Key Points
A decade after the report of the first devices, synthetic biology has developed into an engineering science that provides novel opportunities to understand, diagnose, prevent and treat diseases.
Chemical synthesis and reconstruction of extinct or difficult-to-propagate viral genomes improves our understanding of virulence factors.
The de novo synthesis of deoptimized viral genomes enables the production of safe life vaccines.
Engineering environmentally responsive dominant-lethal genetic circuits into disease-transmitting insects provides a highly specific approach for controlling disease propagation.
The reconstruction of bacterial resistance circuits in mammalian cells enables the integrated discovery of agents to overcome resistance.
Engineered bacteria and synthetic genetic circuits that specifically detect and destroy neoplastic cells will provide momentum to future cancer therapies.
Molecular prostheses that detect disease states and autonomously trigger a therapeutic response in a closed-loop control configuration provide novel opportunities in the treatment of genetic and acquired diseases.
Synthetic gene circuits will provide novel opportunities for future gene and cell-based therapies.
Subject terms: Technology, Gene expression, Systems biology, Synthetic biology, Health sciences
Synthetic biological devices can be engineered to achieve high levels of precision and specificity, which makes them ideally suited for use in clinical settings. Devices are being developed to meet a range of biomedical needs, including specific cancer therapies and metabolic control.
Abstract
Synthetic biology aims to create functional devices, systems and organisms with novel and useful functions on the basis of catalogued and standardized biological building blocks. Although they were initially constructed to elucidate the dynamics of simple processes, designed devices now contribute to the understanding of disease mechanisms, provide novel diagnostic tools, enable economic production of therapeutics and allow the design of novel strategies for the treatment of cancer, immune diseases and metabolic disorders, such as diabetes and gout, as well as a range of infectious diseases. In this Review, we cover the impact and potential of synthetic biology for biomedical applications.
Main
By applying engineering principles to biology, synthetic biology has become the science of reassembling catalogued and standardized biological components in a systematic and rational manner to create and engineer functional biological designer devices, systems and organisms with predictable, useful and novel functions. Synthetic biology is able to use an inventory of biomolecular parts compiled over 50 years of molecular biological and functional genomic research1,2,3,4, as well as technology that has made it possible to analyse5,6, synthesize7,8,9, assemble10, modify11 and transfer12,13 genetic components into living organisms.
Although it has recently become possible to reconstruct a living organism after transfer of a synthetic genome that has been assembled from chemically synthesized nucleic acid pieces13,14, the rational design of state-of-the-art biological circuits with predicable functions remains challenging and is apparently limited to a handful genes15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31.
Synthetic circuitry is composed of basic heterologous control components that fine-tune transgene expression in response to specific exogenous cues or endogenous metabolites32. These gene switches include trigger-inducible protein–DNA33,34,35,36 or aptamer–transcript interactions37,38,39, which control transcription and translation in response to heterologous and endogenous input signals (Fig. 1). The standardized design of gene switches has improved functional compatibility40,41 and has enabled the construction of higher-order networks — including multi-trigger inputs and sequential control42,43, mutual control20,23,44,45 and feedback control16,19,28,46 of circuit components — that are able to provide complex protein expression dynamics with high precision and predictable logic in response to external cues or physiological pathways. Because most control components function in different bacterial and eukaryotic species after minor refinements36,47,48,49, gene switches and network blueprints that were pioneered in bacteria or yeast are often fully functional in mammalian cells. Examples of synthetic networks with similar components and circuit topology in bacterial and mammalian cells include: regulatory cascades42,43, epigenetic toggle switches15,20,23,45, hysteretic circuits22,50,51, molecular timing devices30,52, synthetic eco-sensing systems, synthetic quorum-sensing systems, synthetic hormone systems46,53,54, band-pass filters21,30,55 and different types of oscillators that program rhythmic transgene expression with a tunable frequency and amplitude16,26,27,28,29,56,57. Most of these first-generation synthetic circuits operated in isolation without any interface with the metabolism of the host cell, and they were used to program specific cellular functions using heterologous external input signals58,59,60,61,62.
Figure 1. Mammalian gene expression control strategies.
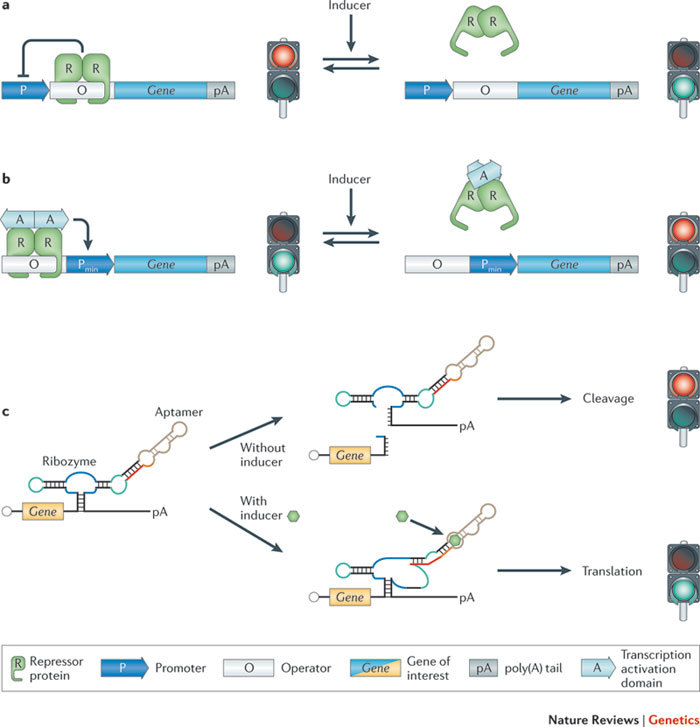
a | Repression-based expression control. A repressor protein binds to its operator and thus prevents activation of the promoter and expression of the gene of interest. In response to an inducer, the repressor dissociates from the operator, the promoter is derepressed, and the gene of interest is expressed. b | Activation-based expression control. A minimal promoter (Pmin) is activated when a chimeric transcription factor that is constructed by fusing a repressor protein to a transcription activation domain binds to its operator. In the presence of an inducer, the repressor protein–transcription-activation domain complex dissociates from its operator, Pmin is no longer activated, and transcription of the gene of interest is prevented. c | mRNA transcript-based expression control. A self-cleaving ribozyme is fused to a small-molecule-binding aptamer and introduced into the 3′ untranslated region (UTR) of a gene of interest. In the absence of the inducer, the ribozyme undergoes self-cleavage, thereby eliminating the poly(A) tail (pA) from the open reading frame and preventing translation. However, in the presence of the inducer, the aptamer undergoes a conformational change, which inactivates the ribozyme and allows translation to occur.
Key challenges that often occur in the biomedical setting are the need for drug–target specificity, precise drug-dosing regimes, minimizing side effects, shortening the diagnosis-to-treatment timelines and avoiding drug-resistance of pathogens. As synthetic biology enables the engineering of complex, high-precision control devices that couple sensing and delivery mechanisms, this emerging science may be able to offer tools that are suited to meeting current biomedical challenges in new ways. A decade after the pioneering synthetic networks were reported18,20, the first successful therapeutic applications in animal models of prominent human diseases are starting to emerge63,64,65. This Review focuses on the recent progress in synthetic biology towards improving human health, including diagnostic applications, design of novel preventive care strategies, progress in drug discovery, design and delivery, and development of novel treatment strategies, such as prosthetic networks. Synthetic biology holds the promise of providing unique opportunities for major advances in the improvement of human health in the twenty-first century4,66,67.
Understanding disease mechanisms
Pathogen mechanisms. The availability of affordable technology for synthesizing and assembling10 sequences for proteins or for viral68 and bacterial69 genomes with increased speed has dramatically improved our understanding of host–pathogen interactions and of disease mechanisms. The synthetic biology principle of 'analysis by synthesis' provides mechanistic insight by combining rapid synthesis, assembly, shuffling and mutation of individual genetic components with straightforward functional analysis. For example, the genome of the H1N1 virus that was responsible for the 1918 Spanish influenza pandemic was synthesized using sequence information from genomic pieces that were extracted from permafrost-conserved tissue samples. Functional analysis of the reconstructed virus provided new insight into the key virulence factors of the pathogen: namely, a haemagglutinin variant that induces membrane fusion without trypsin activation and a modified polymerase that enhances viral replication68. The same study also revealed that a combination of eight genes was responsible for the exceptional virulence of the Spanish influenza strain68,70. This finding may help to identify the pandemic potential of future virus variants71,72,73.
Synthesis and analysis of chimeric viruses have also made a substantial contribution to the understanding of coronavirus zoonoses that were responsible for the severe acute respiratory syndrome (SARS) pandemic of 2002 and 2003. The characterization of the history of the SARS coronavirus, especially its switch in tropism, was particularly challenging, as its direct ancestors could not be propagated in laboratory models. However, after a 30 kb SARS-like bat coronavirus was designed to contain the receptor-binding spike protein of its human homologue, the synthesized chimeric virus was able to replicate in culture and infect mice74. These in vivo studies revealed infection-enhancing mutations in the spike protein and established this surface protein as a key factor that is responsible for tropism switches in coronavirus zoonoses74. Reconstruction of pathogens by DNA synthesis can also be used for the production of diagnostic high-density antigen arrays75,76, such as those used to profile post-Lyme-disease syndrome76 or the humoural immune responses to hepatitis C and the human immunodeficiency virus (HIV)77.
Immune systems. Synthetic biology has recently provided new insight into disorders that are related to deficiencies of the immune system, which is known for its particularly complex control circuits and cellular interaction networks. For example, dysfunction of B-lymphocyte activation underlies several physiological disorders78. Functional reconstitution and analysis of the human B cell antigen receptor (BCR) signalling cascade in insect cells revealed that BCRs are not activated by antigen-specific crosslinking, as presented in textbooks, but instead have an autoinhibitory oligomeric conformation on resting B lymphocytes that shifts to an active dissociated form when antigens bind79. This triggers the signalling cascade, which results in antibody production and the onset of a humoural immune response.
Also, construction of a representation of the complete human peptidome engineered for display on the surface of T7 phages enabled Church and colleagues80 to discover new autoantigens. They used patient-derived autoantibodies to enrich autoantigenic peptides displayed on the phages; they could then identify the antigens by high-throughput sequence analysis80. Knowledge of the antigens that are involved in autoimmune processes is important for understanding disease aetiology, developing accurate diagnostic tests and designing drugs that neutralize autoreactive immune cells80.
Disease prevention
Vaccines. High-throughput and high-precision assembly and engineering of entire genomes from well-defined genetic components using synthetic biology principles has provided new opportunities for the design of attenuated pathogens for use as vaccines. For example, primates immunized with virus-like particles that were produced by selective expression of particular chikungunya virus (CHIKV) structural proteins were protected against viraemia after a high-dose challenge; even immunodeficient mice that were treated with monkey-derived antibodies survived subsequent lethal doses of CHIKV81. DNA synthesis and assembly has also played an essential part in pioneering a safe live vaccine against the poliovirus82. The poliovirus was attenuated by systematic genome-scale changes of adjacent pairs of codons from over- to underrepresented codon sets in viral capsid genes (for example, GCC|GAA is strongly under-represented compared with GCA|GAG, although both encode Ala–Glu). These changes reduced translation and impaired the replication competence and infectivity of the virus. This attenuated poliovirus provides protective immunization in mice and offers a high safety standard given the low probability that all 631 individual changes will revert and thus reconstitute infectious wild-type viruses. The genome-engineering approach used here could represent a general strategy for designing live vaccines against infectious diseases. Other promising vaccination concepts include using antigen-producing immunostimulatory liposomes as genetically programmable synthetic vaccines83 and the production of heat-stable oral algae-based vaccines to protect against Staphylococcus aureus infections84,85.
Vector control. Suppression of insect vector populations using transgenic viral strains that harbour conditional dominant-lethal synthetic circuitry may control the transmission of malaria parasites and dengue viruses and could eventually control the spread of untreatable diseases86,87,88. Mosquitoes that are transgenic for a tetracycline-dependent transactivator (tTA) that is exclusively expressed in the female's indirect flight muscle can only be propagated in the presence of tetracycline, which represses the transcription of this gene. However, the absence of tetracycline leads to the development of a female-specific flightless phenotype87,88. Putting the eggs of this transgenic mosquito into the ecosystem results in male-only releases; female mosquitoes remain grounded and cannot feed, mate or take blood meals, which effectively represents a lethal phenotype. Males do not transmit the disease, but they disseminate the synthetic circuit across the resident wild-type mosquito population87,88 (Fig. 2a).
