so on.17, 20, 21, 22, 23, 24, 25 Picornaviral 3C proteases possess Cys-His-Glu/Asp catalytic triads that share geometry similar with that of the Ser-His-Asp triads found in serine proteases. Surprisingly, in the structure of HAV 3Cpro (with the Cys-His-Asp triad), the aspartic acid is redirected away from the active site. The Tyr143 points toward the catalytic histidine and provides the electrostatic stabilization.26 This observation suggested the possible dyad proteolysis mechanism adopted by HAV 3Cpro. However, recent findings showing that both FMDV 3Cpro and the 3C-like tobacco etch virus protease27 possess the Cys-His-Asp triads with the geometry similar to that of the Ser-His-Asp triad of serine protease, which prompted a reassessment of whether a dyad model is indeed the mechanism used by these family of proteases. Sequence and structural alignments show that the catalytic important motif Gly-X-Cys/Ser-Gly-Gly, serving to position Cys/Ser for nucleophilic attack and to form the oxyanion hole, is structurally conserved in picornaviral 3C proteases and serine proteases. Interestingly, the flexible surface loop β-ribbon is present in all picornaviral 3Cpro, such as FMDV 3Cpro, PV 3Cpro, HRV 3Cpro,CVB3 3Cpro and HAV 3Cpro, and also in several bacteria serine proteases, such as the α-lytic protease,28 Streptomyces griseus protease A29 and S. griseus protease B.30 The β-ribbon, located between the βB2 and βC2 strands, plays an important role in recognizing the P2–P4 region of peptide substrates. Regardless of the differences in length (12–21 aa) and in sequence, the β-ribbons maintain a similar fold and a similar orientation with respect to the substrate binding groove in these proteases.17, 18
Beside the protease activities, picornaviral 3Cpro also has RNA binding activity, implying its role in picornaviral RNA replication.31, 32, 33 By fusing with 3Dpol, the RNA-dependent RNA polymerase that contains a nuclear localization sequence,34 3Cpro can enter nuclei and participate in viral RNA replication. The precursor protein 3CD is a crucial component of the viral replication machinery.35 The PV 3CD functions as a protease better than PV 3Cpro, but lacks polymerase activity.36 The autocatalytic activity enables 3CD to generate 3Cpro and 3Dpol. Collectively, the multifunctional 3Cpro is an important player in many key events on the life cycle of the picornavirus. Therefore, the enzyme is generally considered as an appealing target for antiviral drug design.
In this work, we determined the first crystal structure of EV71 3Cpro and analyzed its protease activities. We found that EV71 3Cpro maintains the typical chymotrypsin fold that is similar to that of other picornaviral 3Cpro. Surprisingly, we found that the β-ribbon of EV71 3Cpro is oriented in an unexpected and previously unobserved open conformation. Our structural analysis revealed two hinge residues located at the base of the β-ribbon. Supported by the mutagenesis studies, we found that the hinge residues likely govern the intrinsic flexibility of the β-ribbon. The mobility of the β-ribbon is relevant to the EV71 3Cpro protease activity. Our results also provide a structural framework for structure-based inhibitor design against EV71 3Cpro.
Results
Crystallization and structure determination
Three recombinant EV71 3Cpro variants (EV71 isolate BJ/CHN/2008) were highly expressed in Escherichia coli with yields of ∼ 2–10 mg of protein per liter of LB culture: N-terminal 6× His-tagged 3Cpro, C-terminal 6× His-tagged 3Cpro and non-tagged 3Cpro. All EV71 3Cpro variants were purified and concentrated to ∼ 12 mg ml− 1 and were subjected to crystallization trials, only the N-terminal 6× His 3Cpro produced protein crystals. The crystals belonged to space group P21, contained five copies of EV71 3Cpro monomer per asymmetric unit (ASU) and diffracted X-ray to a resolution limit of 3.0 Å. The initial phases were obtained by molecular replacement. The phases were further improved by non-crystallographic symmetry averaging and manual model building, especially in the region of 123–133 aa where the novel β-ribbon conformation was observed. The refined model of EV71 3Cpro has R/R free = 0.21/0.26 with good stereochemistry (Table 1 ).
Table 1.
Crystallographic data and model statistics
| Native | |
|---|---|
| Data collection | |
| Space group | P21 |
| Cell dimensions | |
| a, b, c (Å) | 54.667, 75.59, 126.95 |
| α, β, γ (°) | 90, 90.13, 90 |
| Wavelength | 1.00 |
| Resolution (Å) | 3.0 |
| Rsym | 7.7 (38.5) |
| I/σI | 12.18 (2.74) |
| Completeness (%) | 98.0 (92.0) |
| Redundancy | 2.85 |
| Refinement | |
| Resolution range (Å) | 41.5–2.99 |
| No. of reflections | 39,192 |
| Rwork/Rfree | 20.6/25.9 |
| No. of atoms | |
| Protein | 7225 |
| B-factors average | |
| Protein | 82.84 |
| r.m.s.d. | |
| Bond lengths (Å) | 0.008 |
| Bond angles (°) | 1.066 |
Values for the highest-resolution shell are shown in parentheses.
Overall structure
EV71 3Cpro is folded in two topologically equivalent antiparallel six-stranded β-barrel domains (Fig. 1a and b) and is related to other picornaviral 3Cpro structures.17, 20, 22, 37 A long shallow groove for substrate binding is located between the two β-barrel domains. EV71 3Cpro shares ≤ 50% amino acid sequence identity with other picornaviral 3Cpro,17, 20, 24, 25, 37 among which CVB3 3Cpro shares the highest sequence similarity. The structural superimpositions (TOP program) give the r.m.s.d. of 0.87–1.04 Å between EV71 3Cpro (183 aa) and CVB3 3Cpro, HRV 3Cpro and PV 3Cpro for 160 Cα atoms. EV71 3Cpro shares only 24.2% and 29.1% sequence similarities with HAV 3Cpro and FMDV 3Cpro. The structural alignments map 110 Cα atoms of EV71 3Cpro onto structurally equivalent Cα atoms in HAV with r.m.s.d. value of 1.32 Å and map 132 Cα atoms of EV71 3Cpro onto structurally equivalent Cα atoms in FMDV 3Cpro with r.m.s.d. value of 1.40 Å. These data confirm that EV71 3Cpro shares an overall structural similarity with other picornaviral 3Cpro. However, EV71 3Cpro exhibits significant structural differences in the region corresponding to the β-ribbon located between βB2 and βC2 (123–133 aa). In other picornaviral 3Cpro structures, the β-ribbon is located over the substrate binding cleft with its apical tip pointing toward the protease active site. We denote this β-ribbon conformation as “close conformation”. Strikingly, we found that the β-ribbon is flipped open with its tip pointing away from the active site in EV71 3Cpro. We denote this conformation as “open conformation” (Figs. 1a and b and4a and b). In detail, at residue Gly123, the peptide backbone turns in the direction away from the active site, whereas at the corresponding position in the other picornaviral 3Cpro, the peptide backbone turns toward the active site. The opened loop extends for 11 aa and folds back at residue His133. From His133 onward, the backbone runs again in the direction similar to that observed in other picornaviral 3Cpro protease structures (Fig. 4b). Structural alignments of the β-ribbon (123–133 aa) of EV71 3Cpro and the corresponding regions in other picornaviral 3Cpro give r.m.s.d. values of < 2 Å for 11 Cα positions , indicating that the opened β-ribbon still maintains the fold similar to that of the closed β-ribbons. In other words, while the β-ribbon itself remains rigid, the dihedral angles of residues G123 and H133 that are located at the base of the ribbon change dramatically, suggesting that these residues may function as hinges for the open-and-close flipping motion of the β-ribbon. Thus, the nature of the peptide bond restraints of the hinge residues could be the determining factor for the intrinsic flexibility of the β-ribbon.
