Abstract
Background
A multitask deep learning model might be useful in large epidemiologic studies wherein detailed structural assessment of osteoarthritis still relies on expert radiologists’ readings. The potential of such a model in clinical routine should be investigated.
Purpose
To develop a multitask deep learning model for grading radiographic hip osteoarthritis features on radiographs and compare its performance to that of attending-level radiologists.
Materials and Methods
This retrospective study analyzed hip joints seen on weight-bearing anterior-posterior pelvic radiographs from participants in the Osteoarthritis Initiative (OAI). Participants were recruited from February 2004 to May 2006 for baseline measurements, and follow-up was performed 48 months later. Femoral osteophytes (FOs), acetabular osteophytes (AOs), and joint-space narrowing (JSN) were graded as absent, mild, moderate, or severe according to the Osteoarthritis Research Society International atlas. Subchondral sclerosis and subchondral cysts were graded as present or absent. The participants were split at 80% (n = 3494), 10% (n = 437), and 10% (n = 437) by using split-sample validation into training, validation, and testing sets, respectively. The multitask neural network was based on DenseNet-161, a shared convolutional features extractor trained with multitask loss function. Model performance was evaluated in the internal test set from the OAI and in an external test set by using temporal and geographic validation consisting of routine clinical radiographs.
Results
A total of 4368 participants (mean age, 61.0 years ± 9.2 [standard deviation]; 2538 women) were evaluated (15 364 hip joints on 7738 weight-bearing anterior-posterior pelvic radiographs). The accuracy of the model for assessing these five features was 86.7% (1333 of 1538) for FOs, 69.9% (1075 of 1538) for AOs, 81.7% (1257 of 1538) for JSN, 95.8% (1473 of 1538) for subchondral sclerosis, and 97.6% (1501 of 1538) for subchondral cysts in the internal test set, and 82.7% (86 of 104) for FOS, 65.4% (68 of 104) for AOs, 80.8% (84 of 104) for JSN, 88.5% (92 of 104) for subchondral sclerosis, and 91.3% (95 of 104) for subchondral cysts in the external test set.
Conclusion
A multitask deep learning model is a feasible approach to reliably assess radiographic features of hip osteoarthritis.
© RSNA, 2020
Summary
A multitask deep learning model, trained with 15 364 hip joints and five hip osteoarthritis features per radiograph, solved multiple classification tasks simultaneously and assessed those features with a reliability similar to that of attending-level radiologists.
Key Results
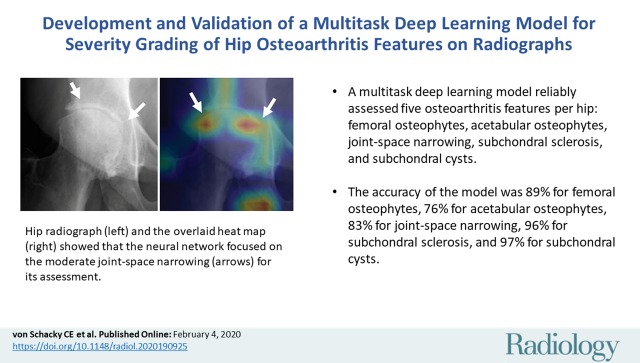
■ A multitask deep learning model can reliably assess five radiographic hip osteoarthritis features per joint on radiographs (femoral osteophytes [FOs], acetabular osteophyte [AOs], joint space narrowing [JSN], subchondral sclerosis, and subchondral cyst).
■ The accuracy of the model for assessing these five features varied depending on the evaluated feature: 89% for FOs, 76% for AOs, 83% for JSN, 96% for subchondral sclerosis, and 97% for subchondral cyst.
Introduction
Osteoarthritis is one of the most common chronic diseases, with more than 230 million individuals affected worldwide (1). Osteoarthritis is a whole-joint disorder that most commonly occurs in the knee and hip joints of middle-age to older people (2). For hip osteoarthritis, incidence, severity, and treatment have increased in the United States as the population has aged (3). The diagnosis and grading of the severity of hip osteoarthritis rely on a variety of clinical findings and findings at imaging. Radiography of the pelvis is the most commonly used primary imaging technique in patients suspected of having hip osteoarthritis (4). Radiographic features of hip osteoarthritis include joint-space narrowing (JSN), osteophytes, subchondral sclerosis, subchondral cysts, and flattening of the femoral head. Altman et al (5) published an atlas of individual radiographic features of osteoarthritis as a guideline for semiquantitatively grading these features. However, accurate assessment of these features is time consuming and requires expertise, and reproducibility in the hands of inexperienced or untrained readers is limited (6,7).
The potential benefit of artificial intelligence in clinical routine radiologic diagnostics remains to be investigated. Artificial intelligence may be particularly useful in the context of large epidemiologic studies that require detailed structural assessment by expert radiologists’ readings. A previous study (6) demonstrated the feasibility of deep learning–based algorithms to evaluate radiographs for the presence or absence of radiographic hip osteoarthritis. However, in clinical practice, the evaluation of disease severity in the affected joint is crucial for patient management. For radiographic knee osteoarthritis, studies have investigated the potential of deep learning to classify radiologic severity on radiographs by using Kellgren-Lawrence scores (8,9). Kellgren-Lawrence scores are condensed grading for the variety of associated radiographic osteoarthritis features. To allow for a thorough radiologic assessment of osteoarthritis, an artificial intelligence that could evaluate the multiple individual features of osteoarthritis on radiographs would be desirable and could help radiologists evaluate radiographs of hip osteoarthritis. In machine learning, this approach translates into multitask learning. Multitask learning is considered a main future trend for deep learning in radiology (10).
The goal of this study was to develop and validate a multitask deep learning approach to automatically extract radiographic features of osteoarthritis in the hip, specifically femoral osteophytes (FOs), acetabular osteophytes (AOs), JSN, subchondral sclerosis, and subchondral cysts, from radiographs and compare its performance to that of attending-level musculoskeletal radiologists.
