Abstract
Pharmacophore-based virtual screening is an effective, inexpensive and fast approach to discovering useful starting points for drug discovery. In this study, we developed a pharmacophore model for the main proteinase of severe acute respiratory syndrome coronavirus (SARS-CoV). Then we used this pharmacophore model to search NCI 3D database including 250, 251 compounds and identified 30 existing drugs containing the pharmacophore query. Among them are six compounds that already exhibited anti-SARS-CoV activity experimentally. This means that our pharmacophore model can lead to the discovery of potent anti-SARS-CoV inhibitors or promising lead compounds for further SARS-CoV main proteinase inhibitor development.
Keywords: SARS-CoV, Pharmacophore, Virtual screening, Drug design
1. Introduction
The infection of the newly emerged severe acute respiratory syndrome coronavirus (SARS-CoV) is characterized by acute flu-like symptoms that progress to acute lung injury or acute respiratory distress syndrome with over 10% of mortality [1]. To date there are no universally recommended therapy for the disease. Many scientists are now making efforts to develop effective drugs against SARS. The combination therapy of corticosteroid with lopinavir, ribavirin and ritonavir can improve clinical response and reduce mortality rates apparently [2], [3]. Cinatl et al. [4] found that ribavirin, azauridine, pyrazofurin and glycyrrhizin are active against SARS-CoV. Barnard et al. [5] reported that calpain inhibitors and β-d-N4-hydroxycytidine exhibit inhibitory effects on SARS-CoV.
Structure-based drug design focuses on two important approaches: one is receptor-based docking technique, another is pharmacophore-based virtual screening technique. A pharmacophore is the 3D arrangement of atoms or functional groups essential for the compound to bind to a specific receptor [6]. The power of a pharmacophore model is to discover new leads by using 3D database pharmacophore searching and guide chemists to synthesize new compounds [7]. Such a pharmacophore-based method has been successfully applied to many drug development programs [8], [9], [10], [11], [12], [13], [14], [15], [16], [17], [18]. The main proteinase of SARS-CoV plays an important role in virus replication and is the primary target for drugs. The aim of this study is to develop 3D pharmacophore models for SARS-CoV main proteinase and expect to provide useful knowledge for anti-SARS drug design.
2. Material and methods
There are two methods to derive a reasonable pharmacophore model. One is from the crystal structure of protein–ligand complex, another is based on molecular modeling of enzyme with its potential inhibitors. Here we used the experimental structure of SARS-CoV main proteinase complexed with its peptide inhibitor CMK (PDB ID 1UK4) [19] and the predicted structures of SARS-CoV main proteinase with six drugs/compounds [20] for establishing pharmacophore models. The structures of CMK peptide and six compounds are shown in Fig. 1 .
Fig. 1.
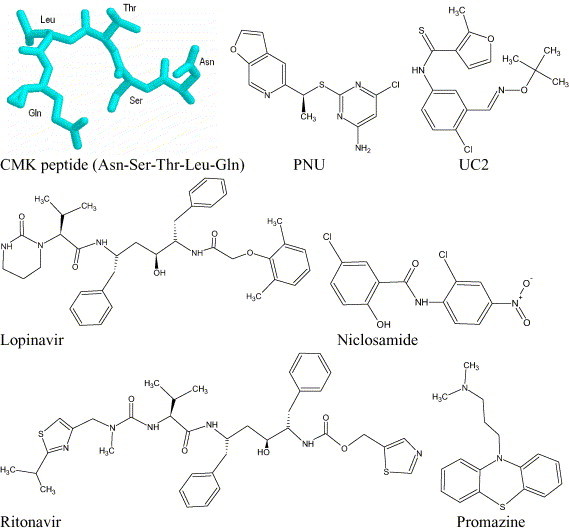
The peptide and compounds used for pharmacophore generation.
The POCKET module in LigBuilder program [21] was employed to obtain the pharmacophore models of SARS-CoV main proteinase. This approach was successfully applied to the identification of novel inhibitors for alanine racemase [22]. The proposed pharmacophore model is a binding-site-derived pharmacophore model, which includes the following pharmacophore features of ligands binding to the enzyme’s active site: a positively charged nitrogen atom (ammonium cation) to represent a hydrogen bond donor (HBD), a negatively charged oxygen atom (as in a carboxyl group) to represent a hydrogen bond acceptor (HBA), and a carbon atom (methane) to represent a hydrophobic center (HPC). A pharmacophore model is generated for each protein–ligand complex.
