Graphical abstract
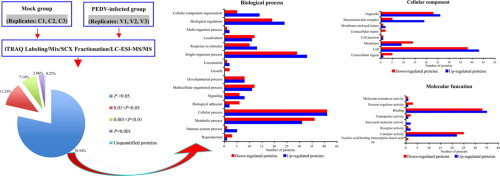
Keywords: PEDV, Infection-associated proteins, Quantitative proteomics
Highlights
-
•
Protein profile of PEDV-infected Vero cells was analyzed by quantitative proteomics.
-
•
126 differentially expressed proteins were identified in PEDV-infected Vero cells.
-
•
The identified proteins provide valuable insight into PEDV-Vero cell interactions.
Abstract
Recent outbreaks of porcine epidemic diarrhea virus (PEDV) have caused widespread concern. The identification of proteins associated with PEDV infection might provide insight into PEDV pathogenesis and facilitate the development of novel antiviral strategies. We analyzed the differential protein profile of PEDV-infected Vero E6 cells using mass spectrometry and an isobaric tag for relative and absolute quantification. A total of 126 proteins were identified that were differentially expressed between the PEDV-infected and mock-infected groups (P < 0.05, quantitative ratio ≥1.2), among which the expression of 58 proteins was up-regulated and that of 68 proteins was down-regulated in the PEDV-infected Vero E6 cells, involving in integrin β2/β3, cystatin-C. The Gene Ontology analysis indicated that the molecular function of the differentially expressed proteins (DEPs) was primarily related to binding and catalytic activity, and that the biological functions in which the DEPs are involved included metabolism, organismal systems, cellular processes, genetic information processing, environmental information processing, and diseases. Among the disease-related functions, certain anti-viral pathways and proteins, such as the RIG-I-like receptor, Rap1, autophagy, mitogen-activated protein kinase, PI3K-Akt and Jak-STAT signaling pathways, and integrin β2/β3 and cystatin-C proteins, represented potential factors in PEDV infection. Our findings provide valuable insight into PEDV-Vero E6 cell interactions.
1. Introduction
The porcine epidemic diarrhea virus (PEDV) is an enveloped, single-stranded positive-sense RNA virus that causes porcine epidemic diarrhea (PED), an acute and highly contagious enteric disease in pigs. PED is characterized by severe diarrhea, vomiting, dehydration, and a mortality rate of up to 90% in suckling piglets (Pensaert and Debouck, 1978). PED was first reported in Belgium and the United Kingdom in 1978, and frequent outbreaks have occurred in various Asian countries (Chen et al., 2010). Since 2007, acute PED outbreaks have continually occurred in Thailand, China, and the USA, which have resulted in substantial economic losses (Puranaveja et al., 2009, Li et al., 2012, Chen et al., 2013, Huang et al., 2013, Marthaler et al., 2013, Stevenson et al., 2013, Yang et al., 2013, Chen et al., 2014). The continued outbreaks of PED, despite control efforts, have caused widespread concern.
The PEDV belongs to the genus Alphacoronavirus, in the family Coronaviridae and order Nidovirales (Belouzard et al., 2012). Previous studies have investigated various control measures to protect against PEDV infection, such as vaccines, diagnostic tools, and therapeutic drugs (Sun et al., 2008, Ren et al., 2011, Sun et al., 2012, Zhu et al., 2013, Guo et al., 2013, Kim and Lee, 2013). Various aspects of PEDV infection remain unclear, for example, swine testis (ST) cells expressing porcine aminopeptidase N of PEDV receptor were not susceptible to PEDV infection. African green monkey kidney (Vero) cells are highly susceptible to PEDV infection, and are widely used for the primary isolation and cultivation of PEDV (Pan et al., 2012, Guo et al., 2014). Therefore, Vero lineages are suitable hosts for understanding the mechanisms of PEDV infection.
Proteomics techniques are effective tools for characterizing protein expression profiles, and have been used widely to investigate disease-associated proteins (Hondermarck et al., 2008, Boja et al., 2011, He et al., 2012, Sun et al., 2013). Among current proteomics methods, quantitative high-throughput proteomics approaches are useful for the analysis of infection-associated proteins of pathogens (Linde et al., 2013, Papachristou et al., 2013, Ye et al., 2013, Zeng et al., 2015). In our current study, we used a quantitative proteomics approach based on an iTRAQ tandem mass spectrometry (MS/MS) technique to identify proteins differentially expressed between PEDV-infected and mock-infected Vero E6 cells. The functions of the differentially expressed proteins (DEPs) were analyzed to determine whether they might be associated with PEDV infection. Our findings provide valuable insight into the changes in cellular processes that occur during PEDV infection.
2. Materials and methods
2.1. Virus, cells, and antibody
The CV777 strain of PEDV, kindly provided by Maurice Pensaert at Ghent University (Merelbeke, Belgium), was used in all of our experiments after being adapted to Vero E6 cells, as previously described (Hofmann and Wyler, 1988). The Vero E6 cell-adapted PEDV, the Vero E6 cells, and the monoclonal antibody against the nucleocapsid protein (Np) of PEDV were stored at the Diarrhea-Related Viruses Section, Division of Swine Infectious Diseases, National Key Laboratory of Veterinary Biotechnology, Harbin Veterinary Research Institute of the Chinese Academy of Agricultural Sciences.
2.2. Viral infection of Vero E6 cells
The Vero E6 cells were cultured in Dulbecco's modified Eagle's medium containing 10% fetal bovine serum (FBS) in 75-cm flasks at 37 °C in a 5% CO2 atmosphere. When the cells reached 70–80% confluence, they were inoculated with the PEDV at a multiplicity of infection of 1 in presence of 5 μg/mL trypsin. At 48 h postinoculation, the cells began to exhibit cytopathic effects (CPEs) of viral infection, but no cells lysis or shedding had occurred. The cells were washed three times with cold phosphate-buffered saline (PBS, pH 7.4). A 1.5-mL aliquot of lysis buffer containing 4% SDS, 1 mM DTT, and 150 mM Tris–HCl (pH 8.0) was added to each flask, and the flasks were incubated at 37 °C for 5 min. The cell lysates were collected using a cell scraper, and boiled for 5 min. Three cell lysate replicates were prepared for the PEDV-infected (V1–V3) and mock-infected (C1–C3) Vero E6 cells, and stored at −80 °C.
2.3. Immunoblotting
Western blotting was performed to confirm PEDV infection by detecting the presence of the Np of PEDV in the Vero E6 cells. Aliquots of the cell lysates were subjected to SDS-PAGE on a 12% acrylamide gel, and the protein bands were transferred to a nitrocellulose membrane using a semi-dry transfer device (Bio-Rad, Hercules, CA, USA). The membrane was blocked using 5% (w/v) nonfat dried milk in PBS at 37 °C for 1 h, before incubation in PBS containing the anti-Np monoclonal antibody (1:2000 dilution) at 37 °C for 1 h. After washing three times with 5% Tween 20 in PBS (PBST), the membrane was incubated in PBST containing a horseradish peroxidase-conjugated goat anti-mouse IgG (1:4000 dilution) at 37 °C for 1 h. After washing three times with PBS, the membrane was incubated with enhanced chemiluminescence detection reagents (Biotopped, Beijing, China) at room temperature for 3 min, and the peroxidase-mediated luminescence was digitally captured using the Molecular Imager ChemiDoc XRS+ System (Bio-Rad) and the Image Lab software (Bio-Rad). To verify the differential expression of the selected DEPs, equivalent volumes of the cell lysate replicates from the PEDV-infected (V1–V3) and mock-infected (C1–C3) Vero E6 cells were pooled into the V and C samples, respectively, and western blotting was performed as described above, with the following exceptions: a 1:1000 dilution of the polyclonal antibodies anti-β tubulin, anti-integrin-β3, anti-cystatin-C, anti-protein S100-A2, anti-apolipoprotein E4, and anti-centrin from rabbit (Beijing Biosynthesis Biotechnology, Beijing, China) was used as the primary antibody, and a 1:5000 dilution of the HRP-conjugated goat anti-rabbit IgG (Sigma–Aldrich, St. Louis, USA) was used as the secondary antibody.
2.4. Protein digestion and iTRAQ labeling
Protein digestion of the samples was performed according to the FASP procedure described by Wiśniewski et al. (2009). An aliquot of each cell lysate containing 200 μg of protein was combined with 30 μL of STD buffer containing 4% SDS, 100 mM DTT, and 150 mM Tris–HCl (pH 8.0). The detergent, DTT, and other low-molecular-weight components were removed by dilution in UA buffer containing 8 M Urea and 150 mM Tris–HCl (pH 8.0) and repeated ultrafiltration using Microcon (30 kDa) ultrafiltration units. The reduction of cysteine residues was blocked by the addition of 100 μL of 0.05 M iodoacetamide to the UA buffer. The samples were incubated for 20 min in darkness before ultrafiltration. The Microcon filters were washed three times with 100 μL of UA buffer, followed by two washes with 100 μL DS buffer containing 50 mM triethyl ammonium bicarbonate (pH 8.5). The final protein suspensions were digested using 2 μg of trypsin (Promega, Madison, WI, USA) in 40 μL of DS buffer overnight at 37 °C, and the digested peptides were collected as the filtrate. The peptide content was quantified based on absorbance at 280 nm using an extinction coefficient of 1.1 for a 0.1 mg/mL solution. The digested peptide mixture was labeled using the 8-plex iTRAQ reagent (Life Technologies, Carlsbad, CA, USA), according to the manufacturer's instructions. Each iTRAQ reagent was dissolved in 70 μL of ethanol, and added to the digested peptide mixture. The samples were labeled as C1-113, C2-114, C3-115, V1-116, V2-117, or V3-118. The samples were multiplexed, and vacuum dried.
