Abstract
Objective Determine the prevalence of the major enteric pathogens in dairy and dairy beef calves with diarrhoea in Australia.
Design Cross‐sectional study.
Methods Faecal samples from 84 Australian dairy and dairy beef properties (597 samples) were screened for rotavirus and coronavirus using real‐time reverse transcription polymerase chain reaction, for Salmonella spp. using selective enrichment faecal culture, and for enterotoxigenic Escherichia coli (K99) and Cryptosporidium parvum using a commercial enzyme‐linked immunosorbent assay. A logistic regression with random effects model was used to compare prevalence of pathogens in dairy and dairy beef operations.
Results Enteric pathogens were isolated from 97.6% of outbreaks and 95.0% of samples. Rotavirus was the most common pathogen identified (477/597, 79.9%) followed by C. parvum (349/597, 58.5%), Salmonella spp. (142/597, 23.8%), coronavirus (129/597, 21.6%) and E. coli K99 (104/597, 17.4%). Multiple pathogens were identified on 96.4% of farms and from 71.0% of samples. Samples from dairy beef properties were more likely to have multiple pathogens than dairy properties (P < 0.05), whereas rotavirus and Salmonella spp. were more likely to be identified in samples collected from dairy beef than dairy properties (P < 0.05).
Conclusion Most outbreaks of calf diarrhoea in dairy and dairy beef operations involve multiple pathogens. Rotavirus and C. parvum were the most frequently identified pathogens across production systems. Salmonella spp. and rotavirus were more frequently identified in dairy beef operations.
Keywords: calves, dairy cattle, diarrhoea, enteric pathogens, prevalence
Abbreviations
- ELISA
enzyme‐linked immunosorbent assay
- qRT‐PCR
real‐time reverse transcription polymerase chain reaction
Infectious diarrhoea is the most significant cause of morbidity and mortality in neonatal dairy calves throughout the world. 1 Calves are at greatest risk of developing diarrhoea within the first month of life and the incidence of diarrhoea decreases with age. 2 , 3 Disease results from the interaction of a number of variables, including pathogen, animal (immunological and nutritional status), environmental and management factors. Significant costs arise from neonatal diarrhoea, including treatment, diagnostics, labour, veterinary intervention and decreased number of herd replacements. 4
There have been many prevalence studies worldwide to determine the major enteric pathogens associated with diarrhoea in dairy calves. 1 , 3 , 4 , 5 , 6 , 7 , 8 , 9 , 10 , 11 In Australia, there are only a few small‐scale studies of individual pathogens 12 , 13 , 14 , 15 and currently there are no published large‐scale surveys of the causative agents of neonatal diarrhoea across the Australian dairy industry.
The pathogens most commonly incriminated in neonatal calf scours include viral (rotavirus and coronavirus), protozoal (Cryptosporidium parvum) and bacterial pathogens (enterotoxigenic Escherichia coli K99 and Salmonella spp.). 4 , 7 , 9 The detection of these pathogens does not prove causality, as all of these pathogens have been identified in both diarrhoeic and normal calves. 7 , 8 , 16 , 17 Identification of the possible causative agent in outbreaks of diarrhoea is important because it allows targeted preventative measures, such as vaccination, and identification of possible risk factors or sources of infection.
The aim of this study was to identify the prevalence of rotavirus, coronavirus, C. parvum, E. coli K99 and Salmonella spp. in outbreaks of diarrhoea in dairy and dairy beef calves aged less than 6 weeks in the main dairy regions of Australia.
Materials and methods
Sample collection
Herd and sample selection. Between May 2007 and September 2008, faecal samples were collected from outbreaks of diarrhoea in dairy and dairy beef calves aged less than 6 weeks. Herds with a minimum of 100 milking cows or rearing a minimum of 15 calves per batch were included in the study. An outbreak of diarrhoea was defined as a minimum of 5% morbidity with calves exhibiting signs of systemic disease (e.g. poor appetite, dehydration, decreased mentation and reduced suckle reflex) and had pasty–watery faeces. Herds were selected on the basis of the client calling a veterinary practice seeking assistance with an outbreak combined with practices actively advising clients to contact them if an outbreak of diarrhoea occurred. Twelve veterinary practices from the six states of Australia (New South Wales, Queensland, South Australia, Tasmania, Victoria and Western Australia), with a large number of dairy herds serviced by the practice were instructed on sample selection, sampling technique, storage and transport protocols.
Practitioners were asked to collect 6–10 faecal samples from individual calves from each outbreak. Signalment data were collected for each calf sampled using a form designed for the purposes of this study. Multiple sampling dates were permitted for smaller and year‐round operations where the minimum number of samples could not be collected at one visit. Practitioners were requested to collect samples from calves early in the course of the disease prior to the onset of antimicrobial therapy. Approximately 25 mL of faecal material was collected from the rectum of calves by direct digital stimulation using a disposable latex glove. Samples were placed into sterile containers and refrigerated until shipping.