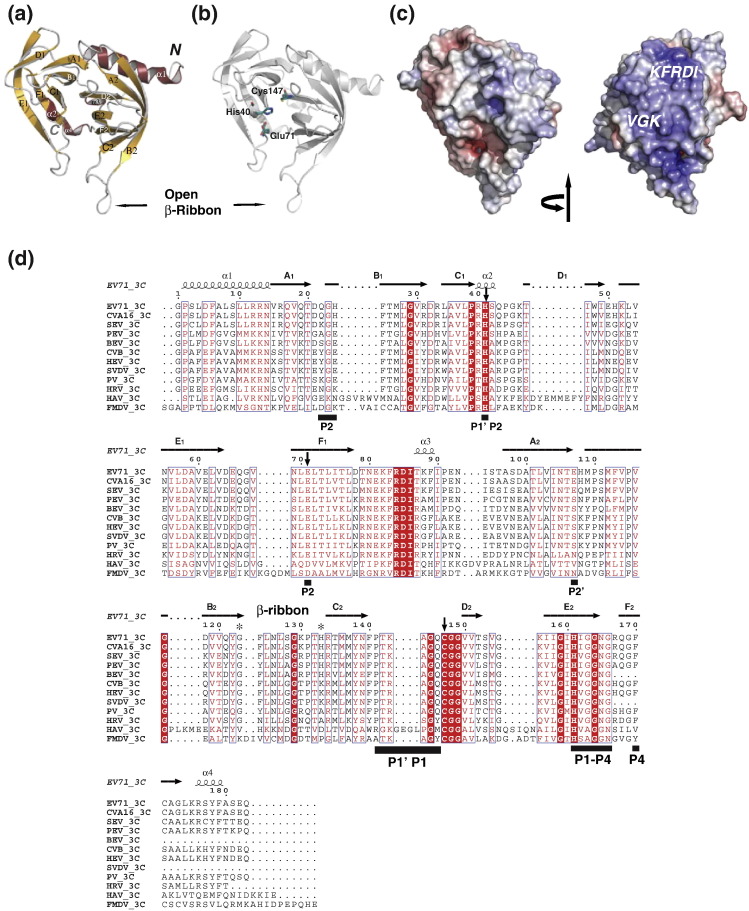
Fig. 1.
Structure of EV71 3Cpro. (a) Ribbon model of EV71 3Cpro with annotated secondary structures. The position of β-ribbon is indicated with black arrow. (b) Ribbon model of EV71 3Cpro (gray) with side chains of catalytic triad, His40, Glu71 and Cys147 (highlighted in stick presentation with deep teal for carbons). The position of β-ribbon is indicated. ⁎There was no electron density for the side chain of residue Glu71. (c) Electrostatic surface potential (ranging from blue = 20 kT/e to red = − 20 kT/e), displayed in two different views (left, “standard view” used in all other figures; right, 180° rotation around the vertical axis). Positions of the conserved RNA binding motifs “KFRDI” and “VGK” are indicated. (d) Structure-based multiple-sequence alignment of 3Cpro of different picornaviruses. The secondary structure is shown on top. Invariant residues in 3Cpro are highlighted with red background; conserved residues are shown in red font. The position of β-ribbon is indicated on the top of the sequences. Hinge residues Gly123 and His133 are indicated by asterisks. Black bars at the bottom of the sequence indicate residues directly involve in substrate binding (based on superimposition with HRV 3C–AG7088 co-structure; PDB code: 1CQQ).
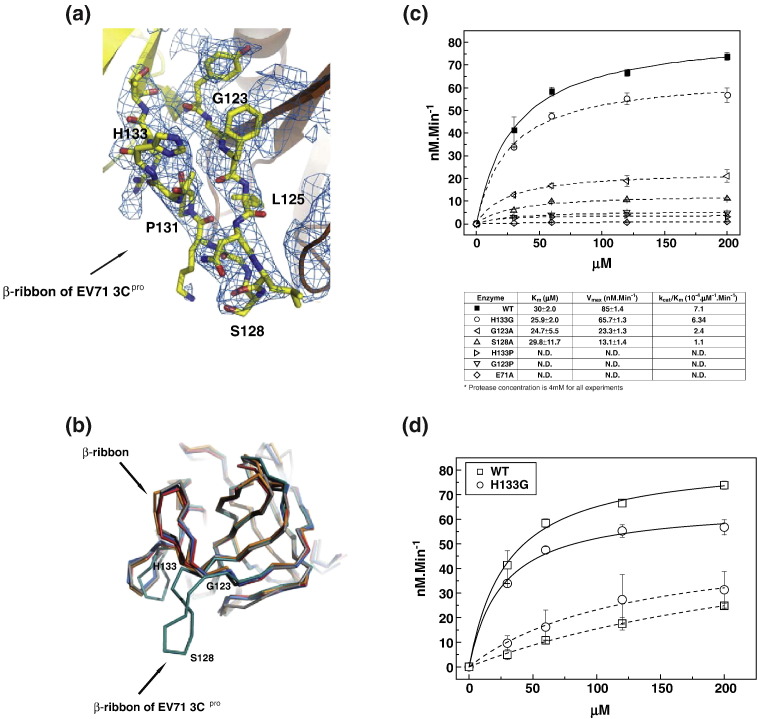
Fig. 4.
The β-ribbon of EV71 3Cpro. (a) Portion of the final 2Fo − Fc electron density (1.0 s contour) at the opened β-ribbon with superimposed final model. (b) Ribbon model (backbone trace) superimpositions of EV71 3Cpro (deep-teal line) and other picornaviruses 3Cpro. Gray line, HRV 3Cpro (PDB code: 1CQQ); blue line, CVB3 3Cpro (PDB code: 2VB0); orange line, PV 3Cpro (PDB code: 1L1N); red line, CVB3 3Cpro (PDB code: 2ZTX). Positions of residues Ser128 sitting on the tip of the β-ribbon and the hinge residues Gly123 and His133 located at the base of the β-ribbon are indicated by the text. (c) Mutagenesis studies reveal the roles of residues in β-ribbon. Proteolytic reactions were performed using WT EV71 3Cpro and EV71 3Cpro mutants as the enzymes and fluorescence-labeled peptide derived from EV71 polyprotein 3B–3C junction as the substrate. Data for WT EV71 3Cpro are fitted with continuous line, and data for EV71 3Cpro mutants are fitted with broken lines. WT EV71 3Cpro (filled squares) serves as positive control; mutant E71A (open diamond) serves as non-active control. Mutants H133P (open right-pointing triangular) and G123P (open down-pointing triangular) are nearly inactive. Whereas, mutant H133G (open circle) is almost as active as WT protease. Mutant G123A (open left-pointing triangular) and mutant S128A (open up-pointing triangular) retain fractions of the enzyme activity. The table below summarizes the values of Km and Vmax and the ratio kcat/Km of the enzymes. (d) Comparison of substrate specificities of WT EV71 3Cpro and mutant H133G. Proteolytic reactions were performed using WT EV71 3Cpro (open square) and EV71 3Cpro mutant H133G (open circle) as the enzymes and fluorescence-labeled peptide derived from the EV71 polyprotein autoprocessing 3B–3C site (continuous lines) and peptide derived from the autoprocessing site in SARS-CoV (broken lines) as the substrates.
