Abstract
Doublecortin like kinase 1 (DCLK1) is a cancer stem cell marker for the colorectal cancer (CRC). It plays critical roles in the oncogenesis, progression and metastasis of CRC. DCLK1 can be an intriguing therapeutic target for CRC treatment. However, the molecular mechanism of how DCLK1 functions is unclear currently. In our research, we aim to apply RNA-Sequencing (RNA Seq) technology, a high throughput massively Next Generation Sequencing approach, to monitor transcriptome changes due to DCLK1 over-expression in the CRC cells. In order to achieve our goal, RNA from quadruplicate samples from two clones of isogenic DCLK1 stable over-expression cells and the parental wild type HCT116 cells was sent for RNA Seq on the Illumina NextSeq500 platform. Differentially expressed (DE) genes were evaluated by t-test (P <0.05 and fold-change ±1.5 or greater) using two methods: (1) FWER; and (2) Benjamani and Hochberg FDR (false discovery rate) which corrects for multiple comparisons. Gene networks and functional analysis were evaluated using Ingenuity Pathways Analysis (IPA). We identified 1463 DE genes common for both DCLK1 overexpression clone A and clone B cells. IPA results indicated that 72 canonical pathways were significantly modified by DCLK1 over-expression (P<0.05), among which 9 out of the top 10 pathways are involved in the cell cycle regulation, indicating that DCLK1 might play its tumorigenesis role via activation of pathways facilitating cell proliferation, repression of pathways inhibiting cells proliferation and function against pathways facilitating cell apoptosis. Cell cycle analysis results confirmed the IPA findings, which demonstrated that DCLK1 over-expression cells had much less G0/G1 cells but much more S and G2/M cells (P<0.05). In conclusion, DCLK1 over-expression significantly modified transcriptome profile of CRC cancer cells. Control of the cell cycle regulation might be one of the critical mechanism for DCLK1 function. Our findings provide more direct evidence for the development of DCLK1 as a therapeutic target for CRC treatment, and will be of great benefit for the discovery of novel therapeutic target within the DCLK1 molecular network for the treatment of colorectal cancer patients.
Background Introduction
Colorectal cancer (CRC) is the third most common cancer diagnosed and the second leading cause of cancer-related deaths in the United States (http://www.cdc.gov/cancer/colorectal/statistics/). Though more chemotherapeutic drugs have been developed, there were still 50% of patients relapsed after surgical and died of metastatic disease due to development of the multiple drug resistance 1. Effect of the same drug varies significantly among different patients since the heterogeneous properties of the CRC. The cancer stem cells (CSCs), which account for 0.05–1% of the tumor mass but can give rise to all of the cell types in the tumor 2, are believed to be able to survive after the chemotherapy. CSCs have been identified in several solid tumors, including CRC. So targeting CSCs for the treatment of CRC attracts a considerable interest. However, the biggest challenge is how to distinguish CSCs from the normal stem cells since they share most of the stem cell marker proteins. Several specific putative markers have been identified for the stem cell populations in the gastrointestinal tract, including doublecortin-like kinase 1 (DCLK1, also known as KIAA0369 3 or DCAMKL14).
DCLK1 was identified as a specific marker for CSCs in the intestine 5. Up-regulated expression of DCLK1 was found in several solid tumors of the digestive system, including CRC5–12, esophageal cancer13, 14, pancreatic cancer15–18, and liver cancer19–21. Up-regulation of DCLK1 facilitates colorectal tumorigenesis by enhancing pluripotency of the intestinal epithelial cells and epithelial mesenchymal transition (EMT) in the mice8, and over-expression of DCLK1 significantly increased the incidence of intestinal polyps compared to the normal control mice5. When DCLK1 expression was specifically knockdown, fewer polyps and decreased dysplasia were observed8 and when DCLK1+ cells were specifically targeted in the developed polyps, the cancer stem cells died and the established polyps were rapidly collapsed5. So DCLK1 plays critical roles in the oncogenesis, progression, invasion and metastasis of CRC. Several DCLK1 downstream targets have been identified, including EMT markers ZEB1, ZEB2, SNAIL, SLUG, pluripotency markers OCT4, SOX2, NANOG, KLF4, and pro-oncogene cMYC, Notch-1 and KRAS. DCLK1 may play its tumorigenesis and invasion role via modulation of microRNA pathways, in which EMT inhibitors miR-200a and let-7a, miR-145 and miR-144 were inhibited 15, 20, 22, 23. miR-137 12, miR-163 21 and miR-539 24 were demonstrated to be the negative regulator of DCLK1. However, how DCLK1 play its diverse functions inside the molecular network of CRC cells is not clearly understood.
In the current research, we applied the RNA-Sequencing (RNA Seq), which is a powerful high throughput approach using the next-generation sequencing (NGS) to reveal how DCLK1 modified the transcriptome of the CRC cells, aiming to establish the molecular network of DCLK1 inside the CRC cells. Our results demonstrated that DCLK1 changed the transcriptome of the CRC cells significantly, and multiple critical pathways were modified by DCLK1 overexpression, which indicates that DCLK1 may become a novel therapeutic target for the more effective treatment of CRC patients.
