Abstract
BACKGROUND
Emerging technologies for rapid identification of microbes demonstrate a shift from traditional biochemical and molecular testing algorithms toward methods using mass spectrometry (MS) for the semiquantitative analysis of microbial proteins and genetic elements. This study was performed to assess the diagnostic accuracy of 2 such technologies, PCR–electrospray ionization (ESI)/MS and MALDI-TOF/MS, with respect to phenotypic and biochemical profiling as a reference standard method. A positive challenge set of blood culture bottles was used to compare PCR-ESI/MS and MALDI-TOF/MS performance on a matched set of samples.
METHODS
We performed characterization of bloodstream infections from blood cultures using the Ibis T5000 PCR-ESI/MS and the Bruker MALDI Biotyper 2.0 (MALDI-TOF/MS) platforms for microbial identification. Diagnostic accuracy was determined by independent comparison of each method to phenotypic and biochemical characterization with Vitek2 analysis as the reference standard identification.
RESULTS
The diagnostic accuracy, represented as positive agreement, at the genus level was 0.965 (0.930–0.984) for PCR-ESI/MS and 0.969 (0.935–0.987) for MALDI-TOF/MS, and at the species level was 0.952 (0.912–0.974) with PCR-ESI/MS and 0.943 (0.902–0.968) for MALDI-TOF/MS. No statistically significant difference was found between PCR-ESI/MS and MALDI-TOF/MS in the ability to rapidly identify microorganisms isolated from blood culture.
CONCLUSIONS
Our results demonstrate that PCR-ESI/MS and MALDI-TOF/MS are equivalent in their ability to characterize bloodstream infections with respect to the reference standard, and highlight key differences in the methods that allow for each method to have a unique niche as a tool for rapid identification of microbes in blood cultures.
Identification of microorganisms is of paramount importance to clinical microbiologists for diagnosis and treatment of bloodstream infections (BSIs).6 Effective management of BSIs requires rapid detection and identification of microorganisms to allow deescalation from broad spectrum to targeted antibiotics, thus reducing overuse of broad spectrum antibiotics. Automated blood culture systems continuously monitor microbial growth and are the primary tools for the identification of BSIs. Their use is followed by routine phenotypic and biochemical evaluation of propagated bacteria and yeast to provide microbial identification (1, 2). Reduction of analysis time would be advantageous, especially for organisms that are fastidious, slow-growing, nonculturable, or occur in polymicrobial infections. Several technologies using molecular methods have developed in recent years [i.e., targeted PCR assays (3, 4) and peptide nucleic acid fluorescent in situ hybridization (5, 6)], and are practical for a subset of microorganisms, but broader strategies that can characterize bacteria and yeast without prior knowledge of genetic targets are desirable.
To meet the need for an analytical technique capable of rapidly detecting the diversity of microorganisms found in BSIs, 2 emerging techniques are currently in development: PCR–electrospray ionization (ESI)/mass spectrometry (MS) and MALDI-TOF/MS. Both apply MS to obtain reproducible, species-specific spectra that can be used to identify microorganisms. PCR-ESI/MS measures the mass/charge ratio (m/z) of PCR amplicons generated from several loci on bacterial and fungal genomes, focusing on conserved and species-specific regions, to identify base compositions comparative to a database of microorganisms. MALDI-TOF/MS relies on proteomic profiling of highly conserved proteins generated from direct ionization of a colony of intact organisms or bacterial protein extract, and correlates this spectral signature to a database of spectra collected from reference strains.
PCR-ESI/MS with the Abbott PLEX-ID (previously Ibis T5000) has proven useful for identification of viruses (7–9), bacteria, and associated antibiotic-resistant pathogens (10–13) and, most recently, pathogens directly from blood culture bottles (14). Bruker MALDI Biotyper (MALDI-TOF/MS) has also demonstrated utility for the identification of bacteria and yeast. Early investigations that used visual comparison and identification (15, 16), which progressed to the use of pattern-matching algorithms (17) and extension of the mass range to detect higher molecular weight proteins, led to the development of the Bruker MALDI Biotyper, with proven utility in the identification of bacteria (18–20), specifically bacteria with limited biochemical reactivity that are typically missed by phenotypic evaluation (21, 22). Results of recent evaluations of bacteria from positive blood cultures, although limited by the need for a large number of cells, show promise for the application of this technique to routine clinical testing (23, 24).
The aim of this study was to estimate the diagnostic accuracy of PCR-ESI/MS and MALDI-TOF/MS independently with respect to reference standard testing via Vitek2 analysis. In addition, we compared PCR-ESI/MS and MALDI-TOF/MS methods directly to evaluate the degree of agreement and comparative diagnostic accuracy of each technique to asses statistical differences between the 2 platforms.
Methods
PATIENTS AND SAMPLES
Procedures were performed in accordance with ethical standards as reviewed by the University of Arizona Human Subjects Protection Program. In this retrospective study we used banked remnant samples from patients at the University Medical Center in Tucson, Arizona, whose routine care included blood culture. No patient recruitment was required. The 273 positive blood culture broths tested during the course of this study were analyzed by the reference standard method before analysis by both PCR/ESI-MS and MALDI-TOF/MS.
Reference standard testing was performed by 11 medical technologists who were proficient in Vitek2 methods and whose minimum qualifications were Bachelor of Science degrees and medical laboratory science board certification. Evaluation of PCR-ESI/MS and MALDI-TOF/MS was undertaken by 3 laboratory scientists who had expertise in medical microbiology and biochemistry and whose minimum qualification was a Master of Science degree or equivalent.
