Abstract
The genus Rotavirus comprises eight species designated A to H and one tentative species, Rotavirus I. In a virus metagenomic analysis of Schreiber's bats sampled in Serbia in 2014 we obtained sequences likely representing novel rotavirus species. Whole genome sequencing and phylogenetic analysis classified the representative strain into a tentative tenth rotavirus species, we provisionally called Rotavirus J. The novel virus shared a maximum of 50% amino acid sequence identity within the VP6 gene to currently known members of the genus. This study extends our understanding of the genetic diversity of rotaviruses in bats.
Keywords: Chiroptera, Viral metagenomics, Semiconductor sequencing, Rotavirus, Astrovirus, Coronavirus, Gemycircularvirus, Retrovirus, Miniopterus schreibersii
Highlights
-
•
Viral metagenomic analysis identified numerous eukaryotic viruses in bat guano.
-
•
Whole genome sequencing was performed to characterize a novel rotavirus strain.
-
•
This novel rotavirus strain likely represents a new rotavirus species, provisionally named Rotavirus J.
1. Introduction
Rotaviruses (RVs, family Reoviridae, genus Rotavirus) are a major cause of acute diarrhea in mammals and birds. At present, eight recognized and one proposed rotavirus species (RVA to RVH and RVI, respectively) are distinguished. Among these, RVA to RVC, RVE, RVH and RVI are known to infect mammals and RVA is the most widespread species in most, if not all, mammalian hosts (Estes and Greenberg, 2013, Matthijnssens et al., 2012, Mihalov-Kovács et al., 2015).
Batborne RVs described so far belong almost exclusively to RVA; sequence analysis of the identified strains uncovered some intriguing details concerning the ecology and evolution of batborne RVAs. For example, a bat strain from Kenya had an unusual VP1 gene and the hypothesis arose that during their evolution mammalian RVs belonging to different RV species may share genes by reassortment (Esona et al., 2010). Furthermore, bats seem to serve as reservoirs of multiple RVA genotypes commonly found in heterologous host species. Consequently, batborne RVAs might pose some veterinary and public health risk (Asano et al., 2016, He et al., 2013, Xia et al., 2014). More recent data indicate that in addition to RVA, RVH may also infect bats (Kim et al., 2016).
Among bats, Schreiber's bat (Miniopterus schreibersii) represents one of the most widespread species complex in the world, living in large colonies. Schreiber's bats are distributed in distinct lineages throughout Oceania, Africa, Southern Europe and South-East Asia (Appleton et al., 2004). Colonies of M. schreibersii are usually large and dense so that members of the colony can save energy during the hibernation period. These bats may roost together with Rhinolophus ferrumequinum, Rhinolophus euryale, Myotis myotis, Myotis blythii, and Myotis emarginatus. M. schreibersii is able to fly large distances (> 500 km) from one roost to another (Hutterer et al., 2005). Overall, these colonial and behavioral characteristics of M. schreibersii may notably influence pathogen dissemination that could lead to high prevalence and maintenance of viruses within colonies (Kemenesi et al., 2014).
Our recent pilot study on fecal virome analysis of the Hungarian bat fauna provided new insight into viral diversity, providing evidence of novel astroviruses and bufaviruses in M. schreibersii (Kemenesi et al., 2014, Kemenesi et al., 2015). To further explore the ecological role of these common bats as virus reservoirs we involved additional geographical locations in our surveys. While we were prepared that new virus diversity may be explored by the method of viral metagenomics, we unexpectedly, identified sequence traces of a novel rotavirus in multiple samples. Sequence and phylogenetic analysis of the complete genome sequence of a selected rotavirus strain provided evidence of a candidate new rotavirus species in these bats.
2. Materials and methods
2.1. Bat guano
Bat guano samples were collected on October 3rd 2014 at cave Pionirska pećina (Beljanica Mt., Serbia; 44° 4′ N, 21°38′ E) during regular bat-ringing activities by experienced chiropterologists (under a license provided by the Ministry of Energetics, Development, and Environmental Protection of the Republic of Serbia, license number: 353-01-2660/2013-08). A mist-net (7 × 2.5 m) was set up at the cave entrance before sunset and remained open until 2 a.m. The trapped bat specimens were removed immediately, identified following Dietz et al. (2009) and held individually in perforated disposable paper bags for maximum of 30 min in order to let them defecate. After collecting fecal samples, bats were aged, sexed, measured, banded and released. A total of 128 Miniopterus schreibersii were captured (45 males and 83 females), and fecal samples were collected from ten specimens (3 males and 7 females). Droppings were stored in RNAlater RNA Stabilization Reagent (QIAGEN) and kept on ice until laboratory processing.
2.2. Semiconductor sequencing
Guano samples were homogenized in 500 μL phosphate buffered saline. After 5 min centrifugation in 10,000 × g, 200 μL of the supernatant was used for nucleic acid extraction, performed with GeneJet Viral DNA and RNA Purification Kit (Thermo Scientific Ltd.), following the manufacturers recommendations. Nucleic acid samples were previously denatured at 97 °C for 5 min in the presence of 10 μM random hexamer tailed by a common PCR primer sequence (Djikeng et al., 2008). Reverse transcription was performed with 1 U AMV reverse transcriptase (Promega), 400 μM dNTP mixture, and 1 × AMV RT buffer (composition at 1 × concentration; 50 mM Tris-HCl [pH 8.3], 50 mM KCl, 10 mM MgCl2, 0.5 mM spermidine and 10 mM DTT) at 42 °C for 45 min following a 5 min incubation at room temperature. Then, 5 μL cDNA was added to 45 μL PCR mixture to obtain a final volume of 50 μL and a concentration of 500 μM for the PCR primer (Djikeng et al., 2008), 200 μM for dNTP mixture, 1.5 mM MgCl2, 1 × Taq DNA polymerase buffer, and 0.5 U of Taq DNA polymerase (Thermo Scientific). The reaction conditions consisted of an initial denaturation step at 95 °C for 3 min, followed by 40 cycles of amplification (95 °C for 30 s, 48 °C for 30 s, 72 °C for 2 min) and terminated at 72 °C for 8 min. 0.1 μg of cDNA was subjected to enzymatic fragmentation and adaptor ligation following the manufacturers recommendations (available at www.neb.com; NEBNext® Fast DNA Fragmentation & Library Prep Set for Ion Torrent™ kit, New England Biolabs). The barcoded adaptors were retrieved from the KAPA Adaptor Kits for Ion Torrent Platforms (Kapa Biosystems). The resulting cDNA libraries were measured on a Qubit® 2.0 equipment using the Qubit® dsDNA BR Assay kit (Invitrogen). The emulsion PCR that produced clonally amplified libraries was carried out according to the manufacturer's protocol using the Ion PGM Template kit on an OneTouch v2 instrument (Life Technologies). Enrichment of the templated beads (on an Ion One Touch ES machine, Life Technologies) and further steps of pre-sequencing set-up were performed according to the 200 bp protocol of the manufacturer. The sequencing protocol recommended for Ion Torrent PGM Sequencing Kit on a 316 chip was strictly followed (Life Technologies).