Materials and Methods
Cell lines and cell culture:
Human colorectal carcinoma cell line HCT116 cells were purchased from ATCC and were maintained in McCoy’s 5A medium (ATCC® 30–2007™) supplemented with 10% FBS in 37 °C incubator with 5% CO2. Isogenic DCLK1 over-expressed cells were established by transfecting human DCLK1 variant 1 cDNA, which is fused with a turboGFP gene at C-terminal (OriGene, Cat # RG217050) into HCT116 cells. In order to avoid the clonal variance, different clones were selected. Control HCT116 cells were established by transfecting pCMV6-AC-GFP Tagged Cloning Vector (Origene, Cat # PS100010) into HCT116 cells. Both DCLK1 over-expressed cells and control HCT116 cells were selected (400ug/ml) and maintained (250ug/ml) using Geneticin.
RNA isolation:
Quadruplicate snap frozen samples of control HCT116 cells and DCLK1 over-expressed cells from two clones were delivered to the Molecular and Genomics Core at University of Mississippi Medical Center. Total cellular RNA was extracted using Trizol® reagent (Invitrogen, Carlsbad, CA) and purified using PureLink™ RNA Mini Kit (Invitrogen, Carlsbad, CA). RNA concentrations were determined by Nano Drop. Quality and integrity of the extracted RNA were assessed using Bio-Rad Experion System. Samples were normalized to specific concentration and stored in −80°C for future use.
RNA sequencing (RNA-Seq):
RNA samples that pass quality parameters (minimum concentration and size range) were used to develop RNA libraries using the TruSeq Stranded Total RNA with Ribo-Zero Kit, Set A (FC-122–2501, San Diego, CA) according to manufacturer’s instructions. Each sample is prepared using 1 µg total RNA. The resulting cDNA libraries were quantified using the Qubit system (Invitrogen, Carlsbad, CA) and checked for quality and size using the Experion DNA 1K chip (BioRad, Hercules, CA). The fragment size generated library was ranging from 220 to 500 bps with a peak at ~250 bps. A portion of each library was diluted to 10 nM and stored at −20°C. 10 µL of 1.2–1.8 nM libraries was diluted and denatured. The libraries were sequenced using NextSeq500 High Output Kit (300 cycles- PE100) on Illumina NextSeq500 platform. The sequencing reads were automatically uploaded and evaluated for quality control using Illumina BaseSpace Onsite Computing platform.
Bioinformatics analysis:
Fastq sequence files generated from the Illumina NextSeq500 sequencer were used for preliminary analysis using applications available on BaseSpace Onsite/ Cloud Computing platform, including TopHat Alignment (read mapping to reference genome-USCS-hg19/ RGSC 6.0/rn6, GRCm38/mm10, etc.) and Cufflinks Assembly & DE (assembly of novel transcripts and differential expression). Additional analysis were performed using commercially available GeneSifter™ software platform (http://www.genesifter.net).
Ingenuity Pathways Analysis (IPA):
Gene networks and functional analysis were evaluated through the use of IPA software (Ingenuity® Systems, https://www.qiagenbioinformatics.com/products/ingenuity-pathway-analysis/). Briefly, significantly differentially expressed annotated genes common to both DCLK1 over-expressed clones, which demonstrated same changing direction, i.e., decrease or increase in both clones, were uploaded to IPA for core analysis. Top canonical pathways modified by DCLK1 over-expression, upstream regulators of DCLK1, etc. wereanalyzed.
Quantitative real time polymerase chain reaction [(q)RT-PCR]:
Total cellular RNA was isolated using the RNeasy Mini Kit (Qiagen, Valencia, CA) according to the manufacturer’s instructions from the HCT116 control cells and the DCLK1 over-expression cells. First strand cDNA was generated using the Reverse Transcription System (Promega, Madison, WI) according to the manufacturer’s instruction. (q)RT-PCR was carried out on the Mx3005P™ thermocycler from Stratagene using the 2x Brilliant II SYBR Green QPCR master mix (Agilent, Santa Clara, CA) according to the manufacturer’s instruction with some modifications. Briefly, 5 µL cDNA from reverse transcription PCR was added to a 25 µL reaction. Primers for human RPS4Y1, DDX3Y, KDM5D, RGS4, SYTL5, and BMP4 were listed in Table 1. Beta-actin was used as an internal control and 2-ΔΔCt 25 approach was applied to analyze the data.
Table 1.
Primers used for the (q)RT-PCR
| RPS4Y1 | Forward | CGCTACCCAGATCCTGTCAT |
| Reverse | GGTGATCACACCAACACGAC | |
| DDX3Y | Forward | CGGTTTTATCAGCCAGCAGT |
| Reverse | ACATGCAGTGCCAGCATCTA | |
| KDM5D | Forward | CCCACCACCCTCTTTCTGTA |
| Reverse | AGACCCAAGGGGAGAAAAGA | |
| RGS4 | Forward | TGAGCACCTTTCTGGCTTTT |
| Reverse | GGCAAGATTGCCAGAGAGAC | |
| SYTL5 | Forward | GCCCAAGGCCATATAGAG |
| Reverse | CCTTGGTCGGGTATGTTCA | |
| BMP4 | Forward | ACGGTGGGAAACTTTTGATG |
| Reverse | CGATCGGCTAATCCTGACAT | |
| Beta Actin | Forward | AAACTGGAACGGTGAAGGTG |
| Reverse | AGAGAAGTGGGGTGGCTTTT |
Fluorescence-activated cell sorting (FACS) and cell cycle analysis.