SAMPLE SELECTION
For this retrospective study, we created a high-prevalence sample set with a diversity of species that could potentially be encountered in the clinical laboratory. Microbes were identified by standard culture methods and Vitek2 analysis before testing. Compared with sets used in previously reported work (14), the sample set used in this study differed by removal of 27 samples that contained mixtures of organisms with different metabolic requirements, to simplify the Biotyper process and to further reduce redundancy, and by addition of 57 organisms, covering 31 additional species. This strategy broadened our scope so that we were able to further assess the accuracy of each method with organisms encompassing a wider range of genus and species diversity. Samples were blinded before evaluation with PCR/ESI-MS and MALDI-TOF/MS. Blood culture broth was used for PCR/ESI-MS; microbial isolates were used for MALDI-TOF/MS.
BLOOD CULTURE BROTHS
Two milliliters of blood from FA and FN blood culture bottles (Fastidious Antibiotic Neutralization, aerobic and anaerobic, respectively; bioMérieux) determined to be positive by the BacT/Alert 3D instrument (bioMérieux) were removed and prepared for cryostorage in the University of Arizona Infectious Disease Research Core's MicroBank biorepository. These specimens were stored at −80 °C within 30 min of bottle positivity until used. PCR-ESI/MS and MALDI-TOF/MS were performed on aliquots from remnant samples. No treatment was given between the reference standard and index testing.
REFERENCE STANDARD METHODS FOR IDENTIFICATION
Aliquots of positive blood culture bottles were subjected to Gram stain and plating on trypticase soy agar supplemented with 5% sheep blood, chocolate agar, MacConkey agar, or Sabouraud agar (Remel), depending on gram-stain results. Phenotypic testing based on determinative protocols described in the Manual of Clinical Microbiology (25) and in accordance with guidelines issued by the CLSI (26–28) was performed as necessary. Microorganism identifications were determined by the Vitek2 instrument using the GP ID (gram-positive identification), GN ID (gram-negative identification) and Vitek2 YST ID (yeast identification) card (bioMérieux).
DNA ISOLATION FROM POSITIVE BLOOD CULTURES
DNA was extracted from 200 μL of remnant blood culture broth by using the Qiagen Biorobot EZ1. The DNA Bacteria protocol was used to perform all extractions, along with the associated EZ1 DNA Blood or EZ1 DNA Tissue kits (Qiagen).
PCR-ESI/MS ANALYSIS
The Ibis T5000 instrument and software package, version 2.6.052, was used (Ibis/Abbott Molecular for PCR/ESI/MS testing). The Sterile Fluid Bacteria and Candida assay was performed according to the manufacturer's specifications, as previously described (14). Briefly, extracted DNA from blood culture bottles was diluted and distributed into the PCR plate containing primer pairs targeted toward detection and characterization of bloodstream pathogens. The extracted DNA was amplified by PCR on either an MJ Research DNA Engine Tetrad 2 (Bio-Rad Laboratories) or an Eppendorf Mastercycler EP gradient S thermocycler, both of which are approved by Abbott Molecular.
The T5000 was used to obtain the base count of each PCR amplicon, which was compared to a reference database of expected amplicons from each primer pair, which was used to identify bacterial/fungal organisms present. The confidence of identification was determined by the software package. For these experiments, all data meeting a threshold of 87.5% confidence for microbial identification were reported; data with <87.5% confidence were recorded as not interpretable and considered discordant.
MALDI-TOF/MS ANALYSIS
Microorganisms were subcultured before identification with Bruker MALDI Biotyper (MALDI-TOF/MS). Aliquots of 50-μL remnant blood culture specimens were enriched in 1 mL of trypticase soy broth with 0.5% sodium chloride, Lim Broth (Todd-Hewitt broth with colistin nalidixic-acid, Becton Dickinson), or chopped meat broth (Becton Dickinson), depending on metabolic requirements. The organisms were incubated in either ambient air, 5% CO2, or in an anaerobic environment using the GasPak™ EZ Anaerobe Container System with Indicator (Becton Dickinson). These enriched cultures were subcultured on trypticase soy agar, chocolate agar, MacConkey agar, Sabouraud agar, or Columbia colistin nalidixic-acid agar (Becton Dickinson) and incubated in the aforementioned conditions. Isolated organisms were frozen in trypticase soy broth containing 20% glycerol (Becton Dickinson) and submitted for MALDI-TOF/MS analysis in batched groupings to the University of Geneva Hospitals, Switzerland. Before analysis by MALDI-TOF/MS, organisms were recultured from trypticase soy broth containing 20% glycerol and incubated at 37 °C for 24 h.
The Bruker MALDI Biotyper version 2.0 platform with library V.3.1.1.0 with 3740 database entries for MALDI-TOF/MS identification was used. A single colony of fresh overnight culture was smeared directly onto a ground-steel MALDI target plate and was covered with 1.5 μL saturated α-cyano-4-hydroxy-cinnamic acid matrix in 50% acetonitrile–2.5% trifluoroacetic acid, and was allowed to dry. In blood cultures containing multiple organisms, colonies exhibiting variant morphologies were selected and analyzed. Negative blood cultures were not evaluated with the Biotyper, because identification by the Biotyper encompasses the entire sample-processing flow, from positivity through subculture and MALDI-TOF/MS analysis. If no growth was observed after 5 days, the sample was considered negative, and was not analyzed by MALDI-TOF/MS. The plate was inserted into the source of a Bruker MicroFlex MALDI-TOF/MS instrument. Spectra were collected from 2000 to 20 000 Da in linear ion mode, using 240 shots of a 20-Hz nitrogen laser for ionization. All spectra were analyzed using the Bruker Biotyper 2.0 software package, and were compared to reference spectra for identification. A score ≥1.7 was considered a confident identification.