Materials and Methods
All participants provided written informed consent. The institutional review boards of the four participating U.S. centers approved this Health Insurance Portability and Accountability Act–compliant study.
Data Sets
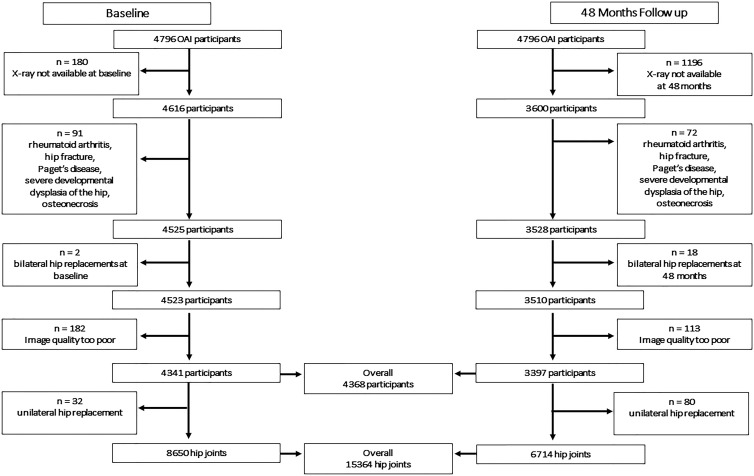
This retrospective study analyzed 15 364 hip joints by using 7738 weight-bearing anterior-posterior pelvic radiographs from 4368 participants of the Osteoarthritis Initiative (OAI), a prospective, observational study. Participants were recruited from February 2004 to May 2006 and formed a consecutive series. A total of 428 participants were excluded because of bilateral hip replacement, hip fracture, rheumatoid arthritis, Paget disease, severe development dysplasia, or poor image quality (Fig 1). Inclusion and exclusion criteria for OAI participants can be found in Appendix E1 (online).
Figure 1:
Flowchart showing participant selection from the Osteoarthritis Initiative (OAI) database.
Radiographs were obtained at baseline (n = 4341) and at 4-year follow-up (n = 3397). The participants were split by using split-sample validation into training, validation, and testing sets of 80% (n = 3494), 10% (n = 437), and 10% (n = 437), respectively.
For joint localization training, bounding boxes were placed at the center of the femoral head in a random subset of hips (n = 8776). Hips were split by using split-sample validation for training, validation, and testing at 80% (n = 7002), 10% (n = 877), and 10% (n = 897), respectively, with unique participants per data set.
Temporal and geographic validation was performed on a test set with clinical routine weight-bearing anterior-posterior pelvic radiographs from the University of California, San Francisco Medical Center (San Francisco, Calif), selected from a subgroup of patients who underwent hip injections in 2013 and 2014 (n = 36) and had clinical osteoarthritis and pain at the time of image acquisition, and a random selection of radiographs from December 2018 to January 2019 (n = 20). This resulted in 104 hip readings.
Image Readings
Clinical readings were specifically performed for this study, whereas OAI readings were performed for an ancillary OAI study (11). Readers were blinded to clinical information and other imaging results. FOs, AOs, JSN, subchondral sclerosis, and subchondral cysts were assessed as described in the Osteoarthritis Research Society International atlas (5). Medial and lateral JSN, superior and inferior AOs, superior and inferior FOs, acetabular and subchondral cysts, and acetabular and subchondral sclerosis were combined; the more severe grade served as the final grade. Subchondral sclerosis and subchondral cysts were graded as present or absent; and osteophytes and JSN were graded as absent, mild, moderate, or severe.
For OAI readings, two musculoskeletal imaging fellowship–trained radiologists (P.M.J. and L.N., with 5 and 7 years of experience reading pelvic radiographs, respectively) read all radiographs independently over 14 months. Initial training and calibration of readings were performed by a rheumatologist (L.N.) and a musculoskeletal imaging fellowship–trained radiologist (T.M.L.) (each with 25 years of experience in reading pelvic radiographs). Radiographs were assessed with both visits paired together in known chronologic order. In cases of agreement, these readings served as ground truth; in cases of disagreement, readings were adjudicated by a radiologist (T.M.L.) to establish ground truth. Interreader reliability between both readers was assessed for both points and each feature by using linearly weighted Cohen κ. When a reader determined that he or she could not accurately classify both joints of the pelvic radiograph for all features, the radiograph was excluded for poor image quality and was reviewed by the second reader, who confirmed this assessment for all radiographs.
For the external test set, readings were performed independently over 3 consecutive days by a musculoskeletal imaging fellowship–trained radiologist (M.P., with 8 years of experience in reading pelvic radiographs) and T.M.L. The median between both served as ground truth.
Model Architecture and Model Training
Preprocessing, model implementation, and evaluation were performed in Python 3.6 (open-source; Python Software Foundation, Wilmington, Del) by using 64-core Intel-Xeon Gold-6130-CPU at 2.10 GHz (Intel, Santa Clara, Calif), 276 GB DDR4-SDRAM and 4-T V100 32 GB graphical processing units (Nvidia, Santa Clara, Calif) running Linux system (Ubuntu 14.04; Canonical, London, England) with CUDA 9.0 (Nvidia).
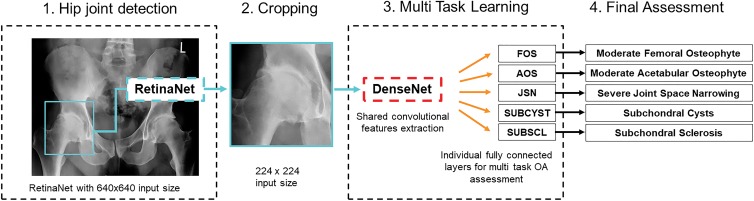
A Microsoft Common Objects in Context pretrained RetinaNet implemented in TensorFlow, version 1.7 (open source, Google Brain, Mountain View, Calif), was trained to depict both hip joints, excluding joints with total hip replacement (12,13). The detected image of one hip was cropped, contrast-stretched, and resized to 224 × 224 pixels. The multitask model was based on an ImageNet pretrained DenseNet-161 serving as a shared convolutional features extractor implemented in PyTorch (open source) 0.41 and fastai 1.03 (open source) using a multitask loss (Fig 2) (14–18). Code can be found online (https://github.com/Rad-190925/Code).
