Abstract
Severe Acute Respiratory Syndrome (SARS) is caused by a novel coronavirus (SARS-CoV). Coronaviruses including SARS-CoV encode an envelope (E) protein, a small, hydrophobic membrane protein. We report that, in planar lipid bilayers, synthetic peptides corresponding to the SARS-CoV E protein forms ion channels that are more permeable to monovalent cations than to monovalent anions. Affinity-purified polyclonal antibodies recognizing the N-terminal 19 residues of SARS-CoV E protein were used to establish the specificity of channel formation by inhibiting the ion currents generated in the presence of the E protein peptides.
Keywords: Severe Acute Respiratory Syndrome (SARS), Coronavirus, Budding, E protein, Ion channel, Conductance, Vpu, Hydrophobic, Monovalent cation, Mouse hepatitis virus (MHV)
Introduction
Severe Acute Respiratory Syndrome (SARS) is a respiratory illness that has recently been reported in Asia, North America, and Europe (Drosten et al., 2003, Ksiazek et al., 2003, Peiris et al., 2003). The World Health Organization (WHO) reported that the cumulative number of probable cases of SARS between 16 November 2002 and 7 August 2003 was 8422 with 916 deaths, just over a 10% death rate (World Health Organization, 2003). Coronaviruses [Order Nidovirales, family Coronaviridae, Genus Coronavirus (Gonzalez et al., 2003)] are enveloped, positive-stranded RNA viruses that bud from the endoplasmic reticulum–Golgi intermediate compartment or the cis-Golgi network (Klumperman et al., 1994, Krijnse-Locker et al., 1994, Tooze et al., 1984). The coronavirus envelope characteristically contains two major viral structural proteins, the spike (S) glycoprotein (for review, see Gallagher and Buchmeier, 2001) and the membrane (M) glycoprotein. A third minor but important membrane protein is the envelope (E) protein (Siddell, 1995a). E protein is a small, 9–12 kDa integral membrane protein (Siddell, 1995b). The N-terminus consists of a short 7–9 amino acid hydrophilic region and a 21–29 amino acid hydrophobic region, followed by a hydrophilic C-terminal region (Shen et al., 2003). E proteins play a part in viral assembly and morphogenesis. Co-expression of E and M proteins, from mouse hepatitis virus (MHV) (Bos et al., 1996, Vennema et al., 1996), transmittable gastroenteritis virus (TGEV), Bovine coronavirus (BCoV) (Baudoux et al., 1998), infectious bronchitis virus (IBV) (Corse and Machamer, 2000), and SARS-CoV (Ho et al., 2004) results in nucleocapsid independent formation of virus-like particles (VLPs). Additionally, MHV and IBV E protein expressed alone results in assembly of E-protein-containing vesicles, with a density similar to that of VLPs (Corse and Machamer, 2000, Maeda et al., 1999). Further evidence of E protein's important role in virus assembly and morphogenesis is that MHV E gene mutants have aberrant morphology with pinched and elongated shapes (Fischer et al., 1998). Recently, MHV recombinant virus was constructed with the entire E gene deleted. The virus was able to replicate to low titer and had a small plaque phenotype, indicating that E protein although important, is not essential for MHV replication in vitro (Kuo and Masters, 2003, Shen et al., 2003). In contrast, E protein expression is essential for TGEV replication, as deletion of the TGEV E protein is lethal and replication of the mutant virus can be rescued by TGEV E protein expressed in trans (Ortego et al., 2002).
Primarily due to their small size, and hydrophobic nature, coronavirus E proteins are candidate members of a family of virus-encoded proteins that form ion channels. (Fischer and Sansom, 2002, Gonzalez and Carrasco, 2003). These include influenza virus proteins, M2 (Duff and Ashley, 1992, Duff et al., 1994, Pinto et al., 1992) BM2 (Mould et al., 2003) and NB (Sunstrom et al., 1996), HIV-1 proteins, Vpu (Ewart et al., 1996), and Vpr (Piller et al., 1996), the 6K protein of alphaviruses (Melton et al., 2002), and, recently, the hepatitis C virus (HCV) protein, p7 (Griffin et al., 2003, Pavlovic et al., 2003, Premkumar et al., 2004). Based on similarities to other virus ion channels, we hypothesized that the SARS-CoV E protein might form ion channels. In this paper, we demonstrate that SARS-CoV E protein does form ion channels, which are more selective for monovalent cations than monovalent anions.
Results
Peptide synthesis and purification
A synthetic 76-residue peptide, corresponding to the full-length SARS-CoV E protein (isolate Tor2 and Urbani), was prepared using solid phase chemistry as described in Materials and methods. A second peptide was also synthesized, corresponding to the N-terminal 40 amino acids of the E protein and encompassing the hydrophobic domain. These are referred to as “full-length peptide” and “N-terminal peptide”, respectively.
MALDI-TOF mass spectrometry of the synthetic peptides revealed that, in addition to the expected products, the crude preparations also contained molecules with lower mass to charge (m/z) ratio, presumably corresponding to truncated peptides generated during chemical synthesis process. After purification to enrich the desired products—as described in Materials and methods—the mass spectra showed a predominant, well-isolated peak at m/z ratio of 8360.1 for the full-length peptide, corresponding to the calculated molecular weight of the SARS E protein (Fig. 1A). Similarly, a prevalent peak at the expected m/z ratio of 4422.3 was seen in the purified preparations of the N-terminal peptide (Fig. 1B). The spectra of the N-terminal peptide indicated that the preparation was clean with respect to low molecular weight contaminants; however, in contrast to the full-length peptide, peaks of decreasing intensity are present, corresponding to products sequentially truncated by single amino acid residues.
Fig. 1.

SARS-CoV full-length and N-terminal E protein mass spectral analysis. Full-length E protein shows a predominant peak at m/z ratio of 8360.1, the expected molecular weight. The N-terminal peptide shows a prevalent peak at 4422.3 m/z, peaks of decreasing intensity are present, corresponding to products sequentially truncated by single amino acid residues.
