Abstract
Background
Toxoplasma gondii is the third most important contributor to health burden caused by food-borne illness. Ingestion of tissue cysts from undercooked meat is an important source of horizontal transmission to humans. However, there is an increasing awareness of the consumption of fresh fruit and vegetables, as a possible source for oocyst transmission, since this stage of the parasite can persist and remain infective in soil and water for long time. Herein, we outline findings related with detection of T. gondii oocysts in vegetables and berry fruits, which are usually raw consumed. The procedure includes the estimation of the number of oocysts.
Methods
Food samples were collected from local producers and supermarket suppliers. Toxoplasma gondii oocysts were concentrated after washing the samples by applying high resolution water filtration and immunomagnetic separation (method 1623.1: EPA 816-R-12-001-Jan 2012), in order to (i) remove potential Cryptosporidium spp. oocysts and Giardia spp. cysts present in the samples; and (ii) select T. gondii oocysts. Toxoplasma gondii oocyst detection and an estimation of their numbers was performed by conventional PCR and real time qPCR, using specific primers for a 183-bp sequence of the T. gondii repetitive DNA region. All PCR-positive DNA samples were purified and sequenced. Restriction enzyme digestion with EcoRV endonuclease confirmed the presence of the T. gondii DNA fragment. In addition, the presence of the parasite was observed by fluorescent microscopy, taking advantage of the oocysts autofluorescence under UV light.
Results
Forty percent of the analysed samples (95% CI: 25.5–56.5%) presented the expected PCR and digested DNA fragments. These fragments were confirmed by sequencing. Microscopic autofluorescence supported the presence of T. gondii-like oocysts. The estimated mean (± SE) oocyst concentration was 23.5 ± 12.1 oocysts/g, with a range of 0.6–179.9 oocysts/g.
Conclusions
Our findings provide relevant evidence of contamination of fresh vegetables and berry fruits with T. gondii oocysts.
Keywords: Toxoplasma gondii, Oocysts, Immunomagnetic separation, PCR, Vegetables, Berry fruits
Background
Toxoplasma gondii is an intracellular protozoan (Apicomplexa: Coccidia) causing human and animal toxoplasmosis [1]. Protozoan infectivity is due to three parasitic stages: an invasive tachyzoite; a bradyzoite in tissue cysts; and an environmental stage, the sporozoite, protected inside mature oocysts [1, 2]. The global prevalence of toxoplasmosis is estimated to be around 30% with 10 million clinical cases [1–4] and it is ranked as the third most important contributor to health burden caused by food-borne illness in Europe [4]. Most infections in humans are asymptomatic. However, severe complications may occur during (i) congenital Toxoplasma infection, such as abortion, stillbirth and hydrocephalus in new-borns [1, 4, 5]; (ii) ocular toxoplasmosis, with retinochoroidal lesions leading to chronic ocular disease [1, 2]; (iii) encephalitis in immunosuppressed patients [1, 4, 5]; and (iv) multivisceral toxoplasmosis due to atypical genotypes in South America [6–8]. A possible implication of T. gondii genetic diversity on the pathogenesis of toxoplasmosis has been postulated [9–11]. Consensually, current opinion is that the majority of horizontal transmissions to humans occurs after ingestion of tissue cysts in infected meat, or through water, raw fruit and vegetables contaminated with sporulated oocysts [2, 12, 13]. In vitro and in vivo experiments using mouse models have shown that infections due to oocysts are clinically more severe when compared to infections caused by tissue cysts (bradyzoites) [14]. However, the relative importance of transmission via tissue cysts versus oocysts is still unclear [2]. In addition, oocysts can remain viable for long periods in the environment and can resist chemical and physical treatment currently applied in water plants, including chlorination and ozone treatment [15–18]. This paved the way for an increasingly awareness related to drinking water and the consumption of raw fruit and vegetables as putative routes for oocyst transmission. So far, the detection of Toxoplasma oocysts in fruit and vegetables has been difficult, and no standardized methods are available. Moreover, findings outlined in the literature on this topic are scarce and/or controversial [19–26]. In this scenario, a laboratory approach was designed based on the experience gained with Method 1623.1/EPA for Cryptosporidium oocyst and Giardia cyst detection [27]. This approach involved: (i) the concentration of oocysts from large volumes of washing water (fruit and vegetables), according to Method 1623.1/EPA; and (ii) subsequent application of PCR for identification and quantification of T. gondii DNA [23, 26, 28–30]. Herein, we outline findings related with the detection of T. gondii oocysts in vegetables and berry fruits, as a contribution to a better comprehension of oocyst transmission to humans.
Methods
Food samples
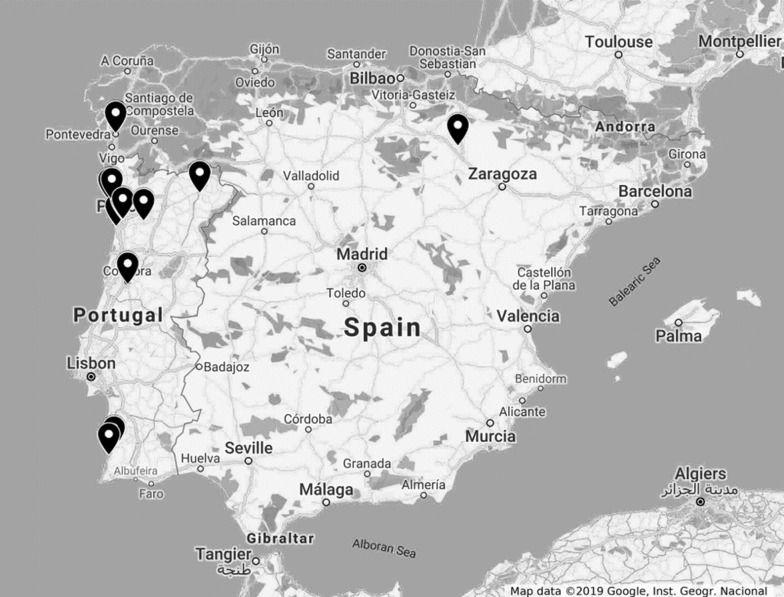
A total of 35 bulk, packaged and ready-to-eat (RTE) vegetables and berry fruits were collected from local producers, provided by retail sellers, or bought in small and large-scale supermarkets between July 2018 and July 2019, in several locations in Portugal and Spain (Fig. 1). The choice of samples was based on expected market preferences of raw fruit and vegetables by Portuguese consumers, described by the Government agency “Instituto Nacional de Estatística” [31]. The food products included different varieties of lettuce (Lactuca sativa), watercress (Nasturtium officinale), coriander (Coriandrum sativum), parsley (Petroselinum crispum) carrots (Daucus carota sativus), arugula (Eruca vesicaria sativa), strawberries (Fragaria ananassa), raspberries (Rubus idaeus) and blueberries (Vaccinium myrtillus). Ready-to-eat mixed salads, with different varieties of lettuce (Lactuca sativa), arugula (Eruca vesicaria sativa), endive (Cichorium endivia), chicória-italiana (Cichorium intybus), carrots (Daucus carota sativus), red cabbage (Brassica oleracea var. capitata f. rubra) and lambʼs lettuce (Valerianella locusta) were also included in this study. Laboratory sample treatment was performed immediately or in less than 24 h after refrigeration.
