Abstract
Since the prevalence of bovine norovirus (BNoV) and their genetic diversity have only been reported in the USA, England, Germany and The Netherlands, this study examined the prevalence and genetic diversity of BNoVs in diarrheic calves in South Korea using 645 diarrheic fecal specimens from calves by RT-PCR and nested PCR assays. Overall, 9.3% of the diarrheic fecal samples tested positive for BNoVs by either RT-PCR or nested PCR, of which 5.9% samples also tested positive for other enteric pathogens including the bovine coronavirus, bovine viral diarrhea virus, bovine torovirus, bovine groups A, B and C rotaviruses, bovine enteric Nebraska-like calicivirus and Escherichia coli. The genetic diversity was determined by direct sequencing of the partial RdRp region of 12 BNoVs detected from the fecal samples by nested PCR. Among the BNoVs examined, one Korean BNoV strain had the highest nucleotide (86.8%) and amino acid (99.1%) identity with the genotype 1 BNoV (GIII-1) strain, while the remaining 11 Korean BNoVs shared a higher nucleotide (88.0–90.5%) and amino acid (93.5–99.1%) identity with the genotype 2 BNoV (GIII-2) strains. The phylogenetic data for the nucleotide and amino acid sequences also demonstrated that one Korean BNoV strain clustered with GIII-1 but the remaining eleven strains clustered with GIII-2. In conclusion, BNoV infections are endemic and there are two distinct genotypes with GIII-2 being the main genotype circulating in the calf population in South Korea.
Keywords: Bovine, Norovirus, Diarrhea, Prevalence, Genetic diversity, Genotypes
1. Introduction
Caliciviruses are nonenveloped, single-stranded RNA viruses with positive-sense genomes of 7.4–8.3 kb (Green et al., 2000). Based on the genomic organization and genetic analysis (Green et al., 2000), the family Caliciviridae is divided into at least four genera: Vesivirus, Lagovirus, Norovirus (NoV), and Sapovirus (SaV) with a new proposed genus, NB-like (Oliver et al., 2006, Smiley et al., 2002). The NoV genome is organized into three open reading frames (ORFs) that encode the nonstructural proteins (ORF1), the major capsid protein (ORF2) and the minor capsid protein (ORF3) (Jiang et al., 1993, Lambden et al., 1993). Based on the genetic divergence in the RNA-dependent RNA polymerase (RdRp) and capsid genes, the NoVs have been tentatively assigned to five genogroups (G) with multiple genotypes (Zheng et al., 2006). The strains of three genogroups, GI, GII, and GIV, are found in humans (GII-11 and GII-19 are porcine), and GIII and GV strains are found in cows and mice (Zheng et al., 2006).
Enteric caliciviruses, morphologically indistinguishable from the human NoVs, were first reported in association with calf diarrhea in the United Kingdom in 1978 and in Germany in 1980 (Almeida et al., 1978, Günther et al., 1984, Woode and Bridger, 1978). In recent years, genomic data has been obtained from the two viruses, Bo/Jena/80/DE and Bo/Newbury2/76/UK, and these strains were assigned into the prototype bovine NoVs (BNoVs) of genotype 1 (GIII-1) and genotype-2 (GIII-2) within GIII, respectively. Further isolates of the genotype 1 or genotype 2 BNoVs have been identified in the United Kingdom, The Netherlands, and United States (Liu et al., 1999, Oliver et al., 2003, Oliver et al., 2004, Smiley et al., 2003, van der Poel et al., 2003, Wise et al., 2004), which have confirmed the existence of the two distinct genotypes.
Although there are reports of the widespread occurrence of divergent human NoV strains, there are only a few reports of the prevalence and genetic diversity of BNoVs from England (Oliver et al., 2003, Woode and Bridger, 1978), Germany (Liu et al., 1999), The Netherlands (van der Poel et al., 2000, van der Poel et al., 2003) and the USA (Smiley et al., 2003, Wise et al., 2004). The aim of this study was to examine the prevalence and genetic diversity of BNoVs in diarrheic calves in South Korea.
