Abstract
Natural products are an important source of new drugs for the treatment of various diseases. However, developing natural product-based new medicines through random moiety modification is a lengthy and costly process, due in part to the difficulties associated with comprehensively understanding the mechanism of action and the side effects. Identifying the protein targets of natural products is an effective strategy, but most medicines interact with multiple protein targets, which complicate this process. In recent years, an increasing number of researchers have begun to screen the target proteins of natural products with chemical proteomics approaches, which can provide a more comprehensive array of the protein targets of active small molecules in an unbiased manner. Typically, chemical proteomics experiments for target identification consist of two key steps: (1) chemical probe design and synthesis and (2) target fishing and identification. In recent decades, five different types of chemical proteomic probes and their respective target fishing methods have been developed to screen targets of molecules with different structures, and a variety of protein identification approaches have been invented. Presently, we will classify these chemical proteomics approaches, the application scopes and characteristics of the different types of chemical probes, the different protein identification methods, and the advantages and disadvantages of these strategies.
Subject terms: Target identification, Target identification
Introduction
Over the last 30 years, natural products have become an important source of new drugs to target various diseases.1,2 In contrast to chemically synthesized drugs, drugs derived from natural products possess remarkable advantages in terms of structural novelty, biocompatibility and functional diversity, stemming from long-term natural selection-based optimizations in their evolution.3 Statistically, among marked drugs approved by the Food and Drug Administration (FDA) from 1939 to 2016, more than 50% are derived from natural products,4 and these compounds are commonly known as natural medicines. For example, elliptinium, a naturally occurring plant alkaloid, has been developed into the anticancer natural medicine Celiptium. It is widely used in multiple cancer therapies, such as breast cancer and renal cell carcinoma.5,6 Retapamulin, another natural medicine derived from pleuromutilin produced by Pleurotus mutilus, an edible mushroom, is the first in a new class of antibacterial drugs known as pleuromutilins to be approved for use in humans.7 In addition to the natural products that have been developed as commercial medications for humans, a number of them are on their way to be patented medicine, such as resveratrol,8–10 curcumin,11–13 oridonin,14,15 etc.16–18 However, developing a natural product-based new medicine from random moiety modification is a lengthy and costly process, due in part to the difficulties associated with comprehensively understanding their mechanism of action (MOA) as well as side effects.19,20
Interactions with intracellular protein targets are the foundation through which natural products exert their pharmacological activity. Therefore, target identification is the initial key step for the discovery and development of new natural medicines,21,22 as this allows the determination of the MOA and side effects. However, further studies on drug and target interaction mechanisms showed that most drugs interact with multiple protein targets rather than a single target.23–25 This multitargeted interaction mode makes identifying the true targets of the natural products being investigated substantially more difficult. Therefore, a target identification method that can comprehensively reveal multiple targets of natural products is urgently needed. Several systematic and nonbiased methods for identifying the targets of natural products, such as transcriptome-wide compound signature profiling, chemical genomics approaches and yeast two-hybrid methods, have been developed in recent decades.26–29 However, these strategies have disadvantages such as narrow applicability and multiple interference.30 With the advancement of molecular biology and the advent of the postgenomic era, an emerging and broadly applicable approach termed chemical proteomics was developed for target identification at the proteomic level.31,32
As an important branch of proteomics, chemical proteomics integrates diverse approaches in synthetic chemistry, cellular biology and mass spectrometry.33 It is an approach to comprehensively fish and identify multiple protein targets of active small molecules, and it consists of two key steps: (1) probe design and synthesis and (2) target fishing and protein identification. In recent decades, five different types of probes and their respective target fishing methods with different scopes and characteristics for chemical proteomics approaches have been developed34 to screen targets for small molecules with different structures. After target fishing, there are also multiple protein identification methods that are suitable for different situations.35–37 Numerous pharmacological studies have applied chemical proteomics to identify drug targets and study their MOA,38–40 especially in the last few years.41 Hence, these studies provides us with a unique background to summarize the recent achievements in this field.
In the present review, we first briefly introduce the chemical proteomics approaches, including their classification and workflow, as well as their advantages and disadvantages. Second, as the initial step of target identification, we provide a glimpse of the synthetic processes of five different types of probes and describe in detail the probes’ application scope and characteristics, as well as their subsequent target enrichment schemes. In the third section of the review, different protein identification methods with distinct scopes, including gel separation and band identification, quantitative proteomics approach, and protein microarray, are described. In the last section, we provide some comments on the future direction of chemical proteomics for the target identification of natural products.