Fig. 1.
Geographical origin of the collected samples. Fruit and vegetables were collected from local producers, provided by retail sellers, or bought in small- and large-scale supermarkets in several locations of Portugal and Spain
Toxoplasma gondii oocyst recovery
The concentration and recovery of T. gondii oocysts, as well as Cryptosporidium spp. oocysts and Giardia spp. cysts from fruit and vegetable samples were performed by Filtration/Immunomagnetic Separation (IMS)/Fluorescence Assay (FA) (Method 1623.1: Cryptosporidium and Giardia in water; US EPA 816-R-12-001-Jan 2012) [27]. Briefly, the fruit and vegetables were vigorously washed by manual swirling, for at least 10 min in large volumes of distilled water (between 10 and 80 l) in 20, 50 or 100 l tanks, accordingly to the size of each sample. Ten litres of distilled water were used to wash an average of 440 g of sample. The weight of each sample varied between 64–3600 g (Table 1); for washing water filtration, a 1 µm Filta-Max® filter (IDEXX, Westbrook, ME, USA) applied to a peristaltic pump at three bar was used. Elution was performed in a Filta-Max® manual wash station, and concentrated into 3 ml of phosphate-buffered solution (PBS) with 0.01% of Tween 20 (Merck KGaA, Darmstadt, Germany), after centrifugation at 1000×g for 10 min at room temperature. Magnetic beads conjugated with specific antibodies (Dynabeads™ GC-Combo; Thermo Fisher Scientific, Waltham, MA, USA) were added to the concentrate, and magnetized Cryptosporidium oocysts and Giardia cysts were removed from the extraneous material using a magnet (data not presented). The remaining suspension, containing putative T. gondii (PTG) oocysts were centrifuged (1000×g for 10 min at room temperature) and resuspended in 2–5 ml of PBS.
Table 1.
Results of Toxoplasma gondii oocyst detection and quantification in the collected vegetables and fruits
| Sample number | Product | Weight (g) | Origin | Collection date | Agricultural system | Product presentation | DNAa | Enzymatic restriction confirmation | Sequence confirmation | Mean no. of oocysts/g |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | Lettuce | 3600 | Aguçadoura (Póvoa do Varzim) | 12-Jul-2018 | Conventional | Bulk | Positive | Yes | Yes | 1.3 |
| 2 | Carrot | 924 | Aguçadoura (Póvoa do Varzim) | 12-Jul-2018 | Conventional | Bulk | Negative | – | – | – |
| 3 | Parsley | 266 | Aguçadoura (Póvoa do Varzim) | 12-Jul-2018 | Conventional | Bulk | Negative | – | – | – |
| 4 | Lettuce | 261 | Aguçadoura (Póvoa do Varzim) | 10-Aug-2018 | Conventional | Bulk | Negative | – | – | – |
| 5 | Lettuce | 261 | Mabegondo-Abegondo (Galiza) | 13-Aug-2018 | Organic | Bulk | Positive | Yes | Yes | 3.6 |
| 6 | Lettuce | 600 | Gondomar (Porto) | 13-Aug-2018 | Organic | Bulk | Positive | Yes | Yes | 2.8 |
| 7 | Strawberry | 64 | São Félix da Marinha (Vila Nova de Gaia) | 05-Sep-2018 | Organic | Bulk | Positive | Yes | Yes | 8.6 |
| 8 | Watercress | 792 | Boavista dos Pinheiros (Odemira) | 27-Sep-2018 | Conventional | Bulk | Positive | Yes | Yes | 0.6 |
| 9 | Lettuce | 1246 | Boavista dos Pinheiros (Odemira) | 27-Sep-2018 | Conventional | Bulk | Positive | Yes | Yes | 1.1 |
| 10 | Lettuce | 2031 | Boavista dos Pinheiros (Odemira) | 27-Sep-2018 | Conventional | Bulk | Negative | – | – | – |
| 11 | Coriander | 749 | Boavista dos Pinheiros (Odemira) | 27-Sep-2018 | Conventional | Bulk | Negative | – | – | – |
| 12 | Parsley | 632 | Boavista dos Pinheiros (Odemira) | 27-Sep-2018 | Conventional | Bulk | Negative | – | – | – |
| 13 | Watercress | 900 | Boavista dos Pinheiros (Odemira) | 02-Oct-2018 | Conventional | RTE | Negative | – | – | – |
| 14 | Mix salad | 800 | Boavista dos Pinheiros (Odemira) | 02-Oct-2018 | Conventional | RTE | Negative | – | – | – |
| 15 | Coriander | 450 | Boavista dos Pinheiros (Odemira) | 02-Oct-2018 | Conventional | Packaged | Positive | Yes | Yes | 3.9 |
| 16 | Parsley | 450 | Boavista dos Pinheiros (Odemira) | 02-Oct-2018 | Conventional | Packaged | Positive | Yes | Yes | 44.5 |
| 17 | Watercress | 900 | Boavista dos Pinheiros (Odemira) | 12-Oct-2018 | Conventional | RTE | Positive | Yes | Yes | 3.2 |
| 18 | Mixed salad | 700 | Boavista dos Pinheiros (Odemira) | 12-Oct-2018 | Conventional | RTE | Positive | Yes | Yes | 6.0 |
| 19 | Coriander | 350 | Boavista dos Pinheiros (Odemira) | 12-Oct-2018 | Conventional | Packaged | Positive | Yes | Yes | 38.3 |
| 20 | Parsley | 350 | Boavista dos Pinheiros (Odemira) | 12-Oct-2018 | Conventional | Packaged | Negative | – | – | – |