Crystal packing
We observed that the opened β-ribbon of EV71 3Cpro is intensely involved in a particular crystal packing interaction. As shown in Fig. 2a (left), five EV71 3Cpro monomers (molecules A–E) within the ASU pack against each other, forming a right-handed helical polymer with the helical axis parallel to the c-axis of the unit cell. Along the c-axis, the helical polymers of the adjacent unit cells are connected, forming the continuous helical polymer, in which the mode of the intermolecular interaction between any adjacent EV71 3Cpro monomers is identical (Fig. 2a, right). In particular, on one side, the N-terminal α1 helix of the lower monomer (molecule D) is packed against the concave surface formed by the βB2, βC2 and βF2 sheets of the upper monomer (molecule E). A hydrogen bond is found between Gly1 of the lower monomer and Gln168 of the upper monomer. Hydrophobic interactions are found between residue Leu4, residue Leu8 on the α1 helix of the lower monomer and residues Val120 and Met137 of the upper monomer (Fig. 2a and b). On the other side, the opened β-ribbon of the upper monomer (molecule E) is packed on the top of the C-terminal β-barrel domain of the lower monomer (molecule D). In detail, three hydrogen bond interactions were observed between the β-ribbon of the upper monomer and the βB2 sheet of the lower monomer (Fig. 2a and c). The Gly123 of the upper EV71 3Cpro monomer is directed way from the position of its counterpart in HRV 3Cpro (Fig. 2c) by a hydrogen bond between the NH group of Gly123 and the carbonyl group of Phe113 of the lower EV71 3Cpro monomer. Two more intermolecular hydrogen bonds are found between Tyr122 of the upper monomer and Phe113 of the lower monomer and between Gln121 of the upper monomer and Ser111 of the lower monomer. These intermolecular hydrogen bonds contribute to maintain the β-ribbon of the upper EV71 3Cpro monomer in the open conformation. The open β-ribbon is further stabilized by hydrophobic interactions between the upper and lower monomers. Residues Leu125 and Pro131 located on the open β-ribbon of the upper EV71 3Cpro monomer form a hydrophobic patch that interact with the hydrophobic surface of the lower EV71 3Cpro monomer comprising of residues Met109, Met112, Val114 and Phe140 (Fig. 2d). In summary, the crystal packing analysis shows that the β-ribbon of the upper EV71 3Cpro monomer is tightly packed on top of the lower EV71 3Cpro monomer by means of a network of hydrogen bonds and hydrophobic interactions. These interactions contribute significantly to the stabilization of the open β-ribbon conformation.
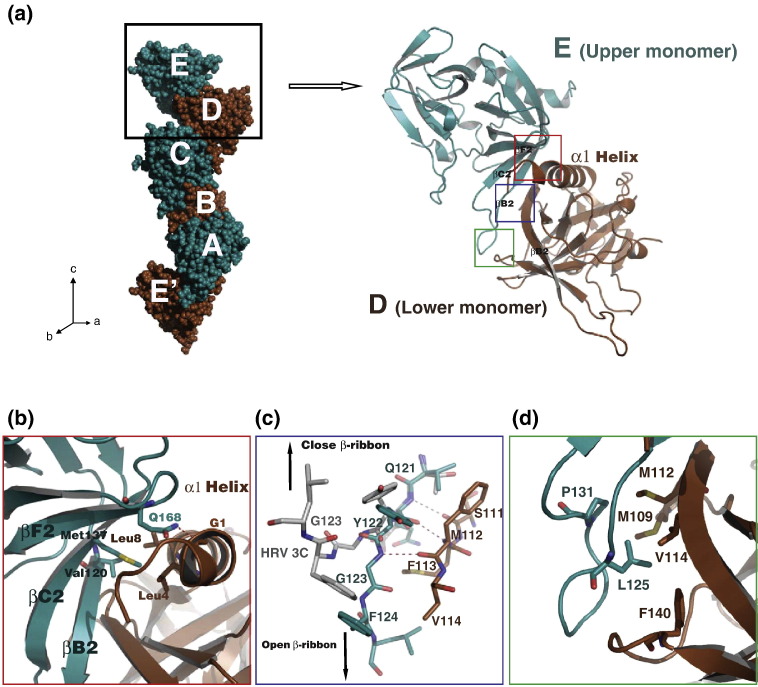
Fig. 2.
Crystal packing stabilizes β-ribbon in open conformation. (a) Left side: five molecules are found in ASU: A, B, C, D and E. Molecule E′ is the corresponding E molecule from the unit cell below. EV71 3Cpro monomers pack against each other, forming the continuous right-handed helical polymers extending alone the c-axis of the unit cell. The intermolecular interaction between any adjacent EV71 3Cpro monomers is identical, which is magnified on the right side. Right side: ribbon model of molecule E/upper monomer (deep teal) and molecular D/lower monomer (brown), showing the intermolecular interaction between the EV71 3Cpro monomers. Detailed crystal packing interactions are magnified in (b) (red box), (c) (blue box) and (d) (green box). (b) The N-terminal portion of the molecule D/lower monomer is packed against the surface of molecule E/upper monomer via hydrophobic interactions and hydrogen bond interaction (red broken line). (c) Stick representations showing the hydrogen-bonding interactions between the β-ribbon of molecule E/upper monomer (carbon atoms in deep teal) and the βB2 sheet of molecule D/lower monomer (carbon atoms in brown). Hydrogen bonds between the monomers are shown in red broken lines. The model of HRV 3Cpro (carbon atoms in gray) is superimposed onto the molecule E/upper monomer. The directions to the tips of the open and close β-ribbons are indicated by arrows. (d) Leu125 and Pro131 located on the open β-ribbon of molecule E/upper monomer (deep teal) form the hydrophobic patch that interacts with the hydrophobic surface of molecule D/lower monomer (brown).
Structure of the active site
In the active site of EV71 3Cpro, the catalytic triads His40, Glu71 and Cys147 have a conformation similar to that observed in other picornaviral 3Cpro (Fig. 1b). However, the active site of EV71 3Cpro is very exposed to the solvent because the β-ribbon is in the open conformation. Poor electron density for the side chain of the catalytic important residue Glu71 did not allow for a definite model, and the residue is therefore associated with very high B-factor (an average B-factor of 121.59 Å2 for Glu71 compare to an average B-factor of 82.84 Å2 for all molecules in ASU ). In order to find out whether or not EV71 3Cpro adopts the catalytic triad proteolytic mechanism, we performed mutagenesis studies. Mutants C147S, H40A and E71A were generated and subjected to protease assays. None of the mutants has detectable protease activities (data not shown), confirming that each member of the catalytic triad is essential for the protease activity of EV71 3Cpro. The catalytic important loop 141–147 aa is partially missing from the electron density map; thus, we could not observe the formation of the oxyanion hole, indicating that this region is disordered.
Structure of the RNA binding site
The KFRDI and VGK motifs were previously characterized as RNA binding motifs in picornaviral 3Cpro 25, 37, 38, 39 and were confirmed as important residues for RNA binding in EV71 3Cpro.31 In our EV71 3Cpro structure, the KFRDI motif is located in a loop connecting the βF1 and the α3 helix. The KFRDI motif forms a basic patch that sits at the opposite face with respect to the protease active site (Fig. 1c). The side chains of apolar residues Phe83 and Ile86 are inserted into the hydrophobic core of the protein, whereas the positively charged side chains of Lys82, Arg84 and Asp85 are exposed to the solvent. Besides Arg84 sitting at the center of the basic patch, the side chains of Arg12, Arg13 and Lys88 are also located at the positively charged surface. The VGK motif is located in the loop between strands βD2 and βE2. The side chain of Val156 is buried in the hydrophobic core, whereas Lys156 has its side chain pointing into the solvent. Based on our structure, more basic residues beside the KFRDI and VGK motifs are found in the putative RNA binding site of EV71 3Cpro, suggesting their involvement in RNA binding. The determination of EV71 3Cpro–RNA co-structure will be important in order to further elucidate the mode of EV71 3Cpro–RNA interaction.
Biochemical characterization of EV71 3Cpro
Size-exclusion chromatography analyses were carried out to determine the oligomerization state of the enzyme. The molecular mass of EV71 3Cpro determined by size-exclusion chromatography is 17 kDa (Fig. 3a, upper-left inset), which is similar to the molecular mass determined by SDS-PAGE (∼ 23 kDa; Fig. 3a, upper-right inset). The data suggest that EV71 3Cpro exists as a monomer in solution, supporting the notion that this class of proteases adopts a monomeric proteolysis mechanism.
