Abstract
Background
In late December 2019, Chinese health authorities reported an outbreak of pneumonia of unknown origin in Wuhan, Hubei Province.
Summary
A few days later, the genome of a novel coronavirus was released (http://virological.org/t/novel-2019-coronavirus-genome/319; Wuhan-Hu-1, GenBank accession No. MN908947) and made publicly available to the scientific community. This novel coronavirus was provisionally named 2019-nCoV, now SARS-CoV-2 according to the Coronavirus Study Group of the International Committee on Taxonomy of Viruses. SARS-CoV-2 belongs to the Coronaviridae family, Betacoronavirus genus, subgenus Sarbecovirus. Since its discovery, the virus has spread globally, causing thousands of deaths and having an enormous impact on our health systems and economies. In this review, we summarize the current knowledge about the epidemiology, phylogenesis, homology modeling, and molecular diagnostics of SARS-CoV-2.
Key Messages
Phylogenetic analysis is essential to understand viral evolution, whereas homology modeling is important for vaccine strategies and therapies. Highly sensitive and specific diagnostic assays are key to case identification, contact tracing, identification of the animal source, and implementation of control measures.
Keywords: COVID-19, SARS-CoV-2, Pandemic, Phylogenesis, Protein modeling, Real-time polymerase chain reaction
Introduction
In December 2019, an outbreak of pneumonia of unknown origin was reported in Wuhan, Hubei Province, China. Most of these cases were epidemiologically linked to the Huanan Seafood Wholesale Market. Inoculation of bronchoalveolar lavage fluid obtained from patients with pneumonia of unknown origin into human airway epithelial cells and Vero E6 and Huh7 cell lines led to the isolation of a novel coronavirus, SARS-CoV-2, previously named 2019-nCov [1].
Coronaviruses belong to the family Coronaviridae and are positive single-stranded RNA viruses surrounded by an envelope. They are divided into four genera: Alpha-, Beta-, Gamma-, and Deltacoronavirus. To date, seven human coronaviruses (HCoVs) have been identified, which fall within the Alpha- and Betacoronavirus genera. The Alphacoronavirus genus includes HCoV-NL63 and HCoV-229E, while the Betacoronavirus genus comprises HCoV-OC43, HCoV-HKU1, SARS-CoV (severe acute respiratory syndrome coronavirus), MERS-CoV (Middle East respiratory syndrome-related coronavirus), and the novel SARS-CoV-2 (severe acute respiratory syndrome coronavirus 2) [2, 3, 4, 5, 6, 7]. The alpha-coronaviruses HCoV-NL63 and HCoV-229E and the betacoronaviruses HCoV-OC43 and HCoV-HKU1 usually cause common colds, but also severe lower respiratory tract infections, especially in the elderly and children [8]. HCoV-NL63 infection has also been significantly associated with croup (laryngotracheitis) [9, 10], and HCoV-OC43 infection with severe lower respiratory tract infection in children [11].
SARS-CoV and MERS-CoV are zoonotic in origin; they cause severe respiratory syndrome and are often fatal [12]. Since the beginning of the epidemic in late December 2019, SARS-CoV-2 has now spread to all continents, and as of March 18, 2020, the WHO communicated 179,111 confirmed cases and 7,426 deaths globally (Situation Report-57). In this review, we try to summarize the most recent knowledge about some epidemiological parameters including clinical symptoms, transmissibility of the virus, and the incubation period. Furthermore, the molecular diagnostics, protein modeling of the spike glycoprotein, and phylogenesis of the virus will be discussed.
SARS-CoV-2 Epidemiology
Clinical Symptoms
Patients infected with SARS-CoV-2 can present a wide range of symptoms ranging from mild to severe. Fever, cough, and shortness of breath are the most common symptoms reported in 83, 82, and 31% of patients [13]. In those patients who develop pneumonia, multiple mottling and ground-glass opacity are described on chest X-ray [1, 13]. Patients that develop acute respiratory distress syndrome may worsen rapidly and die of multiple organ failure [13].
It has also been reported that about 2–10% of the patients with COVID-19 had gastrointestinal symptoms such as vomiting, diarrhea, and abdominal pain [13, 14]. Diarrhea and nausea preceded the development of fever and respiratory symptoms in 10% of patients [13].
Viral Transmission
At present, the exact mechanism of transmission of SARS-CoV-2 is still not completely understood. Human-to-human transmission via droplets is the main route of transmission within a susceptible population. Chinese health authorities reported an R0 of 1.4–2.5 on January 23, 2020, to the WHO International Health Regulations (2005) Emergency Committee. Transmission by asymptomatic carriers cannot be ruled out. Actually, it was reported that an asymptomatic family member who traveled from the epidemic center of Wuhan was most likely responsible for a familial cluster of COVID-19 pneumonia once back home. Her reverse transcription polymerase chain reaction (RT-PCR) result was positive for SARS-CoV-2, but her chest CT images did not show significant alterations [15].