Ion channel formation in planar lipid bilayers
SARS-CoV E protein full-length and N-terminal peptides were tested for their ability to form ion channels in planar lipid bilayers. Initial experiments were done with 500 mM NaCl in the CIS chamber and 50 mM NaCl in the TRANS chamber. Small aliquots of the peptides dissolved in TFE were added to the CIS chamber while stirring to facilitate collision between peptide and bilayer. Spontaneous insertion of ion channel forming peptides into bilayers from TFE solution has been demonstrated previously with a number of small hydrophobic peptides (Ewart et al., 1996, Griffin et al., 2003, Melton et al., 2002, Pavlovic et al., 2003, Premkumar et al., 2004, Schubert et al., 1996b, Sunstrom et al., 1996). In general, ion currents due to SARS-CoV E protein were observed about 5–15 min after peptide was added to the CIS chamber in over 60 experiments. Initiation of channel activity was detected more rapidly and reliably when a holding potential of approximately −100 mV was applied across the bilayer, suggesting that peptide insertion into the bilayer was facilitated by a negative potential (TRANS relative to CIS).
Currents recorded at −88 mV and at −48 mV from a representative experiment with full-length E protein are shown in Fig. 2A, with all points histograms at each potential shown in parts B and C, respectively. In this experiment, the direction of current flow was observed to reverse between +40 and +50 mV and using an interpolated value of +48 mV the conductance was calculated to be 52 pS at −88 mV and 47 pS at −48 mV. Fig. 3A shows typical current traces of full-length E protein ion channel, recorded over a range of holding potentials, showing that the current–voltage (IV) relationship is not linear with conductance being relatively larger at more negative holding potentials (Fig. 3B). If the IV curve is linear, the channel has a constant conductance equal to the slope of the line. Some channels have conductances that vary with voltage and the IV is therefore nonlinear, the channel has larger current at higher potentials. The currents we measure are normally aggregated currents from a number of ion channels and the current amplitude (I) is given by NP o × gamma(V m − V r), where N is the total number of channels in the bilayer, P o is the average open probability of these channels, gamma is single channel conductance, V m is the potential applied across the bilayer membrane, and V r is the potential at which currents reverse direction. As seen with other ion channels, it is not unlikely that P o may be potential-dependent and this would give the nonlinear IV curves we see. Note, however, that the reversal potential is not affected by the voltage dependence of P o.
Fig. 2.
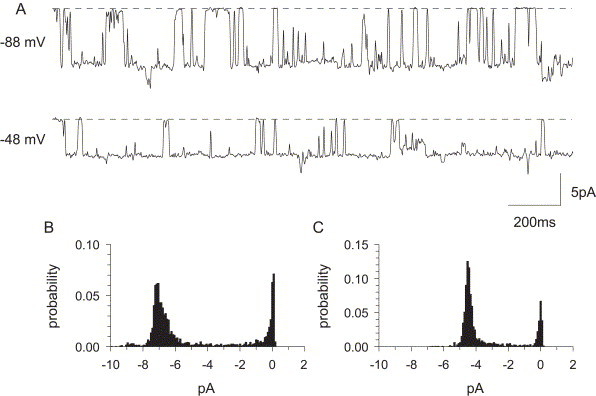
SARS-CoV E protein ion channel currents in lipid bilayers. The CIS chamber contained 500 mM NaCl, 5 mM HEPES buffer (pH 7.2); the TRANS chamber contained 50 mM NaCl, 5 mM HEPES buffer (pH 7.2). (A) Sample traces of raw current data, filtered at 100 Hz. The closed state is shown as a broken line, openings are deviations from the line. Scale bars are 200 ms and 5 pA. Potential was held at −88 or −48 mV. Conductances were 52 or 47 pS, respectively. (B) All points histograms of currents shown in A at −88 and C at −48 mV.
Fig. 3.
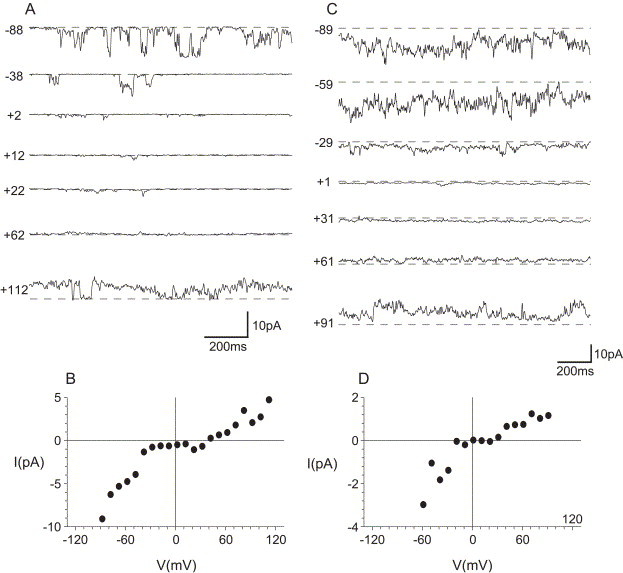
SARS-CoV E protein ion channel activity observed in NaCl or KCl solutions. Data filtered at 100 Hz. (A) The CIS chamber contained 500 mM NaCl, 5 mM HEPES buffer (pH 7.2); the TRANS chamber contained 50 mM NaCl, 5 mM HEPES buffer (pH 7.2). The CIS chamber was earthed and the TRANS chamber was held at various potentials between −88 and +112 mV. The closed state is shown as a broken line, openings are deviations from the line. Scale bars are 200 ms and 10 pA. (B) IV relationship in NaCl solution average current measured against holding potential (mV). (C) The CIS chamber contained 500 mM KCl, 5 mM HEPES buffer (pH 7.2); the TRANS chamber contained 50 mM KCl in 5 mM HEPES buffer (pH 7.2). The CIS chamber was earthed and the TRANS chamber was held at various potentials between −89 and +91 mV. The closed state is shown as a broken line, openings are deviations from the line. Scale bars are 200 ms and 10 pA. (D) IV relationship in KCl solution, average current measured against holding potential (mV).
In seven separate experiments with NaCl solution (Table 1 ), the average reversal potential was +48.3 ± 2.3 mV (mean ± SEM), which is close to the theoretical Na+ equilibrium potential of +54 mV (using activities in the Nernst equation). From this, it can be calculated that the ion channels are about 90 times more permeable to Na+ ions than to Cl− ions. For these seven experiments, the maximum conductance varied between 95 and 164 pS and the average conductance was 130 ± 13 pS.
Table 1.