Samples were then transported on ice to the Livestock Veterinary Teaching and Research Unit, University of Sydney, Camden by a commercial courier. Faeces were stored at 4°C until the time of processing. The subsequent processing of samples is described in detail elsewhere. 18
Pathogen detection
Bacteria and protozoa. Salmonella spp. were detected using selective enrichment faecal cultures. A sterile Dacron swab was used to inoculate faecal material into 10 mL of mannitol selenite broth, which was incubated overnight at 37°C. The swab was then streaked onto xylose lactose deoxycholate agar and incubated overnight at 37°C. Suspect colonies were subcultured on to blood agar plates and incubated overnight at 37°C, then tested with a commercial salmonella latex agglutination kit (Oxoid Salmonella Latex Test, Oxoid Ltd, Cambridge, UK). Salmonella‐suspect colonies underwent biochemical tests (urease, triple sugar iron and O‐nitriphenyl‐β‐D‐galactopyranoside). Salmonella spp. isolates were then streaked onto nutrient agar slopes and submitted to the Institute of Medical and Veterinary Science, Salmonella Reference Laboratory (Adelaide, SA, Australia) for final confirmation and serotyping.
Faecal samples were screened for E. coli K99 and C. parvum using two commercial benchtop kits for enzyme‐linked immunosorbent assay (ELISA; Pourquier® ELISA Calves Diarrhoea and Pourquier® ELISA Cryptosporidium antigen, Institut Pourquier®, Montpellier, France). The ELISA testing was performed according to the manufacturer's instructions. Briefly, 50 µL of dilution buffer and then 50 µL of undiluted faeces were placed into an appropriate number of wells of a microplate coated with the appropriate antibody. The plate was held at room temperature (≈25°C) for 30 min and then washed manually with the supplied wash solution. A unique conjugate (one for each of the pathogens) was then added to separate wells for each sample and held at room temperature for 30 min. Following a final wash, TMB substrate was added to each well and held at room temperature for a further 10 min. A stop solution (0.5 mol/L H2SO4) was added and the absorbance read at 450 nm (Labsystems Multiscan Biochromatic, Labsystems, Basingstoke, UK). The absorbance was transformed, according to the manufacturer's recommendations, to calculate the sample to positive (S/P) ratios. Samples with an S/P ratio <7% were deemed to be positive in accordance with the manufacturer's recommendations.
Viruses. The presence of rotavirus and coronavirus was determined by real‐time reverse transcription polymerase chain reaction (qRT‐PCR) assays as described elsewhere. 18
Data analysis
Data were managed in a Microsoft Access 2003 database (Microsoft Corporation, Redmond, WA, USA) and descriptive statistics calculated using Microsoft Excel 2003, (Microsoft Corporation). Calves were stratified into four age groups for analysis: 0–4 days old, 5–14 days old, 15–21 days old and >21 days old. The age categories were selected based on the pathophysiology of some of the pathogens (i.e. E. coli K99) and the age groups that have been used by previous authors. 4 , 20 The age distributions of calves from dairy and dairy beef properties were compared using chi square (STATA, Statacorp, TX, USA). The majority of calves were in the 5–14‐day‐old age group (69.4% for dairy and 89.5% for dairy beef). The number of calves in the other age groups was unbalanced and sparse. Comparison of pathogen prevalence and number of enteropathogens in dairy and dairy beef operations in 5–14‐day‐old calves was performed using a mixed effects logistic regression model with farm entered as a random effect (STATA). All analyses used alpha = 0.05 for comparative purposes.
Results
A total of 597 samples (540 and 57 from dairy and dairy beef calves, respectively) were collected from 76 dairy and 8 dairy beef calf properties. Only one outbreak of diarrhoea was sampled on each properties. The median number of samples obtained per farm was 7, with an interquartile range of 3.3, a minimum of 1 and maximum of 25.
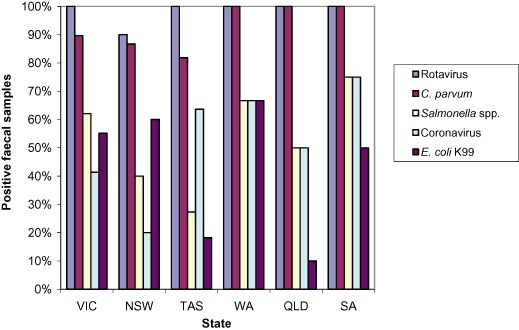
Of the samples collected, 67% were from Victoria and New South Wales. The breakdown of samples and properties by each of the states is shown in Table 1. Rotavirus was identified in at least one sample on each property in every state, except NSW where at least 90% of property samples had at least one sample positive for rotavirus. The percentage of properties with at least one positive sample for each of the pathogens is shown in Figure 1.
Table 1.