2. Materials and methods
2.1. Specimens
From 2004 to 2005, a total of 645 fecal specimens obtained from diarrheic calves in 629 bovine beef herds were collected in South Korea during the spring (407 samples/406 herds), summer (107 samples/98 herds), autumn (73 samples/69 herds) and winter (58 samples/56 herds). The ages of the calves tested ranged from 2 to 90 days. The fecal samples were examined for common bacterial enteric pathogens including Salmonella spp., Clostridium spp., Campylobacter spp. using specific agar media, and any suspect colonies were identified using biochemical tests. The shiga-toxin-producing Escherichia coli was detected using a PCR assay, as described elsewhere (Asakura et al., 1998). For the viral assays, fecal suspensions of each sample were prepared immediately by diluting the feces 1:10 in 0.01 M phosphate-buffered saline (PBS), pH 7.2. The suspensions were then vortexed for 30 s, centrifuged (1200 × g for 20 min), and the supernatants were collected and stored at −80 °C for further testing.
2.2. RNA extraction
The RNA was extracted from a 200 μl starting volume of the centrifuged 10% fecal suspensions using the Trizol-LS (Gibco-BRL, Life Tech, Grand Island, USA) procedure. The total RNA recovered was suspended in 50 μl of RNase free water and stored at −80 °C until needed.
2.3. RT-PCR and nested PCR
The BNoV, bovine coronavirus (BCoV), groups A, B and C bovine rotaviruses (BRV A–C), bovine enteric Nebraska-like calicivirus (BEC-NBV), bovine torovirus (BToV) and bovine viral diarrhea virus (BVDV) were detected using standard one-step RT-PCR with different specific primer sets (Table 1 ) using the RNA extracted from the fecal samples, as described previously (Cho et al., 2001, Park et al., 2006). The mixture for RT-PCR was incubated at 42 °C for 60 min, preheated at 94 °C for 5 min, subjected to 35 cycles of 1 min at 94 °C, 1 min at the required temperature for each primer pair (Table 1), 2 min at 72 °C and a final 7 min incubation at 72 °C. Nested PCR assays with the primer pairs specific to BNoV, BCoV, BRV A, BToV and BEC-NBV were performed in order to increase the sensitivity and specificity of the RT-PCR as described previously (Cho et al., 2001, Park et al., 2006). The mixture for nested PCR was preheated at 94 °C for 5 min, subjected to 30 cycles of 1 min at 94 °C, 1 min at the required temperature for each primer pair (Table 1), 2 min at 72 °C and a final 7 min incubation at 72 °C. As a negative control, the RNA was extracted from the normal feces of a colostrums-deprived calf that had been inoculated with 50 ml sterile PBS. The amplification products were analyzed using 1.5% or 2% agarose gel electrophoresis and visualized by irradiating the ethidium bromide stained samples with UV.
Table 1.
RT-PCR and nested PCR primers for the detection of the bovine norovirus (BNoV), bovine enteric Nebraska-like calicivirus (BEC-NBV), bovine groups A, B and C rotaviruses (BRV A–C), bovine coronavirus (BCoV), bovine torovirus (BToV) and bovine viral diarrhea virus (BVDV) in the fecal specimens from the diarrheic calves
| Target viruses | Target genes | Sequence (5′ → 3′)a | Region | Product size (bp) | Annealing temp (°C) | Source or references |
|---|---|---|---|---|---|---|
| BNoV | RdRp | F: AGTTAYTTTTCCTTYTAYGGBGA | 4543–5074 | 532 | 54 | Smiley et al. (2003) |
| R: AGTGTCTCTGTCAGTCATCTTCAT | ||||||