2.3. Determination of the termini of genomic RNA
To obtain the true sequence of the genome segment ends, a short oligonucleotide (PC3-mod), phosphorylated at the 5′ end and blocked at the 3′ end with dideoxy cytosine, was ligated to the 3′ ends of the genomic RNA in the nucleic acid extract (Lambden et al., 1992, Potgieter et al., 2002). In brief, 5 μL total RNA was combined with 25 μL RNA ligation mixture (consisting of 3.5 μL nuclease free water, 2 μL of 20 μM PC3, 12.5 μL of 34% (w/v) polyethylene glycol 8000, 3 μL of 10 mM ATP, 3 μL 10 × T4 RNA Ligase buffer and 10 U T4 RNA Ligase I (New England Biolabs) and then incubated at 17 °C for 16 h. Following the incubation, the RNA was extracted using the QIAquick Gel Extraction Kit (QIAGEN). Binding of RNA to silica-gel column was performed in the presence of 150 μL QG buffer from the extraction kit and 180 μL isopropanol. All subsequent steps were performed according to the manufacturer's instructions.
Five microliter ligated RNA was heat-denatured in the presence of 1 μL of 20 μM primer (PC2-mod, which is complementary to the PC3-mod oligonucleotide ligated to the 3′ end) at 95 °C for 5 min and then placed on ice slurry. The reverse transcription mixture contained 14 μL nuclease free water, 6 μL 5 × First Strand Buffer, 1 μL of 10 μM dNTP mixture, 1 μL 0.1 M DTT, 20 U RiboLock RNase Inhibitor (Thermo Scientific) and 300 U SuperScript III Reverse Transcriptase (Invitrogen). This mixture was added to the denatured ligated RNA and incubated at 25 °C for 5 min and then 50 °C for 60 min. The reaction was stopped at 70 °C for 15 min.
Subsequently, 2 μL cDNA was added to the PCR mixture, which consisted of 17 μL nuclease free water, 1 μL of 10 μM dNTP mixture, 2.5 μL 10 × DreamTaq Green Buffer (including 20 mM MgCl2), and 2 μL of 20 μM primer pair (i.e. 1 μL PC2 and 1 μL gene-specific primer; see Table 1 ) and 2.5 U DreamTaq DNA polymerase (Thermo Scientific). Gene-specific primers were designed on the basis of preliminary sequence data obtained by semiconductor sequencing. The thermal profile consisted of the following steps: 95 °C 3 min, 40 cycles of 95 °C 30 s, 42 °C 30 s, 72 °C 2 min, final elongation at 72 °C for 8 min. The PCR products were visualized on 1% agarose gel electrophoresis and bands of the expected sizes were excised and cleaned up with Geneaid Gel/PCR DNA fragments Extraction Kit (Geneaid).
Table 1.
Primer sequences used in the study.
| Application | Gene | Orientation | Sequence (5′-3′) | Amplicon length (bp) |
|---|---|---|---|---|
| RNA ligation 5′ and 3′RACE | Universal | (Phos)-GGA TCC CGG GAA TTC GG-(ddC)a | – | |
| Universal | CCG AAT TCC CGG GAT CCa | |||
| VP1 | Fw | CTG CTG AAA CAA TCT TTA AGT GCA A | 270 | |
| Fw | GGA TTG ACT GGT TCA GAA CTA AGG TAT TA | 412 | ||
| Rev | TCT CTT CGA TGA TTT GAG ATG GAG | 198 | ||
| Rev | TCG TCG CAT TCA TTG GAT GTT TTA A | 367 | ||
| VP2 | Fw | GAT GGC GCA GAC TTC GGT ATA C | 233 | |
| Fw | CTC GAT GCA CAG AGA TTA CTC GTC | 208 | ||
| Rev | GAT TTA ACT AAC CGA AGC AAT TCC TTG TA | 204 | ||
| Rev | CTG TTT CTG CTT TTG TTG AGT CTC ATT TC | 280 | ||
| VP3 | Fw | ATG TCT CCG TTT AGA TGG ATA CAG C | 229 | |
| Fw | AAG AGA TAA TTT CGC CGG GTA CTC | 302 | ||
| Rev | TCA ATC GTA ACG TAG AAT GTC TGC TGC | 186 | ||
| Rev | CTT TCA TAA GCA TCA TTT CCC TTC GC | 222 | ||
| VP4 | Fw | ACC CTG TTT TCT TTA CAA ATG CGC A | 243 | |
| Fw | AGT CAG ATG GGT AAT GGC CAT GCA C | 270 | ||
| Rev | CTA TTA TCT TAT TCG AGA GAG GCT TTG TA | 330 | ||
| Rev | GGA GAG AAC GTC AGT ACC AAA TAA TTT CC | 376 | ||
| Rev | GGT TTC ACG TCC GAA TAT TCG CCA CCA | 439 | ||
| VP6 | Fw | CAG CTC CGG CGT CGT TTT TAA TG | 244 | |
| Fw | CTC AAA TGC AAC CGA CAG TAT CA | 328 | ||
| Rev | AGT TGT TCC ATT TGT ACG GGA AGC | 193 | ||
| VP7 | Fw | CTG TCA ATT CGA TAC TGC ACT TTG TTT ATA A | 133 | |
| Fw | GTG TGA GAA AAG ATT CAT CAC AGC CA | 247 | ||
| Rev | CCA TAT AAA CAC GAA CAT TTT GAA ATC GC | 260 | ||
| Rev | TTT CAT ATG TAA ATC CCC TGA ACG AA | 196 | ||
| NSP1 | Fw | GGG AAA AGA TAA ACA ACT TGG AGT GA | 110 | |
| Fw | ATC GAA GAA GCA AGC AAA ACA CGA | 151 | ||
| Rev | GGA AAC AAA GCA ACC ATC TTT CTC TC | 161 | ||
| NSP2 | Fw | CTG GGG ATA GAT TTT TAT CAA TGT GCA | 174 | |
| Fw | CAA GGA AAC AGA AAG AGG AAA TTA CCA | 231 | ||
| Rev | CCT AAT TTC AGC TCT ATC AGC CCC TTG C | 237 | ||
| Rev | GTC ATT CTC CTA CTG CAT CCT GGA GTA | 278 | ||
| NSP3 | Fw | CCA GAC GTT AGA TTC ATG GCT CCA | 168 | |
| Fw | CAG TGA TCG CTT CTA TAG TAA TCA TTG AA | 224 | ||
| Rev | CCA ATT ATC GAC ATT TTC CTC AAG TCC | 203 | ||
| Rev | GGT CAT TTC CTT TGG AAT TCT TTT CTT A | 245 | ||
| NSP4 | Fw | TAA AGA GGA CAT CAT GTA ACT CCA GGA | 120 | |
| Fw | GGG TAA ATA AAA TCT ACA CCA TAC AGG AA | 160 | ||
| Rev | AGT CAT ATC TTT AAA GTA TTG CTG CAT CAT A | 178 | ||
| Rev | CAA CAC CAT ATG TGC GAG TAT TCC TTC | 208 | ||
| NSP5 | Fw | GGC CAA AAC ACT GGA ATC AGC AA | 191 | |
| Fw | CGC ACA GCT CCA ACT TCG ATT GGA A | 236 | ||
| Rev | ATG CCG CGT CGT TTT CTG GAA GGA | 162 | ||
| Rev | GGC CAA TGA TTT AAC GTC CTC TTC A | 200 | ||
| Sequence verification | NSP1 | Fw | CCA CAA ACC GGA CCA AAA GAC GTA CTA | 1074 |
| Rev | CGC CTC GTG TTT TGC TTG CTT CTT C | |||
| NSP5 | Fw | CTC TCT CCA AAA TTA ATC CTT CCA GAA AA | 427 | |
| Rev | CAT GGA GGA GCG TTT TCT TCT GTG GTG TA | |||
| Screening PCR | VP6 | Fw | GGA TTC TCA AAT TAT CTC CAA C | 643 (1st round) |
| Rev | GGA AGT TGA ATA AAA CCT GG | |||
| Fw | CGA TTA CAA CAT TGC TTC | 338 (2nd round) | ||
| Rev | GTT CCA TTC TAG CTG TAT CA |
These oligonucleotides (referred in the text as PC3-mod and PC2-mod, respectively) were adapted from Potgieter et al. (2002).