Another route of possible viral transmission is the oral-fecal route. The scientific literature showed that SARS-CoV and MERS-CoV are viable in environmental conditions that facilitate oral-fecal transmission. SARS-CoV has been detected in sewage water of two Chinese hospitals in which patients with SARS were treated, and MERS-CoV was found to be viable on different surfaces at low temperature and low humidity [16, 17]. SARS-CoV-2 was detected in stool of patients with COVID-19 pneumonia, as well as in respiratory samples [18]. Thus, it is plausible that also SARS-CoV-2 can be transmitted via the oral-fecal route as well as via fomites.
Incubation Period
To know the incubation period of SARS-CoV-2 infection is key for implementing control measures and surveillance. It has been estimated that the median incubation period is 5.1 days (95% CI, 4.5–5.8), and 97.5% of the infected subjects will develop symptoms within 11.5 days (95% CI, 8.2–15.6) of infection. Based on these estimates, it can be assumed that 101 out of 10,000 cases will develop symptoms after 14 days of observation or quarantine [19]. These estimates are consistent with those of other studies that reported a mean incubation period of 6.4 days (95% credible interval: 5.6–7.7), ranging from 2.1 to 11.1 days (2.5th to 97.5th percentile) [20] or 5.2 days (95% CI, 4.1–7.0), with the 95th percentile of the distribution at 12.5 days [21]. Thus, 14-day monitoring is advised following contact with a probable or confirmed SARS-CoV-2 case [22].
Molecular Diagnosis of SARS-CoV-2
Confirmation of cases with suspected SARS-CoV-2 infection is performed by detection of unique viral sequences with nucleic acid amplification tests such as reverse real-time PCR (rRT-PCR). As soon as on January 7, 2020, the Chinese health authorities had declared that a novel coronavirus was responsible for this outbreak of pneumonia in Wuhan, a European network of academic and public laboratories designed an rRT-PCR protocol based on the comparison and alignment of previously available SARS-CoV and bat-related coronavirus genome sequences as well as five sequences derived from the novel coronavirus SARS-CoV-2 made available by the Chinese authorities [23]. Three assays were developed. The first-line assay targets the E gene encoding for the envelope protein, which is common to the Sarbecovirus subgenus, while the second specific assay targets the RdRp gene encoding for RNA-dependent RNA polymerase. This assay contains two probes: one probe, which reacts with the SARS-CoV and SARS-CoV-2 RdRp gene, and a second probe (RdRP_SARSr-P2) which is specific to SARS-CoV-2. Finally, the third additional confirmatory assay targets the nucleocapsid (N) gene. This last assay was not further validated because it is slightly less sensitive [23]. This protocol was adopted in more than 30 European laboratories [24]. Recently, a novel rRT-PCR assay targeting a different region of the RdRp/Hel gene of SARS-CoV-2 has been developed that showed a higher sensitivity and specificity than the RdRp-P2 assay [25].
Currently, several amplification protocols are available on the market and validated for in vitro diagnostic use (CE marked): GeneFinderTM COVID-19 Plus RealAmp Kit (OSANG Healthcare Co., Ltd, South Korea); genesig® Real-Time PCR Coronavirus (COVID-19) (genesig, UK); AllplexTM 2019-nCoV Assay (Seegene, South Korea), etc. Highly sensitive and specific diagnostic assays are key to the identification of cases, contact tracing, identification of the animal source, and implementation of control measures [26, 27, 28].
When performing nucleic acid amplification test assays, it is useful to remind ourselves that several factors can be responsible for a negative result in an infected individual, such as the poor quality of a specimen, the time of specimen collection (specimen collected too early or too late during infection), inappropriate handling or shipment of the specimen, and technical reasons.
SARS-CoV-2 Modeling
Coronavirus entry into the host cell is mediated by the transmembrane spike (S) glycoprotein that forms homotrimers that protrude from the viral surface [29]. The S protein is composed of the two subunits S1 and S2 responsible for binding to the host cell receptor and fusion of the viral and cellular membranes, respectively. Different coronaviruses use different domains within the S1 subunit to enter the cell. These domains are named SA and SB. SARS-CoV and SARS-related coronaviruses interact with the angiotensin-converting enzyme 2 (ACE2) via domain SB to enter target cells [30, 31, 32, 33, 34]. It has recently been shown that SARS-CoV-2 binds the ACE2 receptor via the SB domain similarly to SARS-CoV, and that murine polyclonal antibodies inhibited SARS-CoV-2 entry into the cell mediated by S. These data suggest that cross-neutralizing antibodies targeting conserved S epitopes elicited by vaccination could be used against SARS-CoV-2, SARS-CoV, and SARS-related coronaviruses [35].