| nF: GTCGACGGYCTKGTSTTCCT | 4690–5015 | 326 | 50 | Park et al. (2006) | ||
| nR: CACAGCGACAAATCATGAAA | ||||||
| BEC- | RdRp | F: TTTCTAACYTATGGGGAYGAYG | 4518-5066 | 549 | 52 | Smiley et al. (2003) |
| NBV | -MCP | R: GTCACTCATGTTTCCTTCTCTAAT | ||||
| nF: CGCTCCGTGTGGGATCACGA | 4788–4981 | 194 | 50 | Park et al. (2006) | ||
| nR: GCACGGGCTTCTTCTAGAGA | ||||||
| BRV A | VP7 | F: GCCTTTAAAAGCGAGAATTT | 3–1062 | 1060 | 44 | Chang et al. (1996) |
| R: GGTCACATCATACAAYTC TA | ||||||
| nF: TTTCTAACATCAACACT | 274–930 | 657 | 39 | Park et al. (2006) | ||
| nR: TTGCCACCATTTTTTCCAAT | ||||||
| BRV B | VP7 | F: GGAAATAATCAGAGATG | 1–795 | 795 | 42 | Barman et al. (2004) |
| R: CTACTCGTTTGGCTCCCTCC | ||||||
| BRV C | VP6 | F: TCAAGAAATGGWATGCAACC | 334–918 | 585 | 50 | Park et al. (2006) |
| R: CATAGCMGCTGGTCTWATCA | ||||||
| BCoV | N | F: GCAATCCAGTAGTAGAGCGT | 13–712 | 700 | 50 | Cho et al. (2001) |
| R: CTTAGTGGCATCCTTGCCAA | ||||||
| nF: GCCGATCAGTCCGACCAATG | 152–551 | 400 | 52 | Cho et al. (2001) | ||
| nR: AGAATGTCAGCCGGGGTAG | ||||||
| BToV | M | F: TTCTTACTACACTTTTTGGA | 98–700 | 603 | 49 | Park et al. (2006) |
| R: ACTCAAACTTAACACTAG AC | ||||||
| nF: TATGTACTATGTTTCCAGCT | 152–560 | 409 | 49 | Park et al. (2006) | ||
| nR: CCAACACAAATCCGCAACGC | ||||||
| BVDV | 5′-UTR | F: GCTAGCCATGCCCTTAG | 107–396 | 290 | 47 | Givens et al. (2003) |
| R: CCATGTGCCATGTACAG | ||||||
F: forward primer for RT-PCR; R: reverse primer for RT-PCR; nF: forward primer for nested PCR; nR: reverse primer for nested PCR.
2.4. DNA sequencing
The nested PCR products for a portion of the RdRp gene (326 bp) were selected from different test reactions and sequenced to verify the reaction specificity as well as to obtain the genomic data for phylogenetic analysis. The nested PCR products were purified using a GenClean II kit (BIO 101, Inc., LaJolla, USA) according to the manufacturer's instructions. DNA sequencing was carried out using an automated DNA sequencer (ABI system 3700, Applied Biosystem Inc., Foster City, USA).
2.5. Molecular analysis
The nucleotide (nt) and deduced amino acid (aa) sequences of the partial BNoV RdRp region were compared with those of other known caliciviruses using the DNA Basic module (DNAsis MAX, Alameda, USA) (Table 2 ). Phylogenetic analysis based on the nt and aa alignments were constructed using the neighbor-joining method and the unweighted-pair group method with the average linkages of Molecular Evolutionary Genetics Analysis (MEGA, Version 3.1) with a pairwise distance (Kumar et al., 2004). A sequence similarity search was carried out for the bovine calicivirus the RdRp protein using the LALIGN Query program of the GENESTREAM network server at Institut de Génétque Humaine, Montpellier, FRANCE (http://www.eng.uiowa.edu/∼tscheetz/sequence-analysis/examples/LALIGN/lalign-guess.html).
Table 2.
Name and its Genbank accession numbers of the reference norovirus strains and the other caliciviruses used in phylogenetic analysis
| G_Cluster | Name | Source | G_Cluster | Name | Source |
|---|---|---|---|---|---|
| G1_1 | NV/USA | M87661 | G3_2 | Bo/Dumfries/UK | AY126474 |
| G1_2 | SOV/UK | L07418 | G3_2 | Bo/Aberystwyth24/UK | AY126475 |
| G1_3 | DSV/USA | U04469 | G3_2 | Bo/MA8/KOR | DQ912788 |
| G1_4 | Chiba/JPN | AB042808 | G3_2 | Bo/MA60/KOR | DQ912787 |
| G1_5 | AB318/USA | AF414406 | G3_1 | Bo/MA88/KOR | DQ912789 |
| G1_6 | Hesse/DE | AF093797 | G3_2 | Bo/MA135/KOR | DQ912790 |