Number of properties and samples collected by Australian state
| State | No. of properties | No. of samples | Percentage of total samples |
|---|---|---|---|
| Victoria | 29 | 215 | 36.0 |
| New South Wales | 30 | 184 | 30.8 |
| Tasmania | 11 | 69 | 11.6 |
| Western Australia | 6 | 49 | 8.2 |
| Queensland | 4 | 59 | 9.9 |
| South Australia | 4 | 21 | 3.5 |
Figure 1.
Percentage of properties in each state with at least one positive sample for each of the enteropathogens.
The age of affected calves was recorded for 561 of the 597 samples (94.0%). The median age of the calves sampled was 10 days, with an interquartile range of 7 days, a minimum of 6 h and maximum of 42 days. The majority of calves sampled on both dairy and dairy beef properties were aged 5–14 days (Table 2). The difference in the distribution of ages between dairy and dairy beef sampled was statistically significant (P < 0.05).
Table 2.
Number and percentage of samples in each age category for dairy and dairy beef calves
| Age (days) | Total samples | Dairy samples | Dairy beef samples |
|---|---|---|---|
| 0–4 | 71 (12.6%) | 67 (13.3%) | 4 (7.0%) |
| 5–14 | 401 (71.5%) | 350 (69.4%) | 51 (89.5%) |
| 15–21 | 55 (9.8%) | 53 (10.5%) | 2 (3.5%) |
| >21 | 34 (6.0%) | 34 (6.7%) | 0 (0.0%) |
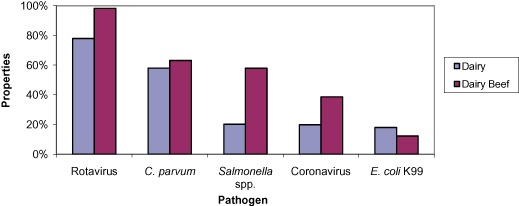
Enteric pathogens were isolated from 97.6% of outbreaks and 95.0% of samples. Rotavirus was the most common pathogen identified (477/597, 79.9%) followed by C. parvum (349/597, 58.5%), Salmonella spp. (142/597, 23.8%), coronavirus (129/597, 21.6%) and E. coli K99 (104/597, 17.4%). A breakdown of the pathogens identified in each system is shown in Figure 2 and Table 3.
Figure 2.
Percentage of samples (n = 597) positive for each enteropathogen tested at dairy and dairy beef farms.
Table 3.
Number of pathogens present in individual faecal samples and farms for each system
| No. of pathogens | Total samples (n = 597) | Dairy samples (n = 540) | Dairy beef samples (n =57) | Dairy farma (n = 76) | Dairy beef farm (n = 8) |
|---|---|---|---|---|---|
| 0 | 30 (5.0%) | 30 (5.6%) | 0 (0.0%) | 2 (2.6%) | 0 (0.0%) |
| 1 | 143 (24.0%) | 139 (25.7%) | 4 (7.0%) | 1 (1.3%) | 0 (0.0%) |
| 2 | 253 (42.4%) | 231(42.8%) | 22 (38.6%) | 11 (14.5%) | 1 (12.5%) |
| 3 | 133 (22.3%) | 115 (21.3%) | 18 (31.6%) | 30 (39.5%) | 1 (12.5%) |
| 4 | 37 (6.2%) | 24 (4.4%) | 13 (22.8%) | 27 (35.5%) | 3 (37.5%) |
| 5 | 1 (0.2%) | 1 (0.2%) | 0 (0.0%) | 5 (6.6%) | 3 (37.5%) |
Farms were analysed as a separate entity to the samples. Figures represent the number of different pathogens present on each farm i.e. a farm with two positive samples, each for a different organism, would be representative of a farm with two pathogens present.
Rotavirus was the most common pathogen identified in each of the four age groups of calves (Figure 3). Rotavirus and Salmonella spp. were more likely to be identified in samples collected from dairy beef than dairy properties (P < 0.05, odds ratio 15.8 and 5.4, respectively), but there was no statistically significant difference between dairy beef and dairy properties in the occurrence of C. parvum, coronavirus or E. coli K99 (P > 0.05) (Table 4).
Figure 3.
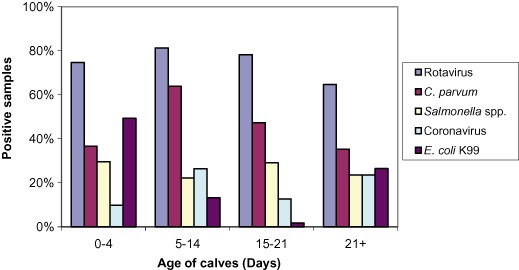
Percentage of samples (n = 561) positive for each enteropathogen by age group.
Table 4.