Previous studies have shown the presence of positive selective pressure on the Nucleocapsid, Spike glycoprotein, and ORF1ab regions, while until now no evidence of a positive selective pressure has been found on the Envelope, Membrane, and other ORF proteins.
In the Nucleocapsid region, significant (p < 0.05) pervasive episodic selection was found in 2 sites. In amino acid position 380 of the Wuhan coronavirus sequence there is a Gln residue instead of an Asn, while in amino acid position 410 there is a Thr residue instead of an Ala. Significant (p < 0.05) pervasive negative selection in 6 sites (14%) has been evidenced and confirmed by FUBAR (Fast Unconstrained Bayesian Approximation) analysis [36].
In the Spike glycoprotein region, significant (p < 0.05) pervasive episodic selection was found in 2 different sites (536th and 644th nucleotide position using the reference sequence). In the 536th amino acid position of the Wuhan coronavirus sequence there is an Asn residue instead of an Asp acid residue, while in amino acid position 644 there is a Thr residue instead of an Ala residue. Significant (p < 0.05) pervasive negative selection in 1,065 sites (87%) has been evidenced and confirmed by FUBAR analysis, suggesting that the S region could be highly conserved [36].
Regarding the sites under positive selective pressure found on the Spike glycoprotein, the results have shown that amino acid position 536 in COVID-19 has an Asn residue, while the Bat SARS-like coronavirus has a Gln residue; the SARS virus, instead, has an Asp residue. In amino acid position 644 of the COVID-19 sequence there is a Thr residue, while the Bat SARS-like virus has a Ser residue; instead, the SARS virus has an Ala residue. Another study highlighted that several key residues responsible for binding of the SARS-CoV receptor-binding domain to the ACE2 receptor were variable in the COVID-19 receptor-binding domain (including Asn439, Asn501, Gln493, Gly485, and Phe486; COVID-19 numbering), and a number of deletion events in amino acid positions 455–457, 463–464, and 485–497 occurred in the bat-derived strains [37].
Also in the ORF1ab region, potential sites under positive selective pressure have been found (p < 0.05). Particularly, in the amino acid position 501, COVID-19 has a Gln residue, the Bat SARS-like coronavirus has a Thr residue, and the SARS virus has an Ala residue. In position 723 of the COVID-19 sequence there is a Ser residue, while the Bat SARS-like virus and the SARS virus have a Gly residue. In amino acid position 1,010, COVID-19 has a Pro residue, the Bat SARS-like coronavirus has a His residue, and the SARS virus has an Ile residue.
As for the residue in position 723 (543 in the nsp3 protein), the COVID-19 sequence displays a Ser, replacing for Gly in the Bat SARS-like and SARS coronaviruses. In this case, it may be argued that this substitution could increase local stiffness of the polypeptide chain both for a steric effect (in contrast to Ser, Gly has no side chain) and for the ability of the Ser side chain to form H-bonds. Moreover, Ser can act as a nucleophile in determined structural environments, such as those of enzymes' active sites, and can be a phosphorylation site. However, within the I-TASSER model, this position is predicted to have low solvent accessibility.
Regarding the amino acid position 1,010 (corresponding to position 192 of the nsp3 protein), the homologous region of the Bat SARS-like coronavirus and SARS virus has a polar and an apolar amino acid, respectively, while COVID-19 has a Pro residue. In this case, it may be speculated that due to the steric bulge and stiffness of Pro, the molecular structure of COVID-19 may undergo a local conformational perturbation compared to the proteins of the other two viruses. In nsp3, the mutation falls near the polyprotein domain similar to a phosphatase present also in the SARS coronavirus (PDB code 2ACF) playing a key role in the replication process of the virus in infected cells [38]. According to the I-TASSER model, the position is partially accessible to the solvent.
The sites under positive selective pressure in this protein may suggest a possible interpretation of some clinical features of this virus compared to SARS and Bat SARS-like coronavirus. This analysis should find which are probably the most common sites undergoing an amino acid change, providing insight into some important proteins of COVID-19 that are involved in the mechanism of viral entry and viral replication. These data should contribute to improving our understanding of how this virus acts in its pathogenicity. Furthermore, to identify a potential molecular target is fundamental to follow the molecular evolution of the virus, which can suggest some interesting sites for a potential therapy or vaccine.
The structural similarity of the region in which the positive selective pressure occurs, and the stabilizing mutation falling in the endosome-associated protein-like domain of the nsp2 protein, should be probable reasons why this virus is more contagious than SARS. Instead, the destabilizing mutation located near the phosphatase domain of the nsp3 protein may explain why viral replication is slower than in SARS with a longer incubation period. Anyway, further studies are needed on this aspect [39].