HCT116 cells and DCLK1 over-expression cells were plated at 2 × 10^6 cells per T75 flask and cultured in the 10%FBS McCoy5A medium for 24 hours. To synchronize cells at G0/G1 phase of cell cycle, the 10% FBS culture medium was replaced with serum-free medium. After serum starvation for 24 hours, the serum free medium was replaced with the McCoy5A medium containing 10% FBS and cells were cultured for 24 hours. Then cells were harvested and prepared for FACS and cell cycle analysis according to the BD online protocol with modifications (https://www.bdbiosciences.com/us/resources/s/detectionki67). Briefly, cells were fixed in 5 ml cold 70% - 80% ethanol, and incubated at −20°C for at least 2 hours. Then cells were washed 3X with 15mL staining buffer (1X PBS with 1% FBS, 0.09% sodium azide). After washing, cells were diluted in the staining buffer at 1X10^7 / mL. Transferred 1X10^6 cells (100uL) into according sample tubes, and cells were processed with no stain control, stained with ki-67 (Invitrogen# 17–5699-42 Anti-Hu Ki-67 Clone 20 Raj1, APC – eBioscience) only, ViaProbe (BD#555815 Via-Probe Cell Viability Solution (7-AAD)) only, or ki-67 and ViaProbe double stain. After 30 minutes incubation (unstained control and ViaProbe only samples on ice, and others at room temperature), cells were washed 2 times with staining buffer, then diluted in 500µL staining buffer, where 20uL ViaProbe was added to the ViaProbe and double stained samples. Incubate 10min at RT in the dark and proceeded to flow analysis on the BD LSRFortessa Cell Analyzer.
Statistical analysis:
Differentially expressed genes were evaluated by t-test [P<0.05 and fold-change ±1.5 or greater] using two methods: (1) FWER; and (2) Benjamani and Hochberg FDR (false discovery rate) which corrects for multiple comparisons.
Results
DCLK1 over-expression was confirmed in the isogenic DCLK1 over-expression clones, and two clones were selected (Clone_A and B, Supplemental Figure S1). Cells were plated as quadruplicate samples for RNA-Seq. Total cellular RNA were extracted with high quality from HCT116 wild type cells and the two DCLK1 over-expression cells (Supplemental Table S1). High quality cDNA libraries were achieved and used for RNA sequencing.
RNA Sequencing and transcripts assembly:
All 12 samples yielded high quality RNA sequencing data with reading length of 101bp per reads with Q score of %>=Q30 above 85% (Supplemental Figure S2). Number of reads ranged from 43,090,698 to 61,861,398 per sample, with an average of 52,476,048 reads per sample (Supplemental Table S2). An average of 50% of the Alignment distributed in the coding region, 30% in the untranslated region, 16% in the intron region and 4% in the intergenic region (Supplemental Figure S3).
Differentially expressed (DE) gene analysis:
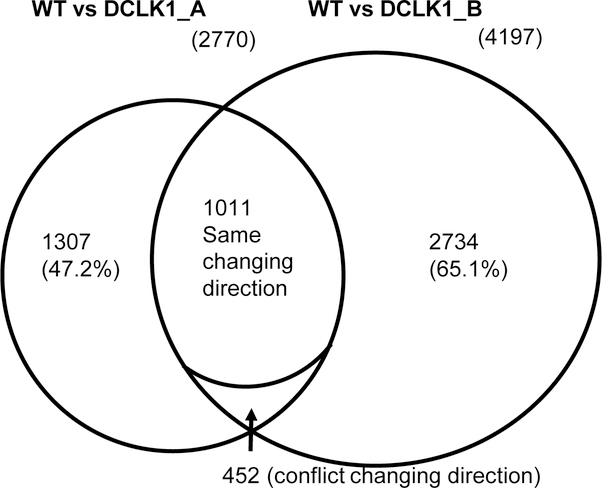
Cufflinks assembly and DE gene analysis identified 3497 significantly DE genes when compared genes identified in the DCLK1 over expressed clone A cells to those in the HCT116 wild type cells with log2(DCLK-A FPKM vs WT FPKM) ranging from −15.67 to 14.88 (6.68E-04<q<4.99E-02), among which 2770 are unique annotated genes. There are 5274 significantly DE genes when compared genes identified in the DCLK1 over expressed clone B cells to those in the HCT116 wild type cells with log2(DCLK-B FPKM vs WT FPKM) ranging from −15.65 to 8.62 (4.16E-04<q<5.00E-02), among which 4197 are unique annotated genes (Figure 1). Differential Expression Heatmap from Cufflinks Assembly & differential expression gene analysis demonstrated that most of the DE genes from quadruplicates of the same cell type clustered more closely compared to the other cell type (Figure 2). When compared the significantly DE genes from clone A and B, there are 1463 common genes that are unique annotated for both clones, among which 1011 genes demonstrated the same changing direction (Figure 1). The top 20 annotated genes common to both clones which were up-regulated due to DCLK1 over expression were shown in Table 2, and which were down-regulated were shown in Table 3. RPS4Y1, DDX3Y and KDM5D were selected as examples of the down-regulated genes, and RGS4, SYTL5, and BMP4 as the up-regulated genes for confirmation using (q)RT-PCR. All of the genes examined were significantly differently expressed in the DCLK1 over-expression cells compared to the wild type cells (P<0.05), which confirmed the RNA-Seq findings (Figure 3).