Logistic regression analysis of calves sampled in 5–14‐day‐old age category, using the odds ratios with dairy calves as the reference
| Variable | Odds ratio | Minimum | Maximum | P value |
|---|---|---|---|---|
| Salmonella spp. | 5.72 | 1.88 | 17.40 | 0.00 |
| Rotavirus | 27.56 | 1.88 | 404.57 | 0.02 |
| Coronavirus | 1.04 | 0.38 | 2.90 | 0.93 |
| Cryptosporidium parvum | 0.96 | 0.33 | 2.80 | 0.94 |
| Eschirichia coli K99 | 1.03 | 0.27 | 4.02 | 0.96 |
| 2 pathogens | 6.67 | 1.46 | 30.48 | 0.01 |
| 3 pathogens | 3.37 | 0.66 | 17.20 | 0.14 |
| 4 pathogens | 6.89 | 0.40 | 119.26 | 0.19 |
Multiple pathogens were identified on 96.4% of farms and from 71.0% of samples (Table 1). Samples from dairy beef properties were more likely to have multiple pathogens (i.e. ≥2 pathogens) than dairy properties (P < 0.05). The odds ratio for 2, 3 and 4 pathogens was 6.7, 3.4 and 6.9, respectively. Dual infections with rotavirus and C. parvum were the most common combined infection occurring in 148 of 597 calves (24.8%).
Discussion
We report the first large‐scale, Australia‐wide study of the prevalence of major enteric pathogens of neonatal diarrhoea in dairy and dairy beef calves. We have characterised the prevalence of the individual pathogens in outbreaks of diarrhoea and demonstrate that the majority of clinical cases are associated with simultaneous infection by multiple pathogens.
The objective of the survey was to determine the prevalence of enteric pathogens associated with outbreaks of diarrhoea. Protocols were designed to increase the likelihood of detecting the primary pathogens contributing to disease. For example, veterinarians were asked to select samples from calves that were showing signs of systemic illness with a duration of not more than 4 days. Isolation of causative agents is increased when animals are sampled early in the disease course and prior to the use of antibiotics that may lead to false negatives for bacterial pathogens. Histopathological evidence demonstrating pathology consistent with the enteropathogen isolated is desirable to establish a causal association and requests for postmortem samples from outbreaks were included in the instructions sent to practitioners. However, the number of postmortem samples collected was very small, possibly because of the extra time required to perform the procedures, the cost and time involved in revisiting a property to necropsy calves that died subsequent to the initial visit, the lack of suitable samples and the reluctance of farm managers to euthanase sick calves. Identification of an enteropathogen from the faeces of a calf does not constitute a definitive aetiological diagnosis. Sampling a group of affected calves provides more robust evidence when similar pathogens are identified in multiple animals.
The prevalence of rotavirus identified in this survey is greater than has been reported in the USA and Europe, 1 , 4 , 5 , 7 , 8 , 11 , 16 , 19 and contrasts with recent studies where the prevalence of C. parvum has been higher than that of rotavirus. 20 This discrepancy may in part reflect the limits of detection of the diagnostic assays use for pathogen detection. qRT‐PCR detection for rotavirus has been shown to be both highly sensitive and specific. 21 Other studies have used less sensitive diagnostic tests, such as ELISA and lateral flow immunochromatography, 1 , 4 , 5 so a greater prevalence is to be expected in the current study. An argument against the use of qRT‐PCR for detection of enteropathogens is that the low viral numbers that can be detected may not be significant. However, this statement is not valid because in most field situations the time of onset of diarrhoea is not known or detected, so the peak of viral shedding may have already passed by the time of sampling. When using qRT‐PCR the pathogen loads can be quantified and high loads identified. In addition, the significance of identifying low viral numbers in an individual sample can be increased by sampling multiple animals. The widespread nature of rotavirus was confirmed by the identification of at least one positive sample in almost all the properties sampled.
The prevalence of C. parvum in the faecal samples in this study is consistent or higher than in other studies that report prevalence ranging from 15% to 59%. 1 , 4 , 5 , 7 , 11 , 19 , 22 , 23 , 24 The prevalence found in each of those studies varies with regard to the target population and the diagnostic modality used, but in agreement with our current study, the peak time for identification of C. parvum in calves is from 1 to 3 weeks of age. 22 , 25 , 26 , 27 Similar to previous studies, 20 , 28 the presence of C. parvum in calves aged less than 4 days was notable, given the experimental and natural pre‐patent period of 2–7 days and 4–8 days, respectively. 29 , 30 Numerous overseas studies have reported the protozoa to be highly prevalent in dairy calves, with infection rates as high as 100% in some herds. 25 , 31 , 32 , 33 C. parvum was widespread and with more than 80% of properties sampled having at least one positive calf, the prevalence is comparable or higher than other herd prevalence data from overseas surveys. 34 , 35 , 36 The commercial ELISA kit used in this study has been evaluated in a recent investigation of four diagnostic techniques and was found to be sensitive (93.6%) and specific (95.9%) for detection of C. parvum in diarrhoeic calves. 37 Previous authors have noted that the use of ELISA for detecting C. parvum is sufficient to detect clinical cases where large amounts of oocysts are shed, but not appropriate for detection of light infections where only a few oocysts are shed. 38 Thus animals shedding small numbers of oocysts may not have been detected.