Figure 2. Synthetic biology for understanding and preventing disease.
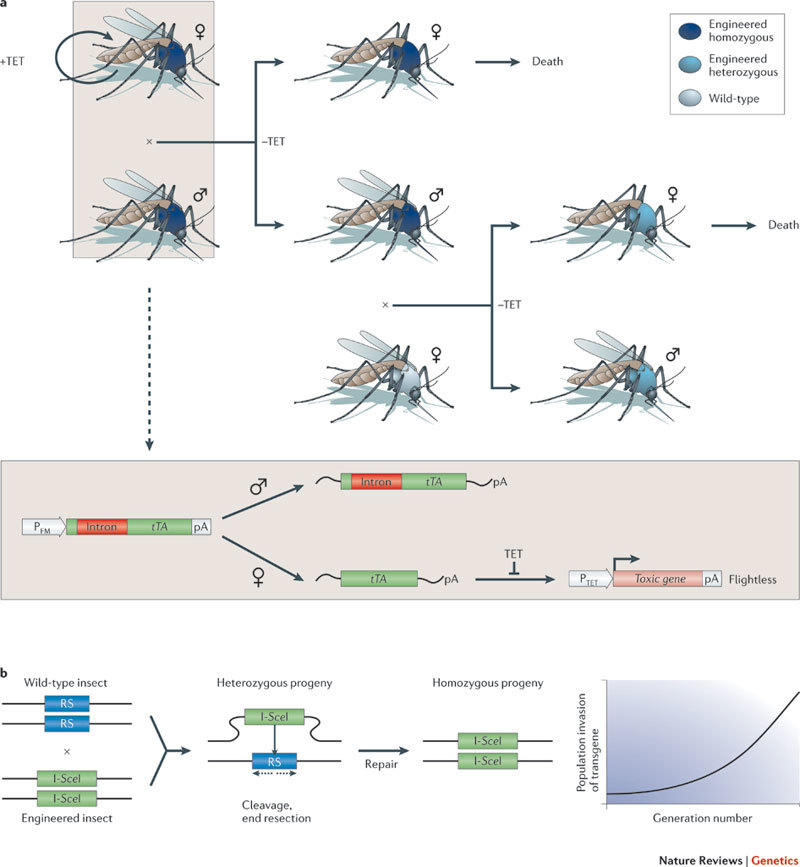
a | A female-specific dominant-lethal gene network for mosquito control. Mosquitoes were engineered to express an intron-containing variant of the tetracycline (TET) transactivator (tTA) under the control of a flight-muscle-specific promoter (PFM). In male mosquitoes, the intron is not spliced out, which prevents correct tTA translation. In female progeny, however, functional tTA translation is restored by sex-specific mRNA splicing. This results in the activation of the tTA-responsive promoter PTET and the expression of a toxic gene that triggers a flightless phenotype. If mosquitoes are raised in the presence of tetracycline (TET), tTA is prevented from activating PTET, which results in a normal phenotype. However, following their release into the TET-free environment, engineered males mate with wild-type females. This transmits the female-specific dominant flightless phenotype and should eventually result in the reduction or extinction of the wild-type population. b | Propagation of a selfish gene converting a heterozygous into a homozygous host. The homing endonuclease I-SceI is expressed and cleaves its cognate restriction site (RS) on the homologous chromosome. Following end resection and repair, the I-SceI expression cassette is inserted into the second chromosome. pA, poly(A) tail.
Similarly, a synthetic homing endonuclease-based gene drive system could be used to spread genetic modification, such as malaria resistance, from engineered mosquitoes to the field population. Homing endonucleases typically produce a single sequence-specific double-strand break in the host genome that is repaired by homologous recombination using the homing endonuclease gene (HEG) as a template. Consequently, the selfish HEG is copied to the broken chromosome in a gene conversion process referred to as 'homing'. Expressing the HEG I-SceI under the control of a male germline promoter enabled efficient homing in transheterozygous males and rapid genetic drive, which led to HEG invasion in caged mosquito populations89 (Fig. 2b). By engineering the sequence specificity of other HEGs (for example, I-AniI or I-CreI), the gene drive concept could, in principle, be used to knock in or knock out gene functions that target the mosquito's ability to serve as a disease vector89.
Field tests of release of insects carrying dominant lethals (RIDL) technology using first-generation tTA-transgenic mosquitoes have already been conducted in Grand Cayman. First, a small-scale release confirmed that transgenic males could survive, mate with wild females and produce transgenic larvae, and then the full field trial showed an 80% reduction in the numbers of wild mosquitoes about 11 weeks after release. As the study site was not isolated and the surrounding areas contained high densities of wild-type mosquitoes, scoring the actual suppression efficiency remains challenging90.
Drug development
Drug discovery. Synthetic mammalian transcription circuits consisting of a chimeric small-molecule-responsive transcription factor and a cognate synthetic promoter were originally designed for future gene-based therapies, and the aim was to adjust therapeutic transgene expression in mammalian cells in response to a pharmacologically active substance34,47,49,91. As most chimeric transcription factors are derived from repressors that manage drug resistance in bacteria (for example, resistance to antibiotics92) and are promiscuous for structurally related compounds, mammalian cells containing such circuitry could also be used in 'reverse mode', as integrated screening devices for the class-specific discovery of new drug candidates33,93 (for example, new antibiotics92) (Fig. 3a). When mammalian cells that are transgenic for the screening circuit are exposed to a compound library, they detect and modulate reporter gene expression in the presence of a non-toxic, cell-permeable and bioavailable molecule that has a class-specific core structure and corresponding drug activity (for example, antibiotic activity) (Fig. 3b). Using the same screening setup, compounds have been detected that lock the transcription factor onto the DNA, which may block induction of antibiotic resistance in pathogens and render them drug-sensitive94 (for example, see Bioversys). Using such compounds alongside the specific antibiotic may offer novel anti-infective treatment opportunities and a new life cycle for established antibiotics (Fig. 3c). Other trigger-inducible transcription control systems can be used in this manner as well, such as those that are responsive to streptogramin47, tetracycline or macrolide antibiotics91, anti-diabetes drugs95 or immunosuppressive lactones96,97.
Figure 3. Mammalian-cell-based drug discovery.
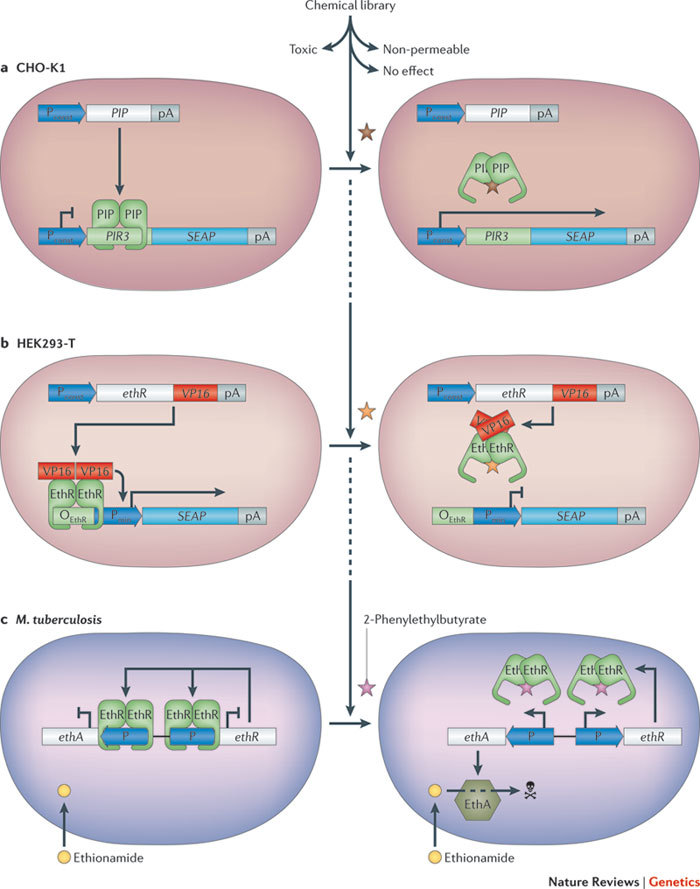
a | Identification of antibiotics. In Chinese hamster ovary (CHO-K1) cells, the streptogramin-responsive repressor (PIP) was expressed by a constitutive promoter (Pconst). PIP binds to its multimeric operator (PIR3) and represses expression of the reporter gene secreted alkaline phosphatase (SEAP). Exposing this screening cell line to a small molecule library only resulted in SEAP production for compounds that were streptogramin-like, cell-permeable and non-toxic (indicated by the brown star). b | Discovery of small molecules that are able to overcome antibiotic resistance. The Mycobacterium tuberculosis antibiotic resistance regulator (EthR) was fused to the herpes-simplex-derived transcriptional activator (VP16) and expressed in human embryonic kidney cells (HEK293-T) under the control of a constitutive promoter (Pconst). When EthR–VP16 binds to its cognate operator (OEthR), the minimal promoter (Pmin) is activated, which results in expression of the reporter gene SEAP. A screen is performed to identify a cell-permeable, non-toxic molecule (indicated by the yellow star) that prevents EthR binding to OEthR, stopping SEAP expression. c | Overcoming resistance to ethionamide in M. tuberculosis. In M. tuberculosis, EthR represses transcription of both the Baeyer–Villiger monooxygenase (EthA) and itself in a negative feedback loop. When 2-phenylethylbutyrate (indicated by the pink star) is added, it prevents EthR binding its target promoter (labelled 'P' in the figure). This derepresses EthA production, thereby turning ethionamide into a cytotoxic compound that kills the mycobacterium. pA, poly(A) tail.
One example of the efficacy of a transcription circuit system involves the bacterial transcriptional repressor EthR. EthR represses transcription of ethA and so prevents EthA-mediated conversion of the last-line-defence antibiotic ethionamide into a pathogen-killing metabolite94. The chemical 2-phenyethylbutyrate, best known for its strawberry flavour, was the first compound found that specifically inactivated EthR and so triggered ethA expression and re-established the sensitivity of Mycobacterium tuberculosis to ethionamide (Fig. 3d). Further work revealed other EthR-inactivating ethionamide booster compounds; these have also been successfully tested in a mouse model of human tuberculosis98. Restoring drug sensitivity by pharmacological inhibition of master resistance regulators may be widely applicable94.
A further example of the use of synthetic circuitry for drug discovery is provided by mammalian cells that are conditionally arrested in the G1 phase of the cell cycle by circuitry controlling the expression of the cycline-dependent kinase inhibitor p27. These cells reproducibly formed a mixture of isogenic subpopulations of proliferation-inhibited cells and proliferating cells that had spontaneously escaped the synthetic cell cycle block99. These cells could be used as a cell-based cancer model and could be used to screen for anticancer compounds that selectively eliminate proliferating cells while leaving arrested ones intact100,101.
Drug production and drug delivery. The synthetic pathways that are created by assembling enzymatic cascades or networks in bacteria, yeast and plants have been instrumental for the large-scale economic production of high-value drug and drug precursor compounds, as well as for the biosynthesis of new secondary metabolites with novel therapeutic activities. Examples include complex polyketides102,103, halogenated alkaloids104,105,106 and the precursors of the anti-malaria drug artemisinin (which is produced by the company Amyris, for example)107 and of the anti-cancer compound taxol108. For production of these compounds, it was necessary to overcome several challenges, including the functional expression of complex biosynthetic enzymes (such as cytochrome P450 monooxygenases109) and the overall orchestration of the multistep pathway to avoid accumulation of (toxic) intermediate products and to ensure metabolic channelling60.
Small-molecule-responsive protein–protein and protein–DNA interactions that are used to pioneer gene switches in mammalian cells36,49,110 have also been successfully re-engineered in the design of trigger-inducible biohybrid materials for drug delivery111,112,113,114,115. Using synthetic protein–polyacrylamide and DNA–polyacrylamide monomers, hydrogels can be produced that dissolve when specific ligands are supplied (Fig. 4). Biopharmaceuticals (for example, vascular endothelial growth factor (VEGF)) supplied during gel formation are loaded into the hydrogel and can be released in a dose-dependent manner after subcutaneous implantation into mice and oral administration of the trigger compound115. It is thought that any trigger-inducible protein–protein and protein–DNA interactions could be used to produce drug-sensing and drug-releasing hydrogels114,116,117.