Figure 2:
Deep learning architecture overview. First (step 1), a RetinaNet was trained to reliably depict right and left hip on a radiograph. For this step, the radiograph was resized to 640 × 640 pixels. Then (step 2), the cropped hip image was resized to 224 × 224 pixels and used as the input for an ImageNet-pretrained DenseNet that served as a shared convolutional features extractor. Fully connected layers were trained for each radiographic osteoarthritis (OA) feature in a multitask deep learning approach (step 3) with hard parameter sharing to assess the severity of femoral osteophytes (FOs), acetabular osteophytes (AOs), joint-space narrowing (JSN), subchondral cysts (SUBCYST), and subchondral sclerosis (SUBSCL) to obtain the final assessment (step 4).
Gradient-weighted class activation maps after the last convolutional layer depicted network decision-making processes (19).
Statistical Analysis
Bounding box placement was evaluated with intersection over union for the validation set and test set and for correct placement on radiographs without bounding boxes by C.E.v.S. (a radiologist in training with 3 years of experience in reading pelvic radiographs). Final model performance was evaluated on the internal test set and external clinical data set. To evaluate the model precision, F1 scores, confusion matrices, and receiver-operating characteristics were calculated by using scikit-learn 0.19 (scikit-learn.org); linearly weighted Cohen κ values with 95% confidence intervals (CIs) were calculated with statsmodels (0.9, statsmodels.org; open source) (20,21). Precision is defined as true-positive results divided by the sum of false-positive and true-positive results. Recall is defined as true-positive results divided by the sum of true-positive and false-negative results. F1 score is defined as the harmonic mean of precision and recall. Receiver-operating characteristics were calculated from the softmax function of absence of a feature. The model was trained, evaluated, and visualized by C.E.v.S., an information technology engineer (7 years of experience in data analysis and visualization).
Results
OAI Data Set and Clinical Data Set
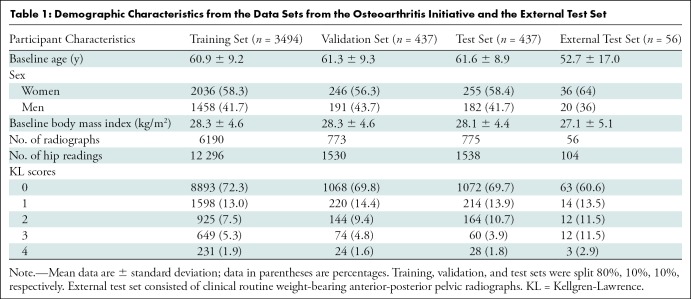
A total of 15 364 hip joints observed on 7738 weight-bearing anterior-posterior pelvic radiographs in 4368 participants in the OAI were included (mean age, 61 years ± 9 [standard deviation]; 1830 men [41.9%] and 2538 women [58.1%]; mean body mass index, 28.3 kg/m2 ± 4.6). In the external test set, participants were mean age 53 years ± 17; 36 participants were women (64.3%). Table 1 provides an overview of participant characteristics. Table 2 summarizes the radiographic osteoarthritis feature grades and their frequencies within the two data sets.
Table 1:
Demographic Characteristics from the Data Sets from the Osteoarthritis Initiative and the External Test Set
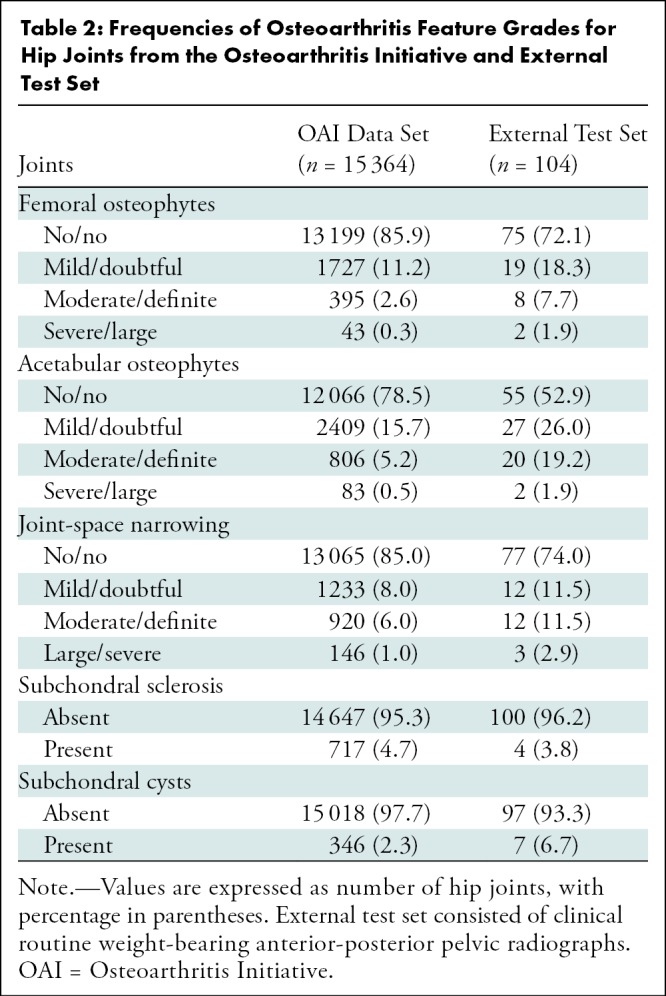
Table 2:
Frequencies of Osteoarthritis Feature Grades for Hip Joints from the Osteoarthritis Initiative and External Test Set
Interreader reliability between the two readers of the training, validation, and test sets from the OAI was assessed for each feature and each point. Baseline visit and follow-up visit radiographs were read together and in known chronologic order. Thirty-five percent (2706 of 7738) of the radiographs were adjudicated by the third reader. For baseline and follow-up, respectively, the recorded linearly weighted Cohen κ values were as follows: 0.77 (95% CI: 0.74, 0.82) and 0.73 (95% CI: 0.69, 0.78) for FOs, 0.71 (95% CI: 0.67, 0.77) and 0.73 (95% CI: 0.69, 0.78) for JSN, 0.56 (95% CI: 0.45, 0.65) and 0.53 for AOs (95% CI: 0.42, 0.63), 0.47 (95% CI: 0.32, 0.61) and 0.48 (95% CI: 0.34, 0.62) for subchondral cyst, and 0.56 (95% CI: 0.47, 0.66) and 0.54 (95% CI: 0.45, 0.64) for subchondral sclerosis. Agreement on presence or absence of each feature was as follows: for FOs, 87.0% (13 372 of 15 364; 78.0% [11 992 of 15 364] for specific grade); for AOs, 77.3% (11 873 of 15 364; 67.4% [10 355 of 15 364] for specific grade); for JSN, 84.8% (13 025 of 15 364; 77.8% [11 955 of 15 364] for specific grade); for subchondral sclerosis, 82.2% (12 634 of 15 364); and for subchondral cyst, 92.8% (14 252 of 15 364).