Amplicons were subjected to Sanger sequencing with the PCR primers using the BigDye Terminator v1.1 Cycle Sequencing Kit (Applied Biosystems). Ethanol precipitated products were run on an ABI PRISM 310 Genetic Analyzer.
2.4. Sanger sequencing of the full-length NSP1 and NSP5 genes
The genome segments encoding NSP1 and NSP5 of RV strains belonging to various RV species may be either mono-, bi- or tricistronic. To validate the results obtained by semiconductor sequencing we performed traditional sequencing. In brief, cDNA production, amplification and Sanger sequencing were carried out with sequence specific primers (Table 1) designed based on the Ion Torrent sequence reads. The experimental protocol was essentially the same as described in the previous section describing the method for determination of genome segment termini.
2.5. RVJ-specific screening RT-PCR assay
Stool samples were homogenized in 500 μL PBS. Following a centrifugation step at 10000 × g for 5 min, the viral RNA was extracted from 200 μL of supernatants using GeneJET Viral DNA and RNA Purification Kit (Thermo Scientific) following the manufacturer's recommendations. Genomic RNA was heat-denaturated at 95 °C for 5 min in the presence of 10 μM gene specific primers. Nested RT-PCR amplification was performed with newly designed primers directed to a 338 nt fragment in the RV VP6 protein region (Table 1). To obtain first round PCR product, 5 μL of the heat-denaturated RNA was reverse transcribed and amplified using QIAGEN One-Step RT-PCR Kit (Qiagen) in a 25 μL final reaction volume. The reaction was performed at 50 °C for 30 min, followed by an initial denaturation at 95 °C for 15 min, and then by 40 cycles of amplification (each cycle included a denaturation step at 94 °C for 30 s, an annealing step at 42 °C for 30 s, and extension step at 72 °C for 1 min). Nested PCR was carried out with the GoTaq DNA Polymerase (Promega). In brief, 3 μL of the first round PCR products were amplified with inner primers for 35 cycles under the following conditions: initial denaturation at 95 °C for 2 min, followed by 40 cycles of amplification (denaturation, 94 °C, 30 s; annealing, 50 °C, 45 s; extension, 72 °C, 1 min). Second round PCR products were analyzed by electrophoresis in 2% agarose gel in TBE buffer stained with GelGreen and then sequenced in both directions using the protocol referred in previous sections.
2.6. Sequence and phylogenetic analysis
For viral metagenomics, raw sequence reads were trimmed and quality controlled using CLC Genomics Workbench (version 9.0; http://www.clcbio.com). The minimal read length parameter was set to 35. Trimmed reads were taxonomically binned using Diamond v0.8.3 versus NCBI-NR (Buchfink et al., 2015). After classification, the output files were analyzed and visualized by MEGAN6 Ultimate Edition (Huson et al., 2016).
The CLC Genomics Workbench software package was utilized to assemble the genome sequence. After visual inspection of sequence mappings a single consensus sequence was created for all 11 genome segments. Further sequence editing and evaluation were carried out by the GeneDoc (Nicholas et al., 1997) and BioEdit software (Hall, 1999) and then analyzed by similarity search using BLAST (Altschul et al., 1990). Multiple alignments were prepared using the TranslatorX online program (Abascal et al., 2010) and manually adjusted in the GeneDoc software (Nicholas et al., 1997), whereas phylogenetic analysis by the maximum-likelihood and the neighbor-joining methods were performed by using the MEGA6 software (Tamura et al., 2013).
For the maximum-likelihood tree the best fit nucleotide substitution models for each gene were selected based on the Bayesian information criterion as implemented in the MEGA software (in particular, TN93 + G was used for NSP3; HKY + G was used for NSP1, NSP4 and NSP5; GTR + G was used for NSP2, VP1, VP2, VP4, VP6, and VP7; GTR + G + I was used for VP3). For the neighbor-joining tree we used the p-distance algorithm.
The coding potential was predicted by using the ORF Finder online platform (http://www.ncbi.nlm.nih.gov/gorf/gorf.html).
2.7. GenBank accession numbers
The whole genome sequence of strain RVJ/Bat-wt/SRB/BO4351/Ms/2014/G1P1 has been deposited under the following accession numbers: KX756619-KX756629.