2.5. Peptide fractionation with strong cation exchange chromatography (SCXC)
The iTRAQ labeled peptides were fractionated by SCXC using the AKTA Purifier system (GE Healthcare, Waukesha, WI, USA). The dried peptide mixture was reconstituted, and acidified by the addition of 2 mL of buffer A containing 10 mM KH2PO4 in 25% acetonitrile (pH 2.7). The samples were loaded onto a 4.6 mm × 100 mm column packed with Polysulfoethyl (5 μm, 200 Å) chromatography resin (PolyLC, Columbia, Maryland, USA). The peptides were eluted at a flow rate of 1 mL/min using a gradient of 0–10% buffer B containing 500 mM KCl and 10 mM KH2PO4 in 25% acetonitrile (pH 2.7). The gradient elution consisted of 10–20% buffer B for 25 min, 20–45% buffer B for 5 min, and 50–100% buffer B for 5 min. The absorbance of the eluate was monitored at 214 nm, and fractions were collected at 1-min intervals. Thirty fractions were combined into ten pools, and desalted using Empore standard density SPE C18 cartridges (Sigma–Aldrich, St. Louis, MO, USA) with a bed diameter of 7 mm and a volume 3 mL. Each fraction was concentrated by centrifugation in a vacuum, and reconstituted in 40 μL of 0.1% (v/v) trifluoroacetic acid. All samples were stored at −80 °C until the MS analysis was performed.
2.6. Liquid chromatography (LC) MS/MS analysis
The LC–MS/MS experiments were performed using a Q Exactive mass spectrometer coupled to a Proxeon Biosystem Easy nanoLC (Thermo Fisher Scientific, Waltham, MA, USA). Ten microliters of each fraction was injected for nanoLC–MS/MS analysis. The peptide mixture (5 μg) was loaded onto a C18-reversed phase column (15 cm × 75 μm) packed with RP-C18 (5 μm) resin in buffer A containing 0.1% formic acid, and eluted with a linear gradient of buffer B (80% acetonitrile and 0.1% formic acid) at a flow rate of 0.25 μL/min for 140 min using the IntelliFlow technology. The eluate underwent electrospray ionization for the MS/MS analysis. The MS/MS instrument was run in the peptide recognition mode, and the spectra were acquired using a data-dependent top-10 method based on the selection of the most abundant precursor ions from the survey scan (300–1800 m/z) for HCD fragmentation. The determination of the target value was based on the predictive automatic gain control, and the dynamic exclusion duration was 60 s. Survey scans were acquired at a resolution of 70, 000 at m/z 200, and the resolution for the HCD spectra was set to 17, 500 at m/z 200. The normalized collision energy was 30 eV, and the underfill ratio, which specifies the minimum percentage of the target value likely to be reached at maximum fill time, was defined as 0.1%.
2.7. Identification and analysis of proteins
The MS/MS spectra were compared to the Uniprot Cercopithecidae database (107 051 sequences, downloaded November 25, 2013) and a decoy database using the MASCOT search engine, version 2.2 (Matrix Science, London, UK), embedded in the Proteome Discoverer 1.4 software (Thermo Electron, San Jose, CA). The following parameters were used for protein identification: a peptide mass tolerance of 20 ppm; an MS/MS tolerance of 0.1 Da; trypsin digestion; a missed cleavage value of 2; the fixed modifications included carbamidomethyl, iTRAQ8plex(K), and iTRAQ8plex(N-term); the variable modification was oxidation; and an FDR value ≤0.01. Protein quantification was performed using the Proteome Discoverer 1.4 software based on the centroided reporter ion peak intensity. The average quantitative value of each protein in samples C1, C2, and C3 (mock-infection group) was used as the internal reference. The value of the quantitative ratio for each protein relative to the internal reference was calculated, and averaged to obtain the quantitative ratio (V/C) of the proteins identified in the treatment groups (Unwin et al., 2010). A protein was considered to be differentially expressed between the PEDV-infected and mock-infected groups based on the following criteria: the protein had to be present in three replicates of both groups, the difference in the level of the protein between the two groups had to be statistically significant (P < 0.05), and the change ratio for the protein had to be ≥1.2 (Yuan et al., 2012). The expression of a protein with a V/C > 1.0 was considered to be up-regulated, and those with a V/C < 1.0 were considered to be down-regulated. The data were analyzed using a two-tailed, paired Student's t test. The statistical analysis was performed using the Excel 2007 software (Microsoft, Redmond, WA, USA). The DEPs were annotated using the Blast2GO, version 2.7.0, program (Ashburner et al., 2000, Quevillon et al., 2005, Götz et al., 2008). The DEPs were blasted against the KEGG Genes database (human). The Gene Ontology categories (GOCs) were retrieved, and mapped to pathways in the KEGG database (Kanehisa et al., 2012).
3. Results
3.1. Identification and analysis of proteins
The Vero E6 cells inoculated with PEDV displayed distinct CPEs at 48 h postinoculation, including cell shrinkage, cell fusion, and a rounded cell morphology, but no cells lysis or shedding was observed (Fig. 1A). The immunoblotting analysis confirmed that the Vero E6 cells were PEDV-infected. The band corresponding to the Np of PEDV was detected in samples V1, V2, and V3, whereas none was detected in samples C1, C2, and C3 (Fig. 1B). The identified peptides, identified proteins, quantified proteins, known/uncharacterized proteins, and the GOC annotations are showed in Table 1 . A total of 3178 proteins, including 15 564 peptides, were identified in the PEDV-infected and mock-infected groups using the iTRAQ-MS/MS approach, among which 3171 (99.78%) were quantified, 1859 (58.50%) were known proteins, and 1319 (41.50%) were uncharacterized/putative proteins. Based on the GOCs, 2061 (64.85%) of the proteins were annotated as biological process, 2495 (78.51%) were annotated as molecular function, and 1917 (60.32%) were annotated as cellular components.
Fig. 1.
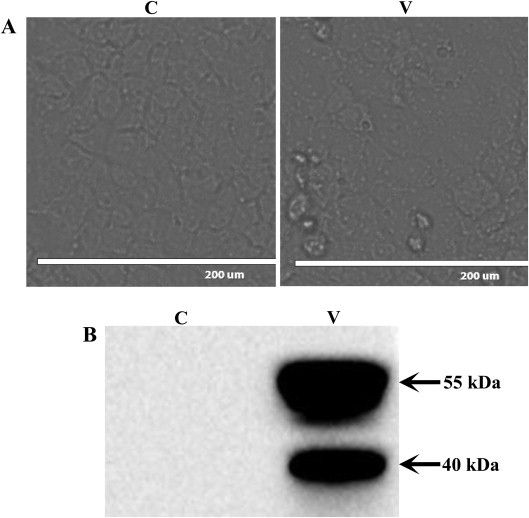
Preparation and identification of PEDV-infected Vero E6 cells. (A) Photomicrographs of PEDV-infected Vero E6 cells or mock-infected Vero E6 cells: C, mock-infected group; V, PEDV-infected group. (B) Verification of PEDV propagation in Vero E6 cells by western blot: C, mock-infected group; V, PEDV-infected group.
Table 1.
The proteins identified from PEDV-infected and mock-infected groups.
| Classification | Number of proteins (percentage) |
|---|---|
| Identified peptides | 15,564 |
| Identified proteins | 3178 (100%) |
| Quantified proteins | 3171 (99.78%) |
| Known proteins | 1859 (58.50%) |
| Uncharacterized proteins | 1319 (41.50%) |
| Annotated proteins in GO categories of biological process | 2061 (64.85%) |
| Annotated proteins in GO categories of molecular function | 2495 (78.51%) |
| Annotated proteins in GO categories of cellular component | 1917 (60.32%) |
The quantification and significance of the identified proteins are shown in Fig. 2 . The changes in the levels of expression between the two groups were analyzed based on statistical significance. Of the 3178 proteins identified, 2496 (78.54%) were not differentially expressed (P > 0.05), and 675 (21.24%) were expressed at statistically different levels between the PEDV-infected and mock-infected Vero E6 cells (P < 0.05), including 357 proteins (11.23%) with a P-value between 0.01 and 0.05, 227 proteins (7.14%) with a P-value between 0.001 and 0.01, and 91 proteins (2.86%) with a P-value <0.001. The proteins with a P-value <0.05 were also filtered based on whether the V/C or C/V was ≥1.2. Based on these criteria, a total of 126 (3.96%) of the 3178 identified proteins were determined to have been differentially expressed between the PEDV-infected and mock-infected groups (Table 2 ). Among the 126 DEPs, 46.03% (58/126) were up-regulated, and 53.97% (68/126) down-regulated. The known proteins and uncharacterized/putative proteins accounted for 69.05% (87/126) and 30.95% (39/126) of the DEPs, respectively. The DEP displaying the greatest increase in expression in the PEDV-infected Vero E6 cells was isoform 2 of the ovarian cancer immunoreactive antigen domain-containing 1 protein (1:2.5), and the DEP displaying the greatest decrease in expression in the PEDV-infected Vero E6 cells was cystatin-C (1:2.2).