Results
Analysis was performed in accordance with STARD (standards for the reporting of diagnostic accuracy studies) guidelines. A total of 273 blood culture bottles were analyzed using PCR-ESI/MS and MALDI-TOF/MS independently. These results were retrospectively compared to Vitek2 biochemical profiles to identify microorganisms. This study was performed from 07/2009 - 10/2010, and remnant blood cultures were collected from June 29, 2008, to April 27, 2010. During this time, there were approximately 2633 patients whose blood was cultured as part of their routine clinical care. A diverse subset of positive cultures was created which were specifically selected by investigators to represent a challenge set to the 2 technologies that contained a range of biologically diverse bacteria and fungi to assess assay specificity. A list of the organisms sampled can be found in Table 1. A flow chart of the testing process is shown in Fig. 1. No adverse events occurred as a result of this testing.
Table 1. Bacteria and yeast that were contained within the blood culture bottles evaluated in this study, and the corresponding number of times that organism was present, including those occasions in which the organism was present in a mixture.
| Organism | No. of samples | Organism | No. of samples |
|---|---|---|---|
| Abiotrophia sp. | 2 | Escherichia coli | 9 |
| Achromobacter denitrificans | 2 | Eubacterium lentum | 3 |
| Achromobacter xylosoxidans | 2 | Fusobacterium sp. | 2 |
| Acinetobacter baumannii | 5 | Haemophilus influenzae | 6 |
| Acinetobacter lwoffii | 1 | Klebsiella oxytoca | 3 |
| Actinomyces israelii | 1 | Klebsiella pneumoniae | 7 |
| Aeromonas hydrophila | 1 | Lactobacillus sp. | 2 |
| Anaerobic diphtheroides | 1 | Listeria monocytogenes | 1 |
| Anaerobic gram+ cocci | 1 | Micrococcus sp. | 7 |
| Bacillus sp. | 3 | Moraxella catarrhalis | 1 |
| Bacteroides capillosus | 1 | Moraxella osloensis | 1 |
| Bacteroides fragilis | 1 | Morganella morganii | 2 |
| Bacteroides gracilis | 1 | Mycobacterium chelonae | 1 |
| Bacteroides thetaiotaomicron | 3 | Mycobacterium fortuitum | 1 |
| Bacteroides uniformis | 1 | Neisseria elongata | 1 |
| Bacteroides vulgatus | 2 | Neisseria sicca | 1 |
| Candida albicans | 4 | Pantoea sp. | 3 |
| Candida glabrata | 2 | Proteus mirabilis | 1 |
| Candida krusei | 1 | Pseudomonas aeruginosa | 6 |
| Candida lusitanae | 1 | Pseudomonas fluorescens | 1 |
| Candida parapsilosis | 1 | Pseudomonas oryzihabitans | 1 |
| Candida tropicalis | 2 | Pseudomonas stutzeri | 1 |
| Capnocytophaga sp. | 2 | Rhizobium radiobacter | 1 |
| Citrobacter freundii | 1 | Roseomonas sp. | 1 |
| Clostridium difficile | 1 | Salmonella sp. | 4 |
| Clostridium paraputrificum | 1 | Serratia marcesens | 2 |
| Clostridium perfringens | 2 | Staphylococcus sp. (coagulase negative) | 18 |
| Corynebacterium sp. | 3 | Staphylococcus aureus | 38 |
| Delftia sp. | 1 | Staphylococcus epidermidis | 1 |
| Enterobacter aerogenes | 1 | Stenotrophomonas maltophilia | 2 |
| Enterobacter cloacae | 9 | Streptococcus gallolyticus | 1 |
| Enterobacter gergoviae | 1 | Streptococcus gordonii | 1 |
| Enterobacter sakazakii | 1 | Streptococcus mitis | 1 |
| Enterococcus casseliflavus | 1 | Streptococcus oralis | 1 |
| Enterococcus faecalis | 13 | Streptococcus sp. (viridans group) | 10 |
| Enterococcus faecium | 15 | Streptococcus sp. (β-hemolytic) | 10 |
| Enterococcus gallinarum | 1 |
Fig. 1. Flow chart for this study.

All selected blood culture bottles were evaluated by each index test and the reference standard, with no exclusions.
Each patient demonstrated characteristics of BSI, including fever (body temperature >38 °C), chills, and hypotension, or had a blood culture ordered at the discretion of the physician. Of those selected patient samples, 36.4% were from inpatient wards, 20.8% from the emergency department, 12.6% from the adult intensive care unit, 10.8% from oncology, 6.9% from the pediatric inpatient ward, 3.9% from the pediatric intensive care unit, 7.4% from adult outpatients, and 1.3% from autopsy. The age range of participating patients was 14 days to 93 years, with a mean age of 46.3 years. The sample set consisted of 38.4% female and 61.6% male patients.
We compared PCR-ESI/MS and MALDI-TOF/MS results directly to reference standard identification to determine the diagnostic accuracy of each method. In doing so, we assessed 3 levels of specificity: gram-stain result, genus identification, and species identification.
PCR-ESI/MS analysis of DNA extracted from 273 blood culture bottles resulted in 100% concordance with the gram-stain classification made by the reference standard. Genus and species concordance are shown in Fig. 2, A and B. Identification was considered concordant if the reference standard identification agreed with the PCR-ESI/MS identification and also when PCR-ESI/MS correctly identified 1 component of the sample in addition to a second identification (false positive with primary ID) or in the case of a missed second organism (false negative with primary ID). These identifications were considered concordant because 1 of the 2 organisms was correctly identified, and the disagreement on the second organism can in most situations be explained, as discussed below. All results that were indeterminate were considered discordant. The combined concordance on the genus level for PCR-ESI/MS was 96.7%, and on the species level was 95.6% (n = 273).
Fig. 2. Concordance seen with respect to the reference standard.