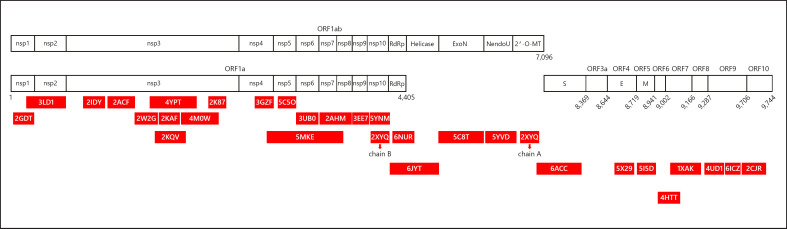
The availability of protein structural information is an essential prerequisite for the interpretation of biological phenomena. In this case, knowledge of the virus's protein structure would greatly enhance the possibility of understanding the biological meaning of the observed mutations. Now, only the X-ray structure of COVID-19 nsp5 protease (PBD code 6LU7) is available, although it is expected that many other structures will become available soon. In the meantime, homology modeling could provide preliminary structural clues. Homology modeling needs structural templates sharing sufficient sequence similarity to the targets. In Figure 1 and Table 1, a list of potential templates for homology modeling of the proteins coded by the COVID-19 genome is displayed. The structures with the largest coverage and the greatest sequence identity have been incorporated into Figure 1 and Table 1. According to this list, it is evident that most of the viral proteins are at modeling distance from PDB structures. This information should be exploited as soon as possible.
Fig. 1.
HHpred mapping of the homologous structures of all proteins coded by the 2019-nCoV genome. Numbers below the open rectangles refer to all open reading frames (ORFs) detected. Red strips represent the PDB (Protein Data Bank) homologous structures. PDB codes are reported within the corresponding red stripes. Details are given in Table 1.
Table 1.
PDB codes of all the homologous models showing similarity with the relative SARS-CoV-2 region, reporting the protein function, the sequence position within the viral ORFs, and the percentage of identity
| PDB code | Protein | Sequence position within virus ORF | % identity |
|---|---|---|---|
| 2GDT | nsp1 from SARS coronavirus | 13–127 | 86% |
| 3LD1 | nsp2a from IBV | 105–564 | 11% |
| 2IDY | nsp3 from SARS coronavirus | 818–929 | 76% |
| 2ACF | nsp3 from SARS coronavirus | 1,025–1,197 | 72% |
| 2W2G | human SARS coronavirus unique domain | 1,231–1,494 | 75% |
| 2KQV | human SARS coronavirus unique domain | 1,368–1,561 | 75% |
| 2KAF | nsp3 from human coronavirus, C-domain | 1,496–1,561 | 70% |
| 4YPT | nsp3 from murine hepatitis virus | 1,449–1,872 | 27% |
| 4M0W | structure papain-like protease from human SARS coronavirus | 1,563–1,881 | 82% |
| 2K87 | putative RNA-binding protein from human SARS coronavirus | 1,906–2,021 | 81% |
| 3GZF | nsp4 from feline coronavirus, C-domain | 3,164–3,260 | 36% |
| 5C5O | protease complex from human SARS-3CL protease complex | 3,264–3,569 | 96% |
| 5MK5 | DHBN domain from human BLM helicase | 3,636–4,042 | 11% |
| 3UB0 | nsp7 and 8 complex from feline coronavirus | 3,857–3,942 | 41% |
| 2AHM | nsps from human SARS coronavirus | 3,942–4,140 | 97% |
| 3EE7 | nsp9 from human SARS coronavirus | 4,141–4,253 | 96% |
| 5YNM | nsp16/nsp10 complex from human betacoronavirus | 4,254–4,394 | 59% |
| 2XYQ chain B | nsp16/nsp10 complex from human SARS coronavirus | 4,264–4,385 | 98% |
| 6NUR | nsp12 from human SARS coronavirus | 4,394–5,329 | 96% |
| 6JYT | nsp13 from human SARS coronavirus | 5,325–5,926 | 100% |
| 5C8T | nsp14/nsp10 complex from human SARS coronavirus | 5,927–6,453 | 95% |
| 5YVD | nsp15 from MERS coronavirus | 6,454–6,799 | 51% |
| 2XYQ chain A | nsp16/nsp10 complex from human SARS coronavirus | 6,800–7,089 | 93% |
| 6ACC | spike glycoprotein from human SARS coronavirus | 7,098–8,312 | 76% |
| 5X29 | protein E from human SARS coronavirus | 8,648–8,710 | 87% |
| 5I5D | domain 245 from salmonella global | 8,724–8,817 | 14% |
| 4HTT | receptor TatC in DDM from Aquifex aeolicus | 8,943–9,022 | 19% |
| 1XAK | ORF7 from human SARS coronavirus | 9,017–9,098 | 88% |
| 4UD1 | nucleocapsid from MERS coronavirus, domain N-term | 9,253–9,420 | 50% |
| 6ICZ | human P complex | 9,422–9,454 | 36% |
| 2CJR | nucleocapsid protein SARS | 9,492–9,609 | 96% |
PDB, Protein Data Bank; ORF, open reading frame.
