Abstract
Diaminoquinazolines represent a privileged scaffold for antimalarial discovery, including use as putative Plasmodium histone lysine methyltransferase inhibitors. Despite this, robust evidence for their molecular targets is lacking. Here we report the design and development of a small-molecule photo-crosslinkable probe to investigate the targets of our diaminoquinazoline series. We demonstrate the effectiveness of our designed probe for photoaffinity labelling of Plasmodium lysates and identify similarities between the target profiles of the probe and the representative diaminoquinazoline BIX-01294. Initial pull-down proteomics experiments identified 104 proteins from different classes, many of which are essential, highlighting the suitability of the developed probe as a valuable tool for target identification in Plasmodium falciparum.
Keywords: malaria, proteomics, photoaffinity probe, pull-down, BIX-01294
Malaria remains one of the deadliest diseases in the developing world. In 2015 there were 212 million cases worldwide, claiming 429 000 lives, mostly from children under 5 years old, with over 90% of deaths occurring in Sub-Saharan Africa.1 Malaria is caused by a protozoan parasite of the Plasmodium spp., of which Plasmodium falciparum is responsible for the highest mortality, and is transmitted to humans by infected mosquitos. While the recommended treatment for malaria relies on artemisinin-based combination therapies, artemisinin resistance is now emerging,2–4 and therefore there is an urgent need for new treatments, especially drugs with novel mechanisms of action.
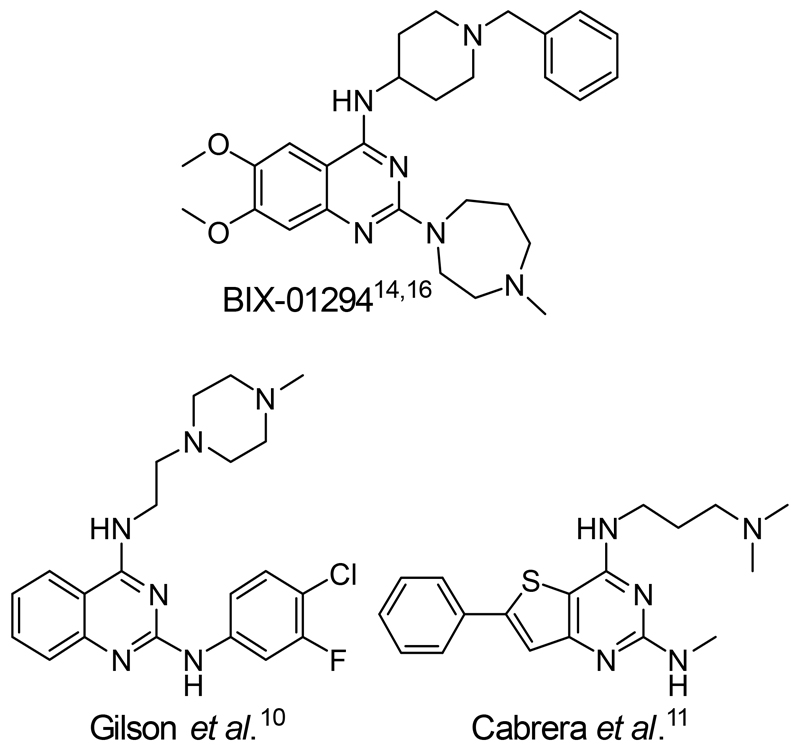
Compounds bearing the heteroaromatic quinazoline core have long been shown to possess anti-Plasmodium activity (Figure 1).5–8 Diaminoquinazolines specifically were found as one of the 68 parent structures of active chemotypes obtained in a screen of almost two million compounds, carried out by GlaxoSmithKline in the search of new starting points for antimalarial drug discovery.9 Furthermore, a number of groups have optimised diaminoquinazolines10 and related scaffolds11 (Figure 1) for potent and in vivo anti-malarial activity. Despite this, in many cases the precise targets of this class of compounds remain ill-defined, a rare exception being a 2,4-unsubstituted diaminoquinazoline scaffold that was previously reported to target Plasmodium dihydrofolate reductase (PDHFR).12,13 We have explored the diaminoquinazoline scaffold as a means to validate the Plasmodium histone lysine methyltransferase (PHKMT) enzymes as future antimalarial epigenetic drug targets.14–17 Given the ability of diaminoquinazoline analogues to inhibit a range of human HKMT enzymes – G9a,18,19,20 GLP,21 SETD822 and EZH223 – we previously sought to ‘repurpose’ this chemotype as a PHKMT inhibitor. Our compounds were found to exhibit rapid and irreversible asexual cycle blood-stage-independent cytotoxic activity at nM concentrations. Comparable potency against resistant strains (including artemisinin) and clinical isolates of P. falciparum and P. vivax, and oral efficacy in in vivo mouse models of P. berghei and P. falciparum infection were also achieved.14,16 Furthermore, these compounds were effective on other therapeutically-relevant parasite life-cycle stages, inhibiting the progression of sexual gametocytes, essential for transmission of the disease,16 and showing an unprecedented ability to re-awaken the dormant liver-stage forms (hypnozoites) responsible for relapsing malaria.24 Relating such phenotypic activity to target-based activity has remained a challenge however, particularly in the absence of isolated enzymes for robust biochemical/biophysical characterisation. Of the ten predicted PHKMTs, to date, our study on PfSET7 represents the only successful report of recombinant expression of an active PHKMT, suitable for biochemical characterisation.25 In the absence of the full complement of purified PHKMTs – required for robust biochemical characterisation of our lead inhibitors – we have used indirect approaches to confirm on-target activity, including monitoring histone methylation levels in treated cells14,15 and SAR comparisons15,17 to those observed for inhibitors of homologous targets. However, such approaches do not unambiguously confirm the PHKMTs to be the target of this compound class, nor do they discount the potential role of unrelated off-targets in the phenotypic effects observed.
Figure 1.
