Abstract
Background
Mesenchymal stem/stromal cells (MSCs) are multipotent cells with a promising application potential in regenerative medicine and immunomodulation. However, MSCs cultured in vitro exhibit functional heterogeneity. The underlying molecular mechanisms that define MSC heterogeneity remain unclear.
Methods
We investigated the gene expression profile via single-cell RNA sequencing (scRNA-seq) of human primary Wharton’s jelly-derived MSCs (WJMSCs) cultured in vitro from three donors. We also isolated CD142+ and CD142− WJMSCs based on scRNA-seq data and compared their proliferation capacity and “wound healing” potential in vitro. Meanwhile, we analyzed publicly available adipose-derived MSC (ADMSCs) scRNA-seq data and performed transcriptome comparison between WJMSCs and ADMSCs at the single-cell level.
Results
GO enrichment analysis of highly variable genes (HVGs) obtained from WJMSCs revealed that these genes are significantly enriched in extracellular region with binding function, involved in developmental process, signal transduction, cell proliferation, etc. Pathway analysis showed that these HVGs are associated with functional characteristics of classic MSCs, such as inflammation mediated by chemokine and cytokine signaling, integrin signaling, and angiogenesis. After regressing out the batch and cell cycle effects, these HVGs were used for dimension reduction and clustering analysis to identify candidate subpopulations. Differentially expressed gene analysis revealed the existence of several distinct subpopulations of MSCs that exhibit diverse functional characteristics related to proliferation, development, and inflammation response. In line with our data, sorted CD142+ and CD142− WJMSCs showed distinct proliferation capacity as well as “wound healing” potential. Although WJMSCs and ADMSCs were derived from different tissues and were displaying different differentiation potencies, their HVGs were largely overlapped and had similar functional enrichment.
Conclusion
HVGs identified in MSCs are associated with classic MSC function. Regarding therapeutic potential, these genes are associated with functional characteristics, on which the MSC clinical application were theoretically based, such as development and inflammation response. Altogether, these HVGs hold the potential to be used as candidate markers for further potency association studies.
Keywords: MSCs, scRNA-seq, Highly variable genes, Functional enrichment, Potency
Background
Mesenchymal stem/stromal cells (MSCs) are multipotent with self-renewal capacity and can be derived from various tissues, including the bone marrow [1], adipose tissue [2], umbilical cord [3, 4], and placenta [5]. Besides the multi-lineage potential to differentiate into various cell types, such as chondrocytes, osteocytes, adipocytes, myocytes, and neuronal cells [6, 7], MSCs could modulate immune cell response via interaction with lymphocytes from both the innate and adaptive immune system to deliver immunosuppressive and anti-inflammatory effects after homing to sites of inflammation in vivo [8, 9]. Furthermore, human MSCs could be cultured in large scale and have minimal functional loss after long-term cryopreservation [10, 11]. Therefore, MSCs demonstrate promising utilization potential and are ideal cell types in both fundamental and translational biology fields, such as developmental biology, cellular therapy, immunomodulation, and regenerative medicine [12, 13]. Currently, more than 700 clinical trials have been registered in ClinicalTrials.gov (http://www.clinicaltrials.gov), which utilize MSCs for cellular therapy. Transplantation of MSCs demonstrates no obvious adverse effect, regardless of allogeneic or autologous cell origin, and has been extensively explored in treatment of various disease types, such as bone and cartilage defects [14, 15], cardiovascular disease [16, 17], neurological degeneration [18, 19], liver disorder [20], and immunological diseases [21, 22] with encouraging clinical outcomes. Several MSC-based products have been approved or conditionally approved in certain country or district to treat disorders, such as graft versus host disease (GvHD) and Crohn’s related enterocutaneous fistular disease [23].
The minimal criteria for defining multipotent MSCs were published by the International Society for Cellular Therapy (ISCT) in 2006 [24], which is widely accepted and adopted in both basic research and industrial application. However, it only defines basic morphological and functional characteristic. More and more research works have recognized that MSC populations exhibit tissue-to-tissue functional variation [25, 26], as well as inter-population heterogeneity when using current markers to define MSCs, which makes it difficult to predict cell population dynamics and functional alterations after extended culture or exposed to extrinsic factors [27, 28]. Functional heterogeneity coupled with large-scale expansion in clinical manufacturing process may explain, in part, why data across MSC-based clinical trials are largely incongruent [29]. During MSC culturing and passaging, the competition and balance between different subpopulations may change, resulting in decrease in proportion or even loss of certain subpopulations and ultimately leading to alterations in cell function and treatment outcomes in clinical studies [30, 31]. Therefore, there is an urgent need to elucidate whether a certain MSC subtype or a cocktail of defined population of different subtypes can demonstrate effectiveness during cellular therapy and tissue engineering. Investigation into the underlying molecular mechanisms that define MSC heterogeneity will facilitate subtype identification and improve methods for cell isolation and expansion in vitro. By in-depth analysis of cell quality attributes, it will also help to interpret the results from clinical trials and eventually improve clinical efficacy of MSC products.
Recently, single-cell RNA sequencing (scRNA-seq) technology, which allows massive parallel analysis of gene expression profiles at single-cell level, has become a powerful tool in investigating tissue and cell heterogeneity. It provides unprecedented opportunities for identifying subpopulations that share a common gene expression profile in a heterogeneous cell population [32]. Here, we investigated the gene expression profile via scRNA-seq of human primary WJMSCs cultured in vitro from three donors. In contrast to other source-derived MSCs, WJMSCs, which isolated from previously discarded umbilical cord tissues, bear higher proliferation rate and the strongest immunomodulatory effect, making them an attractive alternative source of MSCs for clinical research and application [33, 34]. Meanwhile, we analyzed publicly available ADMSC scRNA-seq data [35] and performed transcriptome comparison between WJMSCs and ADMSCs at the single-cell level.
Methods
Cell isolation and culture
Human umbilical cord tissues were collected from naturally delivered full-term newborns (n = 3, two females and one male). WJs were isolated from the umbilical cord after dissection and mechanically dissociated into tissue explants of approximately 2 mm2, which were then seeded into T75 flasks and cultured in UltraCULTURE™ Serum-free Medium (LONZA) at 37 °C with 5% CO2 in a humidified atmosphere. After the cell density reached about 80% confluence, cells were dissociated with TrypLE™ Select (Thermo Fisher Scientific) incubated at 37 °C for 5 min. The collected cells were immediately used for single-cell library construction, sub-cultured into a new culture dish, tri-lineage differentiation potency test, or freezing in liquid nitrogen for long-term banking.
scRNA-seq and analysis
scRNA-seq experiment was performed using the Chromium Single Cell Gene Expression Solution, V2 Chemistry (10x Genomics), following the manufacturer’s protocol. Briefly, the collected cells were washed with PBS twice and resuspended in 500 μl PBS, targeting the required 500 cells/μl concentration. We pipetted 6.4 μl cell suspension, targeting the recovery of about 2000 cells per sample. Single-cell RNA-seq libraries were obtained following the 10x Genomics recommended protocol, using the reagents included in the Chromium Single Cell 3′ v2 Reagent Kit. Libraries were sequenced on the BGISEQ-500 (BGI) instrument [36] using 26 cycles (cell barcode and UMI [37]) for read1 and 108 cycles (sample index and transcript 3′ end) for read2.
scRNA-seq data analysis was available in (Additional file 1) for details. Briefly, the scRNA-seq data was processed using cellranger-2.0.0 for mapping, outlier cells using the median absolute deviation from the median total library size (logarithmic scale), and total gene numbers (logarithmic scale), as well as mitochondrial percentage, as implemented in scran package, using a cutoff of 3 [38]. Any gene expressed across all the cells by average UMI less than 0.1 was removed. Cell cycle phase assignment and confound effect removal, highly variable gene identification, linear and nonlinear dimension reduction, and clustering and differential expression analysis were performed using the Seurat package [39].
Bulk RNA-seq and analysis
Total mRNA was extracted using the TRIzol (Invitrogen) reagent, as described previously [40]. All protocols for BGISEQ-500 library construction, preparation, sequencing, and quality control were provided by BGI.
Reads were mapped to the human genome (GRCh38) using HISAT2 with default parameters [41]. Raw counts of sequencing reads for the feature of genes were extracted by featureCounts [42]. Transcripts per million (TPM) [43] was used to normalize for sequencing depth and gene length for relative quantity, and then we ranked the gene abundance according to the TPM values. SRA files of bulk RNA-seq data of MSCs derived from adipose tissues (GEO: GSE76081) were downloaded from NCBI, and the same processing pipeline was used for read mapping and normalization.
Functional enrichment analysis
GO-slim and protein class and pathway overrepresentation test enrichment analyses were performed using PANTHER™ Version 14.1 according to Mi et al. [44] via a test type of Fisher’s exact, applying the Benjamini–Hochberg false discovery rate (FDR) correction for multiple testing.
Lineage differentiation potency evaluation
Using the marker genes listed in (Additional file 2: Table S1), we calculated the osteogenic, adipogenic, chondrogenic, neurogenic, and myogenic “scores” according to [45]. Specifically, the score was defined as a single numeric value representative of the expression of multiple marker genes and the sum of log normalized expression across all markers in a category. Housekeeping genes were also used and named as “housekeeping score.”
Tri-lineage differentiation
For osteogenic differentiation, MSCs were seeded into a 24-well plate at the density of 5 × 103 cells/well. When the cells reached 70% confluency, the medium was replaced with osteogenic differentiation medium (MEM-alpha medium (Thermo Fisher Scientific), 10% FBS (Hyclone), 1% Pen-Strep (Thermo Fisher Scientific), 100 nM dexamethasone (Sigma), 10 mM sodium β-glycero phosphate (Sigma), 0.05 mM ascorbic acid (Sigma)) and kept for 3 weeks. To assess the osteogenic differentiation, Alizarin Red S staining (Sigma) was performed for the calcium-rich extracellular matrix.
For adipogenic differentiation, 1 × 104 cells were seeded per well. The cells at confluence were then treated with adipogenic differentiation medium (high-glucose DMEM (Thermo Fisher Scientific), 10% FBS (Hyclone), 0.5 mM IBMX (Sigma), 1 μM dexamethasone (Sigma), 1.7 μM insulin (Sigma), 0.2 mM indomethacin (Sigma)) for 3 weeks. The cells were fixed with 4% formaldehyde solution, and lipid droplets of the resultant differentiated cells were detected using Oil Red O staining (Sigma).
For chondrogenic differentiation, the StemPro™ Chondrogenesis Differentiation Kit was used and performed according to the manufacturer protocol. Briefly, 1.6 × 107 cells were resuspended in MSC medium. Micromass cultures were generated by seeding 5-μL droplets of cell solution in the center of 96-well plate wells. After cultivating micromass cultures for 2 h under high humidity conditions, we added warmed chondrogenesis media to culture vessels and incubated them in 37 °C incubator with 5% CO2. After 3 weeks, the cells were fixed with 4% formaldehyde solution and stained with Alcian Blue (Sigma).
FACS-based cell sorting
For CD142− and CD142+ cells isolation, MSCs were stained for 30 min on ice with PE mouse anti-human CD142 antibody (BD Biosciences). PE mouse IgG1 and κ Isotype Control (BD Biosciences) were used as an isotype control. Then, the stained cells were washed, analyzed, and sorted on a BD FACSAria II (BD Biosciences).