3. Results and discussion
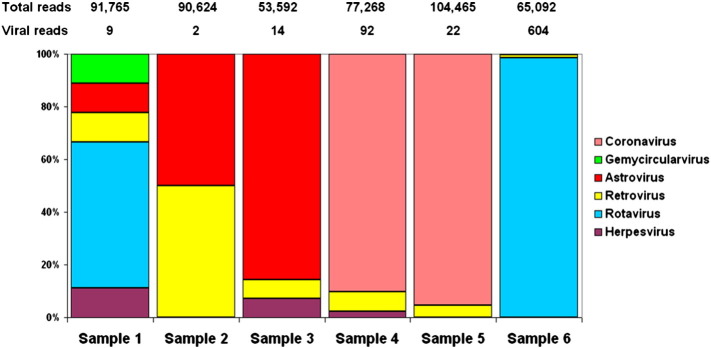
To explore the viral diversity six fecal specimens collected from apparently healthy adult M. schreibersii bats were processed for viral metagenomics. In these samples various amounts of sequence reads mapped onto known eukaryotic viral sequences (range, ≪ 0.1% to 0.9%; Fig. 1 ). When evaluating the results of viral metagenomics data we need to point out that sample processing did not include virion enrichment step and it is not clear whether each of the relevant sequence reads originate from intact virions. Consequently, the presence of potential endogenous viral sequence elements may have affected the overall landscape of viral diversity. For example retrovirus specific reads, which may represent endogenous viral genomic traits from genomic DNA of the host species, were detected in all samples. Overall the rate of eukaryotic virus specific sequence reads was low, likely because we omitted virus particle enrichment procedures in our sample processing protocol. Nonetheless, various eukaryotic viruses were detected in all six selected fecal samples. Herpesvirus, astrovirus and coronavirus sequences were detected in at least three samples (herpesvirus in sample 1, 3, and 4; astrovirus in sample 1 to 3; coronavirus in sample 4 to 6). Rotavirus and gemycircularvirus sequences were found in two and one samples, respectively (both viruses in sample 1; rotavirus without gemycircularvirus in sample 6). In one specimen (i.e. sample 6) RV sequences were the most abundant genomic representatives (98.5%); however, these sequence reads were distributed among various RV species (incl. RVB, RVG, RVH and RVI). To clarify this ambiguous situation, the library DNA that contained the most abundant RV-specific reads was resequenced at a greater sequencing depth. The resulting > 1.3 Million sequence reads were subjected to de novo assembly.
Fig. 1.
Distribution of viral sequence reads in six bat fecal specimens.
As a result, the consensus genome sequence of strain BO4351/Ms/2014 could be assembled from a total of 36,630 sequence reads at 131 X (segment 3) to 457 X (segment 11) average coverage. Once the consensus rotavirus gene sequences were assembled for all 11 genomic segments, the 5′ and 3′ ends of each segment were validated by an independent method. The resulting genome of BO4351/Ms/2014 was 18,135 bp in length (range, 3533 bp for segment 1 and 620 bp for segment 11). Terminal sequences at the 5′ ends showed relatively conserved structure with stable nucleotides at positions 1, 2 and 4 and some variations at positions 3, 5, and 6 (segments 1 and 2, GGCACA; segments 3 and 4, GGCATT; segments 5, 7 and 9, GGAAAT; segments 6 and 10, GGCAAA), while at the 3′ ends the variation was less (TAYACCC) (see details in Table 2 ).
Table 2.
Assignment and some features of the genome segments of the candidate new bat rotavirus, BO4351/Ms/2014.
| Genome segmenta | Assignment based on the main gene product | Positions of start and stop codons |
Sequences at genome segment termini |
||
|---|---|---|---|---|---|
| Start | Stop | 5′ end | 3′ end | ||
| Segment 1 | VP1 | 7 | 3513 | GGCACA | TATACCC |
| Segment 2 | VP2 | 21 | 2981 | GGCACA | TACACCC |
| Segment 3 | VP4 | 10 | 2490 | GGCATT | TATACCC |
| Segment 4 | VP3 | 9 | 2156 | GGCATT | TACACCC |
| Segment 5 | NSP1 | 50 | 1255 | GGAAAT | TACACCC |
| Segment 6 | VP6 ORF-X |
33 118 |
1220 789 |
GGCAAA | TATACCC |
| Segment 7 | NSP3 | 49 | 1044 | GGAAAT | TACACCC |
| Segment 8 | NSP2 | 59 | 958 | GGAAAA | TACACCC |
| Segment 9 | VP7 | 8 | 745 | GGAAAT | TATACCC |
| Segment 10 | NSP4 | 27 | 659 | GGCAAA | TATACCC |
| Segment 11 | NSP5 ORF-Y |
58 212 |
555 433 |
GGAATT | TATACCC |
Order of genome segments was defined on the basis of their size.
Each segment had non-translated regions at both 5′ end (length range, 6 to 57 nt) and 3′ end (length range, 20 to 84 nt). Encoded proteins were assigned based on significant hits through the Blast engine and conserved peptide motifs. With this approach we found the equivalents of the major structural (VP1 to VP4, VP6 and VP7) and non-structural (NSP1 to NSP5) proteins of RVs (Table 2, Table 3 ). The encoded structural and non-structural proteins were assigned to particular RNA segments based on the size of full-length genome segments. Additional putative ORFs were predicted to be encoded on segments coding for VP6 and NSP5; however, these putative proteins shared no conserved protein motifs with those of known from other rotavirus species.
Table 3.
Comparison of the genome size and the coding potential of different RV species.