Deep Learning Model Performance on OAI Test Set
After training on 7002 bounding boxes on 3544 radiographs, the RetinaNet for hip joint localization accurately placed bounding boxes in 100% of validation joints (444 of 444 radiographs and 877 of 877 bounding boxes) and 100% of test joints (451 of 451 radiographs and 897 of 897 bounding boxes), with an excellent intersection over union of 0.91 ± 0.07 and 0.91 ± 0.06 for the validation and test sets, respectively. The RetinaNet model was then applied on an additional 3962 radiographs in 7861 hips; it correctly placed the bounding boxes for all cases.
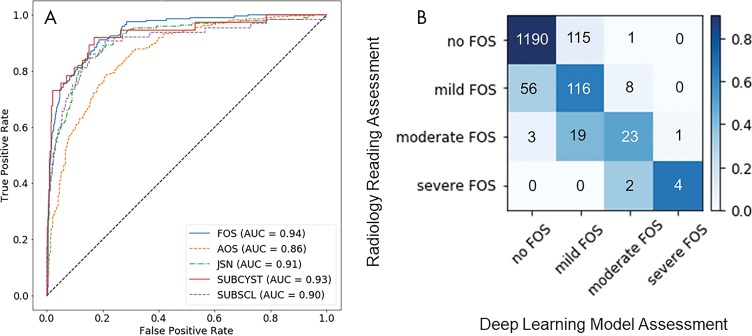
With use of the DenseNet to assess overall presence or absence of each feature, the overall accuracy and area under the receiver operating characteristic curve (AUC) were as follows, respectively: 88.9% (1368 of 1538) and 0.94 (95% CI: 0.92, 0.96) for FOs, 80.1% (1232 of 1538) and 0.86 (95% CI: 0.83, 0.89) for AOs, 86.1% (1324 of 1538) and 0.91 (95% CI: 0.89, 0.94) for JSN, 95.8% (1473 of 1538) and 0.90 (95% CI: 0.85, 0.95) for subchondral sclerosis, and 97.6% (1501 of 1538) and 0.93 (95% CI: 0.88, 0.98) for subchondral cysts. The receiver-operating characteristic curves for all features are shown on Figure 3, A.
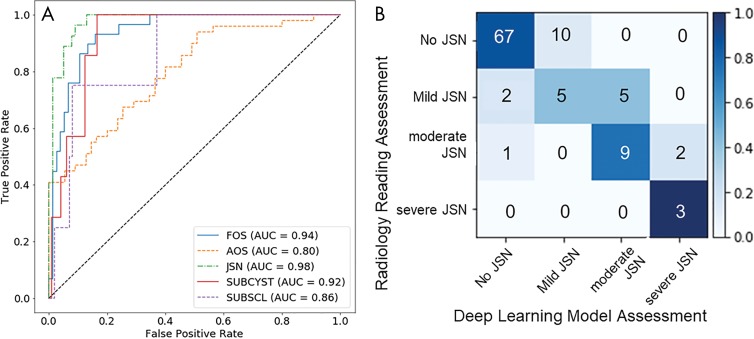
Figure 3:
Performance of the deep learning model on the test set from the Osteoarthritis Initiative. A, Receiver operating characteristic curves for the detection of radiographic osteoarthritis features: femoral osteophytes (FOS), acetabular osteophytes (AOS), joint-space narrowing (JSN), subchondral cysts (SUBCYST), and subchondral sclerosis (SUBSCL). FOS showed the highest area under the receiver operating characteristic curve (0.94). B, Confusion matrix of the deep learning model for grading FOS. The overall accuracy was 86.7% (1333 of 1538), with a linearly weighted Cohen κ value of 0.62 (95% confidence interval: 0.49, 0.76). Most grading discrepancies occurred between two neighboring grades, whereas only 0.3% (four of 1538) of the cases demonstrated discrepancies between nonneighboring grades.