Salmonella spp. pose a significant animal and public health threat, 39 with human salmonellosis often linked to consumption of animal‐derived food products. 40 In this study, Salmonella spp. were isolated from 23.8% of individual samples and from 25–75% of properties in each state. The number of positive herds in this survey was less than that found in a large‐scale North American study of 110 dairy farms in which Salmonella spp. were isolated on 90% of farms. 41 The prevalence established in our survey is comparable to or greater than that reported in studies across Europe, America and New Zealand. 4 , 7 , 11 , 19 , 42 , 43 The higher incidence seen in our survey may have been related to the study design as it is probable that farmers were more likely to involve private practitioners in calf diarrhoea outbreaks if there was significant mortality. Outbreaks of disease caused by Salmonella spp. (and also with E. coli K99) often involve significant mortality compared with other enteric pathogens. 8 Previous studies that included healthy animals in the sample pool are likely to demonstrate a lower prevalence of Salmonella spp., as many authors have noted that intestinal carriage of Salmonella spp. is minimal in healthy calves. 44 , 45 , 46
Enterotoxigenic E. coli (K99) causes diarrhoea in young calves and can cause significant mortality as a result of marked dehydration and electrolyte losses. 47 The percentage of positive samples in this study is comparable to overseas prevalence studies. 1 , 4 , 5 , 7 , 19 , 42 Identification of E. coli K99 in faeces and its capacity to cause disease has been shown to be time dependent. 4 , 48 As expected, 50% of calves infected with E. coli K99 were less than 4 days old, which corresponds to the ability of the pilus antigens to bind to the immature enterocytes. 47 Other authors have identified a potential, prolonged isolation of E. coli K99 with concurrent C. parvum or rotavirus infection (≤14 days). 49 , 50 , 51 The surprising results in our study was the detection of E. coli K99 in a large proportion of calves aged more than 21 days old. We are unable to explain this finding from a pathophysiological standpoint. Perhaps it is related to inaccuracies in the diagnostic test or to the transient passage of coliforms in altered intestinal environments. According to the manufacturer of the ELISA kit used in this study, the sensitivity and specificity for the detection of E. coli K99 is 100% and 91.3%, respectively, when compared with PCR. We were unable to find external validation of the kit in the published literature.
Diarrhoea caused by coronavirus is typically more severe than rotaviral diarrhoea because the former also invades the large intestine. The prevalence of coronavirus observed in diarrhoeic calves in other countries and in the current study is lower than for rotavirus. However, the prevalence of coronavirus was, overall, relatively high when compared with European and North American surveys. 1 , 4 , 5 , 7 , 42
Identification of all five of the major calf scour enteropathogens for which the diagnostic tests were used in samples from every state emphasises the widespread dissemination of these organisms. Only 5% of samples did not have an identified pathogen.
Most of the prevalence surveys of enteropathogens in calves fail to detect pathogens in 10–30% of faecal samples, 1 , 4 , 7 , 8 , 19 which is an important source of frustration for clinicians trying to establish the causative agent in order to identify control methods. The higher pathogen detection achieved in this study likely reflects the use of the qRT‐PCR assay, which has a lower limit of detection than antigen detection assays. The emphasis placed on collecting samples from calves early in the disease course may have also facilitated pathogen detection. In a recent review of diarrhoeic calf submissions to diagnostic laboratories in Australia, no pathogen was isolated from 25–52% of samples, and the higher failure to detect any pathogen was mainly because of incomplete diagnostic testing, with only 17.2% of samples (37/215) tested for all the major calf scour pathogens. 52 Such results highlight the misunderstandings and errors in diagnosis that often occur in the identification of enteropathogens in the field in the absence of laboratory investigation. Underutilisation of diagnostic laboratories likely reflects their policy of full cost recovery.
The majority of samples in this study contained mixed infections, with two or more pathogens isolated. The percentage of calves with mixed infections (71.0%) was higher than in previous reports (5–28%). 1 , 4 , 7 , 8 , 16 Previous surveys comparing diarrhoeic and healthy calves have shown that mixed infections are more commonly detected in diarrhoeic animals, 7 , 53 although this has still yet to be proven in controlled experimental studies. 20 The significant number of multiple infections highlights the need for preventative measures to include generic as well as pathogen‐specific protection strategies.
Given the high incidences of rotavirus and Cryptosporidium, it was not surprising that these pathogens were the most common combined infection, a finding consistent with other studies. 1 , 4 , 5 , 10 , 11
Although the number of dairy beef properties sampled was relatively small, the number of calves raised in these facilities is proportionally higher than in most dairy operations. Dairy beef operations grow and finish dairy/dairy‐cross steers and heifers for the domestic and export beef market. The dairy beef property is very different from a dairy farm, with calves coming from multiple sources, thus providing opportunities for a diversity of pathogens to be introduced. This situation was reflected by the significantly greater proportion of dairy beef samples from which multiple pathogens were isolated. The logistic regression performed to investigate the pathogen profile of dairy and dairy beef operations was limited to the age group that accounted for nearly 90% of the dairy beef sample submissions (i.e. 5–14 days), because the data for the other age groups was sparse and unbalanced. It would have been ideal to have more animals in the different age groups so that age and system could be evaluated in the same model. The age distribution of calves in dairy beef operations is inherently slightly different to dairy operations because of the restrictions in Australia on the transport of calves aged less than 4 days.