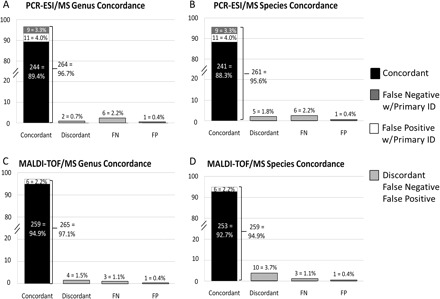
(A), PCR-ESI/MS identification on the genus level. FN, false negative; FP, false positive. (B), PCR-ESI/MS identification on the species level. (C), MALDI-TOF/MS identification on the genus level. (D), MALDI-TOF/MS identification on the species level.
Diagnostic accuracy was measured as positive and negative agreement values with corresponding 95% CIs calculated by using the score method incorporating continuity correction, which is more accurate at high values (29). Positive agreement was defined as the true-positive rate, calculated as the ratio of “true positives” to the total number of positives (true-positive rate = true positives/positives), and negative agreement was defined as the false-positive rate, calculated as the ratio of false positives to the total number of negatives [false-positive rate = 1 − (false positives/negatives)]. Positive and negative agreement values are shown in Table 2.
Table 2. Positive and negative agreement values for PCR-ESI/MS and MALDI-TOF/MS on the genus and species level, considering 2 different methods for determining accuracy.a.
| Method 1a |
Method 2b |
|||
|---|---|---|---|---|
| Value | CI | Value | CI | |
| PCR-ESI/MS | ||||
| Genus | ||||
| Positive agreement | 0.965 | (0.930–0.984) | 0.922 | (0.875–0.952) |
| Negative agreement | 0.978 | (0.868–0.999) | 0.786 | (0.652–0.880) |
| Species | ||||
| Positive agreement | 0.952 | (0.912–0.974) | 0.908 | (0.859–0.941) |
| Negative agreement | 0.978 | (0.868–0.999) | 0.786 | (0.652–0.980) |
| MALDI-TOF/MS | ||||
| Genus | ||||
| Positive agreement | 0.969 | (0.935–0.987) | 0.969 | (0.933–0.986) |
| Negative agreement | 0.978 | (0.868–0.999) | 0.863 | (0.731–0.939) |
| Species | ||||
| Positive agreement | 0.943 | (0.902–0.968) | 0.941 | (0.900–0.967) |
| Negative agreement | 0.978 | (0.868–0.999) | 0.863 | (0.731–0.939) |
Method 1 considers “false positives with primary ID” and “false negatives with primary ID” as concordant, because at least 1 organism was correctly identified.
Method 2 considers “false positives with primary ID” as false positives and “false negatives with primary ID” as false negatives.
MALDI-TOF/MS analysis of subcultured microorganisms from 273 blood culture bottles resulted in 100% concordance with the gram-stain classification made with the reference standard. Genus and species identification concordance can be seen in Fig. 2, C and D. A combined concordance of 97.1% was observed on the genus level, and 94.9% on the species level (n = 273). The diagnostic accuracy was measured as positive and negative agreement values with corresponding 95% CIs, and is shown in Table 2.
No subset analysis was performed with respect to the sample population; test reproducibility was not measured in these experiments owing to feasibility in terms of time and cost of repeating the procedures for such a large sample set. Reproducibility has been previously assessed for MALDI-TOF/MS (22, 30), and for PCR-ESI/MS is measured by correct identification of an internal positive control included in every PCR.
Discussion
Both PCR-ESI/MS and MALDI-TOF/MS methods showed high diagnostic accuracy, as demonstrated by the positive and negative agreement. A more traditional sensitivity and specificity could not be calculated for this study because the positive set of blood culture bottles was investigator-selected to challenge the 2 techniques. Although these results show clinically applicable positive and negative agreement, it is important to note that agreement rates would likely be higher if only commonly encountered organisms were considered. A closer look at discordant and false-positive results provides more information on the accuracy and common errors associated with the 2 techniques. All discordant, false-positive, and false-negative results are illustrated in Table 3. There was 1 false positive with sample 154, which was discordant utilizing both techniques. PCR-ESI/MS identification of Ralstonia picketti is a known blood culture bottle contaminant, resulting in residual DNA following bottle sterilization before use (31), and MALDI-TOF/MS identification of Staphylococcus warneri is a common contaminant that is seen in several instances with MALDI-TOF/MS, as discussed below.
Table 3. List of samples that showed discordance by PCR-ESI/MS and MALDI-TOF/MS compared to the reference standard.