Chemical structure of key diaminoquinazolines (and related analogues) with potent anti-malarial activity.
Chemical proteomics has emerged as a versatile tool for profiling the targets of a given inhibitor.26–29 Photoaffinity labelling is a particularly powerful methodology30 which allows for covalent linking of an inhibitor bearing an appropriate photoactive group to its molecular targets, when exposed to UV irradiation. Downstream identification (ID) of the resulting protein-inhibitor adducts provides direct experimental evidence for target and off-target engagement. Given the promise of our putative PHKMT series, and the wider interest in diaminoquinazolines as anti-malarial scaffolds, we sought to develop an appropriate diaminoquinazoline photoaffinity probe for target ID studies.
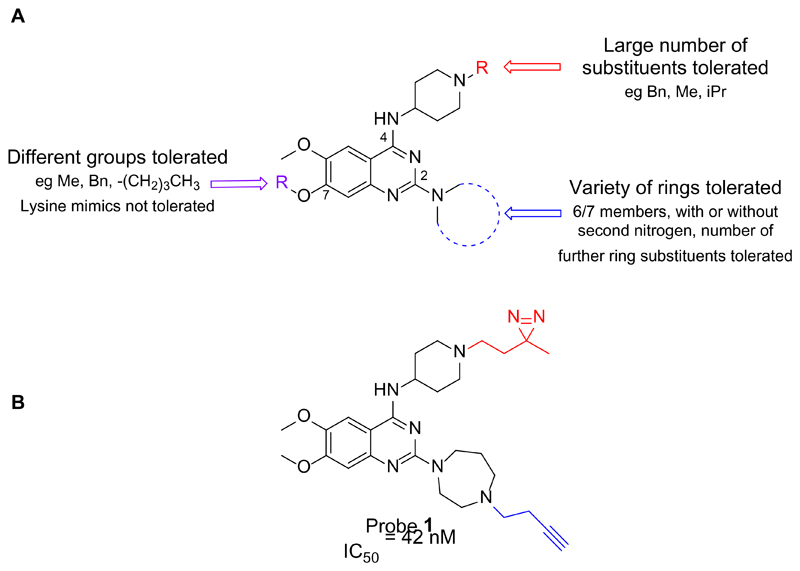
Taking advantage of the structure-activity relationship (SAR) studies previously carried out by our group (Figure 2A), which have led to the establishment of the key features needed for improved parasite activity and selectivity over human cell lines,15,17 we designed and synthesized probe 1 (Figure 2B). The diazirine subunit was selected as a photo-crosslinking group due to its small size, which should have a minimal impact on target-binding, coupled with its established effectiveness in photoaffinity-labelling methodologies.31,32 A terminal alkyne was added to the diazepane ring to allow, after incubation with the desired proteome, visualization, enrichment and identification of probe-labelled proteins through ‘click’ chemistry. Probe 1 was tested against P. falciparum 3D7 strains and showed comparable antimalarial activity (IC50 = 42 ± 20 nM) to that of the parent compound BIX-01294 (IC50 = 50 ± 12 nM). Combined with our extensive SAR,15,17 this result provides confidence that probe 1 is relevant to study the targets of BIX-01294.
Figure 2.
(A) SAR of diaminoquinazolines for parasite-killing activity.15,17 The quinazoline core, N-H at position 4, and basic nitrogen at position 2 are all essential for activity. A large number of substituents can be tolerated on the piperidyl ring at position 4, a number of rings with or without substituents can be introduced at position 2, and there is also tolerance of some substituents at position 7. (B) Chemical structure of diaminoquinazoline probe 1, bearing a photo-crosslinkable diazirine (red) and an alkyne handle (blue). The IC50 of the probe was measured using a 3-day SYBR Green I assay.
The synthesis of probe 1 was carried out according to routes previously developed for this chemical series (see Supporting Information for synthetic details). To confirm that our designed probe was able to undergo effective photoaffinity labelling, we incubated the probe with lysate from saponin-treated pellets derived from 30-40 hour post invasion, blood stage Plasmodium falciparum (3D7 strain) cultures. Labelling used a protocol developed from previously reported methods,32–35 as depicted in Figure 3. Treatment of the lysate with probe 1 (0-100 μM), followed by UV irradiation and copper catalysed click chemistry with a fluorescent reporter tag (TAMRA-azide, AzT– Figure S2A) led to dose-dependent labelling of multiple bands in the lysate, visualised by in-gel fluorescence scanning as shown in Figure 4A.
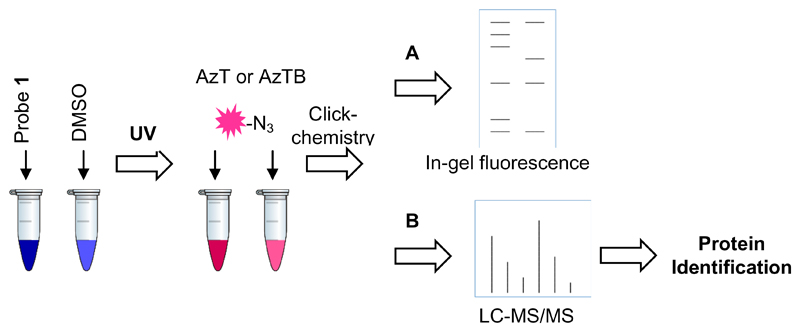
Figure 3.
Work-flow for labelling of Plasmodium falciparum lysate with probe 1. Lysate is incubated with the probe, irradiated with UV and then the capture reagent (AzT or AzTB) is added and conjugated to the bound probe by copper-catalysed click chemistry. After protein precipitation, proteins are analysed by either A) gel electrophoresis and visualisation of the fluorophore by in-gel fluorescence; or B) pull-down with neutravidin beads, reduction, alkylation, on-bead digestion and LC-MS/MS analysis for protein identification.