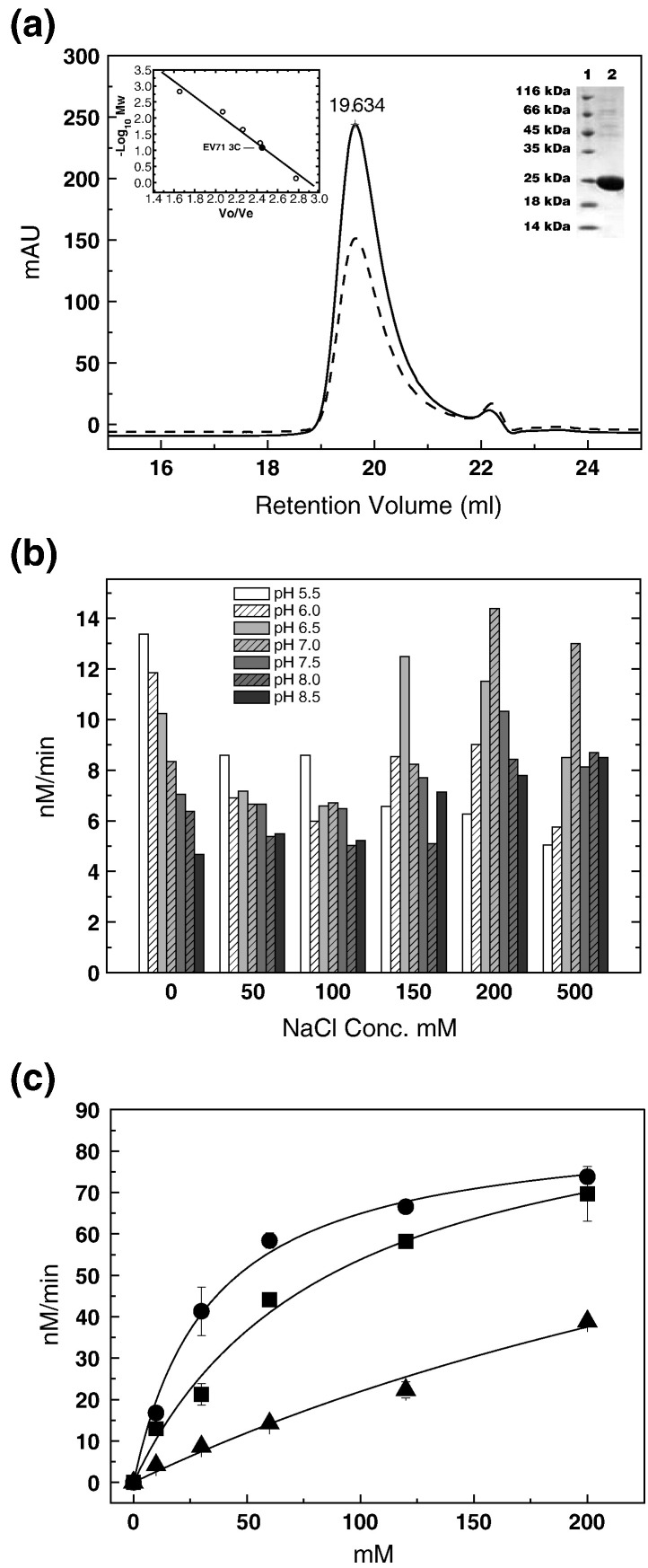
Fig. 3.
Biochemical characterization of EV71 3Cpro. (a) Size-exclusion chromatography and SDS-PAGE analysis of EV71 3Cpro protein. Chromatographic profiles are as follows: continuous line is UV absorbance at 280nm, and broken line is UV absorbance at 260nm. Upper-left inset: the − log (MW) values of standard proteins for size-exclusion column calibration (Superose 6 10/300GL) are plotted as the function of Ve/Vo (○). The data are fitted linearly to derive standard curve. The molecular mass of EV71 3Cpro is estimated at ∼ 17 kDa (●), indicating that EV71 3Cpro exists as monomer in solution. Upper-right inset: SDS-PAGE analysis of purified EV71 3Cpro. Lane 1, molecular mass standards; lane 2, EV71 3Cpro. (b) Optimization of protease assay conditions. The initial velocities of proteolytic reactions are measure in different reaction buffers with various combinations of salt (0–500 mM) and pH (pH 5.5–8.5). The best combination of salt and pH is determined as NaCl 200 mM (pH 7.0). (c) Substrate specificity of EV71 3Cpro. Fluorescence-quenching-pair-based proteolytic reactions are performed using different substrates: (●) substrate derived from the EV71 autoprocessing 3B–3C junction, (▪) substrate derived from the first cleavage site (251/252 aa) in CstF-64 and (▲) substrate derived from the autoprocessing site of a 3C-like protease in SARS-CoV. The concentration of the protease was 4 μM for all experiments.
In order to determine the optimal reaction conditions for EV71 3Cpro protease, we performed grid searches by varying salt concentrations and pH values. As shown in Fig. 3b, the combination of salt concentration and pH value that gives the best proteolytic activity is 200 mM NaCl and pH 7.0. Cations optimization was also performed by replacing sodium with potassium or by including magnesium, calcium, and so on, and only marginal differences were observed. Zinc was the only cation that negatively affected the protease activity of EV71 3Cpro (data not shown). This is likely due to the metal binding property of the catalytic Cys147. The optimized reaction buffer was used throughout the following experiments.
It has been reported that EV71 3Cpro cleaves most efficiently on the 3B–3C junction of EV71 polyprotein.16 Recently, CstF-64 was identified as a novel cellular protein that serves as the target for EV71 3Cpro.15 Therefore, we designed experiments to compare the enzyme activities of EV71 3Cpro against the enzyme activities of the EV71 polyprotein and the host target. The peptide corresponding to the EV71 polyprotein autoprocessing site 3B–3C (with the Q–G junction), the peptide corresponding to the internal cleavage site (251/252 aa with the Q–G junction) of CstF-64 and the peptide derived from the severe acute respiratory syndrome-coronavirus (SARS-CoV) autoprocessing site (with the Q–S junction) were used as the substrates in the protease assays. The protease concentration (4 μM) was kept constant in all protease assays. The kinetic parameters V max and K m were determined to calculate the specificity constant k cat/K m. As shown in Fig. 3c, EV71 3Cpro displayed the best enzyme activity against the autoprocessing 3B–3C site (K m = 30 ± 2 μM, V max = 85 ± 1.4 nM min− 1 and k cat/K m = 7.1 × 10− 4 μM− 1 min− 1). Moderate enzyme activity was measured against the internal cleavage site of CstF-64 (K m = 87 ± 15 μM, V max = 91 ± 7 nM min− 1 and k cat/K m = 2.6 × 10− 4 μM− 1 min− 1), and poor enzyme activity was measured in digesting the autoprocessing site of SARS-CoV (K m and V max could not be precisely determined by Michaelis–Menten equation fitting; thus, the k cat/K m was calculated from the linear part of the rate-versus-substrate concentration plot, k cat/K m = 0.5 ± 0.02 × 10− 4 μM− 1 min− 1). The data suggest that EV71 3Cpro has substrate specificity and enzyme activity against the autoprocessing 3B–3C site greater than against the host target CstF-64 internal cleavage site. Although both substrates have Q–G junctions and P4–Ala residues, different residues at P2, P3, P4 and P2′ are also important for substrate recognition. The autoprocessing site of SARS-CoV with the Q–S junction is not the preferred substrate for EV71 3Cpro.