| G2_1 | Hawaii/USA | U07611 | G3_2 | Bo/MA140/KOR | DQ912791 |
| G2_2 | Msham/UK | X81879 | G3_2 | Bo/MA156/KOR | DQ912792 |
| G2_3 | Toronto/CAN | U02030 | G3_2 | Bo/MA242/KOR | DQ912793 |
| G2_3 | BB321/USA | AF414415 | G3_2 | Bo/MA259/KOR | DQ912794 |
| G2_4 | VA98387/USA | AY038600 | G3_2 | Bo/MA271/KOR | DQ912795 |
| G2_5 | MOH99/HUN | AF397156 | G3_2 | Bo/SA296/KOR | DQ912796 |
| G2_6 | SU17/JPN | AB039779 | G3_2 | Bo/MA302/KOR | DQ912797 |
| G2_7 | GN273/USA | AF414409 | G3_2 | Bo/MA421/KOR | DQ912798 |
| G2_11 | Sw43/JPN | AB074892 | G4_1 | FLD560/USA | AF414426 |
| G2_19 | QW218/USA | AY823307 | G4_1 | SCD624/ USA | AF414427 |
| G3_1 | Bo/Jena/DE | AJ011099 | G5 | Murine1/USA | AY228235 |
| G3_2 | Bo/Newbury2/UK | AF097917 | Human sapovirus | SaV | U65427 |
| G3_2 | Bo/CV95-OH/USA | AF542083 | NB-like virus | NBV | AY082891 |
| G3_2 | Bo/CV186-OH/USA | AF542084 | Lagovirus | RHDV | M67473 |
| G3_2 | Bo/Penrith55/UK | AY126476 | Vesivirus | FCV | M86379 |
3. Results
3.1. Prevalence of BNoVs in calves with diarrhea in South Korea
Eighteen (2.8%) out of 645 fecal samples (18/629 herds; 2.9%) tested positive using a 1-step RT-PCR assay, targeting a 532 bp fragment of the RdRp region of the BNoV. Nested PCR, targeting a 326 bp fragment of the RdRp region of the BNoV, detected 60 (9.3%) positive fecal samples from 60 herds (9.4%).
3.2. Other enteric pathogens
Of the 60 BNoV-positive fecal specimens from 60 calf herds with diarrhea, 22 fecal samples (3.4%) from 22 herds (3.5%) tested positive for the BNoV alone, while the 38 fecal samples (5.9%) from the other 38 herds (5.9%) also tested positive for other enteric pathogens (Table 3 ). Of the concurrent infections of the BNoV with the other enteric pathogens, BRV A was the most common [15 fecal samples (2.2%) from 15 herds (2.3%)]. Interestingly, one diarrheic fecal sample tested positive for four enteric pathogens including BNoV, BEC-NBV, BRV A and BVDV. In addition, 397 fecal specimens from 395 herds that tested negative for BNoV also tested positive for other enteric pathogens (Table 3). No enteric pathogens were detected in 189 fecal samples from 175 herds.
Table 3.
Summary of the enteric pathogens present in diarrheic fecal specimens obtained from calves
| Enteric pathogens presenta | No. of farms (%)b | No. of calves (%)c |
|---|---|---|
| BNoV alone | 22 (3.5) | 22 (3.4) |
| BNoV plus BEC-NBV | 4 (0.6) | 4 (0.6) |
| BNoV plus BRV A | 15 (2.4) | 15 (2.3) |
| BNoV plus BRV B | 1 (0.2) | 1 (0.2) |
| BNoV plus BRV C | 3 (0.5) | 3 (0.5) |
| BNoV plus BCoV | 1 (0.2) | 1 (0.2) |
| BNoV plus BToV | 3 (0.5) | 3 (0.5) |
| BNoV plus BVDV | 3 (0.5) | 3 (0.5) |
| BNoV plus BRV A, BRV C | 1 (0.2) | 1 (0.2) |
| BNoV plus BRV A, BVDV | 2 (0.3) | 2 (0.3) |
| BNoV plus, BRV C, BcoV | 1 (0.2) | 1 (0.2) |
| BNoV plus BcoV, BVDV | 1 (0.2) | 1 (0.2) |
| BNoV plus BToV, BVDV | 1 (0.2) | 1 (0.2) |
| BNoV plus BEC-NBV, BRV A, BVDV | 1 (0.2) | 1 (0.2) |
| BNoV plus BRV C, E. coli (STX) | 1 (0.2) | 1 (0.2) |
| Other enteric pathogens detected | 395 (62.8) | 397 (61.6) |
| No enteric pathogens detected | 175 (27.8) | 189 (29.3) |
| Total | 629 (100) | 645 (100) |
BNoV: bovine norovirus; BEC-NBV: bovine enteric Nebraska-like calicivirus; BRV A–C: groups A–C bovine rotaviruses; BCoV: bovine coronavirus; BToV: bovine torovirus; BVDV: bovine viral diarrhea virus.
Number of positive herds.