Reversal potentials (mV)
| SARS E protein | NaCl |
KCl |
||
|---|---|---|---|---|
| Vr | Gamma | Vr | Gamma | |
| Full length | 48.3 ± 2.3 (n = 7) | 130 ± 13 pS | 34.5 ± 2.5 (n = 4) | 83.4 ± 26 pS |
| TM domain | 46.3 ± 2.5 (n = 7) | 35 ± 7 pS | 39.5 ± 3.6 (n = 5) | 93 ± 36 pS |
The channel formed by the E protein also conducted potassium ions, as shown in Fig. 3C over a range of holding potentials. In the experiment shown in Fig. 3D, with 500 mM KCl in the CIS chamber and 50 mM KCl in the TRANS chamber, the currents reversed at +31 mV. In four similar experiments, the average reversal potential was +34.5 ± 2.5 mV (Table 1) indicating that the SARS-CoV E protein ion channel is about nine times more permeable to K+ ions than to Cl− ions (Table 1). In those four experiments, the maximum conductance varied between 24 and 166 pS and the average conductance was 83.4 ± 26 pS (Table 1). The reversal potential is considerably less positive in KCl than in NaCl solution, indicating that the channel is relatively less selective for K+ than for Na+ ions.
The “N-terminal peptide”—corresponding to the first 40 amino acids of the SARS-CoV E protein—formed monovalent cation-selective channels with properties similar to those of the full-length peptide. The mean reversal potentials (V r) for currents formed by the full-length and N-terminal peptides measured in NaCl solutions were not significantly different from each other (see Table 1), nor were the V r measured in KCl solutions significantly different. However, for both peptides, comparison of the V r measured in KCl with that measured in NaCl solutions revealed a significant difference (P < 0.003). The ion selectivity series for both peptides was Na+ > K+ > Cl− with the channels being approximately 5–10 fold more permeable to Na+ than K+ and approximately 10-fold more permeable to K+ than Cl−. Given the above similarities, we conclude that pore-forming structure and selectivity filter for the SARS-CoV E protein are encoded within the first 40 amino acids of the N-terminal, as might be predicted from the hydropathy profile of the E protein.
In 10 control experiments in which no protein was added to the CIS chamber, no ion channel activity was detected, even during observation periods lasting for over 1 h. Therefore, channel formation was dependant on addition of peptide samples and was not an artifact of the lipids, buffers, or solvents alone.
Inhibition of peptide ion channels by N-terminal epitope-specific antibodies
As described above, mass spectral analysis established the presence and predominance of the full-length peptide (m/z ratio of 8360.1) in purified preparations. However, less intense peaks were also present indicating presence of minor contamination by smaller molecules not removed by the purification steps. These were mainly apparent below an m/z ratio of approximately 6000. Therefore, it was important to show that channel formation was specific to the full-length peptide and not the contaminants. To achieve this, we synthesized a third peptide corresponding to the first 19 N-terminal amino acids of the SARS E protein and used it to raise and purify polyclonal antibodies recognizing this domain (see Materials and methods). Previous results with other ion-channel-forming proteins have shown that such epitope-specific antibodies can inhibit channel activity, or be used to selectively remove the channel-forming species from preparations (Ewart et al., 1996, Melton et al., 2002, Sunstrom et al., 1996).
When the purified antibody was used to probe Western blots a ladder of four discrete immunoreactive bands was seen in lanes in which either the purified full-length (Fig. 4A) or N-terminal peptide (Fig. 4B) had been run, indicating that these peptides form aggregates. The aggregates were not disrupted by addition of reducing agent, β-mercaptoethanol, nor were they significantly affected by boiling or not boiling the samples prior to PAGE (not shown). In the blot shown in Fig. 4, relatively large quantities of peptide were loaded per lane and the smallest bands detected (presumably monomeric peptide molecules) ran just above or below the 11 kDa marker for the full-length and N-terminal peptides, respectively. The observation that these putative monomers do not run at their theoretical molecular weights is not an unusual phenomenon for small hydrophobic peptides, where it is thought that the uniform binding of SDS may be affected, altering the relative rate of migration through polyacrylamide gels. For example, chemically cross-linked homo-oligomers of the HIV-1 protein Vpu do not run at their calculated molecular weights (Maldarelli et al., 1993). Importantly, the Western blots did not show any immunoreaction with lower molecular weight bands that might correlate to truncated peptides or contaminating species. By this criterion, the purified antibody was judged to be a specific reagent for binding to the N-terminal epitope of full-length and N-terminal peptides. Further, the anti-N-terminal antibody did not cross-react with a nonspecific protein antigen, such as the hepatitis C (HCV) p7 protein (Fig. 4, lanes 3).
Fig. 4.
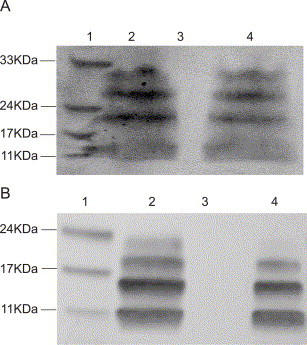
SARS-CoV full-length E and N-terminal peptide Western blot. Lane 1: marker; lane 2: without β-mercaptoethanol; lane 3: Hepatitis C p7; lane 4: with β-mercaptoethanol. (A) About 150 μg/lane full-length E protein detected with anti-N terminal polyclonal antibody. (B) About 30 μg/lane E protein N-terminal peptide detected with anti-N terminal polyclonal antibody.
In bilayer experiments, addition of approximately threefold molar excess of affinity-purified antibody recognizing the N-terminal domain of SARS-CoV E protein to the CIS chamber significantly reduced (P ≥ 0.005, n = 4 experiments) the current amplitude in NaCl solution as illustrated for one of the experiments in Fig. 5 . In contrast, addition of the antibody to the TRANS chamber did not affect ion channel conductance (Fig. 5). In separate experiments (not shown), the antibody did not affect ion channel conductance due to HCV p7 in planar lipid bilayers.
Fig. 5.