| 21 | Mixed salad | 400 | Boavista dos Pinheiros (Odemira) | 04-Feb-2019 | Conventional | RTE | Positive | Yes | Yes | 179.9 |
| 22 | Lettuce | 494 | Torre de Bera (Coimbra) | 04-Mar-2019 | Organic | Bulk | Negative | – | – | – |
| 23 | Lettuce | 1100 | Torre de Bera (Coimbra) | 04-Mar-2019 | Organic | Bulk | Negative | – | – | – |
| 24 | Strawberry | 500 | Macedo de Cavaleiros (Bragança) | 27-May-2019 | Conventional | Bulk | Positive | Yes | Yes | 2.5 |
| 25 | Strawberry | 500 | Macedo de Cavaleiros (Bragança) | 27-May-2019 | Conventional | Bulk | Positive | Yes | Yes | 3.1 |
| 26 | Mixed salad | 800 | Milagro (Navarra) | 17-Jun-2019 | Conventional | RTE | Negative | – | – | – |
| 27 | Mixed salad | 800 | Milagro (Navarra) | 17-Jun-2019 | Conventional | RTE | Negative | – | – | – |
| 28 | Arugula | 500 | Milagro (Navarra) | 17-Jun-2019 | Conventional | RTE | Negative | – | – | – |
| 29 | Raspberry | 500 | Brejão (Odemira) | 17-Jun-2019 | Conventional | Packaged | Negative | – | – | – |
| 30 | Blueberry | 500 | Laundos (Póvoa do Varzim) | 17-Jun-2019 | Conventional | Packaged | Negative | – | – | – |
| 31 | Lettuce | 250 | São Pedro da Cova (Gondomar) | 18-Jun-2019 | Organic | Bulk | Negative | – | – | – |
| 32 | Lettuce | 270 | São Pedro da Cova (Gondomar) | 18-Jun-2019 | Organic | Bulk | Positive | Yes | Yes | 53.3 |
| 33 | Mixed salad | 800 | Milagro (Navarra) | 15-Jul-2019 | Conventional | RTE | Negative | – | – | – |
| 34 | Blueberry | 625 | Santa Leucádia (Baião) | 15-Jul-2019 | Organic | Packaged | Negative | – | – | – |
| 35 | Raspberry | 625 | Santa Leucádia (Baião) | 15-Jul-2019 | Conventional | Packaged | Negative | – | – | – |
a529-bp REP
Abbreviations: REP: repetitive region; RTE: ready-to-eat
Toxoplasma gondii DNA detection and oocysts number estimation
DNA extraction
Two-hundred microliters of PTG suspension was centrifuged at 1000×g for 10 min at room temperature and the pellet resuspended in 200 ml lysis buffer (QIAamp® DNA Mini Kit; Qiagen, Hilden, Germany). The disruption of the oocyst cell wall was performed with 4 freeze (− 20 °C)/thaw (95 °C) cycles. Prior to overnight incubation at 56 °C with 20 µl proteinase K (Qiagen), samples were treated with 1 ml of InhibitEX buffer (Qiagen) that efficiently removes PCR inhibitors commonly present in environmental samples. The commercial kit, QIAamp® DNA Mini Kit (Qiagen), was used for DNA isolation according to the manufacturer’s instructions.
Conventional PCR and sequencing
Specific primers (FW: 5′-AGC CAC AGA AGG GAC AGA AG-3′ and REV: 5′-TCC AGG AAA AGC AGC CAA G-3′) targeting a 183-bp sequence of the 529-bp repetitive sequence of T. gondii [23, 26, 28–30] were designed (Primer3Plus and BLAST®) for DNA detection according to the MIQE (minimum information for publication of quantitative real-time PCR experiments) guidelines [32, 33]. These primers do not bind to any site of the H. hammondi 529-bp repetitive sequence (GenBank: EU493285.1), and no significant similarity was found between the primers’ nucleotides and H. hammondi repetitive sequence. PCR amplification was performed with an initial polymerase activation step (3 min at 95 °C), followed by 35 cycles of denaturation (30 s at 95 °C), annealing (30 s at 62 °C) and extension (30 s at 72 °C), followed by a final extension step of 10 min at 72 °C. The amplification reaction mixture consisted of 12.5 μl DreamTaq™ Hot Start Green PCR Master Mix 2× (Thermo Fisher Scientific), 600 nM of each primer (Eurofins Genomics, Ebersberg, Germany) and 5 μl of template DNA in a 25 μl reaction volume. Amplifications were performed in a C1000 Touch™ thermal cycler (Bio-Rad, Hercules, CA, USA). PCR products (183 bp) were observed in a Gel DocTM XR+ (Bio-Rad) and analysed using Image LabTM Software (Bio-Rad). In all PCR experiments, a positive control (genomic DNA isolated from 105T. gondii ME49 oocysts) and a negative control (water template) were used. Additionally, all of the negative samples were retested for the presence of PCR inhibitors by adding 1 μl of T. gondii-positive control to the 5 μl of DNA template. Positive PCR fragments were purified (IllustraTM GFXTM PCR DNA and Gel Band Purification kit; GE HealthCare, IL, Chicago, USA) from a low melting 2% agarose gel (Lonza, Basel, Switzerland) and sequenced using Sanger sequencing services from GATC Biotech (Eurofins Genomics, Ebersberg, Germany). Sequence comparison was made with already published sequences (GenBank: AF146527.1) using the NCBI Basic Local Alignment Search Tool (BLAST).