3. Results and discussion
Using CMK peptide and six compounds in Fig. 1, we generated a set of seven eight-point pharmacophore models for SARS-CoV main proteinase, which is listed in Table 1 . These hypotheses exhibit different features due to the diversities of the compounds involved. Based on these models, we extracted a common four-point pharmacophore distance pattern shown in Fig. 2 , P1 is HBA, HBD and HPC, P2 is HBA and HPC, P3 is HBA and HBD, P4 is HBA and HBD. Such a pharmacophore distance pattern was subsequently used for the 3D database search.
Table 1.
The eight-point pharmacophore models obtained by LigBuilder from seven peptide and compounds
| Pharmacophore | Peptide/compounds | Features |
|---|---|---|
| 1 | CMK peptide | HBD HBD HPC HBD HPC HBD HPC HBA |
| 2 | Lopinavir | HBA HBD HBA HBD HBA HBA HBA HBA |
| 3 | Ritonavir | HPC HPC HBD HBD HBD HBA HBD HBA |
| 4 | Niclosamide | HBA HBD HBA HBA HBA HBD HPC HBA |
| 5 | Promazine | HBA HBD HBA HPC HBA HBA HBA HBA |
| 6 | PNU | HBA HPC HBA HBD HBD HBA HBA HBA |
| 7 | UC2 | HPC HBD HBD HBA HBD HPC HPC HBA |
HBA = hydrogen bond acceptor, HBD = hydrogen bond donor, HPC = hydrophobic center.
Fig. 2.
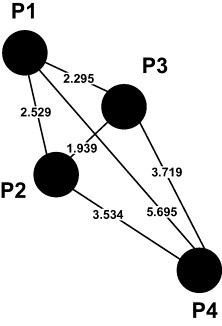
Four-point pharmacophore distance pattern for SARS-CoV main proteinase. Here, P1 is HBA, HBD and HPC, P2 is HBA and HPC, P3 is HBA and HBD, P4 is HBA and HBD.
The pharmacophore searching in 3D database was conducted in the 3D NCI database, which has 250, 251 open structures ready for searching. Table 2 summarizes the results for similarity search of above-mentioned four-point pharmacophore model with constraints: (1) the distance ranges for P1P2, P1P3, P2P3, P1P4, P2P4 and P3P4 (Fig. 2) are 2–3, 2–3, 1.5–2.5, 5–6, 3–4 and 3–4 Å, respectively; (2) the compound is drug; (3) the antiviral probability is over 70%; (4) the compounds labeled as “No Name” is not included. After review of these hitlists, 30 drugs were selected for further analysis, their chemical structures and bioactivities (documented in anti-HIV and anti-opportunistic infection chemical compound database, which contains approximately 100,000 compounds, http://www.apps1.niaid.nih.gov/struct_search/an/an_search.htm) are shown in Fig. 3 and Table 3 , respectively. It is noted that almost all drugs exhibited anti-HIV activity, and some of them have activities against Mycobacterium tuberculosis (puromycin), influenza virus (5-bromo-2′-deoxycytidine), hepatitis virus (dideoxyguanosine, 2′,3′-dideoxycytidine and ribavirin), dengue virus (azauridine and ribavirin), rhinovirus and poliovirus (ribavirin). This should be very meaningful in consideration of the following facts: (1) there are some links between SARS-CoV and HIV and HBV [23], [24], [25]; (2) similar structure patterns exist in SARS-CoV main proteinase with rhinovirus 3c protease, poliovirus 3c proteinase, HAV 3c protease, HCV Ns3 protease and dengue virus Ns3 protease [26]; (3) SARS-CoV has clinically similar symptoms with influenza virus/M. tuberculosis, such as fever, cough, pains, pneumonia and death [27].
Table 2.