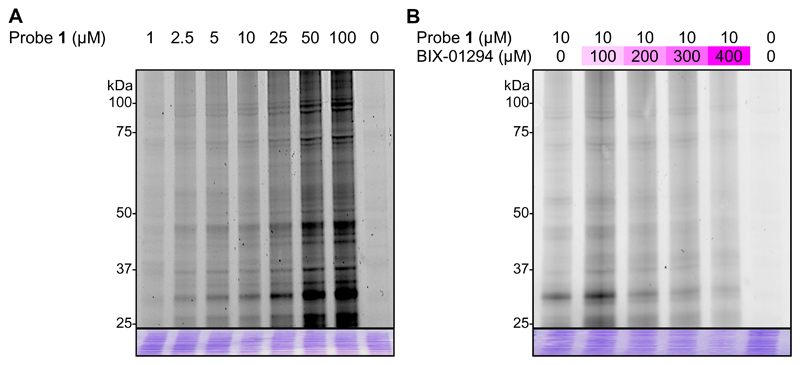
Figure 4.
(A) Dose-dependent labelling of Plasmodium falciparum lysate by probe 1. (B) Competition of probe 1 at 10 μM by increasing concentrations of BIX-01294 in Plasmodium falciparum lysates. Plasmodium falciparum lysate was incubated with BIX-01294, before the probe was added and the samples subjected to photo-crosslinking with UV (365 nm) and copper-catalysed click-chemistry with AzT. Fluorescence is shown in greyscale. All lanes contained 30 μg of total protein and equal loading was confirmed by coomassie brilliant blue staining (image in blue). Numbers in the left side indicate the molecular weights (in kDa) of proteins.
While this result demonstrates the effectiveness of our probe to label a variety of proteins within the lysate, the workflow employed was not able to distinguish between specific versus non-specific targets, given the irreversible nature of photo-crosslinking. To confirm which of the bands identified relate to specific targets, we used a representative competitor compound - BIX-01294 (Pf3D7 IC50 = 50 nM ± 12 nM) - at increasing concentrations (0-400 μM) in the labelling experiments. Due to the reversible nature of BIX-01294 binding compared with the irreversible nature of photo-crosslinking, high equivalents of the competitor were required. For specific BIX-01294 targets, a reduction in labelling would be expected. As depicted in Figure 4B there is a general attenuation in fluorescence with increasing concentrations of BIX-01294. In general terms, this suggests that probe 1 and BIX-01294 have similar proteome target profiles, as anticipated from the SAR, and suggests probe 1 to be a good tool for the identification of the targets of this compound series.
In order to identify the targets engaged by probe 1, the workflow was altered so that the labelled proteins could be enriched prior to their identification by LC-MS/MS. To that end, the AzT tag was replaced with an azide-TAMRA-biotin35,36 (AzTB, Figure S2B). This versatile reagent contains a TAMRA fluorophore for protein visualization, but also a biotin subunit to allow for purification of the probe targets by neutravidin ‘pull-down’. The lysate was incubated with either probe 1 (50 μM) or dimethyl sulfoxide (DMSO) as a control. For these experiments, no competitive inhibitor was used. Photo-crosslinking and conjugation to AzTB, was then followed by incubation with neutravidin beads and stringent washing. The captured proteins were subjected to on-bead trypsin digestion and the obtained peptides were analysed by nano LC-MS/MS on a high resolution Orbitrap mass spectrometer.37 Both probe and control samples were carried out in duplicate, and hits identified in less than two independent runs were not considered for further analysis.38
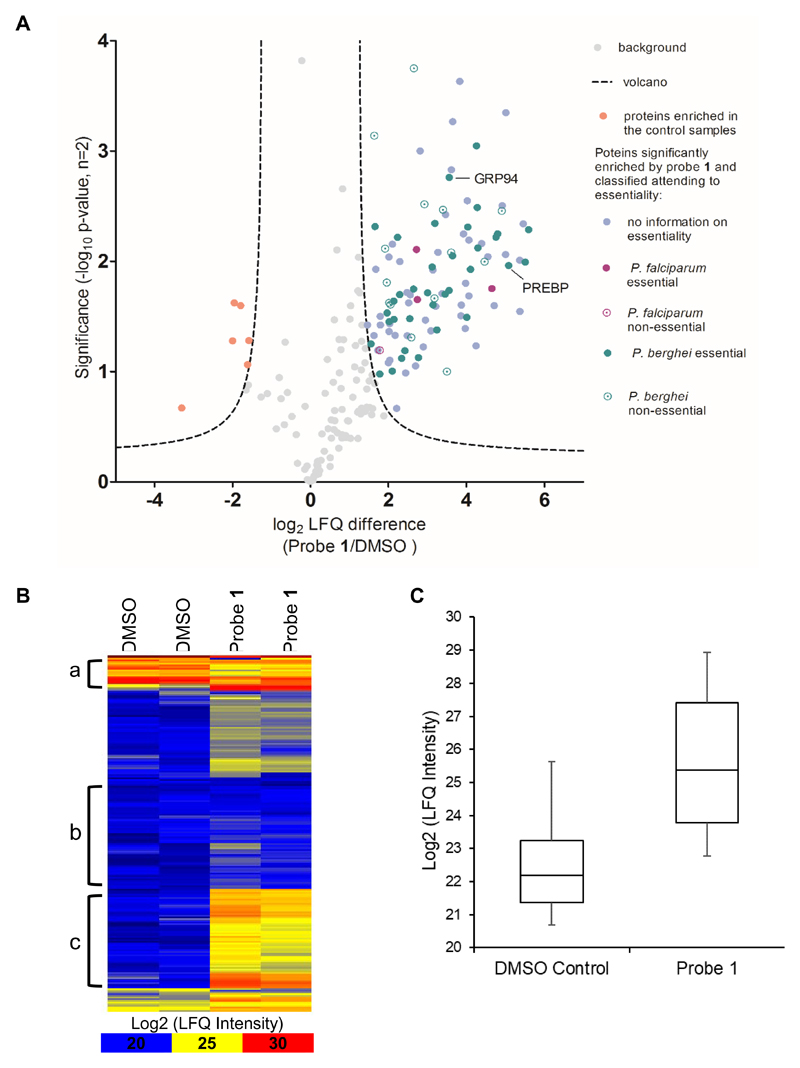
Following this criterion, 205 proteins were identified, of which 104 were found to be significantly enriched by probe 1 compared to the DMSO control samples (Figure 5A, Table S1). Figure 5B shows the heatmap of all the proteins identified with regard to the relative abundance of the detected peptides, color-coded from blue to red, calculated using label-free quantification (LFQ) and showing a good correlation between duplicates. For instance, proteins in groups a and b are not significantly different in the probe samples when compared to the controls regardless of their intensity, while section c shows higher intensity for the detected peptides when probe 1 was used. This was confirmed when the 104 significantly enriched proteins were analysed in terms of their relative abundance in probe and control samples separately (Figure 5C).