Quantitative real-time PCR
A quantitative (qPCR) method based on a standard curve was used to estimate the initial number of oocysts per gram of vegetable or fruit. Briefly, a standard template was obtained after purification of the positive control (T. gondii ME49 oocysts) PCR product and DNA concentration was measured with a NanoDrop 1000 spectrophotometer (ThermoFisher Scientific, Waltham, MA, USA). The DNA copy number/µl was calculated and two series of a ten-fold serial dilution of the DNA (from 1 × 106 to 1 copy/μl) were prepared. qPCR was carried out in a final volume of 20 µl, using 10 μl of SsoAdvanced™ Universal SYBR® Green Supermix (2×) (Bio-Rad) and 350 nM of each specific primer (previously used for conventional PCR) and 2 μl of the DNA sample or standard dilution. The plates were sealed and centrifuged at 1000×g for 1 min, at room temperature. The cycling conditions for the qPCR CFX96 Touch real-time instrument (Bio-Rad) were as follows: initial denaturation at 98 °C for 3 min; followed by 35 cycles at 98 °C for 15 s; then 62 °C for 30 s. Melting curve analysis was also performed at the end of each PCR run (65–95 °C at 0.5 °C/5 s). qPCR of the standard dilution gradient and the DNA isolated from fruit and vegetable samples were run in duplicate and in the same plate. The absolute quantification of the initial DNA copy number/μl and estimation of the number of oocysts per gram of each environmental sample was calculated by comparing the quantification cycle threshold (Cq) values of the environmental samples with the standard curve taking into account the variation of the 529-bp repetitive sequence and the ploidy of the T. gondii oocysts.
Restriction enzyme digestion
Purified positive PCR fragments were subject to restriction enzyme digestion with the endonuclease EcoRV (New England Biolabs, Ipswich, MA, USA). The reaction was performed in a volume of 10 μl with 5 μl of DNA, 1 μl of the enzyme and 1 μl of 10× NEBuffer 3.1, according to the manufacturer’s instructions. Each sample was incubated at 37 °C for 1 h. Additionally, purified T. gondii PCR fragments and uncut experimental DNA were used as controls. The digested fragments (74 bp and 109 bp) were electrophoresed on a 2% high-resolution agarose gel (Sigma-Aldrich) and analysed using a Gel DocTM XR+ (Bio-Rad).
Epifluorescence microscopy
Twenty-five microliters of the PTG suspension was diluted (1:2) and air-dried onto a 2-well SuperStick™ Slide (Waterborne™, Inc., New Orleans, LA, USA) chemically treated to increase adhesion of the oocysts. Fixation was performed for 2 min with absolute methanol (Panreac Química SLU, Castellar del Vallès, Spain) and the slides were allowed to dry completely. As a positive control, a slide was prepared with 105T. gondii ME49 oocysts (kindly provided by J. T. Dubey). The background fluorescence was reduced by adding Evans Blue dye solution (BlockOut Counterstain; Waterborne™, Inc.), slides were rinsed with PBS and completely air-dried before being mounted with DPX mounting media (Merck KGaA, Darmstadt, Germany). Oocysts were visualized using a Nikon fluorescence microscope (Nikon Optiphot 2; Melville, NY, USA), under bright field and ultraviolet (UV) filter block (excitation 335 nm; emission 450 nm), based on the autofluorescent nature of the oocyst wall due to Tyr-rich proteins [18, 34]. Images were captured at 400× magnification using a PowerShot A630 digital camera (Canon, Amstelveen, Holland).
Statistical analysis
Graphpad Prism version 7.02 (San Diego, CA, USA) was used for statistical analysis. The 95% confidence intervals (95% CI) for the population proportions were calculated using the modified Wald method [35]. Fisher’s exact test was used to find out differences for the different categorical variables. The non-parametric Mann-Whitney U-test was used to compare the number of oocysts per gram in each food group. A two-tailed P-value of less than 0.05 was considered significant.
Results
Sampling
A total of 25 kg of fresh products distributed in 35 samples were analysed and results are shown in Table 1. These comprised 18 bulk samples, corresponding to 14.5 kg and 17 samples of packaged and RTE corresponding to 10.5 kg; 28 samples (21.7 kg) of fresh vegetables (lettuce, watercress, parsley, coriander, arugula, carrots and mixed salads) and 7 (3.3 kg) of berry fruits (strawberry, raspberry and blueberry); 27 samples (21.3 kg) were provided from conventional agriculture systems and 8 samples (3.7 kg) from organic agriculture.
Toxoplama gondii DNA detection
Toxoplasma gondii detection was performed by conventional PCR and a 183-bp gel band corresponding to the specific DNA fragment of the T. gondii repetitive region (GenBank: AF146527.1) was observed in 14 out of the 35 (40.0%) analysed samples (95% CI: 6.5–25.5%). Statistical analysis of contingency tables showed no significant differences (P > 0.05) for the different categorical variables, such us the type of products, the season when they were collected, the type of agriculture system applied and Bulk or RTE products (Table 2). Sequence analysis confirmed that the obtained amplicon of all positive samples had more than 95% nucleotide similarity with the 529-bp T. gondii repetitive region (GenBank: AF146527.1) (Table 1). Moreover, two bands of expected sizes (109 bp and 74 bp) were observed after EcoRV DNA enzymatic digestion (Fig. 2), confirming T. gondii DNA sequence.
Table 2.
Prevalence rate of Toxoplasma gondii in different groups of fruit and vegetables
| Samples | Toxoplasma gondii DNA detection | P-value | |
|---|---|---|---|
| Negative (n, %) | Positive (n, %) | ||
| Total | 21 (60.0) | 14 (40.0) | |
| Product presentation | |||
| Bulk | 9 (50.0) | 9 (50.0) | P = 0.499 |
| RTE/packaged | 11 (64.7) | 6 (35.3) | |
| Type of product | |||
| Vegetables | 16 (57.1) | 12 (42.9) | P = 1.000 |
| Berry fruits | 4 (57.1) | 3 (42.9) | |
| Agricultural system | |||
| Conventional | 16 (59.3) | 11 (40.7) | P = 0.700 |
| Organic | 4 (50.0) | 4 (50.0) | |
| Season | |||
| Spring/summer | 12 (63.2) | 7 (36.8) | P = 0.506 |
| Autumn/winter | 8 (50.0) | 8 (50.0) | |
Notes: Contingency tables were constructed in order to compare categorical variables. The Fisher’s exact test was used and P > 0.05 indicates non-significant associations
Abbreviations: n, number of samples; %, percentage
Fig. 2.