Figure 4. Drug delivery.
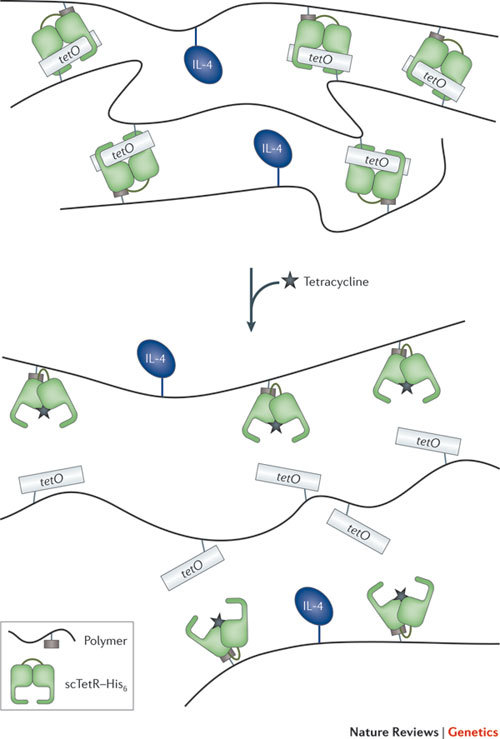
Interactive biohybrid material based on the interaction of a repressor protein with its cognate DNA operator motif. Homodimeric tetracycline repressor (TetR) is converted into a single-chain repressor (scTetR) by connecting two TetR subunits through a flexible peptide linker, and it is tagged with six histidines (scTetR–His6). This molecule is coupled to a polymer and is mixed with a polyacrylamide that has copies of a tetracycline operator (tetO) attached to it. scTetR binds to tetO so crosslinks are formed, making a hydrogel. When tetracycline is added, scTetR releases tetO, and the gel is dissolved. This can be used to release another molecule that was attached to the polymer — in this case, the cytokine interleukin 4 (IL-4).
Novel treatments for infections
Breaking bacterial resistance by designer phages. Biofilms are surface-associated bacterial communities that are encased in a hydrated extracellular polymeric substance (EPS) matrix that is composed of polysaccharides, proteins, nucleic acids and lipids. They are crucial to the pathogenesis of many clinically important bacteria and exhibit resistance both to the immune system and to antimicrobial treatments, making them difficult to eradicate118,119. Collins and colleagues120 successfully engineered bacteriophage T7 to constitutively express DspB: an enzyme that hydrolyses β-1,6-N-acetyl-D-glucosamine, which is an adhesin that is required for biofilm formation and integrity in Staphylococcus spp. and Escherichia coli clinical isolates. The initial infection of a bacterial biofilm with this bacteriophage (known as T7DspB) results in rapid multiplication of the phage and expression of DspB. Following lysis, T7DspB and DspB are released into the biofilm, which leads to re-infection and degradation of β-1,6-N-acetyl-D-glucosamine. During the process of T7DspB infection, bacterial biofilm cell counts are reduced by 99.997% — over two orders of magnitude greater than when a non-enzymatic phage is used120.
In a follow-up study, bacteriophage M13 was engineered to express LexA3, which suppresses the SOS DNA repair system that bacteria require to counteract antibiotic-induced oxidative stress121,122,123. Infection by this designer phage sensitizes E. coli to quinolone antibiotics. Use of this phage increases the survival of mice that are infected with E. coli, decreases the survival of antibiotic-resistant bacteria, persister cells and biofilm cells and reduces the number of antibiotic-resistant bacteria that arise from an antibiotic-treated population. It also acts as a strong adjuvant for other bactericidal antibiotics124. The designer phage platform can be used to produce other antibiotic adjuvants124.
Although it was once abandoned after the introduction of antibiotics, phage therapy is currently being revisited in several clinical trials around the world as the prevalence of multidrug-resistant pathogens is dramatically increasing. Although phage therapy may face clinical challenges associated with development of bacterial phage resistance, phage neutralization by the immune system and pharmacokinetics, the field will certainly receive an impetus from designer phages125.
Engineered probiotic bacteria decrease pathogen virulence. Bacteria can communicate with each other using a chemical language known as quorum sensing. Individual bacteria produce and secrete signalling molecules (called autoinducers) that are common to multiple species or are species-specific. These molecules accumulate as the population grows and can bind to receptors that coordinate colony-wide gene expression or manipulate the behaviour of other bacterial populations. For example, Vibrio cholerae produces cholera autoinducer 1 (CAI-1) and autoinducer 2 (AI-2), which trigger repression of key virulence factors. Feeding infant mice with a probiotic E. coli that naturally produces AI-2 and has been engineered to constitutively synthesize CAI-1 significantly increased the animals' survival rate after ingesting V. cholerae126. This suggests that such an approach could be an economic strategy to prevent infectious diseases. Unlike antibiotics, quorum-sensing-based interventions do not kill pathogens but reprogram their behaviour; this strategy may be free of selection pressure and therefore may be less prone to develop resistance. In another study, commensal bacteria were equipped with synthetic circuitry to stimulate glucose-dependent insulin production in intestinal epithelial cells127.
Cancer therapies
Despite decades of progress in cancer therapy, a major challenge remains: how to specifically target and selectively kill neoplastic cells that develop within native and implanted tissue and relocate within the organism to form metastasis. Therefore, therapeutic strategies that are designed to eliminate cancer cells must be extremely precise to exclusively target diseased tissue while leaving normal tissue intact. Although native cytotoxicity or the constitutive expression of anticancer compounds have demonstrated some potential in animal studies and human clinical trials128, trigger-inducible drug expression circuits delivered by tumour-invasive bacteria or tumour-transducing viral particles may improve cancer therapy. Synthetic biologists have recently designed a few anti-cancer devices that provide precise timing, location and dosing of drug production by external cues and could provide greater intra-tumoural effects while minimizing systemic toxicity.
Bacterial synthetic devices. After intravenous injection or oral administration, many bacterial species (for example, E. coli and Salmonella spp.) naturally sense and self-propel towards tumours. These bacteria have also been engineered to selectively invade and proliferate in tumour tissues and to produce cytotoxic compounds as well as reporter proteins for non-invasive follow-up monitoring of tumour regression128. These bacteria express flagella to penetrate tissue and chemotactic receptors to promote migration towards aspartate produced by viable cancer cells, ribose released by necrotic tissue or hypoxic regions generated by the hyper-metabolic activities of neoplastic cells. After they have reached the tumour site, the bacteria then either proliferate in the extracellular space or invade the tumour cells. In either situation, selective cytotoxicity was engineered by expressing toxins, cytokines, tumour antigens, pro-apoptotic factors or prodrug-converting enzymes128. Non-invasive E. coli has successfully been programmed to invade cultured tumour cells in a hypoxia-responsive or population-density-dependent manner. The corresponding circuitries consist of the anaerobically induced formate dehydrogenase promoter driving the Yersinia pseudotuberculosis invasin gene (inv), which mediates invasion using specific integrin receptors that are typically expressed on tumour cells. Population-density-dependent invasion requires an engineered quorum-sensing circuit that triggers inv expression after the bacterial population has reached a threshold size at the tumour site. This circuitry consists of quorum-sensing receptor LuxR that co-induces luxI (which encodes the enzyme producing the quorum-sensing messenger autoinducer 1 (AI-1)), and inv. AI-1-triggered, LuxR-mediated expression of luxI represents a positive feedback loop that amplifies inv expression and AI-1 production; this coordinates and broadcasts the invasion order across the entire population129 (Fig. 5a).
Figure 5. Bacterial and viral cancer therapy.
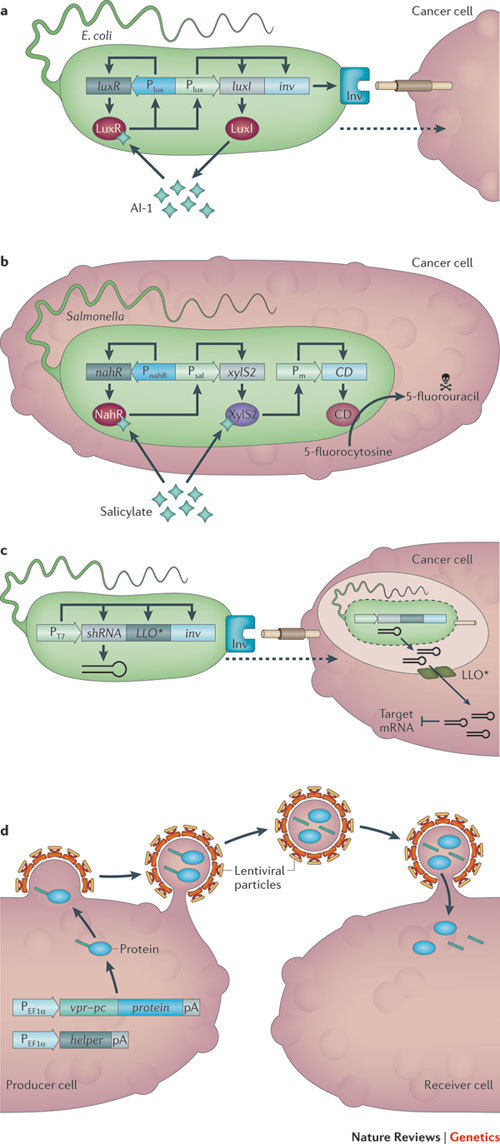
a | Population-density-dependent invasion of cancer cells. After intravenous injection, Escherichia coli accumulates in cancer tissue, where it reaches high population densities. E. coli is engineered to link the quorum-sensing receptor LuxR to an autoinducer 1 (AI-1)-inducible promoter (Plux). Plux is also used to drive luxI and the invasin gene inv. LuxI produces AI-1, generating a positive feedback loop that coordinates invasion throughout the population. b | Acetylsalicylic acid (Aspirin)-triggered killing of cancer cells after invasion of Salmonella spp. Salmonella spp. naturally invade cancer cells after intravenous injection. Salmonella spp. were engineered with a Pseudomonas putida-derived signal-amplifying two-level cascade in which NahR controls salicylate promoter (Psal)-driven xylS2 expression and XylS2 then triggers a XylS2-dependent promoter (Pm)-driven expression of the cytosine deaminase (labelled CD in the figure). Salicylate induces both NahR-based Psal and XylS2-mediated Pm activation. Mammalian cells are resistant to 5-fluorocytosine because they lack cytosine deaminase, which converts 5-fluorocytosine into the toxic cancer therapeutic 5-fluorouracil. c | Invasive bacteria suppress oncogene expression. E. coli is engineered to constitutively co-express a catenin β-1-specific short hairpin RNA (shRNA), Listeria monocytogenes listeriolysin (LLO*) and inv under control of the bacteriophage T7 promoter (PT7). They invade cancer cells (using the Inv protein), escape from the phagosome (using LLO*) and knock down the catenin β-1 oncogene (using shRNA). d | Therapeutic protein transduction. Lentiviral particles are produced using an integrase-negative helper vector (designated 'helper' in the figure) and a constitutive expression vector encoding the protein of interest (designated 'protein' in the figure) fused to viral protein R (VPR) and a protease cleavage site (PC). This can be delivered to any target cell in the absence of viral nucleic acids and proteins. An example application is described in the main text. pA, poly(A) tail. PEF1α, elongation factor 1 alpha (EF1α) promoter.
Tumour-invading bacteria have also been engineered for trigger-inducible drug expression after entering tumour cells. In addition to L-arabinose-130 and γ-irradiation-induced131 drug expression, a synthetic salicylate-triggered expression device has been used to control expression of drug components following systemic administration of the trigger molecule in mice in tumour cells that have been invaded by Salmonella spp.132. The device is based on a circuit that is derived from Pseudomonas putida, which controls expression of cytosine deaminase in a salicylate-inducible manner132. Mammalian cells normally lack cytosine deaminase, which means that they are resistant to 5-fluorocytosine because this enzyme is needed to convert 5-fluorocytosine into the cytotoxic molecule 5-fluorouracil. Tumour-bearing mice were injected with attenuated Salmonella enterica engineered with the P. putida-derived circuit and then treated with 5-fluorocytosine. The mice showed significant tumour regression when fed with acetylsalicylic acid (Aspirin)132, which is rapidly converted to salicylate after intake by the animal (Fig. 5b).