Figure 5.
(A) Volcano plot from analysis with Perseus 1.5.6.0 using t-test significance testing of duplicate results with FDR = 0.001 (False Discovery Rate) and S0 = 2. 104 proteins were enriched in the samples containing probe 1 (right-hand side: blue, purple and green) compared to the 6 significantly relevant proteins obtained in the DMSO controls (left-hand side: pink). (B) Heatmap for all proteins found in the two probe and the two control samples, in terms of their relative abundance log2(LFQ intensity) from Euclidian hierarchical row clustering with Perseus 1.5.6.0. High intensity is shown in red and low intensity in blue. As an example, group a) shows proteins heavily enriched in both probe 1 and DMSO control samples; group b) shows proteins enriched in neither sample; and group c) shows proteins heavily enriched in the samples with probe 1 compared to the control. (C) Box plots of the 104 proteins significantly enriched for probe 1. The plots show the spread of intensities of the identified proteins around the median, for the control samples (left) and the probe samples (right), with 80% of the data contained within the marked box. Intensities for probe 1 are significant compared to the background from the control.
In order to assess the importance of the significantly enriched proteins, we analysed the identified genes for essentiality, using the PhenoPlasm database (http://phenoplasm.org/).39 Out of the 104 enriched proteins there is only phenotypic information regarding disruption of four of them in P. falciparum (Figure 5A, purple datasets, Table S1). It is worth mentioning that three of those genes were found to be essential for parasite survival (filled circles in Figure 5A): PfnPrx is a peroxiredoxin with a widespread distribution throughout P.falciparum genome, which localizes to the nucleus where it protects the parasite’s DNA from oxidative damage;40 NAPL is one of the two existing nucleosome assembly proteins in P. falciparum, and is responsible for histone shuttling in the cytoplasm, playing therefore an essential role in chromatin structure organization;41 PfHSP110c is a cytoplasmic heat shock protein very important for parasite survival during febrile episodes, by avoiding the aggregation of Asn-rich proteins, widely present in the parasite genome, and prone to aggregate especially at elevated temperatures.42
Although the genes identified in our proteomic studies are from P. falciparum, the PhenoPlasm database provides information of all the studies available for a given gene ID across the Plasmodium spp.39 Given that a much more thorough functional profiling of P. berghei has been carried out compared to P. falciparum (2700 genes versus to 398, respectively), we analysed the enriched proteins for phentypic information in P.bergei. Thus, out of the 104 significantly enriched proteins for probe 1, phenotypic information for 49 additional genes was found in P.berghei (Figure 5A, green data sets, Table S1), and 35 of them were found to be essential (filled vs empty circles). Gene ontology (GO) analysis of these and the P.falciparum essential genes using the Protein ANalysis THrough Evolutionary Relationships (PANTHER) classification system, 43–45 showed that the identified proteins belong to different functional classes and display a variety of molecular functions (Figure S4). A closer look to the most significant and strongly shifted essential hits (filled circles in Figure 5A with significance and LFQ difference higher than 2.20 and 3.50, respectively) revealed a high abundance of protein synthesis-related proteins, including ribosomal proteins and elongation factors. These may arise from specific or non-specific pull-down of the ribosome, which is a common background protein in such studies due to its high abundance, especially in schizonts. Interestingly, a number of signficant and strongly shifted hits were identified that are involved in transcriptional regulation, such as the pre-binding protein (PREBP) or endoplasmin putative protein (GRP94) (see hits indicated in Figure 5A). PREBP is a transcription factor involved in activation of gene expression at all stages of the intraerythrocytic life cycle of the parasite,46 while GRP94 has been reported to be a non-histone target of the PHKMTs.47 The absence of PHKMTs from the target list generated using probe 1 is notable, and we speculate that the Plasmodium lysates used in this study may be inappropriate for identification of putative PHKMT targets of this series. The conditions used to generate lysate, limited stability, or low abundance of PHKMTs in the sample may all contribute, and studies are ongoing to investigate these potential limitations. Alternatively, the PHKMTs may prove not to be a major target of this diaminoquinazoline series.
In summary, we have reported the design and development of small-molecule photo-crosslinkable probe 1 to investigate the targets of our diaminoquinazoline series by photoaffinity labelling. Thus far our data identifies design rules to generate effective photoaffinity probes based on this scaffold and provides an initial proteomic dataset for target ID. A large number of the identified prospective targets represent essential proteins in P. falciparum or P. bergei and further work is needed to interrogate and validate the prospective targets that have emerged. This report lays the groundwork for such an endeavour, which is ongoing in our laboratories. The absence of the PfHKMTs from this target list may be due to limitations in the method and a number of additional techniques and approaches (such as target-based “pull down”) are being used to investigate this aspect further. Alternatively, the series may have alternative targets, further justifying the need for unbiased target ID studies in order to elucidate the primary targets that underpin their exciting antimalarial activities.