Real-time quantitative polymerase chain reaction (real-time qPCR)
Total RNA was isolated using TRIzol™ Reagent (Invitrogen). PrimeScript™ RT reagent Kit with gDNA Eraser (Perfect Real Time) (TAKARA) and TB Green® Premix Ex Taq™ II (Tli RNAseH Plus) (TAKARA) were used for cDNA synthesis and qPCR according to manufacturer’s protocol, respectively. Applied Biosystems StepOnePlus Real-Time PCR System (Thermo Fisher Scientific) was used for data collection and analysis. Data were representative of three independent experiments with two replicates for each and normalized to inner reference GAPDH. The primers for real-time qPCR are listed in Table 1.
Table 1.
List of primers for real-time qPCR
| Name | Forward primer (5′-3′) | Reverse primer (5′-3′) | Size (bp) |
|---|---|---|---|
| TGFB1 | CAATTCCTGGCGATACCTCAG | GCACAACTCCGGTGACATCAA | 86 |
| SPARC | AGCACCCCATTGACGGGTA | GGTCACAGGTCTCGAAAAAGC | 105 |
| COL4A1 | GGGATGCTGTTGAAAGGTGAA | GGTGGTCCGGTAAATCCTGG | 113 |
| COL1A1 | GTGCGATGACGTGATCTGTGA | CGGTGGTTTCTTGGTCGGT | 119 |
| COL5A1 | TACCCTGCGTCTGCATTTCC | GCTCGTTGTAGATGGAGACCA | 97 |
| CCL2 | GATCTCAGTGCAGAGGCTCG | TGCTTGTCCAGGTGGTCCAT | 153 |
| CXCL8 | ACTGAGAGTGATTGAGAGTGGAC | AACCCTCTGCACCCAGTTTTC | 112 |
| MKI67 | GCCTGCTCGACCCTACAGA | GCTTGTCAACTGCGGTTGC | 127 |
| GAPDH | CTGGGCTACACTGAGCACC | AAGTGGTCGTTGAGGGCAATG | 101 |
Wound healing assay
Wound healing assays were performed as previously described [46]. Briefly, when sorted MSC cultures were approximately 70% confluent, serum-free DMEM/F12 medium supplemented with insulin, transferring, selenium (ITS-X, Invitrogen), and antibiotics were used to substitute the MSC medium. After 72 h, the conditioned medium was collected and filtered (0.2 μ filter) to remove all cellular debris. Human skin fibroblasts were seeded into 24-well plates and cultured at 37 °C and 5% CO2 to form a confluent monolayer. Sterile pipette tips were used to scratch a “wound field.” Then, the conditioned medium was added to the culture. Wound closure was monitored by collecting digitized images (Olympus CKX41) after the scratch was performed. ImageJ was used to obtain the wound area for each image. Data were presented as percentage of wound area compared to the original scratch for each given time point.
Results
Overview of the single-cell RNA sequencing data
To investigate into inter-population heterogeneity in primary cultured WJMSCs at the single-cell transcriptome level, primary cells isolated from three human umbilical cords (named as UC1, UC2, and UC3, respectively) were collected and used for scRNA-seq. A total of about 5 × 108 raw reads with high quality for each donor was obtained (Additional file 3: Figure S1A). Mapping these reads to human GRCh38 genome, an average of about 56.90% and 61.03% reads was mapped confidently to the transcriptome and exonic regions, respectively (Additional file 3: Figure S1B, C). Briefly, a total of 6878 cells (filtered matrix) was obtained from the three donors, with an average of 2293 cells for each, with 209,769 mean reads, 38,983 median unique molecular identifier (UMI) counts, and 6279 median genes per cell (Additional file 3: Figure S1D-F), suggesting that our data were of high quality.
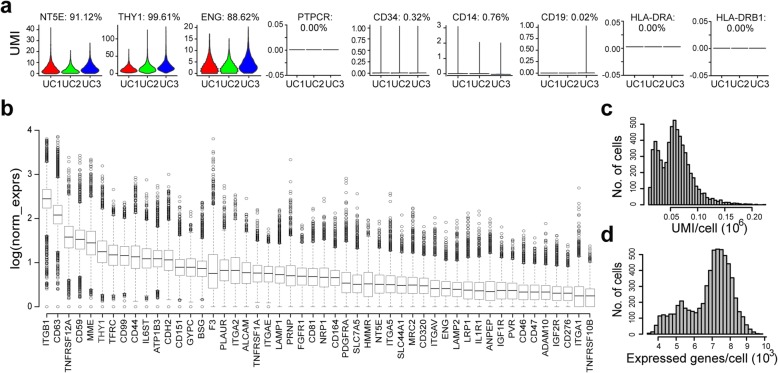
According to the minimal criteria proposed by the International Society for Cellular Therapy in 2006 [24], MSCs must express three positive markers, i.e., CD105 (ENG), CD73 (NT5E), and CD90 (THY1), and lack expression of several negative genes, including CD45 (PTPRC), CD34, CD14 or CD11b (ITGAM), CD79a (CD79A) or CD19, and HLA-DR (HLA-DRA and HLA-DRB1). When we looked at the expression of those markers in our raw data, we saw the expression of those positive markers (UMI > 0), while negative genes were not expressed (UMI = 0) in most cells (Fig. 1a). Next, we ranked cluster of differentiation (CD) genes by average normalized expression or percentage of cells with at least one UMI across all cells (Additional file 2: Table S2 and Table S3). Classic cell surface markers for MSC definition, including ENG, NT5E, and THY1, as expected, belong to the top 50 highly expressed CDs (Fig. 1b). Among the CDs, integrins, such as ITGB1, ITGA1, ITGA2, and ITGA5, which play important roles in MSC morphology, migration, proliferation, differentiation, and survival [47–49], are also highly expressed in WJMSC population (Fig. 1b and Additional file 2: Table S2). To strengthen and validate our scRNA-seq data, we found that most top 100 CDs ranked by average normalized expression in scRNA-seq are also highly expressed in bulk RNA-seq data, including MSC positive markers ENG, THY1, and NT5E (Additional file 2: Table S2). In addition, we assayed the tri-lineage capability of the cultured WJMSCs for scRNA-seq, and the results confirmed that they have the potency to differentiate into osteoblasts, adipocytes, and chondroblasts in vitro (Additional file 3: Figure S1G).
Fig. 1.
Overview of WJMSCs single-cell RNA-seq data. a Expression of marker genes in the three samples. Number on the top showing percentage of cells with at least one UMI. b Boxplot showing top 50 cluster of differentiation (CD) genes ranked by average normalized expression. c Distribution of UMI cross cells after pre-processing to filter out low-quality cells. d Distribution of expressed genes after pre-processing to filter out low-abundance genes with mean-based method (genes with means more than 0.1 were retained)
For further analysis, we filtered the outlier cells using the median absolute deviation from the median total library size (logarithmic scale) and total gene numbers (logarithmic scale), as well as mitochondrial percentage for each donor [38]. Totally, 702 outlier cells were removed and 6176 single cells were kept by median absolute deviation method. Considering none or low abundant expressed genes across cells, we also integrated these three data together and removed any gene with an average expression less than 0.1 UMI. Finally, 6176 high-quality single cells with 11,458 expressed genes were passed on to downstream analysis. Across the cells, the number of UMI per cell ranged from 13,121 to 221,432, and the number of genes from 3543 to 9775 (Fig. 1c, d).
Highly variable genes identified in WJMSCs
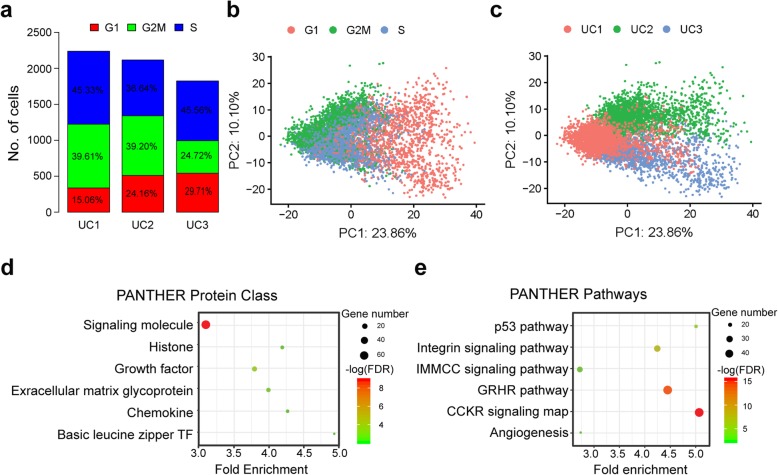
Considering cell cycle effect may influence gene expression, we first assigned cell cycle phases’ state to each cell. The results showed that an average of 22.98%, 34.51%, and 42.51% cells was assigned to G1, G2/M, and S cell cycle phase, respectively (Fig. 2a), suggesting that in vitro cultured WJMSCs are highly proliferated population. Principle components (PCs) analyzed without removing unwanted sources of variation demonstrated that PC1, counting for 23.86% variance, is mainly caused by cell cycle effect (Fig. 2b), while PC2 is counting for 10.10% variance (Fig. 2c), which results from donor-to-donor variation or batch effect. Thus, we selected overlapped highly variable genes among each phase for each donor as mentioned in (Additional file 1) and totally got 770 genes defined as HVGs for the following analysis (Additional file 3: Fig. S2A-D and Additional file 2: Table S4).
Fig. 2.
Heterogeneity and highly variable genes in WJMSCs. a Phases of cell cycle assigned for each of the three samples. b, c Cell cycle effects (b) and batch effects (c) represent the dominant source of heterogeneity in primary cultured WJMSC population. d Results of pathway enrichment analysis for highly variable genes identified in WJMSCs. e Results of protein class enrichment analysis for highly variable genes identified in WJMSCs. IMMC, inflammation mediated by chemokine and cytokine; GRHR, gonadotropin-releasing hormone receptor
Highly variable genes (HVGs) exhibiting high variability across cells represent heterogeneous features within a cell population [50, 51]. Here, we investigated gene functional enrichment of HVGs identified in the WJMSC population. Interestingly, the protein class analysis demonstrated that those genes were overrepresented in signaling molecules, growth factors, extracellular matrix protein, chemokine, histone, and basic leucine zipper transcription factor (Fig. 2d). Besides, the pathway analysis exhibited that these highly variable gene-expressed cross cells were enriched in integrin signaling pathway, inflammation mediated by chemokine and cytokine signaling pathway, gonadotropin-releasing hormone receptor pathway, p53 pathway, and angiogenesis (Fig. 2e). Furthermore, GO enrichment analysis showed that those HVGs are significantly enriched in the extracellular region (Additional file 3: Figure S2E) with binding function, such as protein binding and cytokine receptor binding (Additional file 3: Figure S2G), involved in biological processes like developmental process, signal transduction, cellular component morphogenesis, cell communication, and cell proliferation (Additional file 3: Figure S2F). Micro-environmental interaction is crucial for morphogenesis, cell differentiation, homeostasis, and cell growth [52, 53]. Therefore, variations in the expression of those extracellular functioning genes identified in our analysis could influence interaction of MSCs with micro-environment and cell fate determination [54, 55]. Furthermore, our results showed that highly variable genes in WJMSCs were enriched in distinct biological functions of MSCs, such as anti-inflammation (Fig. 2e), regeneration (Additional file 3: Figure S2F), and wound healing (Additional file 3: Figure S2G), some of which can potentially be selected as candidate markers for a matrix assay to test their therapeutic efficacy in clinical application.