RNAi is a potent and highly conserved mechanism for the targeted knockdown of mRNA translation by small RNAs. Non-pathogenic E. coli was engineered to express a short RNA hairpin that triggers RNAi against catenin β-1, which is a colon-cancer-specific oncogene133. These bacteria, which were also engineered to express proteins to mediate cellular invasion and escape from the phagosome, were administered orally or intravenously and significantly reduced catenin β-1 levels in the intestinal epithelium and in human colon cancer xenografts in mice133 (Fig. 5c). Combining various bacterial anti-cancer treatment strategies may increase safety, specificity and efficiency in future clinical trials.
Viral synthetic devices. Viruses have also been successfully engineered to transduce specific cells by expressing epitopes that are recognized by particular cell-surface receptors and to express prodrug convertases and cytokines for use in cancer therapy134. Most of these oncolytic viruses carry coding viral nucleic acids, which may cause side effects owing to recombination with the host chromosome or proviral elements that are already in the host cell. Recently, synthetic viral particles have been designed that lack coding nucleic acids and that exclusively package therapeutic proteins, which can be released in a dose-dependent manner135. For example, viral particles carrying linamarase from Manihot esculenta were injected into human breast cancer xenografts in mice that had been treated with the non-toxic natural product linamarin; these viruses triggered efficient tumour regression owing to the cyanide produced by linamarase-mediated conversion of linamarin135 (Fig. 5d). Similarly, protein-carrying viral nanoparticles have been used to deliver site-specific DNA recombinases, such as FLP, to precisely integrate or excise genetic components on the host chromosome136. They might also be used to deliver native or chimeric transcription factors that could transiently control the expression of target genes that are involved in therapeutic interventions, lineage control or induction of pluripotency137.
A transformation sensor for cancer therapy. Gene therapy advances for cancer include virus-mediated delivery of cytotoxic effector genes controlled by cancer-specific promoters138,139 or delivery of chimeric adaptor proteins to link tyrosine kinase signalling to the apoptosis-inducing caspase machinery140. Most promoters and control circuits that coordinate simple reactions such as these are inherently noisy and only allow linear responses, which means limited control of specificity and efficacy. However, using two internal input signals can improve fidelity, mediate sharp response profiles and ensure robust biochemical processes141. Using decision-making circuits as blueprints, Nissim and Bar-Ziv142 designed a tunable dual promoter integrator (DPI) to target cancer cells precisely. The DPI consists of two native promoters that are concurrently activated by two independent transcription factors. Each cancer-sensing promoter produces a different fusion protein in proportion to its activity, and these two proteins assemble together as a chimeric transcription factor. This transcription factor then activates a synthetic promoter that controls expression of the herpes simplex virus type 1 thymidine kinase (TK1), which is cytotoxic in the presence of nucleotide analogues, such as ganciclovir (Fig. 6a). The DPI could be optimized for a specific cancer cell type by using different combinations of input promoters and effector genes, as well as by modulating the assembly efficiency and half life of the chimeric transactivator components. So far, a set of three promoters have been characterized in detail, but the DPI design may accommodate other suitable promoters.
Figure 6. Synthetic genetic cancer classifiers.
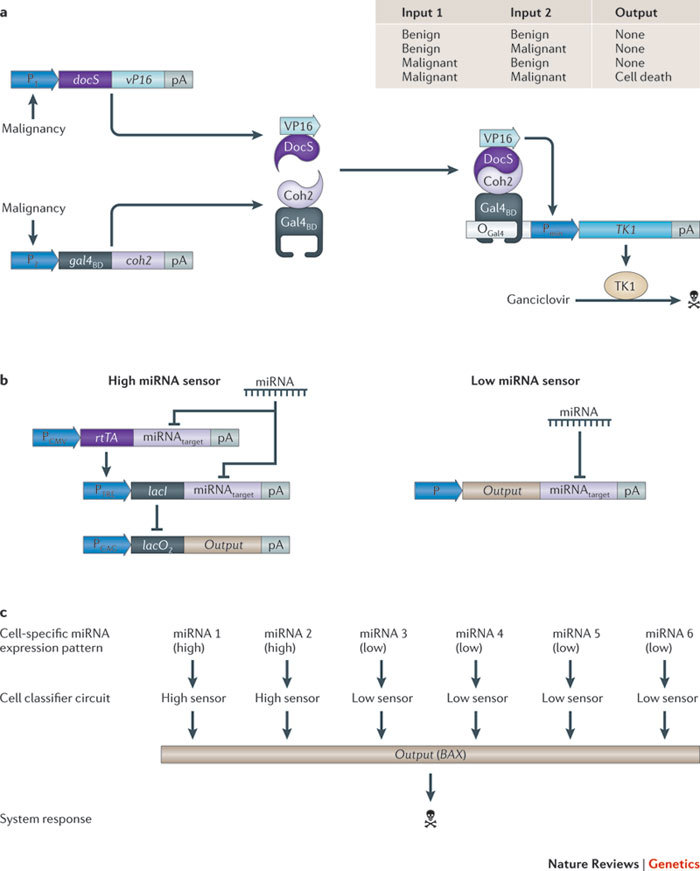
a | A transformation-sensing cancer kill switch can consist of a two-input, transformation-sensing device with 'AND' logic. The device constantly monitors the transformation state of a cell and produces a kill signal when two malignancy markers occur. Two independent malignancy-sensitive promoters drive expression of two chimeric proteins (DocS–VP16 and Gal4BD–Coh2). When they are simultaneously expressed, both proteins dimerize to form a synthetic transcription factor that binds Gal4 operator sites (OGal4), induces downstream minimal promoters (Pmin) and triggers expression of the herpes simplex virus type 1 thymidine kinase (TK1). In the presence of ganciclovir, the system is cytotoxic. b | A microRNA (miRNA)-based cancer classifier that discriminates cancer cells from non-transformed cells by scoring high and low expression profiles of a set of cancer-specific miRNAs. The classifier consists of high and low miRNA sensors that exclusively promote output gene expression if the specific input miRNAs are expressed at high or low levels, respectively. In the high miRNA sensor, high-target miRNA concentrations prevent translation of mRNAs encoding the reverse tetracycline-dependent transactivator (rtTA) and the repressor of the lactose operon (LacI). This results in derepression of transcription of the output gene (labelled 'Output' in the figure). In the low miRNA sensor, the output-gene-encoding mRNA is only translated when low-target miRNA concentrations are present. c | By combining different high and low miRNA sensors, the classifier can be customized to sense predetermined profiles of high and low miRNA levels, such as the ones that are typically produced by cancer cells and respond with expression of the apoptosis-inducing human BCL2-associated X protein (BAX). pA, poly(A) tail.
The recently developed 'cell-type classifier' is conceptually similar to the DPI, as it can also be programmed to destroy cells that express a specific set of neoplastic markers31. The cell-type classifier combines transcription and translation control components in a single scalable synthetic circuit that senses expression levels of a set of (currently up to six) endogenous microRNAs (miRNAs); it triggers an apoptosis-inducing response only if those levels match a preset profile. The cell-type classifier combines sensor modules for the detection of highly and lowly expressed miRNAs (Fig. 6b). For clinical implementation, both the DPI and the cell-type classifier must either be delivered to the cancer tissue, or they must provide a fail-safe mechanism that constantly eliminates transforming cells from engineered tissue implants.
Other emerging tools for biomedicine
Novel treatment strategies will require new technologies to sense and control disease. Synthetic biologists have designed new devices that could sense key physiological activities and have found new ways to dose therapeutic interventions precisely in response to external physical cues. Such synthetic devices could have wide-ranging biomedical applications.
RNA controllers of cell proliferation. Thus far, synthetic control devices that are designed to interface with host metabolism and to reprogram cellular behaviour have largely been limited to heterologous transcription factors. RNA controllers may represent an alternative. They are straightforward to design and can be integrated into a single expression unit containing sensors (aptamers), gene-regulatory components (ribozymes) and effector transgenes39,143,144. The inherent modularity and compatibility of RNA-based control components enables them to be independently optimized or exchanged. For example, an RNA control device consisting of a drug-responsive aptamer linked to a ribozyme in the 3′ untranslated region (UTR) of a cytokine expression unit enabled trigger-inducible inactivation of ribozyme-mediated transcript cleavage and full transgene expression in the presence of the input signal145. This synthetic RNA control device was applied to control proliferation of engineered primary human T cells and enabled external control of the expansion of transgenic T cells that are implanted into mice. Synthetic RNA control devices could provide the advance that is necessary to enable T cell therapy145; by contrast, state-of-the-art, trigger-inducible expansion of engineered T cells using chimeric antigen receptors has only led to moderate proliferation and poor survival of T cells in clinical trials146,147.
Another use for synthetic RNA is the design of programmable sensor–actuator devices that convert levels of an intracellular protein into a discrete high or low transgene-expression state148. The RNA devices consist of a three-exon, two-intron minigene followed by the transgene. The introns contain protein-sensing aptamers, and the central exon includes a stop codon. Binding of the protein to the aptamers controls splicing of the minigene; when the central exon is spliced out, the transgene is expressed at high levels, and when it remains unspliced, the transgene is expressed at low levels. Such a device was configured to sense subunits of nuclear factor kappa B (NFκB) or β-catenin (which are neoplastic markers) and to express the herpes simplex virus thymidine kinase. Thymidine kinase renders cells susceptible to ganciclovir, so this device only operated as a cancer kill switch in the presence of the cancer markers and gangcicolvir148. The modular configuration of the RNA sensor–actuator device allows it to be tailored to different intracellular proteins and even to multi-protein input using specific intronic aptamers. Also, responsiveness and performance can be tuned by placing the aptamers at different locations within the introns. The availability of compact RNA sensor–actuators that are easy to design and to alter and that control transgene expression in response to intracellular levels of key proteins may also improve the ability to link metabolic disease states with gene-based therapeutic interventions.
Optogenetic devices in blood glucose homeostasis. Light is becoming increasingly popular as a traceless, molecule-free input signal for triggering transgene expression in living systems. Bacteria have been engineered to record projected images with gigapixel resolution149,150,151 and to adjust transgene expression in response to multi-chromatic input150, and now genetic light switches have also been designed to control gene expression152 and shape of mammalian cells153.
Devices that convert light pulses into transcription may foster novel therapeutic opportunities in future gene- and cell-based therapies and may improve the manufacturing of difficult-to-produce protein pharmaceuticals, such as cancer therapeutics. An illustrative example is light-controlled expression of the glucagon-like peptide 1 (GLP1), which is a promising drug candidate for the treatment of type 2 diabetes65 (Fig. 7a). An optogenetic device that enables light-triggered gene expression in human cells was designed. This involves ectopic expression of melanopsin in human embryonic kidney cells and functional rewiring of signalling downstream of melanopsin; the cascade integrates blue-light-pulse-triggered photoreception and produces a reversible and sustained intensity-dependent transcription response. When placed in hollow fibre containers and implanted into mice, transgene expression in the engineered light-sensitive cells could be controlled remotely by an optical fibre65. Illuminating mice that carried subcutaneous implants of microencapsulated photo-responsive cells also enabled transdermal control of transgene expression and of corresponding protein levels in the blood of treated animals. This system was able to attenuate glycaemic excursions and to control glucose homeostasis in a mouse model of human type 2 diabetes65.
Figure 7. Advanced therapeutic and prosthetic networks.