Number of positive calves.
3.3. Genetic diversity of BNoVs
The genetic diversity of the BNoVs and their genetic similarity to the other genogroup strains were investigated by sequencing 326 bp of the nt 4690 to nt 5015 RdRp region from the 12 BNoVs detected from the diarrheic fecal samples. The alignment indicated that the Korean Bo/MA88/KOR strain clustered with the GIII-1 prototype, the Bo/Jena/DE strain, whereas the remaining 11 Korean BNoVs clustered with the GIII-2 prototype, the Bo/Newbury2/UK strain (Fig. 1 ). Among the 11 Korean BNoVs, 10 Korean BNoVs clustered together on a separate branch within GIII-2, whereas the Bo/MA156/KOR, which belongs to GIII-2, clustered with the Bo/Dumfries/UK strain (Fig. 1). Among the BNoVs examined, the Korean Bo/MA88/KOR strain had the highest nt (86.8%) and aa (99.1%) identity with the GIII-1 prototype, the Bo/Jena/DE strain, but shared 75.2–78.2% nt and 85.2–90.7% aa identity with the GIII-2 BNoVs. The remaining 11 Korean BNoVs shared a higher nt (88.0–90.5%) and aa (93.5–99.1%) identity with the GIII-2 prototype, Bo/Newbery2/UK strain, than those from the GIII-1 prototype, Bo/Jena/DE strain, sharing a 73.6–79.1% nt and 84.3–89.8% aa identity. In addition, the GIII-1 BNoV strains, one Korean strain Bo/MA88/KOR and one German strain (Bo/Jena/DE; Liu et al., 1999), showed a low aa identity with the human sapovirus (SaV) (24.1%), BEC-NBV (18.5%), rabbit hemorrhagic disease virus (RHDV) (21.3%), and feline calicivirus (FCV) (27.8%) strains, whereas there was slightly higher aa identity with the human GI, GII and GIV NoV (56.5–72.2%), porcine GII-2 NoV (59.3–60.2%), and murine GV NoV (50.0–50.9%) strains. The GIII-2 strains (Bo/MA88/KOR and Bo/Jena/DE) also had a low aa identity with the human SaV (22.2–23.1%), BEC-NBV (17.6–18.5%), RHDV (22.2–23.1%), and FCV (25.9%) strains, whereas there was slightly higher aa identity with the human GI, GII and GIV NoV (53.7–75.0%), porcine GII-2 NoV (56.5–60.2%), and murine GV NoV (52.8–54.6%) strains.
Fig. 1.
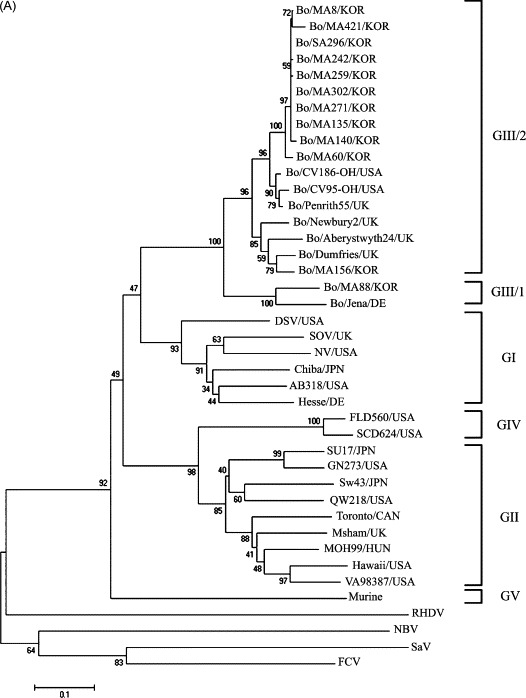
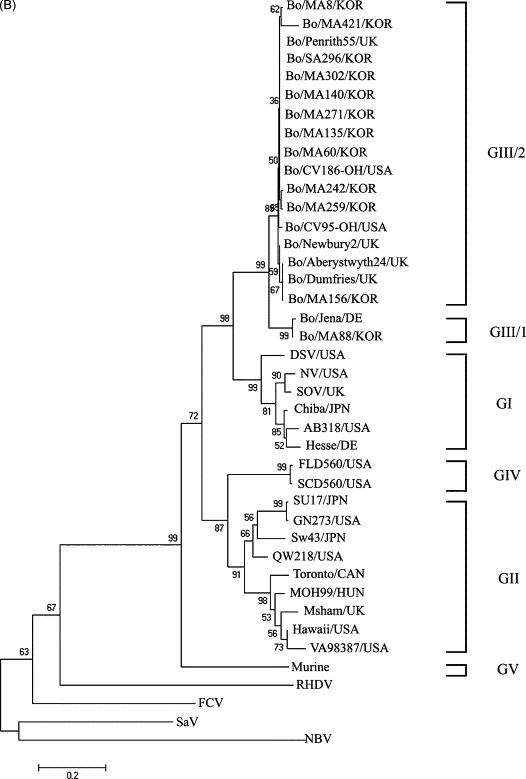
The phylogenetic trees of the partial nucleotide (A) and amino acid (B) sequences of the RNA dependent RNA polymerase of caliciviruses were constructed using the neighbor-joining method of Molecular Evolutionary Genetics Analysis (Kumar et al., 2004). Table 3 gives the name and GenBank accession numbers of each virus.