Chemical proteomics in target identification
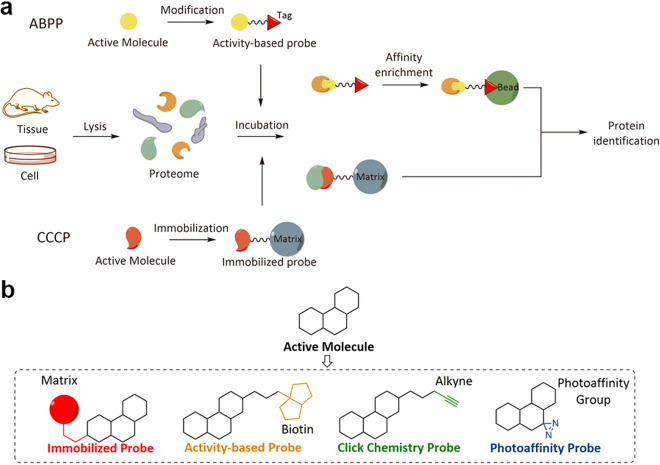
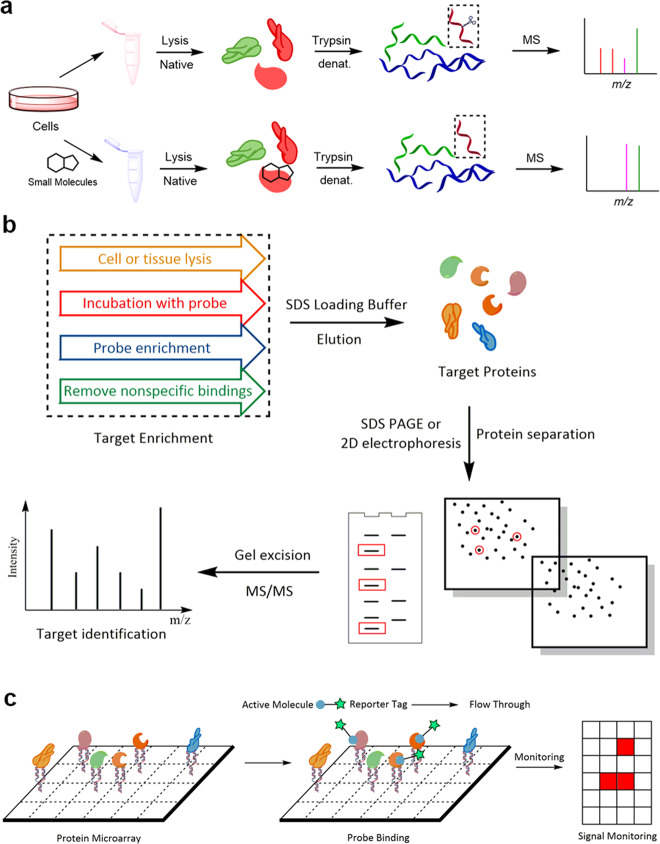
Chemical proteomics is a postgenomic version of classical drug affinity chromatography that is coupled to subsequent high-resolution MS and bioinformatic analyses.42 As illustrated in Fig. 1a, chemical proteomics approaches can be divided into two categories according to their different workflows, namely, activity-based protein profiling (ABPP) and compound-centric chemical proteomics (CCCP).23
Fig. 1.
a Comparison of activity-based probe profiling and compound-centric chemical proteomics. b General molecular structures of different types of chemical proteomics probes
ABPP is a technology that combines activity-based probe and proteomics technologies to identify protein targets of small bioactive molecules to help elucidate their MOA and side effects.31,43 In a typical ABPP experiment44 (Fig. 1a), probes derived from the parent molecules are first designed and synthesized based on a structure–activity relationship (SAR) study of the parent molecules. The probes should be synthesized as follows: (i) the probes should retain the pharmacological activity of their parent molecules to ensure the accuracy of subsequent target identification; (ii) the probes should allow for easy enrichment of bound protein targets. Next, the probes are incubated with effector living cells, lysate or tissue homogenates, allowing them to completely bind their target proteins. After enrichment with chemical and biochemical techniques, the protein targets are identified through proteomics approaches. The final step is to validate the target information via SPR, MST, ITC, etc. and the corresponding pharmacological effects with the appropriate biological function assays.
CCCP originates from classic drug affinity chromatography, which has been in use for decades.45–48 Along with the development of proteomics techniques, CCCP merges the classic method with modern proteomics to identify protein targets of small bioactive molecules at the proteome level. Unlike ABPP (Fig. 1a), the first step of CCCP is to immobilize the drug molecule on a matrix,23 such as magnetic or agarose beads. The probe synthesis and immobilization processes will be described in detail in the next section. Similar to ABPP, immobilization should not influence the pharmacological activity of the drug of interest. Subsequently, lysates from cells or tissues are incubated with the affinity matrix, followed by extensive washing to remove nonspecific binders. After complete elution, the enriched proteins are identified with proteomics approaches, and the target information and corresponding pharmacological effects must also be confirmed.