| Genome segment | Rotavirus A, Wa |
Rotavirus A, 02V0002G3 |
Rotavirus B, Bang373 |
Rotavirus C, Bristol |
Rotavirus D, 05V0049 |
Rotavirus F, 03V0568 |
Rotavirus G, 03V0567 |
Rotavirus H, J19 |
Rotavirus I, KE135/2012 |
Rotavirus J, BO4351a |
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Size (nt) | Protein (aa) | Size (nt) | Protein (aa) | Size (nt) | Protein (aa) | Size (nt) | Protein (aa) | Size (nt) | Protein (aa) | Size (nt) | Protein (aa) | Size (nt) | Protein (aa) | Size (nt) | Protein (aa) | Size (nt) | Protein (aa) | Size (nt) | Protein (aa) | |
| 1 | 3302 | VP1 (1088) | 3305 | VP1 (1089) | 3511 | VP1 (1160) | 3309 | VP1 (1090) | 3274 | VP1 (1079) | 3296 | VP1 (1086) | 3526 | VP1 (1160) | 3538 | VP1 (1167) | 3518 | VP1 (1162) | 3533 | VP1 (1168) |
| 2 | 2717 | VP2 (890) | 2732 | VP2 (895) | 2847 | VP2 (934) | 2736 | VP2 (884) | 2801 | VP2 (913) | 2769 | VP2 (904) | 3014 | VP2 (991) | 2969 | VP2 (973) | 3002 | VP2 (982) | 3010 | VP2 (986) |
| 3 | 2591 | VP3 (835) | 2583 | VP3 (829) | 2341 | VP3 (763) | 2283 | VP3 (693) | 2366 | VP4 (777) | 2246 | VP4 (738) | 2364 | VP4 (772) | 2512 | VP4 (823) | 2371 | VP4 (777) | 2512 | VP4 (826) |
| 4 | 2359 | VP4 (775) | 2354 | VP4 (770) | 2306 | VP4 (750) | 2166 | VP4 (744) | 2104 | VP3 (685) | 2174 | VP3 (694) | 2352 | VP3 (768) | 2204 | VP3 (719) | 2161 | VP3 (701) | 2200 | VP3 (715) |
| 5 | 1567 | NSP1 (486) | 2122 | NSP1 (577) | 1276 | NSP1–1 (107) NSP1–2 (321) NSP1–3 (65) | 1353 | VP6 (395) | 1872 | NSP1 (574) | 1791 | NSP1 (547) | 1295 | NSP1–1 (106) NSP1–2 (324) | 1307 | NSP1 (395) | 1485 | NSP1–1 (79) NSP1–2 (390) | 1322 | NSP1 (401) |
| 6 | 1356 | VP6 (397) | 1348 | VP6 (397) | 1269 | VP6 (391) | 1350 | NSP3 (402) | 1353 | VP6 (398) | 1314 | VP6 (396) | 1267 | VP6 (391) | 1287 | VP6 (396) | 1278 | VP6 (395) | 1277 | VP6 (395) ORF-X (223) |
| 7 | 1074 | NSP3 (310) | 1089 | NSP3 (304) | 1179 | NSP3 (347) | 1270 | VP6 (394) | 1242 | NSP3 (370) | 1309 | NSP3 (370) | 1052 | NSP3 (300) | 1004 | NSP3 (297) | 1018 | NSP2 (301) | 1108 | NSP3 (331) |
| 8 | 1062 | VP7 (326) | 1066 | VP7 (329) | 1007 | NSP2 (301) | 1063 | VP7 (332) | 1026 | NSP2 (310) | 1068 | NSP2 (318) | 1012 | NSP2 (282) | 932 | NSP2 (262) | 954 | NSP3 (273) | 1017 | NSP2 (299) |
| 9 | 1059 | NSP2 (317) | 1042 | NSP2 (315) | 814 | VP7 (249) | 1037 | NSP2 (312) | 1025 | VP7 (316) | 990 | VP7 (295) | 825 | VP7 (247) | 820 | VP7 (258) | 858 | VP7 (268) | 793 | VP7 (245) |
| 10 | 750 | NSP4 (175) | 724 | NSP4 (168) | 751 | NSP4 (219) | 730 | NSP5 (212) | 765 | NSP4 (127) ORF2 (93) | 706 | NSP5 (218) | 801 | NSP4 (187) | 739 | NSP4 (213) | 751 | NSP4 (219) | 743 | NSP4 (210) |
| 11 | 664 | NSP5 (197) NSP6 (92) | 699 | NSP5 (208) | 631 | NSP5 (170) | 615 | NSP4 (150) | 672 | NSP5 (195) | 678 | NSP4 (169) | 678 | NSP5 (181) | 649 | NSP5 (176) | 593 | NSP5 (157) | 620 | NSP5 (165) ORF-Y (73) |
| Total | 18,501 | 19,064 | 17,932 | 17,912 | 18,500 | 18,341 | 18,186 | 17,961 | 17,989 | 18,135 | ||||||||||
Abbreviated name of BO4351/Ms/2014.
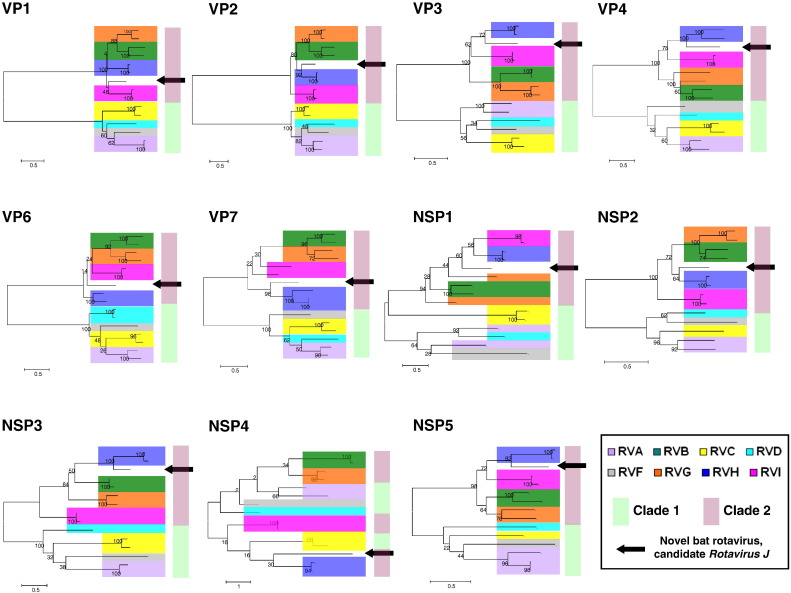
In the phylogenetic analyses cognate sequences of representative RVA to RVH strains were included, except for RVE, for which no sequence information is available. Neighbor-joining and maximum-likelihood trees provided similar topologies, clearly distinguishing clade 1 and clade 2 RV strains. The novel batborne RV consistently clustered with clade 2 RV strains, and in particular, with porcine and human RVH strains. One exception was found when analyzing the NSP4 tree, where the limited bootstrap support at the deepest nodes prevented the separation of the two major RV clades (Fig. 2 ). Consistent with the phylogenetic analyses, the greatest nucleotide and amino acid sequence identities for the novel batborne RV were seen when compared to reference RVH strains (range, 41 (nt%) and 14 (aa%) for NSP4; 63 (nt%) and 64 (aa%) for VP1) (Table 4 ).
Fig. 2.
Phylogenetic trees obtained for the genes encoding all major structural proteins (VP1 to VP4, VP6, and VP7) and non-structural proteins (NSP1 to NSP5) with representative strains of RVA to RVI. Alignments were created using the TranslatorX online platform (http://translatorx.co.uk/). Phylogenetic trees were prepared using the maximum likelihood method as implemented in Mega6 (http://www.megasoftware.net/). Bootstrap values are shown at the branch nodes. Calibration bars are proportional to the genetic distance.
Table 4.
Percentile nucleotide (nt) and amino acid (aa) sequence based identities between the novel batborne RV strain, BO4351/Ms/2014, and reference RVA-RVD and RVF-RVI strains.