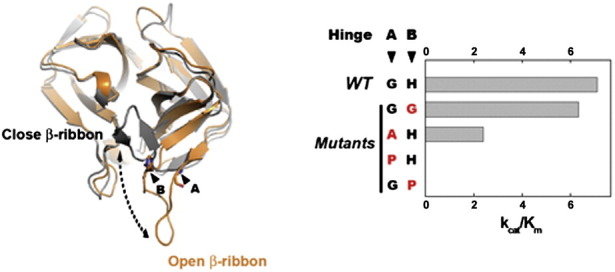
Mutagenesis studies revealed the mobile nature of the β-ribbon
The unusually open β-ribbon conformation is observed in EV71 3Cpro structure; however, in such case, the residues located on the opened β-ribbon cannot be positioned to form parts of S2–S4 substrate binding pockets. In HRV 3Cpro–AG7088 co-structure, the conserved Ser128 located on the tip of the β-ribbon is involved in substrate recognition by forming a hydrogen bond between its side-chain oxygen and backbone NH group of the inhibitor.22 We found that EV71 3Cpro mutant S128A, in which the serine side-chain oxygen is missing, suffered from significant losses of the enzyme activity (Fig. 4c), confirming that the residue is important for substrate recognition. Therefore, we speculate that the EV71 3Cpro must also maintain the close β-ribbon conformation in solution. Supported by the structure of EV71 3Cpro, residues G123 and His133 located at the base of the β-ribbon may function as hinges. Thus, it is possible that the β-ribbon of EV71 3Cpro can undergo open and close motions around the hinges. To test this idea, we introduced a series of amino acid substitutions at the hinge positions. At one hinge, mutant G123P was inactive, whereas mutant G123A recovered a considerable fraction of activities. The K m value for mutant G123A is similar to that of wild-type (WT) protease, but the V max is reduced, indicating that the mutant has a similar substrate binding affinity, but suffers the losses of enzyme turnover rate. At the other hinge, no enzyme activity was observed for mutant H133P, whereas H133G had close-to-WT protease activity (Fig. 4c; K m, V max and k cat/K m values are listed in the table below). In general, when residues with more peptide bond restraints were introduced to the hinge positions, less protease activity was observed. When residues with less peptide bond restraints were introduced to the hinge positions, more protease activity was observed. In other words, the β-ribbon with the relaxed hinge favors the protease activity.
Subsequently, we compared the substrate specificities of mutant H133G (with the relaxed β-ribbon) and WT protease in processing different substrates. As shown in Fig. 4d, the WT protease cleaves the EV71 autoprocessing 3B–3C site with the specificity constant k cat/K m of 7.1 × 10− 4 μM− 1 min− 1 (K m = 30 ± 2 μM; V max = 85 ± 1 nM min− 1); whereas the specificity constant for cleaving the autoprocessing site in SARS-CoV is 0.39 ± 0.02 × 10− 4 μM− 1 min− 1. Mutant H133G cleaves the autoprocessing 3B–3C site with the specificity constant k cat/K m of 6.34 × 10− 4 μM− 1 min− 1 (K m = 25.9 ± 2 μM; V max = 65.7 ± 1 nM min− 1), and the mutant H133G cleaves the substrate of the autoprocessing site from SARS-CoV with the specificity constant of 0.69 ± 0.04 × 10− 4 μM− 1 min− 1. The data indicate that the substitution of His133 by glycine revealed no significant changes in substrate specificity.
Discussion
In the co-crystal structures of HRV 3Cpro–inhibitors, the β-ribbon adopts slightly different orientations when different inhibitors are bound.37 Two very dissimilar β-ribbon conformations were found in different crystal forms of FMDV 3Cpro.17, 18 These studies indicate that the β-ribbon is intrinsically flexible. Our structural studies discovered a novel open conformation of the β-ribbon in EV71 3Cpro, providing a clear snapshot of the β-ribbon at the wide open extremity (Fig. 4a and b). The similar unusual β-ribbon conformation was first found in one of the crystal forms of FMDV 3Cpro, in which parts of the opened β-ribbon of FMDV 3Cpro were missing from the electron density map. It was proved that the unusual β-ribbon conformation of FMDV 3Cpro was due to the crystallization artifacts. We found that the β-ribbon of EV71 3Cpro is also involved in the extensive crystallographic packing interactions in the current crystal form. The crystal packing apparently contributes to the stabilization of the open β-ribbon conformation. Moreover, our structural studies demonstrated that although the orientation of the β-ribbon with respect to the body of the EV71 3Cpro changed dramatically, the β-ribbon itself still maintains a rigid fold. Importantly, two hinge residues were discovered at the base of the β-ribbon, which govern the flexibility of the β-ribbon. One of the hinge residue Gly123, an amino acid with minimal peptide bond restraints, is highly conserved (Fig. 1d), suggesting that the dynamic β-ribbon is a common feature of picornaviral 3Cpro. Residues such as Leu127 and Ser128 in HRV 3Cpro and Cys142 in FMDV 3Cpro located on the β-ribbon are usually important for substrate recognition (P2–P4). However, in the case of the open β-ribbon conformation, these residues are directed away from the substrate binding cleft and, thus, cannot participate in the substrate recognition. In the mutagenesis study, we determined that residue Ser128 located on the tip of the β-ribbon of EV71 3Cpro is involved in substrate recognition; however, the residue is located far away from the substrate binding cleft in the crystal structure of EV71 3Cpro due to the open β-ribbon conformation. Thus, the observed open β-ribbon conformation may only be transient, and the conformation was stabilized by crystallization artifact. We believe that EV71 3Cpro may adopt both open and close conformations. The protease may adopt the close β-ribbon conformation during substrate binding. Our mutagenesis studies also demonstrated that the increased mobility of the β-ribbon is correlated to the improved protease activity. However, both mutant with the restrained β-ribbon (G123A) and the mutant with relaxed β-ribbon (H133G) have the K m value similar to that of WT protease, suggesting the similar substrate binding affinity. Thus, the data suggest that the mobility of the β-ribbon may be important to the product production steps subsequent to the initial substrate binding.
In summary, our data suggest a scenario where the substrate recognition and hydrolysis involve the movement of the β-ribbon. The β-ribbon is in motion in the absence of the substrate. The motion mode of the β-ribbon can be described as a moving clamp that flips between the open and close positions around the hinge residues. The mobility of the β-ribbon is determined by the peptide bond restraints of the hinge residues. The transiently opened β-ribbon results in the more exposed substrate binding cleft, thus may increase the substrate accessibility. When the substrate is bound, the β-ribbon can interact with the N-terminal side of the substrate and then stabilizes at the close conformation, leading to the enzyme–substrate complex. The β-ribbon may adopt different orientations upon substrate binding, depending on the structure of the substrate. Subsequently, the mobility of the β-ribbon may be required for the efficient disassociation of the N-terminal of the cleaved substrate after the acylation half of the reaction.