| PCR-ESI/MS and MALDI-TOF/MS combined discordance | |||
|---|---|---|---|
| Sample ID | Discordance | True identificationa | Discordant identification |
| 154 | Both FPb | No growth | PCR-ESI/MS: Ralstonia picketti |
| MALDI-TOF/MS: Staphylococcus warneri | |||
| MALDI-TOF/MS discordance | |||
|---|---|---|---|
| Sample ID | Discordance | True identification | Discordant identification |
| 125 | FN | Anaerobic diphtheroides | Unable to identify |
| 2027 | FN | Mycobacterium chelonae | Unable to identify |
| 2634 | FN | Fusobacterium sp. | Unable to identify |
| 145 | FP w/primary ID | Clostridium perfringens | Bacteroides vulgatus |
| 171 | FP w/primary ID | Clostridium perfringens | Enterococcus faecalis |
| 177 | FP w/primary ID | Pseudomonas aeruginosa | Staphylococcus epidermidis |
| 221 | FP w/primary ID | Streptococcus sp. (Viridans) | Staphylococcus hominis |
| 226 | FP w/primary ID | Pseudomonas aeruginosa | Staphylococcus aureus |
| 252 | FP w/primary ID | Candida albicans | Staphylococcus aureus |
| 329 | FP w/primary ID | Enterococcus faecalis | Staphylococcus saprophyticus |
| 259 | Discordant genus and species | Streptococcus sp. (β-hemolytic) | Staphylococcus aureus |
| 264 | Discordant genus and species | Actinomyces israelii | Clostridium butyricum |
| 429 | Discordant genus and species | Corynebacterium sp. | Actinomyces viscosus |
| 1609 | Discordant genus and species | Abiotrophia sp. | Gamella haemolysans |
| 249 | Discordant species | Bacteroides thetaiotaomicron | Bacteroides fragilis |
| 2566 | Discordant species | Achromobacter xylosoxidans | Achromobacter ruhlandii |
| 2889 | Discordant species | Aeromonas hydrophilia | Aeromonas caviae |
| 2573 | Discordant species | Achromobacter xylosoxidans | Achromobacter ruhlandii |
| 296 | Discordant species | Streptococcus sp. (Viridans) | Streptococcus australis |
| 440 | Discordant species | Streptococcus oralis/S. mitis | Streptococcus pneumoniae |
| PCR/ESI-MS discordance | |||
|---|---|---|---|
| Sample ID | Discordance | True identification | Discordant identification |
| 231 | FN | Candida albicans | Unable to identify |
| 1362 | FN | Candida lusitanae | Unable to identify |
| 2096 | FN | Bacteroides uniformis | Unable to identify |
| 2964 | FN | Eubacterium lentum | Unable to identify |
| 3138 | FN | Eubacterium lentum | Unable to identify |
| 3666 | FN | Eubacterium lentum | Unable to identify |
| 89 | FN w/primary ID | Streptococcus agalactiae | (Staphylococcus epidermidis) |
| 124 | FN w/primary ID | Streptococcus sp. (Viridans) | (Staphylococcus aureus) |
| 135 | FN w/primary ID | Streptococcus sp. (Viridans) | (Staphylococcus epidermidis) |
| 200 | FN w/primary ID | Streptococcus sp. (β-hemolytic) | (Staphylococcus sp. (Coagulase negative) and Enterococcus sp.) |
| 218 | FN w/primary ID | Streptococcus sp. (Viridans) | (Streptococcus pneumoniae) |
| 228 | FN w/primary ID | Staphylococcos epidermidis | (Enterococcus faecalis) |
| 271 | FN w/primary ID | Streptococcus agalactiae | (Staphylococcus sp., coagulase negative) |
| 223 | FN w/primary ID | Klebsiella pneumoniae | (Stenotrophomonas maltophilia) |
| 242 | FN w/primary ID | Staphylococcus epidermidis | (Streptococcus pyogenes) |
| 1272 | FN w/primary ID | Streptococcus sp. (Viridans) | (Neisseria sicca) |
| 76 | FP w/primary ID | Pantoea agglomerans | Erwinia tasmaniensis |
| 302 | FP w/primary ID | Haemophilus influenzae | Mannheimia haemolytica |
| 1647 | FP w/primary ID | Haemophilus influenzae | Mannheimia haemolytica |
| 276 | FP w/primary ID | Micrococcus luteus | Acidothermus cellulolyticus |
| 309 | FP w/primary ID | Staphylococcus aureus | Caudatispora biapiculata |
| 323 | FP w/primary ID | Lactobacillus casei | Bifidobacterium animalis |
| 341 | FP w/primary ID | Citrobacter freundii | Clostridium ramnosum |
| 482 | FP w/primary ID | Moraxella atlantae | Methylococcus capsulatus |
| 1499 | FP w/primary ID | Morganelli morganii | Aeromonas hydrophilia |
| 2889 | FP w/primary ID | Aeromonas hydrophilia | Pseudomonas entomophilia |
| 1033 | Discordant genus and species | Pantoea sp. | Citrobacter braakii |
| 1050 | Discordant genus and species | Pantoea sp. | Citrobacter braakii |
| 30 | Discordant species | Streptococcus gallolyticus | Streptococcus pneumoniae |
| 202 | Discordant species | Staphylococcus sp. (Coagulase negative) | Staphylococcus aureus |
| 2997 | Discordant species | Moraxella catarrhalis | Moraxella nonliquifaciens |
True identification indicates that the organism was identified by the reference standard. Discordant identification indicates that the organism was identified by the index test. Organism name in parentheses indicate the organism that was missed in a mixture.
FP, false positive; FN, false negative.
The MALDI-TOF/MS discordance list shows false negatives in samples 125, 2027, and 2634 that may be difficult to identify owing to low bacterial biomass in culture, as well as being more resistant to protein extraction and ionization due to cell wall complexity. The false positives seen when the primary identification is correct (samples 145, 171, 177, 221, 226, and 329) were predominantly instances in which a Staphylococcus sp. was identified as an additional organism. During the course of these experiments, organisms isolated from representative blood culture bottles underwent multiple passages, offering several opportunities for inadvertent environmental contamination. We believe these false positives were caused by this type of error, and would not be detected if MALDI-TOF/MS was performed immediately after positivity of the blood culture, or in the first pass after removal from cryostorage, as they were not initially seen in subculture before Vitek2 analysis. Discordance observed with MALDI-TOF/MS was due to a limited database, as well as limitations in protein extraction methods for resistant organisms, (e.g., Actinobacteria, Mycobacteria, and those possessing a strong capsule or slime layer).