Summary of NCI database search by four-point pharmacophores
| Pharmacophore features in Fig. 2 | Hits | |||
|---|---|---|---|---|
| P1 |
P2 |
P3 |
P4 |
|
| HBD | HBA | HBD | HBD | 987 |
| HBA | HBA | HBA | HBA | 794 |
| HPC | HBA | HBA | HBD | 286 |
| HPC | HPC | HBA | HBD | 305 |
| HPC | HBA | HBD | HBD | 298 |
| HBA | HBA | HBD | HBD | 881 |
| HBA | HBA | HBA | HBD | 895 |
HBA = hydrogen bond acceptor, HBD = hydrogen bond donor, HPC = hydrophobic center.
Fig. 3.
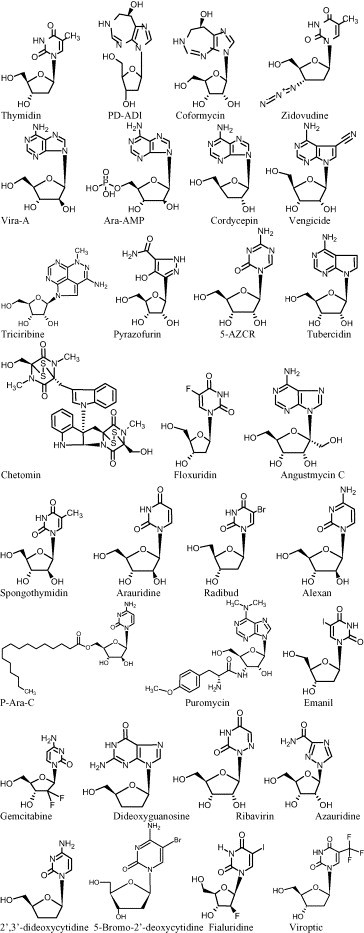
Chemical structures of 30 drugs obtained by four-point pharmacophore search in NCI 3D database.
Table 3.
Thirty drugs obtained by four-point pharmacophore search in NCI 3D database
| Name | NSC number | Formula | Bioactivity documented in HIV/OI therapeutics database (http://www.apps1.niaid.nih.gov/struct_search/an/an_search.htm) |
|---|---|---|---|
| PD-ADI | 218321 | C11H16N4O4 | HIV |
| Coformycin | 277817 | C11H16N4O5 | HIV |
| Zidovudine | 602670 | C10H13N5O4 | HIV, HSV, human cytomegalovirus, vaccinia virus, cowpox virus |
| Vira-A | 404241 | C10H13N5O4 | HIV, HSV, human cytomegalovirus, varicella-zoster virus, vaccinia virus, cowpox virus |
| Angustmycin C | 53104 | C11H15N5O5 | |
| ARA-AMP | 259272 | C10H14N5O7P | HIV, HSV, vaccinia virus |
| Cordycepin | 63984 | C10H13N5O3 | HIV |
| Triciribine | 154020 | C13H16N6O4 | HIV, human cytomegalovirus, HSV |
| 5-AZCR | 102816 | C8H12N4O5 | HIV |
| Puromycin | 3055 | C22H29N7O5 | HIV, M. tuberculosis |
| CHETOMIN | 289491 | C31H30N6O6S4 | HIV |
| Vengicide | 99843 | C12H13N5O4 | Human cytomegalovirus |
| Spongothymidin | 68929 | C10H14N2O6 | HIV, HSV, varicella-zoster virus |
| Arauridine | 68928 | C9H12N2O6 | |
| P-Ara-C | 135962 | C25H43N3O6 | HIV |
| Pyrazofurin | 143095 | C9H13N3O6 | HIV, vaccinia virus, West Nile virus |
| Thymidin | 21548 | C10H14N2O5 | HIV, varicella-zoster virus |
| Radibud | 38297 | C9H11BrN2O5 | |
| Alexan | 63878 | C9H13N3O5 | HIV, HSV, human cytomegalovirus, varicella-zoster virus, measles virus |
| Floxuridin | 27640 | C9H11FN2O5 | HIV, HSV, vaccinia virus |
| Gemcitabine | 613327 | C9H12ClF2N3O4 | HIV, cowpox virus, vaccinia virus |
| Dideoxyguanosine | 619072 | C10H13N5O3 | HIV, HBV |
| 2′,3′-Dideoxycytidine | 606170 | C9H13N3O3 | HIV, HBV |
| 5-Bromo-2′-deoxycytidine | 61765 | C9H12BrN3O4 | HSV |
| Ribavirin | 163039 | C8H12N4O5 | HIV, HSV, HCV, influenza virus, dengue virus, measles virus, respiratory syncytial virus, rhinovirus, polio virus, vaccinia virus, cowpox virus |