4. Discussion
Based on a comparison of the aa and nt sequences of the partial RdRp region, the 12 BNoV strains that was detected in diarrheic fecal samples from calves in South Korea were classified as Norovirus GIII/1 (Bo/Jena/DE strain-like, 1 strain) and GIII/2 (Bo/Newbury2/UK strain-like, 11 strains). These results highlight the existence of the two distinct genotypes within the group and confirming the genotype 2 as the main genotype (Oliver et al., 2003, Smiley et al., 2003, van der Poel et al., 2000, van der Poel et al., 2003, Wise et al., 2004). The phylogenetic data for the nt and aa sequences of the partial RdRp region also support the findings (Oliver et al., 2003, Smiley et al., 2003, van der Poel et al., 2000, van der Poel et al., 2003, Wise et al., 2004) that the Korean BNoVs were classified as Norovirus GIII with the main genotype being GIII/2. These results, along with other reports (Oliver et al., 2003, Smiley et al., 2003, van der Poel et al., 2000, van der Poel et al., 2003, Wise et al., 2004), suggest GIII/2 to be the dominant genotype of BNoVs.
The aa identity within the GIII/2 BNoVs ranged from 93.5% to 99.1% (88.0–90.5% nt identity), in which genetic divergence was observed not only in the Korean BNoVs but also between the Korean BNoVs and BNoVs reported in the other countries, which indicates the genetic variability in the BNoVs belonging to GIII/2. This aa identity is similar to findings reported by Oliver et al. (2003) and Wise et al. (2004) who showed that the aa identity within the GIII BNoVs in England and USA shared a 92–100% and 91.2–100% homology with the other known strain in the partial RdRp region. Like the other RNA viruses, the NoVs show broad genomic sequence diversity between the circulating strains, which is partly due to the poor template fidelity of their RNA polymerases (Zheng et al., 2006). Therefore, it is expected that more genetic clusters will be identigied within GIII in the future when more sequence data of the BNoVs circulating in cattle becomes available.
The fecal prevalence of BNoV infections in diarrheic calves have been reported to be 8.0% in England (Oliver et al., 2003), 31.1% in The Netherlands, and 72.0% in the Ohio, USA (Smiley et al., 2003). Based on the nested PCR assay, the BNoVs were detected in 9.3% of the fecal samples examined in this study. This suggests that BNoV infection are endemic in diarrheic calves in South Korea in a similar manner to the prevalence of BNoV infections in England (Oliver et al., 2003).
Although it was reported that BNoVs induce diarrhea in experimental calves (Bridger et al., 1984), the pathogenicity and mechanisms of BNoVs infections are largely obscure. Therefore, other bovine enteric pathogens might play a role in the clinical and pathological presentation of this disease because many other enteric viruses have been found to be associated with diarrhea in cattle and cause lesions of villous atrophy (BRVs, BCoVs, BToVs, BEC-NBV, etc.). In this study, 5.9% of the diarrheic fecal samples testing positive to the BNoVs also tested positive to other pathogens including the BEC-NBV, BRVs A, B and C, BCoV, BToV, BVDV and shiga-toxin-producing E. coli. This suggests that a number of enteric pathogens, either singly or in combination, can influence the clinical course of BNoV infections (Almeida et al., 1978, Reynolds et al., 1986, Woode and Bridger, 1978).