| Encoded protein | RVA |
RVB |
RVC |
RVD |
RVF |
RVG |
RVH |
RVI |
||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| nt | aa | nt | aa | nt | aa | nt | aa | nt | aa | nt | aa | nt | aa | nt | aa | |
| VP1 | 40 | 25 | 58 | 57 | 41 | 24 | 41 | 25 | 42 | 24 | 60 | 59 | 63 | 64 | 61 | 59 |
| VP2 | 34 | 14 | 54 | 46 | 35 | 14 | 34 | 12 | 36 | 14 | 56 | 47 | 62 | 61 | 54 | 45 |
| VP3 | 38 | 17 | 46 | 32 | 38 | 19 | 36 | 16 | 38 | 18 | 46 | 31 | 56 | 49 | 51 | 36 |
| VP4 | 33 | 11 | 41 | 25 | 35 | 14 | 35 | 12 | 35 | 12 | 42 | 25 | 48 | 34 | 44 | 26 |
| VP6 | 35 | 15 | 50 | 39 | 34 | 17 | 34 | 13 | 32 | 11 | 49 | 39 | 55 | 49 | 52 | 46 |
| VP7 | 38 | 16 | 42 | 22 | 36 | 16 | 36 | 14 | 37 | 16 | 43 | 25 | 52 | 37 | 49 | 29 |
| NSP1 | 32 | < 10 | 39 | 21 | 30 | < 10 | 31 | < 10 | 31 | < 10 | 39 | 18 | 47 | 34 | 44 | 26 |
| NSP2 | 38 | 16 | 56 | 48 | 35 | 17 | 38 | 17 | 38 | 17 | 55 | 47 | 63 | 59 | 56 | 46 |
| NSP3 | 36 | 15 | 44 | 27 | 34 | 11 | 38 | 18 | 33 | 11 | 40 | 26 | 56 | 49 | 40 | 22 |
| NSP4 | 34 | 10 | 36 | 15 | 32 | 12 | 34 | 12 | 33 | 12 | 36 | 10 | 41 | 14 | 35 | 13 |
| NSP5 | 38 | 13 | 43 | 28 | 38 | 13 | 34 | 11 | 32 | 11 | 47 | 25 | 50 | 39 | 46 | 31 |
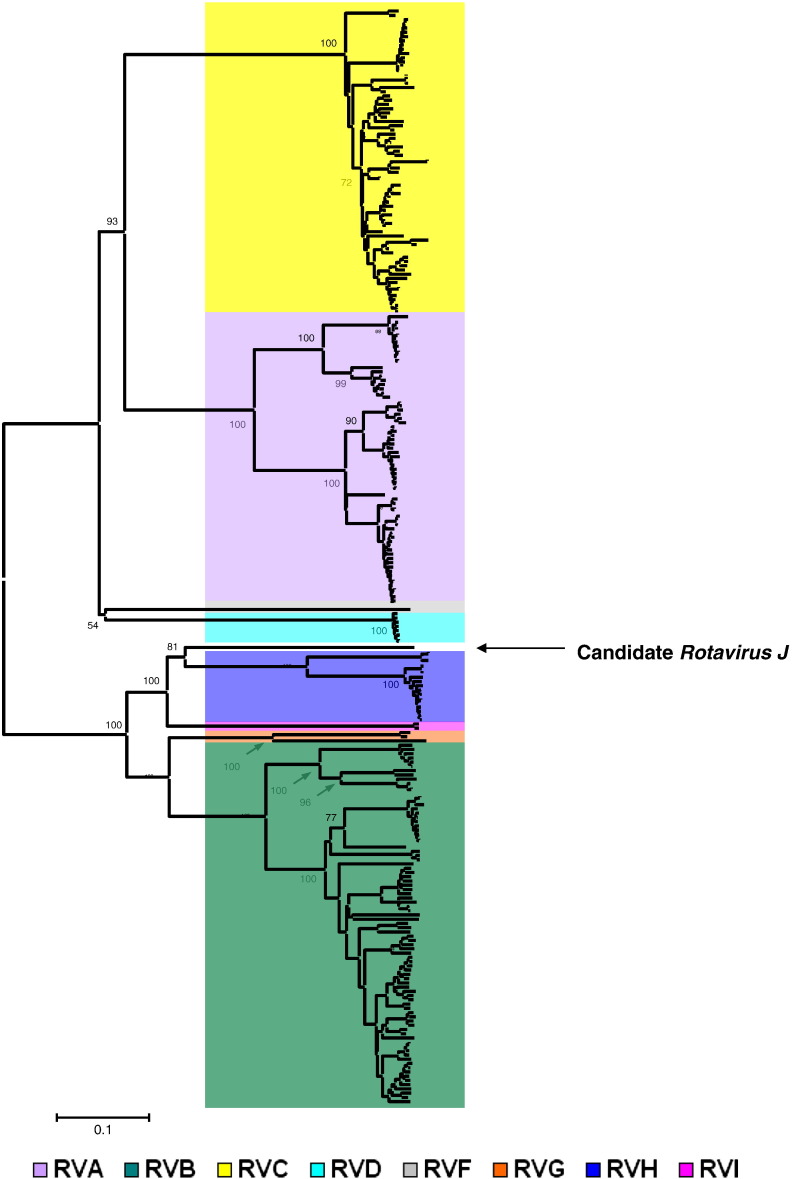
To place the novel batborne strain, BO4351/Ms/2014, into the latest RV taxonomic framework (Matthijnssens et al., 2012; http://www.ictvonline.org), additional VP6 gene sequences were selected from GenBank to represent a broader genetic diversity of various RV species (Fig. 3 ). In this analysis, again, BO4351/Ms/2014 was most closely related to the major genetic lineage containing RVH strains (49–50%, aa) and showed lower similarity to other clade 2 RVs (RVB, 39%, RVG, 39%). The genetic relationship of BO4351/Ms/2014 to clade 1 RVs was marginal (max. identity with RVC, 17%) (Fig. 4 , Table 4). Thus, applying the official species demarcation sequence cut-off value, which is 53% identity at the amino acid level, we conclude that the novel batborne RV strain represents a new RV species, tentatively called Rotavirus J (RVJ). The reference strain was therefore designated as RVJ/Bat-wt/SRB/BO4351/Ms/2014/G1P1.
Fig. 3.
Phylogenetic analysis of the VP6 gene. A total of 258 representative amino acid sequences were selected to provide a more comprehensive phylogenetic analysis for the VP6 gene. Color codes are indicated below the tree. Bootstrap values at the deeper nodes are shown. Calibration bar is proportional to the genetic distance.
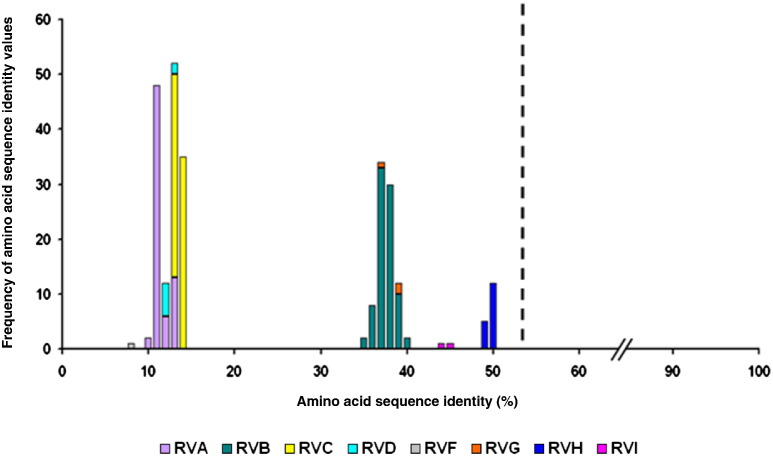
Fig. 4.
Similarity plot prepared from amino acid sequences of the VP6 protein. Dashed line indicates the rotavirus species demarcation sequence identity cut-off value determined by Matthijnssens et al. (2012). Color codes are indicated below the plot.