Figure 1. Differentially expressed(DE) unique annotated genes identified in the isogeneic DCLK1 over-expression cells compared to the HCT116 wild type cells.
Numbers of unique annotated DE genes to each DCLK1 over-expression clone, common DE genes to both clones, and common genes with different expression pattern were identified.
Figure 2. Differential Expression Heatmap from Cufflinks Assembly & differential expression gene analysis.
A. HCT116 wild type cells (WT) compared to the DCLK1 over expression clone_A. B. WT cells compared to the DCLK1 over expression clone_B.
Table 2.
Top 20 differentially expressed genes down-regulated in both DCLK1 over expressed Clone A and B compared to control cells (WT)
| Gene Symbol | Entrez Gene Name | Expr Log Ratio_AvsWT | Expr Log Ratio_BvsWT |
|---|---|---|---|
| RPS4Y1 | ribosomal protein S4, Y-linked 1 | −15.67 | −15.65 |
| DDX3Y | DEAD-box helicase 3, Y-linked | −11.58 | −11.56 |
| KDM5D | lysine demethylase 5D | −9.71 | −9.69 |
| USP9Y | ubiquitin specific peptidase 9, Y-linked | −9.53 | −9.51 |
| UTY | ubiquitously transcribed tetratricopeptide repeat containing, Y-linked | −9.04 | −9.02 |
| BEX4 | brain expressed X-linked 4 | −4.72 | −5.3 |
| MECOM | MDS1 and EVI1 complex locus | −4.24 | −1.56 |
| TNC | tenascin C | −3.98 | −2.24 |
| DAPP1 | dual adaptor of phosphotyrosine and 3-phosphoinositides 1 | −3.35 | −2.17 |
| OLAH | oleoyl-ACP hydrolase | −3.12 | −2.29 |
| CALB2 | calbindin 2 | −3.07 | −0.52 |
| TUSC3 | tumor suppressor candidate 3 | −2.81 | −5.25 |
| FXYD3 | FXYD domain containing ion transport regulator 3 | −2.78 | −0.8 |
| MAL2 | mal, T cell differentiation protein 2 (gene/pseudogene) | −2.74 | −1.18 |
| KLK5 | kallikrein related peptidase 5 | −2.7 | −1.46 |
| FXYD4 | FXYD domain containing ion transport regulator 4 | −2.56 | −1.87 |
| ATP2C2 | ATPase secretory pathway Ca2+ transporting 2 | −2.48 | −1.55 |
| KLK11 | kallikrein related peptidase 11 | −2.48 | −2.05 |
| HEPH | hephaestin | −2.43 | −2.52 |
Table 3.
Top 20 differentially expressed genes up-regulated in both DCLK1 over expressed Clone A and B compared to control cells (WT)
| Gene Symbol | Entrez Gene Name | Expr Log Ratio_AvsWT | Expr Log Ratio_BvsWT |
|---|---|---|---|
| RGS4 | regulator of G protein signaling 4 | 3.99 | 2.01 |
| SYTL5 | synaptotagmin like 5 | 3.03 | 1.85 |
| FGF9 | fibroblast growth factor 9 | 2.76 | 3.06 |
| CPA4 | carboxypeptidase A4 | 2.55 | 0.47 |
| BMP4 | bone morphogenetic protein 4 | 2.51 | 1.48 |
| HTRA1 | HtrA serine peptidase 1 | 2.46 | 1.2 |
| VSNL1 | visinin like 1 | 2.2 | 1.77 |
| TNIK | TRAF2 and NCK interacting kinase | 2.14 | 0.97 |
| COL24A1 | collagen type XXIV alpha 1 chain | 2.02 | 1.53 |
| METTL7A | methyltransferase like 7A | 1.85 | 1.36 |
| PDLIM5 | PDZ and LIM domain 5 | 1.77 | 0.5 |
| RND2 | Rho family GTPase 2 | 1.73 | 0.91 |
| GRIK4 | glutamate ionotropic receptor kainate type subunit 4 | 1.71 | 1.44 |
| IQGAP2 | IQ motif containing GTPase activating protein 2 | 1.61 | 1.04 |
| TESC | tescalcin | 1.55 | 1.25 |
| CXCR4 | C-X-C motif chemokine receptor 4 | 1.54 | 1.19 |
| SPRY2 | sprouty RTK signaling antagonist 2 | 1.54 | 0.55 |
| GSTA1 | glutathione S-transferase alpha 1 | 1.52 | 2 |
| PLEKHS1 | pleckstrin homology domain containing S1 | 1.46 | 1.19 |
| DDIT4L | DNA damage inducible transcript 4 like | 1.39 | 0.59 |
Figure 3. qRT-PCR validation of exampled differentially expressed(DE) genes identified by RNA-Seq.