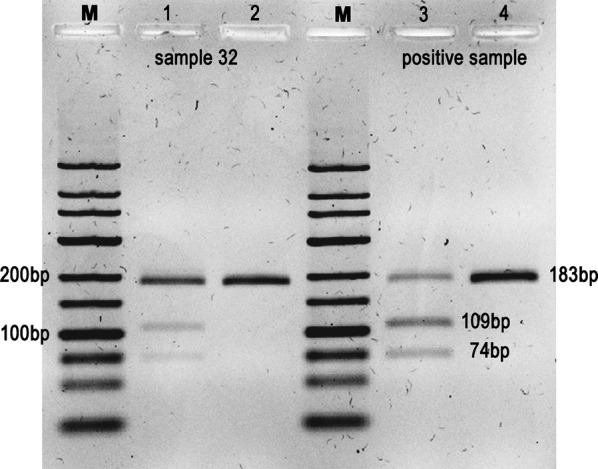
Representative image of a 2% high resolution agarose gel with T. gondii uncut, positive, 183 bp PCR fragments (Lane 2 and Lane 4) and subject to restriction enzyme digestion with the endonuclease EcoRV (Lane 1 and Lane 3). A ready-to-use DNA marker (Lanes M) suitable for sizing linear double-stranded DNA fragments from 25–700 bp was used to confirm the expected digested fragments of 74 bp and 109 bp
Estimation of the number of T. gondii oocysts
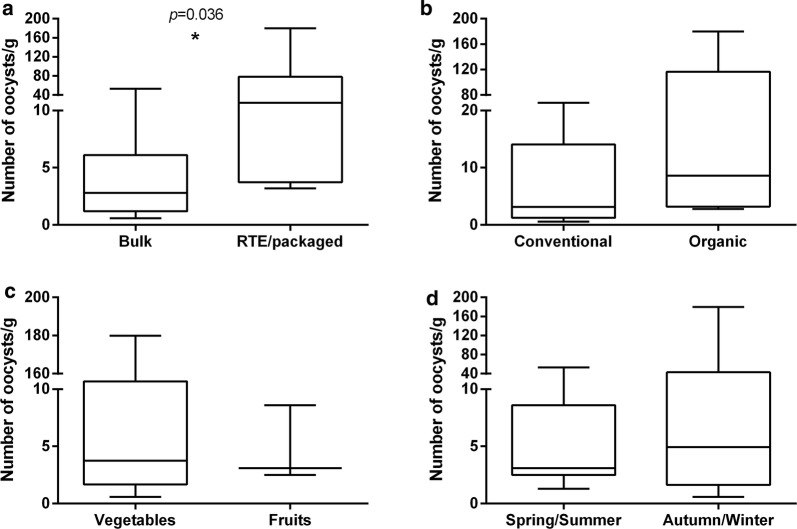
Real-time qPCR was used to estimate the initial number of oocysts present in each gram of vegetable or fruit. Quantification was performed by using as a standard, a T. gondii DNA gradient of a known initial number of DNA copies/µl. Amplification efficiencies were calculated and were always between 90–110%. In addition, melt curve analysis allowed confirmation of the specific T. gondii amplified sequence. The diagnostic melting temperature (Tm) peak was 86.5 ± 1 °C. The concentration of Toxoplasma from the fruit and vegetables varied between 0.6–179.9 oocysts/g (Table 1), with a mean number (± SE) of 23.5 ± 12.1 oocysts/g. A significantly higher number of oocysts/g was found from RTE/packaged samples (45.9 ± 27.8 oocysts/g) when compared with the mean number of oocysts/g present in bulk products (8.5 ± 5.6 oocyst/g) (Mann-Whitney U-test: P = 0.036) (Fig. 3a). No statistically significant differences (P > 0.05) were found when comparing the number of oocysts/g in products originally from conventional or organic agriculture, between fruit and vegetables or the season when the products were collected (Fig. 3b–d).
Fig. 3.
Box and whiskers diagrams comparing the estimated number of oocysts per gram, between bulk and RTE/packaged products (a), conventional and organic agriculture system (b), fruit and vegetables (c) and food items collected in spring/summer and autumn/winter (d), quantified by qPCR. The non-parametric Mann-Whitney U-test was used to compare the estimated number of oocysts per gram between the groups. *P = 0.036
Toxoplasma gondii oocysts autofluorescence
Direct visualization of T. gondii oocyst autofluorescence was performed using an epifluorescent microscope. In slides from all PCR-positive samples we observed T. gondii-like oocysts, based on morphology and size (between 10–15 µm in diameter), and the pale blue glow when illuminated with a UV light source. The majority of the samples presented unsporulated oocysts. However, sporulated oocysts were observed in one of the strawberry samples.
Discussion
Water and food matrices may be accidentally contaminated by environmental oocysts [12, 13]. Thus far, there are no standard methods or consensual tools described for oocyst identification and quantification from these matrices. The lack of standardised methods probably sustains the exclusion of T. gondii from regular surveillance systems [12, 13, 26]. In order to estimate the prevalence of T. gondii oocysts in fresh fruit and vegetables, we have designed an approach described herein. For molecular detection of the parasite, we have selected a consensual specific DNA fragment of the T. gondii repetitive region [28]. The 529-bp DNA fragment was amplified in 40% of all samples, with an estimated mean of 23.5 oocysts per gram of product. Our findings indicate a higher prevalence compared to other studies [19–26]. However, critical analysis of the data in Table 3, evidences major differences related to: (i) sampling design; (ii) DNA target strategy; and of relevance to this particular study (iii) the choice of oocyst recovery method. In our opinion, the choice of the Filta-Max® System (IDEXX), for high-resolution filtration of large volumes of washing water, increases oocyst recovery. Consequently, it increases the sensitivity of the detection method. Complementary to this, the execution of Method 1623.1 [27] seems to be an advisable strategy for removing Cryptosporidium spp. oocysts and Giardia spp. cysts and separating them from potential T. gondii oocysts present in the sample washing water. Indeed, high-grade filtration followed by immunomagnetic separation of water samples has been described as a more accurate method for parasite concentration when compared with membrane filtration and centrifugation [36] or sucrose gradient separation [37].
Table 3.