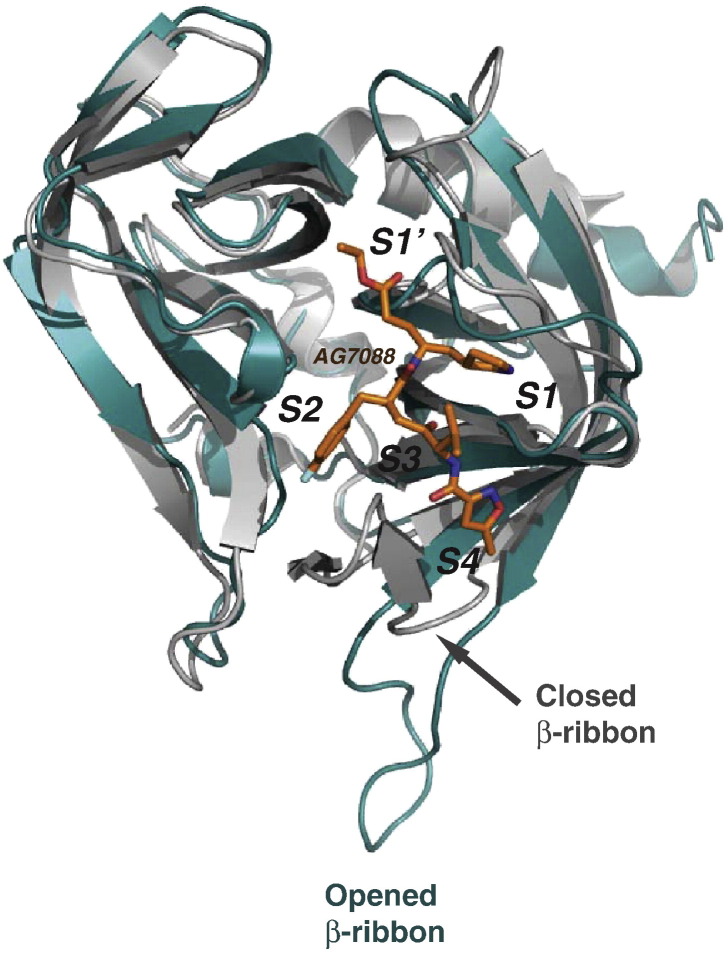
AG7088 (Rupintrivir) is an HRV 3Cpro inhibitor designed to combat common cold caused by rhinovirus infections. Interestingly, there are increasing evidences showing that AG7088 is also effective against enteroviruses.16, 40 In order to shed light on the structural basis of these observations, we modeled AG7088 into substrate binding cleft of EV71 3Cpro by superimposing the EV71 3Cpro structure onto the HRV 3Cpro–AG7088 co-crystal structure (Fig. 5 ). The modeled AG7088 is fitted to the substrate binding cleft of EV71 3Cpro similarly as observed in HRV 3Cpro–AG7088 interactions, suggesting that HRV 3Cpro and EV71 3Cpro share the similar architecture of substrate binding cleft and similar mechanisms for substrate recognition.
Fig. 5.
Superimposition of EV71 3Cpro and HRV 3Cpro–AG7088 structures. Ribbon model of EV71 3Cpro structure (deep teal) is superimposed onto the ribbon model of HRV 3Cpro–AG7088 co-structure (PDB code: 1CQQ). HRV 3Cpro is colored gray, and AG7088 is colored by elements (carbon atoms in orange). The substrate binding pockets, S1′–S4, are indicated. The opened β-ribbon of EV71 3Cpro and the closed β-ribbon of HRV 3Cpro are indicated.
EV71 3Cpro has no known cellular homologue in humans. Marginal differences in the sequences of 3Cpro proteins are observed among enteroviruses and coxsackieviruses causing HFMD.40 This limited sequence diversity suggests the significant structural similarity, which may offer the possibility of targeting the protease with single-drug therapies. Recently, based on the architecture of AG7088, specific inhibitors against EV71 3Cpro were designed.16 Our structural insight into EV71 3Cpro can certainly provide valuable information for further structure-based inhibitor developments.
Materials and Methods
Constructs and proteins preparation
The cDNA encoding the EV71 3Cpro (1–183 aa) was amplified by RT-PCR experiment using the genomic RNA of EV71 (EV71 isolate BJ/CHN/2008; GenBank accession number: HQ615421) as template. The sequence of the resulting cDNA was verified by DNA sequencing and was inserted into pET28a or pET21a vectors (Novagen) to generate plasmids expressing N-terminal 6× His-tagged, non-tagged and C-terminal 6× His-tagged EV71 3Cpro variants. The plasmid expressing the N-terminal-tagged EV71 3Cpro variant encodes full-length EV71 3Cpro (183 aa) and the N-terminal extension: MGSSHHHHHHSSGLVPRGSHM, containing 6× His, a linker region and a thrombin cleavage site (underlined). The non-tagged EV71 3Cpro variant was obtained by thrombin digestion followed by additional purification steps to remove thrombin contamination. The construct expressing the C-terminal EV71 3Cpro variant encodes full-length EV71 3Cpro plus the C-terminal extension: AAALEHHHHHH. Plasmids expressing EV71 3Cpro mutants were generated using the site-directed mutagenesis methods (QuikChange™) and were verified by DNA sequencing. The plasmids were transformed into E. coli Rosetta competent cells (Novagen). The bacteria cultures were grown at 37 °C to an OD600 = 1.0 and were cooled down to 18 °C before induction (0.5 mM IPTG). The bacteria cultures were shaken at 18 °C overnight and were subsequently harvested and disrupted by ultrasonication. All recombinant EV71 3Cpro variants and mutants were purified to homogeneity using multiple chromatographic steps following standard protocols. Briefly, batches of EV71 3Cpro lysate were initially purified using affinity chromatography (Ni-NTA; Qiagen) followed by cation-exchange chromatography (HighTrap SP; GE Healthcare). The final step of purification was size-exclusion chromatography (Sephacryl S-200 HR 26/60 column; GE Healthcare).
Size-exclusion experiments
Superose 6 10/300GL column (GE Healthcare) was pre-equilibrated with a buffer containing 50 mM Tris–HCl (pH 7.5), 150 mM NaCl and 1 mM DTT. The column was then calibrated using gel-filtration molecular mass standards (Bio-Rad), containing thyroglobulin (670 kDa), γ-globulin (158 kDa), ovalbumin (45 kDa), myoglobin (17 kDa) and vitamin B12 (1.35 kDa). The retention volumes of the standards were measured, and the values of V e (retention volume)/V o (void volume) were plotted as the function of − log10(MW) to derive the standard curve. EV71 3Cpro was loaded on the column to determine V e. The molecular weight of EV71 3Cpro was calculated using the standard curve.
Protease activity assay
The synthetic peptide substrates were attached with a fluorescence quenching pair, Dabcyl and Edans. The fluorogenic peptides Dabcyl-RTATVQGPSLDFE-Edans corresponding to the EV71 polyprotein autoprocessing site between 3B–3C, Dabcyl-MQASMQGGVPAPE-Edans derived from the cleavage site (251/252 aa) in CstF-64 and Dabcyl-TSAVLQSGFRKM-Edans derived from the N-terminal autoprocessing site of a 3C-like protease from SARS-CoV were used to measure the protease activities. Fluorescence experiments were performed with SpectraMax® M5 Multi-Mode Microplate Reader. We measured 100 μl of reaction mixture containing 50 mM Tris–HCl (pH 7.0), 200 mM NaCl, 2 mM DTT and 4 μM EV71 3Cpro or its mutants and fluorogenic peptides of different concentrations in a 96-well plate (Greiner) at 30 °C. The instrument was pre-calibrated by Edans standard to calculate the relationship between relative fluorescence unit and concentration. The relative fluorescence unit was collected using an excitation wavelength of 340nm and by monitoring the emission 500nm and was then converted to substrate concentration. The initial velocities of the reactions are plotted versus substrate concentrations. Values are averaged from three independent experiments. The data were fitted to Michaelis–Menten equation to calculate K m, V max and k cat/K m (MicroCal Origin). For data that the K m and V max values cannot be precisely determined by Michaelis–Menten equation fitting, the k cat/K m was calculated from the linear part of the rate-versus-substrate concentration plot.
Crystallization and structure determination
The EV71 3Cpro variants were concentrated to ∼ 12 mg ml− 1 prior to the crystallization trials, only N-terminal 6× His-tagged EV71 3Cpro resulted in useful crystals. The optimal crystallization of EV71 3Cpro was achieved by mixing 0.8 μl protein with 1.0 μl buffer containing 100 mM Tris–HCl (pH 7.7), 200 mM sodium citrate and 20% polyethylene glycol 3350 in a hanging-drop vapor diffusion system at 22 °C. Cryocooling was achieved by soaking the crystals for 30–60 s in reservoir solution containing 15% ethylene glycol and by flash-freezing them in liquid nitrogen. Complete data to a resolution limit of 3.0 Å on native crystals were collected at the PXIII beamline of the Swiss Light Source (SLS; Villigen, Switzerland). The data sets were processed with the XDS Package.41 The space group was identified as P21, and five molecules were found in the ASU. The initial phases were obtained by the program Phaser v2.1,42 using the structure of CVB3 3Cpro [Protein Data Bank (PDB) ID: 2VB0] structure as search model. The electron density map was further improved by non-crystallographic symmetry averaging. The manual model building was carried out with the program Coot v0.6.43 Prior to model building and refinement, we randomly omitted 5% of the reflections for monitoring the R free value. Refinement was performed with PHENIX44 and included overall anisotropic B-factor and bulk solvent corrections, individual B-factor refinement, simulated annealing and positional refinement. Data collection and model statistics are summarized in Table 1. Figures were prepared with PyMOL (DeLano Scientific).