There are a number of reasons for the increased probability of identifying rotavirus or Salmonella in dairy beef samples. Dairy bull calves are an unwanted byproduct of current dairy production systems, so are less likely to receive colostrum and thus passive transfer of immunity than the heifer calves that are retained as herd replacements. Co‐mingling of dairy beef calves during farm collection and transport also provides more opportunity for mixed pathogen exposure and stress may contribute to immunosuppression. Although rotavirus and Salmonella are prevalent on a significant number of farms, there are likely to be strain variations between farms. Thus, antibodies derived from maternal colostrum may only be specific for strains on a particular property. In order to maximise the number of animals that pass through dairy beef enterprises, there is a need for all available housing to be maximally stocked, which can lead to failure of appropriate disinfection between the removal of older calves and the introduction of young ones, resulting in enteric pathogens being passed between different groups of calves.
The results of this study provide valuable insight into the pathogens associated with outbreaks of diarrhoea in dairy calves across Australia. The prevalence of enteropathogens was comparable to or higher than the prevalence reported overseas. Rotavirus and C. parvum were the most prevalent enteropathogens identified in calves aged less than 6 weeks. Salmonella spp. and rotavirus were more likely to be isolated from diarrhoeic calves from dairy beef rather than dairy operations. Disease prevention and management is important from a calf and public health perspective, because of the zoonotic potential of Cryptosporidium and Salmonella spp.
Acknowledgments
Funding for the project was provided by Intervet/Schering‐Plough Animal Health. The study would not have been possible without the veterinary practices and their clients who provided the samples. The authors are indebted to Dr Xingnian Gu for expert assistance during the development of the coronavirus PCR assay and to Yogini Lele and the staff of the Virology Laboratory at EMAI for technical assistance during this study.
References
- 1. Lanz Uhde F, Kaufmann T, Sager H et al Prevalence of four enteropathogens in the faeces of young diarrhoeic dairy calves in Switzerland. Vet Rec 2008;163:362–366. [DOI] [PubMed] [Google Scholar]
- 2. Frank NA, Kaneene JB. Management risk factors associated with calf diarrhea in Michigan dairy herds. J Dairy Sci 1993;76:1313–1323. [DOI] [PubMed] [Google Scholar]
- 3. Garcia A, Ruiz‐Santa‐Quiteria JA, Orden JA et al Rotavirus and concurrent infections with other enteropathogens in neonatal diarrheic dairy calves in Spain. Comp Immunol Microbiol Infect Dis 2000;23:175–183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. De la Fuente R, Garcia A, Ruiz‐Santa‐Quiteria JA et al Proportional morbidity rates of enteropathogens among diarrheic dairy calves in central Spain. Prev Vet Med 1998;36:145–152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Gulliksen SM, Jor E, Lie KI et al Enteropathogens and risk factors for diarrhea in Norwegian dairy calves. J Dairy Sci 2009;92:5057–5066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Morgan JH, Grady EA, Pocock DH. A longitudinal study of enteric pathogens in dairy calves. In: Proceedings of a Society for Veterinary Epidemiology and Preventive Medicine meeting, Queens University, Belfast, 4–6 April 1990.
- 7. Reynolds DJ, Morgan JH, Chanter N et al Microbiology of calf diarrhoea in southern Britain. Vet Rec 1986;119:34–39. [DOI] [PubMed] [Google Scholar]
- 8. Snodgrass DR, Terzolo HR, Sherwood D et al Aetiology of diarrhoea in young calves. Vet Rec 1986;119:31–34. [DOI] [PubMed] [Google Scholar]
- 9. Waltner‐Toews D, Martin SW, Meek AH. An epidemiological study of selected calf pathogens on Holstein dairy farms in south‐western Ontario. Can J Vet Res 1986;50:307–313. [PMC free article] [PubMed] [Google Scholar]
- 10. Kaba J, Kita J, Piwowarczyk A, Pawinski J, Witkowski L. Epidemiology of neonatal calf diarrhea in large dairy herds. Med Wet 2006;62:665–668. [Google Scholar]
- 11. Langoni H, Linhares AC, de Avila FA, Da Silva AV, Elias AO. Contribution to the study of diarrhea etiology in neonate dairy calves in Sao Paulo state, Brazil. Braz J Vet Res Anim Sci 2004;41:313–319. [Google Scholar]