Fig. 2.
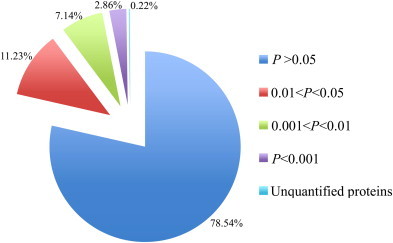
The quantitation and significance of the 3178 identified proteins from PEDV-infected and mock-infected groups.
Table 2.
The differentially expressed protein lists between PEDV-infected and mock-infected groups.
| No. | Protein name | UniProtKB accession no. | GO annotation |
P value | Average V/Cb | ||
|---|---|---|---|---|---|---|---|
| Molecular function | Cellular component | Biological process | |||||
| 1 | Cystatin-C or Cystatin-3 | G7PH52 | enzyme regulator activity | – | metabolic process; regulation of biological process | 6.98E−04 | 0.46 |
| 2a | Osteopontin precursor | F7F5L5 | – | – | – | 9.11E−03 | 0.50 |
| 3 | Retinol dehydrogenase 10 | G7MZK0 | nucleotide binding; catalytic activity | cytoplasm; membrane | development; metabolic process; reproduction; cell differentiation | 5.22E−03 | 0.59 |
| 4 | Testis cDNA clone | Q4R3Z6 | – | – | – | 1.36E−03 | 0.59 |
| 5 | Overexpressed in colon carcinoma 1 protein | H9FAZ7 | – | – | – | 6.48E−04 | 0.61 |
| 6 | Cytochrome b-245 light chain | H9F3U1 | – | membrane | – | 1.11E−02 | 0.62 |
| 7a | Centrin-2 | F7HKU5 | metal ion binding; nucleotide binding; catalytic activity | – | – | 1.26E−03 | 0.66 |
| 8a | Kidney-specific cadherin | F6VCT3 | metal ion binding | membrane | – | 6.70E−05 | 0.67 |
| 9a | Putative WD repeat-containing protein 33 | F6SBK9 | protein binding | nucleus | – | 1.36E−03 | 0.67 |
| 10 | Apolipoprotein E4 | D5G333 | – | extracellular | transport; metabolic process | 9.85E−03 | 0.68 |
| 11 | Receptor-type tyrosine-protein phosphatase T isoform 1 | H9F9X9 | catalytic activity | membrane | metabolic process | 3.56E−03 | 0.68 |
| 12a | Putative neutral and basic amino acid transport protein rBAT-like isoform 3 | F7HIT7 | catalytic activity | – | metabolic process | 8.18E−05 | 0.69 |
| 13a | Mitochondrial ornithine aminotransferase | F7BGF3 | catalytic activity | cytoplasm; mitochondrion | metabolic process; cell organization and biogenesis | 3.51E−02 | 0.69 |
| 14 | Lysosomal protective protein | G7N4N3 | catalytic activity | nucleus | metabolic process | 2.85E-05 | 0.70 |
| 15a | Putative low-density lipoprotein receptor-related protein 2 | F7H113 | metal ion binding; protein binding | cytoplasm; endosome; endoplasmic reticulum; Golgi; membrane | metabolic process; transport; cell proliferation; development | 1.02E−04 | 0.70 |
| 16 | Alpha-adducin isoform c | H9FPQ1 | metal ion binding | cytoskeleton | – | 3.58E−03 | 0.71 |
| 17 | Integrin beta 2 | H9Z8N5 | receptor activity; protein binding | membrane | cell communication; regulation of biological process; response to stimulus; development | 1.82E−04 | 0.71 |
| 18 | Dipeptidyl peptidase 2 preproprotein | H9EXB4 | catalytic activity | membrane | metabolic process | 4.22E−03 | 0.71 |
| 19a | Estrogen sulfotransferase | F6RUQ2 | catalytic activity | nucleus; cytoplasm; membrane | metabolic process | 4.73E−03 | 0.71 |
| 20a | Putative protein EGK_14077 | F6PJM4 | – | – | – | 2.34E−03 | 0.72 |
| 21a | Putative legumain | F6S082 | catalytic activity | cytoplasm; endosome | response to stimulus; metabolic process; regulation of biological process; cell death | 2.53E−03 | 0.73 |
| 22 | Metallothionein-1E | F6PYY1 | metal ion binding | nucleus; cytoplasm | regulation of biological process; response to stimulus | 4.86E−03 | 0.74 |
| 23 | Trophoblast glycoprotein | H9F4Q1 | protein binding | membrane | – | 6.68E−04 | 0.74 |
| 24a | Putative proactivator polypeptide isoform X6 | F7F376 | – | cytoplasm; vacuole | metabolic process | 6.40E−03 | 0.74 |
| 25 | Cathepsin D (Predicted) | A9L947 | catalytic activity | – | metabolic process | 1.33E−04 | 0.75 |
| 26 | Aldehyde dehydrogenase family 1 member L1 | F7HB04 | catalytic activity | cytoplasm; mitochondrion | metabolic process | 9.07E−03 | 0.75 |
| 27 | Carbonic anhydrase 2 | G7MZP3 | catalytic activity; metal ion binding | – | metabolic process | 1.83E−03 | 0.75 |
| 28 | Erythrocyte band 7 integral membrane protein isoform a | F7HP19 | – | cytoskeleton; membrane; extracellular | cell organization and biogenesis | 4.40E−05 | 0.75 |
| 29a | Putative laminin subunit beta-1 | F7HPY4 | catalytic activity; motor activity; signal transducer activity; protein binding; structural molecule activity; nucleotide binding | extracellular; nucleus; cytoplasm; membrane; cytoskeleton; mitochondrion | metabolic process; regulation of biological process; cell communication; response to stimulus; transport; cellular component movement; development; cell differentiation; cell organization and biogenesis | 1.92E−03 | 0.76 |
| 30a | Putative tissue alpha-L-fucosidase | F7HDC0 | catalytic activity | – | metabolic process | 6.56E−03 | 0.76 |
| 31 | Similar to human bone marrow stromal cell antigen 1 | I7GP78 | catalytic activity | – | – | 2.77E−02 | 0.77 |
| 32 | Solute carrier family 17, member 5 | A9X190 | – | membrane | transport | 8.00E−03 | 0.77 |
| 33 | Galectin | G7N3S9 | – | – | – | 2.88E−03 | 0.78 |
| 34 | Sulfhydryl oxidase 2 | H9F332 | catalytic activity | membrane | metabolic process | 4.97E−03 | 0.78 |
| 35 | Folate receptor alpha | F7BP60 | – | – | – | 2.98E−03 | 0.78 |
| 36a | Putative protein EGK_10171 | G7NLB0 | – | – | – | 4.60E−03 | 0.79 |
| 37 | Lysosome-associated membrane glycoprotein 2 isoform B | F7BCK9 | – | membrane | – | 1.61E−03 | 0.79 |
| 38 | RNA-binding motif protein 12B | G7MZR8 | nucleotide binding | membrane | – | 2.10E−03 | 0.79 |
| 39 | Similar to human synaptobrevin-like 1 (SYBL1) | I7G8H0 | – | membrane | transport | 8.83E−03 | 0.79 |
| 40a | Putative polyadenylate-binding protein-interacting protein 1 isoform 2 | F7H0R0 | DNA binding; RNA binding; protein binding | – | metabolic process | 1.17E−04 | 0.79 |
| 41a | Putative versican core protein-like isoform 8 | F7C5T5 | metal ion binding; protein binding | extracellular | cell differentiation; cellular component movement; development | 5.67E−05 | 0.79 |
| 42a | Putative N-acetylglucosamine-6-sulfatase | G7PIY5 | catalytic activity | cytoplasm; vacuole | metabolic process | 1.14E−04 | 0.80 |
| 43 | ACADSB | Q5IFK6 | catalytic activity; nucleotide binding | – | metabolic process | 3.47E−02 | 0.80 |
| 44 | Clusterin | Q5ISQ2 | – | cell death | 2.54E−03 | 0.80 | |
| 45 | Cathepsin Z | G7N4B3 | catalytic activity | – | metabolic process | 6.96E−03 | 0.80 |
| 46a | Endoplasmic reticulum resident protein 28 | F7GNV5 | – | cytoplasm; endoplasmic reticulum; organelle lumen | transport | 4.49E−02 | 0.80 |
| 47a | Putative transducin-like enhancer protein 4 isoform 12 | F6PZC8 | protein binding | nucleus | metabolic process; regulation of biological process | 2.22E−03 | 0.81 |
| 48a | T-cell immunoglobulin and mucin domain-containing protein 1 | F7GB98 | protein binding | – | – | 4.71E−05 | 0.81 |
| 49a | Putative N-acetylgalactosamine-6-sulfatase | G7NQ90 | catalytic activity | – | metabolic process | 1.58E−02 | 0.81 |
| 50a | Uncharacterized protein | F7GQ07 | catalytic activity | – | metabolic process | 3.01E−04 | 0.81 |
| 51a | Putative galactokinase | F7DF76 | nucleotide binding; catalytic activity | cytoplasm | metabolic process | 4.72E−02 | 0.81 |