Characteristics of candidate subpopulations in WJMSCs
To remove batch and cell cycle effects, we scaled the data and performed linear regression to regress the effects out before candidate subpopulation clustering. Here, we used regularized negative binomial regression method to perform normalization and variance stabilization of our scRNA-seq data, which is an appropriate distribution to model UMI count data from a “homogeneous” single-cell population suggested by [56]. The results of nonlinear dimensional reduction performed by UAMP showed that the cells were obviously separated by cell cycle and batch effects before regression, while cells were well mixed after regression and scaling (Additional file 3: Figure S3A, B), implying that those unwanted sources of variation have been effectively removed.
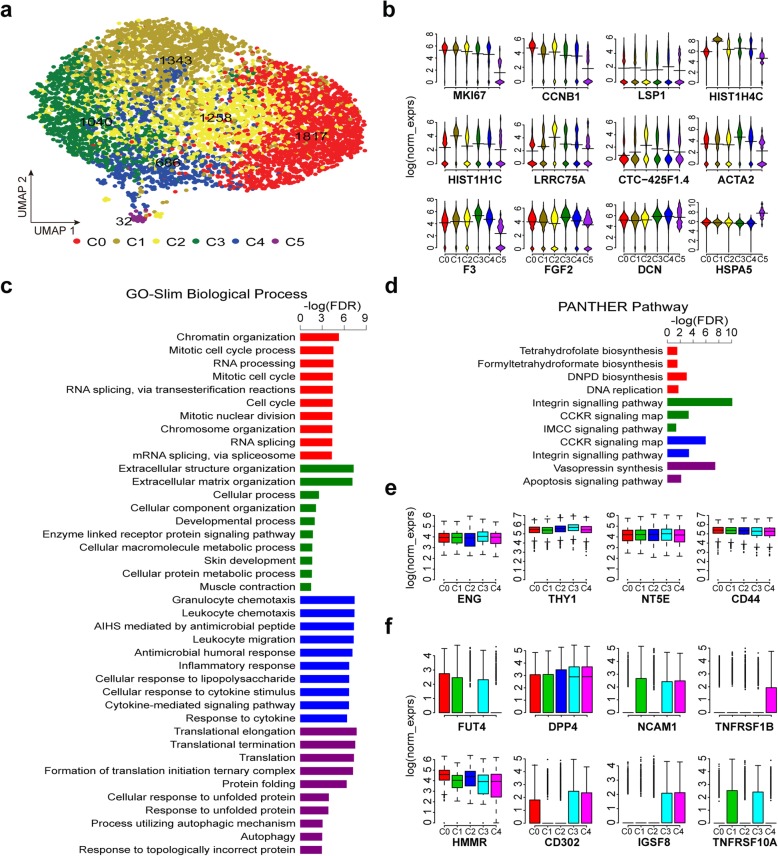
Next, we performed cell cluster analysis by a graph-based clustering approach [57], and six candidate clusters in primary cultured WJMSCs were identified (Fig. 3a). To study the molecular and functional characteristics of these candidate subpopulations in WJMSCs, we performed differentially expressed gene (DEG) analysis among the six clusters (C0–C5) (Additional file 2: Table S5-S6 and Fig. 3b). Intriguingly, MKI67 (marker of proliferation Ki-67), a gene strongly associated with cell proliferation and growth, is expressed at higher levels in subpopulations C0 and C1 compared with others, implying that subpopulations C0 and C1 possess a higher proliferative capacity. Results of GO enrichment analysis showed that DEGs upregulated in C0 were significantly enriched in DNA replication pathway and cell cycle process as well (Fig. 3c, d). Besides, several histone genes, such as HIST1H4C and HIST1H1C, exhibited higher expression levels in subpopulations C1 (Fig. 3b). Contrarily, cells in C5 displayed aging characteristics, although a proportion of which is very small in the total populations, and almost all these cells are assigned to G1 phase belonging to UC1 sample (Additional file 3: Figure S3C). We thought that cells in subpopulation C5 may have experienced mutation or replicative senescence during expansion, and they were removed from the following analysis. Across those candidate subpopulations, several markers of MSCs showed a similar expression level (Fig. 3e). Meanwhile, we noted that collagen and chemokine genes across these subpopulations were differentially expressed. Specially, the expression of collagen genes was much higher in C3 while the expression of chemokine genes was higher in C4 (Additional file 3: Figures S3D, E). Furthermore, we presented several candidate surface markers according their differential expression pattern across these candidate subpopulations (Fig. 3f, Additional file 2: Table S6). Whether they could be used efficiently to sort those subpopulations with stable phenotypes and functions needs to be assayed.
Fig. 3.
Candidate subpopulations with different functional characteristics. a UMAP visualizing the results of cell clustering. b Bean plots showing expression of several representative DEGs among the six subpopulations. c Pathways significantly enriched for the genes differentially expressed in one subpopulation compared to others. d GO-slim biological process enriched for the genes differentially expressed in one subpopulation compared to others. For c and d, only the top 10 terms with lowest FDR (FDR ≤ 0.05) were present. e Boxplots showing expression of classic MSC marker genes in subpopulations. f Example of candidate markers showing different expression pattern among the five subpopulations (C0–C4). C0, red; C1, olive; C2, yellow; C3, green; C4, blue; C5, purple. IMCC, inflammation mediated by chemokine and cytokine; DNPD, de novo pyrimidine deoxyribonucleotide
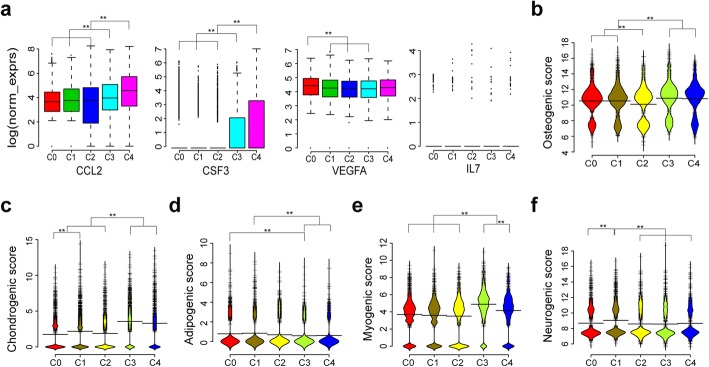
In terms of MSC function, on which the MSC clinical application were theoretically based, the DEGs upregulated in subpopulation C3 were enriched in extracellular structure organization, developmental process, skin development, and muscle contraction, while DEGs upregulated in subpopulation C4 were associated with immunomodulation function (Fig. 3c). The secretome analysis revealed that increased levels of some cytokines, such as CCL2, GCSF (CSF3), VEGF, and IL-7, are positively correlated with immunosuppression [58]. Among these subpopulation, the expression levels of CCL2 and CSF3 are highest in the C4 subpopulation (Fig. 4a), implicating its immunomodulation therapeutic potential. Besides, the lineage differentiation score among these subpopulations were different, indicating their distinct differentiation propensity to osteogenic, chondrogenic, adipogenic, myogenic, or neurogenic cells (Fig. 4b–f).
Fig. 4.
Candidate subpopulations showing different predicted potencies on differentiation and immunosuppression. a Boxplots showing expression of genes correlated with PBMC suppression across the five candidate subpopulations (C0–C4). b–f Bean plots showing distribution of log (norm_exprs) values of osteogenic score (b), chondrogenic score (c), adipogenic score (d), myogenic score (e), and neurogenic score (f) across the five candidate subpopulations (C0–C4). Wilcoxon rank sum test were performed for significant test, ** P < 0.001
To support our data, we isolated CD142+ WJMSCs and CD142− WJMSCs by flow cytometric sorting. Considering that CD142− cells account for about 20% of the total cells (Additional file 3: Fig S4A), a comparable number of cells with the highest fluorescence intensity were set as gate for CD142+ cells (Additional file 3: Fig S4B). According to our scRNA-seq analysis, CD142 (F3), which is a transmembrane protein and plays multiple important roles in wound healing [59], is expressed at the highest level in subpopulations C3 and lowest in C0 (Fig. 3b). Besides, DEGs, such as ITGA5, MME, FGF2, SPARC, FN1, TIMP1, COL4A1, FLNA, COL1A1, COL3A1, and COL5A1, which are functionally enriched in skin development and wound healing, were upregulated in C3 compared to others (Fig. 3c, Additional file 2: Table S5). Therefore, the C3 subpopulation is predicted to have greater potency in wound healing. In line with the scRNA-seq data, the RT-qPCR results showed that expression levels of SPARC, COL4A1, COL1A1, and COL5A1 were higher in CD142+ cells (Additional file 3: Fig S4C). Compared to CD142+ MSCs, CD142− MSCs showed higher MKI67 expression, indicating CD142− cells might possess a higher proliferative capacity (Additional file 3: Fig S4C). Indeed, we saw more cells from CD142− MSCs than CD142+ MSCs after culturing in vitro for 72 h (Additional file 3: Fig S4D). Then, we compared the wound healing potential between CD142+ WJMSCs and CD142− WJMSCs. The results revealed that fibroblasts cultured in conditioned media from CD142+ cells presented higher rate of wound closure than the conditioned media from CD142− cells (Additional file 3: Fig S4E and S4F). Therefore, these data demonstrated that CD142+ and CD142− WJMSCs showed distinct proliferation capacity as well as “wound healing” potency.