Current available data on the detection of Toxoplasma gondii oocysts in fresh fruit and vegetables, under non-experimental conditions
| References | Product | Origin | Sample size | Oocyst recovery method | Detection method | PCR target | T. gondii prevalence (%) | No. of oocysts | Sequence confirmation | Genotyping analysis |
|---|---|---|---|---|---|---|---|---|---|---|
| Al-Megrin (2010) [19] | Spinach, radish, leek, parsley, basil, green onion, dill, lettuce, cabbage, watercress, coriander, mint | Saudi Arabia |
n = 470 (250 g) |
Washing, centrifugation | Microscopy | na | 6.6 | nd | No | No |
| Lass et al. (2012) [20] | Lettuce, strawberries, carrots, radishes | Poland |
n = 216 One lettuce; 1000 g strawberries; 500 g carrots; 20 radishes |
Washing, centrifugation, flocculation (CaCO3) | Real-time qPCR | B1 gene | 9.7 | 1–20/sample | No | Yes (type I and II) |
| Haq et al. (2014) [21] | Cabbage, carrot, chili, coriander, cucumber, lettuce, mint, radish, tomato | Pakistan |
n = 500 (200–250 g) |
Washing, centrifugation, sedimentation or flotation | Microscopy | na | 1.9 | nd | No | No |
| Lalonde & Gajadhar (2016) [22] | Pre-packaged or bulk spinach, lettuce, chard, collards, dandelion greens, rapini, mixed salads | Canada |
n = 470 (250 g) |
Washing, centrifugation, flotation (Sheather’s sucrose solution) | Real-time PCR | 18S rDNA | 0.26 | nd | Yes | No |
| Marchioro et al. (2016) [23] | Organic and nonorganic: crisp lettuce, regular lettuce, chicory, rocket, parsley | Brazil |
n = 1171 (35 g) |
Washing, filtration (0.3 mm cellulose ester membrane), centrifugation | Conventional PCR | B1 gene/ 529 bp REP | 2.9/0.8 | nd | No | No |
| Caradonna et al. (2016) [24] | Ready-to-eat packaged salads | Italy |
n = 238 (50 g) |
Washing, centrifugation | Microscopy, qPCR, sequencing | B1 gene | 0.8 | 62–554/g | Yes | Yes (type I) |
| Ferreira et al. (2018) [25] | Crisp lettuce, arugula, chicory, chives, purple lettuce, spinach, chard | Brazil |
n = 470 (250 g) |
Washing, filtration (two layers of gauze), centrifugation | Conventional PCR | 529 bp REP | 12.9 | nd | No | No |
| Slany et al. (2019) [26] | Carrot, cucumber, lettuce | Czech Republic |
n = 648 (100 g) |
Washing, centrifugation | Triplex qPCR | B1 gene and 529 bp REP | 9.6 | 1–900/g | No | Type II |
| Present study | RTE/packaged and bulk (organic and nonorganic: lettuce, watercress, coriander, parsley, carrots, arugula, strawberries, raspberries, blueberries, mixed salads) | Portugal, Spain |
n = 35 (64–3600 g) |
IDEXX Filta-Max® filtration EPA method1623.1 | Microscopy, conventional PCR, qPCR, restriction enzyme, sequencing | 529 bp REP | 40.0 | 1–180/g | Yes | No |
Abbreviations: na, not applicable; nd, not determined; REP, repetitive region
In our study, T. gondii DNA fragments were confirmed by sequencing and by DNA digestion with a restriction enzyme. Enzymatic digestion with EcoRV endonuclease suggests specificity and excludes cross-amplification of non-target organisms, such as Hammondia hammondi; this closely related apicomplexan presents a homologous region with around 84% sequence identity with the 529-bp repeat region of T. gondii [38]. The selected endonuclease EcoRV does not bind to any recognition site of the H. hammondi repetitive sequence, consequently, DNA cleavage does not occur.
Slides from PCR-positive samples were examined under epifluorescent microscopy and autofluorescent structures, compatible with T. gondii unsporulated oocysts in morphology and size (between 10–15 µm in diameter) were observed. A sporulated oocyst was observed in one sample. Data concerning the direct microscopic examination of oocysts in fresh vegetables are scarce [19, 21]. Indeed, no clear references to the observation of T. gondii sporulated oocysts has yet been described in fruit or vegetables. On the other hand, T. gondii oocysts are morphologically indistinguishable from the oocysts of related coccidian parasites, not infectious to humans, such as Hammondia spp. and Neospora caninum [13]; debris present in the samples may affect the ability to visualize small fluorescent structures such as autofluorescent oocysts, and so far there are no commercial antibodies to implement a specific immunomagnetic separation protocol. Therefore, and in the absence of standard methods for oocysts observation, T. gondii confirmation always requires molecular identification. Our findings suggest no significant seasonal variation or differences between the type of agriculture system applied. Additionally, no significant differences between the prevalence of T. gondii in the leafy vegetables or fruits were observed. Although a statistically significant difference in the proportion, has not been observed between bulk/RTE and positive T. gondii DNA amplification, curiously, a remarkably higher concentration of Toxoplasma oocysts was found in RTE and packaged fruit and vegetables, when compared to bulk products. So far, we have no clear explanation for this result. However, it is reasonable to speculate about the management of the washing systems used in food industry. Trevisan et al. [39] referred that managing water sustainably in agriculture means increased utilization of wastewater for irrigation, reusing freshwater to wash the products and therefore increasing the chances for RTE/packaged fruit and vegetables to become contaminated [39]. Nevertheless, testing water samples from washing/packing facilities and the water used for irrigation of bulk products are warranted in order to decipher the degree of Toxoplasma oocyst contamination on fruit and vegetables. In addition, genotyping should be carried out in all positive-PCR samples, for risk assessment evaluation.
Conclusions
Our findings sustain that consumption of raw fruit and vegetables may be a source of T. gondii infection in humans. They also emphasise the need of a procedure accepted with a consensus as the “gold standard” method for the recovery, detection and quantification of T. gondii oocysts and validated for use with fresh fruit and vegetables. Major improvements are still required for routine application at the industrial level or for food testing in laboratories for detection of T. gondii oocysts.
Acknowledgements
The authors would like to thank J. P. Dubey for kindly providing the T. gondii ME49 oocysts used in this work and Dr Margarida Saraiva for her critical advice regarding the conception and design of the experimental procedure and discussion of the results. The authors would also like to acknowledge SONAE MC SGPS, S.A., Vitacress Portugal S.A., and HORPOZIM association for providing some of the fruit and vegetables used in this study.
Abbreviations
- BLAST
basic local alignment search tool
- CI
confidence interval
- FA
fluorescence assay
- IMS
immunomagnetic separation
- MIQE
minimum information for publication of quantitative real-time PCR experiments
- PBS
phosphate-buffered solution
- PTG
putative T. gondii
- qPCR
quantitative PCR
- RE
restriction enzyme
- RTE
ready-to-eat
- SE
standard error
- Tm
melting temperature
Authors’ contributions
CSM, SS, AC and JMC participated in the design the experiments, the conception of the study, drafted and review the manuscript. CSM carried out the experiments and CSM and JMC analysed the data. All authors read and approved the final manuscript.