| 52 | Signal transducer and activator of transcription | Q9N145 | protein binding; DNA binding; signal transducer activity | nucleus; cytoplasm; membrane | cell differentiation; development; metabolic process; regulation of biological process; transport; cell communication; response to stimulus; cell proliferation; reproduction | 7.92E−03 | 0.81 |
| 53 | Delta(14)-sterol reductase | G7PPJ4 | catalytic activity | membrane | metabolic process | 3.51E−02 | 0.81 |
| 54a | Putative disabled homolog 2 isoform X4 | F7GRX9 | protein binding | – | – | 4.59E−03 | 0.81 |
| 55 | Pyridoxal-dependent decarboxylase domain-containing protein 1 | F7GW28 | catalytic activity | – | metabolic process | 1.59E−02 | 0.81 |
| 56 | Rho GTPase-activating protein 29 | H9FI14 | metal ion binding | membrane | cell communication; regulation of biological process; response to stimulus | 1.51E−02 | 0.82 |
| 57 | Protein DB83 | F7H2H1 | – | membrane; cytoplasm | – | 2.25E−03 | 0.82 |
| 58 | DORA reverse strand protein | G7Q0P8 | catalytic activity | – | metabolic process | 1.01E−02 | 0.82 |
| 59 | Bifunctional ATP-dependent dihydroxyacetone kinase/FAD-AMP lyase (Cyclizing) | G7PPY4 | catalytic activity; nucleotide binding | membrane | metabolic process | 4.96E−03 | 0.82 |
| 60 | Cadherin 6 | Q5ISM2 | metal ion binding | membrane | – | 6.61E−03 | 0.82 |
| 61a | Putative mitochondrial delta(3,5)-Delta(2,4)-dienoyl-CoA isomerase | F6TNT1 | catalytic activity; protein binding | cytoplasm; mitochondrion | metabolic process | 5.83E−03 | 0.82 |
| 62 | Fatty acid-binding protein, heart | G7MI71 | transporter activity | – | transport | 2.36E−03 | 0.82 |
| 63a | Putative NADH dehydrogenase 1 α subcomplex subunit 8 | F7B3T9 | – | cytoplasm; mitochondrion; membrane | – | 1.78E−03 | 0.83 |
| 64 | Endoplasmic reticulum resident protein 58 | G7NJK7 | protein binding | cytoplasm; endoplasmic reticulum; organelle lumen | – | 5.49E−03 | 0.83 |
| 65 | Agrin | H9FU64 | transporter activity; protein binding | extracellular; membrane | transport; cell communication; regulation of biological process; response to stimulus; cell organization and biogenesis | 9.20E−03 | 0.83 |
| 66a | Toll-interacting protein | F7DRQ6 | protein binding | – | – | 2.53E−02 | 0.83 |
| 67a | Putative tetraspanin-3 isoform X2 | F6SRI3 | – | membrane | – | 3.37E−03 | 0.83 |
| 68a | Putative protein EGK_14027 | G7MM35 | enzyme regulator activity; protein binding | cytoplasm; cytoskeleton; membrane | metabolic process; regulation of biological process | 6.05E−03 | 0.83 |
| 69 | Inosine-5′-monophosphate dehydrogenase | F6VXT4 | nucleotide binding; catalytic activity; protein binding; metal ion binding | nucleus; cytoplasm; membrane | metabolic process; cell proliferation; response to stimulus | 3.12E−04 | 1.20 |
| 70 | 78 kDa glucose-regulated protein | F7C3R1 | nucleotide binding; catalytic activity; protein binding | nucleus; cytoplasm; endoplasmic reticulum; cell surface; membrane | cell organization and biogenesis; cell communication; regulation of biological process; response to stimulus; metabolic process; development; cell death | 9.49E−05 | 1.20 |
| 71a | Uncharacterized protein | F6W6U2 | – | – | – | 1.40E−02 | 1.20 |
| 72a | Follistatin-related protein 1 | G7MKF2 | metal ion binding; protein binding | – | – | 3.29E−04 | 1.20 |
| 73a | Putative retrotransposon-like protein 1 | F7G2J3 | RNA binding; catalytic activity | – | metabolic process | 6.78E−04 | 1.20 |
| 74a | Heat shock protein 105 kDa | F6S529 | nucleotide binding | – | – | 6.60E−05 | 1.21 |
| 75a | Putative protein midA homolog | F7GTQ7 | – | – | – | 2.96E−03 | 1.21 |
| 76 | Similar to human S-adenosylhomocysteine hydrolase-like 1 | I7GBN2 | catalytic activity | – | metabolic process; transport | 9.34E−03 | 1.21 |
| 77 | Transferrin receptor 1 | F6UX47 | catalytic activity | extracellular; cytoplasm; endosome; membrane; cell surface | metabolic process; cellular homeostasis; cell differentiation; development; regulation of biological process | 4.64E−04 | 1.22 |
| 78a | Exocyst complex component 6B | F7GZR4 | – | cytoplasm | transport | 5.75E−03 | 1.22 |
| 79 | Protein S100-A2 | H9F670 | metal ion binding | membrane | – | 4.75E−02 | 1.22 |
| 80a | Ephrin type-A receptor 2 | F7E018 | nucleotide binding; catalytic activity; receptor activity; signal transducer activity; protein binding | membrane | development; cell differentiation; metabolic process; cell communication; regulation of biological process; response to stimulus; cell death; cell organization and biogenesis; cell proliferation; cellular component movement | 3.02E−02 | 1.22 |
| 81a | Putative nucleolar RNA helicase 2-like isoform 3 | F6SQP8 | nucleotide binding; RNA binding; catalytic activity | nucleus | – | 1.12E−03 | 1.23 |
| 82a | Putative polyadenylate-binding protein 1-like isoform X2 | F7EU27 | nucleotide binding; RNA binding | – | – | 1.77E−03 | 1.23 |
| 83a | Uncharacterized protein | F7CEU8 | nucleotide binding | – | – | 5.52E−03 | 1.23 |
| 84 | Sequestosome-1 isoform 1 | I2CY26 | metal ion binding | – | – | 8.51E−03 | 1.23 |
| 85 | NEDD8 ultimate buster 1 isoform 2 | H9EZG1 | protein binding | – | metabolic process; regulation of biological process; defense response; response to stimulus | 2.32E−02 | 1.23 |
| 86 | Thioredoxin domain-containing protein 9 | F6YEB0 | – | nucleus; cytoplasm; cytoskeleton | cellular homeostasis; regulation of biological process | 3.79E−02 | 1.23 |
| 87 | Four and a half LIM domains protein 2 | F7GXH4 | protein binding; metal ion binding | nucleus; cytoskeleton | metabolic process; regulation of biological process; cell differentiation; response to stimulus; cell death; development | 2.24E−05 | 1.23 |
| 88 | Similar to human DKFZP564M182 protein | I7G2J1 | RNA binding; structural molecule activity | cytoplasm; ribosome | metabolic process | 2.95E−03 | 1.23 |
| 89 | Similar to human hypothetical protein FLJ10634 | I7GMW8 | nucleotide binding | – | – | 1.69E−02 | 1.23 |
| 90a | Putative hexokinase-2 | F6Y855 | nucleotide binding; catalytic activity | cytoplasm; membrane; mitochondrion | metabolic process | 3.26E−04 | 1.24 |
| 91 | Ribosomal protein L37 | F7FYI2 | RNA binding; structural molecule activity; metal ion binding | cytoplasm; ribosome; cytosol | metabolic process | 3.51E−03 | 1.24 |
| 92 | Pyruvate kinase | F7FI39 | metal ion binding; catalytic activity | – | metabolic process | 1.07E−02 | 1.24 |
| 93 | Protein phosphatase 1B isoform 2 | H9EM08 | metal ion binding; catalytic activity | – | metabolic process | 1.40E−02 | 1.24 |
| 94a | Glycogen synthase kinase-3 alpha | G7PXQ8 | nucleotide binding; catalytic activity | membrane | metabolic process | 4.34E−03 | 1.24 |
| 95 | Phosphoserine aminotransferase | F7H2J8 | catalytic activity | – | metabolic process | 1.01E−03 | 1.25 |
| 96 | Histone H2A | H9FCA2 | DNA binding; protein binding | chromosome; nucleus; membrane | cell organization and biogenesis; metabolic process | 2.02E−03 | 1.25 |
| 97 | EF-hand domain-containing protein D2 | H9FC53 | metal ion binding | – | – | 6.88E−04 | 1.25 |