Materials and Methods
Lysate Labelling and In-Gel Fluorescence
For dose-dependent studies, saponin-treated pellets derived from Plasmodium falciparum cultures, 30-40 hours post invasion, were used to produce a lysate in PBS (100 μL, 3 mg mL-1, Supporting Information), which was then incubated with probe 1 (1-100 μM) at 0 °C for 1 hour. For competition experiments, lysates were first treated with the desired concentration of BIX 01294 (100-400 μM) at 0 °C for 30 min before adding probe 1 (10 μM), and incubated for an additional 1 h at the same temperature. For those samples to which no probe was added, DMSO was used as a vehicle. Stock solutions of the probe and BIX-01294 were prepared so that the same amount of DMSO was added in all samples, keeping it always below 2%. The lysates were then irradiated (365 nm, 0 °C, 20 min) and conjugated to AzT by treatment with 6 μL of a pre-mixed solution containing 1 μL of 10 mM AzT in DMSO (0.1 mM final concentration), 2 μL of 50 mM CuSO4 in water (1 mM final concentration), 2 μL of 50 mM tris(2-carboxyethyl)phosphine (TCEP) in water (1 mM final concentration) and 1 μL of 10 mM tris[(1-benzyl-1H-1,2,3-triazol-4-yl)methyl]amine (TBTA) in DMSO (0.1 mM final concentration). After 1 h shaking at room temperature, 1 μL of 500 mM ethylenediaminetetraacetic acid (EDTA) in water (5 mM final concentration) was added and the lysates vortexed. To precipitate the proteins, methanol (200 μL), chloroform (50 μL), and water (100 μL) were added, and the mixture vortexed and centrifuged (16,000 g, 10 min). The solvent was removed, and the pellets were washed with methanol (2 x 200 μL) and stored at -80 °C until analysis. The protein pellets were dissolved in 25 μL of 2% sodium dodecyl sulfate (SDS) and 10 mM dithiothreitol (DTT) in phosphate buffered saline (PBS), and the solution was diluted with PBS (75 μL) and added to 25 μL of 4x loading buffer (LDS) and 10 mM DTT. Samples were boiled at 95 °C for 10 min and 30 μg were loaded into each gel lane and resolved using SDS-polyacrylamide gel electrophoresis (SDS-PAGE). Images were acquired using an Amersham Typhoon FLA 7000 fluorescence scanner. Fluorescent images are shown in grey-scale.
Lysate Labelling and Pull Down for Proteomics
Plasmodium falciparum lysates (200 μL, 3 mg mL-1) were incubated with probe 1 (50 μM) or DMSO for control experiments at 0 °C for 1 h, irradiated (365 nm, 0 °C, 20 min) and conjugated to the trifunctional capture reagent AzTB as described above. After 1 h shaking at room temperature, 2 μL of 500 mM EDTA in water (5 mM final concentration) was added and the lysates vortexed. Proteins were precipitated by adding methanol (400 μL), chloroform (100 μL) and water (200 μL). The mixture was vortexed and centrifuged (16,000 g, 10 min). The solvent was removed, and the pellets washed with methanol (2 x 400 μL). Pellets were stored at -80 °C overnight and then dissolved in 50 μL of 2% SDS and 10 mM DTT in PBS, and the solution was diluted with PBS (450 μL). Neutravidin agarose beads (50 μL) were added to the samples after being washed with 0.2% SDS in PBS (3 x 200 μL), and the samples were then shaken at room temperature for 90 min. The beads were pelleted (2000 g, 2 min) and the supernatant removed. The beads were washed sequentially in 1% SDS in PBS (3 x 400 μL), 4M Urea in PBS (2 x 400 μL), 50 mM ammonium bicarbonate (4 x 400 μL). For each wash step the beads were gently vortexed for 1 min followed by pelleting in a microcentrifuge (2000 g, 2 min). The beads were re-suspended with 50 mM ammonium bicarbonate (200 μL) and 10 μL of 100 mM DTT were added (5 mM final concentration). The samples were incubated at 55 °C for 30 min with shaking. After centrifugation (2000 g, 2 min) and removal of the supernatant, the beads were washed with 50 mM ammonium bicarbonate (200 μL, 2000 g, 2 min) and then re-suspended with 200 μL of 50 mM ammonium bicarbonate. 15 μL of 100 mM iodoacetamide (7.5 mM final concentration) were added, and the samples were incubated at room temperature for 30 min in the dark. After centrifugation (2000 g, 2 min) and removal of the supernatant, the beads were washed with 50 mM ammonium bicarbonate (200 μL, 2000 g, 2 min), re-suspended with 100 μL of 50 mM ammonium bicarbonate, and 5 μL of trypsin in 50 mM ammonium bicarbonate (20 μg trypsin in 100 μL of 50 mM ammonium bicarbonate) were added. The samples were incubated at 37 °C overnight with shaking. The beads were pelleted by centrifugation (2000 g, 2 min) and the supernatant collected. Beads were washed with 100 μL of 50 mM ammonium bicarbonate for 10 min, pelleted (2000 g, 2 min), and the supernatant collected. Next, beads were washed with 100 μL of 1.5% trifluoroacetic acid (TFA) for 10 min, precipitated (2000 g, 2 min), and the supernatant collected.
Stage-tip purification of peptides
For LC-MS/MS analysis the supernatants containing the peptides were combined and stage-tipped.48 P200 pipette tips were fitted with 3 layers of SDB-XC extraction disks (Empore®) cut out using an in-house constructed tool. The pipettes were inserted into the hole of a microcentifuge tube lid. The tips were activated by addition of methanol (150 μL) and the tips centrifuged (2000 g, 2 min). The tip was washed with LC-MS/MS grade water (150 μL). The peptide solution was loaded into the tip and the tip centrifuged again. The water wash was repeated. The peptides were eluted into a clean microcentrifuge tube by addition of 79% acetonitrile in water (60 μL). The peptides were dried in a Savant SPD1010 SpeedVac® Concentrator (Thermo Scientific) and stored at -80°C until LC-MS/MS analysis. Peptides were reconstituted in 25 μL of 2% acetonitrile in water with 0.5% trifluoroacetic acid for LC-MS/MS analysis (see Supporting Information).