The overall accuracy achieved for grading FOs, AOs, and JSN on a scale with four severity grades was 86.7% (1333 of 1538), 69.9% (1075 of 1538), and 81.7% (1257 of 1538), respectively. For FOs, the precision for each grade was 95.3% (1190 of 1249) for absent, 46.4% (116 of 250) for mild, 68% (23 of 34) for moderate, and 80% (four of five) for severe. For AOs, the precision for each grade was 88.8% (891 of 1003) for absent, 33.0% (117 of 415) for mild, 38.6% (44 of 114) for moderate, and 50% (three of six) for severe. For JSN, the precision for each grade was 92.2% (1168 of 1267) for absent, 28.4% (46 of 162) for mild, 34% (32 of 95) for moderate, and 79% (11 of 14) for severe. Most grading discrepancies occurred between two neighboring grades, as demonstrated on Figure 3, B. Grading discrepancies between nonneighboring grades occurred in 0.2% (four of 1538) of the joints for FOs, 3.0% (47 of 1538) for AOs, and 2.5% (39 of 1538) for JSN.
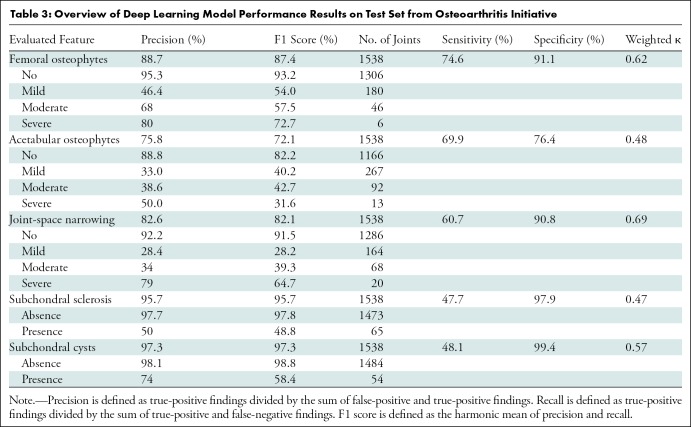
Overall, grading FOs showed the highest reliability, with a linearly weighted κ value of 0.62 (95% CI: 0.49, 0.76). Grading of AOs and JSN showed a reliability of 0.52 (95% CI: 0.34, 0.62) and 0.64 (95% CI: 0.57, 0.80), respectively. Grading subchondral sclerosis had the lowest reliability: 0.47 (95% CI: 0.36, 0.58). Grading subchondral cysts showed a reliability of 0.57 (95% CI: 0.45, 0.70). Overall, the reliability of the deep learning model was moderate to good. An overview of the results with precision, recall, F1 score, κ values, and sensitivity and specificity are shown in Table 3.
Table 3:
Overview of Deep Learning Model Performance Results on Test Set from Osteoarthritis Initiative
Deep Learning Model Performance on External Test Set Consisting of Clinical Routine Radiographs and Comparison with Radiologists
Applying the RetinaNet to detect hip joints for hip joint localization accurately placed bounding boxes in all joints (n = 104). As demonstrated by the receiver-operating characteristics in Figure 4, A, the model had an AUC of 0.8 or greater for all assessing the presence of absence of each evaluated feature.
Figure 4:
Performance of the deep learning model on the external test set consisting of clinical routine radiographs compared with radiologists. A, Receiver-operating characteristic curves for the detection of radiographic osteoarthritis features: femoral osteophytes (FOs), acetabular osteophytes (AOs), joint-space narrowing (JSN), subchondral cysts (SUBCYST), and subchondral sclerosis (SUBSCL). JSN showed the highest area under the receiver operating characteristic curve (0.98). B, Confusion matrix of the deep learning model for grading JSN. The overall accuracy was 80.8% (84 of 104), and the interreader reliability with the ground truth as assessed by two radiology readers was good, with a linearly weighted Cohen κ of 0.74 (95% confidence interval: 0.62, 0.85).
The accuracy and AUC were 82.7% (86 of 104) and 0.94 (95% CI: 0.89, 0.98) for grading FOs, 65.4% (68 of 104) and 0.8 (95% CI: 0.72, 0.88) for AOs, 80.8% (84 of 104) and 0.98 (95% CI: 0.96, 1) for JSN, 88.5% (92 of 104) and 0.86 (95% CI: 0.70, 1) for subchondral sclerosis, and 91.3% (95 of 104) and 0.92 (95% CI: 0.86, 0.98) for subchondral cysts.
The interreader reliability between deep learning model assessment and ground truth as measured by linearly weighted Cohen κ was 0.64 (95% CI: 0.49, 0.79) for FOs, 0.53 (95% CI: 0.39, 0.66) for AOs, 0.74 (95% CI: 0.63, 0.85) for JSN, 0.29 (95% CI: 0.02, 0.57) for subchondral sclerosis, and 0.53 (95% CI: 0.27, 0.79) for subchondral cysts.
To compare, the agreement between reader 1 and reader 2 was 68.3% (71 of 104) for FOs, 55.8% (58 of 104) for AOs, 71.1% (74 of 104) for JSN, 51.9% (54 of 104) for subchondral sclerosis, and 96.2% (100 of 104) for subchondral cysts.
The interreader reliability between the two readers as measured by linearly weighted Cohen κ was 0.58 (95% CI: 0.45, 0.71) for FOs, 0.45 (95% CI: 0.33, 0.58) for AOs, 0.62 (95% CI: 0.49, 0.75) for JSN, 0.07 (95% CI: 0.0, 0.14) for subchondral sclerosis, and 0.76 (95% CI: 0.53, 0.98) for subchondral cysts. Table 4 gives an overview of these results.