| Azauridine | 32074 | C8H11N3O6 | West Nile virus, cowpox virus, vaccinia virus, dengue virus, Japanese encephalitis virus, Yellow fever virus |
| Fialuridine | 678514 | C9H10FIN2O5 | Cowpox virus, vaccinia virus, HSV, varicella-zoster virus |
| Emanil | 39661 | C9H11IN2O5 | HIV, HSV, cowpox virus, vaccinia virus, varicella-zoster virus |
| Tubercidin | 56408 | C11H14N4O4 | HIV, vaccinia virus, human cytomegalovirus |
| Viroptic | 75520 | C10H11F3N2O5 | HSV, cowpox virus, vaccinia virus |
Indeed, among the 30 drugs are six compounds that already exhibited anti-SARS-CoV activity experimentally: azauridine, pyrazofurin, ribavirin, 2′,3′-dideoxycytidine, dideoxyguanosine, and 5-bromo-2′-deoxycytidine [4], [5]. This shows that our pharmacophore model can lead to the discovery of potent anti-SARS-CoV inhibitors or at least provide some useful clues. Fig. 4 shows the mappings of the six compounds into the four-point pharmacophore model, which are mapped to 1–3 HBA, 1–2 HBD and 0–1 HPC. In addition, most of the remaining compounds have remarkable similarities with one of the above six compounds, for example, azauridine with 5-AZCR, alexan, arauridine and spongothymidin; 5-bromo-2′-deoxycytidine with emanil, fialuridine, floxuridin, radibud, thymidin and viroptic; dideoxyguanosine with angustmycin C, cordycepin, tubercidin, vengicide and vira-A. The superpositions for these compounds are shown in Fig. 5 . In summary, our results indicate that the existing 30 drugs identified by our pharmacophore model could be potential inhibitors against SARS-CoV, or at least good lead compounds for anti-SARS-CoV drug design.
Fig. 4.
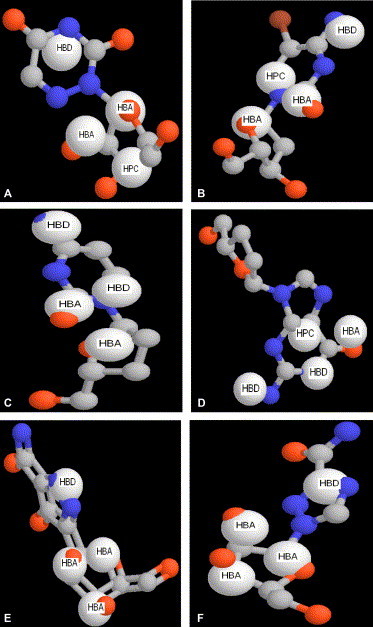
The mappings of six compounds that experimentally exhibited anti-SARS-CoV activity into the four-point pharmacophore model: (A) azauridine, (B) 5-bromo-2′-deoxycytidine, (C) dideoxycytidine, (D) dideoxyguanosine, (E) pyrazofurin, (F) ribavirin.
Fig. 5.
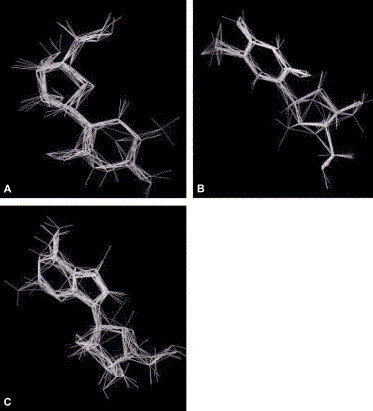
Superpositions of anti-SARS-CoV compounds with other compounds. (A) azauridine with 5-AZCR, alexan, arauridine and spongothymidin; (B) 5-bromo-2′-deoxycytidine with email, fialuridine, floxuridin, radibud, thymidin and viroptic; (C) dideoxyguanosine with angustmycin C, cordycepin, tubercidin, vengicide and vira-A.
Acknowledgements
We wish to thank the Hong Kong Innovation and Technology Fund for supporting the present research.