To determine whether RVJ infection was common among M. schreibersii in the cave under investigation, a nested PCR assay was developed targeting a sequence region that is conserved within the VP6 coding gene of both RVH and RVJ. By adapting the nested PCR assay that amplified a 338 bp long fragment (spanning nucleotide position 137 to 474), another four stool samples were found to be positive for RVJ. All PCR products obtained in the 2nd round PCR were bidirectionally sequenced. The low sequence variation within these short segments (data not shown) suggested the presence of the same virus strain within the colony. Notably, given that RVs have been detected exclusively in birds and mammals, the data presented here suggests that bats may be a true host species of RVJ, although further studies are required to confirm this hypothesis.
It is important to note that by morphological examination all tested animals were confirmed as adult specimens. Immune competence and pathogenicity need to be clarified for most viruses harbored by bats, although asymptomatic virus shedding seems to be common. Further studies are needed to clarify the pathogenicity, prevalence and effect of the virus on bat colonies. Since bats seems to possess special immune characteristics (Zhang et al., 2013), these features may contribute to an altered response to rotavirus infection and explain the high rate of fecal virus shedding in adult M. schreibersii specimens.
Recent years have witnessed considerable sequence data accumulation in public data bases pointing out the enormous genetic diversity within the Rotavirus genus. Viral metagenomics largely contributed to our understanding of this genetic diversity (Asano et al., 2016, He et al., 2013, Kluge et al., 2016, Li et al., 2011, Marton et al., 2015, Mihalov-Kovács et al., 2015, Theuns et al., 2016, Xia et al., 2014). Until the early 2000s RVA to RVG were considered as the only extant RV species (Estes and Greenberg, 2013, Matthijnssens et al., 2012). Sequence independent amplification followed by cloning and sequencing led to the discovery of a novel human RV species that, together with closely related porcine origin strains, was classified into RVH (Matthijnssens et al., 2012, Wakuda et al., 2011, Yang et al., 2004). A newly described member of the Rotavirus genus, RVI, was identified in the fecal viromes of seals and dogs (Li et al., 2011, Mihalov-Kovács et al., 2015).
In this study we described a novel RV detected in M. schreibersii bats from Serbia in 2014. This novel batborne RV belongs to clade 2 RVs, which also includes RVB, RVG, RVH and RVI (Kindler et al., 2013, Mihalov-Kovács et al., 2015). Of interest, the novel strain was closely related to representative strains of RVH suggesting that these RVs had diverged from a common ancestor. Nonetheless, molecular classification indicated that the Serbian batborne RV strain could be the member of a novel RV species that we propose here as Rotavirus J. New sequence information of the complete RVJ genome should enable the design of sophisticated nucleic acid based diagnostic assays and the production of recombinant protein for serological assays that will help describe further details about the ecology, epizootiology and evolution of the novel RV. Of particular interest, given that many batborne viruses are capable of causing severe disease in humans it will be important to study whether or not the novel RVJ strains pose any occupational risk for professional chiropterologists or individuals coming into contact with bats and their excreta.
Acknowledgements
Financial support was obtained from the Momentum (Lendület) Program (awarded by the Hungarian Academy of Sciences), from the Ministry of Education, Science and Technological Development of Serbia (Grant No. 173003) and from TÁMOP (4.2.4.A/2-11-1-2012-0001). S.M. was a recipient of the János Bolyai fellowship (awarded by the Hungarian Academy of Sciences), K.K. was supported by the Szentágothai Talent Program (awarded by the Szentágothai Research Centre, University of Pécs). Research activity of G.K. and F.J. was supported by the ÚNKP-16-3-III and ÚNKP-16-4-III – New Excellence Program of the Ministry of Human Capacities. The present scientific contribution is dedicated to the 650th anniversary of the foundation of the University of Pécs, Hungary.
Contributor Information
Krisztián Bányai, Email: bkrota@hotmail.com.
Ferenc Jakab, Email: jakabf@gamma.ttk.pte.hu.
References
- Abascal F., Zardoya R., Telford M.J. TranslatorX: multiple alignment of nucleotide sequences guided by amino acid translations. Nucleic Acids Res. 2010;38:W7–13. doi: 10.1093/nar/gkq291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Altschul S.F., Gish W., Miller W., Myers E.W., Lipman D.J. Basic local alignment search tool. J. Mol. Biol. 1990;215:403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- Appleton B.R., McKenzie J.A., Christidis L. Molecularsystematics and biogeography of the bent-wing bat complex Miniopterus schreibersii (Kuhl, 1817) (Chiroptera: Vespertilionidae) Mol. Phylogenet. Evol. 2004;31:431–439. doi: 10.1016/j.ympev.2003.08.017. [DOI] [PubMed] [Google Scholar]
- Asano K.M., Gregori F., Hora A.S., Scheffer K.C., Fahl W.O., Iamamoto K., Mori E., Silva F.D., Taniwaki S.A., Brandão P.E. Group A rotavirus in Brazilian bats: description of novel T15 and H15 genotypes. Arch. Virol. 2016;161:3225–3230. doi: 10.1007/s00705-016-3010-9. [DOI] [PubMed] [Google Scholar]
- Buchfink B., Xie C., Huson D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods. 2015;12:59–60. doi: 10.1038/nmeth.3176. [DOI] [PubMed] [Google Scholar]
- Dietz C., von Helversen O., Nill D. A and C Black Publishers Ltd.; London, UK: 2009. Bats of Britain, Europe and Northwest Africa. [Google Scholar]
- Djikeng A., Halpin R., Kuzmickas R., Depasse J., Feldblyum J., Sengamalay N., Afonso C., Zhang X., Anderson N.G., Ghedin E., Spiro D.J. Viral genome sequencing by random priming methods. BMC Genomics. 2008;9:5. doi: 10.1186/1471-2164-9-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Esona M.D., Mijatovic-Rustempasic S., Conrardy C., Tong S., Kuzmin I.V., Agwanda B., Breiman R.F., Banyai K., Niezgoda M., Rupprecht C.E., Gentsch J.R., Bowen M.D. Reassortant group A rotavirus from straw-colored fruit bat (Eidolon helvum) Emerg. Infect. Dis. 2010;16:1844–1852. doi: 10.3201/eid1612.101089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Estes M.K., Greenberg H.B. Rotaviruses. In: Knipe D.M., Howley P.M., editors. Fields virology. sixth ed. Wolters Kluwer - Lippincott Williams & Wilkins; Philadelphia: 2013. pp. 1347–1401. [Google Scholar]