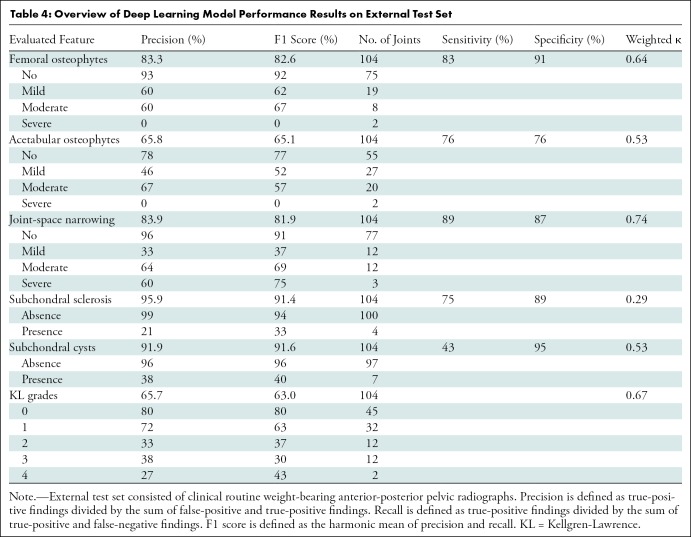
Table 4:
Overview of Deep Learning Model Performance Results on External Test Set
An example of the grading for JSN is given on Figure 4, B.
Model Interpretation and Visualization
Gradient-weighted class activation maps after the last convolutional layer of the model were overlaid with the radiograph to show the relevance of specific areas for the model classification. As demonstrated in Figure 5, the model focused on the region of the osteoarthritis abnormality for its assessment. These findings indicate that the model learned to assess the correct features instead of learning image correlations. Figure 6, A, B, shows examples of grading discrepancies that included radiographs of poor image quality from the OAI data set, possibly causing incorrect classifications. Figure 6, C, D, shows one of the two joints with grading discrepancies between nonneighboring grades of the external test set.
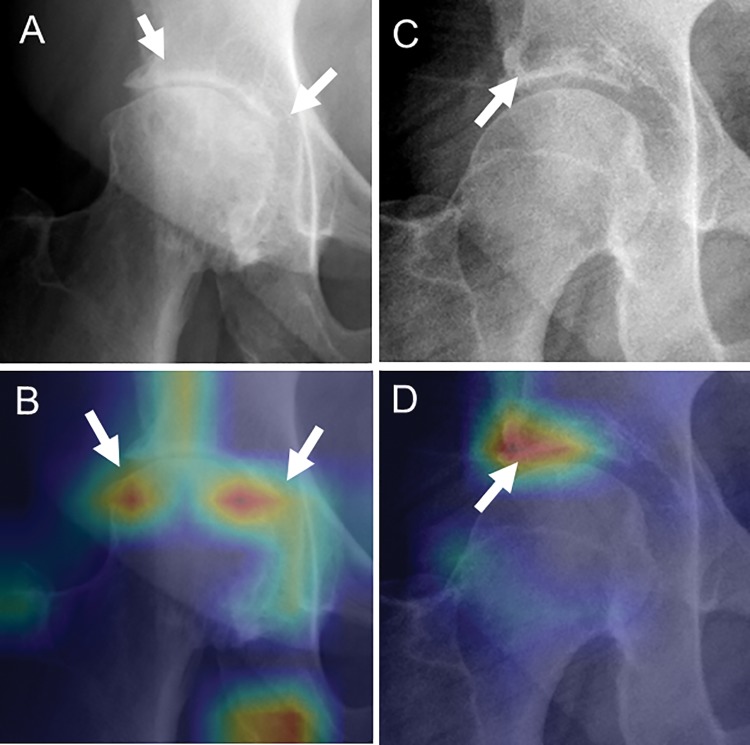
Figure 5:
Heat maps for osteoarthritis feature assessment. A, C, Radiographs and, B, D, gradient-weighted class activation maps with activation after the last convolutional layer of the DenseNet overlaid with the radiograph serving as heat maps (red indicating higher activation, blue indicating lower activation). The heat maps show that the neural network focused on the region of the abnormality for its assessment. A, B, Moderate joint-space narrowing (arrows). The heat map in B shows that the whole joint-space region was considered in determining narrowing of the joint space (arrows). C, D, Acetabular subchondral cyst (arrow). The heat map shows that the neural network focused its attention on this area.
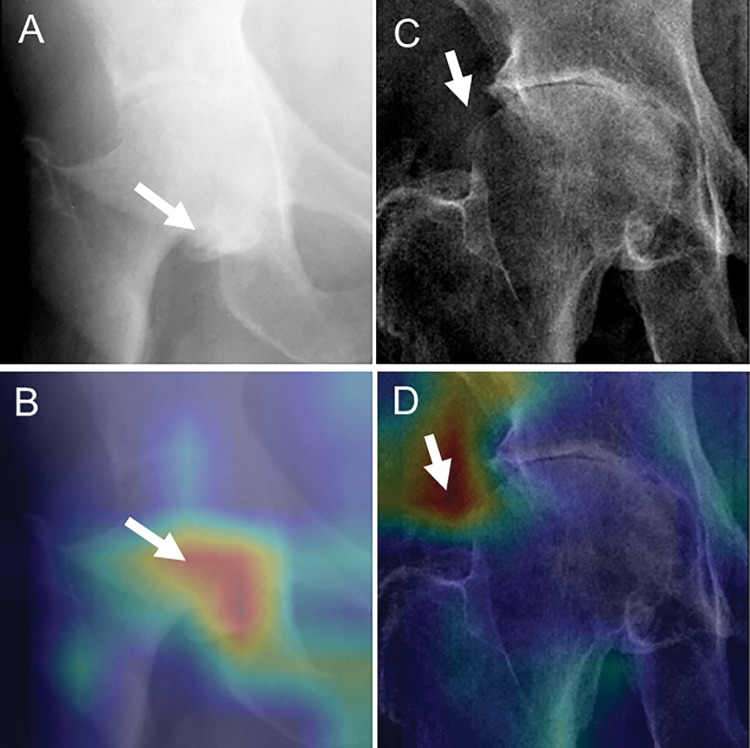
Figure 6:
Examples of grading discrepancies. A, C, Radiographs and, B, D, gradient-weighted class activation maps with activation after the last convolutional layer serving as heat maps (red indicating higher activation, blue indicating lower activation). A, B, Radiographs with poor image quality from the Osteoarthritis Initiative that possibly led to incorrect classification of this radiograph (ground truth: moderate femoral osteophyte [arrow]; model assessment: no osteophyte). C, D, One of two cases of grading discrepancies between nonneighboring grades from the external test set that were graded as mild femoral osteophyte by both readers but as severe femoral osteophyte by the deep learning model. Arrow points to a hyperdense area that led the model to come to this decision as shown by the heat map.