SARS-CoV-2 Phylogeny
Phylogenetic analysis of the SARS-CoV-2 genomes showed that the novel coronavirus responsible for the pneumonia outbreak in Wuhan, China, belongs to the Betacoronavirus genus, subgenus Sarbecovirus [37]. Within the Betacoronavirus genus, 2019-nCov (SARS-CoV-2) is distant from SARS-CoV (about 79% identity) and MERS-CoV (about 50% identity) responsible for the 2002–2003 [4] and 2012 [7] epidemics, respectively, but closely related (88% identity) to the two bat-derived (SARS)-like coronaviruses bat-SL-CoVZC45 and bat-SL-CoVZXC21 [37].
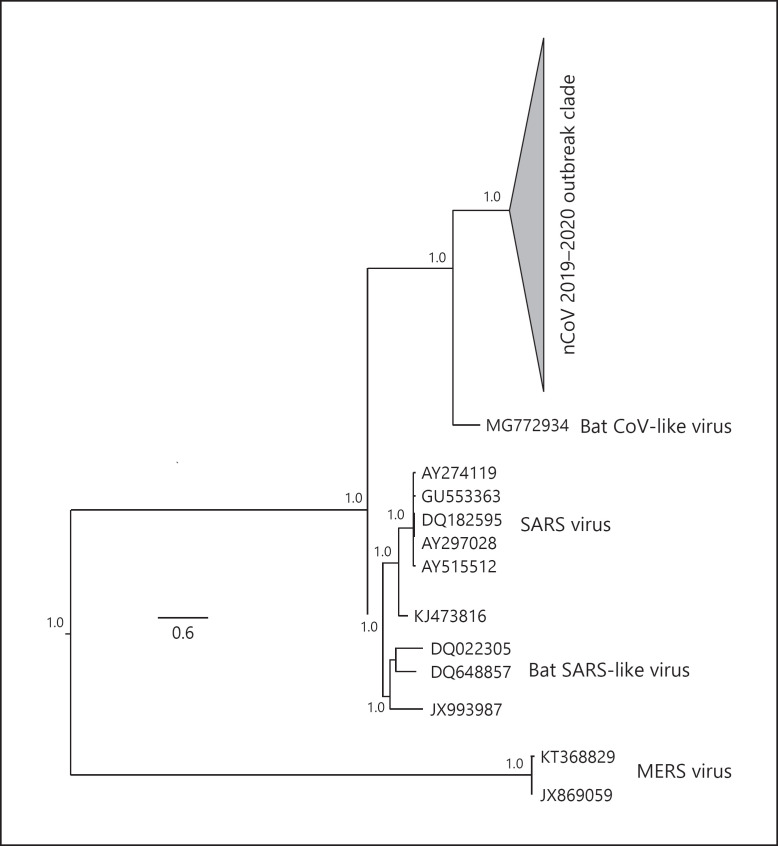
The origin of the virus is still unclear; however, genomic analysis suggests that SARS-CoV-2 is most closely related to viruses previously identified in bats (Fig. 2). It is plausible that there were other intermediate animal transmissions before its introduction into humans. However, there is no evidence of snakes as an intermediary [36].
Fig. 2.
Phylogenetic tree of all the 2019-nCov sequences available on February 9, 2020 (n = 74, cartooned outbreak clade), plus n = 1 Bat CoV-like sequence, n = 5 Human SARS, n = 4 Bat SARS-like sequences, and n = 2 from MERS. Numbers along the branches represent a bootstrap value >0.90. The scale bar represents expected substitutions per nucleotide site.
Using 74 publicly shared novel coronavirus (nCoV) genomes, we examined genetic diversity to infer the date of the common ancestor and the rate of spread. The high similarity of the genomes suggests they share a recent common ancestor. Otherwise, we would expect a greater number of differences between the samples. The jump from bats to humans most likely occurred in late November or early December 2019 (November 25, 2019; 95% HPD: September 28, 2019; December 21, 2019) [40].
Previous research on related coronaviruses suggests that these viruses accumulate between 1 and 3 changes in their genome per month (rates of 3 × 10−4 to 1 × 10−3 per site per year). Molecular clock calibration estimated the evolutionary rate of the SARS-CoV-2 whole genome sequences at 6.58 × 10−3 substitutions per site per year (95% HPD: 5.2 × 10−3 to 8.1 × 10−3).