The PCR-ESI/MS discordance list shows false negatives in 6 samples (231, 1362, 2096, 2964, 3138, and 3666). Three of these 6 false-negative samples were due to the presence of Eubacterium lentum as identified by the reference standard method, an identification that was missed each time this organism was presented. E. lentum is environmental in origin and is not commonly isolated from blood culture, thus this result demonstrates another opportunity for database adjustment to ensure identification of this organism in future samples. PCR-ESI/MS demonstrated several instances in which false negatives were seen in a mixture, where 1 organism was correctly identified, as in samples 89, 124, 135, 200, 228, and 271. As discussed in a previous report (14), an error can occur when combinations of staphylococci and streptococci, or staphylococci and enterococci are present. In these situations, 1 organism is preferentially identified. Other missed identifications were possibly due to large differences in bacterial load, such that only 1 of the 2 organisms is detected in the PCR. Additional false negatives with primary ID in samples 223, 242, and 1272 were likely due to large differences in organism burden, resulting in 1 organism being missed in the identification. False-positive results were observed when a second organism was correctly identified (false positive with primary ID) can be split into 2 groups. The first group contains samples 76, 302, and 1647, in which the second identification is due to an incompletely populated database for Pantoea sp. and Hemophilus influenzae, resulting in a mixed identification. In addition, discordance on both the genus and species levels was seen for Pantoea sp. in samples 1033 and 1050, in which the PCR-ESI/MS identification was Citrobacter braakii. This identification is very genetically similar, because both organisms belong to the Enterobacteriaceae family, which could be resolved in the future by populating the database with isolates from this family. The remaining group, including samples 276, 309, 323, 341, 482, 1499, and 2889, all contained an organism that was identified by PCR-ESI/MS, but at a significantly lower confidence level than the primary identification. Because PCR-ESI/MS is a DNA-based technique, it is possible to detect organisms that are nonviable by culture or represent DNA contamination. These identifications may be true positives based on DNA present, or may be true false positives in mixture. Discordance was also observed in samples 30, 202, and 2997, and the organisms in each sample were misidentified as genetically similar, possibly because of low copy numbers that resulted in incomplete profiles for identification.
COMPARISON OF PCR-ESI/MS TO MALDI-TOF/MS
The clearest comparison of PCR-ESI/MS and MALDI-TOF/MS is demonstrated when positive and negative agreement are compared for each technique. These values were calculated by using samples in which false negatives were detected in the case of positive agreement, and samples in which false positives were detected in the case of negative agreement. The performance of each technique was compared by using the McNemar test, and results indicated that PCR-ESI/MS and MALDI-TOF/MS were not statistically different on either the genus or species level (P > 0.05).
Although both techniques show similar performance, there are several findings of merit that must be considered in the comparison. Although the resulting identification of a microorganism is the same regardless of the technique used, the information required to come to these data is very different for each technique. PCR-ESI/MS uses genetic information whereas MALDI-TOF/MS uses proteomic information to identify organisms present; this difference highlights the varying utilities of each instrument. Genetic identification allows for discrimination down to the representative strain type of each organism during routine work, detects silent mutations, and allows access to antibiotic resistance genes, providing a mechanism for rapid antibiotic susceptibility testing. Proteomic information however, in many instances discriminates down to species-level functioning to provide organism identification. Although several studies have been performed using MALDI-TOF/MS to discriminate clonal/strain types (32–34), this procedure is not performed routinely and would not be applied if used for high-throughput clinical identification. PCR-ESI/MS and MALDI-TOF/MS show similar performance for routine use, but PCR-ESI/MS offers extended utility for epidemiology and infection control. Furthermore, utilizing genetic information offers the potential for identifying uncultivatable organisms.
An additional aspect that should be considered regarding these methods is the time to result. PCR-ESI/MS requires approximately 4–6 h from the time of blood culture positivity, and can resolve mixtures in 1 assay, but this method requires the batching of 6 samples at a time, and may allow direct analysis on whole blood, removing the requirement for blood culture. Conversely, MALDI-TOF/MS currently requires subculture before identification. The time required for this step varies from an additional 12 h from positivity up to and beyond 72 h (35), depending on the organism. MALDI-TOF/MS does not require batching, although owing to the subculture requirement, a batched format may be more convenient. Several recent reports have described the use of MALDI-TOF/MS for identification of microorganisms without subculture (23, 24, 36, 37), and the recently released Sepsityper by Bruker Daltonics (www.bdal.com) has demonstrated the potential to reduce the time to identification. In these studies results have shown successful identification for approximately 80% of blood cultures and therefore demonstrate that more developmental work is required to increase this identification rate before clinical implementation. Although PCR-ESI/MS is currently the faster technique for microorganism identification, recently described MALDI-TOF/MS methods may soon reduce the time to identification to less than an hour from blood culture positivity. The identification times observed for these 2 MS techniques offer a stark comparison to current reports of requirements of approximately 40 h for identification with gold standard methods (38).
The costs of these 2 techniques also differ substantially. Both require the purchase of a dedicated mass spectrometer and software package at a cost that can be considered equivalent for the purposes of this discussion, despite the expected differences in the overall costs of each method. However, the day-to-day cost for consumables greatly varies. PCR-ESI/MS requires DNA extraction and a kit that contains reagents necessary for a PCR, including buffers, enzyme, and primers, resulting in a cost of $50 to $100 per sample, which is comparable to the cost of existing molecular assays. Alternatively, MALDI-TOF/MS requires only media to culture the organism, <1 μg matrix, and tips, the cost of which is approximately $3 to $7 per sample.
In conclusion, the results of this study, overall, demonstrate that both PCR-ESI/MS and MALDI-TOF/MS show high diagnostic accuracy compared to biochemical and phenotypic identification as the reference standard used in the clinical laboratory. Both methods show promise for routine and, in some cases, epidemiological use in hospital settings. These 2 techniques show no statistically significant differences in performance. The results we report highlight key differences between the 2 techniques, both of which will fill unique niches in the clinical microbiology laboratory.
Acknowledgments:
Our sincere thanks go to members of the University of Arizona Infectious Disease Research Core whose efforts allow banking of specimens.