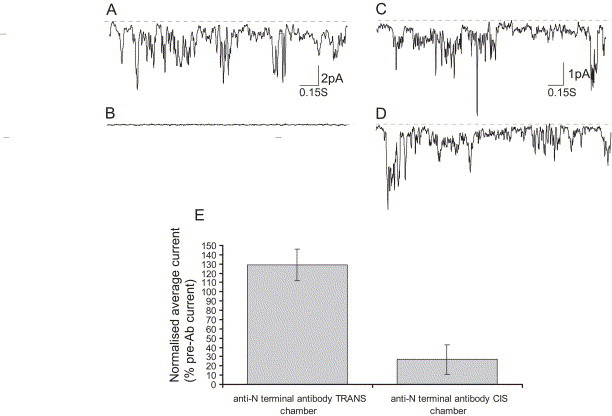
SARS-CoV E protein ion channel activity is blocked by anti-N-terminal affinity-purified polyclonal antibody. CIS chamber contained 500 mM NaCl, 5 mM HEPES buffer (pH 7.2); the TRANS chamber contained 50 mM NaCl, 5 mM HEPES buffer (pH 7.2). Representative current traces at holding potential of −60 mV. The closed state is shown as a broken line, openings are deviations from the line. Scale bars are 0.15 s and 2 pA. (A) E protein ion channel activity before addition of antibody. (B) E protein ion channel activity after addition of ≤20 μg anti-N-terminal antibody to the CIS chamber (n = 4). (C) E protein ion channel activity before addition of antibody. (D) E protein ion channel activity after addition of ≤20 μg anti-N-terminal antibody to the TRANS chamber (n = 4).
Taken together, antibody-specific inhibition of the ion conductances in planar lipid bilayers in an orientation-dependant manner (as shown in Fig. 5), and lack of detectable immunoreaction of the antibodies with low molecular weight species on Western blots confirms that ion channel formation in these experiments is due to peptides containing the SARS-CoV E protein N-terminal sequence and is not an artifact due to significantly truncated peptides or other contaminating molecules.
Discussion
We have shown that synthetic peptides corresponding to the full-length SARS-CoV E protein and its N-terminal domain form cation-selective ion channels in planar lipid bilayers. Affinity-purified polyclonal antibodies to the N-terminal domain of the SARS-CoV E protein specifically block the ion channel activity, supporting our conclusion that the ion channel activity is due to the SARS-CoV E protein and not to minor contaminants in the preparations. Addition of the anti-N-terminal antibody to the CIS chamber reduced average bilayer current generated by the SARS-CoV E protein by 73% ± 16 (1 SEM), whereas addition of the anti-N-terminal antibody to the TRANS chamber did not significantly reduce conductance. As well as confirming specificity, this result indicates that the N-terminus faces into the CIS chamber. With the exception of HCV p7, the viral ion channels are generally thought to span the bilayer once, as a single alpha helix, so it is tempting to assume that the C-terminus would be located in the TRANS chamber. However, very recently, evidence was presented that the SARS-CoV E protein transmembrane domain can form a palindromic transmembrane helical hairpin and incorporation of the helical hairpin into lipid vesicles increased membrane curvature by deforming lipid bilayers (Arbely et al., 2004), a known function of viral ion channel family proteins (Gonzalez and Carrasco, 2003). Clearly, if the SARS-CoV E protein forms such a helical hairpin under the conditions of our bilayer experiments, both the N- and C-termini would be located in the CIS chamber. From energetic considerations, it would seem that incorporation of a helical hairpin into lipid bilayers would be favored over the alternative, which requires a large number of hydrophilic amino acid residues to cross the bilayer into the TRANS chamber. Unfortunately, an antibody was not available to us with which to orient the C-terminus.
SARS-CoV E protein N-terminal first 40 amino acids were sufficient for the formation of ion channels with similar selectivity as the full-length E protein. Therefore, the C-terminal of the SARS-CoV E protein is not essential for ion channel conductance. It has also been shown that the Vpu C-terminal hydrophilic domain does not participate in ion channel conductance, and is not essential for regulation of virus release, although the presence of the cytoplasmic domain improves the function of Vpu, indicating that it may have a regulatory role (Schubert et al., 1996a, Schubert et al., 1996b). Likewise, it is possible that the C-terminal of coronavirus E proteins may have regulatory roles. Supporting this idea, site-directed mutations to a cluster of charged amino acids in the C-terminal of MHV E protein have been found to impair viral assembly and maturation (Fischer et al., 1998).
SARS-CoV E protein ion channels have similar ion-selectivity to previously described viral ion channels, Vpu, NB, 6K, and p7, which also form cation-selective ion channels in planar lipid bilayers (Ewart et al., 1996, Griffin et al., 2003, Melton et al., 2002, Pavlovic et al., 2003, Piller et al., 1996, Premkumar et al., 2004, Schubert et al., 1996a, Sunstrom et al., 1996). Channels formed by Vpu and p7 are equally selective for Na+ and K+ ions (Ewart et al., 1996, Premkumar et al., 2004). The NB ion channel is selective for Na+ at pH 5.5 to 6.5 (Sunstrom et al., 1996). The SARS-CoV E protein is more selective for Na+ than for K+ ions, and this has also been reported for 6K ion channels (Melton et al., 2002). The SARS-CoV E protein is 90 times more selective for Na+ than Cl− ions, more selective than reported for Vpu, 6K, or p7 ion channels (Ewart et al., 1996, Melton et al., 2002, Premkumar et al., 2004).
The large range of conductance states observed with the SARS E peptides have also been reported with other viral ion channels (Ewart et al., 1996, Melton et al., 2002, Premkumar et al., 2004, Sunstrom et al., 1996, Tosteson et al., 1994). It is possible that the larger conductances are due to opening of many clustered ion channels in the bilayer. Additionally, increased aggregation states of ion channels, due to formation of homo-oligomeric complexes, can result in variable ion conductances (Becker et al., 2004). Experiments are underway to determine the homo-oligomeric state of the SARS-CoV E protein.
The question of whether the ion channel activity of E protein is involved in the known physiological roles of these proteins in coronavirus replication remains to be investigated. The best-characterized virus ion channels are the M2 channel from influenza A and the Vpu ion channel from HIV-1. M2 ion channel activity has been linked to uncoating of the virion in the early endosome (Mould et al., 2000) and mediating transport of viral proteins through the excretory pathway during viral assembly late in infection (Mould et al., 2000, Sakaguchi et al., 1996). The hydrophobic N-terminal α-helical domain of Vpu self-associates in membranes and plays a part in the enhancement of budding and release of new virus particles from infected cells (Klimkait et al., 1990)—an activity that correlates with ion channel formation (Ewart et al., 1996, Schubert et al., 1996a, Schubert et al., 1996b).