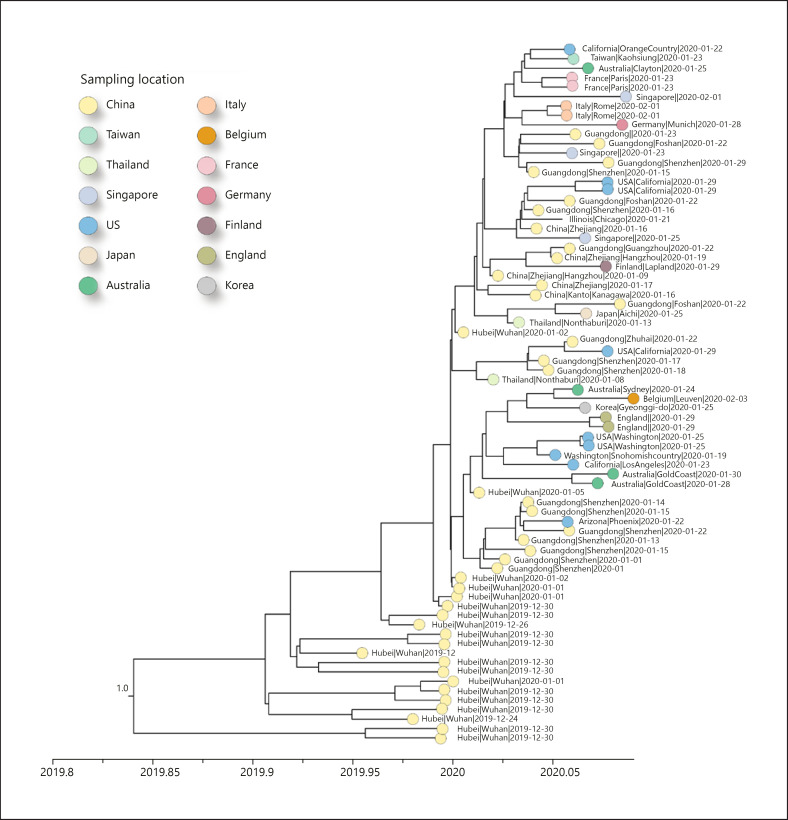
The outbreak first started in Wuhan, China, but cases have been identified in many East and South-East Asian countries, the USA, Australia, the Middle East, and Europe. Vietnam, Japan, and Germany have reported transmission within the country, albeit always with a known link to Wuhan, China (Fig. 3).
Fig. 3.
Maximum clade credibility phylogeny estimated using all the complete and near-complete 2019-nCov genome sequences available up to February 9, 2020 (n = 74). Clade posterior probabilities are shown at well-supported nodes. The colors represent different locations.
Conclusions
This study is a picture of the current research on molecular evolution, epidemiology, and diagnostics in response to the outbreak of COVID-19. Many studies have been published within different scientific disciplines with the intent to control and prevent this pandemic. Phylogenetic analysis and homology modeling add new knowledge together with epidemiological and diagnostic methods. Studies exploring the genome and the structure of the viral proteins are essential in order to define prevention and control measures to minimize the impact of the outbreak. All this knowledge will pave the way for the development of a vaccine and antiviral therapy.
Disclosure Statement
The authors have no conflicts of interest to declare.
Funding Sources
There was no funding for this review.
Author Contributions
M. Ciotti, S. Angeletti, and M. Ciccozzi: concept and writing; S. Pascarella and M. Bianchi: protein modeling paragraph; M. Ciotti, S. Bernardini, and M. Minieri: molecular diagnostics paragraph; M. Giovannetti, D. Benvenuto, and M. Ciccozzi: phylogenesis; M. Ciotti and C. Sagnelli: epidemiology.