Supplementary Material
Synthetic procedures for the probe, cell culture procedures and LC-MS/MS proteomic methods can be found in the Supporting Information, along with figures of the capture reagents, scatter plots for the proteomic data and protein class analysis. Proteomics pull-down data can be found in Supporting Information Table S1.
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE49 partner repository with the dataset identifier PXD008164
Acknowledgements
AL acknowledges the Faculty of Natural Sciences at Imperial College London for a Schrodinger Scholarship. AR-Z acknowledges Alfonso Martín Escudero Foundation for a postdoctoral fellowship. KSH, JMS and EWT acknowledge support from Medicines for Malaria Venture and JB is supported by the EPSRC Global Challenge Research Fund (EP/P510798/1) and the Wellcome Trust via an Investigator Award (100993/Z/13/Z).
References
- [1].WHO. World Malaria Report 2016. 2016 http://www.who.int/malaria/publications/world-malaria-report-2016/report/en/
- [2].Fairhurst RM, Dondorp AM. Artemisinin-Resistant Plasmodium falciparum Malaria. Microbiol Spectr. 2016;4:409–429. doi: 10.1128/microbiolspec.EI10-0013-2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [3].Ashley EA, Dhorda M, Fairhurst RM, Amaratunga C, Lim P, Suon S, Sreng S, Anderson JM, Mao S, Sam B, Sopha C, et al. Spread of Artemisinin Resistance in Plasmodium falciparum Malaria. N Engl J Med. 2014;371:411–423. doi: 10.1056/NEJMoa1314981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [4].Lu F, Culleton R, Zhang M, Ramaprasad A, von Seidlein L, Zhou H, Zhu G, Tang J, Liu Y, Wang W, Cao Y, et al. Emergence of Indigenous Artemisinin-Resistant Plasmodium falciparum in Africa. N Engl J Med. 2017;376:991–993. doi: 10.1056/NEJMc1612765. [DOI] [PubMed] [Google Scholar]
- [5].Gellis A, Primas N, Hutter S, Lanzada G, Remusat V, Verhaeghe P, Vanelle P, Azas N. Looking for new antiplasmodial quinazolines: DMAP-catalyzed synthesis of 4-benzyloxy- and 4-aryloxy-2-trichloromethylquinazolines and their in vitro evaluation toward Plasmodium falciparum. Eur J Med Chem. 2016;119:34–44. doi: 10.1016/j.ejmech.2016.04.059. [DOI] [PubMed] [Google Scholar]
- [6].Verhaeghe P, Azas N, Gasquet M, Hutter S, Ducros C, Laget M, Rault S, Rathelot P, Vanelle P. Synthesis and antiplasmodial activity of new 4-aryl-2-trichloromethylquinazolines. Bioorg Med Chem Lett. 2008;18:396–401. doi: 10.1016/j.bmcl.2007.10.027. [DOI] [PubMed] [Google Scholar]
- [7].Warhurst DC. Antimalarial drugs II: Current antimalarials and new drug development. Trends in Pharmacological Sciences. 6:302–304. doi: 10.1016/0165-6147(85)90140-3. [DOI] [Google Scholar]
- [8].Madapa S, Tusi Z, Mishra A, Srivastava K, Pandey SK, Tripathi R, Puri SK, Batra S. Search for new pharmacophores for antimalarial activity. Part II: synthesis and antimalarial activity of new 6-ureido-4-anilinoquinazolines. Bioorg Med Chem. 2009;17:222–234. doi: 10.1016/j.bmc.2008.11.005. [DOI] [PubMed] [Google Scholar]
- [9].Gamo F-J, Sanz LM, Vidal J, de Cozar C, Alvarez E, Lavandera J-L, Vanderwall DE, Green DVS, Kumar V, Hasan S, Brown JR, et al. Thousands of chemical starting points for antimalarial lead identification. Nature. 2010;465:305–310. doi: 10.1038/nature09107. [DOI] [PubMed] [Google Scholar]
- [10].Gilson PR, Tan C, Jarman KE, Lowes KN, Curtis JM, Nguyen W, Di Rago AE, Bullen HE, Prinz B, Duffy S, Baell JB, et al. Optimization of 2-Anilino 4-Amino Substituted Quinazolines into Potent Antimalarial Agents with Oral in Vivo Activity. J Med Chem. 2017;60:1171–1188. doi: 10.1021/acs.jmedchem.6b01673. [DOI] [PubMed] [Google Scholar]
- [11].González Cabrera D, Le Manach C, Douelle F, Younis Y, Feng T-S, Paquet T, Nchinda AT, Street LJ, Taylor D, de Kock C, Wiesner L, et al. 2,4-Diaminothienopyrimidines as Orally Active Antimalarial Agents. J Med Chem. 2014;57:1014–1022. doi: 10.1021/jm401760c. [DOI] [PubMed] [Google Scholar]
- [12].Hansch C, Fukunaga JY, Jow PYC, Hynes JB. Quantitative structure-activity relation of antimalarial and dihydrofolate reductase inhibition by quinazolines and 5-substituted benzyl-2,4-diaminopyrimidines. J Med Chem. 1977;20:96–102. doi: 10.1021/jm00211a020. [DOI] [PubMed] [Google Scholar]
- [13].Plouffe D, Brinker A, McNamara C, Henson K, Kato N, Kuhen K, Nagle A, Adrian F, Matzen JT, Anderson P, Nam TG, et al. In silico activity profiling reveals the mechanism of action of antimalarials discovered in a high-throughput screen. Proc Natl Acad Sci USA. 2008;105:9059–9064. doi: 10.1073/pnas.0802982105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [14].Malmquist NA, Moss TA, Mecheri S, Scherf A, Fuchter MJ. Small-molecule histone methyltransferase inhibitors display rapid antimalarial activity against all blood stage forms in Plasmodium falciparum. Proc Natl Acad Sci USA. 2012;109:16708–16713. doi: 10.1073/pnas.1205414109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [15].Sundriyal S, Malmquist NA, Caron J, Blundell S, Liu F, Chen X, Srimongkolpithak N, Jin J, Charman SA, Scherf A, Fuchter MJ. Development of diaminoquinazoline histone lysine methyltransferase inhibitors as potent blood-stage antimalarial compounds. ChemMedChem. 