In summary, this study identified the existence of two distinct genotypes of BNoVs within the GIII, and confirmed the GIII/2 BNoVs to be the principal genotype, which is analogous to the previously reported BNoV genogroup and genotypes. In addition, these BNoVs cause endemic infections in diarrheic calves in South Korea.
Acknowledgments
This study was supported by the National Veterinary Research and Quarantine Service (NVRQS), Ministry of Agriculture and Forestry, Korea Health 21 R&D (01-PJ10-PG6-01GM02-002) by the Ministry of Health and Welfare, and the Regional Technology Innovation Program (RTI05-01-01) of the Ministry of Commerce, Industry and Energy (MOCIE), Republic of Korea. The authors acknowledge a graduate fellowship provided by the Korean Ministry of Education and Human Resources Development through the Brain Korea 21 project.
References
- Almeida J.D., Craig C.R., Hall T.E. Multiple viruses present in the faeces of a scouring calf. Vet. Rec. 1978;102:170–171. doi: 10.1136/vr.102.8.170. [DOI] [PubMed] [Google Scholar]
- Asakura H., Makino S., Shirahata T., Tsukamoto T., Kurazono H., Ikeda T., Takeshi K. Detection and genetical characterization of Shiga toxin-producing Escherichia coli from wild deer. Microbiol Immunol. 1998;42:815–822. doi: 10.1111/j.1348-0421.1998.tb02356.x. [DOI] [PubMed] [Google Scholar]
- Barman P., Ghosh S., Das S., Varghese V., Chaudhuri S., Sarkar S., Krishnan T., Bhattacharya S.K., Chakrabarti A., Kobayashi N., Naik T.N. Sequencing and sequence analysis of VP7 and NSP5 genes reveal emergence of a new genotype of bovine group B rotavirus in India. J. Clin. Microbiol. 2004;42:1818–2816. doi: 10.1128/JCM.42.6.2816-2818.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bridger J.C., Hall G.A., Brown J.F. Characterization of calici-like virus (Newbury agent) found in association with astrovirus in bovine diarrhea. Infect. Immun. 1984;43:133–138. doi: 10.1128/iai.43.1.133-138.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang K.O., Parawani A.V., Saif L.J. The characterization of VP7(G type) and VP4(P type) genes of bovine group A rotavirus from field samples using RT-PCR and RFLP analysis. Arch. Virol. 1996;141:1727–1739. doi: 10.1007/BF01718295. [DOI] [PubMed] [Google Scholar]
- Cho K.O., Hasoksuz M., Nielsen P.R., Chang K.O., Lathrop S., Saif L.J. Cross-protection studies between respiratory and calf diarrhea and winter dysentery coronavirus strains in calves and RT-PCR and nested PCR for their detection. Arch. Virol. 2001;146:2401–2419. doi: 10.1007/s007050170011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Givens M.D., Heath A.M., Carson R.L., Brock K.V., Edens M.S., Wenzel J.G., Stringfellow D.A. Analytical sensitivity of assays used for detection of bovine viral diarrhea virus in semen samples from the Southeastern United States. Vet. Microbiol. 2003;96:145–155. doi: 10.1016/s0378-1135(03)00213-x. [DOI] [PubMed] [Google Scholar]
- Green K.Y., Ando T., Balayan M.S., Berke T., Clarke I.N., Estes M.K., Matson D.O., Nakata S., Neill J.D., Studdert M.J., Thiel H.J. Taxonomy of the caliciviruses. J. Infect. Dis. 2000;18:S322–S330. doi: 10.1086/315591. [DOI] [PubMed] [Google Scholar]
- Günther H., Otto P., Heilman P. Studies into diarrhoea of young calves. Sixth communication: detection and determination of pathogenicity of a bovine corona virus and an undefined icosahedric virus. Arch. Exp. Vet. Med. Leipzig. 1984;38:781–792. [PubMed] [Google Scholar]
- Jiang X., Wang M., Wang K., Estes M.K. Sequence and genomic organization of Norwalk virus. Virology. 1993;195:51–61. doi: 10.1006/viro.1993.1345. [DOI] [PubMed] [Google Scholar]
- Kumar S., Tamura K., Nei M. MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform. 2004;5:150–163. doi: 10.1093/bib/5.2.150. [DOI] [PubMed] [Google Scholar]
- Lambden P.R., Caul E.O., Ashley C.R., Clarke I.N. Sequence and genome organization of a human small round-structured (Norwalk-like) virus. Science. 1993;259:516–519. doi: 10.1126/science.8380940. [DOI] [PubMed] [Google Scholar]