The exact mechanism of Vpu-enhanced virus budding is unknown (for review, see Montal, 2003). Evidence suggests that Vpu plays a passive role in enhancement of viral budding, perhaps through generalized affects on the cellular environment, indirectly enhancing Gag trafficking to the plasma membrane (Deora et al., 2000, Gottlinger et al., 1993, Marassi et al., 1999, Montal, 2003, Schubert et al., 1996a). Gottlinger et al. (1993) have shown that Vpu can significantly enhance the release of Gag from heterologous retroviruses, such as HIV-2, visna virus, and Moloney murine leukemia virus, which do not normally express Vpu. Furthermore, Vpu can partially compensate for a Sindbis 6K-deleted virus (Gonzalez and Carrasco, 2001). Taken together, the data suggests that Vpu does not directly interact with Gag during virus budding (Gottlinger et al., 1993). Co-expression of Gag and Vpu results in the redistribution of Gag from the cytosol to the plasma membrane (Deora et al., 2000). In the absence of Vpu, budding has been observed from internal membranes (Klimkait et al., 1990).
Although Vpu does not directly interact with Gag, Vpu has been shown to associate with cellular proteins (Geraghty et al., 1994, Handley et al., 2001, Hsu et al., 2004). Recently, it has been shown that Vpu shares homology with the N-terminal domain of the mammalian background K+ channel, TASK-1. Interaction of TASK-1 and Vpu results in Vpu abolishing TASK-1 currents. Overexpression of TASK-1 impaired Vpu-enhanced viral budding; however, the N-terminal domain of TASK-1 was able to enhance HIV-1 particle release (Hsu et al., 2004). Different levels of TASK-1 expression in various cell lines (Hsu et al., 2004) may explain why Vpu viral enhancement is cell type dependent (Geraghty et al., 1994, Gottlinger et al., 1991, Yao et al., 1992). Furthermore, Vpu and Gag have been shown to bind to viral protein U-binding Protein (Ubp) (Callahan et al., 1998, Geraghty et al., 1994). Ubp may play a role in targeting Gag to the plasma membrane (Handley et al., 2001).
Vpu and SARS-CoV E protein have similar ion channel selectivity and both play a role in virus budding. E protein has been shown to be important in coronavirus assembly and morphogenesis (Bos et al., 1996, Fischer et al., 1998, Kuo and Masters, 2003, Vennema et al., 1996). Conceivably, E protein ion channel could change the cellular milieu at the budding site to enhance viral morphogenesis and assembly. The scientific challenge remains to determine how E protein ion channel activity mediates budding in the coronavirus lifecycle.
Materials and methods
Peptide synthesis and purification
A synthetic peptide corresponding to the full-length SARS-CoV (isolate Tor2 and Urbani GenBank accession number NC004718 and AY278741, respectively) E protein (MYSFVSEETGTLIVNSVLLFLAFVVFLLVTLAILTALRLCAYCCNIVNVSLVKPTVYVYSRVKNLNSSEGVPDLLV) and a second peptide corresponding to the first 40 amino acids in the N-terminal transmembrane domain of the E protein “N-terminal peptide” (MYSFVSEETGTLIVNSVLLFLAFVVFLLVTLAILTALRLC) were synthesized on a SYMPHONY/MULTIPLEX (Protein Technologies inc. Woburn, MS) multiple peptide synthesizer, using FMOC chemistry and solid phase peptide synthesis at the Biomolecular Research Facility, John Curtin School of Medical Research following manufacturer's instructions.
To enrich the appropriate full-length product from the crude synthesis reactions, the following procedure, relying on the differential solubility of the smaller molecules and full-length peptide, was devised. The crude preparation was suspended at 12 mg/ml in 70% CH3CN, 0.1% TFA, and vortexed for 10 min. This suspension was centrifuged at 10 000 × g for 10 min at 20 °C. The supernatant was discarded and the insoluble fraction was further extracted two more times with 70% CH3CN, 0.1% TFA, as above. The insoluble material containing the E protein was dried in a Speedvac and the weight of the final product was used to calculate the yield. The purified peptides were analyzed by Bruker Omniflex MALDI-TOF mass spectrometry in a HABA matrix at 2.5 mg/ml in methanol at a 1:1 ratio and spectra were obtained in the positive linear mode. A predominant peak at m/z ratio of 8360.1 was seen, as was expected for the calculated molecular weight of full-length E protein. Similarly, a prevalent peak at 4422.3 m/z was seen for the N-terminal peptide (see Fig. 1).
Raising and purifying antibodies against SARS-CoV E protein
A synthetic peptide corresponding to SARS-CoV E protein N-terminal 19 residues (MYSFVSEETGTLIVNSVLC) was coupled to a polylysine core via the terminal cysteine residue to prepare multiple antigenic peptides (MAP) (Lu et al., 1991). Individual, crossbred New Zealand White rabbit were immunized with the MAP peptides. Initial immunization was with 200 μg of peptide in Freund's complete adjuvant (Imject, Pierce, Rockford, IL). Boosters of 200 μg of peptide were given every 4 weeks in Freund's incomplete adjuvant (Imject, Pierce), until sufficient antibody titers were detected by Western blotting. About 15 ml of blood was collected by ear vein cannulation 2, 4, 6 and 8 weeks post-immunization and antisera prepared. Antiserum was assayed for antibody production by western blotting with the SARS-CoV full-length purified peptide. Specific antibodies were purified from antisera by SulfoLink columns (Pierce) following manufacturer's instructions.
Western blots
For Western blots, samples were prepared in loading buffer (60 mM Tris–HCl pH 8.3, 6 M urea, 5% SDS, 10% glycerol, 0.2% bromophenol blue, ±100 mM β-mercaptoethanol) and run with molecular weight markers (MBI Fermentas, Hanover, MD) on 4–20% gradient polyacrylamide gels (Gradipore, NSW, Australia). Peptides were transferred to polyvinylidene difluoride membranes (Invitrogen, Vic, Australia), using a semidry transfer apparatus (Amersham Biosciences, Vic, Australia). Nonspecific sites were blocked with skim milk proteins in Tris-buffered saline containing 1% Tween 20. Primary antibody was polyclonal anti-N terminal (1:250), which was detected with goat anti-rabbit IgG alkaline phosphatase-conjugated antibody (Dako, NSW, Australia). Color development was detected with Western Blue-stabilized substrate for alkaline phosphatase (Promega, NSW).