6 Nonstandard abbreviations:
- BSI
bloodstream infection
- ESI
electrospray ionization
- MS
mass spectrometry.
Footnotes
Author Contributions: All authors confirmed they have contributed to the intellectual content of this paper and have met the following 3 requirements: (a) significant contributions to the conception and design, acquisition of data, or analysis and interpretation of data; (b) drafting or revising the article for intellectual content; and (c) final approval of the published article.
Authors' Disclosures or Potential Conflicts of Interest: Upon manuscript submission, all authors completed the Disclosures of Potential Conflict of Interest form. Potential conflicts of interest:
Employment or Leadership: V.H. Wysocki, University of Arizona.
Consultant or Advisory Role: None declared.
Stock Ownership: None declared.
Honoraria: J. Schrenzel, Bruker Daltonics, bioMérieux, and Abbott.
Research Funding: D.M. Wolk, Abbott Molecular.
Expert Testimony: None declared.
Role of Sponsor: The funding organizations played no role in the design of study, choice of enrolled patients, review and interpretation of data, or preparation or approval of manuscript.
References
- 1. Chen JR, Lee SY, Yang BH, Lu JJ. Rapid identification and susceptibility testing using the VITEK 2 system using culture fluids from positive BacT/ALERT blood cultures. J Microbiol Immunol Infect 2008;41:259–64. [PubMed] [Google Scholar]
- 2. Ling TK, Liu ZK, Cheng AF. Evaluation of the VITEK 2 system for rapid direct identification and susceptibility testing of gram-negative bacilli from positive blood cultures. J Clin Microbiol 2003;41:4705–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Raja S, Ching J, Xi L, Hughes SJ, Chang R, Wong W et al. Technology for automated, rapid, and quantitative PCR or reverse transcription-PCR clinical testing. Clin Chem 2005;51:882–90. [DOI] [PubMed] [Google Scholar]
- 4. Wolk DM, Struelens MJ, Pancholi P, Davis T, Della-Latta P, Fuller D et al. Rapid detection of Staphylococcus aureus and methicillin-resistant S. aureus (MRSA) in wound specimens and blood cultures: multicenter preclinical evaluation of the Cepheid Xpert MRSA/SA skin and soft tissue and blood culture assays. J Clin Microbiol 2009;47:823–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Forrest GN, Mankes K, Jabra-Rizk MA, Weekes E, Johnson JK, Lincalis DP, Venezia RA. Peptide nucleic acid fluorescence in situ hybridization-based identification of Candida albicans and its impact on mortality and antifungal therapy costs. J Clin Microbiol 2006;44:3381–3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Forrest GN, Mehta S, Weekes E, Lincalis DP, Johnson JK, Venezia RA. Impact of rapid in situ hybridization testing on coagulase-negative staphylococci positive blood cultures. J Antimicrob Chemother 2006;58:154–8. [DOI] [PubMed] [Google Scholar]
- 7. Sampath R, Hofstadler SA, Blyn LB, Eshoo MW, Hall TA, Massire C et al. Rapid identification of emerging pathogens: coronavirus. Emerg Infect Dis 2005;11:373–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Blyn LB, Hall TA, Libby B, Ranken R, Sampath R, Rudnick K et al. Rapid detection and molecular serotyping of adenovirus by use of PCR followed by electrospray ionization mass spectrometry. J Clin Microbiol 2008;46:644–51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Sampath R, Hall TA, Massire C, Li F, Blyn LB, Eshoo MW et al. Rapid identification of emerging infectious agents using PCR and electrospray ionization mass spectrometry. Ann NY Acad Sci 2007;1102:109–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Ecker DJ, Sampath R, Blyn LB, Eshoo MW, Ivy C, Ecker JA et al. Rapid identification and strain-typing of respiratory pathogens for epidemic surveillance. Proc Natl Acad Sci U S A 2005;102:8012–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Ecker JA, Massire C, Hall TA, Ranken R, Pennella TT, Agasino IC et al. Identification of Acinetobacter species and genotyping of Acinetobacter baumannii by multilocus PCR and mass spectrometry. J Clin Microbiol 2006;44:2921–32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Hall TA, Sampath R, Blyn LB, Ranken R, Ivy C, Melton R et al. Rapid molecular genotyping and clonal complex assignment of Staphylococcus aureus isolates by PCR coupled to electrospray ionization-mass spectrometry. J Clin Microbiol 2009;47:1733–41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Wolk DM, Blyn LB, Hall TA, Sampath R, Ranken R, Ivy C et al. Pathogen profiling: rapid molecular characterization of Staphylococcus aureus by PCR/ESI-MS and correlation with phenotype. J Clin Microbiol 2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Kaleta EJ, Clark AE, Johnson DR, Gamage DC, Wysocki VH, Cherkaoui A et al. Use of polymerase chain reaction coupled to electrospray ionization mass spectrometry for rapid identification of bacteria and yeast bloodstream pathogens from blood culture bottles. J Clin Microbiol 2011;49:345–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Claydon MA, Davey SN, Edwards-Jones V, Gordon DB. The rapid identification of intact microorganisms using mass spectrometry. Nat Biotechnol 1996;14:1584–6. [DOI] [PubMed] [Google Scholar]
- 16. Holland RD, Wilkes JG, Rafii F, Sutherland JB, Persons CC, Voorhees KJ, Lay JO. Rapid identification of intact whole bacteria based on spectral patterns using matrix-assisted laser desportion/ionization with time-of-flight mass spectrometry. Rapid Commun Mass Spectrom 1996;10:1227–32. [DOI] [PubMed] [Google Scholar]
- 17. Jarman KH, Daly DS, Petersen CE, Saenz AJ, Valentine NB, Wahl KL. Extracting and visualizing matrix-assisted laser desorption/ionization time-of-flight mass spectral fingerprints. Rapid Commun Mass Spectrom 1999;13:1586–94. [DOI] [PubMed] [Google Scholar]