3A. DE genes which are down-regulated. 3B. DE genes which are up-regulated. Expression of Genes of interest were normalized to GAPDH before comparing to the wild type (WT) cells. Data was expressed as mean±SEM from 3 independent experiments. All of the genes expressed in the DCLK1_Clone_A (DCLK1_A) and DCLK_Clone_B (DCLK1_B) were significant different compared to the WT cells (P<0.05, two-sided student t test). NoCt: no expression can be detected.
Modification of canonical pathways due to DCLK1 over-expression:
IPA analysis identified 72 canonical pathways which were significantly modified by DCLK1 over-expression (P<0.05). According to the Z score, except for the pathways which cannot predict the activity according to the knowledge database, 13 pathways were inhibited and 18 were activated. The most intriguing pathways which are modified by DCLK1 are those which play critical roles in regulating cell proliferation (Figure 4), which actively interact with each other to regulate cell proliferation (Figure 5). Several pathways which involves in enhancing cell proliferation are activated, including the cyclins and cell cycle regulation pathway, the mitotic roles of polo-like kinase pathway, the estrogen-mediated S-phase entry pathway, etc., and some pathways which involves in repressing cell proliferation are inhibited, such as the cell cycle: G2/M DNA damage checkpoint regulation pathway and the role of CHK proteins in cell cycle checkpoint control pathway. Expression of multiple genes involved in the regulation of the same pathway demonstrated coordinated pattern. Using genes function in the cyclins and cell cycle regulation pathway as an example, multiple cyclins, cyclin dependent kinases (CDK), CDK inhibitors, and the key cell cycle regulators E2F, which are involved in promoting cell proliferation were up-regulated significantly, and HDAC9, which involved in the inhibition of cell cycle was significantly repressed (Figure 6). Genes in the other pathways demonstrated similar pattern. So it’s most likely that through activating pathways facilitating cell proliferation and inhibiting pathways arresting cell proliferation, DCLK1 modifies the cell proliferation rate and results in tumorigenesis of CRC. Our cell cycle analysis results confirmed the scenario (Figure 7). DCLK1 over-expression cells demonstrated significantly much less cells in the G0/G1 phase, but much more cells in the S and G2 phase compared to the control HCT116 cells(P<0.01).
Figure 4. Top ten canonical pathways modified by DCLK1 over expression.
1011 differentially expressed genes which were common to both of the two DCLK1 over-expression clones and demonstrated consistent changing were used for Ingenuity Pathways Analysis.
Figure 5.
Overlapping canonical pathways modified by DCLK1 over-expression which play critical roles in the regulation of cell proliferation.
Figure 6. Heatmap of genes involved in the cyclins and cell cycle regulation pathway.
CCN: cyclin; CDK: cyclin dependant kinase; CDKN: cyclin dependant kinase inhibitor; E2F: E2F transcription factor; HDAC: histone deacetylase
Figure 7. Cell cycle analysis of HCT116 wild type (WT) and DCLK1 isogeneic cells using FACS.
7A: WT cells; 7B: DCLK1 Clone_A cells; and 7C: DCLK1 Clone_B cells. 7D: Quantitation of cell cycle analysis from 7A, B and C.
Conclusion and Discussion
Our results demonstrated that DCLK1 over-expression significantly modified transcriptome profile of CRC. 1463 genes common to both DCLK1 over-expression Clone_A and Clone_B were identified to be significantly differentially expressed, and 72 canonical pathways were significantly modified by DCLK1 over-expression. Control of multiply critical biological pathways which are involved in the regulation of cell proliferation might be one of the critical underlying molecular mechanisms for DCLK1 to play its function during oncogenesis, progression, invasion and metastasis of CRC. Moreover, our previous findings also demonstrated that DCLK1 significantly increased the chemoresistance of CRC cells to 5-Fu treatment 26. Therefore, DCLK1 can be developed as a novel therapeutic target for the treatment of colorectal cancer.
Growth out of control is one of the most basic properties for all cancer cells. Numerous oncogenic genes play their roles through manipulation of the cell cycle regulation, i.e., promoting cell proliferation without cell cycle check point regulation. Correlation of DCLK1 and cell cycle regulation has not been well understood. Deng et al 24 identified that miR-539, a tumor suppressor microRNA, promotes cell cycle arrest in G0/G1 phase by directly inhibition of DCLK1 in non-small cell lung cancer (NSCLC). With DCLK1 over-expressed, the cell cycle arrest effect of miR-539 was cancelled. When LRRK2-IN-1, a DCLK1 kinase inhibitor, was applied to the colorectal cancer cells and pancreatic cancer cells, cells were induced to be arrested in the S, G1 and G2/M phase, indicating that DCLK1 might be involved in promoting cell proliferation by regulating multiple check points during the cell cycle regulation 27. Our pathway analysis demonstrated that DCLK1 over-expression significantly activated several pathways which involves in enhancing cell proliferation, including the cyclins and cell cycle regulation pathway, the ATM signaling pathway, the mitotic roles of polo-like kinase pathway, the Aryl Hydrocarbon Receptor Signaling pathway, the estrogen-mediated S-phase entry pathway, etc., and some pathways which involves in repressing cell proliferation are highly inhibited, such as the cell cycle: G2/M DNA damage checkpoint regulation pathway and the role of CHK proteins in cell cycle checkpoint control pathway.