Discussion
A multitask deep learning model that reliably grades severity of hip osteoarthritis features might be desirable, particularly for large epidemiologic studies that require detailed structural assessment by expert radiologists. Therefore, we developed and validated a multitask deep learning model to automatically assess femoral osteophytes (FOs), acetabular osteophytes (AOs), joint-space narrowing (JSN), subchondral sclerosis, and subchondral cyst. The accuracy of the model for assessing these five features varied depending on the evaluated feature and was 86.7% for FOs, 69.9% for AOs, 81.7% for JSN, 95.8% for subchondral sclerosis, 97.6% for subchondral cyst on an internal test set and 82.7% for FOs, 65.4% for AOs, 80.8% for JSN, 88.5% for subchondral sclerosis, and 91.3% for subchondral cyst on the external test set.
Previous studies investigated the potential of deep learning to assess osteoarthritis in radiographs. Xue et al (6) used a VGG-16 model for binary hip osteoarthritis classification of pelvic radiographs without preprocessing into normal or abnormal and achieved high diagnostic accuracy. Two other studies used approaches to assess the severity of knee osteoarthritis with Kellgren-Lawrence scores on radiographs from the Multicenter Osteoarthritis Study and the OAI: Tiulpin et al (9) used a Siamese network architecture to evaluate the medial and lateral sides of the knee for a final Kellgren-Lawrence grade classification achieving moderate grading reliability; and Norman et al (8) used DenseNets and additionally capitalized on demographic information, leading to higher sensitivity and specificity for grading knee osteoarthritis. However, Kellgren-Lawrence scores represent a condensed grading and do not accurately reflect the variety of imaging features that are associated with osteoarthritis (22). Further studies investigated the potential of deep learning models to assess more than one feature. One of these studies evaluated the potential of an AlexNet-based model as a convolutional feature extractor to detect abnormalities, anterior cruciate ligament tears, and meniscal tears at MRI without grading severity (23).
We used a multitask deep learning approach to grade the severity of radiographic hip osteoarthritis features with hard parameter sharing on the basis of DenseNet to achieve grading reliabilities similar to and succeeding those previously reported for Kellgren-Lawrence scores.
Our deep learning model showed varying grading reliabilities depending on the evaluated radiographic feature. We recorded higher reliabilities for grading of FOs and JSN and lower reliabilities for the assessment of AOs, subchondral cyst, and subchondral sclerosis in both our deep learning model and our radiology readings. Similarly, in previous clinical studies, interreader reliability was lower for the radiographic assessment of subchondral sclerosis and AOs and interreader reliabilities were higher for JSN and FOs (24,25). Overall, these findings indicated that some osteoarthritis imaging features are likely more challenging to evaluate for both deep learning models and radiologists. However, the lower performance of the deep learning model could potentially be related to less reliable radiology readings for specific features because these served as ground truth. Artificial intelligence may be particularly useful in the context of large epidemiologic studies requiring detailed structural assessment by expert radiologists’ readings. Further studies are warranted to advance current deep learning models and to investigate the effect of deep learning to aid diagnostics in radiology. However, some studies already show potential benefits of deep learning–aided diagnostics for some applications (23,26).
Our study had limitations. First, ground truth was obtained from radiology readings. Although expert radiology readings are the best standard of reference for many applications, they might contain variability. Second, the deep learning model used only a single frontal-view pelvic radiograph for its assessment, even though in clinical practice additional views, such as dedicated hip radiographs, are acquired for osteoarthritis evaluation. Third, the agreement for mild and moderate JSN was low, even though this is the defining feature to differentiate between Kellgren-Lawrence grades 2 and 3. Fourth, the model did not assess other underlying abnormalities that are relevant to osteoarthritis grading, such as avascular necrosis, protrusion acetabuli, and inflammatory arthritis. Finally, low-quality studies were excluded, which possibly affected the diagnostic performance of the model.
In conclusion, our study demonstrated the feasibility of a multitask deep learning approach to grading hip osteoarthritis features on radiographs and showed that its performance was similar to that of expert radiologists. This model may be useful in large epidemiologic studies for structural assessment of hip osteoarthritis features.
APPENDIX
Study supported by the National Institutes of Health (R01AR064771, R00AR070902, NIBIB 5T32EB001631) and used public-use data sets from the Osteoarthritis Initiative, which was funded by the National Institutes of Health (N01-AR-2-2258, N01-AR-2-2259, N01-AR-2-2260, N01-AR-2-2261, N01-AR-2-2262). Study supported by Merck Research Laboratories, Novartis Pharmaceuticals, GlaxoSmithKline, and Pfizer.
Disclosures of Conflicts of Interest: C.E.v.S. disclosed no relevant relationships. J.H.S. disclosed no relevant relationships. F.L. disclosed no relevant relationships. E.O. disclosed no relevant relationships. P.M.J. disclosed no relevant relationships. L.N. disclosed no relevant relationships. M.P. disclosed no relevant relationships. S.C.F. disclosed no relevant relationships. M.C.N. disclosed no relevant relationships. T.M.L. disclosed no relevant relationships. V.P. disclosed no relevant relationships.