References
- 1.Lew T.W., Kwek T.K., Tai D., Earnest A., Loo S., Singh K. Acute respiratory distress syndrome in critically ill patients with severe acute respiratory syndrome. J. Am. Med. Assoc. 2003;290(3):374–380. doi: 10.1001/jama.290.3.374. [DOI] [PubMed] [Google Scholar]
- 2.Chan K.S., Lai S.T., Chu C.M., Tsui E., Tam C.Y., Wong M.M. Treatment of severe acute respiratory syndrome with lopinavir/ritonavir: a multicentre retrospective matched cohort study. Hong Kong Med. J. 2003;9(6):399–406. [PubMed] [Google Scholar]
- 3.Chu C.M., Cheng V.C., Hung I.F., Wong M.M., Chan K.H., Chan K.S., HKU/UCH SARS Study Group Role of lopinavir/ritonavir in the treatment of SARS: initial virological and clinical findings. Thorax. 2004;59(3):252–256. doi: 10.1136/thorax.2003.012658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Cinatl J., Morgenstern B., Bauer G., Chandra P., Rabenau H., Doerr H.W. Glycyrrhizin, an active component of liquorice roots, and replication of SARS-associated coronavirus. Lancet. 2003;361:2045–2046. doi: 10.1016/S0140-6736(03)13615-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Barnard D.L., Hubbard V.D., Burton J., Smee D.F., Morrey J.D., Otto M.J. Inhibition of severe acute respiratory syndrome-associated coronavirus (SARSCoV) by calpain inhibitors and beta-d-N4-hydroxycytidine. Antiviral Chem. Chemother. 2004;15(1):15–22. doi: 10.1177/095632020401500102. [DOI] [PubMed] [Google Scholar]
- 6.Mason J.S., Good A.C., Martin E.J. 3-D pharmacophores in drug discovery. Curr. Pharm. Des. 7. 2001;7:567–597. doi: 10.2174/1381612013397843. [DOI] [PubMed] [Google Scholar]
- 7.Dror O., Shulman-Peleg A., Nussinov R., Wolfson H.J. Predicting molecular interactions in silico: I. A guide to pharmacophore identification and its applications to drug design. Curr. Med. Chem. 2004;11(1):71–90. doi: 10.2174/0929867043456287. [DOI] [PubMed] [Google Scholar]
- 8.Neamati N., Hong H., Mazumder A., Wang S., Sunder S., Nicklaus M.C. Depsides and depsidones as inhibitors of HIV-1 integrase: discovery of novel inhibitors through 3D database searching. J. Med. Chem. 1997;40(6):942–951. doi: 10.1021/jm960759e. [DOI] [PubMed] [Google Scholar]
- 9.Hong H., Neamati N., Wang S., Nicklaus M.C., Mazumder A., Zhao H. Discovery of HIV-1 integrase inhibitors by pharmacophore searching. J. Med. Chem. 1997;40(6):930–936. doi: 10.1021/jm960754h. [DOI] [PubMed] [Google Scholar]
- 10.Hong H., Neamati N., Winslow H.E., Christensen J.L., Orr A., Pommier Y. Identification of HIV-1 integrase inhibitors based on a four-point pharmacophore. Antiviral Chem. Chemother. 1998;9(6):461–472. doi: 10.1177/095632029800900602. [DOI] [PubMed] [Google Scholar]
- 11.Nicklaus M.C., Neamati N., Hong H., Mazumder A., Sunder S., Chen J. HIV-1 integrase pharmacophore: discovery of inhibitors through three-dimensional database searching. J. Med. Chem. 1997;40(6):920–929. doi: 10.1021/jm960596u. [DOI] [PubMed] [Google Scholar]
- 12.Clement O.O., Freeman C.M., Hartmann R.W., Handratta V.D., Vasaitis T.S., Brodie A.M. Three dimensional pharmacophore modeling of human CYP17 inhibitors. Potential agents for prostate cancer therapy. J. Med. Chem. 2003;46(12):2345–2351. doi: 10.1021/jm020576u. [DOI] [PubMed] [Google Scholar]
- 13.Debnath A.K. Generation of predictive pharmacophore models for CCR5 antagonists: study with piperidine- and piperazine-based compounds as a new class of HIV-1 entry inhibitors. J. Med. Chem. 2003;46(21):4501–4515. doi: 10.1021/jm030265z. [DOI] [PubMed] [Google Scholar]