Cyclins and cyclin-dependent kinases, together with tumor suppressors P53 and Rb accurately and precisely control cell proliferation under normal conditions. However, when balance among these control factors inside the cell is disturbed and favors out of control growth of the cell, tumorigenesis will be the inevitable consequence, and multiple chemotherapeutic drugs have been developed which through inhibition of the CDKs activity to induce cancer cell apoptosis 28, 29. ATM is a multiple functional key factors during oncogenesis of multiple tumors. It regulates cell cycle checkpoints and its activation is required for DNA double strand break repair 30, and it is correlated with increased chemoresistance and radioresistance 31. The polo-like kinase is required for proper mitosis procedure inside the cell. Without normal functional polo-like kinase, the chromosomes cannot align up on the metaphase plate and the mitotic spindle cannot be properly formed, so mitosis cannot happen. Polo-like kinase is up-regulated in multiple cancers, including colorectal cancer 32 and it increases chemoresistance of colorectal cancer cells to oxaliplatin 33. Aryl hydrocarbon receptor (AhR) is a cytosolic, ligand-activated receptor and it plays crucial roles in maintain cellular homeostasis. By binding to different ligands, it functions in the oncogenesis, progression and metastasis of multiple cancers, including colorectal cancer, and multiple chemotherapeutic drugs targeting AhR have been developed 34. Suppression of immune recognition of cancer cells might be one of the major molecular mechanism applied by AhR 35. Estrogen belongs to the steroid hormone family and by binding to the estrogen receptor-α (ERα), it up-regulates expression of Myc and cyclin D1, which in turn activate cycline E and CDK2 complex and facilitating transition from G1 to S phase. Normally estrogen does not induce DNA damage, but it does required for cell prolifereation 36. Correlation between DCLK1 and all of the above pathways was not revealed previously, except that it was reported in the tuft cells, ATM activation in responding to DNA damage is DCLK1 dependent 37. Our pathway analysis using the RNA Seq data first clearly identified that DCLK1 can activate the above mentioned critical pathways, which may contribute to the initiation, progression, invasion and metastasis, and chemoresistance of CRC.
Moreover, G2/M DNA damage checkpoint regulation pathway and the role of CHK proteins in cell cycle checkpoint control pathway were significantly repressed by DCLK1. Checkpoints during cell cycle allow DNA repair and prevent cells with damaged DNA to proliferate. They are pivotal for the induction of cell apoptosis and prevention of oncogenesis 38. CHK proteins, which are checkpoint kinases, are responsible to keep the integrity of cellular DNA. More important for the p53-defficient cells, which can pass the G1/S checkpoint, CHK proteins can induce cell cycle arrest at S and G2/M phase 39. With defect checkpoint regulation and CHK proteins, cells will be at much higher risk for tumor formation 40. Our results demonstrated that with DCLK1 over-expression, the critical cell checkpoint responding to DNA damage is repressed. The CRC cells can go through cell proliferation without efficient checkpoint regulation, which benefit tumor growth and also survival of CRC cells after chemotherapy treatment.
In conclusion, DCLK1 over-expression significantly changes the transcriptome profile of human CRC cells. Multiple canonical pathways are modified, which favor tumorigenesis, progression, invasion and metastasis, and chemoresistance of CRC. RNA-Seq is a powerful high throughput approaches and when combined with diverse bioinformatics tool, it can be successfully applied to reveal whole genome profile of cells. Currently, we only confirmed that with over-expression of DCLK1, the cell cycle was modified significantly. In the future experiment, in order to further confirm function of DCLK1 in the regulation of cell cycle, cells with transient knockdown of DCLK1 expression or isogenic cells with DCLK1 knockout will be used to determine the cell proliferation rate compared to the wild type cells, and cell cycle analysis will be applied to see whether without DCLK1 expression, distribution of the cells in the cell cycle will be affected. RNA Seq technology will be applied to determine the transcriptome change with knockdown/knockout of DCLK1 gene expression. With this data, it will be more convincible about functions of DCLK1 in the regulation of multiple critical pathways. Combined with its role in the chemoresistance, DCLK1 can be development into an intriguing therapeutic target for human CRC treatment.
Supplementary Material
Acknowledgement
This work was supported by the Mississippi INBRE, funded by an Institutional Development Award (IDeA) from the National Institute of General Medical Sciences of the National Institutes of Health under grant number P20GM103476. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of General Medical Sciences or the National Institutes of Health.
The work performed through the UMMC Molecular and Genomics Facility is supported, in part, by funds from the NIGMS, including Mississippi INBRE (P20GM103476), Center for Psychiatric Neuroscience (CPN)-COBRE (P30GM103328), Obesity, Cardiorenal and Metabolic Diseases- COBRE (P20GM104357), and Mississippi Center of Excellence in Perinatal Research (MS-CEPR)-COBRE (P20GM121334). The content of the manuscript is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Special acknowledgments to Dr. Jonathan Lindner at University of Southern Mississippi for his great help with the FACS, Dr. Sweta Khanal and Dr. Alex Flynt at University of Southern Mississippi for his great help with the pathway heatmap establishment.