2014;9:2360–2373. doi: 10.1002/cmdc.201402098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [16].Malmquist NA, Sundriyal S, Caron J, Chen P, Witkowski B, Menard D, Suwanarusk R, Renia L, Nosten F, Jiménez-Díaz MB, Angulo-Barturen I, et al. Histone Methyltransferase Inhibitors Are Orally Bioavailable, Fast-Acting Molecules with Activity against Different Species Causing Malaria in Humans. Antimicrob Agents Chemother. 2015;59:950–959. doi: 10.1128/AAC.04419-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [17].Sundriyal S, Chen PB, Lubin AS, Lueg GA, Li F, White AJP, Malmquist NA, Vedadi M, Scherf A, Fuchter MJ. Histone lysine methyltransferase structure activity relationships that allow for segregation of G9a inhibition and anti-Plasmodium activity. MedChemComm. 2017;8:1069–1092. doi: 10.1039/C7MD00052A. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [18].Kubicek S, O'Sullivan RJ, August EM, Hickey ER, Zhang Q, Teodoro Miguel L, Rea S, Mechtler K, Kowalski JA, Homon CA, Kelly TA, et al. Reversal of H3K9me2 by a Small-Molecule Inhibitor for the G9a Histone Methyltransferase. Mol Cell. 2007;25:473–481. doi: 10.1016/j.molcel.2007.01.017. [DOI] [PubMed] [Google Scholar]
- [19].Vedadi M, Barsyte-Lovejoy D, Liu F, Rival-Gervier S, Allali-Hassani A, Labrie V, Wigle TJ, Dimaggio PA, Wasney GA, Siarheyeva A, Dong A, et al. A chemical probe selectively inhibits G9a and GLP methyltransferase activity in cells. Nat Chem Biol. 2011;7:566–574. doi: 10.1038/nchembio.599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [20].Liu F, Barsyte-Lovejoy D, Li F, Xiong Y, Korboukh V, Huang X-P, Allali-Hassani A, Janzen WP, Roth BL, Frye SV, Arrowsmith CH, et al. Discovery of an in Vivo Chemical Probe of the Lysine Methyltransferases G9a and GLP. J Med Chem. 2013;56:8931–8942. doi: 10.1021/jm401480r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [21].Xiong Y, Li F, Babault N, Wu H, Dong A, Zeng H, Chen X, Arrowsmith CH, Brown PJ, Liu J, Vedadi M, et al. Structure-activity relationship studies of G9a-like protein (GLP) inhibitors. Bioorg Med Chem. 2017;25:4414–4423. doi: 10.1016/j.bmc.2017.06.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [22].Ma A, Yu W, Xiong Y, Butler KV, Brown PJ, Jin J. Structure-activity relationship studies of SETD8 inhibitors. MedChemComm. 2014;5:1892–1898. doi: 10.1039/C4MD00317A. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [23].Curry E, Green I, Chapman-Rothe N, Shamsaei E, Kandil S, Cherblanc FL, Payne L, Bell E, Ganesh T, Srimongkolpithak N, Caron J, et al. Dual EZH2 and EHMT2 histone methyltransferase inhibition increases biological efficacy in breast cancer cells. Clin Epigenet. 2015;7:1–12. doi: 10.1186/s13148-015-0118-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [24].Dembele L, Franetich JF, Lorthiois A, Gego A, Zeeman AM, Kocken CH, Le Grand R, Dereuddre-Bosquet N, van Gemert GJ, Sauerwein R, Vaillant JC, et al. Persistence and activation of malaria hypnozoites in long-term primary hepatocyte cultures. Nat Med. 2014;20:307–312. doi: 10.1038/nm.3461. [DOI] [PubMed] [Google Scholar]
- [25].Chen PB, Ding S, Zanghi G, Soulard V, DiMaggio PA, Fuchter MJ, Mecheri S, Mazier D, Scherf A, Malmquist NA. Plasmodium falciparum PfSET7: enzymatic characterization and cellular localization of a novel protein methyltransferase in sporozoite, liver and erythrocytic stage parasites. Sci Rep. 2016;6 doi: 10.1038/srep21802. 21802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [26].Heal WP, Dang THT, Tate EW. Activity-based probes: discovering new biology and new drug targets. Chem Soc Rev. 2011;40:246–257. doi: 10.1039/C0CS00004C. [DOI] [PubMed] [Google Scholar]
- [27].Wright MH, Sieber SA. Chemical proteomics approaches for identifying the cellular targets of natural products. Nat Prod Rep. 2016;33:681–708. doi: 10.1039/C6NP00001K. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [28].Li H, van der Linden WA, Verdoes M, Florea BI, McAllister FE, Govindaswamy K, Elias JE, Bhanot P, Overkleeft HS, Bogyo M. Assessing subunit dependency of the Plasmodium proteasome using small molecule inhibitors and active site probes. ACS Chem Biol. 2014;9:1869–1876. doi: 10.1021/cb5001263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [29].Ismail HM, Barton V, Phanchana M, Charoensutthivarakul S, Wong MH, Hemingway J, Biagini GA, O'Neill PM, Ward SA. Artemisinin activity-based probes identify multiple molecular targets within the asexual stage of the malaria parasites Plasmodium falciparum 3D7. Proc Natl Acad Sci USA. 2016;113:2080–2085. doi: 10.1073/pnas.1600459113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [30].Smith E, Collins I. Photoaffinity labeling in target- and binding-site identification. Future Med Chem. 2015;7:159–183. doi: 10.4155/fmc.14.152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [31].Brunner J. New photolabeling and crosslinking methods. Annu Rev Biochem. 1993;62:483–514. doi: 10.1146/annurev.bi.62.070193.002411. [DOI] [PubMed] [Google Scholar]