Accession number
The coding sequence of EV71 isolate BJ/CHN/2008 3C protein has been deposited in GenBank with the accession number HQ615421.
Coordinates and structure factors have been deposited in the PDB with accession number 3OSY.
Acknowledgements
We thank Prof. Dr. Hopfner Karl-Peter for careful reading the manuscript. We thank the SLS (Villigen) for beam time allowance, and we thank the staffs of beamline X 06DA (PXIII) at SLS for help with data collection. We thank Dr. Ezequiel Panepucci for the careful proofreading of the manuscript. This work was supported by the National Basic Research Program of China (973 Program), program no. 2011CB504902; the National Natural Science Foundation of China, program no. 81071349 and an intramural grant from the Institute of Pathogen Biology, Chinese Academy of Medical Sciences and Peking Union Medical College (no. 2008IPB201) to S.C. National Science and Technology Major Project of China, project No. 2009ZX10004-303.
Edited by R. Huber
Contributor Information
Sheng Cui, Email: cuisheng2007@yahoo.com.cn.
Meitian Wang, Email: meitian.wang@psi.ch.
Qi Jin, Email: jinqi@ipbcams.ac.cn.
References
- 1.Chen C.Y., Chang Y.C., Huang C.C., Lui C.C., Lee K.W., Huang S.C. Acute flaccid paralysis in infants and young children with enterovirus 71 infection: MR imaging findings and clinical correlates. Am. J. Neuroradiol. 2001;22:200–205. [PMC free article] [PubMed] [Google Scholar]
- 2.Lee T.C., Guo H.R., Su H.J., Yang Y.C., Chang H.L., Chen K.T. Diseases caused by enterovirus 71 infection. Pediatr. Infect. Dis. J. 2009;28:904–910. doi: 10.1097/INF.0b013e3181a41d63. [DOI] [PubMed] [Google Scholar]
- 3.McMinn P., Stratov I., Nagarajan L., Davis S. Neurological manifestations of enterovirus 71 infection in children during an outbreak of hand, foot, and mouth disease in Western Australia. Clin. Infect. Dis. 2001;32:236–242. doi: 10.1086/318454. [DOI] [PubMed] [Google Scholar]
- 4.Ho M., Chen E.R., Hsu K.H., Twu S.J., Chen K.T., Tsai S.F. An epidemic of enterovirus 71 infection in Taiwan. Taiwan Enterovirus Epidemic Working Group. N. Engl. J. Med. 1999;341:929–935. doi: 10.1056/NEJM199909233411301. [DOI] [PubMed] [Google Scholar]
- 5.Zhang S.B., Liao H., Huang C.H., Tan Q.Y., Zhang W.L., Huang Y. [Serum types of enterovirus and clinical characteristics of 237 children with hand, foot and mouth disease in Shenzhen] Zhongguo Dangdai ErKe ZaZhi. 2008;10:38–41. [PubMed] [Google Scholar]
- 6.AbuBakar S., Chee H.Y., Al-Kobaisi M.F., Xiaoshan J., Chua K.B., Lam S.K. Identification of enterovirus 71 isolates from an outbreak of hand, foot and mouth disease (HFMD) with fatal cases of encephalomyelitis in Malaysia. Virus Res. 1999;61:1–9. doi: 10.1016/s0168-1702(99)00019-2. [DOI] [PubMed] [Google Scholar]
- 7.Alexander J.P., Jr., Baden L., Pallansch M.A., Anderson L.J. Enterovirus 71 infections and neurologic disease—United States, 1977–1991. J. Infect. Dis. 1994;169:905–908. doi: 10.1093/infdis/169.4.905. [DOI] [PubMed] [Google Scholar]
- 8.Kehle J., Roth B., Metzger C., Pfitzner A., Enders G. Molecular characterization of an Enterovirus 71 causing neurological disease in Germany. J. NeuroVirol. 2003;9:126–128. doi: 10.1080/13550280390173364. [DOI] [PubMed] [Google Scholar]
- 9.Yan, Central China province reports surge in HFMD deaths http://news.xinhuanet.com/english, 2010.
- 10.The Chinese Center for Disease Control and Prevention, and the World Health Organization, Report on the Hand, Foot and Mouth Disease Outbreak in Fuyang City, Anhui Province and the Prevention and Control in China (May 2008) World Health Organization Representative Office in China, 2008.
- 11.Zhang D., Lu J., Lu J. Enterovirus 71 vaccine: close but still far. Int. J. Infect. Dis. 2010;14:e739–e743. doi: 10.1016/j.ijid.2009.12.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yang F., Jin Q., He Y., Li L., Hou Y. The complete genome of Enterovirus 71 China strain. Sci. China, Ser. C: Life Sci. 2001;44:178–183. doi: 10.1007/BF02879323. [DOI] [PubMed] [Google Scholar]
- 13.Hwang Y.C., Chen W., Yates M.V. Use of fluorescence resonance energy transfer for rapid detection of enteroviral infection in vivo. Appl. Environ. Microbiol. 2006;72:3710–3715. doi: 10.1128/AEM.72.5.3710-3715.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Bishop N.E., Anderson D.A. RNA-dependent cleavage of VP0 capsid protein in provirions of hepatitis A virus. Virology. 1993;197:616–623. doi: 10.1006/viro.1993.1636. [DOI] [PubMed] [Google Scholar]
- 15.Weng K.F., Li M.L., Hung C.T., Shih S.R. Enterovirus 71 3C protease cleaves a novel target CstF-64 and inhibits cellular polyadenylation. PLoS Pathog. 2009;5:e1000593. doi: 10.1371/journal.ppat.1000593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kuo C.J., Shie J.J., Fang J.M., Yen G.R., Hsu J.T., Liu H.G. Design, synthesis, and evaluation of 3C protease inhibitors as anti-enterovirus 71 agents. Bioorg. Med. Chem. 2008;16:7388–7398. doi: 10.1016/j.bmc.2008.06.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Birtley J.R., Knox S.R., Jaulent A.M., Brick P., Leatherbarrow R.J., Curry S. Crystal structure of foot-and-mouth disease virus 3C protease. New insights into catalytic mechanism and cleavage specificity. J. Biol. Chem. 2005;280:11520–11527. doi: 10.1074/jbc.M413254200. [DOI] [PubMed] [Google Scholar]
- 18.Sweeney T.R., Roqué-Rosell N., Birtley J.R., Leatherbarrow R.J., Curry S. Structural and mutagenic analysis of foot-and-mouth disease virus 3C protease reveals the role of the beta-ribbon in proteolysis. J. Virol. 2007;81:115–124. doi: 10.1128/JVI.01587-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zunszain P.A., Knox S.R., Sweeney T.R., Yang J., Roqué-Rosell N., Belsham G.J. Insights into cleavage specificity from the crystal structure of foot-and-mouth disease virus 3C protease complexed with a peptide substrate. J. Mol. Biol. 2009;395:375–389. doi: 10.1016/j.jmb.2009.10.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lee C.C., Kuo C.J., Ko T.P., Hsu M.F., Tsui Y.C., Chang S.C. Structural basis of inhibition specificities of 3C and 3C-like proteases by zinc-coordinating and peptidomimetic compounds. J. Biol. Chem. 2009;284:7646–7655. doi: 10.1074/jbc.M807947200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Fuhrmann C.N., Kelch B.A., Ota N., Agard D.A. The 0.83 Å resolution crystal structure of alpha-lytic protease reveals the detailed structure of the active site and identifies a source of conformational strain. J. Mol. Biol. 2004;338:999–1013. doi: 10.1016/j.jmb.2004.03.018. [DOI] [PubMed] [Google Scholar]