- 18. Ilina EN, Borovskaya AD, Malakhova MM, Vereshchagin VA, Kubanova AA, Kruglov AN et al. Direct bacterial profiling by matrix-assisted laser desorption-ionization time-of-flight mass spectrometry for identification of pathogenic Neisseria. J Mol Diagn 2009;11:75–86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Nagy E, Maier T, Urban E, Terhes G, Kostrzewa M. Species identification of clinical isolates of Bacteroides by matrix-assisted laser-desorption/ionization time-of-flight mass spectrometry. Clin Microbiol Infect 2009;15:796–802. [DOI] [PubMed] [Google Scholar]
- 20. Alispahic M, Hummel K, Jandreski-Cvetkovic D, Nobauer K, Razzazi-Fazeli E, Hess M, Hess C. Species-specific identification and differentiation of Arcobacter, Helicobacter and Campylobacter by full-spectral matrix-associated laser desorption/ionization time of flight mass spectrometry analysis. J Med Microbiol 2010;59:295–301. [DOI] [PubMed] [Google Scholar]
- 21. Mellmann A, Cloud J, Maier T, Keckevoet U, Ramminger I, Iwen P et al. Evaluation of matrix-assisted laser desorption ionization-time-of-flight mass spectrometry in comparison to 16S rRNA gene sequencing for species identification of nonfermenting bacteria. J Clin Microbiol 2008;46:1946–54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Mellmann A, Bimet F, Bizet C, Borovskaya AD, Drake RR, Eigner U et al. High interlaboratory reproducibility of matrix-assisted laser desorption ionization-time of flight mass spectrometry-based species identification of nonfermenting bacteria. J Clin Microbiol 2009;47:3732–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Prod’hom G, Bizzini A, Durussel C, Bille J, Greub G. Matrix-assisted laser desorption ionization-time of flight mass spectrometry for direct bacterial identification from positive blood culture pellets. J Clin Microbiol 2010;48:1481–3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Stevenson LG, Drake SK, Murray PR. Rapid identification of bacteria in positive blood culture broths by matrix-assisted laser desorption ionization-time of flight mass spectrometry. J Clin Microbiol 2010;48:444–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Murray PR, Baron EJ, Pfaller MA, Tenover FC, Yolken RH, eds. Manual of clinical microbiology. Washington (DC): American Society of Microbiology; 2010. 1773 p. [Google Scholar]
- 26. CLSI. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically: approved standard—eighth edition. CLSI document M07–A8. Wayne (PA): CLSI; 2009. Vol. 29, no. 2. [Google Scholar]
- 27. CLSI. Performance standards for antimicrobial disk susceptibility tests: approved standard—tenth edition. CLSI document M02–A10. Wayne (PA): CLSI; 2009. Vol. 29, no. 1. [Google Scholar]
- 28. CLSI. Performance standards for antimicrobial susceptibility testing; twentieth informational supplement. CLSI Document M100–S20. [Wayne (PA)]: CLSI; 2010. Vol. 30, no. 1. [Google Scholar]
- 29. Newcombe RG. Two-sided confidence intervals for the single proportion: comparison of seven methods. Stat Med 1998;17:857–72. [DOI] [PubMed] [Google Scholar]
- 30. Eigner U, Holfelder M, Oberdorfer K, Betz-Wild U, Bertsch D, Fahr AM. Performance of a matrix-assisted laser desorption ionization-time-of-flight mass spectrometry system for the identification of bacterial isolates in the clinical routine laboratory. Clin Lab 2009;55:289–96. [PubMed] [Google Scholar]
- 31. Boutros N, Gonullu N, Casetta A, Guibert M, Ingrand D, Lebrun L. Ralstonia pickettii traced in blood culture bottles. J Clin Microbiol 2002;40:2666–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Dubois D, Leyssene D, Chacornac JP, Kostrzewa M, Schmit PO, Talon R et al. Identification of a variety of Staphylococcus species by matrix-assisted laser desorption ionization-time of flight mass spectrometry. J Clin Microbiol 2010;48:941–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Seibold E, Maier T, Kostrzewa M, Zeman E, Splettstoesser W. Identification of Francisella tularensis by whole-cell matrix-assisted laser desorption ionization-time of flight mass spectrometry: fast, reliable, robust, and cost-effective differentiation on species and subspecies levels. J Clin Microbiol 2010;48:1061–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Barbuddhe SB, Maier T, Schwarz G, Kostrzewa M, Hof H, Domann E et al. Rapid identification and typing of listeria species by matrix-assisted laser desorption ionization-time of flight mass spectrometry. Appl Environ Microbiol 2008;74:5402–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Peters RP, van Agtmael MA, Danner SA, Savelkoul PH, Vandenbroucke-Grauls CM. New developments in the diagnosis of bloodstream infections. Lancet Infect Dis 2004;4:751–60. [DOI] [PubMed] [Google Scholar]
- 36. La SB, Raoult D. Direct identification of bacteria in positive blood culture bottles by matrix-assisted laser desorption ionisation time-of-flight mass spectrometry. PLoS One 2009;4:e8041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Christner M, Rohde H, Wolters M, Sobottka I, Wegscheider K, Aepfelbacher M. Rapid identification of bacteria from positive blood culture bottles by use of matrix-assisted laser desorption-ionization time of flight mass spectrometry fingerprinting. J Clin Microbiol 2010;48:1584–91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Barenfanger J, Drake C, Kacich G. Clinical and financial benefits of rapid bacterial identification and antimicrobial susceptibility testing. J Clin Microbiol 1999;37:1415–8. [DOI] [PMC free article] [PubMed] [Google Scholar]