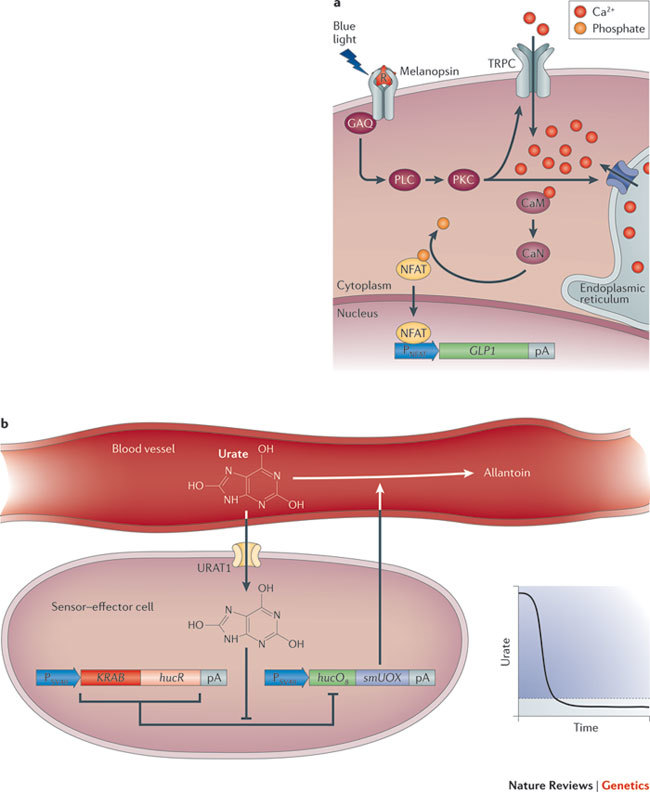
a | Light-triggered transcription control of blood glucose homeostasis. The synthetic phototransduction cascade consists of rewired melanopsin and nuclear factor of activated T cells (NFAT) control circuits. Photo-isomerization of the 11-cis-retinal chromophore (R) by blue light (~480 nm) activates melanopsin. This sequentially turns on Gaq-type G protein (GAQ), phospholipase C (PLC) and phosphokinase C (PKC) and triggers Ca2+ ion influx via transient receptor potential channels (TRPCs) and possibly also from the endoplasmic reticulum. This Ca2+ ion surge activates calmodulin (CaM) to calcineurin (CaN), which dephosphorylates NFAT. NFAT then translocates into the nucleus, where it binds to specific promoters (PNFAT) and coordinates transgene transcription. When linked to the glucagon-like peptide (GLP1), this mechanism allowed light-controlled blood glucose homeostasis to be achieved in a mouse model of type 2 diabetes. b | Prosthetic network for the treatment of tumour lysis syndrome and gout. Implanted sensor–effector cells are used to monitor serum urate levels constantly: they import urate via a transgenic human uric acid transporter (URAT1). Urate prevents binding of the uric acid-sensitive transsilencer (KRAB–HucR, which is the uricase regulator linked to a KRAB domain) to its operator (hucO8). This operator controls expression of secretion-engineered urate oxidase (smUOX), so smUOX is expressed when urate concentration reaches pathological levels. smUOX mediates conversion of urate into allantoin. Expression of smUOX stops when urate concentration reaches oxidative-stress-protective urate levels. pA, poly(A) tail. Part a is modified, with permission, from Ref. 65 © (2011) American Academy for the Advancement of Science.
Prosthetic networks. Prosthetic networks are synthetic sensor–effector devices that act as molecular prostheses. When engineered into cells and functionally connected to host metabolism, they sense, monitor and score disease-relevant metabolites, process off-level concentrations and coordinate adjusted diagnostic, preventive or therapeutic responses in a seamless, automatic and self-sufficient manner.
An example of the use of a prosthetic network is the sensing of metabolites to improve control of urate homeostasis (Fig. 7b). Moderate levels of uric acid, which scavenges radicals, are deemed to be beneficial. However, a transient surge in uric acid that is released by dying cells during cancer therapy leads to tumour lysis syndrome, and chronic hyperuricaemia can result in gout. Humans are particularly sensitive to imbalances of urate homeostasis because they lack uricolytic activity. A prosthetic network that constantly monitors blood urate concentrations and restores urate homeostasis by controlled expression of a urate oxidase — which reduces excessive urate concentration while preserving levels that are suitable for radical scavenging — could represent a treatment strategy for hyperuricaemic disorders64. In brief, human cells that contain such a prosthetic network have recently been designed by combining: the uric acid sensor HucR154, which manages oxidative stress protection in Deinococcus radiodurans; the human uric acid transporter URAT1 (also known as SLC22A12), which increases the intracellular uric acid levels and thus the sensitivity of the prosthetic circuit; and a secretion-engineered urate oxidase (smUOX) that is clinically licensed for the treatment of the tumour lysis syndrome155. The prosthetic uric-acid-responsive expression network (UREX) was able to sense uric acid concentrations precisely and to activate secretion of smUOX when the uric acid concentration was at pathologic levels. Secretion of smUOX is stopped as soon as the uric acid concentration has returned to the homeostasis level. This was impressively demonstrated when UREX-transgenic cells were implanted into urate-oxidase-deficient mice, which develop acute hyperuricaemia with symptoms that are similar to human gout. UREX was able to degrade urate and restore urate homeostasis in the blood, resulting in the dissolution of uric acid crystal deposits in the kidney of treated animals64. Its straightforward design may allow UREX to serve as a blueprint for the assembly of other prosthetic networks that sense metabolic disturbances and circulating pathologic metabolites.
An artificial insemination device. Artificial insemination is standard practice to facilitate both human reproduction and livestock breeding. Because of broad variations in oestrus expression and ovulation timing, coordinating sperm delivery with female oestrus is still a major challenge. Ovulation is triggered at a specific time when the pituitary gland releases luteinizing hormone, which binds to the luteinizing hormone receptor (LHR) and coordinates the release of the oocyte. By integrating synthetic signalling cascades with advanced biomaterials, Kemmer and colleagues63 designed an artificial insemination device that coordinates sperm delivery with oestrus control. The artificial insemination device consists of cellulose sulphate capsules containing bull sperm and sensor cells63,156,157. The sensor cells are engineered to express LHR constitutively so that when it is activated, it triggers expression of cellulase that can be secreted. After implantation into the cow's uterus, the sensor cell line constantly monitors the animal's luteinizing hormone levels, and the oestrus-triggered surge in luteinizing hormone levels leads to the production of secreted cellulase, which degrades the implanted capsule and results in the timely delivery of the sperm and successful conception. Fine tuning of the designer cascade could enable its use in other species, including in humans.
Perspectives and conclusions
In recent years, synthetic biology has substantially advanced strategies for classical biomedical applications, such as pathogen characterization68,70,74, disease analysis78,79,80, diagnostics75,76,77, screening assays33,92,94,100,101, drug production105,106,107,108,158 and vaccination81,82,85. This progress may imminently develop into shorter drug discovery94,98 and drug development timelines, increased precision of drug delivery112,114 and production of new and more affordable medicines104,105,106,107,108,109. Ultimately, sophisticated therapeutic sensor–effector devices that can sense disturbances, seek out pathological conditions and restore function are on the roadmap. Such therapeutic networks that connect diagnostic input with therapeutic output may provide all-in-one diagnostic, preventive and therapeutic solutions in future gene- and cell-based therapies. Matching diagnostic outcome with high-end therapies has recently become a focus of the pharmaceutical industry, which has declared personalized medicine as the treatment strategy of the future. Tools that will have a tremendous impact in future biomedical applications include: using light-activated triggers to bring about a precise therapeutic response in cells65, programming bacteria to seek and destroy cancer cells128,132,133 and using synthetic circuitry to keep crucial metabolites at homeostatic levels64,65, to manage disease-controlled expansion145 or to eliminate specific cell populations31,148,159. Recent work has shown that this is, in principle, possible and that some devices are working as expected and are producing a therapeutic impact in animal models of human diseases64,65. Implants consisting of engineered microencapsulated cells represent a way of introducing prosthetic networks with a predefined function instead of directly targeting the host cells with the genetic material. Although implants containing cells with engineered prosthetic networks are certainly the most promising way forward, they will limit biomedical applications to extracellular disease metabolites that can be therapeutically addressed through the vascular system.
However, there is still a long way to go until synthetic-biology-based biomedical devices will be a clinical reality. Placing therapeutic circuits in specific cells of a patient and making sure that there will be no interference with human metabolism are the most important challenges. Therefore, clinical use of synthetic-biology-based devices and therapeutic scenarios will face the same scientific, ethical and legal issues as any gene- and cell-based therapy, but they may offer more complex control dynamics and are therefore expected to have a higher therapeutic impact. Although none of the synthetic devices, prosthetic networks and related products that are pioneered by synthetic biologists have yet been used in the clinics or in clinical trials, the stage is set — with novel treatment strategies available and the commitment of the pharmaceutical industry in place — for synthetic biology to deliver the biomedicines of the twenty-first century.
Acknowledgements
Work in the laboratory of W.W. is supported by the European Research Council (ERC) under the European Community's Seventh Framework Programme (FP7/2007-2013) ERC Grant agreement number 259043-CompBioMat and the Excellence Initiative of the German Federal and State Governments (EXC 294). Work in the laboratory of M.F. is supported by the Swiss National Science Foundation (grant number 3100A0-112549) and in part by the European Community Framework 7 (Persist).
Glossary
- Aptamer
An oligonucleic acid that binds to a specific target, such as a chemical compound, a protein or a nucleic acid. Aptamers were found to control riboswitches in natural systems, but they can also be selected in vitro.
- Epigenetic toggle switches
Genetic circuits that can be switched between two stable expression states (for example, an 'on' and an 'off' state) by a transient stimulus. In the absence of a switching stimulus, the expression state is locked and inherited across cell generations.
- Hysteretic circuits
Genetic circuits whose response dynamics depend on a combination of past and present states.
- Eco-sensing
The capacity of a species to sense and score its surrounding ecosystem, for example, to identify the type and population density of neighbouring species.
- Quorum sensing
A small-molecule-based chemical language by which bacteria communicate within and across populations (the 'quorum'). Production and response to quorum-sensing molecules is correlated with population density.
- Band-pass filters
These are devices that are selectively induced within a specific concentration range of the input signal. At lower or higher trigger levels, the band-pass filter shuts down output signals.
- Immunostimulatory liposomes
Liposomes that are decorated with antigenic peptides or proteins that elicit an immune response.
- Gene drive system
These are molecular devices that promote the spreading of a specific gene throughout a target population by taking advantage of a mechanism that multiplies the specific gene in the host genome. Gene drive systems produce non-Mendelian patterns of inheritance.
- Polyketides
These constitute a group of secondary metabolites produced through linear decarboxylative condensation of acetyl-CoA with several malonyl-CoA-derived extender units to a polyketide chain. Many pharmacologically active compounds, such as antibiotics and anti-cancer drugs, belong to the polyketide class.
- SOS DNA repair
Genetically encoded repair program protecting against DNA damage. In prokaryotes, the repair program is coordinated by LexA and RecA.
- Persister cells
Dormant individual cells within a bacterial population that show a high tolerance to antimicrobials.
- Pharmacokinetics
The action of drugs in the body over a period of time. It covers absorption of the drug as well as its distribution, tissue localization, biotransformation and excretion.
- Commensal bacteria
Commensal bacteria live in close contact with the host. In this special type of symbiosis, one partner is benefited, whereas the other is neither benefited nor harmed.
- Melanopsin
A vitamin-A-dependent, G-protein-coupled receptor that is expressed in intrinsically photosensitive retinal ganglion cells
- Prosthetic networks
Networks that replace existing cellular functionality that is ill-driven or out of order. They represent molecular prostheses for non-functional cellular activity; they differ from other synthetic networks that add useful functionality but do not replace non-functional cellular networks.
Biographies
Wilfried Weber is Professor of Synthetic Biology at the University of Freiburg, Germany. Wilfried studied Biochemistry and Biotechnology and earned his Ph.D. with Martin Fussenegger at the Eigenössische Technische Hochschule (ETH) Zurich, Switzerland, where he also started his independent research group. His research interest focuses on applying synthetic biology principles for drug discovery as well as for the design of interactive biohybrid materials for biomedical applications.
Martin Fussenegger is Professor of Biotechnology and Bioengineering at the Department of Biosystems Science and Engineering (D-BSSE) of the ETH Zurich in Basel, Switzerland. Martin graduated with Werner Arber at the Biocenter of the Basel University, did his Ph.D. at the Max Planck Institute of Biology, Tübingen, Germany, and a postdoctoral fellowship at the MPI for Infection Biology, Berlin, Germany. He joined the ETH Zurich as junior group leader and served in different management positions as professor at the ETH Zurich, most recently he moved to Basel to coordinate the build-up of the D-BSSE. His research focuses on mammalian cell engineering to reach therapeutic goals.