| 98a | Putative ATP-dependent RNA helicase DDX10 | F7BIN5 | nucleotide binding; catalytic activity | – | – | 1.20E−04 | 1.25 |
| 99a | Putative pterin-4-alpha-carbinolamine dehydratase | F7F694 | catalytic activity; protein binding | nucleus; cytoplasm | metabolic process; cell organization and biogenesis | 1.85E−02 | 1.26 |
| 100 | Integrin beta 3 | F7FG54 | receptor activity; protein binding | membrane; cell surface | cell organization and biogenesis; cell communication; regulation of biological process; response to stimulus; development; cellular component movement; cell proliferation; coagulation | 2.60E−04 | 1.26 |
| 101 | Similar to human exocyst complex component 7 | I7GLB8 | – | cytoplasm | transport | 5.30E−03 | 1.27 |
| 102a | Putative leucine-rich repeat flightless-interacting protein 1 | F6S3D2 | – | – | – | 1.48E−03 | 1.27 |
| 103a | Uncharacterized protein | G7P460 | – | – | – | 8.80E−04 | 1.28 |
| 104 | Eukaryotic initiation factor 4A-I isoform 1 | H9FAB5 | nucleotide binding; RNA binding; catalytic activity | cytoplasm | metabolic process | 2.51E−03 | 1.29 |
| 105a | Phosphatidylinositol transfer protein beta isoform isoform 2 | F7G7C8 | – | – | transport | 1.70E−02 | 1.29 |
| 106 | Phosphatidylinositol-3,4,5-trisphosphate 5-phosphatase 2 | G7NE91 | protein binding | – | metabolic process | 4.14E−03 | 1.30 |
| 107a | Putative tripartite motif-containing protein 47 (TRIM47) | F7HF49 | protein binding; metal ion binding | intracellular | – | 2.65E−02 | 1.31 |
| 108 | UDP-N-acetylhexosamine pyrophosphorylase | F7CXU2 | catalytic activity | – | metabolic process | 3.83E−02 | 1.32 |
| 109a | Putative protein EGK_20713 | F7B9G5 | protein binding | – | – | 3.42E−03 | 1.32 |
| 110a | Putative 60S ribosomal protein L23a-like | F7HD49 | nucleotide binding; structural molecule activity | cytoplasm; ribosome | metabolic process | 3.89E−04 | 1.33 |
| 111 | Mitochondrial import inner membrane translocase subunit Tim13 | F6R7Z6 | metal ion binding | cytoplasm; mitochondrion | cell organization and biogenesis; transport | 1.98E−04 | 1.33 |
| 112 | Poly [ADP-ribose] polymerase 9 isoform c | H9ZE68 | catalytic activity | membrane | – | 3.46E−03 | 1.34 |
| 113a | Serpin B6 | F7BRQ5 | enzyme regulator activity | extracellular | – | 7.69E−04 | 1.35 |
| 114 | EH domain-containing protein 4 | F7HAE5 | catalytic activity; metal ion binding; protein binding; nucleotide binding | cytoplasm; endoplasmic reticulum; endosome; membrane | metabolic process; transport; cell organization and biogenesis | 2.73E−04 | 1.35 |
| 115 | Calponin-like integrin-linked kinase-binding protein | G7PQP7 | protein binding; catalytic activity | membrane | cell communication; regulation of biological process; response to stimulus; metabolic process | 5.57E−04 | 1.35 |
| 116a | Putative protein EGK_00649 | G8F2R2 | transporter activity | membrane | transport | 2.83E−03 | 1.36 |
| 117 | Ribosomal RNA-processing protein 8 | F7C3H4 | catalytic activity; protein binding | nucleus; organelle lumen; cytoplasm; membrane; chromosome | metabolic process; regulation of biological process; cell communication; response to stimulus; cell death | 3.59E−03 | 1.36 |
| 118 | Microtubule-associated protein 1B | G7P7P5 | protein binding; catalytic activity | cytoskeleton; membrane | cell organization and biogenesis | 3.37E−05 | 1.37 |
| 119 | FMRP-interacting protein 2 | G7NGR9 | – | – | – | 3.95E−04 | 1.42 |
| 120a | Putative centrosomal protein of 89 kDa isoform X1 | F7H5V4 | protein binding | – | – | 8.26E−04 | 1.44 |
| 121a | Ubiquitin-conjugating enzyme | F6ULI1 | catalytic activity | – | – | 3.58E−03 | 1.49 |
| 122 | Nucleolar complex protein 14 | G7NVR8 | – | membrane | – | 4.14E−07 | 1.52 |
| 123 | Ubiquitin-like protein ISG15 | F7GS84 | protein binding | extracellular | metabolic process; regulation of biological process; defense response; response to stimulus | 1.59E−05 | 1.76 |
| 124a | Putative caspase-7 isoform X1 | G7PDZ9 | catalytic activity | cytoplasm | metabolic process; cell death | 2.21E−04 | 1.76 |
| 125a | Putative tRNA pseudouridine synthase Pus10 | H9FK14 | – | – | – | 2.56E−04 | 2.21 |
| 126 | OCIA domain-containing protein 1 isoform 2 | H9FWA7 | – | membrane | – | 1.20E−02 | 2.50 |
The names of the uncharacterized/putative proteins were annotated again and corrected by the blastp search in GenBank database of NCBI based on the primary identified protein sequences from the UniProtKB.
The average V/C ratio >1 represents the up-regulated proteins; the average V/C ratio <1 represents the down-regulated proteins.
3.2. GO annotations of the DEPs
The Gene Ontology (GO) database has been widely used for describing protein function in a standardized format. According to their GOCs, the 126 DEPs were annotated as cellular component, biological process, or molecular function. The GO annotations are shown in Table 2, and distributions of the GO annotations are shown in Fig. 3 . Seventy-eight DEPs were distributed among 16 groups of biological processes (Fig. 3A). The metabolic process (GO:0008152), cellular process (GO:0009987), single-organism process (GO:0044699), and biological regulation (GO:0065007) groups contained the highest proportions of the biological process DEPs. There were more up-regulated proteins in the cellular component organization group (GO:0071840) than down-regulated proteins. Seventy-four DEPs were distributed among eight cellular component groups (Fig. 3B), among which the organelle (GO:0043226) and cell (GO:0005623) groups contained the highest proportion of cellular component DEPs. There were more down-regulated DEPs in the membrane group (GO:0016020) than up-regulated DEPs, and there were more up-regulated DEPs in the macromolecular complex group (GO:0032991) than down-regulated DEPs. Ninety-seven DEPs were distributed among eight molecular function groups (Fig. 3C), among which the binding (GO:0005488) and catalytic activity (GO:0003824) groups contained the greatest proportion of molecular function DEPs.
Fig. 3.
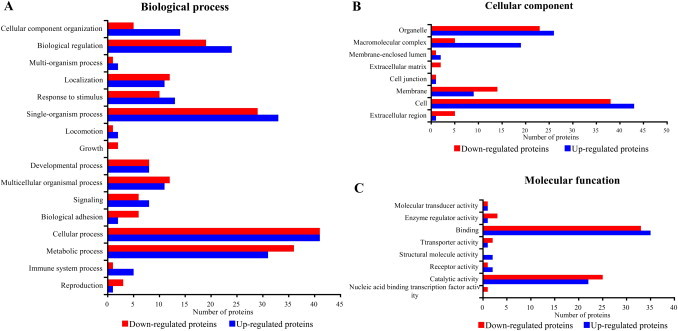
The Gene Ontology (GO) categories of the differentially expressed proteins at level 2. (A) Biological process GO categories; (B) cellular component GO categories; (C) molecular function GO categories.
3.3. KEGG pathway analysis of the DEPs
The kyoto encyclopedia of genes and genomes (KEGG) pathway is a collection of pathway maps that represent molecular interactions and reaction networks in cells. Seventy-five of the 126 DEPs identified were annotated, and mapped to a total of six KEGG pathway categories, which included the metabolism, organismal systems, cellular processes, genetic information processing, environmental information processing, and diseases pathway categories (Fig. 4 ). The annotations in the metabolism, organismal systems, and diseases pathway categories represented 32, 25, and 36 pathway groups, respectively (Fig. 4A, B and F).
Fig. 4.
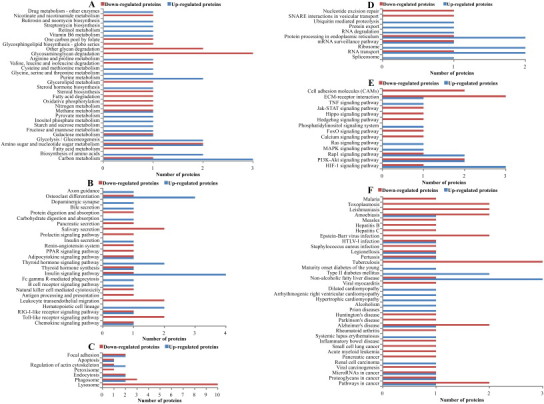
Analysis of the KEGG pathway of the differentially expressed proteins. (A) Metabolism; (B) organismal systems; (C) cellular processes; (D) genetic information processing; (E) environmental information processing; (F) diseases.