Single-cell transcriptome comparison between WJMSCs and ADMSCs
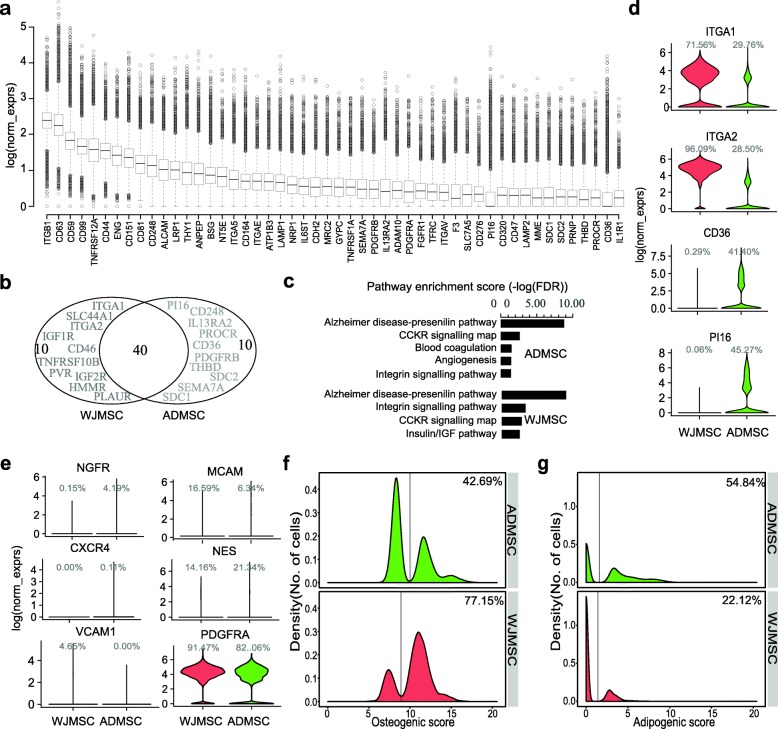
To provide insights into the heterogeneity of MSCs, several previous studies have compared gene expression of MSCs isolated from different sources using bulk-cell transcriptomic profiles [60–65]. However, even MSCs derived from the same tissue exhibited inter-population functional heterogeneity, such as different differentiation potency and proliferation capacity. Bulk RNA-seq measures the average expression of genes, which is the sum of cell type-specific gene expression weighted by cell type proportions. Bulk transcriptome comparisons may hide some meaningful information that can help to elucidate the underlying mechanisms of functional heterogeneity. Thus, here, we compared the transcriptome data at the single-cell level between WJMSCs and ADMSCs. As expected, a lot of highly expressed classic MSC surface markers are shared between these two MSCs, including ENG, NT5E, THY1, and CD44 (Fig. 5a, b and Additional file 2: Table S3). Meanwhile, some unshared CDs were identified (Fig. 5b), which suggest a phenotypic diversity between WJMSCs and ADMSCs. These unshared genes are involved in different cell signaling pathways inferred from the pathway enrichment analysis of the top 50 CDs (Fig. 5c). Not surprisingly, some of these unshared CDs, although ranked in the top 50 genes by average expression (Fig. 1a and Fig. 5a), are only expressed at high levels in a small proportion of the MSCs (Fig. 5d). Some of the unshared CDs are expressed (or not expressed) in the majority of the MSCs derived from one tissue but are expressed only in a small proportion of the MSCs in the other one, such as CD36, which plays an important role in the formation of intracellular lipid droplets [66], as well as ITGA1, ITGA2, and PI16 (Fig. 5d and Additional file 2: Table S3). Those CD genes hold the potential to be used as markers for subpopulation sorting for further physiological and functional research. Accordingly, we also found that several markers, which have been reported to identify special MSC subpopulations with different biological functions, are expressed weakly in a small portion of MSCs, including CD271 (NGFR) [67, 68], CD146 (MCAM) [69], CXCR4, NES [70], and CD106 (VCAM1) [71], except PDGFRA, which are highly expressed in most cells both in WJMSCs and ADMSCs (Fig. 5e and Additional file 2: Table S3).
Fig. 5.
Transcriptome comparison at the single-cell level between WJMSCs and ADMSCs. a Boxplot showing the top 50 CD genes ranked by average normalized expression in ADMSCs. b Venn diagram showing top 50 ADMSC CD genes overlap with the top 50 WJMSC CD genes, unshared genes were highlighted. c Pathway enrichment of top 50 CD genes expressed in ADMSCs and WJMSCs. d Example of CD genes showing different expression percentage between ADMSCs and WJMSCs. e Violin plots showing expression of markers for reported MSC subpopulations in ADMSCs and WJMSCs. f, g Density distribution showing osteogenic score (f) and adipogenic score (g) between ADMSCs and WJMSCs. Percentage indicating proportion of cells assigned to the right side of the line
Increasing reports demonstrated that MSCs derived from different sources exhibited distinct biological properties, such as proliferative capacity, multi-lineage differentiation ability, and immunomodulation potency [72], although they all meet the minimal criteria for defining multipotent MSCs. As regards the differentiation ability, we evaluated it by calculating the lineage differentiation score using single-cell gene expression data from WJMSCs and ADMSCs. Interestingly, density distributions of lineage score displayed two major peaks, while only one peak was observed in housekeeping genes analysis (Fig. 5f, g and Additional file 3: S5A-D), which indicates the existence of multiple subpopulations in MSCs. The density distribution of osteogenic score showed that 77.15% of the cells in WJMSCs have high osteogenic score vs. 42.69% in ADMSCs (Fig. 5f). On the contrary, more cells (54.84%) in ADMSCs have higher adipogenic score than in WJMSCs (22.12%) (Fig. 5g). These results suggested that WJMSCs have the propensity towards the osteogenic lineage while ADMSCs are inclined to differentiate into adipose cells, which are in line with previous studies [33]. Moreover, the difference in other lineage differentiation potencies also existed, such as the chondrogenic and myogenic potential (Additional file 2: Figures S5A, C).
Recently, human skeletal stem cells (SSCs) and adipose progenitor cells were identified [73, 74]. Human SSCs with PDPN +CD146 (MCAM) −CD73 (NT5E) + CD164 + phenotype have the ability to generate progenitors of the bone, cartilage, and stroma, but not fat [73]. Human adipose progenitor cells which expressed DPP4 are able to give rise to committed ICAM1+ preadipocytes [74]. To determine whether cultured MSCs consist of cells with phenotype as SSCs or adipose progenitors, we analyzed single-cell expression data with the above marker genes. Notably, the proportion of cells which expressed SSC markers in WJMSCs is much higher than in ADMSCs (Additional file 3: Figures S5E, F), while more cells with adipose progenitor’s markers exist in ADMSCs (Additional file 3: Figures S5G, H). These results further indicated that bulk-cell variations among MSCs from different sources may originate from the composition diversity of distinct subpopulations.
Discussion
MSCs are promising cell therapy products with great potential in promoting tissue regeneration and modulating inflammation. However, significant variations were reported on MSCs that were isolated from different donors and different tissue sites. Unrefined, non-standardized isolation and culture techniques become the challenges of standardization in processes of MSC product manufacturing and quality management. Even in a “homogeneous” population, which is defined by the classic minimal criteria, including cell size [75], morphology [76–78], proliferation capacity [75], differentiation potency [27], and immunomodulation capacity [76, 77], these cells still display phenotype and function heterogeneity among individual cells [79]. In previous clinical trials, functional variation and heterogeneity in MSCs are potentially the main reasons which lead to inconsistent or controversial results [29, 80]. The underlying molecular mechanisms that lead to MSC functional variation and heterogeneity at the cell population level remain unknown, which require further investigation and elucidation.
Recently, several studies have been performed to investigate into the heterogeneity of cultured MSCs by single-cell transcriptomic analysis [35, 81–83]. Huang et al. profiled the transcriptomes of 361 single MSCs derived from two umbilical cords (UC-MSCs) that were harvested at different passages and stimulated with or without inflammatory cytokines. Following the analysis, they concluded that in vitro expanded UC-MSCs are a well-organized population with limited heterogeneity, which is mainly caused by distinct distribution in cell cycle phases [81]. However, the number of cells sequenced for each condition is small (~ 50 cells per condition), and they did not remove the cell cycle effects for the subpopulations identification. Besides, the marker (HMMR) they used to sort the cells to confirm their hypothesis may be unable to isolate different subpopulations. Liu et al. performed a large-scale single-cell transcriptomic sequencing of 24,370 cultured ADMSCs from three donors [35]. They regressed out batch and cell cycle effects before candidate subpopulation classification; however, the results they exhibited in the report were limited to the data analysis pipeline. Wang et al. sequenced a total of 103 single hWJMSCs from three umbilical cords and 63 single hBMMSC cells from two different donors and just focused on gene expression comparison between the two different source-derived MSCs [82]. Thus, the cellular transcriptomic heterogeneity within a MSC population cultured in vitro still has not been comprehensively investigated at the single-cell level.
In this study, we dissected the gene expression heterogeneity of human primary WJMSCs cultured in vitro using scRNA-seq. Single-cell RNA sequencing technologies can offer an unbiased approach for understanding the extent, basis, and function of gene expression variation between seemingly identical cells, revealing complex and rare cell populations, uncovering regulatory relationships between genes, and tracking the trajectories of distinct cell lineages in development [32, 84]. In primary WJMSCs, we found that the HVGs are significantly enriched in extracellular region with binding function, involved in developmental process, signal transduction, cell proliferation, etc. For example, MKI67, a marker of proliferation, was identified as one of the HVGs, implying a different proliferate capacity among individual cells. Indeed, we showed that CD142− WJMSCs with higher expression of MKI67 presented higher proliferation capacity than CD142+ WJMSCs cultured in vitro (Additional file 3: Fig S4C). In terms of therapeutic potential, these genes are associated with functional characteristics of MSCs, such as integrin signaling pathway, angiogenesis, and inflammation mediated by chemokine and cytokine signaling pathway (Fig. 2e). Integrin signaling pathway plays a critical role in homing of MSCs to the bone, osteogenic differentiation, and bone formation, and even some integrins are suggested as targets to promote bone formation and repair [85–87]. Several integrin genes were identified in our data with highly variable expression across the cells, such as ITGA5 and ITGB1, which respectively encode α5 and β1 and together form the α5β1 integrin, a cell surface receptor for fibronectin implicated in the control of osteoblastogenesis [87, 88]. Pro-angiogenesis is one of the important biological properties of MSCs, implicating in promoting wound healing and tissue repair [89, 90]. Genes related to the angiogenesis, such as ANGPT1, PDGFRA, and VEGFA., were identified in our HVGs (Additional file 2: Table S4). Studies have reported that ANGPT1 gene-modified human MSCs could promote angiogenesis and reduce acute pancreatitis in rats [91], while PDGFRA+ MSCs have enhanced skin repair/regeneration potential [92]. VEGFA, and other two cytokines, CXCL5 and CXCL8 (IL-8), were required for the angiogenic activity of MSCs and have been selected as an assay matrix for angiogenic potency assay for MultiStem product [93, 94]. Furthermore, in vitro co-culture assays demonstrated that the increased levels of VEGFA and chemokine CCL2 in MSCs were positively correlated with PBMC suppression [58]. Chemokines, a family of small cytokines, are recognized as key mediators of MSCs migration and immunosuppression [8, 95]. Notably, most of the chemokines expressed in primary WJMSCs were highly heterogeneous, including the abovementioned CCL2, CXCL5, and CXCL8. These results indicated that highly variable genes within WJMSCs are associated with classic MSC functional properties and suggested the existence of potential subpopulations with different gene expression patterns.
Although cultured MSCs meet the minimal criteria with classic phenotype, increasing reports demonstrated that many cell surface membrane proteins are not uniformly expressed in MSCs [96]. Several subpopulations with a different phenotype, property, and therapeutic potential have been identified in MSCs derived from different tissues. Some subpopulations express CXCR4 and have a propensity to migrate to sites of tissue injury [97], while some express VCAM-1 (CD106) and show priority in immunosuppression [71]. However, these above-reported markers only weakly expressed in WJMSCs (Fig. 4e) may be unable to serve as effective markers to isolate these subpopulations in WJMSCs. Here, we classified WJMSCs into several candidate subpopulations (C0–C5) with different functional characteristics. Among these candidate subpopulations, DEG analysis indicated that C0 and C1 show greater proliferation ability while C3 and C4 have greater osteogenic and chondrogenic differentiation potency. Myogenic score is also significantly higher in C3, implying its potential in myocardial repair application. As to immunomodulation, we found that most of the chemokines and some immune-related cytokines detected in WJMSCs upregulate in C4. Furthermore, our analysis demonstrated that DEGs, including F3 (CD142), ITGA5, MME, FGF2, SPARC, FN1, TIMP1, COL4A1, FLNA, IGFBP7, COL1A1, COL3A1, and COL5A1, which are functionally enriched in skin development and wound healing, are expressed at the highest level in the C3 cluster (Fig. 3b). To support our analysis, we isolated CD142+ and CD142− WJMSCs to compare their potency in wound healing (Additional file 3: Fig S4). And the results showed that CD142+ WJMSCs exhibited lower proliferation capacity and higher wound healing potency than CD142− cells. Many preclinical and clinical studies have been performed to test the regenerative properties of human MSCs in different pathologies, one of which is wound healing [98]. Although the underlying mechanism of MSCs for the enhancement of wound healing remains to be elucidated, several studies have shown that MSCs could secrete some cytokines and growth factors, including TGFB1, FGF2, CCL2, CXCL8, and IGFBP7, to accelerate cell migration and enhance normal wound healing [46, 99]. Similarly, extracellular matrix (ECM) components, such as collagen types I, V, VI, and XII and fibronectin, along with SPARC, play important roles in regulating cell migration as well [46]. Using RT-qPCR assays, we confirmed that most of the abovementioned wound healing-related genes, TGFB1, SPARC, COL4A1, COL1A1, and COL5A1, were expressed at higher levels in CD142+ than in CD142− WJMSCs (Additional file 3: Fig S4C). Taken together, these candidate subpopulations identified in primary WJMSCs would be valuable for further biological characterization via experimental investigations and clinical researches.