Ion channel recording
The SARS-CoV full-length and N-terminal E protein were resuspended at 1 mg/ml in 2,2,2-trifluoroethanol and their ability to form ion channels was tested on a Warner bilayer rig (Warner Instruments, Inc. 1125 Dixwell Avenue, Hamden, CT 06514) as follows: A lipid mix of 3:1:1, 1-palmitoyl-2-oleolyl phosphatidyl ethanolamine/1-palmitoyl-2-oleolyl phosphatidyl serine/1-palmitoyl-2-oleolyl phosphatidyl choline in CHCl3 were dried under N2 gas and resuspended in n-decane. Bilayers were painted across a circular hole of approximately 100-μm diameter in a Delrin cup separating aqueous solution in the CIS and TRANS chambers. For testing E protein selectivity for Na+ and Cl− ions, the solution in the CIS chamber consisted of 500 mM NaCl and 5 mM HEPES (pH 7.2), while the solution in the TRANS chamber contained 50 mM NaCl and 5 mM HEPES (pH 7.2). For testing E protein selectivity for K+ and Cl− ions, the solution in the CIS chamber consisted of 500 mM KCl, and 5 mM HEPES (pH 7.2), while the solution in the TRANS chamber contained 50 mM KCl and 5 mM HEPES (pH 7.2). Typically, 3 μg of purified synthetic full-length or N-terminal E protein was added to the CIS chamber while stirring until ion channel activity was detected. The CIS chamber was earthed and the TRANS chamber was held at various holding potentials ranging between +100 to −100 mV. Currents were amplified using a Warner model BD-525D amplifier with sampling rate of 5 kHz and filtered at 1 kHz before being digitally recorded directly using the Data Collect software developed by Mr. Bernie Keys (BioResearch Electronics, Canberra, Australia).
In some experiments in which ion channel activity was detected, affinity-purified anti-N-terminal polyclonal antibody was added to the CIS or TRANS chamber to a final concentration of about 10–20 μg/ml, while stirring for 30 s. A t test (SPSS Software, SPSS Inc. Chicago, IL) was used to determine whether there was significant difference between the mean current recorded before and after addition of the antibody.
Acknowledgments
The authors would like to thank Frank Bowden for his critical reading of the manuscript; Victoria Hodgson for her assistance with data analysis; and Louise Cengia, Lisa Teakle, and Tammy Gomersall for their technical assistance.
References
- Arbely E., Khattari Z., Brotons G., Akkawi M., Salditt T., Arkin I.T. A highly unusual palindromic transmembrane helical hairpin formed by SARS coronavirus E protein. J. Mol. Biol. 2004;341:769–779. doi: 10.1016/j.jmb.2004.06.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baudoux P., Carrat C., Besnardeau L., Charley B., Laude H. Coronavirus pseudoparticles formed with recombinant M and E proteins induce alpha interferon synthesis by leukocytes. J. Virol. 1998;72:8636–8643. doi: 10.1128/jvi.72.11.8636-8643.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Becker C.F., Oblatt-Montal M., Kochendoerfer G.G., Montal M. Chemical synthesis and single channel properties of tetrameric and pentameric TASP proteins derived from the transmembrane domain of HIV Virus protein u-Vpu. J. Biol. Chem. 2004:17483–17489. doi: 10.1074/jbc.M313212200. [DOI] [PubMed] [Google Scholar]
- Bos E.C., Luytjes W., van der Meulen H.V., Koerten H.K., Spaan W.J. The production of recombinant infectious DI-particles of a murine coronavirus in the absence of helper virus. Virology. 1996;218:52–60. doi: 10.1006/viro.1996.0165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Callahan M.A., Handley M.A., Lee Y.H., Talbot K.J., Harper J.W., Panganiban A.T. Functional interaction of human immunodeficiency virus type 1 Vpu and Gag with a novel member of the tetratricopeptide repeat protein family. J. Virol. 1998;72:5189–5197. doi: 10.1128/jvi.72.6.5189-5197.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corse E., Machamer C.E. Infectious bronchitis virus E Protein is targeted to the Golgi complex and directs release of virus-like particles. J. Virol. 2000;74:4319–4326. doi: 10.1128/jvi.74.9.4319-4326.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deora A., Spearman P., Ratner L. The N-terminal matrix domain of HIV-1 Gag is sufficient but not necessary for viral protein U-mediated enhancement of particle release through a membrane-targeting mechanism. Virology. 2000;269:305–312. doi: 10.1006/viro.1999.0094. [DOI] [PubMed] [Google Scholar]
- Drosten C., Gunther S., Preiser W., van der Werf S., Brodt H.-R., Becker S., Rabenau H., Panning M., Kolesnikova L., Fouchier R.A.M., Berger A., Burguiere A.-M., Cinatl J., Eickmann M., Escriou N., Grywna K., Kramme S., Manuguerra J.-C., Muller S., Rickerts V., Sturmer M., Vieth S., Klenk H.-D., Osterhaus A.D.M.E., Schmitz H., Doerr H.W. Identification of a novel coronavirus in patients with Severe Acute Respiratory Syndrome. N. Engl. J. Med. 2003;348:1967–1976. doi: 10.1056/NEJMoa030747. [DOI] [PubMed] [Google Scholar]
- Duff K.C., Ashley R.H. The transmembrane domain of influenza A M2 protein forms amantadine-sensitive proton channels in planar lipid bilayers. Virology. 1992;190:485–489. doi: 10.1016/0042-6822(92)91239-q. [DOI] [PubMed] [Google Scholar]
- Duff K.C., Gilchrist P.J., Saxena A.M., Bradshaw J.P. Neutron diffraction reveals the site of amantadine blockade in the influenza A M2 ion channel. Virology. 1994;202:287–293. doi: 10.1006/viro.1994.1345. [DOI] [PubMed] [Google Scholar]
- Ewart G.D., Sutherland T., Gage P.W., Cox G.B. The Vpu protein of human immunodeficiency virus type 1 forms cation-selective ion channels. J. Virol. 1996;70:7108–7115. doi: 10.1128/jvi.70.10.7108-7115.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fischer W.B., Sansom M.S. Viral ion channels: structure and function. Biochim. Biophys. Acta. 2002;1561:27–45. doi: 10.1016/s0304-4157(01)00009-0. [DOI] [PubMed] [Google Scholar]