- 12. Becher KA, Robertson ID, Fraser DM, Palmer DG, Thompson RCA. Molecular epidemiology of Giardia and Cryptosporidium infections in dairy calves originating from three sources in Western Australia. Vet Parasitol 2004;123:1–9. [DOI] [PubMed] [Google Scholar]
- 13. Jerrett IV, Snodgrass DR. Cryptosporidia associated with outbreaks of neonatal calf diarrhoea. Aust Vet J 1981;57:434–435. [DOI] [PubMed] [Google Scholar]
- 14. Huang JA, Nagesha HS, Snodgrass DR, Holmes IH. Molecular and serological analyses of 2 bovine rotaviruses (B‐11 and B‐60) causing calf scours in Australia. J Clin Microbiol 1992;30:85–92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Swiatek DL, Palombo EA, Lee A et al Detection and analysis of bovine rotavirus strains circulating in Australian calves during 2004 and 2005. Vet Micribiol 2010;140:56–62. [DOI] [PubMed] [Google Scholar]
- 16. Sherwood D, Snodgrass DR, Lawson GHK. Prevalence of enterotoxigenic Escherichia coli in calves in Scotland and northern England. Vet Rec 1983;113:208–212. [DOI] [PubMed] [Google Scholar]
- 17. Chinsangaram J, Schore CE, Guterbock W, Weaver LD, Osburn BI. Prevalence of group‐A and group‐B rotaviruses in the feces of neonatal dairy calves from California. Comp Immunol Microbiol Infect Dis 1995;18:93–103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Izzo M, Kirkland P, Mohler V, Gunn A, House J. Comparison of three diagnostic techniques for detection of rotavirus and coronavirus in calf faeces in Australia. Aust Vet J (in press). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Gatti R, Nigrelli AD, Gelmetti D. Prevalence of cryptosporidia, rotavirus, E. coli K99 and Salmonella sp. in newborn calves: results of a diagnostic study in Mantova province. Atti Soc Ital Buiatria 1985;17. [Google Scholar]
- 20. De la Fuente R, Luzon M, Ruiz‐Santa‐Quiteria JA et al Cryptosporidium and concurrent infections with other major enteropathogens in 1 to 30‐day‐old diarrheic dairy calves in central Spain. Vet Parasitol 1999;80:179–185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Gutierrez‐Aguirre I, Steyer A, Boben J et al Sensitive detection of multiple rotavirus genotypes with a single reverse transcription‐real‐time quantitative PCR assay. J Clin Microbiol 2008;46:2547–2554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Castro‐Hermida JA, Gonzalez‐Losada YA, Ares‐Mazas E. Prevalence of and risk factors involved in the spread of neonatal bovine cryptosporidiosis in Galicia (NW Spain). Vet Parasitol 2002;106:1–10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Trotz‐Williams LA, Jarvie BD, Martin SW, Leslie KE, Peregrine AS. Prevalence of Cryptosporidium parvum infection in south‐western Ontario and its association with diarrhea in neonatal dairy calves. Can Vet J 2005;46:349–351. [PMC free article] [PubMed] [Google Scholar]
- 24. Olson ME, Guselle NJ, O'Handley RM et al Giardia and Cryptosporidium in dairy calves in British Columbia. Can Vet J 1997;38:703–706. [PMC free article] [PubMed] [Google Scholar]
- 25. Huetnik REC, Van der Giessen JWB, Noordhuizen JPTM. Epidemiology of Cryptosporidium spp. and Giardia duodenalis on a dairy farm. Vet Parasitol 2001;102:53–67. [DOI] [PubMed] [Google Scholar]
- 26. Nydam DV, Wade SE, Schaaf SL. Number of Cryptosporidium parvum oocysts or Giardia spp. cysts shed by dairy calves after natural infection. Am J Vet Med 2001;62:1612–1215. [DOI] [PubMed] [Google Scholar]
- 27. Barwick RS, Mohammed HO, White ME, Bryant RB. Prevalence of Giardia spp. and Cryptosporidium spp. on dairy farms in south‐eastern New York state. Prev Vet Med 2003;59:1–11. [DOI] [PubMed] [Google Scholar]
- 28. Fagan JG, Dwyer PJ, Quinlan JG. Factors that may affect the occurrence of enteropathogens in the faeces of diarrhoeic calves in Ireland. Ir Vet J 1995;48:17–21. [Google Scholar]
- 29. Markovics A, Pipano E. Shedding of cryptosporidial oocysts by naturally infected calves. Isr J Vet Med 1987;43:46–49. [Google Scholar]
- 30. Tzipori S, Smith M, Halpin R et al Experimental cryptosporidiosis in calves: Clinical manifestations and pathological finding. Vet Rec 1983;112:116–120. [DOI] [PubMed] [Google Scholar]
- 31. De Graaf DC, Vanopdenbosch E, Ortega‐Mora LM, Abbassi H, Peeters JE. A review of the importance of cryptosporidiosis in farm animals. Int J Parasitol 1999;29:1269–1287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. O'Handley RM, Cockwill C, McAllister TA et al Duration of naturally acquired giardiasis and cryptosporidiosis in dairy calves and their association with diarrhoea. J Am Vet Med 1999;214:391–396. [PubMed] [Google Scholar]