- 22.Matthews D.A., Dragovich P.S., Webber S.E., Fuhrman S.A., Patick A.K., Zalman L.S. Structure-assisted design of mechanism-based irreversible inhibitors of human rhinovirus 3C protease with potent antiviral activity against multiple rhinovirus serotypes. Proc. Natl Acad. Sci. USA. 1999;96:11000–11007. doi: 10.1073/pnas.96.20.11000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Hopfner K.P., Lang A., Karcher A., Sichler K., Kopetzki E., Brandstetter H. Coagulation factor IXa: the relaxed conformation of Tyr99 blocks substrate binding. Structure. 1999;7:989–996. doi: 10.1016/s0969-2126(99)80125-7. [DOI] [PubMed] [Google Scholar]
- 24.Bergmann E.M., Cherney M.M., Mckendrick J., Frormann S., Luo C., Malcolm B.A. Crystal structure of an inhibitor complex of the 3C proteinase from hepatitis A virus (HAV) and implications for the polyprotein processing in HAV. Virology. 1999;265:153–163. doi: 10.1006/viro.1999.9968. [DOI] [PubMed] [Google Scholar]
- 25.Mosimann S.C., Cherney M.M., Sia S., Plotch S., James M.N. Refined X-ray crystallographic structure of the poliovirus 3C gene product. J. Mol. Biol. 1997;273:1032–1047. doi: 10.1006/jmbi.1997.1306. [DOI] [PubMed] [Google Scholar]
- 26.Allaire M., Chernaia M.M., Malcolm B.A., James M.N. Picornaviral 3C cysteine proteinases have a fold similar to chymotrypsin-like serine proteinases. Nature. 1994;369:72–76. doi: 10.1038/369072a0. [DOI] [PubMed] [Google Scholar]
- 27.Phan J., Zdanov A., Evdokimov A.G., Tropea J.E., Peters H.K., 3rd, Kapust R.B. Structural basis for the substrate specificity of tobacco etch virus protease. J. Biol. Chem. 2002;277:50564–50572. doi: 10.1074/jbc.M207224200. [DOI] [PubMed] [Google Scholar]
- 28.Fujinaga M., Delbaere L.T., Brayer G.D., James M.N. Refined structure of alpha-lytic protease at 1.7 Å resolution. Analysis of hydrogen bonding and solvent structure. J. Mol. Biol. 1985;184:479–502. doi: 10.1016/0022-2836(85)90296-7. [DOI] [PubMed] [Google Scholar]
- 29.James M.N., Sielecki A.R., Brayer G.D., Delbaere L.T., Bauer C.A. Structures of product and inhibitor complexes of Streptomyces griseus protease A at 1.8 Å resolution. A model for serine protease catalysis. J. Mol. Biol. 1980;144:43–88. doi: 10.1016/0022-2836(80)90214-4. [DOI] [PubMed] [Google Scholar]
- 30.Fujinaga M., Read R.J., Sielecki A., Ardelt W., Laskowski M., Jr, James M.N. Refined crystal structure of the molecular complex of Streptomyces griseus protease B, a serine protease, with the third domain of the ovomucoid inhibitor from turkey. Proc. Natl Acad. Sci. USA. 1982;79:4868–4872. doi: 10.1073/pnas.79.16.4868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Shih S.R., Chiang C., Chen T.C., Wu C.N., Hsu J.T., Lee J.C. Mutations at KFRDI and VGK domains of enterovirus 71 3C protease affect its RNA binding and proteolytic activities. J. Biomed. Sci. 2004;11:239–248. doi: 10.1007/BF02256567. [DOI] [PubMed] [Google Scholar]
- 32.Peters H., Kusov Y.Y., Meyer S., Benie A.J., Bäuml E., Wolff M. Hepatitis A virus proteinase 3C binding to viral RNA: correlation with substrate binding and enzyme dimerization. Biochem. J. 2005;385:363–370. doi: 10.1042/BJ20041153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kusov Y.Y., Gauss-Muller V. In vitro RNA binding of the hepatitis A virus proteinase 3C (HAV 3Cpro) to secondary structure elements within the 5′ terminus of the HAV genome. RNA. 1997;3:291–302. [PMC free article] [PubMed] [Google Scholar]
- 34.Andino R., Rieckhof G.E., Achacoso P.L., Baltimore D. Poliovirus RNA synthesis utilizes an RNP complex formed around the 5′-end of viral RNA. EMBO J. 1993;12:3587–3598. doi: 10.1002/j.1460-2075.1993.tb06032.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Parsley T.B., Towner J.S., Blyn L.B., Ehrenfeld E., Semler B.L. Poly (rC) binding protein 2 forms a ternary complex with the 5′-terminal sequences of poliovirus RNA and the viral 3CD proteinase. RNA. 1997;3:1124–1134. [PMC free article] [PubMed] [Google Scholar]
- 36.Harris K.S., Reddigari S.R., Nicklin M.J., Hämmerle T., Wimmer E. Purification and characterization of poliovirus polypeptide 3CD, a proteinase and a precursor for RNA polymerase. J. Virol. 1992;66:7481–7489. doi: 10.1128/jvi.66.12.7481-7489.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Matthews D.A., Smith W.W., Ferre R.A., Condon B., Budahazi G., Sisson W. Structure of human rhinovirus 3C protease reveals a trypsin-like polypeptide fold, RNA-binding site, and means for cleaving precursor polyprotein. Cell. 1994;77:761–771. doi: 10.1016/0092-8674(94)90059-0. [DOI] [PubMed] [Google Scholar]
- 38.Leong L.E., Walker P.A., Porter A.G. Human rhinovirus-14 protease 3C (3Cpro) binds specifically to the 5′-noncoding region of the viral RNA. Evidence that 3Cpro has different domains for the RNA binding and proteolytic activities. J. Biol. Chem. 1993;268:25735–25739. [PubMed] [Google Scholar]
- 39.Walker P.A., Leong L.E., Porter A.G. Sequence and structural determinants of the interaction between the 5′-noncoding region of picornavirus RNA and rhinovirus protease 3C. J. Biol. Chem. 1995;270:14510–14516. doi: 10.1074/jbc.270.24.14510. [DOI] [PubMed] [Google Scholar]
- 40.Zhang X.N., Song Z.G., Jiang T., Shi B.S., Hu Y.W., Yuan Z.H. Rupintrivir is a promising candidate for treating severe cases of Enterovirus-71 infection. World J. Gastroenterol. 2010;16:201–209. doi: 10.3748/wjg.v16.i2.201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kabsch W. XDS. Acta Crystallogr., Sect. D: Biol. Crystallogr. 2010;66:125–132. doi: 10.1107/S0907444909047337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.McCoy A.J., Grosse-Kunstleve R.W., Adams P.D., Winn M.D., Storoni L.C., Read R.J. Phaser crystallographic software. J. Appl. Crystallogr. 2007;40:658–674. doi: 10.1107/S0021889807021206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Emsley P., Cowtan K. Coot: model-building tools for molecular graphics. Acta Crystallogr., Sect. D: Biol. Crystallogr. 2004;60:2126–2132. doi: 10.1107/S0907444904019158. [DOI] [PubMed] [Google Scholar]
- 44.Adams P.D., Afonine P.V., Bunkóczi G., Chen V.B., Davis I.W., Echols N. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr., Sect. D: Biol. Crystallogr. 2010;66:213–221. doi: 10.1107/S0907444909052925. [DOI] [PMC free article] [PubMed] [Google Scholar]