Funding
This study was supported by the Horizon 2020 project SafeConsume (Grant Agreement No. 727580).
Availability of data and materials
Data supporting the conclusions of this article are included within the article. The datasets used and analysed in the present study are available from the corresponding author upon reasonable request.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Cláudia S. Marques, Email: claudiamarques1978@gmail.com
Susana Sousa, Email: adelaide.sousa@insa.min-saude.pt.
António Castro, Email: antonio.castro@insa.min-saude.pt.
José Manuel Correia da Costa, Email: jose.costa@insa.min-saude.pt.
References
- 1.Dubey JP. Toxoplasmosis of animals and humans. 2. Boca Raton: CRC Press, Taylor & Francis Group; 2010. [Google Scholar]
- 2.Robert-Gangneux F, Dardé ML. Epidemiology of and diagnostic strategies for toxoplasmosis. Clin Microbiol Rev. 2012;25:264–296. doi: 10.1128/CMR.05013-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Montoya JG, Liesenfeld O. Toxoplasmosis. Lancet. 2004;363:1965–1976. doi: 10.1016/S0140-6736(04)16412-X. [DOI] [PubMed] [Google Scholar]
- 4.WHO. The burden of foodborne diseases in the WHO European Region. Copenhagen: WHO; 2017. http://www.euro.who.int/__data/assets/pdf_file/0005/402989/50607-WHO-Food-Safety-publicationV4_Web.pdf?ua=1. Accessed 17 Feb 2020.
- 5.Koutsoumanis K, Allende A, Alvarez-Ordóñez A, Bolton D, Bover-Cid S, Chemaly M, et al. Public health risks associated with food-borne parasites. EFSA J. 2018;16:e05495. doi: 10.2903/j.efsa.2018.5495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Demar M, Ajzenberg D, Maubon D, Djossou F, Panchoe D, Punwasi W, et al. Fatal outbreak of human toxoplasmosis along the Maroni river: epidemiological, clinical, and parasitological aspects. Clin Infect Dis. 2007;45:88–95. doi: 10.1086/521246. [DOI] [PubMed] [Google Scholar]
- 7.Demar M, Hommel D, Djossou F, Peneau C, Boukhari R, Louvel D, et al. Acute toxoplasmoses in immunocompetent patients hospitalized in an intensive care unit in French Guiana. Clin Microbiol Infect. 2012;18:221–231. doi: 10.1111/j.1469-0691.2011.03648.x. [DOI] [PubMed] [Google Scholar]
- 8.Pomares C, Devillard S, Holmes TH, Olariu TR, Press CJ, Ramirez R, et al. Genetic characterization of Toxoplasma gondii DNA samples isolated from humans living in north America: an unexpected high prevalence of atypical genotypes. J Infect Dis. 2018;218:1783–1791. doi: 10.1093/infdis/jiy375. [DOI] [PubMed] [Google Scholar]
- 9.Boothroyd JC, Grigg ME. Population biology of Toxoplasma gondii and its relevance to human infection: do different strains cause different disease? Curr Opin Microbiol. 2002;5:438–442. doi: 10.1016/S1369-5274(02)00349-1. [DOI] [PubMed] [Google Scholar]
- 10.Xiao J, Yolken RH. Strain hypothesis of Toxoplasma gondii infection on the outcome of human diseases. Acta Physiol. 2015;213:828–845. doi: 10.1111/apha.12458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Galal L, Hamidović A, Dardé ML, Mercier M. Diversity of Toxoplasma gondii strains at the global level and its determinants. Food Waterborne Parasitol. 2019;15:e00052. doi: 10.1016/j.fawpar.2019.e00052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Dumètre A, Dardé ML. How to detect Toxoplasma gondii oocysts in environmental samples? FEMS Immunol Med Microbiol. 2003;27:651–661. doi: 10.1016/S0168-6445(03)00071-8. [DOI] [PubMed] [Google Scholar]
- 13.Shapiro K, Bahia-Oliveira L, Dixon B, Dumètre A, de Wit LA, VanWormer E, et al. Environmental transmission of Toxoplasma gondii: oocysts in water, soil and food. Food Waterborne Parasitol. 2019;15:e00049. doi: 10.1016/j.fawpar.2019.e00049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Dubey JP. Comparative infectivity of oocysts and bradyzoites of Toxoplasma gondii for intermediate (mice) and definitive (cats) hosts. Vet Parasitol. 2006;140:69–75. doi: 10.1016/j.vetpar.2006.03.018. [DOI] [PubMed] [Google Scholar]
- 15.Dumètre A, Le Bras C, Baffet M, Meneceur P, Dubey JP, Derouin F, et al. Effects of ozone and ultraviolet radiation treatments on the infectivity of Toxoplasma gondii oocysts. Vet Parasitol. 2008;153:209–213. doi: 10.1016/j.vetpar.2008.02.004. [DOI] [PubMed] [Google Scholar]
- 16.Dumètre A, Dubey JP, Ferguson DJP, Bongrand P, Azas N, Puech PH. Mechanics of the Toxoplasma gondii oocyst wall. Proc Natl Acad Sci USA. 2013;110:11535–11540. doi: 10.1073/pnas.1308425110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Mirza Alizadeh A, Jazaeri S, Shemshadi B, Hashempour-Baltork F, Sarlak Z, Pilevar Z, et al. A review on inactivation methods of Toxoplasma gondii in foods. Pathog Glob Health. 2018;112:306–319. doi: 10.1080/20477724.2018.1514137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Freppel W, Ferguson DJP, Shapiro K, Dubey JP, Puech P-H, Dumètre A. Structure, composition, and roles of the Toxoplasma gondii oocyst and sporocyst walls. Cell Surf. 2019;5:e100016. doi: 10.1016/j.tcsw.2018.100016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Al-Megrin WAI. Prevalence intestinal parasites in leafy vegetables in Riyadh, Saudi Arabia. Int J Trop Med. 2010;5:20–23. doi: 10.3923/ijtmed.2010.20.23. [DOI] [Google Scholar]