- Liu B.L., Lambden P.R., Gunther H., Otto P., Elschner M., Clarke I.N. Molecular characterization of a bovine enteric calicivirus: relationship to the Norwalk-like viruses. J. Virol. 1999;73:819–825. doi: 10.1128/jvi.73.1.819-825.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oliver S.L., Asobayire E., Dastjerdi A.M., Bridger J.C. Genomic characterization of the unclassified bovine enteric virus Newbury agent-1 (Newbury1) endorses a new genus in the family Caliciviridae. Virology. 2006;350:240–250. doi: 10.1016/j.virol.2006.02.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oliver S.L., Brown D.W., Green J., Bridger J.C. A chimeric bovine enteric calicivirus: evidence for genomic recombination in genogroup III of the Norovirus genus of the Caliciviridae. Virology. 2004;326:231–239. doi: 10.1016/j.virol.2004.06.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oliver S.L., Dastjerdi A.M., Wong S., El Attar L., Gallimore C., Brown D.W., Green J., Bridger J.C. Molecular characterization of bovine enteric caliciviruses: a distinct third genogroup of noroviruses (Norwalk-like viruses) unlikely to be of risk to humans. J. Virol. 2003;77:2789–2798. doi: 10.1128/JVI.77.4.2789-2798.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park S.J., Jeong C., Yoon S.S., Choy H.E., Saif L.J., Park S.H., Kim Y.J., Jeong J.H., Park S.I., Kim H.H., Lee B.J., Cho H.S., Kim S.K., Kang M.I., Cho K.O. Detection and characterization of bovine coronaviruses in fecal specimens of adult cattle with diarrhea during the warmer seasons. J. Clin. Microbiol. 2006;44:3178–3188. doi: 10.1128/JCM.02667-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reynolds D.J., Morgan J.H., Chanter N., Jones P.W., Bridger J.C., Debney T.G., Bunch K.J. Microbiology of calf diarrhea in southern Britain. Vet. Rec. 1986;12:34–39. doi: 10.1136/vr.119.2.34. [DOI] [PubMed] [Google Scholar]
- Smiley J.R., Hoet A.E., Traven M., Tsunemitsu H., Saif L.J. Reverse transcription-PCR assays for detection of bovine enteric caliciviruses (BEC) and analysis of the genetic relationships among BEC and human caliciviruses. J. Clin. Microbiol. 2003;41:3089–3099. doi: 10.1128/JCM.41.7.3089-3099.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smiley J.R., Chang K.O., Hayes J., Vinje J., Saif L.J. Characterization of an enteropathogenic bovine calicivirus representing a potentially new calicivirus genus. J. Virol. 2002;76:10089–10098. doi: 10.1128/JVI.76.20.10089-10098.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Poel W.H., van der Heide R., Verschoor F., Gelderblom H., Vinje J., Koopmans M.P. Epidemiology of Norwalk-like virus infections in cattle in The Netherlands. Vet. Microbiol. 2003;92:297–309. doi: 10.1016/s0378-1135(02)00421-2. [DOI] [PubMed] [Google Scholar]
- van der Poel W.H., Vinje J., van Der Heide R., Herrera M.I., Vivo A., Koopmans M.P. Norwalk-like calicivirus genes in farm animals. Emerg. Infect. Dis. 2000;6:36–41. doi: 10.3201/eid0601.000106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wise A.G., Monroe S.S., Hanson L.E., Grooms D.L., Sockett D., Maes R.K. Molecular characterization of noroviruses detected in diarrheic stools of Michigan and Wisconsin dairy calves: circulation of two distinct subgroups. Virus Res. 2004;100:165–177. doi: 10.1016/j.virusres.2003.11.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woode G.N., Bridger J.C. Isolation of small viruses resembling astroviruses and caliciviruses from acute enteritis of calves. J. Med. Microbiol. 1978;11:441–452. doi: 10.1099/00222615-11-4-441. [DOI] [PubMed] [Google Scholar]
- Zheng D., Ando T., Fankhauser R.l., Beard R.S., Glass R.I., Monroe S.S. Norovirus classification and proposed strain nomenclature. Virology. 2006;346:312–323. doi: 10.1016/j.virol.2005.11.015. [DOI] [PubMed] [Google Scholar]