Related links
FURTHER INFORMATION
Competing interests
The authors declare no competing financial interests.
References
- 1.Arkin AP, Schaffer DV. Network news: innovations in 21st century systems biology. Cell. 2011;144:844–849. doi: 10.1016/j.cell.2011.03.008. [DOI] [PubMed] [Google Scholar]
- 2.Isalan M, et al. Evolvability and hierarchy in rewired bacterial gene networks. Nature. 2008;452:840–845. doi: 10.1038/nature06847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Roy S, et al. Identification of functional elements and regulatory circuits by Drosophila modENCODE. Science. 2010;330:1787–1797. doi: 10.1126/science.1198374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Smolke CD, Silver PA. Informing biological design by integration of systems and synthetic biology. Cell. 2011;144:855–859. doi: 10.1016/j.cell.2011.02.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Jasny BR, Zahn LM. Genome-sequencing anniversary. A celebration of the genome, part I. Science. 2011;331:546. doi: 10.1126/science.331.6017.546-a. [DOI] [PubMed] [Google Scholar]
- 6.Nielsen R, Paul JS, Albrechtsen A, Song YS. Genotype and SNP calling from next-generation sequencing data. Nature Rev. Genet. 2011;12:443–451. doi: 10.1038/nrg2986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kosuri S, et al. Scalable gene synthesis by selective amplification of DNA pools from high-fidelity microchips. Nature Biotech. 2010;28:1295–1299. doi: 10.1038/nbt.1716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Matzas M, et al. High-fidelity gene synthesis by retrieval of sequence-verified DNA identified using high-throughput pyrosequencing. Nature Biotech. 2010;28:1291–1294. doi: 10.1038/nbt.1710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Quan J, et al. Parallel on-chip gene synthesis and application to optimization of protein expression. Nature Biotech. 2011;29:449–452. doi: 10.1038/nbt.1847. [DOI] [PubMed] [Google Scholar]
- 10.Lartigue C, et al. Creating bacterial strains from genomes that have been cloned and engineered in yeast. Science. 2009;325:1693–1696. doi: 10.1126/science.1173759. [DOI] [PubMed] [Google Scholar]
- 11.Barak M, et al. Evidence for large diversity in the human transcriptome created by Alu RNA editing. Nucleic Acids Res. 2009;37:6905–6915. doi: 10.1093/nar/gkp729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cartier N, et al. Hematopoietic stem cell gene therapy with a lentiviral vector in X-linked adrenoleukodystrophy. Science. 2009;326:818–823. doi: 10.1126/science.1171242. [DOI] [PubMed] [Google Scholar]
- 13.Gibson DG, et al. Creation of a bacterial cell controlled by a chemically synthesized genome. Science. 2010;329:52–56. doi: 10.1126/science.1190719. [DOI] [PubMed] [Google Scholar]
- 14.Bedau M, et al. Life after the synthetic cell. Nature. 2010;465:422–424. doi: 10.1038/465422a. [DOI] [PubMed] [Google Scholar]
- 15.Burrill DR, Silver PA. Making cellular memories. Cell. 2010;140:13–18. doi: 10.1016/j.cell.2009.12.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Danino T, Mondragon-Palomino O, Tsimring L, Hasty J. A synchronized quorum of genetic clocks. Nature. 2010;463:326–330. doi: 10.1038/nature08753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ellis T, Wang X, Collins JJ. Diversity-based, model-guided construction of synthetic gene networks with predicted functions. Nature Biotech. 2009;27:465–471. doi: 10.1038/nbt.1536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Elowitz MB, Leibler S. A synthetic oscillatory network of transcriptional regulators. Nature. 2000;403:335–338. doi: 10.1038/35002125. [DOI] [PubMed] [Google Scholar]
- 19.Fung E, et al. A synthetic gene-metabolic oscillator. Nature. 2005;435:118–122. doi: 10.1038/nature03508. [DOI] [PubMed] [Google Scholar]
- 20.Gardner TS, Cantor CR, Collins JJ. Construction of a genetic toggle switch in Escherichia coli. Nature. 2000;403:339–342. doi: 10.1038/35002131. [DOI] [PubMed] [Google Scholar]
- 21.Greber, D. & Fussenegger, M. An engineered mammalian band-pass network. Nucleic Acids Res. 38, e174 (2010). [DOI] [PMC free article] [PubMed]
- 22.Kramer BP, Fussenegger M. Hysteresis in a synthetic mammalian gene network. Proc. Natl Acad. Sci. USA. 2005;102:9517–9522. doi: 10.1073/pnas.0500345102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kramer BP, et al. An engineered epigenetic transgene switch in mammalian cells. Nature Biotech. 2004;22:867–870. doi: 10.1038/nbt980. [DOI] [PubMed] [Google Scholar]
- 24.Leisner M, Bleris L, Lohmueller J, Xie Z, Benenson Y. Rationally designed logic integration of regulatory signals in mammalian cells. Nature Nanotechnol. 2010;5:666–670. doi: 10.1038/nnano.2010.135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rinaudo K, et al. A universal RNAi-based logic evaluator that operates in mammalian cells. Nature Biotech. 2007;25:795–801. doi: 10.1038/nbt1307. [DOI] [PubMed] [Google Scholar]
- 26.Stricker J, et al. A fast, robust and tunable synthetic gene oscillator. Nature. 2008;456:516–519. doi: 10.1038/nature07389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Swinburne IA, Miguez DG, Landgraf D, Silver PA. Intron length increases oscillatory periods of gene expression in animal cells. Genes Dev. 2008;22:2342–2346. doi: 10.1101/gad.1696108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Tigges M, Marquez-Lago TT, Stelling J, Fussenegger M. A tunable synthetic mammalian oscillator. Nature. 2009;457:309–312. doi: 10.1038/nature07616. [DOI] [PubMed] [Google Scholar]
- 29.Toettcher JE, Mock C, Batchelor E, Loewer A, Lahav G. A synthetic-natural hybrid oscillator in human cells. Proc. Natl Acad. Sci. USA. 2010;107:17047–17052. doi: 10.1073/pnas.1005615107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Weber W, et al. A synthetic time-delay circuit in mammalian cells and mice. Proc. Natl Acad. Sci. USA. 2007;104:2643–2648. doi: 10.1073/pnas.0606398104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Xie Z, Wroblewska L, Prochazka L, Weiss R, Benenson Y. Multi-input RNAi-based logic circuit for identification of specific cancer cells. Science. 2011;333:1307–1311. doi: 10.1126/science.1205527. [DOI] [PubMed] [Google Scholar]
- 32.Nandagopal N, Elowitz MB. Synthetic biology: integrated gene circuits. Science. 2011;333:1244–1248. doi: 10.1126/science.1207084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Aubel D, Fussenegger M. Mammalian synthetic biology—from tools to therapies. Bioessays. 2010;32:332–345. doi: 10.1002/bies.200900149. [DOI] [PubMed] [Google Scholar]
- 34.Tigges M, Fussenegger M. Recent advances in mammalian synthetic biology — design of synthetic transgene control networks. Curr. Opin. Biotechnol. 2009;20:449–460. doi: 10.1016/j.copbio.2009.07.009. [DOI] [PubMed] [Google Scholar]
- 35.Weber W, Fussenegger M. Synthetic gene networks in mammalian cells. Curr. Opin. Biotechnol. 2010;692:235–249. doi: 10.1016/j.copbio.2010.07.006. [DOI] [PubMed] [Google Scholar]
- 36.Weber W, Fussenegger M. Molecular diversity—the toolbox for synthetic gene switches and networks. Curr. Opin. Chem. Biol. 2011;15:414–420. doi: 10.1016/j.cbpa.2011.03.003. [DOI] [PubMed] [Google Scholar]
- 37.Benenson Y. RNA-based computation in live cells. Curr. Opin. Biotechnol. 2009;20:471–478. doi: 10.1016/j.copbio.2009.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Wieland M, Fussenegger M. Ligand-dependent regulatory RNA parts for synthetic biology in eukaryotes. Curr. Opin. Biotechnol. 2010;21:760–765. doi: 10.1016/j.copbio.2010.06.010. [DOI] [PubMed] [Google Scholar]
- 39.Win MN, Liang JC, Smolke CD. Frameworks for programming biological function through RNA parts and devices. Chem. Biol. 2009;16:298–310. doi: 10.1016/j.chembiol.2009.02.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Canton B, Labno A, Endy D. Refinement and standardization of synthetic biological parts and devices. Nature Biotech. 2008;26:787–793. doi: 10.1038/nbt1413. [DOI] [PubMed] [Google Scholar]
- 41.Greber D, Fussenegger M. Mammalian synthetic biology: engineering of sophisticated gene networks. J. Biotechnol. 2007;130:329–345. doi: 10.1016/j.jbiotec.2007.05.014. [DOI] [PubMed] [Google Scholar]
- 42.Hooshangi S, Thiberge S, Weiss R. Ultrasensitivity and noise propagation in a synthetic transcriptional cascade. Proc. Natl Acad. Sci. USA. 2005;102:3581–3586. doi: 10.1073/pnas.0408507102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Kramer BP, Fischer M, Fussenegger M. Semi-synthetic mammalian gene regulatory networks. Metab. Eng. 2005;7:241–250. doi: 10.1016/j.ymben.2005.02.005. [DOI] [PubMed] [Google Scholar]
- 44.Greber D, El-Baba MD, Fussenegger M. Intronically encoded siRNAs improve dynamic range of mammalian gene regulation systems and toggle switch. Nucleic Acids Res. 2008;36:e101. doi: 10.1093/nar/gkn443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Yao G, Tan C, West M, Nevins JR, You L. Origin of bistability underlying mammalian cell cycle entry. Mol. Syst. Biol. 2011;7:485. doi: 10.1038/msb.2011.19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Weber W, Daoud-El Baba M, Fussenegger M. Synthetic ecosystems based on airborne inter- and intrakingdom communication. Proc. Natl Acad. Sci. USA. 2007;104:10435–10440. doi: 10.1073/pnas.0701382104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Fussenegger M, et al. Streptogramin-based gene regulation systems for mammalian cells. Nature Biotech. 2000;18:1203–1208. doi: 10.1038/81208. [DOI] [PubMed] [Google Scholar]
- 48.Gitzinger M, Kemmer C, El-Baba MD, Weber W, Fussenegger M. Controlling transgene expression in subcutaneous implants using a skin lotion containing the apple metabolite phloretin. Proc. Natl Acad. Sci. USA. 2009;106:10638–10643. doi: 10.1073/pnas.0901501106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Karlsson M, Weber W, Fussenegger M. De novo design and construction of an inducible gene expression system in mammalian cells. Methods Enzymol. 2011;497:239–253. doi: 10.1016/B978-0-12-385075-1.00011-1. [DOI] [PubMed] [Google Scholar]
- 50.Atkinson MR, Savageau MA, Myers JT, Ninfa AJ. Development of genetic circuitry exhibiting toggle switch or oscillatory behavior in Escherichia coli. Cell. 2003;113:597–607. doi: 10.1016/s0092-8674(03)00346-5. [DOI] [PubMed] [Google Scholar]
- 51.Ozbudak EM, Thattai M, Lim HN, Shraiman BI, Van Oudenaarden A. Multistability in the lactose utilization network of Escherichia coli. Nature. 2004;427:737–740. doi: 10.1038/nature02298. [DOI] [PubMed] [Google Scholar]
- 52.Weber W, Kramer BP, Fussenegger M. A genetic time-delay circuitry in mammalian cells. Biotechnol. Bioeng. 2007;98:894–902. doi: 10.1002/bit.21463. [DOI] [PubMed] [Google Scholar]
- 53.Brenner K, Karig DK, Weiss R, Arnold FH. Engineered bidirectional communication mediates a consensus in a microbial biofilm consortium. Proc. Natl Acad. Sci. USA. 2007;104:17300–17304. doi: 10.1073/pnas.0704256104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.You L, Cox RS, Weiss R, Arnold FH. Programmed population control by cell–cell communication and regulated killing. Nature. 2004;428:868–871. doi: 10.1038/nature02491. [DOI] [PubMed] [Google Scholar]