The annotations in metabolism pathways category included the carbohydrate, energy, lipid, nucleotide, amino acid, glycan biosynthesis, cofactors and vitamins, biosynthesis of other secondary metabolites, and xenobiotics pathway groups (Fig. 4A). The annotations in the organismal systems category included the Toll-like receptor (TLR) signaling (ko04620), RIG-I-like receptor (RLR) signaling (ko04622), and natural killer cell mediated cytotoxicity (ko04650) pathway groups (Fig. 4B), which represent pathways related primarily to the immune response to virus infection. The largest number of DEPs in the cellular process category were mapped to the lysosome (ko04142) pathway group, all ten of which were down-regulated DEPs (Fig. 4C). The annotations in the genetic information processing category included pathway groups related to DNA replication and repair, transcription, translation, and the folding, sorting, and degradation of proteins (Fig. 4D). The annotations in the environmental information processing category included the PI3K-Akt signaling (ko04151), mitogen-activated protein kinase (MAPK) signaling (ko04010), Jak-STAT signaling (ko04630), TNF signaling (ko04668), and cell adhesion molecule (ko04514) pathway groups (Fig. 4E), all of which represented signal transduction and signaling-molecule interactions that have been shown to be associated with virus infection. The annotations in the diseases category included the human T-cell leukemia virus infection, Epstein–Barr virus infection, hepatitis C, hepatitis B, and measles pathway groups (Fig. 4F), all of which are associated with virus infection involving in three down-regulated proteins and one up-regulated protein. Overall, more disease pathway groups were assigned to a single down-regulated DEP than those assigned to up-regulated DEPs. The integrin (β2 and β3 subunits) protein was annotated to the largest number of pathway groups (28), which included the organismal systems, environmental information processing, cellular processes, and diseases categories.
3.4. Verification of differential expression
The β tubulin as loading control, three down-regulated DEPs cystatin-C, apolipoprotein E4 and centrin-2, two up-regulated DEPs integrin-β3 and protein S100-A2, were selected to verify differential expression between the PEDV-infected and mock-infected Vero E6 cells. The immunoblotting analysis showed that the ratios of these proteins between the PEDV-infected and mock-infected groups were consistent with those obtained using the quantitative proteomics analysis (Fig. 5 ).
Fig. 5.
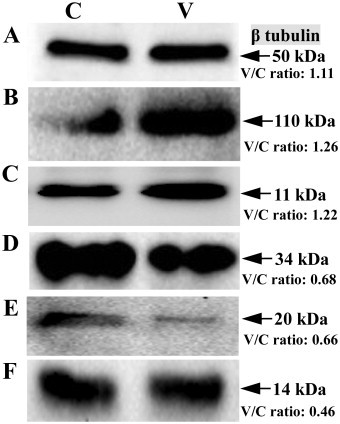
Verification of the selected differential expression proteins by western blot. (A) β tubulin; (B) Integrin β3; (C) Protein S100-A2; (D) Apolipoprotein E4; (E) Centrin-2; (F) Cystatin-C. The iTRAQ quantitative ratio “V/C ratio” is shown in Figure.
4. Discussion
In our study, PEDV infection significantly alters protein expression in Vero E6 cells. The differentially expressed proteins (DEPs) annotated to virus infection-associated signaling pathways, autophagy, and virus entry-associated proteins were analyzed further to assess their potential roles in PEDV infection. In mammals, the first line of defense against virus infection is the innate immune system. Early antiviral responses are initiated upon the recognition of pathogen-associated molecular patterns (PAMPs) by pattern recognition receptors (PRRs), resulting in the production of interferons for the innate immune response and the maturation of dendritic cells for establishing acquired immunity (Yokota et al., 2010). The PRRs are grouped into the TLRs, RLRs, and nucleotide binding-oligomerization domain-like receptors. Our results showed that PEDV infection induced the DEPs that participated in six signaling pathways involved in viral infection, including the RLR, Rap1, PI3K-Akt, MAPK, Jak-STAT, and TLR signaling pathways.
The PEDV is an enteric virus that infects the intestinal epithelial cells (IEC) of swine, causing severe diarrhea. Hirata et al. (2007) reported the RIG-I signaling pathway plays an important role in antiviral innate immunity mechanisms in IECs. Sheikh et al. (2013) reported the Rap1A signaling pathway was associated with secretory diarrhea. The Jak-STAT signaling pathway regulates the adaptive and innate mechanisms related to mucosal immunity (Heneghan et al., 2013, Wang et al., 2013). Our results showed that DEPs induced by PEDV infection in Vero E6 cells involved in the RLR, Rap1, and Jak-STAT signaling pathways. It has been reported that the TLR, MAPK, and PI3K-Akt signaling pathways play roles in host cell responses to coronaviruses (Mizutani et al., 2004, Mizutani et al., 2005, Mazaleuskaya et al., 2012). Our results demonstrate that the recognition of viral PAMPs by PRRs may mediate antiviral signaling in PEDV-infected Vero E6 cells.
Integrins are cell surface α/β heterodimeric glycoproteins that contribute to a variety of cellular functions (Stewart and Nemerow, 2007). Combinations of the various isotypes of the α and β subunits of integrins generate more than 20 different integrin proteins. Previous studies have shown that various integrin molecules are used as receptors for virus attachment (Stewart and Nemerow, 2007, Sun et al., 2013). In our current study, the expression of integrin-β2 and -β3 was down-regulated and up-regulated, respectively, in response to PEDV infection. The upregulation of integrin-β3 expression is consistent with that observed in response to dengue virus infection (Zhang et al., 2007). Our pathway analysis revealed that both integrin-β2 and -β3 are involved in 28 pathways that contribute to organismal systems, environmental information processing, cellular processes, and diseases.
The integrin αvβ3 protein has been shown to serve as an entry receptor for various viruses (Guerrero et al., 2000, Neff et al., 2000, Chu and Ng, 2004, Parry et al., 2005, Wang et al., 2005), some of which bind the integrin through an RGD sequence in a viral structural protein to initiate infection (Stewart and Nemerow, 2007). The S protein of PEDV is a glycoprotein peplomer on the viral surface that plays an important role in receptor-mediated binding and cell membrane fusion. In our study, the integrin recognized sequences of PEDV S protein was analyzed based on Ruoslahti's (1996) report. The results indicated that four conserved integrin-recognized amino acid motifs (Asp-Gly-Glu, Lys-Gly-Glu, Arg-Leu-Asp, and Leu-Asp-Val) were found in the S proteins of various PEDV strains (data not shown). These data suggest that integrin proteins may act as an infection associated protein for the attachment and entry of PEDV.
Autophagy is an essential component of host defenses against viral infection (Dong and Levine, 2013). Maier and Britton (2012) reported that β-coronaviruses induced autophagy. In our study, more DEPs were mapped to the autophagy pathway group than any of the other pathway groups. Fifteen DEPs were mapped to the lysosome and phagosome pathways. Of the 15 proteins, 12 (80%) were down-regulated DEPs. Although the autophagy pathway plays an antiviral role in virus-infected cells, the autophagy machinery is exploited by certain viruses for viral evasion and propagation. Our results showed that PEDV infection induced the downregulation of the expression of many autophagy-associated proteins. Therefore, PEDV infection might inhibit autophagy in Vero E6 cells, thus facilitating virus replication. Previous studies have shown that the microtubule-associated protein 1B is a useful biomarker protein for autophagy (Dong and Levine, 2013). We found that the expression of MAP1B was up-regulated 1.37-fold in the PEDV-infected Vero E6 cells. These results suggest that the PEDV induces autophagy.
Cystatin-C has been shown to reduce the replication of certain viruses, including the poliovirus, rhinovirus, and human coronaviruses OC43 and 229E (Korant et al., 1986, Collins and Grubb, 1991). The cleavage of S protein has been shown to be essential for the induction of cell-to-cell fusion and coronavirus entry into cells (Sturman et al., 1985). Shirato et al. (2011) reported the transmembrane type II serine protease 2 enhanced infection of PEDV in Vero cells by increasing virus release. In our study, the reduced expression of cystatin-C might facilitate PEDV replication and release through the activation of cysteine-associated proteases in Vero E6 cells. Apolipoprotein E4, galectin, clusterin, and transferrin receptor 1 have also been shown to be associated with virus infection (Hishiki et al., 2010, Peng et al., 2011, Martin and Uprichard, 2013, Tripathi et al., 2013), and may therefore function as infection-associated proteins in PEDV-infected Vero E6 cells. Additionally, the decreased in vitro expression of the adherens junction protein, such as cadherin, might be associated with a reduced integrity of PEDV-infected intestinal epithelial cells in vivo.
To the best of our knowledge, our study represents the analysis of the interactions between PEDV and Vero E6 cells using a quantitative proteomics technique. PEDV infection-associated pathways and proteins are described and discussed based on the bioinformatics analysis of the differentially expressed proteins. Our analysis of Vero E6 cell responses to PEDV infection identified relevant targets for subsequent in-depth studies of PEDV pathogenesis, expand the current knowledge base regarding the interaction between the PEDV and the host cell, and provide useful basic information about other coronaviruses. Although the Vero E6 cells are highly susceptible to PEDV infection and facilitate experimental design and performance for proteomics, the Vero E6 cell line is an interferon-deficient cell line and not a pig cell line. So, the detailed functions of these pathways and proteins in PEDV infection require further verification in the actual host cells of PEDV.