ADMSCs can be easily isolated from the stromal vascular fraction (SVF) by enzymatic digestion or non-enzymatic digestion of adipose tissue [100, 101], which are the most widely studied MSCs after BM-MSCs. During wound healing, hair regrowth, and tissue reconstruction, there are several similar bio-molecular pathways that are essential, like angiogenesis, cell migration and recruitment, and cell growth and morphogenesis, which can be enhanced by many growth factors such as TGF-β, FGF family, VEGF, interleukin (IL), PDGF, and IGF. It is well known that these are typical characteristics and biological functions of ADMSCs. Therefore, alone or in combination with hyaluronic acid, platelet-rich plasma (PRP), or fat graft, ADMSCs or AD-SVFs have been tested in many clinical settings to improve wound healing, hair regrowth, and breast reconstruction and demonstrated an encouraging therapeutic potential [98, 102–106]. However, challenges of developing potency assays for those MSC-like products still hinder their clinical applications, which include variability of tissue sources, different isolated and culture methods, largely undefined mechanisms of action in humans, and lack of reference standards [58, 94, 100, 107, 108]. By comparing gene expression between ADMSCs and WJMSCs at the single-cell level, we found that HVGs are largely overlapped between them (Additional file 3: Figure S6A). These shared HVGs include genes, such as HGF, FGF2, FGF7, VEGFA, TGFB2, IGF2, IGFBP3-7, CSF3, PDGFA, PTGES, and PTGS2, coding proteins which are involved in bio-molecular pathways that could promote cell proliferation, differentiation, and neo-angiogenesis and suppress apoptotic cues, to favor wound healing and hair regrowth processes [98, 105, 109]. Furthermore, functional enrichment analysis of HVGs from ADMSCs showed similar results as those from WJMSCs (Additional file 3: Figures S6B-G), though they exhibited distinct differentiation propensity (Additional file 3: Figure S5). Altogether, we inferred that these HVGs should play critical roles in MSC functional heterogeneity and may serve as candidate markers for further potency association studies. Further studies in cell-to-cell variability in transcriptome, proteome, secretome, and epigenome on MSCs derived from different tissues will increase our understanding of the heterogeneity associated with MSC function and facilitate the development of MSC release criteria for clinical application.
Conclusions
In summary, our results reveal that highly variable genes within MSCs are significantly enriched in the extracellular region with binding function, involved in developmental process, signal transduction, and cell proliferation. Several candidate subpopulations exhibiting distinct function can be obtained in MSCs applying these HVGs for dimension reduction and clustering. Regarding therapeutic potential, these HVGs are associated with functional characteristics, on which the MSC clinical application was theoretically based, such as development and inflammation response. Altogether, our study suggests that these HVGs hold the potential to be used as candidate markers for further potency association studies.
Supplementary information
Additional file 1: Supplemental methods describing scRNA-seq data analysis for detail, including methods about Quality control, Removal of cell cycle effect, Highly Variable genes identification, Linear and nonlinear dimension reduction, Clustering the cells and Differential expression analysis.
Additional file 2: Supplemental tables with six tables. Table S1. Marker genes used for potency score analysis. Table S2. Top100 CD genes ranked by mean expression in WJMSCs and ADMSCs. Table S3. CD genes ranked by percentage of cells expressed the genes (at least one UMI). Table S4. List of HVGs identified in WJMSCs. Table S5. DEGs in each cluster by comparing it to all of the others. Table S6. Results of differential gene expression analysis between two different clusters.
Additional file 3: Supplemental figures with five figures. Figure S1. Quality of the WJMSCs single-cell RNA-seq data. (A) Number of reads were sequenced for each of the three samples. Percentage of reads mapped to exonic (B) and mapped to transcriptome (C) for each of the three samples. (D) Number of cells obtained for each of the three samples. Boxplot showing number of expressed genes per cell (E) and number of UMI per cell (F) for each of the three samples. (G) Tri-lineage differentiation potency of primary cultured WJMSCs used for scRNA-seq. Figure S2. Highly variable genes identification in WJMSCs and GO enrichment analysis. (A) Venn diagram showing overlap of top 2000 highly variable genes among different phases for sample UC1. (B) Venn diagram showing overlap of top 2000 highly variable genes among different phases for sample UC2. (C) Venn diagram showing overlap of top 2000 highly variable genes among different phases for sample UC3. (D) Venn diagram showing overlap of highly variable genes among samples. Results of GO-slim cellular component enrichment analysis (E), GO-slim biological process enrichment analysis (F), and GO-slim functional molecular enrichment analysis for highly variable genes. Figure S3. Candidate subpopulations identified in WJMSCs. (A) and (B) UMAP showing dimension reduction before and after batch (A) and cell cycle effect (B) removal. Left, before removal; right, after removal. (C) Histogram showing number of cells for each phase of cell cycle and sample in the candidate subpopulations. (D) Violin plots showing distribution of log normalized expression (log (norm_exprs)) values of collagen genes across the six candidate subpopulations (C0–C5). (E) Violin plots showing distribution of log (norm_exprs) values of chemokines genes across the six candidate subpopulations (C0–C5). Figure S4. Wound healing potency for CD142+ and CD142− WJMSCs. (A) CD142 analysis by flow cytometry for WJMSCs. (B) Example of gate setting for CD142− (left gate) and CD142+ (right gate) cells sorting. (C) qPCR-based expression fold-changes for genes upregulated in C3 plus CCL2, CXCL8 and MKI67 (n = 3) between CD142+ and CD142− cells. (D) Proliferation for CD142− and CD142+ cells cultured in vitro. n = 4 for each time point. (E) Representative images of wound healing assays for conditioned media from CD142- and CD142+ cells cultured fibroblast, respectively. (F) Wound closure comparison between CD142+ and CD142− cells conditioned media for 24 h (n = 9). Data shown are means ± SD, *p < 0.05. Paired two-tailed Student’s t-test were performed for significant test. Figure S5. Differentiation potency compared between ADMSCs and WJMSCs. Density distribution showing chondrogenic score(A), neurogenic score (B), myogenic score(C), and housekeeping score (D) between ADMSCs and WJMSCs. Percentage indicating proportion of cells assigned to the right side of the line. (E) Violin plots showing marker genes of SSC expressed in ADMSCs and WJMSCs. (F) Percentage of cells expressed SSC marker genes in ADMSCs and WJMSCs. (G) Violin plots showing marker genes of adipose progenitors expressed in ADMSCs and WJMSCs. (H) Percentage of cells expressed marker genes of adipose progenitors in ADMSCs and WJMSCs. Figure S6. Functional enrichment of highly variable genes identified in ADMSCs. (A) Venn diagram showing overlap of HVGs between ADMSCs and WJMSCs. Barplots showing results of GO-slim cellular component (B), GO-slim molecular function (C), Go-slim biological process (D), Reactome pathways (E), Pathways (F), and protein class (G) enrichment analysis for HVGs identified in ADMSCs.
Acknowledgements
We thank the members of the China National GeneBank for technical support.
Abbreviations
- ADMSCs
Adipose-derived MSCs
- CD
Cluster of differentiation
- DEGs
Differentially expressed genes
- DMEM
Dulbecco’s modified Eagle’s medium
- ECM
Extracellular matrix
- FBS
Fetal bovine serum
- GO
Gene ontology
- GvHD
Graft versus host disease
- HVGs
Highly variable genes
- IBMX
3-Isobutyl-1-methylxanthine
- ISCT
International Society for Cellular Therapy
- MSCs
Mesenchymal stem/stromal cells
- PCs
Principle components
- scRNA-seq
Single-cell RNA sequencing
- SSC
Skeletal stem cell
- UAMP
Uniform Manifold Approximation and Projection
- UMI
Unique molecular identifiers
- WJMSCs
Wharton’s jelly-derived MSCs
Authors’ contributions
C.S. and X.Z. contributed to the conception and design of the study. C.S., L.W., T.H., W.Y. and J.L. performed the experiments. C.S. did the data analysis and prepared the figures of the manuscript. C.S., H.W. and X.Z. wrote the main manuscript. All authors read and approved the final manuscript.
Funding
This work was supported by the Shenzhen Municipal Government of China (No. CYFWT201507021005 to X.Z.) and Science, Technology and Innovation Commission of Shenzhen Municipality under grant no. KQJSCX20170322143848 413 to X.Z.
Availability of data and materials
The data that support the findings of this study have been deposited in the CNSA (https://db.cngb.org/cnsa/) of CNGBdb with accession number CNP0000562. All data analyzed during this study have been included in this published article and its supplementary information files.
Ethics approval and consent to participate
This study was approved by the Ethics Committee of BGI-IRB (NO. BGI-IRB 19015-1).
Consent for publication
Written informed consent was obtained from the donors for recording/using their individual details (age, gender, weight, and height) for research purposes.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information accompanies this paper at 10.1186/s13287-020-01660-4.