- Fischer F., Stegen C.F., Masters P.S., Samsonoff W.A. Analysis of constructed E gene mutants of mouse hepatitis virus confirms a pivotal role for E protein in coronavirus assembly. J. Virol. 1998;72:7885–7894. doi: 10.1128/jvi.72.10.7885-7894.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gallagher T.M., Buchmeier M.J. Coronavirus spike proteins in viral entry and pathogenesis. Virology. 2001;279:371–374. doi: 10.1006/viro.2000.0757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geraghty R.J., Talbot K.J., Callahan M., Harper W., Panganiban A.T. Cell type-dependence for Vpu function. J. Med. Primatol. 1994;23:146–150. doi: 10.1111/j.1600-0684.1994.tb00115.x. [DOI] [PubMed] [Google Scholar]
- Gonzalez M.E., Carrasco L. Human immunodeficiency virus type 1 VPU protein affects Sindbis virus glycoprotein processing and enhances membrane permeabilization. Virology. 2001;279:201–209. doi: 10.1006/viro.2000.0708. [DOI] [PubMed] [Google Scholar]
- Gonzalez M.E., Carrasco L. Viroporins. FEBS Lett. 2003;552:28–34. doi: 10.1016/s0014-5793(03)00780-4. [DOI] [PubMed] [Google Scholar]
- Gonzalez J.M., Gomez-Puertas P., Cavanagh D., Gorbalenya A.E., Enjuanes L. A comparative sequence analysis to revise the current taxonomy of the family Coronaviridae. Arch. Virol. 2003;148:2207–2235. doi: 10.1007/s00705-003-0162-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gottlinger H.G., Dorfman T., Sodroski J.G., Haseltine W.A. Effect of mutations affecting the p6 gag protein on human immunodeficiency virus particle release. Proc. Natl. Acad. Sci. U.S.A. 1991;88:3195–3199. doi: 10.1073/pnas.88.8.3195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gottlinger H.G., Dorfman T., Cohen E.A., Haseltine W.A. Vpu protein of human immunodeficiency virus type 1 enhances the release of capsids produced by gag gene constructs of widely divergent retroviruses. Proc. Natl. Acad. Sci. U.S.A. 1993;90:7381–7385. doi: 10.1073/pnas.90.15.7381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Griffin S.D., Beales L.P., Clarke D.S., Worsfold O., Evans S.D., Jaeger J., Harris M.P., Rowlands D.J. The p7 protein of hepatitis C virus forms an ion channel that is blocked by the antiviral drug, Amantadine. FEBS Lett. 2003;535:34–38. doi: 10.1016/s0014-5793(02)03851-6. [DOI] [PubMed] [Google Scholar]
- Handley M.A., Paddock S., Dall A., Panganiban A.T. Association of Vpu-binding protein with microtubules and Vpu-dependent redistribution of HIV-1 Gag protein. Virology. 2001;291:198–207. doi: 10.1006/viro.2001.1166. [DOI] [PubMed] [Google Scholar]
- Ho Y., Lin P.-H., Liu C.Y.Y., Lee S.-P., Chao Y.-C. Assembly of human Severe Acute Respiratory Syndrome coronavirus-like particles. Biochem. Biophys. Res. Commun. 2004;318:833–838. doi: 10.1016/j.bbrc.2004.04.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsu K., Seharaseyon J., Dong P., Bour S., Marban E. Mutual functional destruction of HIV-1 Vpu and host TASK-1 channel. Mol. Cell. 2004;14:259–267. doi: 10.1016/s1097-2765(04)00183-2. [DOI] [PubMed] [Google Scholar]
- Klimkait T., Strebel K., Hoggan M.D., Martin M.A., Orenstein J.M. The human immunodeficiency virus type 1-specific protein vpu is required for efficient virus maturation and release. J. Virol. 1990;64:621–629. doi: 10.1128/jvi.64.2.621-629.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klumperman J., Locker J.K., Meijer A., Horzinek M.C., Geuze H.J., Rottier P.J. Coronavirus M proteins accumulate in the Golgi complex beyond the site of virion budding. J. Virol. 1994;68:6523–6534. doi: 10.1128/jvi.68.10.6523-6534.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krijnse-Locker J., Ericsson M., Rottier P., Griffiths G. Characterization of the budding compartment of mouse hepatitis virus: evidence that transport from the RER to the Golgi complex requires only one vesicular transport step. J. Cell Biol. 1994;124:55–70. doi: 10.1083/jcb.124.1.55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ksiazek T.G., Erdman D., Goldsmith C.S., Zaki S.R., Peret T., Emery S., Tong S., Urbani C., Comer J.A., Lim W., Rollin P.E., Dowell S.F., Ling A.-E., Humphrey C.D., Shieh W.-J., Guarner J., Paddock C.D., Rota P., Fields B., DeRisi J., Yang J.-Y., Cox N., Hughes J.M., LeDuc J.W., Bellini W.J., Anderson L.J., the SARS Working Group A novel coronavirus associated with Severe Acute Respiratory Syndrome. N. Engl. J. Med. 2003;348:1953–1966. doi: 10.1056/NEJMoa030781. [DOI] [PubMed] [Google Scholar]
- Kuo L., Masters P.S. The small envelope protein E is not essential for murine coronavirus replication. J. Virol. 2003;77:4597–4608. doi: 10.1128/JVI.77.8.4597-4608.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu Y.A., Clavijo P., Galantino M., Shen Z.Y., Liu W., Tam J.P. Chemically unambiguous peptide immunogen: preparation, orientation and antigenicity of purified peptide conjugated to the multiple antigen peptide system. Mol. Immunol. 1991;28:623–630. doi: 10.1016/0161-5890(91)90131-3. [DOI] [PubMed] [Google Scholar]
- Maeda J., Maeda A., Makino S. Release of coronavirus E protein in membrane vesicles from virus-infected cells and E protein-expressing cells. Virology. 1999;263:265–272. doi: 10.1006/viro.1999.9955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maldarelli F., Chen M.Y., Willey R.L., Strebel K. Human immunodeficiency virus type 1 Vpu protein is an oligomeric type I integral membrane protein. J. Virol. 1993;67:5056–5061. doi: 10.1128/jvi.67.8.5056-5061.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marassi F.M., Ma C., Gratkowski H., Straus S.K., Strebel K., Oblatt-Montal M., Montal M., Opella S.J. Correlation of the structural and functional domains in the membrane protein Vpu from HIV-1. Proc. Natl. Acad. Sci. U.S.A. 1999;96:14336–14341. doi: 10.1073/pnas.96.25.14336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Melton J.V., Ewart G.D., Weir R.C., Board P.G., Lee E., Gage P.W. Alphavirus 6 K proteins form ion channels. J. Biol. Chem. 2002:46923–46931. doi: 10.1074/jbc.M207847200. [DOI] [PubMed] [Google Scholar]