Acknowledgments
This work is supported by the National Natural Science Foundation of China (grant no. 31472209), the State National Key Laboratory of Veterinary Biotechnology (grant no. SKLVBF201506/201302), and the Program for New Century Excellent Talents in Heilongjiang Provincial University (grant no. 1252-NCET-016).
Contributor Information
Dongbo Sun, Email: dongbosun@126.com.
Li Feng, Email: fl@hvri.ac.cn.
References
- Ashburner M., Ball C.A., Blake J.A., Botstein D., Butler H., Cherry J.M., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Harris M.A., Hill D.P., Issel-Tarver L., Kasarskis A., Lewis S., Matese J.C., Richardson J.E., Ringwald M., Rubin G.M., Sherlock G. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 2000;25:25–29. doi: 10.1038/75556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belouzard S., Millet J.K., Licitra B.N., Whittaker G.R. Mechanisms of coronavirus cell entry mediated by the viral spike protein. Viruses. 2012;4:1011–1033. doi: 10.3390/v4061011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boja E., Hiltke T., Rivers R., Kinsinger C., Rahba A., Mesri M., Rodriguez H. Evolution of clinical proteomics and its role in medicine. J. Proteome Res. 2011;10:66–84. doi: 10.1021/pr100532g. [DOI] [PubMed] [Google Scholar]
- Chen J., Liu X., Shi D., Shi H., Zhang X., Li C., Chi Y., Feng L. Detection and molecular diversity of spike gene of porcine epidemic diarrhea virus in China. Viruses. 2013;5:2601–2613. doi: 10.3390/v5102601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J., Wang C., Shi H., Qiu H., Liu S., Chen X., Zhang Z., Feng L. Molecular epidemiology of porcine epidemic diarrhea virus in China. Arch. Virol. 2010;155:1471–1476. doi: 10.1007/s00705-010-0720-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Q., Li G., Stasko J., Thomas J.T., Stensland W.R., Pillatzki A.E., Gauger P.C., Schwartz K.J., Madson D., Yoon K.J., Stevenson G.W., Burrough E.R., Harmon K.M., Main R.G., Zhang J. Isolation and characterization of porcine epidemic diarrhea viruses associated with the 2013 disease outbreak in US swine. J. Clin. Microbiol. 2014;52:234–243. doi: 10.1128/JCM.02820-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chu J.J., Ng M.L. Interaction of West Nile virus with alpha v beta 3 integrin mediates virus entry into cells. J. Biol. Chem. 2004;279:54533–54541. doi: 10.1074/jbc.M410208200. [DOI] [PubMed] [Google Scholar]
- Collins A.R., Grubb A. Inhibitory effects of recombinant human cystatin C on human coronaviruses. Antimicrob. Agents Chemother. 1991;35:2444–2446. doi: 10.1128/aac.35.11.2444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dong X., Levine B. Autophagy and viruses: adversaries or allies? J. Innate Immun. 2013;5:480–493. doi: 10.1159/000346388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Götz S., García-Gómez J.M., Terol J., Williams T.D., Nagaraj S.H., Nueda M.J., Robles M., Talón M., Dopazo J., Conesa A. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008;36:3420–3435. doi: 10.1093/nar/gkn176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guerrero C.A., Méndez E., Zárate S., Isa P., López S., Arias C.F. Integrin alpha(v)beta(3) mediates rotavirus cell entry. Proc. Natl. Acad. Sci. U.S.A. 2000;97:14644–14649. doi: 10.1073/pnas.250299897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo D., Zhu Q., Feng L., Sun D. Screening and antiviral analysis of phages that display peptides with an affinity to subunit C of porcine aminopeptidase. Monoclon. Antib. Immunodiagn. Immunother. 2013;32:326–329. doi: 10.1089/mab.2013.0038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo D., Zhu Q., Zhang H., Sun D. Proteomic analysis of membrane proteins of vero cells: Exploration of potential proteins responsible for virus entry. DNA Cell Biol. 2014;33:20–28. doi: 10.1089/dna.2013.2193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He Y., Li W., Liao G., Xie J. Mycobacterium tuberculosis-specific phagosome proteome and underlying signaling pathways. J. Proteome Res. 2012;11:2635–2643. doi: 10.1021/pr300125t. [DOI] [PubMed] [Google Scholar]
- Heneghan A.F., Pierre J.F., Kudsk K.A. JAK-STAT and intestinal mucosal immunology. JAKSTAT. 2013;2:e25530. doi: 10.4161/jkst.25530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirata Y., Broquet A.H., Menchén L., Kagnof M.F. Activation of innate immune defense mechanisms by signaling through RIG-I/IPS-1 in intestinal epithelial cells. J. Immunol. 2007;179:5425–5432. doi: 10.4049/jimmunol.179.8.5425. [DOI] [PubMed] [Google Scholar]
- Hishiki T., Shimizu Y., Tobita R., Sugiyama K., Ogawa K., Funami K., Ohsaki Y., Fujimoto T., Takaku H., Wakita T., Baumert T.F., Miyanari Y., Shimotohno K. Infectivity of hepatitis C virus is influenced by association with apolipoprotein E isoforms. J. Virol. 2010;84:12048–12057. doi: 10.1128/JVI.01063-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hofmann M., Wyler R. Propagation of the virus of porcine epidemic diarrhea in cell culture. J. Clin. Microbiol. 1988;26:2235–2239. doi: 10.1128/jcm.26.11.2235-2239.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hondermarck H., Tastet C., El Yazidi-Belkoura I., Toillon R.A., Le Bourhis X. Proteomics of breast cancer: the quest for markers and therapeutic targets. J. Proteome Res. 2008;7:1403–1411. doi: 10.1021/pr700870c. [DOI] [PubMed] [Google Scholar]
- Huang Y.W., Dickerman A.W., Piñeyro P., Li L., Fang L., Kiehne R., Opriessnig T., Meng X.J. Origin, evolution, and genotyping of emergent porcine epidemic diarrhea virus strains in the United States. MBio. 2013;4:e00737–e813. doi: 10.1128/mBio.00737-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kanehisa M., Goto S., Sato Y., Furumichi M., Tanabe M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 2012;40:109–114. doi: 10.1093/nar/gkr988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim Y., Lee C. Ribavirin efficiently suppresses porcine nidovirus replication. Virus Res. 2013;171:44–53. doi: 10.1016/j.virusres.2012.10.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korant B.D., Towatari T., Ivanoff L., Petteway S.J., Brzin J., Lenarcic B., Turk V. Viral therapy: prospects for protease inhibitors. J. Cell Biochem. 1986;32:91–95. doi: 10.1002/jcb.240320202. [DOI] [PubMed] [Google Scholar]
- Li W., Li H., Liu Y., Pan Y., Deng F., Song Y., Tang X., He Q. New variants of porcine epidemic diarrhea virus, China, 2011. Emerg. Infect. Dis. 2012;18:1350–1353. doi: 10.3201/eid1808.120002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linde M.E., Colquhoun D.R., Mohien C.U., Kole T., Aquino V., Cotter R., Edwards N., Hildreth J.E., Graham D.R. The conserved set of host proteins incorporated into HIV-1 virions suggests a common egress pathway in multiple cell types. J. Proteome Res. 2013;12:2045–2054. doi: 10.1021/pr300918r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maier H.J., Britton P. Involvement of autophagy in coronavirus replication. Viruses. 2012;4:3440–3451. doi: 10.3390/v4123440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marthaler D., Jiang Y., Otterson T., Goyal S., Rossow K., Collins J. Complete genome sequence of porcine epidemic diarrhea virus strain USA/Colorado/2013 from the United States. Genome Announc. 2013;1:e00555–e613. doi: 10.1128/genomeA.00555-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin D.N., Uprichard S.L. Identification of transferrin receptor 1 as a hepatitis C virus entry factor. Proc. Natl. Acad. Sci. U.S.A. 2013;110:10777–10782. doi: 10.1073/pnas.1301764110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mazaleuskaya L., Veltrop R., Ikpeze N., Martin-Garcia J., Navas-Martin S. Protective role of Toll-like Receptor 3-induced type I interferon in murine coronavirus infection of macrophages. Viruses. 2012;4:901–923. doi: 10.3390/v4050901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mizutani T., Fukushi S., Murakami M., Hirano T., Saijo M., Kurane I., Morikawa S. Tyrosine dephosphorylation of STAT3 in SARS coronavirus-infected Vero E6 cells. FEBS Lett. 2004;577:187–192. doi: 10.1016/j.febslet.2004.10.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mizutani T., Fukushi S., Saijo M., Kurane I., Morikawa S. JNK and PI3k/Akt signaling pathways are required for establishing persistent SARS-CoV infection in Vero E6 cells. Biochim. Biophys. Acta. 2005;1741:4–10. doi: 10.1016/j.bbadis.2005.04.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neff S., Mason P.W., Baxt B. High-efficiency utilization of the bovine integrin alpha(v)beta(3) as a receptor for foot-and-mouth disease virus is dependent on the bovine beta(3) subunit. J. Virol. 2000;74:7298–7306. doi: 10.1128/jvi.74.16.7298-7306.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pan Y., Tian X., Li W., Zhou Q., Wang D., Bi Y., Chen F., Song Y. Isolation and characterization of a variant porcine epidemic diarrhea virus in China. Virol. J. 2012;9:195. doi: 10.1186/1743-422X-9-195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Papachristou E.K., Roumeliotis T.I., Chrysagi A., Trigoni C., Charvalos E., Townsend P.A., Pavlakis K., Garbis S.D. The shotgun proteomic study of the human ThinPrep cervical smear using iTRAQ mass-tagging and 2D LC-FT-Orbitrap-MS: the detection of the human papillomavirus at the protein level. J. Proteome Res. 2013;12:2078–2089. doi: 10.1021/pr301067r. [DOI] [PubMed] [Google Scholar]