- 20.Lass A, Pietkiewicz H, Szostakowska B, Myjak P. The first detection of Toxoplasma gondii DNA in environmental fruits and vegetables samples. Eur J Clin Microbiol Infect Dis. 2012;31:1101–1108. doi: 10.1007/s10096-011-1414-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Haq SU, Maqbool A, Khan UJ, Yasmin G, Sultana R. Parasitic contamination of vegetables eaten raw in Lahore. Pak J Zool. 2014;46:1303–1309. [Google Scholar]
- 22.Lalonde LF, Gajadhar AA. Detection of Cyclospora cayetanensis, Cryptosporidium spp., and Toxoplasma gondii on imported leafy green vegetables in Canadian survey. Food Waterborne Parasitol. 2016;2:8–14. doi: 10.1016/j.fawpar.2016.01.001. [DOI] [Google Scholar]
- 23.Marchioro AA, Tiyo BT, Colli CM, De Souza CZ, Garcia JL, Gomes ML, et al. First detection of Toxoplasma gondii DNA in the fresh leafs of vegetables in south America. Vector Borne Zoonotic Dis. 2016;16:624–626. doi: 10.1089/vbz.2015.1937. [DOI] [PubMed] [Google Scholar]
- 24.Caradonna T, Marangi M, Del Chierico F, Ferrari N, Reddel S, Bracaglia G, et al. Detection and prevalence of protozoan parasites in ready-to-eat packaged salads on sale in Italy. Food Microbiol. 2017;67:67–75. doi: 10.1016/j.fm.2017.06.006. [DOI] [PubMed] [Google Scholar]
- 25.Ferreira FP, Caldart ET, Freire RL, Mitsuka-Breganó R, de Freitas FM, Miura AC, et al. The effect of water source and soil supplementation on parasite contamination in organic vegetable gardens. Rev Bras Parasitol Vet. 2018;27:327–337. doi: 10.1590/S1984-296120180050. [DOI] [PubMed] [Google Scholar]
- 26.Slany M, Dziedzinska R, Babak V, Kralik P, Moravkova M, Slana I. Toxoplasma gondii in vegetables from fields and farm storage facilities in the Czech Republic. FEMS Microbiol Lett. 2019;366:fnz170. doi: 10.1093/femsle/fnz170. [DOI] [PubMed] [Google Scholar]
- 27.EPA. Method 1623.1: Cryptosporidium and Giardia in water by filtration/IMS/FA. Washington: United States Environmental Protection Agency; 2012. https://nepis.epa.gov/Exe/ZyPDF.cgi/P100J7G4.PDF?Dockey=P100J7G4.PDF. Accessed 17 Feb 2020.
- 28.Homan WL, Vercammen M, De Braekeleer J, Verschueren H. Identification of a 200- to 300-fold repetitive 529 bp DNA fragment in Toxoplasma gondii, and its use for diagnostic and quantitative. Int J Parasitol. 2000;30:69–75. doi: 10.1016/S0020-7519(99)00170-8. [DOI] [PubMed] [Google Scholar]
- 29.Wells B, Shaw H, Innocent G, Guido S, Hotchkiss E, Parigi M, et al. Molecular detection of Toxoplasma gondii in water samples from Scotland and a comparison between the 529 bp real-time PCR and ITS1 nested PCR. Water Res. 2015;87:175–181. doi: 10.1016/j.watres.2015.09.015. [DOI] [PubMed] [Google Scholar]
- 30.Lalle M, Possenti A, Dubey JP, Pozio E. Loop-mediated isothermal amplification-lateral-flow dipstick (LAMP-LFD) to detect Toxoplasma gondii oocyst in ready-to-eat salad. Food Microbiol. 2018;70:137–142. doi: 10.1016/j.fm.2017.10.001. [DOI] [PubMed] [Google Scholar]
- 31.Instituto Nacional de Estatística. Balança alimentar portuguesa—2012–2016. Lisboa-Portugal; 2017. https://www.ine.pt/xportal/xmain?xpid=INE&xpgid=ine_publicacoes&PUBLICACOESpub_boui=289818234&PUBLICACOESmodo=2&xlang=pt. Accessed 17 Feb 2020.
- 32.Bustin S, Huggett J. qPCR primer design revisited. Biomol Detect Quantif. 2017;14:19–28. doi: 10.1016/j.bdq.2017.11.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem. 2009;55:611. doi: 10.1373/clinchem.2008.112797. [DOI] [PubMed] [Google Scholar]
- 34.Lindquist HDA, Bennett JW, Hester JD, Ware MW, Dubey JP, Everson WV. Autofluorescence of Toxoplasma gondii and related coccidian oocysts. J Parasitol. 2003;89:865–867. doi: 10.1645/GE-3147RN. [DOI] [PubMed] [Google Scholar]
- 35.Agresti A, Coull BA. Approximate is better than “exact” for interval estimation of binomial proportions. Am Stat. 1998;52:119–126. [Google Scholar]
- 36.Dumètre A, Dardé ML. Immunomagnetic separation of Toxoplasma gondii oocysts using a monoclonal antibody directed against the oocyst wall. J Microbiol Methods. 2005;61:209–217. doi: 10.1016/j.mimet.2004.11.024. [DOI] [PubMed] [Google Scholar]
- 37.Moulin L, Richard F, Stefania S, Goulet M, Gosselin S, Gonçalves A, et al. Contribution of treated wastewater to the microbiological quality of Seine river in Paris. Water Res. 2010;44:5222–5231. doi: 10.1016/j.watres.2010.06.037. [DOI] [PubMed] [Google Scholar]
- 38.Schares G, Herrmann DC, Beckert A, Schares S, Hosseininejad M, Pantchev N, et al. Characterization of a repetitive DNA fragment in Hammondia hammondi and its utility for the specific differentiation of H. hammondi from Toxoplasma gondii by PCR. Mol Cell Probes. 2008;22:244–251. doi: 10.1016/j.mcp.2008.04.003. [DOI] [PubMed] [Google Scholar]
- 39.Trevisan C, Torgerson PR, Robertson LJ. Foodborne parasites in Europe: present status and future Trends. Trends Parasitol. 2019;35:695–703. doi: 10.1016/j.pt.2019.07.002. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Data supporting the conclusions of this article are included within the article. The datasets used and analysed in the present study are available from the corresponding author upon reasonable request.