Footnotes
Conflict of Interests: The authors have nothing to disclose.
References
- 1.De Dosso S, Sessa C, Saletti P. Adjuvant therapy for colon cancer: present and perspectives. Cancer Treat Rev 2009; 35(2): 160–6. [DOI] [PubMed] [Google Scholar]
- 2.Yamakage K, Omori Y, Piccoli C, Yamasaki H. Growth control of 3T3 fibroblast cell lines established from connexin 43-deficient mice. Mol Carcinog 1998; 23(2): 121–8. [DOI] [PubMed] [Google Scholar]
- 3.Omori Y, Suzuki M, Ozaki K, Harada Y, Nakamura Y, Takahashi E et al. Expression and chromosomal localization of KIAA0369, a putative kinase structurally related to Doublecortin. J Hum Genet 1998; 43(3): 169–77. [DOI] [PubMed] [Google Scholar]
- 4.May R, Riehl TE, Hunt C, Sureban SM, Anant S, Houchen CW. Identification of a novel putative gastrointestinal stem cell and adenoma stem cell marker, doublecortin and CaM kinase-like-1, following radiation injury and in adenomatous polyposis coli/multiple intestinal neoplasia mice. Stem Cells 2008; 26(3): 630–7. [DOI] [PubMed] [Google Scholar]
- 5.Nakanishi Y, Seno H, Fukuoka A, Ueo T, Yamaga Y, Maruno T et al. Dclk1 distinguishes between tumor and normal stem cells in the intestine. Nat Genet 2013; 45(1): 98–103. [DOI] [PubMed] [Google Scholar]
- 6.Gagliardi G, Goswami M, Passera R, Bellows CF. DCLK1 immunoreactivity in colorectal neoplasia. Clinical and experimental gastroenterology 2012; 5: 35–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Arruda-Olson AM, Bursi F, Gerber Y, May RH, Roger VL, Pellikka PA. Three-dimensional echocardiography for evaluating left ventricular function in patients with ST elevation myocardial infarction: a pilot study. Mayo Clin Proc 2008; 83(3): 372–3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chandrakesan P, Weygant N, May R, Qu D, Chinthalapally HR, Sureban SM et al. DCLK1 facilitates intestinal tumor growth via enhancing pluripotency and epithelial mesenchymal transition. Oncotarget 2014; 5(19): 9269–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Westphalen CB, Asfaha S, Hayakawa Y, Takemoto Y, Lukin DJ, Nuber AH et al. Long-lived intestinal tuft cells serve as colon cancer-initiating cells. J Clin Invest 2014; 124(3): 1283–95. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kantara C, O’Connell MR, Luthra G, Gajjar A, Sarkar S, Ullrich RL et al. Methods for detecting circulating cancer stem cells (CCSCs) as a novel approach for diagnosis of colon cancer relapse/metastasis. Lab Invest 2015; 95(1): 100–12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Mirzaei A, Tavoosidana G, Modarressi MH, Rad AA, Fazeli MS, Shirkoohi R et al. Upregulation of circulating cancer stem cell marker, DCLK1 but not Lgr5, in chemoradiotherapy-treated colorectal cancer patients. Tumour Biol 2015; 36(6): 4801–10. [DOI] [PubMed] [Google Scholar]
- 12.Sakaguchi M, Hisamori S, Oshima N, Sato F, Shimono Y, Sakai Y. miR-137 Regulates the Tumorigenicity of Colon Cancer Stem Cells through the Inhibition of DCLK1. Mol Cancer Res 2016; 14(4): 354–62. [DOI] [PubMed] [Google Scholar]
- 13.Vega KJ, May R, Sureban SM, Lightfoot SA, Qu D, Reed A et al. Identification of the putative intestinal stem cell marker doublecortin and CaM kinase-like-1 in Barrett’s esophagus and esophageal adenocarcinoma. J Gastroenterol Hepatol 2012; 27(4): 773–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Whorton J, Sureban SM, May R, Qu D, Lightfoot SA, Madhoun M et al. DCLK1 is detectable in plasma of patients with Barrett’s esophagus and esophageal adenocarcinoma. Dig Dis Sci 2015; 60(2): 509–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Sureban SM, May R, Lightfoot SA, Hoskins AB, Lerner M, Brackett DJ et al. DCAMKL-1 regulates epithelial-mesenchymal transition in human pancreatic cells through a miR-200a-dependent mechanism. Cancer Res 2011; 71(6): 2328–38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Bailey JM, Alsina J, Rasheed ZA, McAllister FM, Fu YY, Plentz R et al. DCLK1 marks a morphologically distinct subpopulation of cells with stem cell properties in preinvasive pancreatic cancer. Gastroenterology 2014; 146(1): 245–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Rao CV, Mohammed A. New insights into pancreatic cancer stem cells. World J Stem Cells 2015; 7(3): 547–55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Qu D, Johnson J, Chandrakesan P, Weygant N, May R, Aiello N et al. Doublecortin-like kinase 1 is elevated serologically in pancreatic ductal adenocarcinoma and widely expressed on circulating tumor cells. PLoS One 2015; 10(2): e0118933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ali N, Allam H, Bader T, May R, Basalingappa KM, Berry WL et al. Fluvastatin interferes with hepatitis C virus replication via microtubule bundling and a doublecortin-like kinase-mediated mechanism. PloS one 2013; 8(11): e80304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sureban SM, Madhoun MF, May R, Qu D, Ali N, Fazili J et al. Plasma DCLK1 is a marker of hepatocellular carcinoma (HCC): Targeting DCLK1 prevents HCC tumor xenograft growth via a microRNA-dependent mechanism. Oncotarget 2015; 6(35): 37200–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wang W, Zhang H, Wang L, Zhang S, Tang M. miR-613 inhibits the growth and invasiveness of human hepatocellular carcinoma via targeting DCLK1. Biochem Biophys Res Commun 2016; 473(4): 987–92. [DOI] [PubMed] [Google Scholar]