References
- 1.Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, et al. China Novel Coronavirus Investigating and Research Team A Novel Coronavirus from Patients with Pneumonia in China, 2019. N Engl J Med. 2020 Feb;382((8)):727–33. doi: 10.1056/NEJMoa2001017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Almeida JD, Tyrrell DA. The morphology of three previously uncharacterized human respiratory viruses that grow in organ culture. J Gen Virol. 1967 Apr;1((2)):175–8. doi: 10.1099/0022-1317-1-2-175. [DOI] [PubMed] [Google Scholar]
- 3.Kapikian AZ, James HD, Jr, Kelly SJ, Dees JH, Turner HC, McIntosh K, et al. Isolation from man of “avian infectious bronchitis virus-like” viruses (coronaviruses) similar to 229E virus, with some epidemiological observations. J Infect Dis. 1969 Mar;119((3)):282–90. doi: 10.1093/infdis/119.3.282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Peiris JS, Guan Y, Yuen KY. Severe acute respiratory syndrome. Nat Med. 2004 Dec;10((12 Suppl)):S88–97. doi: 10.1038/nm1143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.van der Hoek L, Pyrc K, Jebbink MF, Vermeulen-Oost W, Berkhout RJ, Wolthers KC, et al. Identification of a new human coronavirus. Nat Med. 2004 Apr;10((4)):368–73. doi: 10.1038/nm1024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Woo PC, Lau SK, Chu CM, Chan KH, Tsoi HW, Huang Y, et al. Characterization and complete genome sequence of a novel coronavirus, coronavirus HKU1, from patients with pneumonia. J Virol. 2005 Jan;79((2)):884–95. doi: 10.1128/JVI.79.2.884-895.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Zaki AM, van Boheemen S, Bestebroer TM, Osterhaus AD, Fouchier RA. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N Engl J Med. 2012 Nov;367((19)):1814–20. doi: 10.1056/NEJMoa1211721. [DOI] [PubMed] [Google Scholar]
- 8.Wevers BA, van der Hoek L. Recently discovered human coronaviruses. Clin Lab Med. 2009 Dec;29((4)):715–24. doi: 10.1016/j.cll.2009.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.van der Hoek L, Sure K, Ihorst G, Stang A, Pyrc K, Jebbink MF, et al. Croup is associated with the novel coronavirus NL63. PLoS Med. 2005 Aug;2((8)):e240. doi: 10.1371/journal.pmed.0020240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Choi EH, Lee HJ, Kim SJ, Eun BW, Kim NH, Lee JA, et al. The association of newly identified respiratory viruses with lower respiratory tract infections in Korean children, 2000-2005. Clin Infect Dis. 2006 Sep;43((5)):585–92. doi: 10.1086/506350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zhu Y, Li C, Chen L, Xu B, Zhou Y, Cao L, et al. A novel human coronavirus OC43 genotype detected in mainland China. Emerg Microbes Infect. 2018 Oct;7((1)):173. doi: 10.1038/s41426-018-0171-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cui J, Li F, Shi ZL. Origin and evolution of pathogenic coronaviruses. Nat Rev Microbiol. 2019 Mar;17((3)):181–92. doi: 10.1038/s41579-018-0118-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J, et al. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus–infected pneumonia in Wuhan, China. JAMA. 2020 Feb;323((11)):1061. doi: 10.1001/jama.2020.1585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020 Feb;395((10223)):507–13. doi: 10.1016/S0140-6736(20)30211-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Bai Y, Wei LYT, Tian F, Jin D-Y, Chen L, Wang M. Presumed Asymptomatic Carrier Transmission of COVID-19. JAMA. 2020 doi: 10.1001/jama.2020.2565. [Epub ahead of print] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Wang XW, Li J, Guo T, Zhen B, Kong Q, Yi B, et al. Concentration and detection of SARS coronavirus in sewage from Xiao Tang Shan Hospital and the 309th Hospital of the Chinese People's Liberation Army. Water Sci Technol. 2005;52((8)):213–21. [PubMed] [Google Scholar]
- 17.van Doremalen N, Bushmaker T, Munster VJ. Stability of Middle East respiratory syndrome coronavirus (MERS-CoV) under different environmental conditions. Euro Surveill. 2013 Sep;18((38)):20590. doi: 10.2807/1560-7917.es2013.18.38.20590. [DOI] [PubMed] [Google Scholar]
- 18.Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H, et al. Washington State 2019-nCoV Case Investigation Team First Case of 2019 Novel Coronavirus in the United States. N Engl J Med. 2020 Mar;382((10)):929–36. doi: 10.1056/NEJMoa2001191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lauer SA, Grantz KH, Bi Q, Jones FK, Zheng Q, Meredith HR, et al. The Incubation Period of Coronavirus Disease 2019 (COVID-19) From Publicly Reported Confirmed Cases: estimation and Application. Ann Intern Med. 2020 doi: 10.7326/M20-0504. [Epub ahead of print] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Backer JA, Klinkenberg D, Wallinga J. Incubation period of 2019 novel coronavirus (2019-nCoV) infections among travellers from Wuhan, China, 20-28 January 2020. Euro Surveill. 2020 Feb;25((5)) doi: 10.2807/1560-7917.ES.2020.25.5.2000062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Li Q, Guan X, Wu P, Wang X, Zhou L, Tong Y, et al. Early Transmission Dynamics in Wuhan, China, of Novel Coronavirus-Infected Pneumonia. N Engl J Med. 2020 Jan;:NEJMoa2001316. doi: 10.1056/NEJMoa2001316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.European Centre for Disease Prevention and Control Algorithm for the management of contacts of probable or confirmed COVID-19 cases [2020 Feb 25] Available from: https://www.ecdc.europa.eu/en/publications-data/algorithm-management-contacts-probableor-confirmed-covid-19-cases.