- Hall T.A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp. Ser. 1999;41:95–98. [Google Scholar]
- He B., Yang F., Yang W., Zhang Y., Feng Y., Zhou J., Xie J., Feng Y., Bao X., Guo H., Li Y., Xia L., Li N., Matthijnssens J., Zhang H., Tu C. Characterization of a novel G3P[3] rotavirus isolated from a lesser horseshoe bat: a distant relative of feline/canine rotaviruses. J. Virol. 2013;87:12357–12366. doi: 10.1128/JVI.02013-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huson D.H., Beier S., Flade I., Górska A., El-Hadidi M., Mitra S., Ruscheweyh H.J., Tappu R. MEGAN community edition - interactive exploration and analysis of large-scale microbiome sequencing data. PLoS Comput. Biol. 2016;12 doi: 10.1371/journal.pcbi.1004957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hutterer R., Ivanova T., Meyer-Cords C., Rodrigues L. Bundesamt für Naturschutz; Bonn: 2005. Bat Migrations in Europe—A Review of Banding Data and Literature; p. 180. [Google Scholar]
- Kemenesi G., Dallos B., Görföl T., Boldogh S., Estók P., Kurucz K., Kutas A., Földes F., Oldal M., Németh V., Martella V., Bányai K., Jakab F. Molecular survey of RNA viruses in Hungarian bats: discovering novel astroviruses, coronaviruses, and caliciviruses. Vector Borne Zoonotic Dis. 2014;14:846–855. doi: 10.1089/vbz.2014.1637. [DOI] [PubMed] [Google Scholar]
- Kemenesi G., Dallos B., Görföl T., Estók P., Boldogh S., Kurucz K., Oldal M., Marton S., Bányai K., Jakab F. Genetic diversity and recombination within bufaviruses: detection of a novel strain in Hungarian bats. Infect. Genet. Evol. 2015;33:288–292. doi: 10.1016/j.meegid.2015.05.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim H.K., Yoon S.W., Kim D.J., Koo B.S., Noh J.Y., Kim J.H., Choi Y.G., Na W., Chang K.T., Song D., Jeong D.G. Detection of severe acute respiratory syndrome-like, Middle East respiratory syndrome-like bat coronaviruses and group H rotavirus in faeces of Korean bats. Transbound. Emerg. Dis. 2016;63:365–372. doi: 10.1111/tbed.12515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kindler E., Trojnar E., Heckel G., Otto P.H., Johne R. Analysis of rotavirus species diversity and evolution including the newly determined full-length genome sequences of rotavirus F and G. Infect. Genet. Evol. 2013;14:58–67. doi: 10.1016/j.meegid.2012.11.015. [DOI] [PubMed] [Google Scholar]
- Kluge M., Campos F.S., Tavares M., de Amorim D.B., Valdez F.P., Giongo A., Roehe P.M., Franco A.C. Metagenomic survey of viral diversity obtained from feces of Subantarctic and South American fur seals. PLoS One. 2016;11 doi: 10.1371/journal.pone.0151921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lambden P.R., Cooke S.J., Caul E.O., Clarke I.N. Cloning of noncultivatable human rotavirus by single primer amplification. J. Virol. 1992;66:1817–1822. doi: 10.1128/jvi.66.3.1817-1822.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li L., Shan T., Wang C., Côté C., Kolman J., Onions D., Gulland F.M., Delwart E. The fecal viral flora of California sea lions. J. Virol. 2011;85:9909–9917. doi: 10.1128/JVI.05026-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marton S., Mihalov-Kovács E., Dóró R., Csata T., Fehér E., Oldal M., Jakab F., Matthijnssens J., Martella V., Bányai K. Canine rotavirus C strain detected in Hungary shows marked genotype diversity. J. Gen. Virol. 2015;96:3059–3071. doi: 10.1099/jgv.0.000237. [DOI] [PubMed] [Google Scholar]
- Matthijnssens J., Otto P.H., Ciarlet M., Desselberger U., Van Ranst M., Johne R. VP6-sequence-based cutoff values as a criterion for rotavirus species demarcation. Arch. Virol. 2012;157:1177–1182. doi: 10.1007/s00705-012-1273-3. [DOI] [PubMed] [Google Scholar]
- Mihalov-Kovács E., Gellért Á., Marton S., Farkas S.L., Fehér E., Oldal M., Jakab F., Martella V., Bányai K. Candidate new rotavirus species in sheltered dogs, Hungary. Emerg. Infect. Dis. 2015;21:660–663. doi: 10.3201/eid2104.141370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicholas K.B., Nicholas H.B., Jr., Deerfield D.W.I.I. GeneDoc: analysis and visualization of genetic variation. Embnet News. 1997;4:14. [Google Scholar]
- Potgieter A.C., Steele A.D., van Dijk A.A. Cloning of complete genome sets of six dsRNA viruses using an improved cloning method for large dsRNA genes. J. Gen. Virol. 2002;83(Pt 9):2215–2223. doi: 10.1099/0022-1317-83-9-2215. [DOI] [PubMed] [Google Scholar]
- Tamura K., Stecher G., Peterson D., Filipski A., Kumar S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013;30:2725–2729. doi: 10.1093/molbev/mst197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Theuns S., Conceição-Neto N., Zeller M., Heylen E., Roukaerts I.D., Desmarets L.M., Van Ranst M., Nauwynck H.J., Matthijnssens J. Characterization of a genetically heterogeneous porcine rotavirus C, and other viruses present in the fecal virome of a non-diarrheic Belgian piglet. Infect. Genet. Evol. 2016;43:135–145. doi: 10.1016/j.meegid.2016.05.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wakuda M., Ide T., Sasaki J., Komoto S., Ishii J., Sanekata T., Taniguchi K. Porcine rotavirus closely related to novel group of human rotaviruses. Emerg. Infect. Dis. 2011;17:1491–1493. doi: 10.3201/eid1708.101466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xia L., Fan Q., He B., Xu L., Zhang F., Hu T., Wang Y., Li N., Qiu W., Zheng Y., Matthijnssens J., Tu C. The complete genome sequence of a G3P[10] Chinese bat rotavirus suggests multiple bat rotavirus inter-host species transmission events. Infect. Genet. Evol. 2014;28:1–4. doi: 10.1016/j.meegid.2014.09.005. [DOI] [PubMed] [Google Scholar]
- Yang H., Makeyev E.V., Kang Z., Ji S., Bamford D.H., van Dijk A.A. Cloning and sequence analysis of dsRNA segments 5, 6 and 7 of a novel non-group A, B, C adult rotavirus that caused an outbreak of gastroenteritis in China. Virus Res. 2004;106:15–26. doi: 10.1016/j.virusres.2004.05.011. [DOI] [PubMed] [Google Scholar]
- Zhang G., Cowled C., Shi Z., Huang Z., Bishop-Lilly K.A., Fang X., Wynne J.W., Xiong Z., Baker M.L., Zhao W., Tachedjian M., Zhu Y., Zhou P., Jiang X., Ng J., Yang L., Wu L., Xiao J., Feng Y., Chen Y., Sun X., Zhang Y., Marsh G.A., Crameri G., Broder C.C., Frey K.G., Wang L.F., Wang J. Comparative analysis of bat genomes provides insight into the evolution of flight and immunity. Science. 2013;339:456–460. doi: 10.1126/science.1230835. [DOI] [PMC free article] [PubMed] [Google Scholar]