Abbreviations:
- AO
- acetabular osteophyte
- AUC
- area under receiver-operating characteristic curve
- CI
- confidence interval
- FO
- femoral osteophyte
- JSN
- joint-space narrowing
- OAI
- Osteoarthritis Initiative
References
- 1.GBD 2015 Disease and Injury Incidence and Prevalence Collaborators . Global, regional, and national incidence, prevalence, and years lived with disability for 310 diseases and injuries, 1990-2015: a systematic analysis for the Global Burden of Disease Study 2015. Lancet 2016;388(10053):1545–1602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Prieto-Alhambra D, Judge A, Javaid MK, Cooper C, Diez-Perez A, Arden NK. Incidence and risk factors for clinically diagnosed knee, hip and hand osteoarthritis: influences of age, gender and osteoarthritis affecting other joints. Ann Rheum Dis 2014;73(9):1659–1664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kim C, Linsenmeyer KD, Vlad SC, et al. Prevalence of radiographic and symptomatic hip osteoarthritis in an urban United States community: the Framingham osteoarthritis study. Arthritis Rheumatol 2014;66(11):3013–3017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Altman R, Alarcón G, Appelrouth D, et al. The American College of Rheumatology criteria for the classification and reporting of osteoarthritis of the hip. Arthritis Rheum 1991;34(5):505–514. [DOI] [PubMed] [Google Scholar]
- 5.Altman RD, Gold GE. Atlas of individual radiographic features in osteoarthritis, revised. Osteoarthritis Cartilage 2007;15(Suppl A):A1–A56. [DOI] [PubMed] [Google Scholar]
- 6.Xue Y, Zhang R, Deng Y, Chen K, Jiang T. A preliminary examination of the diagnostic value of deep learning in hip osteoarthritis. PLoS One 2017;12(6):e0178992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hayashi D, Roemer FW, Guermazi A. Recent advances in research imaging of osteoarthritis with focus on MRI, ultrasound and hybrid imaging. Clin Exp Rheumatol 2018;36 Suppl 114(5):43–52. [PubMed] [Google Scholar]
- 8.Norman B, Pedoia V, Noworolski A, Link TM, Majumdar S. Applying densely connected convolutional neural networks for staging osteoarthritis severity from plain radiographs. J Digit Imaging 2019;32(3):471–477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Tiulpin A, Thevenot J, Rahtu E, Lehenkari P, Saarakkala S. Automatic knee osteoarthritis diagnosis from plain radiographs: a deep learning-based approach. Sci Rep 2018;8(1):1727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Soffer S, Ben-Cohen A, Shimon O, Amitai MM, Greenspan H, Klang E. Convolutional neural networks for radiologic images: a radiologist’s guide. Radiology 2019;290(3):590–606. [DOI] [PubMed] [Google Scholar]
- 11.Joseph GB, Hilton JF, Jungmann PM, et al. Do persons with asymmetric hip pain or radiographic hip OA have worse pain and structure outcomes in the knee opposite the more affected hip? Data from the Osteoarthritis Initiative. Osteoarthritis Cartilage. 2016 Mar;24(3):427–435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lin T, Goyal P, Girshick R, He K, Dollár P. Focal loss for dense object detection. 2017 IEEE International Conference on Computer Vision (ICCV), Venice, Italy, October 22–29, 2017. Piscataway, NJ: IEEE, 2017; 2999–3007.
- 13.Lin TY, Maire M, Belongie S, et al. Microsoft COCO: Common Objects in Context. Cham, Switzerland: Springer International, 2014; 740–755. [Google Scholar]
- 14.Paszke A, Gross S, Chintala S, et al. Automatic differentiation in PyTorch. NIPS-W2017. https://openreview.net/forum?id=BJJsrmfCZ. Accessed January 9, 2020.
- 15.Huang G, Liu Z, van der Maaten L, Weinberger KQ. Densely connected convolutional networks. 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, July 21–26, 2017. Piscataway, NJ: IEEE, 2017; 2261–2269.
- 16.Verma V, Lamb A, Beckham C, et al. Manifold mixup: learning better representations by interpolating hidden states. https://arxiv.org/abs/1806.05236. Published 2018. Accessed January 9, 2020.
- 17.Subramanian V. Deep Learning with PyTorch: A Practical Approach to Building Neural Network Models Using PyTorch. Birmingham, England: Packt Publishing, 2018. [Google Scholar]
- 18.Kingma DP, Ba J. Adam: A Method for Stochastic Optimization. arXiv e-prints [serial online]. https://arxiv.org/abs/1412.6980. Published 2014. Accessed December 1, 2014.
- 19.Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D. Grad-CAM: visual explanations from deep networks via gradient-based localization. 2017 IEEE International Conference on Computer Vision (ICCV), Venice, Italy, October 22–29, 2017. Piscataway, NJ: IEEE, 2017; 618–626.
- 20.Pedregosa F, Varoquaux G, Gramfort A, et al. Scikit-learn: machine learning in Python. J Mach Learn Res 2011;12:2825–2830. [Google Scholar]
- 21.Powers DMW. Evaluation: from precision, recall and F-measure to ROC, informedness, markedness & correlation. J Mach Learn Technol 2011;2(1):37–63. [Google Scholar]
- 22.Lindsey R, Daluiski A, Chopra S, et al. Deep neural network improves fracture detection by clinicians. Proc Natl Acad Sci USA 2018;115(45):11591–11596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Liu F, Zhou Z, Samsonov A, et al. Deep learning approach for evaluating knee MR images: achieving high diagnostic performance for cartilage lesion detection. Radiology 2018;289(1):160–169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kellgren JH, Lawrence JS. Radiological assessment of osteo-arthrosis. Ann Rheum Dis 1957;16(4):494–502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Günther KP, Sun Y. Reliability of radiographic assessment in hip and knee osteoarthritis. Osteoarthritis Cartilage 1999;7(2):239–246. [DOI] [PubMed] [Google Scholar]
- 26.Bien N, Rajpurkar P, Ball RL, et al. Deep-learning-assisted diagnosis for knee magnetic resonance imaging: Development and retrospective validation of MRNet. PLoS Med 2018;15(11):e1002699. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.