- 23.Corman VM, Landt O, Kaiser M, Molenkamp R, Meijer A, Chu DK, et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 2020 Jan;25((3)):2000045. doi: 10.2807/1560-7917.ES.2020.25.3.2000045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Reusken C, Broberg EK, Haagmans B, Meijer A, Corman VM, Papa A, et al. Laboratory readiness and response for novel coronavirus (2019-nCoV) in expert laboratories in 30 EU/EEA countries, January 2020. Euro Surveill. 2020 Feb;25((6)):2000082. doi: 10.2807/1560-7917.ES.2020.25.6.2000082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Chan JF, Yip CC, To KK, Tang TH, Wong SC, Leung KH, et al. Improved molecular diagnosis of COVID-19 by the novel, highly sensitive and specific COVID-19-RdRp/Hel real-time reverse transcription-polymerase chain reaction assay validated in vitro and with clinical specimens. J Clin Microbiol. 2020 doi: 10.1128/JCM.00310-20. [Epub ahead of print] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Peiris JS, Lai ST, Poon LL, Guan Y, Yam LY, Lim W, et al. SARS study group Coronavirus as a possible cause of severe acute respiratory syndrome. Lancet. 2003 Apr;361((9366)):1319–25. doi: 10.1016/S0140-6736(03)13077-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Cheng VC, Lau SK, Woo PC, Yuen KY. Severe acute respiratory syndrome coronavirus as an agent of emerging and reemerging infection. Clin Microbiol Rev. 2007 Oct;20((4)):660–94. doi: 10.1128/CMR.00023-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Chan JF, Lau SK, To KK, Cheng VC, Woo PC, Yuen KY. Middle East respiratory syndrome coronavirus: another zoonotic betacoronavirus causing SARS-like disease. Clin Microbiol Rev. 2015 Apr;28((2)):465–522. doi: 10.1128/CMR.00102-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Tortorici MA, Veesler D. Structural insights into coronavirus entry. Adv Virus Res. 2019;105:93–116. doi: 10.1016/bs.aivir.2019.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Ge XY, Li JL, Yang XL, Chmura AA, Zhu G, Epstein JH, et al. Isolation and characterization of a bat SARS-like coronavirus that uses the ACE2 receptor. Nature. 2013 Nov;503((7477)):535–8. doi: 10.1038/nature12711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kirchdoerfer RN, Wang N, Pallesen J, Wrapp D, Turner HL, Cottrell CA, et al. Stabilized coronavirus spikes are resistant to conformational changes induced by receptor recognition or proteolysis. Sci Rep. 2018 Oct;8((1)):15701. doi: 10.1038/s41598-018-34171-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Song W, Gui M, Wang X, Xiang Y. Cryo-EM structure of the SARS coronavirus spike glycoprotein in complex with its host cell receptor ACE2. PLoS Pathog. 2018 Aug;14((8)):e1007236. doi: 10.1371/journal.ppat.1007236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Li W, Moore MJ, Vasilieva N, Sui J, Wong SK, Berne MA, et al. Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature. 2003 Nov;426((6965)):450–4. doi: 10.1038/nature02145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Li F, Li W, Farzan M, Harrison SC. Structure of SARS coronavirus spike receptor-binding domain complexed with receptor. Science. 2005 Sep;309((5742)):1864–8. doi: 10.1126/science.1116480. [DOI] [PubMed] [Google Scholar]
- 35.Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell. 2020 Mar;180:1–12. doi: 10.1016/j.cell.2020.02.058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Benvenuto D, Giovanetti M, Ciccozzi A, Spoto S, Angeletti S, Ciccozzi M. The 2019-new coronavirus epidemic: evidence for virus evolution. J Med Virol. 2020 Apr;92((4)):455–9. doi: 10.1002/jmv.25688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020 Feb;395((10224)):565–74. doi: 10.1016/S0140-6736(20)30251-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Saikatendu KS, Joseph JS, Subramanian V, Clayton T, Griffith M, Moy K, et al. Structural basis of severe acute respiratory syndrome coronavirus ADP-ribose-1′'-phosphate dephosphorylation by a conserved domain of nsP3. Structure. 2005 Nov;13((11)):1665–75. doi: 10.1016/j.str.2005.07.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Angeletti S, Benvenuto D, Bianchi M, Giovanetti M, Pascarella S, Ciccozzi M. COVID-2019: the role of the nsp2 and nsp3 in its pathogenesis. J Med Virol. 2020 doi: 10.1002/jmv.25719. [Epub ahead of print] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Benvenuto D, Giovanetti M, Salemi M, Prosperi M, De Flora C, Junior Alcantara LC, et al. The global spread of 2019-nCoV: a molecular evolutionary analysis. Pathog Glob Health. 2020 doi: 10.1080/20477724.2020.1725339. [Epub ahead of print] [DOI] [PMC free article] [PubMed] [Google Scholar]