References
- 1.Prockop DJ. Marrow stromal cells as stem cells for nonhematopoietic tissues. Science. 1997;276(5309):71–74. doi: 10.1126/science.276.5309.71. [DOI] [PubMed] [Google Scholar]
- 2.Zuk PA, Zhu M, Ashjian P, De Ugarte DA, Huang JI, Mizuno H, et al. Human adipose tissue is a source of multipotent stem cells. Mol Biol Cell. 2002;13(12):4279–4295. doi: 10.1091/mbc.e02-02-0105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Romanov YA, Svintsitskaya VA, Smirnov VN. Searching for alternative sources of postnatal human mesenchymal stem cells: candidate MSC-like cells from umbilical cord. Stem Cells. 2003;21(1):105–110. doi: 10.1634/stemcells.21-1-105. [DOI] [PubMed] [Google Scholar]
- 4.Wang HS, Hung SC, Peng ST, Huang CC, Wei HM, Guo YJ, et al. Mesenchymal stem cells in the Wharton's jelly of the human umbilical cord. Stem Cells. 2004;22(7):1330–1337. doi: 10.1634/stemcells.2004-0013. [DOI] [PubMed] [Google Scholar]
- 5.Fukuchi Y, Nakajima H, Sugiyama D, Hirose I, Kitamura T, Tsuji K. Human placenta-derived cells have mesenchymal stem/progenitor cell potential. Stem Cells. 2004;22(5):649–658. doi: 10.1634/stemcells.22-5-649. [DOI] [PubMed] [Google Scholar]
- 6.Gao F, Chiu SM, Motan DA, Zhang Z, Chen L, Ji HL, et al. Mesenchymal stem cells and immunomodulation: current status and future prospects. Cell Death Dis. 2016;7:e2062. doi: 10.1038/cddis.2015.327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Pittenger MF, Mackay AM, Beck SC, Jaiswal RK, Douglas R, Mosca JD, et al. Multilineage potential of adult human mesenchymal stem cells. Science. 1999;284(5411):143–147. doi: 10.1126/science.284.5411.143. [DOI] [PubMed] [Google Scholar]
- 8.Ren G, Zhang L, Zhao X, Xu G, Zhang Y, Roberts AI, et al. Mesenchymal stem cell-mediated immunosuppression occurs via concerted action of chemokines and nitric oxide. Cell Stem Cell. 2008;2(2):141–150. doi: 10.1016/j.stem.2007.11.014. [DOI] [PubMed] [Google Scholar]
- 9.Aggarwal S, Pittenger MF. Human mesenchymal stem cells modulate allogeneic immune cell responses. Blood. 2005;105(4):1815–1822. doi: 10.1182/blood-2004-04-1559. [DOI] [PubMed] [Google Scholar]
- 10.Parekkadan B, Milwid JM. Mesenchymal stem cells as therapeutics. Annu Rev Biomed Eng. 2010;12:87–117. doi: 10.1146/annurev-bioeng-070909-105309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ikebe C, Suzuki K. Mesenchymal stem cells for regenerative therapy: optimization of cell preparation protocols. Biomed Res Int. 2014;2014:951512. doi: 10.1155/2014/951512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Baksh D, Song L, Tuan RS. Adult mesenchymal stem cells: characterization, differentiation, and application in cell and gene therapy. J Cell Mol Med. 2004;8(3):301–316. doi: 10.1111/j.1582-4934.2004.tb00320.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Abdallah BM, Kassem M. Human mesenchymal stem cells: from basic biology to clinical applications. Gene Ther. 2008;15(2):109–116. doi: 10.1038/sj.gt.3303067. [DOI] [PubMed] [Google Scholar]
- 14.Wakitani S, Imoto K, Yamamoto T, Saito M, Murata N, Yoneda M. Human autologous culture expanded bone marrow mesenchymal cell transplantation for repair of cartilage defects in osteoarthritic knees. Osteoarthr Cartil. 2002;10(3):199–206. doi: 10.1053/joca.2001.0504. [DOI] [PubMed] [Google Scholar]
- 15.Krampera M, Pizzolo G, Aprili G, Franchini M. Mesenchymal stem cells for bone, cartilage, tendon and skeletal muscle repair. Bone. 2006;39(4):678–683. doi: 10.1016/j.bone.2006.04.020. [DOI] [PubMed] [Google Scholar]
- 16.Chen SL, Fang WW, Ye F, Liu YH, Qian J, Shan SJ, et al. Effect on left ventricular function of intracoronary transplantation of autologous bone marrow mesenchymal stem cell in patients with acute myocardial infarction. Am J Cardiol. 2004;94(1):92–95. doi: 10.1016/j.amjcard.2004.03.034. [DOI] [PubMed] [Google Scholar]
- 17.Ranganath SH, Levy O, Inamdar MS, Karp JM. Harnessing the mesenchymal stem cell secretome for the treatment of cardiovascular disease. Cell Stem Cell. 2012;10(3):244–258. doi: 10.1016/j.stem.2012.02.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Mazzini L, Ferrero I, Luparello V, Rustichelli D, Gunetti M, Mareschi K, et al. Mesenchymal stem cell transplantation in amyotrophic lateral sclerosis: a phase I clinical trial. Proc Natl Acad Sci U S A. 2010;223(1):229–237. doi: 10.1016/j.expneurol.2009.08.007. [DOI] [PubMed] [Google Scholar]
- 19.Karussis D, Karageorgiou C, Vaknin-Dembinsky A, Gowda-Kurkalli B, Gomori JM, Kassis I, et al. Safety and immunological effects of mesenchymal stem cell transplantation in patients with multiple sclerosis and amyotrophic lateral sclerosis. Arch Neurol. 2010;67(10):1187–1194. doi: 10.1001/archneurol.2010.248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kharaziha P, Hellstrom PM, Noorinayer B, Farzaneh F, Aghajani K, Jafari F, et al. Improvement of liver function in liver cirrhosis patients after autologous mesenchymal stem cell injection: a phase I-II clinical trial. Eur J Gastroenterol Hepatol. 2009;21(10):1199–1205. doi: 10.1097/MEG.0b013e32832a1f6c. [DOI] [PubMed] [Google Scholar]
- 21.Ghannam S, Bouffi C, Djouad F, Jorgensen C, Noel D. Immunosuppression by mesenchymal stem cells: mechanisms and clinical applications. Stem Cell Res Ther. 2010;1(1):2. doi: 10.1186/scrt2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Le Blanc K, Frassoni F, Ball L, Locatelli F, Roelofs H, Lewis I, et al. Mesenchymal stem cells for treatment of steroid-resistant, severe, acute graft-versus-host disease: a phase II study. Lancet. 2008;371(9624):1579–1586. doi: 10.1016/S0140-6736(08)60690-X. [DOI] [PubMed] [Google Scholar]
- 23.Galipeau J, Sensebe L. Mesenchymal stromal cells: clinical challenges and therapeutic opportunities. Cell Stem Cell. 2018;22(6):824–833. doi: 10.1016/j.stem.2018.05.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Dominici M, Le Blanc K, Mueller I, Slaper-Cortenbach I, Marini F, Krause D, et al. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy. 2006;8(4):315–317. doi: 10.1080/14653240600855905. [DOI] [PubMed] [Google Scholar]
- 25.Jin HJ, Bae YK, Kim M, Kwon SJ, Jeon HB, Choi SJ, et al. Comparative analysis of human mesenchymal stem cells from bone marrow, adipose tissue, and umbilical cord blood as sources of cell therapy. Int J Mol Sci. 2013;14(9):17986–18001. doi: 10.3390/ijms140917986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Yoo KH, Jang IK, Lee MW, Kim HE, Yang MS, Eom Y, et al. Comparison of immunomodulatory properties of mesenchymal stem cells derived from adult human tissues. Cell Immunol. 2009;259(2):150–156. doi: 10.1016/j.cellimm.2009.06.010. [DOI] [PubMed] [Google Scholar]
- 27.Russell KC, Phinney DG, Lacey MR, Barrilleaux BL, Meyertholen KE, O'Connor KC. In vitro high-capacity assay to quantify the clonal heterogeneity in trilineage potential of mesenchymal stem cells reveals a complex hierarchy of lineage commitment. Stem Cells. 2010;28(4):788–798. doi: 10.1002/stem.312. [DOI] [PubMed] [Google Scholar]
- 28.Samsonraj RM, Rai B, Sathiyanathan P, Puan KJ, Rotzschke O, Hui JH, et al. Establishing criteria for human mesenchymal stem cell potency. Stem Cells. 2015;33(6):1878–1891. doi: 10.1002/stem.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Phinney DG. Functional heterogeneity of mesenchymal stem cells: implications for cell therapy. J Cell Biochem. 2012;113(9):2806–2812. doi: 10.1002/jcb.24166. [DOI] [PubMed] [Google Scholar]
- 30.Bustos ML, Huleihel L, Kapetanaki MG, Lino-Cardenas CL, Mroz L, Ellis BM, et al. Aging mesenchymal stem cells fail to protect because of impaired migration and antiinflammatory response. Am J Respir Crit Care Med. 2014;189(7):787–798. doi: 10.1164/rccm.201306-1043OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sethe S, Scutt A, Stolzing A. Aging of mesenchymal stem cells. Ageing Res Rev. 2006;5(1):91–116. doi: 10.1016/j.arr.2005.10.001. [DOI] [PubMed] [Google Scholar]
- 32.Tang F, Barbacioru C, Wang Y, Nordman E, Lee C, Xu N, et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat Methods. 2009;6(5):377–382. doi: 10.1038/nmeth.1315. [DOI] [PubMed] [Google Scholar]
- 33.Li X, Bai J, Ji X, Li R, Xuan Y, Wang Y. Comprehensive characterization of four different populations of human mesenchymal stem cells as regards their immune properties, proliferation and differentiation. Int J Mol Med. 2014;34(3):695–704. doi: 10.3892/ijmm.2014.1821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Troyer DL, Weiss ML. Wharton's jelly-derived cells are a primitive stromal cell population. Stem Cells. 2008;26(3):591–599. doi: 10.1634/stemcells.2007-0439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Liu X, Xiang Q, Xu F, Huang J, Yu N, Zhang Q, et al. Single-cell RNA-seq of cultured human adipose-derived mesenchymal stem cells. Sci Data. 2019;6:190031. doi: 10.1038/sdata.2019.31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Natarajan KN, Miao Z, Jiang M, Huang X, Zhou H, Xie J, et al. Comparative analysis of sequencing technologies for single-cell transcriptomics. Genome Biol. 2019;20(1):70. doi: 10.1186/s13059-019-1676-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Islam S, Zeisel A, Joost S, La Manno G, Zajac P, Kasper M, et al. Quantitative single-cell RNA-seq with unique molecular identifiers. Nat Methods. 2014;11(2):163–166. doi: 10.1038/nmeth.2772. [DOI] [PubMed] [Google Scholar]
- 38.Lun AT, McCarthy DJ, Marioni JC. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Research. 2016;5:2122. doi: 10.12688/f1000research.9501.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Butler A, Hoffman P, Smibert P, Papalexi E, Satija R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol. 2018;36(5):411–420. doi: 10.1038/nbt.4096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Sun C, Zhang J, Zheng D, Wang J, Yang H, Zhang X. Transcriptome variations among human embryonic stem cell lines are associated with their differentiation propensity. PLoS One. 2018;13(2):e0192625. doi: 10.1371/journal.pone.0192625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Pertea M, Kim D, Pertea GM, Leek JT, Salzberg SL. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat Protoc. 2016;11(9):1650–1667. doi: 10.1038/nprot.2016.095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30(7):923–930. doi: 10.1093/bioinformatics/btt656. [DOI] [PubMed] [Google Scholar]
- 43.Li B, Dewey CN. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics. 2011;12:323. doi: 10.1186/1471-2105-12-323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Mi H, Muruganujan A, Huang X, Ebert D, Mills C, Guo X, et al. Protocol update for large-scale genome and gene function analysis with the PANTHER classification system (v.14.0) Nat Protoc. 2019;14(3):703–721. doi: 10.1038/s41596-019-0128-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Schwalie PC, Dong H, Zachara M, Russeil J, Alpern D, Akchiche N, et al. A stromal cell population that inhibits adipogenesis in mammalian fat depots. Nature. 2018;559(7712):103–108. doi: 10.1038/s41586-018-0226-8. [DOI] [PubMed] [Google Scholar]