- 55.Basu S, Gerchman Y, Collins CH, Arnold FH, Weiss R. A synthetic multicellular system for programmed pattern formation. Nature. 2005;434:1130–1134. doi: 10.1038/nature03461. [DOI] [PubMed] [Google Scholar]
- 56.Aubel D, Fussenegger M. Watch the clock — engineering biological systems to be on time. Curr. Opin. Genet. Dev. 2010;20:634–643. doi: 10.1016/j.gde.2010.09.003. [DOI] [PubMed] [Google Scholar]
- 57.Tigges M, Denervaud N, Greber D, Stelling J, Fussenegger M. A synthetic low-frequency mammalian oscillator. Nucleic Acids Res. 2010;38:2702–2711. doi: 10.1093/nar/gkq121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Agapakis CM, Silver PA. Synthetic biology: exploring and exploiting genetic modularity through the design of novel biological networks. Mol. Biosyst. 2009;5:704–713. doi: 10.1039/b901484e. [DOI] [PubMed] [Google Scholar]
- 59.Haynes KA, Silver PA. Eukaryotic systems broaden the scope of synthetic biology. J. Cell Biol. 2009;187:589–596. doi: 10.1083/jcb.200908138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Khalil AS, Collins JJ. Synthetic biology: applications come of age. Nature Rev. Genet. 2010;11:367–379. doi: 10.1038/nrg2775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Lu TK, Khalil AS, Collins JJ. Next-generation synthetic gene networks. Nature Biotech. 2009;27:1139–1150. doi: 10.1038/nbt.1591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Purnick PE, Weiss R. The second wave of synthetic biology: from modules to systems. Nature Rev. Mol. Cell Biol. 2009;10:410–422. doi: 10.1038/nrm2698. [DOI] [PubMed] [Google Scholar]
- 63.Kemmer C, et al. A designer network coordinating bovine artificial insemination by ovulation-triggered release of implanted sperms. J. Control. Release. 2011;150:23–29. doi: 10.1016/j.jconrel.2010.11.016. [DOI] [PubMed] [Google Scholar]
- 64.Kemmer C, et al. Self-sufficient control of urate homeostasis in mice by a synthetic circuit. Nature Biotech. 2010;28:355–360. doi: 10.1038/nbt.1617. [DOI] [PubMed] [Google Scholar]
- 65.Ye H, Daoud-El Baba M, Peng RW, Fussenegger M. A synthetic optogenetic transcription device enhances blood-glucose homeostasis in mice. Science. 2011;332:1565–1568. doi: 10.1126/science.1203535. [DOI] [PubMed] [Google Scholar]
- 66.Fears R, ter Meulen V. The potential of synthetic biology: a view from the European Academies Science Advisory Council. Nature Rev. Microbiol. 2011;9:222. doi: 10.1038/nrmicro2498-c1. [DOI] [PubMed] [Google Scholar]
- 67.Ruder WC, Lu T, Collins JJ. Synthetic biology moving into the clinic. Science. 2011;333:1248–1252. doi: 10.1126/science.1206843. [DOI] [PubMed] [Google Scholar]
- 68.Tumpey TM, et al. Characterization of the reconstructed 1918 Spanish influenza pandemic virus. Science. 2005;310:77–80. doi: 10.1126/science.1119392. [DOI] [PubMed] [Google Scholar]
- 69.Gibson DG, et al. Complete chemical synthesis, assembly, and cloning of a Mycoplasma genitalium genome. Science. 2008;319:1215–1220. doi: 10.1126/science.1151721. [DOI] [PubMed] [Google Scholar]
- 70.Tumpey TM, et al. A two-amino acid change in the hemagglutinin of the 1918 influenza virus abolishes transmission. Science. 2007;315:655–659. doi: 10.1126/science.1136212. [DOI] [PubMed] [Google Scholar]
- 71.Cohen J, Enserink M. Infectious diseases. As swine flu circles globe, scientists grapple with basic questions. Science. 2009;324:572–573. doi: 10.1126/science.324_572. [DOI] [PubMed] [Google Scholar]
- 72.Smith GJ, et al. Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature. 2009;459:1122–1125. doi: 10.1038/nature08182. [DOI] [PubMed] [Google Scholar]
- 73.Wimmer E, Mueller S, Tumpey TM, Taubenberger JK. Synthetic viruses: a new opportunity to understand and prevent viral disease. Nature Biotech. 2009;27:1163–1172. doi: 10.1038/nbt.1593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Becker MM, et al. Synthetic recombinant bat SARS-like coronavirus is infectious in cultured cells and in mice. Proc. Natl Acad. Sci. USA. 2008;105:19944–19949. doi: 10.1073/pnas.0808116105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Burbelo PD, Ching KH, Bush ER, Han BL, Iadarola MJ. Antibody-profiling technologies for studying humoral responses to infectious agents. Expert Rev. Vaccines. 2010;9:567–578. doi: 10.1586/erv.10.50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Chandra A, Wormser GP, Marques AR, Latov N, Alaedini A. Anti-Borrelia burgdorferi antibody profile in post-Lyme disease syndrome. Clin. Vaccine Immunol. 2011;18:767–771. doi: 10.1128/CVI.00002-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Burbelo PD, et al. LIPS arrays for simultaneous detection of antibodies against partial and whole proteomes of HCV, HIV and EBV. Mol. Biosyst. 2011;7:1453–1462. doi: 10.1039/c0mb00342e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Ferrari S, et al. Mutations of the Igbeta gene cause agammaglobulinemia in man. J. Exp. Med. 2007;204:2047–2051. doi: 10.1084/jem.20070264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Yang J, Reth M. Oligomeric organization of the B-cell antigen receptor on resting cells. Nature. 2010;467:465–469. doi: 10.1038/nature09357. [DOI] [PubMed] [Google Scholar]
- 80.Larman HB, et al. Autoantigen discovery with a synthetic human peptidome. Nature Biotech. 2011;29:535–541. doi: 10.1038/nbt.1856. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Akahata W, et al. A virus-like particle vaccine for epidemic Chikungunya virus protects nonhuman primates against infection. Nature Med. 2010;16:334–338. doi: 10.1038/nm.2105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Coleman JR, et al. Virus attenuation by genome-scale changes in codon pair bias. Science. 2008;320:1784–1787. doi: 10.1126/science.1155761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Amidi M, et al. Optimization and quantification of protein synthesis inside liposomes. J. Liposome Res. 2010;20:73–83. doi: 10.3109/08982100903402954. [DOI] [PubMed] [Google Scholar]
- 84.Dance A. From pond scum to pharmacy shelf. Nature Med. 2010;16:146–149. doi: 10.1038/nm0210-146. [DOI] [PubMed] [Google Scholar]
- 85.Dreesen IA, Charpin-El Hamri G, Fussenegger M. Heat-stable oral alga-based vaccine protects mice from Staphylococcus aureus infection. J. Biotechnol. 2010;145:273–280. doi: 10.1016/j.jbiotec.2009.12.006. [DOI] [PubMed] [Google Scholar]
- 86.Fu G, et al. Female-specific insect lethality engineered using alternative splicing. Nature Biotech. 2007;25:353–357. doi: 10.1038/nbt1283. [DOI] [PubMed] [Google Scholar]
- 87.Fu G, et al. Female-specific flightless phenotype for mosquito control. Proc. Natl Acad. Sci. USA. 2010;107:4550–4554. doi: 10.1073/pnas.1000251107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Wise de Valdez MR, et al. Genetic elimination of dengue vector mosquitoes. Proc. Natl Acad. Sci. USA. 2011;108:4772–4775. doi: 10.1073/pnas.1019295108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Windbichler N, et al. A synthetic homing endonuclease-based gene drive system in the human malaria mosquito. Nature. 2011;473:212–215. doi: 10.1038/nature09937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Subbaraman N. Science snipes at Oxitec transgenic-mosquito trial. Nature Biotech. 2011;29:9–11. doi: 10.1038/nbt0111-9a. [DOI] [PubMed] [Google Scholar]
- 91.Weber W, et al. Macrolide-based transgene control in mammalian cells and mice. Nature Biotech. 2002;20:901–907. doi: 10.1038/nbt731. [DOI] [PubMed] [Google Scholar]
- 92.Aubel D, et al. Design of a novel mammalian screening system for the detection of bioavailable, non-cytotoxic streptogramin antibiotics. J. Antibiot. 2001;54:44–55. doi: 10.7164/antibiotics.54.44. [DOI] [PubMed] [Google Scholar]
- 93.Weber W, Fussenegger M. The impact of synthetic biology on drug discovery. Drug Discov. Today. 2009;14:956–963. doi: 10.1016/j.drudis.2009.06.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Weber W, et al. A synthetic mammalian gene circuit reveals antituberculosis compounds. Proc. Natl Acad. Sci. USA. 2008;105:9994–9998. doi: 10.1073/pnas.0800663105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Tascou S, et al. Stringent rosiglitazone-dependent gene switch in muscle cells without effect on myogenic differentiation. Mol. Ther. 2004;9:637–649. doi: 10.1016/j.ymthe.2004.02.013. [DOI] [PubMed] [Google Scholar]
- 96.Chong H, Ruchatz A, Clackson T, Rivera VM, Vile RG. A system for small-molecule control of conditionally replication-competent adenoviral vectors. Mol. Ther. 2002;5:195–203. doi: 10.1006/mthe.2002.0531. [DOI] [PubMed] [Google Scholar]
- 97.Weber W, et al. Streptomyces-derived quorum-sensing systems engineered for adjustable transgene expression in mammalian cells and mice. Nucleic Acids Res. 2003;31:e71. doi: 10.1093/nar/gng071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Willand N, et al. Synthetic EthR inhibitors boost antituberculous activity of ethionamide. Nature Med. 2009;15:537–544. doi: 10.1038/nm.1950. [DOI] [PubMed] [Google Scholar]
- 99.Fussenegger M, Schlatter S, Datwyler D, Mazur X, Bailey JE. Controlled proliferation by multigene metabolic engineering enhances the productivity of Chinese hamster ovary cells. Nature Biotech. 1998;16:468–472. doi: 10.1038/nbt0598-468. [DOI] [PubMed] [Google Scholar]
- 100.Gonzalez-Nicolini V, Fussenegger M. In vitro assays for anticancer drug discovery—a novel approach based on engineered mammalian cell lines. Anticancer Drugs. 2005;16:223–228. doi: 10.1097/00001813-200503000-00001. [DOI] [PubMed] [Google Scholar]
- 101.Gonzalez-Nicolini V, Fux C, Fussenegger M. A novel mammalian cell-based approach for the discovery of anticancer drugs with reduced cytotoxicity on non-dividing cells. Invest. New Drugs. 2004;22:253–262. doi: 10.1023/B:DRUG.0000026251.00854.77. [DOI] [PubMed] [Google Scholar]
- 102.Li JW, Vederas JC. Drug discovery and natural products: end of an era or an endless frontier? Science. 2009;325:161–165. doi: 10.1126/science.1168243. [DOI] [PubMed] [Google Scholar]
- 103.Medema MH, Breitling R, Bovenberg R, Takano E. Exploiting plug-and-play synthetic biology for drug discovery and production in microorganisms. Nature Rev. Microbiol. 2011;9:131–137. doi: 10.1038/nrmicro2478. [DOI] [PubMed] [Google Scholar]
- 104.Noel JP. Chemical biology: synthetic metabolism goes green. Nature. 2010;468:380–381. doi: 10.1038/468380a. [DOI] [PubMed] [Google Scholar]
- 105.Runguphan W, Maresh JJ, O'Connor SE. Silencing of tryptamine biosynthesis for production of nonnatural alkaloids in plant culture. Proc. Natl Acad. Sci. USA. 2009;106:13673–13678. doi: 10.1073/pnas.0903393106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Runguphan W, Qu X, O'Connor SE. Integrating carbon-halogen bond formation into medicinal plant metabolism. Nature. 2010;468:461–464. doi: 10.1038/nature09524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Ro DK, et al. Production of the antimalarial drug precursor artemisinic acid in engineered yeast. Nature. 2006;440:940–943. doi: 10.1038/nature04640. [DOI] [PubMed] [Google Scholar]
- 108.Ajikumar PK, et al. Isoprenoid pathway optimization for Taxol precursor overproduction in Escherichia coli. Science. 2010;330:70–74. doi: 10.1126/science.1191652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Chang MC, Eachus RA, Trieu W, Ro DK, Keasling JD. Engineering Escherichia coli for production of functionalized terpenoids using plant P450s. Nature Chem. Biol. 2007;3:274–277. doi: 10.1038/nchembio875. [DOI] [PubMed] [Google Scholar]