- Montal M. Structure–function correlates of Vpu, a membrane protein of HIV-1. FEBS Lett. 2003;552:47–53. doi: 10.1016/s0014-5793(03)00849-4. [DOI] [PubMed] [Google Scholar]
- Mould J.A., Drury J.E., Frings S.M., Kaupp U.B., Pekosz A., Lamb R.A., Pinto L.H. Permeation and activation of the M2 ion channel of influenza A virus. J. Biol. Chem. 2000;275:31038–31050. doi: 10.1074/jbc.M003663200. [DOI] [PubMed] [Google Scholar]
- Mould J.A., Paterson R.G., Takeda M., Ohigashi Y., Venkataraman P., Lamb R.A., Pinto L.H. Influenza B Virus BM2 Protein Has Ion Channel Activity that Conducts Protons across Membranes. Dev. Cell. 2003;5:175–184. doi: 10.1016/s1534-5807(03)00190-4. [DOI] [PubMed] [Google Scholar]
- Ortego J., Escors D., Laude H., Enjuanes L. Generation of a replication-competent, propagation-deficient virus vector based on the transmissible gastroenteritis coronavirus genome. J. Virol. 2002;76:11518–11529. doi: 10.1128/JVI.76.22.11518-11529.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pavlovic D., Neville D.C., Argaud O., Blumberg B., Dwek R.A., Fischer W.B., Zitzmann N. The hepatitis C virus p7 protein forms an ion channel that is inhibited by long-alkyl-chain iminosugar derivatives. Proc. Natl. Acad. Sci. U.S.A. 2003;100:6104–6108. doi: 10.1073/pnas.1031527100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peiris J., Lai S., Poon L., Guan Y., Yam L., Lim W., Nicholls J., Yee W., Yan W., Cheung M. Coronavirus as a possible cause of Severe Acute Respiratory Syndrome. Lancet. 2003;361:1319–1325. doi: 10.1016/S0140-6736(03)13077-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piller S.C., Ewart G.D., Premkumar A., Cox G.B., Gage P.W. Vpr protein of human immunodeficiency virus type 1 forms cation-selective channels in planar lipid bilayers. Proc. Natl. Acad. Sci. U.S.A. 1996;93:111–115. doi: 10.1073/pnas.93.1.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pinto L.H., Holsinger L.J., Lamb R.A. Influenza virus M2 protein has ion channel activity. Cell. 1992;69:517–528. doi: 10.1016/0092-8674(92)90452-i. [DOI] [PubMed] [Google Scholar]
- Premkumar A., Wilson L., Ewart G.D., Gage P.W. Cation-selective ion channels formed by p7 of hepatitis C virus are blocked by hexamethylene amiloride. FEBS Lett. 2004;557:99–103. doi: 10.1016/s0014-5793(03)01453-4. [DOI] [PubMed] [Google Scholar]
- Sakaguchi T., Leser G.P., Lamb R.A. The ion channel activity of the influenza virus M2 protein affects transport through the Golgi apparatus. J. Cell Biol. 1996;133:733–747. doi: 10.1083/jcb.133.4.733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schubert U., Bour S., Ferrer-Montiel A.V., Montal M., Maldarell F., Strebel K. The two biological activities of human immunodeficiency virus type 1 Vpu protein involve two separable structural domains. J. Virol. 1996;70:809–819. doi: 10.1128/jvi.70.2.809-819.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schubert U., Ferrer-Montiel A.V., Oblatt-Montal M., Henklein P., Strebel K., Montal M. Identification of an ion channel activity of the Vpu transmembrane domain and its involvement in the regulation of virus release from HIV-1-infected cells. FEBS Lett. 1996;398:12–18. doi: 10.1016/s0014-5793(96)01146-5. [DOI] [PubMed] [Google Scholar]
- Shen X., Xue J.H., Yu C.Y., Luo H.B., Qin L., Yu X.J., Chen J., Chen L.L., Xiong B., Yue L.D., Cai J.H., Shen J.H., Luo X.M., Chen K.X., Shi T.L., Li Y.X., Hu G.X., Jiang H.L. Small envelope protein E of SARS: cloning, expression, purification, CD determination, and bioinformatics analysis. Acta Pharmacol. Sin. 2003;24:505–511. [PubMed] [Google Scholar]
- Siddell S. The coronaviridae an introduction. In: Siddell S.G., editor. The Coronaviridae. Plenum; New York, NY: 1995. pp. 1–10. [Google Scholar]
- Siddell S. The small-membrane protein. In: Siddell S.G., editor. The Coronaviridae. Plenum; New York, NY: 1995. pp. 181–189. [Google Scholar]
- Sunstrom N.A., Premkumar L.S., Premkumar A., Ewart G., Cox G.B., Gage P.W. Ion channels formed by NB, an influenza B virus protein. J. Membr. Biol. 1996;150:127–132. doi: 10.1007/s002329900037. [DOI] [PubMed] [Google Scholar]
- Tooze J., Tooze S., Warren G. Replication of coronavirus MHV-A59 in sac-cells: determination of the first site of budding of progeny virions. Eur. J. Cell Biol. 1984;33:281–293. [PubMed] [Google Scholar]
- Tosteson M.T., Pinto L.H., Holsinger L.J., Lamb R.A. Reconstitution of the influenza virus M2 ion channel in lipid bilayers. J. Membr. Biol. 1994;142:117–126. doi: 10.1007/BF00233389. [DOI] [PubMed] [Google Scholar]
- Vennema H., Godeke G., Rossen J., Voorhout W., Horzinek M., Opstelten D., Rottier P. Nucleocapsid-independent assembly of coronavirus-like particles by co-expression of viral envelope protein genes. EMBO J. 1996;15:2020–2028. doi: 10.1002/j.1460-2075.1996.tb00553.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- World Health Organization World Health Organization (WHO) Summary table of SARS cases by country, 1 November 2002–7 August 2003. Wkly. Epidemiol. Rec. 2003;78:310–311. [Google Scholar]
- Yao X.J., Gottlinger H., Haseltine W.A., Cohen E.A. Envelope glycoprotein and CD4 independence of vpu-facilitated human immunodeficiency virus type 1 capsid export. J. Virol. 1992;66:5119–5126. doi: 10.1128/jvi.66.8.5119-5126.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]