- 46.Walter MN, Wright KT, Fuller HR, MacNeil S, Johnson WE. Mesenchymal stem cell-conditioned medium accelerates skin wound healing: an in vitro study of fibroblast and keratinocyte scratch assays. Exp Cell Res. 2010;316(7):1271–1281. doi: 10.1016/j.yexcr.2010.02.026. [DOI] [PubMed] [Google Scholar]
- 47.Docheva D, Popov C, Mutschler W, Schieker M. Human mesenchymal stem cells in contact with their environment: surface characteristics and the integrin system. J Cell Mol Med. 2007;11(1):21–38. doi: 10.1111/j.1582-4934.2007.00001.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hamidouche Z, Fromigue O, Ringe J, Haupl T, Vaudin P, Pages JC, et al. Priming integrin alpha5 promotes human mesenchymal stromal cell osteoblast differentiation and osteogenesis. Proc Natl Acad Sci U S A. 2009;106(44):18587–18591. doi: 10.1073/pnas.0812334106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Olivares-Navarrete R, Hyzy SL, Park JH, Dunn GR, Haithcock DA, Wasilewski CE, et al. Mediation of osteogenic differentiation of human mesenchymal stem cells on titanium surfaces by a Wnt-integrin feedback loop. Biomaterials. 2011;32(27):6399–6411. doi: 10.1016/j.biomaterials.2011.05.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Pijuan Sala B, Diamanti E, Shepherd M, Laurenti E, Wilson NK, Kent DG, et al. Comprehensive integration of single-cell data. Blood. 2019;177(7):1888–1902. doi: 10.1016/j.cell.2019.05.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Yip SH, Sham PC, Wang J. Evaluation of tools for highly variable gene discovery from single-cell RNA-seq data. Brief Bioinform. 2019;20(4):1583–89. [DOI] [PMC free article] [PubMed]
- 52.Frantz C, Stewart KM, Weaver VM. The extracellular matrix at a glance. J Cell Sci. 2010;123(Pt 24):4195–4200. doi: 10.1242/jcs.023820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Rozario T, DeSimone DW. The extracellular matrix in development and morphogenesis: a dynamic view. Dev Biol. 2010;341(1):126–140. doi: 10.1016/j.ydbio.2009.10.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Guilak F, Cohen DM, Estes BT, Gimble JM, Liedtke W, Chen CS. Control of stem cell fate by physical interactions with the extracellular matrix. Cell Stem Cell. 2009;5(1):17–26. doi: 10.1016/j.stem.2009.06.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Even-Ram S, Artym V, Yamada KM. Matrix control of stem cell fate. Cell. 2006;126(4):645–647. doi: 10.1016/j.cell.2006.08.008. [DOI] [PubMed] [Google Scholar]
- 56.Hafemeister C, Satija R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 2019;20(1):296. [DOI] [PMC free article] [PubMed]
- 57.Macosko EZ, Basu A, Satija R, Nemesh J, Shekhar K, Goldman M, et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell. 2015;161(5):1202–1214. doi: 10.1016/j.cell.2015.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Chinnadurai R, Rajan D, Qayed M, Arafat D, Garcia M, Liu Y, et al. Potency analysis of mesenchymal stromal cells using a combinatorial assay matrix approach. Cell Rep. 2018;22(9):2504–2517. doi: 10.1016/j.celrep.2018.02.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Hoffman M. The tissue factor pathway and wound healing. Semin Thromb Hemost. 2018;44(2):142–150. doi: 10.1055/s-0037-1606181. [DOI] [PubMed] [Google Scholar]
- 60.Lee RH, Kim B, Choi I, Kim H, Choi HS, Suh K, et al. Characterization and expression analysis of mesenchymal stem cells from human bone marrow and adipose tissue. Cell Physiol Biochem. 2004;14(4–6):311–324. doi: 10.1159/000080341. [DOI] [PubMed] [Google Scholar]
- 61.Ma J, Wu J, Han L, Jiang X, Yan L, Hao J, et al. Comparative analysis of mesenchymal stem cells derived from amniotic membrane, umbilical cord, and chorionic plate under serum-free condition. Stem Cell Res Ther. 2019;10(1):19. doi: 10.1186/s13287-018-1104-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Meng X, Sun B, Xiao Z. Comparison in transcriptome and cytokine profiles of mesenchymal stem cells from human umbilical cord and cord blood. Gene. 2019;696:10–20. doi: 10.1016/j.gene.2019.02.017. [DOI] [PubMed] [Google Scholar]
- 63.Fong CY, Chak LL, Biswas A, Tan JH, Gauthaman K, Chan WK, et al. Human Wharton’s jelly stem cells have unique transcriptome profiles compared to human embryonic stem cells and other mesenchymal stem cells. Stem Cell Rev. 2011;7(1):1–16. doi: 10.1007/s12015-010-9166-x. [DOI] [PubMed] [Google Scholar]
- 64.Alhattab D, Jamali F, Ali D, Hammad H, Adwan S, Rahmeh R, et al. An insight into the whole transcriptome profile of four tissue-specific human mesenchymal stem cells. Regen Med. 2019;14(9):841–65. [DOI] [PubMed]
- 65.Taşkiran EZ, Karaosmanoğlu B. Transcriptome analysis reveals differentially expressed genes between human primary bone marrow mesenchymal stem cells and human primary dermal fibroblasts. Turk J Biol. 2019;43(1):21–7. [DOI] [PMC free article] [PubMed]
- 66.Durandt C, van Vollenstee FA, Dessels C, Kallmeyer K, de Villiers D, Murdoch C, et al. Novel flow cytometric approach for the detection of adipocyte subpopulations during adipogenesis. J Lipid Res. 2016;57(4):729–742. doi: 10.1194/jlr.D065664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Battula VL, Treml S, Bareiss PM, Gieseke F, Roelofs H, de Zwart P, et al. Isolation of functionally distinct mesenchymal stem cell subsets using antibodies against CD56, CD271, and mesenchymal stem cell antigen-1. Haematologica. 2009;94(2):173–184. doi: 10.3324/haematol.13740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Kohli N, Al-Delfi IRT, Snow M, Sakamoto T, Miyazaki T, Nakajima H, et al. CD271-selected mesenchymal stem cells from adipose tissue enhance cartilage repair and are less angiogenic than plastic adherent mesenchymal stem cells. Sci Rep. 2019;9(1):3194. doi: 10.1038/s41598-019-39715-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Espagnolle N, Guilloton F, Deschaseaux F, Gadelorge M, Sensebe L, Bourin P. CD146 expression on mesenchymal stem cells is associated with their vascular smooth muscle commitment. J Cell Mol Med. 2014;18(1):104–114. doi: 10.1111/jcmm.12168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Morrison SJ, Scadden DT. The bone marrow niche for haematopoietic stem cells. Nature. 2014;505(7483):327–334. doi: 10.1038/nature12984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Yang ZX, Han ZB, Ji YR, Wang YW, Liang L, Chi Y, et al. CD106 identifies a subpopulation of mesenchymal stem cells with unique immunomodulatory properties. PLoS One. 2013;8(3):e59354. doi: 10.1371/journal.pone.0059354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Via AG, Frizziero A, Oliva F. Biological properties of mesenchymal stem cells from different sources. Muscles Ligaments Tendons J. 2012;2(3):154–162. [PMC free article] [PubMed] [Google Scholar]
- 73.Chan CKF, Gulati GS, Sinha R, Tompkins JV, Lopez M, Carter AC, et al. Identification of the human skeletal stem cell. Cell. 2018;175(1):43–56. doi: 10.1016/j.cell.2018.07.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Merrick D, Sakers A. Identification of a mesenchymal progenitor cell hierarchy in adipose tissue. Science. 2019;364 (6438):eaav2501. [DOI] [PMC free article] [PubMed]
- 75.Majore I, Moretti P, Hass R, Kasper C. Identification of subpopulations in mesenchymal stem cell-like cultures from human umbilical cord. Science. 2009;7:6. doi: 10.1186/1478-811X-7-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Marklein RA, Klinker MW, Drake KA, Polikowsky HG, Lessey-Morillon EC, Bauer SR. Morphological profiling using machine learning reveals emergent subpopulations of interferon-gamma-stimulated mesenchymal stromal cells that predict immunosuppression. Cytotherapy. 2019;21(1):17–31. doi: 10.1016/j.jcyt.2018.10.008. [DOI] [PubMed] [Google Scholar]
- 77.Klinker MW, Marklein RA, Lo Surdo JL, Wei CH, Bauer SR. Morphological features of IFN-gamma-stimulated mesenchymal stromal cells predict overall immunosuppressive capacity. Proc Natl Acad Sci U S A. 2017;114(13):E2598–607. [DOI] [PMC free article] [PubMed]
- 78.Marklein RA, Lo Surdo JL, Bellayr IH, Godil SA, Puri RK, Bauer SR. High content imaging of early morphological signatures predicts long term mineralization capacity of human mesenchymal stem cells upon osteogenic induction. Stem Cells. 2016;34(4):935–947. doi: 10.1002/stem.2322. [DOI] [PubMed] [Google Scholar]
- 79.Pevsner-Fischer M, Levin S, Zipori D. The origins of mesenchymal stromal cell heterogeneity. Stem Cell Rev. 2011;7(3):560–568. doi: 10.1007/s12015-011-9229-7. [DOI] [PubMed] [Google Scholar]
- 80.Zhang J, Huang X, Wang H, Liu X, Zhang T, Wang Y, et al. The challenges and promises of allogeneic mesenchymal stem cells for use as a cell-based therapy. Stem Cell Res Ther. 2015;6:234. doi: 10.1186/s13287-015-0240-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Huang Y, Li Q, Zhang K, Hu M, Wang Y, Du L, et al. Single cell transcriptomic analysis of human mesenchymal stem cells reveals limited heterogeneity. Cell Death Dis. 2019;10(5):368. [DOI] [PMC free article] [PubMed]
- 82.Wang Y, Barrett AN, Fong CY, Subramanian A, Liu W, Feng Y, et al. Human Wharton’s jelly mesenchymal stem cells show unique gene expression compared with bone marrow mesenchymal stem cells using single-cell RNA-sequencing. Cell Death Dis. 2019;28(3):196–211. doi: 10.1089/scd.2018.0132. [DOI] [PubMed] [Google Scholar]
- 83.Khong SML, Lee M, Kosaric N, Khong DM, Dong Y, Hopfner U, et al. Single-cell transcriptomics of human mesenchymal stem cells reveal agerelated cellular subpopulation depletion and impaired regenerative function. Stem Cells. 2019;37(2):240–6. [DOI] [PMC free article] [PubMed]
- 84.Gurtner GC, Hwang B, Lee JH, Bang D. Single-cell RNA sequencing technologies and bioinformatics pipelines. Stem Cells. 2018;50(8):96. doi: 10.1038/s12276-018-0071-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Olivares-Navarrete R, Rodil SE, Hyzy SL, Dunn GR, Almaguer-Flores A, Schwartz Z, et al. Role of integrin subunits in mesenchymal stem cell differentiation and osteoblast maturation on graphitic carbon-coated microstructured surfaces. Biomaterials. 2015;51:69–79. doi: 10.1016/j.biomaterials.2015.01.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Di Maggio N, Martella E, Frismantiene A, Resink TJ, Schreiner S, Lucarelli E, et al. Extracellular matrix and alpha5beta1 integrin signaling control the maintenance of bone formation capacity by human adipose-derived stromal cells. Sci Rep. 2017;7:44398. doi: 10.1038/srep44398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Marie PJ. Targeting integrins to promote bone formation and repair. Nat Rev Endocrinol. 2013;9(5):288–295. doi: 10.1038/nrendo.2013.4. [DOI] [PubMed] [Google Scholar]