- 110.Fussenegger M. The impact of mammalian gene regulation concepts on functional genomic research, metabolic engineering, and advanced gene therapies. Biotechnol. Prog. 2001;17:1–51. doi: 10.1021/bp000129c. [DOI] [PubMed] [Google Scholar]
- 111.Christen EH, et al. Conditional DNA–protein interactions confer stimulus-sensing properties to biohybrid materials. Adv. Funct. Mater. 2011;21:2861–2867. [Google Scholar]
- 112.Ehrbar M, Schoenmakers R, Christen EH, Fussenegger M, Weber W. Drug-sensing hydrogels for the inducible release of biopharmaceuticals. Nature Mater. 2008;7:800–804. doi: 10.1038/nmat2250. [DOI] [PubMed] [Google Scholar]
- 113.Ehrick JD, et al. Genetically engineered protein in hydrogels tailors stimuli-responsive characteristics. Nature Mater. 2005;4:298–302. doi: 10.1038/nmat1352. [DOI] [PubMed] [Google Scholar]
- 114.Jakobus Kathrin, Wend Sabrina, Weber Wilfried. Synthetic mammalian gene networks as a blueprint for the design of interactive biohybrid materials. Chem. Soc. Rev. 2012;41(3):1000–1018. doi: 10.1039/c1cs15176b. [DOI] [PubMed] [Google Scholar]
- 115.Kämpf MM, et al. A gene therapy technology-based hydrogel for the trigger inducible release of bipharmaceuticals in mice. Adv. Funct. Mater. 2010;20:2534–2538. [Google Scholar]
- 116.Zhao HF, Boyd J, Jolicoeur N, Shen SH. A coumermycin/novobiocin-regulated gene expression system. Hum. Gene Ther. 2003;14:1619–1629. doi: 10.1089/104303403322542266. [DOI] [PubMed] [Google Scholar]
- 117.Rivera VM, et al. Regulation of protein secretion through controlled aggregation in the endoplasmic reticulum. Science. 2000;287:826–830. doi: 10.1126/science.287.5454.826. [DOI] [PubMed] [Google Scholar]
- 118.Allison KR, Brynildsen MP, Collins JJ. Metabolite-enabled eradication of bacterial persisters by aminoglycosides. Nature. 2011;473:216–220. doi: 10.1038/nature10069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Lee HH, Molla MN, Cantor CR, Collins JJ. Bacterial charity work leads to population-wide resistance. Nature. 2010;467:82–85. doi: 10.1038/nature09354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Lu TK, Collins JJ. Dispersing biofilms with engineered enzymatic bacteriophage. Proc. Natl Acad. Sci. USA. 2007;104:11197–11202. doi: 10.1073/pnas.0704624104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Kohanski MA, DePristo MA, Collins JJ. Sublethal antibiotic treatment leads to multidrug resistance via radical-induced mutagenesis. Mol. Cell. 2010;37:311–320. doi: 10.1016/j.molcel.2010.01.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Kohanski MA, Dwyer DJ, Collins JJ. How antibiotics kill bacteria: from targets to networks. Nature Rev. Microbiol. 2010;8:423–435. doi: 10.1038/nrmicro2333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Kohanski MA, Dwyer DJ, Hayete B, Lawrence CA, Collins JJ. A common mechanism of cellular death induced by bactericidal antibiotics. Cell. 2007;130:797–810. doi: 10.1016/j.cell.2007.06.049. [DOI] [PubMed] [Google Scholar]
- 124.Lu TK, Collins JJ. Engineered bacteriophage targeting gene networks as adjuvants for antibiotic therapy. Proc. Natl Acad. Sci. USA. 2009;106:4629–4634. doi: 10.1073/pnas.0800442106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Fernebro J. Fighting bacterial infections-future treatment options. Drug Resist. Updat. 2011;14:125–139. doi: 10.1016/j.drup.2011.02.001. [DOI] [PubMed] [Google Scholar]
- 126.Duan F, March JC. Engineered bacterial communication prevents Vibrio cholerae virulence in an infant mouse model. Proc. Natl Acad. Sci. USA. 2010;107:11260–11264. doi: 10.1073/pnas.1001294107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Duan F, Curtis KL, March JC. Secretion of insulinotropic proteins by commensal bacteria: rewiring the gut to treat diabetes. Appl. Environ. Microbiol. 2008;74:7437–7438. doi: 10.1128/AEM.01019-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Forbes NS. Engineering the perfect (bacterial) cancer therapy. Nature Rev. Cancer. 2010;10:785–794. doi: 10.1038/nrc2934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Anderson JC, Clarke EJ, Arkin AP, Voigt CA. Environmentally controlled invasion of cancer cells by engineered bacteria. J. Mol. Biol. 2006;355:619–627. doi: 10.1016/j.jmb.2005.10.076. [DOI] [PubMed] [Google Scholar]
- 130.Nguyen VH, et al. Genetically engineered Salmonella typhimurium as an imageable therapeutic probe for cancer. Cancer Res. 2010;70:18–23. doi: 10.1158/0008-5472.CAN-09-3453. [DOI] [PubMed] [Google Scholar]
- 131.Ganai S, Arenas RB, Forbes NS. Tumour-targeted delivery of TRAIL using Salmonella typhimurium enhances breast cancer survival in mice. Br. J. Cancer. 2009;101:1683–1691. doi: 10.1038/sj.bjc.6605403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Royo JL, et al. In vivo gene regulation in Salmonella spp. by a salicylate-dependent control circuit. Nature Methods. 2007;4:937–942. doi: 10.1038/nmeth1107. [DOI] [PubMed] [Google Scholar]
- 133.Xiang S, Fruehauf J, Li CJ. Short hairpin RNA-expressing bacteria elicit RNA interference in mammals. Nature Biotech. 2006;24:697–702. doi: 10.1038/nbt1211. [DOI] [PubMed] [Google Scholar]
- 134.Cattaneo R, Miest T, Shashkova EV, Barry MA. Reprogrammed viruses as cancer therapeutics: targeted, armed and shielded. Nature Rev. Microbiol. 2008;6:529–540. doi: 10.1038/nrmicro1927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Link N, et al. Therapeutic protein transduction of mammalian cells and mice by nucleic acid-free lentiviral nanoparticles. Nucleic Acids Res. 2006;34:e16. doi: 10.1093/nar/gnj014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Voelkel C, et al. Protein transduction from retroviral Gag precursors. Proc. Natl Acad. Sci. USA. 2010;107:7805–7810. doi: 10.1073/pnas.0914517107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Zhang F, et al. Efficient construction of sequence-specific TAL effectors for modulating mammalian transcription. Nature Biotech. 2011;29:149–153. doi: 10.1038/nbt.1775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Dong JG, Fernandez-Maculet JC, Yang SF. Purification and characterization of 1-aminocyclopropane-1-carboxylate oxidase from apple fruit. Proc. Natl Acad. Sci. USA. 1992;89:9789–9793. doi: 10.1073/pnas.89.20.9789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Dorer DE, Nettelbeck DM. Targeting cancer by transcriptional control in cancer gene therapy and viral oncolysis. Adv. Drug Deliv. Rev. 2009;61:554–571. doi: 10.1016/j.addr.2009.03.013. [DOI] [PubMed] [Google Scholar]
- 140.Howard PL, Chia MC, Del Rizzo S, Liu FF, Pawson T. Redirecting tyrosine kinase signaling to an apoptotic caspase pathway through chimeric adaptor proteins. Proc. Natl Acad. Sci. USA. 2003;100:11267–11272. doi: 10.1073/pnas.1934711100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Mebratu Y, Tesfaigzi Y. How ERK1/2 activation controls cell proliferation and cell death: is subcellular localization the answer? Cell Cycle. 2009;8:1168–1175. doi: 10.4161/cc.8.8.8147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Nissim L, Bar-Ziv RH. A tunable dual-promoter integrator for targeting of cancer cells. Mol. Syst. Biol. 2010;6:444. doi: 10.1038/msb.2010.99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Link KH, Breaker RR. Engineering ligand-responsive gene-control elements: lessons learned from natural riboswitches. Gene Ther. 2009;16:1189–1201. doi: 10.1038/gt.2009.81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Wieland M, Benz A, Klauser B, Hartig JS. Artificial ribozyme switches containing natural riboswitch aptamer domains. Angew. Chem. Int. Edn Engl. 2009;48:2715–2718. doi: 10.1002/anie.200805311. [DOI] [PubMed] [Google Scholar]
- 145.Chen YY, Jensen MC, Smolke CD. Genetic control of mammalian T-cell proliferation with synthetic RNA regulatory systems. Proc. Natl Acad. Sci. USA. 2010;107:8531–8536. doi: 10.1073/pnas.1001721107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Carpenito C, et al. Control of large, established tumor xenografts with genetically retargeted human T cells containing CD28 and CD137 domains. Proc. Natl Acad. Sci. USA. 2009;106:3360–3365. doi: 10.1073/pnas.0813101106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Sadelain M, Brentjens R, Riviere I. The promise and potential pitfalls of chimeric antigen receptors. Curr. Opin. Immunol. 2009;21:215–223. doi: 10.1016/j.coi.2009.02.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Culler SJ, Hoff KG, Smolke CD. Reprogramming cellular behavior with RNA controllers responsive to endogenous proteins. Science. 2010;330:1251–1255. doi: 10.1126/science.1192128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Levskaya A, et al. Synthetic biology: engineering Escherichia coli to see light. Nature. 2005;438:441–442. doi: 10.1038/nature04405. [DOI] [PubMed] [Google Scholar]
- 150.Tabor JJ, Levskaya A, Voigt CA. Multichromatic control of gene expression in Escherichia coli. J. Mol. Biol. 2011;405:315–324. doi: 10.1016/j.jmb.2010.10.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Tabor JJ, et al. A synthetic genetic edge detection program. Cell. 2009;137:1272–1281. doi: 10.1016/j.cell.2009.04.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Yazawa M, Sadaghiani AM, Hsueh B, Dolmetsch RE. Induction of protein-protein interactions in live cells using light. Nature Biotech. 2009;27:941–945. doi: 10.1038/nbt.1569. [DOI] [PubMed] [Google Scholar]
- 153.Levskaya A, Weiner OD, Lim WA, Voigt CA. Spatiotemporal control of cell signalling using a light-switchable protein interaction. Nature. 2009;461:997–1001. doi: 10.1038/nature08446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Wilkinson SP, Grove A. HucR, a novel uric acid-responsive member of the MarR family of transcriptional regulators from Deinococcus radiodurans. J. Biol. Chem. 2004;279:51442–51450. doi: 10.1074/jbc.M405586200. [DOI] [PubMed] [Google Scholar]
- 155.Legoux R, et al. Cloning and expression in Escherichia coli of the gene encoding Aspergillus flavus urate oxidase. J. Biol. Chem. 1992;267:8565–8570. [PubMed] [Google Scholar]
- 156.Fluri DA, Kemmer C, Daoud-El Baba M, Fussenegger M. A novel system for trigger-controlled drug release from polymer capsules. J. Control. Release. 2008;131:211–219. doi: 10.1016/j.jconrel.2008.07.036. [DOI] [PubMed] [Google Scholar]
- 157.Weber W, Rimann M, Schafroth T, Witschi U, Fussenegger M. Design of high-throughput-compatible protocols for microencapsulation, cryopreservation and release of bovine spermatozoa. J. Biotechnol. 2006;123:155–163. doi: 10.1016/j.jbiotec.2005.11.008. [DOI] [PubMed] [Google Scholar]
- 158.Noel D, et al. Short-term BMP-2 expression is sufficient for in vivo osteochondral differentiation of mesenchymal stem cells. Stem Cells. 2004;22:74–85. doi: 10.1634/stemcells.22-1-74. [DOI] [PubMed] [Google Scholar]
- 159.Nissim L, Beatus T, Bar-Ziv R. An autonomous system for identifying and governing a cell's state in yeast. Phys. Biol. 2007;4:154–163. doi: 10.1088/1478-3975/4/3/002. [DOI] [PubMed] [Google Scholar]