- 33. Olson ME, O'Handley RM, Ralston B, Thompson RCA. Emerging issues of Cryptosporidium and Giardia infections in cattle. Trends Parasitol 2004;20:185–191. [DOI] [PubMed] [Google Scholar]
- 34. Ruest N, Faubert GM, Couture Y. Prevalence and geographical distribution of Giardia spp. and Cryptosporidium spp. in dairy farms in Quebec. Can Vet J 1998;39:697–700. [PMC free article] [PubMed] [Google Scholar]
- 35. Brook E, Hart CA, French N, Christley R. Prevalence and risk factors for Cryptosporidium spp. infection in young calves. Vet Parasitol 2008;152:46–52. [DOI] [PubMed] [Google Scholar]
- 36. Hamnes IS, Gjerde B, Robertson L. Prevalence of Giardia and Cryptosporidium in dairy calves in three areas of Norway. Vet Parasitol 2006;140:204–216. [DOI] [PubMed] [Google Scholar]
- 37. Geurden T, Claerebout E, Vercruysse J, Berkvens D. A Bayesian evaluation of four immunological assays for the diagnosis of clinical cryptosporidiosis in calves. Vet J 2008;176:400–402. [DOI] [PubMed] [Google Scholar]
- 38. McCluskey BJ, Greiner EC, Donovan GA. Patterns of Cryptosporidium oocyst shedding in calves and a comparison of two diagnostic methods. Vet Parasitol 1995;60:185–190. [DOI] [PubMed] [Google Scholar]
- 39. Mead PS, Slutsker L, Dietz V et al Food‐related illness and death in the United States. Emerg Infect Dis 1999;5:607–625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Bopp CA, Brenner FW, Fields PI, Wells JG, Strockbine NA. Escherichia, Shigella and Salmonella In: Murray R, editor. Manual of clinical microbiology. 8th edn. ASM Press, Washington DC, 2003;654–671. [Google Scholar]
- 41. Fossler CP, Wells SJ, Kaneene JB et al Prevalence of Salmonella spp on conventional and organic dairy farms. J Am Vet Med 2004;225:567–573. [DOI] [PubMed] [Google Scholar]
- 42. Bulgin MS, Anderson BC, Ward ACS, Evermann JF. Infectious agents associated with neonatal calf disease in south‐western Idaho and eastern Oregon. J Am Vet Med 1982;180:1222–1226. [PubMed] [Google Scholar]
- 43. Grinberg A, Pomroy WE, Weston JF, Ayanegui‐Alcerreca A, Knight D. The occurrence of Cryptosporidium parvum, Campylobacter and Salmonella in newborn dairy calves in the Manawatu region of New Zealand. NZ Vet J 2005;53:315–320. [DOI] [PubMed] [Google Scholar]
- 44. Busato A, Hofer D, Lentze T, Gaillard C, Burnens A. Prevalence and infection risks of zoonotic enteropathogens in Swiss cow‐calf farms. Vet Microbiol 1999;69:251–263. [DOI] [PubMed] [Google Scholar]
- 45. Lance SE, Miller GY, Hancock DD, Bartlett PC, Heider LE. Salmonella infections in neonatal dairy calves. J Am Vet Med 1992;201:864–868. [PubMed] [Google Scholar]
- 46. Naciri M, Paul Lefay M, Mancassola R, Poirier P, Chermette R. Role of Cryptosporidium parvum as a pathogen in neonatal diarrhoea complex in suckling and dairy calves in France. Vet Parasitol 1999;85:245–257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Butler DG, Clarke RC. Diarrhoea and dysentery in calves In: Gyles CL, editor. Escherichia coli in domestic animals and humans. CAB International, Oxon, 1994;91–116. [Google Scholar]
- 48. Acres SD, Laing CJ, Saunders JR, Radostitis OM. Acute undifferentiated neonatal diarrhoea in beef calves. I: Occurrence and distribution of infectious agents. Can J Comp Med 1975;39:116–132. [PMC free article] [PubMed] [Google Scholar]
- 49. McDonough SP, Stull CL, Osburn BI. Enteric pathogens in intensively reared calves. Am J Vet Res 1994;55:1516–1520. [PubMed] [Google Scholar]
- 50. Runnels PL, Moon HW, Matthews PJ, Whipp SC, Woode GN. Effect of microbial and host variables on the interaction of rotavirus and Escherichia coli infections in gnobiotic calves. Am J Vet Res 1986;47:1542–1550. [PubMed] [Google Scholar]
- 51. Snodgrass DR, Smith ML, Krautil FL. Interaction of rotavirus and enterotoxigenic Escherichia coli in conventionally‐reared dairy calves. Vet Microbiol 1982;7:51–60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Gunn A. Calf scours in southern Australia Meat and Livestock Australia, 2003.
- 53. Morin M, Lariviee S, Lallier R et al Diarrhoea of newborn calves. II: Agents responsible for the disease on Quebec dairy farms. Med Vet Quebec 1980;10:60–65. [Google Scholar]