- 22.Sureban SM, May R, Mondalek FG, Qu D, Ponnurangam S, Pantazis P et al. Nanoparticle-based delivery of siDCAMKL-1 increases microRNA-144 and inhibits colorectal cancer tumor growth via a Notch-1 dependent mechanism. J Nanobiotechnology 2011; 9: 40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Sureban SM, May R, Qu D, Weygant N, Chandrakesan P, Ali N et al. DCLK1 regulates pluripotency and angiogenic factors via microRNA-dependent mechanisms in pancreatic cancer. PLoS One 2013; 8(9): e73940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Deng H, Qianqian G, Ting J, Aimin Y. miR-539 enhances chemosensitivity to cisplatin in non-small cell lung cancer by targeting DCLK1. Biomed Pharmacother 2018; 106: 1072–1081. [DOI] [PubMed] [Google Scholar]
- 25.Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001; 25(4): 402–8. [DOI] [PubMed] [Google Scholar]
- 26.Li L, Jones K, Mei H. Doublecotin-Like Kinase 1 Increases Chemoresistance of Colorectal Cancer Cells through the Anti-Apoptosis Pathway. J Stem Cell Res Ther 2019; 9(3). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Weygant N, Qu D, Berry WL, May R, Chandrakesan P, Owen DB et al. Small molecule kinase inhibitor LRRK2-IN-1 demonstrates potent activity against colorectal and pancreatic cancer through inhibition of doublecortin-like kinase 1. Mol Cancer 2014; 13: 103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wenzel ES, Singh ATK. Cell-cycle Checkpoints and Aneuploidy on the Path to Cancer. In Vivo 2018; 32(1): 1–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Whittaker SR, Mallinger A, Workman P, Clarke PA. Inhibitors of cyclin-dependent kinases as cancer therapeutics. Pharmacol Ther 2017; 173: 83–105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Jin MH, Oh DY. ATM in DNA repair in cancer. Pharmacol Ther 2019: 107391. [DOI] [PubMed]
- 31.Zhang P, Wei Y, Wang L, Debeb BG, Yuan Y, Zhang J et al. ATM-mediated stabilization of ZEB1 promotes DNA damage response and radioresistance through CHK1. Nat Cell Biol 2014; 16(9): 864–75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Han T, Lin J, Wang Y, Fan Q, Sun H, Tao Y et al. Forkhead box D1 promotes proliferation and suppresses apoptosis via regulating polo-like kinase 2 in colorectal cancer. Biomed Pharmacother 2018; 103: 1369–1375. [DOI] [PubMed] [Google Scholar]
- 33.Xie Y, Liu Y, Li Q, Chen J. Polo-like kinase 2 promotes chemoresistance and predicts limited survival benefit from adjuvant chemotherapy in colorectal cancer. Int J Oncol 2018. [DOI] [PMC free article] [PubMed]
- 34.Kolluri SK, Jin UH, Safe S. Role of the aryl hydrocarbon receptor in carcinogenesis and potential as an anti-cancer drug target. Arch Toxicol 2017; 91(7): 2497–2513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Xue P, Fu J, Zhou Y. The Aryl Hydrocarbon Receptor and Tumor Immunity. Front Immunol 2018; 9: 286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Foster JS, Henley DC, Bukovsky A, Seth P, Wimalasena J. Multifaceted regulation of cell cycle progression by estrogen: regulation of Cdk inhibitors and Cdc25A independent of cyclin D1-Cdk4 function. Mol Cell Biol 2001; 21(3): 794–810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Chandrakesan P, May R, Weygant N, Qu D, Berry WL, Sureban SM et al. Intestinal tuft cells regulate the ATM mediated DNA Damage response via Dclk1 dependent mechanism for crypt restitution following radiation injury. Sci Rep 2016; 6: 37667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Yin L, Chang C, Xu C. G2/M checkpoint plays a vital role at the early stage of HCC by analysis of key pathways and genes. Oncotarget 2017; 8(44): 76305–76317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Qiu Z, Oleinick NL, Zhang J. ATR/CHK1 inhibitors and cancer therapy. Radiother Oncol 2018; 126(3): 450–464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kastan MB, Bartek J. Cell-cycle checkpoints and cancer. Nature 2004; 432(7015): 316–23. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.