- Parry C., Bell S., Minson T., Browne H. Herpes simplex virus type 1 glycoprotein H binds to alphavbeta3 integrins. J. Gen. Virol. 2005;86:7–10. doi: 10.1099/vir.0.80567-0. [DOI] [PubMed] [Google Scholar]
- Peng G., Sun D., Rajashankar K.R., Qian Z., Holmes K.V., Li F. Crystal structure of mouse coronavirus receptor-binding domain complexed with its murine receptor. Proc. Natl. Acad. Sci. U.S.A. 2011;108:10696–10701. doi: 10.1073/pnas.1104306108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pensaert M.B., Debouck P. A new coronavirus-like particle associated with diarrhea in swine. Arch. Virol. 1978;58:243–247. doi: 10.1007/BF01317606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Puranaveja S., Poolperm P., Lertwatcharasarakul P., Kesdaengsakonwut S., Boonsoongnern A., Urairong K., Kitikoon P., Choojai P., Kedkovid R., Teankum K., Thanawongnuwech R. Chinese-like strain of porcine epidemic diarrhea virus, Thailand. Emerg. Infect. Dis. 2009;15:1112–1115. doi: 10.3201/eid1507.081256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quevillon E., Silventoinen V., Pillai S., Harte N., Mulder N., Apweiler R., Lopez R. InterProScan: protein domains identifier. Nucleic Acids Res. 2005;33:116–120. doi: 10.1093/nar/gki442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ren X., Suo S., Jang Y.S. Development of a porcine epidemic diarrhea virus M protein-based ELISA for virus detection. Biotechnol. Lett. 2011;33:215–220. doi: 10.1007/s10529-010-0420-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruoslahti E. RGD and other recognition sequences for integrins. Annu. Rev. Cell Dev. Biol. 1996;12:697–715. doi: 10.1146/annurev.cellbio.12.1.697. [DOI] [PubMed] [Google Scholar]
- Sheikh I.A., Koley H., Chakrabarti M.K., Hoque K.M. The Epac1 signaling pathway regulates Cl− secretion via modulation of apical KCNN4c channels in diarrhea. J. Biol. Chem. 2013;288:20404–20415. doi: 10.1074/jbc.M113.467860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shirato K., Matsuyama S., Ujike M., Taguchi F. Role of proteases in the release of porcine epidemic diarrhea virus from infected cells. J. Virol. 2011;85:7872–7880. doi: 10.1128/JVI.00464-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stevenson G.W., Hoang H., Schwartz K.J., Burrough E.R., Sun D., Madson D., Cooper V.L., Pillatzki A., Gauger P., Schmitt B.J., Koster L.G., Killian M.L., Yoon K.J. Emergence of porcine epidemic diarrhea virus in the United States: clinical signs, lesions, and viral genomic sequences. J. Vet. Diagn. Invest. 2013;25:649–654. doi: 10.1177/1040638713501675. [DOI] [PubMed] [Google Scholar]
- Stewart P.L., Nemerow G.R. Cell integrins: commonly used receptors for diverse viral pathogens. Trends Microbiol. 2007;15:500–507. doi: 10.1016/j.tim.2007.10.001. [DOI] [PubMed] [Google Scholar]
- Sturman L.S., Ricard C.S., Holmes K.V. Proteolytic cleavage of the E2 glycoprotein of murine coronavirus: activation of cell-fusing activity of virions by trypsin and separation of two different 90 K cleavage fragments. J. Virol. 1985;56:904–911. doi: 10.1128/jvi.56.3.904-911.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun D., Shi H., Chen J., Shi D., Zhu Q., Zhang H., Liu S., Wang Y., Qiu H., Feng L. Generation of a mouse scFv library specific for porcine aminopeptidase N using the T7 phage display system. J. Virol. Methods. 2012;182:99–103. doi: 10.1016/j.jviromet.2012.03.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun D., Zhang H., Guo D., Sun A., Wang H. Shotgun proteomic analysis of plasma from dairy cattle suffering from footrot: characterization of potential disease-associated factors. PLoS ONE. 2013;8:e55973. doi: 10.1371/journal.pone.0055973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun D.B., Feng L., Shi H.Y., Chen J.F., Cui X.C., Chen H.Y., Liu S.W., Tong Y.E., Wang Y.F., Tong G.Z. Identification of two novel B cell epitopes on porcine epidemic diarrhea virus spike protein. Vet. Microbiol. 2008;131:73–81. doi: 10.1016/j.vetmic.2008.02.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tripathi S., Batra J., Cao W., Sharma K., Patel J.R., Ranjan P., Kumar A., Katz J.M., Cox N.J., Lal R.B., Sambhara S., Lal S.K. Influenza A virus nucleoprotein induces apoptosis in human airway epithelial cells: implications of a novel interaction between nucleoprotein and host protein Clusterin. Cell Death Dis. 2013;4:e562. doi: 10.1038/cddis.2013.89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Unwin R.D., Griffiths G.R., Whetton A.D. Simultaneous analysis of relative protein expression levels across multiple samples using iTRAQ isobaric tags with 2D nano LC–MS/MS. Nat. Protoc. 2010;5:1574–1582. doi: 10.1038/nprot.2010.123. [DOI] [PubMed] [Google Scholar]
- Wang R., Nan Y., Yu Y., Zhang Y.J. Porcine reproductive and respiratory syndrome virus Nsp1β inhibits interferon-activated JAK/STAT signal transduction by inducing karyopherin-α1 degradation. J. Virol. 2013;87:5219–5228. doi: 10.1128/JVI.02643-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X., Huang D.Y., Huong S.M., Huang E.S. Integrin alphavbeta3 is a coreceptor for human cytomegalovirus. Nat. Med. 2005;11:515–521. doi: 10.1038/nm1236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wiśniewski J.R., Zougman A., Nagaraj N., Mann M. Universal sample preparation method for proteome analysis. Nat. Methods. 2009;6:359–362. doi: 10.1038/nmeth.1322. [DOI] [PubMed] [Google Scholar]
- Yang X., Huo J.Y., Chen L., Zheng F.M., Chang H.T., Zhao J., Wang X.W., Wang C.Q. Genetic variation analysis of reemerging porcine epidemic diarrhea virus prevailing in central China from 2010 to 2011. Virus Genes. 2013;46:337–344. doi: 10.1007/s11262-012-0867-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ye Y., Yan G., Luo Y., Tong T., Liu X., Xin C., Liao M., Fan H. Quantitative proteomics by amino acid labeling in foot-and-mouth disease virus (FMDV)-infected cells. J. Proteome Res. 2013;12:363–377. doi: 10.1021/pr300611e. [DOI] [PubMed] [Google Scholar]
- Yokota S., Okabayashi T., Fujii N. The battle between virus and host: modulation of Toll-like receptor signaling pathways by virus infection. Mediators Inflamm. 2010;2010:184328. doi: 10.1155/2010/184328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuan J., Gao H., Sui J., Duan H., Chen W.N., Ching C.B. Cytotoxicity evaluation of oxidized single-walled carbon nanotubes and graphene oxide on human hepatoma HepG2 cells: an iTRAQ-coupled 2D LC–MS/MS proteome analysis. Toxicol. Sci. 2012;126:149–161. doi: 10.1093/toxsci/kfr332. [DOI] [PubMed] [Google Scholar]
- Zeng, S., Zhang, H., Ding, Z., Luo, R., An, K., Liu, L., Bi, J., Chen, H., Xiao, S., Fang, L., 2015. Proteome analysis of porcine epidemic diarrhea virus (PEDV)-infected vero cells. Proteomics 2015 Jan 20, http://dx.doi.org/10.1002/pmic.201400458, PubMed PMID: 25604190 (epub ahead of print). [DOI] [PMC free article] [PubMed]
- Zhang J.L., Wang J.L., Gao N., Chen Z.T., Tian Y.P., An J. Up-regulated expression of beta3 integrin induced by dengue virus serotype 2 infection associated with virus entry into human dermal microvascular endothelial cells. Biochem. Biophys. Res. Commun. 2007;356:763–768. doi: 10.1016/j.bbrc.2007.03.051. [DOI] [PubMed] [Google Scholar]
- Zhu Q., Guo D., Feng L., Sun D. Expression and purification of the scFv from hybridoma cells secreting a monoclonal antibody against S Protein of PEDV. Monoclon. Antib. Immunodiagn. Immunother. 2013;32:41–46. doi: 10.1089/mab.2012.0089. [DOI] [PMC free article] [PubMed] [Google Scholar]