- [32].MacKinnon AL, Taunton J. Target Identification by Diazirine Photo-Cross-linking and Click Chemistry. Curr Protoc Chem Bio. 2009;1:55–73. doi: 10.1002/9780470559277.ch090167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [33].Heal WP, Wright MH, Thinon E, Tate EW. Multifunctional protein labeling via enzymatic N-terminal tagging and elaboration by click chemistry. Nat Protocols. 2012;7:105–117. doi: 10.1038/nprot.2011.425. [DOI] [PubMed] [Google Scholar]
- [34].Wright MH, Clough B, Rackham MD, Rangachari K, Brannigan JA, Grainger M, Moss DK, Bottrill AR, Heal WP, Broncel M, Serwa RA, et al. Validation of N-myristoyltransferase as an antimalarial drug target using an integrated chemical biology approach. Nat Chem. 2014;6:112. doi: 10.1038/nchem.1830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [35].Thinon E, Serwa RA, Broncel M, Brannigan JA, Brassat U, Wright MH, Heal WP, Wilkinson AJ, Mann DJ, Tate EW. Global profiling of co- and post-translationally N-myristoylated proteomes in human cells. Nat Commun. 2014;5 doi: 10.1038/ncomms5919. 4919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [36].Wright MH, Clough B, Rackham MD, Rangachari K, Brannigan JA, Grainger M, Moss DK, Bottrill AR, Heal WP, Broncel M, Serwa RA, et al. Validation of N-myristoyltransferase as an antimalarial drug target using an integrated chemical biology approach. Nature chemistry. 2014;6:112–121. doi: 10.1038/nchem.1830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [37].Broncel M, Serwa RA, Ciepla P, Krause E, Dallman MJ, Magee AI, Tate EW. Multifunctional reagents for quantitative proteome-wide analysis of protein modification in human cells and dynamic profiling of protein lipidation during vertebrate development. Angew Chem Int Ed Engl. 2015;54:5948–5951. doi: 10.1002/anie.201500342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [38].Tyanova S, Temu T, Sinitcyn P, Carlson A, Hein MY, Geiger T, Mann M, Cox J. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat Methods. 2016;13:731–740. doi: 10.1038/nmeth.3901. [DOI] [PubMed] [Google Scholar]
- [39].Sanderson T, Rayner JC. PhenoPlasm: a database of disruption phenotypes for malaria parasite genes. Wellcome Open Res. 2017;2:45. doi: 10.12688/wellcomeopenres.11896.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [40].Richard D, Bartfai R, Volz J, Ralph SA, Muller S, Stunnenberg HG, Cowman AF. A genome-wide chromatin-associated nuclear peroxiredoxin from the malaria parasite Plasmodium falciparum. J Biol Chem. 2011;286:11746–11755. doi: 10.1074/jbc.M110.198499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [41].Gill J, Yogavel M, Kumar A, Belrhali H, Jain SK, Rug M, Brown M, Maier AG, Sharma A. Crystal structure of malaria parasite nucleosome assembly protein: distinct modes of protein localization and histone recognition. J Biol Chem. 2009;284:10076–10087. doi: 10.1074/jbc.M808633200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [42].Muralidharan V, Oksman A, Pal P, Lindquist S, Goldberg DE. Plasmodium falciparum heat shock protein 110 stabilizes the asparagine repeat-rich parasite proteome during malarial fevers. Nat Commun. 2012;3 doi: 10.1038/ncomms2306. 1310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [43].Mi H, Huang X, Muruganujan A, Tang H, Mills C, Kang D, Thomas PD. PANTHER version 11: expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids Res. 2017;45:D183–D189. doi: 10.1093/nar/gkw1138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [44].Mi H, Muruganujan A, Casagrande JT, Thomas PD. Large-scale gene function analysis with the PANTHER classification system. Nat Protocols. 2013;8:1551–1566. doi: 10.1038/nprot.2013.092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [45].Mi H, Muruganujan A, Thomas PD. PANTHER in 2013: modeling the evolution of gene function, and other gene attributes, in the context of phylogenetic trees. Nucleic Acids Res. 2013;41:D377–386. doi: 10.1093/nar/gks1118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [46].Komaki-Yasuda K, Okuwaki M, Nagata K, Kawazu S, Kano S. Identification of a novel and unique transcription factor in the intraerythrocytic stage of Plasmodium falciparum. PLoS One. 2013;8:e74701. doi: 10.1371/journal.pone.0074701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [47].Kaur I, Zeeshan M, Saini E, Kaushik A, Mohmmed A, Gupta D, Malhotra P. Widespread occurrence of lysine methylation in Plasmodium falciparum proteins at asexual blood stages. Sci Rep. 2016;6 doi: 10.1038/srep35432. 35432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [48].Rappsilber J, Ishihama Y, Mann M. Stop and Go Extraction Tips for Matrix-Assisted Laser Desorption/Ionization, Nanoelectrospray, and LC/MS Sample Pretreatment in Proteomics. Anal Chem. 2003;75:663–670. doi: 10.1021/ac026117i. [DOI] [PubMed] [Google Scholar]
- [49].Vizcaino JA, Csordas A, del-Toro N, Dianes JA, Griss J, Lavidas I, Mayer G, Perez-Riverol Y, Reisinger F, Ternent T, Xu QW, et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 2016;44:D447–456. doi: 10.1093/nar/gkv1145. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.