- 14.Keller P.A., Birch C., Leach S.P., Tyssen D., Griffith R. Novel pharmacophore-based methods reveal gossypol as a reverse transcriptase inhibitor. J. Mol. Graph Model. 2003;21(5):365–373. doi: 10.1016/s1093-3263(02)00183-3. [DOI] [PubMed] [Google Scholar]
- 15.Faragalla J., Bremner J., Brown D., Griffith R., Heaton A. Comparative pharmacophore development for inhibitors of human and rat 5-alpha-reductase. J. Mol. Graph Model. 2003;22(1):83–92. doi: 10.1016/S1093-3263(03)00138-4. [DOI] [PubMed] [Google Scholar]
- 16.Doddareddy M.R., Jung H.K., Lee J.Y., Lee Y.S., Cho Y.S., Koh H.Y. First pharmacophoric hypothesis for T-type calcium channel blockers. Bioorg. Med. Chem. 2004;12(7):1605–1611. doi: 10.1016/j.bmc.2004.01.034. [DOI] [PubMed] [Google Scholar]
- 17.Mustata G.I., Brigo A., Briggs J.M. HIV-1 integrase pharmacophore model derived from diverse classes of inhibitors. Bioorg. Med. Chem. Lett. 2004;14(6):1447–1454. doi: 10.1016/j.bmcl.2004.01.027. [DOI] [PubMed] [Google Scholar]
- 18.Sun H., Greeley D.N., Chu X.J., Cheung A., Danho W., Swistok J. A predictive pharmacophore model of human melanocortin-4 receptor as derived from the solution structures of cyclic peptides. Bioorg. Med. Chem. 2004;12(10):2671–2677. doi: 10.1016/j.bmc.2004.03.017. [DOI] [PubMed] [Google Scholar]
- 19.Yang H., Yang M., Ding Y., Liu Y., Lou Z., Zhou Z. The crystal structures of severe acute respiratory syndrome virus main protease and its complex with an inhibitor. Proc. Natl. Acad. Sci. USA. 2003;100:13190–13195. doi: 10.1073/pnas.1835675100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.a Zhang X.W., Yap Y.L. Old drugs as lead compounds for a new disease? Binding analysis of SARS coronavirus main proteinase with HIV, psychotic and parasite drugs. Bioorg. Med. Chem. 2004;12(10):2517–2521. doi: 10.1016/j.bmc.2004.03.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wang R., Gao Y., Lai L. LigBuilder: a multipurpose program for structure-based drug design. J. Mol. Model. (Online) 2000;6:498–516. [Google Scholar]
- 22.Mustata G.I., Briggs J.M. A structure-based design approach for the identification of novel inhibitors: application to an alanine racemase. J. Comput. Aided Mol. Des. 2002;16(12):935–953. doi: 10.1023/a:1023875514454. [DOI] [PubMed] [Google Scholar]
- 23.Kliger Y., Levanon E.Y. Cloaked similarity between HIV-1 and SARS-CoV suggests an anti-SARS strategy. BMC Microbiol. 2003;3:20–26. doi: 10.1186/1471-2180-3-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.b Zhang X.W., Yap Y.L. Structural similarity between HIV-1 gp41 and SARS-CoV S2 proteins suggests an analogous membrane fusion mechanism. J. Mol. Struct. (Theochem.) 2004;677:73–76. doi: 10.1016/j.theochem.2004.02.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Robertson M. FgI2: link between hepatitis B and SARS? Drug Discov. Today. 2003;8:768–770. doi: 10.1016/S1359-6446(03)02836-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.c Zhang X.W., Yap Y.L. Exploring the binding mechanism of the main proteinase in SARS-associated coronavirus and its implication to anti-SARS drug design. Bioorg. Med. Chem. 2004;12:2219–2223. doi: 10.1016/j.bmc.2004.02.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Oxford J., Balasingam S., Lambkin R. A new millennium conundrum: how to use a powerful class of influenza anti-neuraminidase drugs (NAIs) in the community. J. Antimicrob. Chemother. 2004;53(2):133–136. doi: 10.1093/jac/dkh037. [DOI] [PMC free article] [PubMed] [Google Scholar]