- 88.Park SJ, Gadi J, Cho KW, Kim KJ, Kim SH, Jung HS, et al. The forkhead transcription factor Foxc2 promotes osteoblastogenesis via up-regulation of integrin beta1 expression. Bone. 2011;49(3):428–438. doi: 10.1016/j.bone.2011.05.012. [DOI] [PubMed] [Google Scholar]
- 89.Wu Y, Chen L, Scott PG, Tredget EE. Mesenchymal stem cells enhance wound healing through differentiation and angiogenesis. Stem Cells. 2007;25(10):2648–2659. doi: 10.1634/stemcells.2007-0226. [DOI] [PubMed] [Google Scholar]
- 90.Watt SM, Gullo F, van der Garde M, Markeson D, Camicia R, Khoo CP, et al. The angiogenic properties of mesenchymal stem/stromal cells and their therapeutic potential. Br Med Bull. 2013;108:25–53. doi: 10.1093/bmb/ldt031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Hua J, He ZG, Qian DH, Lin SP, Gong J, Meng HB, et al. Angiopoietin-1 gene-modified human mesenchymal stem cells promote angiogenesis and reduce acute pancreatitis in rats. Int J Clin Exp Pathol. 2014;7(7):3580–3595. [PMC free article] [PubMed] [Google Scholar]
- 92.Iinuma S, Aikawa E, Tamai K, Fujita R, Kikuchi Y, Chino T, et al. Transplanted bone marrow-derived circulating PDGFRalpha+ cells restore type VII collagen in recessive dystrophic epidermolysis bullosa mouse skin graft. J Immunol. 2015;194(4):1996–2003. doi: 10.4049/jimmunol.1400914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Lehman N, Cutrone R, Raber A, Perry R, Van't Hof W, Deans R, et al. Development of a surrogate angiogenic potency assay for clinical-grade stem cell production. Cytotherapy. 2012;14(8):994–1004. doi: 10.3109/14653249.2012.688945. [DOI] [PubMed] [Google Scholar]
- 94.Galipeau J, Krampera M, Barrett J, Dazzi F, Deans RJ, DeBruijn J, et al. International Society for Cellular Therapy perspective on immune functional assays for mesenchymal stromal cells as potency release criterion for advanced phase clinical trials. Cytotherapy. 2016;18(2):151–159. doi: 10.1016/j.jcyt.2015.11.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Hocking AM. The role of chemokines in mesenchymal stem cell homing to wounds. Adv Wound Care (New Rochelle) 2015;4(11):623–630. doi: 10.1089/wound.2014.0579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Mo M, Wang S, Zhou Y, Li H, Wu Y. Mesenchymal stem cell subpopulations: phenotype, property and therapeutic potential. Cell Mol Life Sci. 2016;73(17):3311–3321. doi: 10.1007/s00018-016-2229-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Cheng Z, Ou L, Zhou X, Li F, Jia X, Zhang Y, et al. Targeted migration of mesenchymal stem cells modified with CXCR4 gene to infarcted myocardium improves cardiac performance. Mol Ther. 2008;16(3):571–579. doi: 10.1038/sj.mt.6300374. [DOI] [PubMed] [Google Scholar]
- 98.Gentile P, Scioli MG, Bielli A, Orlandi A, Cervelli V. Concise review: the use of adipose-derived stromal vascular fraction cells and platelet rich plasma in regenerative plastic surgery. Stem Cells. 2017;35(1):117–134. doi: 10.1002/stem.2498. [DOI] [PubMed] [Google Scholar]
- 99.Latifi-Pupovci H, Kuci Z, Wehner S, Bonig H, Lieberz R, Klingebiel T, et al. In vitro migration and proliferation (“wound healing”) potential of mesenchymal stromal cells generated from human CD271(+) bone marrow mononuclear cells. J Transl Med. 2015;13:315. doi: 10.1186/s12967-015-0676-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Gentile P, Calabrese C, De Angelis B, Pizzicannella J, Kothari A, Garcovich S. Impact of the different preparation methods to obtain human adipose-derived stromal vascular fraction cells (AD-SVFs) and human adipose-derived mesenchymal stem cells (AD-MSCs): enzymatic digestion versus mechanical centrifugation. Int J Mol Sci. 2019;20(21):5471. [DOI] [PMC free article] [PubMed]
- 101.Gentile P, Piccinno MS, Calabrese C. Characteristics and Potentiality of Human Adipose-Derived Stem Cells (hASCs) Obtained from Enzymatic Digestion of Fat Graft. Cells. 2019;8(3):E282. [DOI] [PMC free article] [PubMed]
- 102.Scioli MG, Bielli A, Gentile P, Cervelli V, Orlandi A. Combined treatment with platelet-rich plasma and insulin favours chondrogenic and osteogenic differentiation of human adipose-derived stem cells in three-dimensional collagen scaffolds. J Tissue Eng Regen Med. 2017;11(8):2398–2410. doi: 10.1002/term.2139. [DOI] [PubMed] [Google Scholar]
- 103.Gentile P, Scioli MG, Orlandi A, Cervelli V. Breast reconstruction with enhanced stromal vascular fraction fat grafting: what is the best method? Plast Reconstr Surg Glob Open. 2015;3(6):e406. doi: 10.1097/GOX.0000000000000285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Gentile P. Autologous cellular method using micrografts of human adipose tissue derived follicle stem cells in androgenic alopecia. Int J Mol Sci. 2019;20(14):3446. [DOI] [PMC free article] [PubMed]
- 105.Gentile P, Garcovich S. Advances in regenerative stem cell therapy in androgenic alopecia and hair loss: Wnt pathway, growth-factor, and mesenchymal stem cell signaling impact analysis on cell growth and hair follicle development. Cells. 2019;8(5):E466. [DOI] [PMC free article] [PubMed]
- 106.Gentile P, Kothari A, Casella D, Calabrese C. Fat graft enhanced with adipose-derived stem cells in aesthetic breast augmentation: clinical, histological, and instrumental evaluation. Aesthet Surg J. 2019. Epub ahead of print. [DOI] [PubMed]
- 107.Hematti P. Characterization of mesenchymal stromal cells: potency assay development. Transfusion (Paris) 2016;56(4):32s–35s. doi: 10.1111/trf.13569. [DOI] [PubMed] [Google Scholar]
- 108.Galipeau J, Krampera M. The challenge of defining mesenchymal stromal cell potency assays and their potential use as release criteria. Cytotherapy. 2015;17(2):125–127. doi: 10.1016/j.jcyt.2014.12.008. [DOI] [PubMed] [Google Scholar]
- 109.Gentile P, Scioli MG, Bielli A, De Angelis B, De Sio C, De Fazio D, et al. Platelet-rich plasma and micrografts enriched with autologous human follicle mesenchymal stem cells improve hair re-growth in androgenetic alopecia. biomolecular pathway analysis and clinical evaluation. Biomedicines. 2019;7(2):E27. [DOI] [PMC free article] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: Supplemental methods describing scRNA-seq data analysis for detail, including methods about Quality control, Removal of cell cycle effect, Highly Variable genes identification, Linear and nonlinear dimension reduction, Clustering the cells and Differential expression analysis.
Additional file 2: Supplemental tables with six tables. Table S1. Marker genes used for potency score analysis. Table S2. Top100 CD genes ranked by mean expression in WJMSCs and ADMSCs. Table S3. CD genes ranked by percentage of cells expressed the genes (at least one UMI). Table S4. List of HVGs identified in WJMSCs. Table S5. DEGs in each cluster by comparing it to all of the others. Table S6. Results of differential gene expression analysis between two different clusters.
Additional file 3: Supplemental figures with five figures. Figure S1. Quality of the WJMSCs single-cell RNA-seq data. (A) Number of reads were sequenced for each of the three samples. Percentage of reads mapped to exonic (B) and mapped to transcriptome (C) for each of the three samples. (D) Number of cells obtained for each of the three samples. Boxplot showing number of expressed genes per cell (E) and number of UMI per cell (F) for each of the three samples. (G) Tri-lineage differentiation potency of primary cultured WJMSCs used for scRNA-seq. Figure S2. Highly variable genes identification in WJMSCs and GO enrichment analysis. (A) Venn diagram showing overlap of top 2000 highly variable genes among different phases for sample UC1. (B) Venn diagram showing overlap of top 2000 highly variable genes among different phases for sample UC2. (C) Venn diagram showing overlap of top 2000 highly variable genes among different phases for sample UC3. (D) Venn diagram showing overlap of highly variable genes among samples. Results of GO-slim cellular component enrichment analysis (E), GO-slim biological process enrichment analysis (F), and GO-slim functional molecular enrichment analysis for highly variable genes. Figure S3. Candidate subpopulations identified in WJMSCs. (A) and (B) UMAP showing dimension reduction before and after batch (A) and cell cycle effect (B) removal. Left, before removal; right, after removal. (C) Histogram showing number of cells for each phase of cell cycle and sample in the candidate subpopulations. (D) Violin plots showing distribution of log normalized expression (log (norm_exprs)) values of collagen genes across the six candidate subpopulations (C0–C5). (E) Violin plots showing distribution of log (norm_exprs) values of chemokines genes across the six candidate subpopulations (C0–C5). Figure S4. Wound healing potency for CD142+ and CD142− WJMSCs. (A) CD142 analysis by flow cytometry for WJMSCs. (B) Example of gate setting for CD142− (left gate) and CD142+ (right gate) cells sorting. (C) qPCR-based expression fold-changes for genes upregulated in C3 plus CCL2, CXCL8 and MKI67 (n = 3) between CD142+ and CD142− cells. (D) Proliferation for CD142− and CD142+ cells cultured in vitro. n = 4 for each time point. (E) Representative images of wound healing assays for conditioned media from CD142- and CD142+ cells cultured fibroblast, respectively. (F) Wound closure comparison between CD142+ and CD142− cells conditioned media for 24 h (n = 9). Data shown are means ± SD, *p < 0.05. Paired two-tailed Student’s t-test were performed for significant test. Figure S5. Differentiation potency compared between ADMSCs and WJMSCs. Density distribution showing chondrogenic score(A), neurogenic score (B), myogenic score(C), and housekeeping score (D) between ADMSCs and WJMSCs. Percentage indicating proportion of cells assigned to the right side of the line. (E) Violin plots showing marker genes of SSC expressed in ADMSCs and WJMSCs. (F) Percentage of cells expressed SSC marker genes in ADMSCs and WJMSCs. (G) Violin plots showing marker genes of adipose progenitors expressed in ADMSCs and WJMSCs. (H) Percentage of cells expressed marker genes of adipose progenitors in ADMSCs and WJMSCs. Figure S6. Functional enrichment of highly variable genes identified in ADMSCs. (A) Venn diagram showing overlap of HVGs between ADMSCs and WJMSCs. Barplots showing results of GO-slim cellular component (B), GO-slim molecular function (C), Go-slim biological process (D), Reactome pathways (E), Pathways (F), and protein class (G) enrichment analysis for HVGs identified in ADMSCs.
Data Availability Statement
The data that support the findings of this study have been deposited in the CNSA (https://db.cngb.org/cnsa/) of CNGBdb with accession number CNP0000562. All data analyzed during this study have been included in this published article and its supplementary information files.