As mentioned above, the chemical proteomics approach may possess many advantages, such as being unbiased and allowing high-throughput at the proteome level, but it also has limitations. With chemical proteomics, the mass of nonspecifically bound proteins and the active metabolites, in addition to the true target proteins, may also be identified, leading to potential false-positive results. Furthermore, proteins that are insoluble in the buffers (e.g., PBS, Tris-HCl) used during the target enrichment process may pass unnoticed through the matrix. Comparing the two chemical proteomics approaches, in contrast to ABPP, the activation state of the identified proteins cannot be detected with CCCP, but CCCP is a more unbiased approach, allowing it to even identify targets with no enzymatic function, thereby facilitating the discovery of novel targets.
Probe design and synthesis
Designing and synthesizing the probe is the initial and pivotal step for target identification in chemical proteomics approaches. Generally, a probe consists of three parts, which are responsible for its respective functions: (i) a reactive group, which is derived from the parent drug molecule and ensures that it retains its pharmacological activity and ability to bind or modify protein targets; (ii) a reporter tag, such as biotin, an alkyne or a fluorescence group, for target enrichment or detection; (iii) a linker, sometimes cleavable, to connect the reactive group and the reporter tag, and it should be long enough to avoid steric hindrance.49,50 However, the structure of the probe may not always be constant. For example, in different chemical proteomics strategies for target identification, the probe might have one or even two of the components omitted. In this section, we will describe the diverse types of probes applied in chemical proteomics target profiling, as well as their design and synthesis, characteristics and scope of application.
Immobilized probe
In earlier studies, bioactive natural products were covalently immobilized on biocompatible inert resins, such as agarose and magnetic beads, to serve as bait for fish for target proteins in the active proteome (Fig. 1b). Due to the intrinsic properties of the beads, such as their macroscopic size and magnetism, the probe-fished proteins can be easily enriched, which is convenient for subsequent target identification. In the structure of bioactive natural products, various groups, such as sulfhydryl, amino and carboxyl groups, can be utilized for attachment to different active resins, which are commercially available.
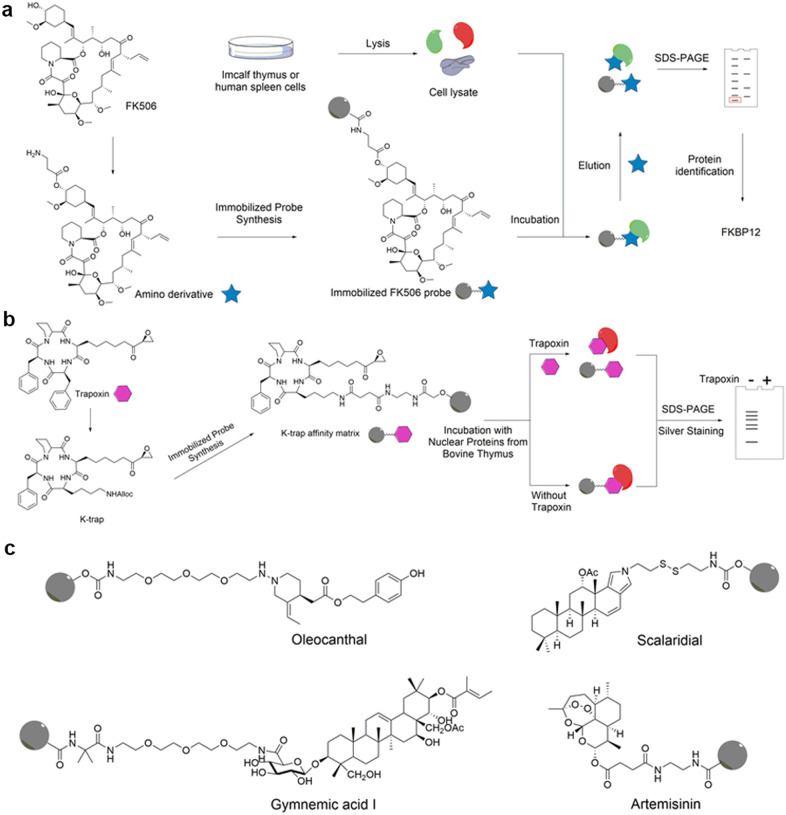
For example, Schreiber et al.46 immobilized FK506 (tacrolimus), a natural immunosuppressant, to identify its protein targets in lysates obtained from calf thymus and human spleen cells. As shown in Fig. 2a, FK506 affinity matrices were prepared using an FK506 amino derivative. After complete incubation with cytosolic extracts of bovine thymus and human spleen, the matrix was competitively eluted with FK506, and a 14 K protein was enriched and identified. This led to the identification of a FK506-binding protein of 12 K (FKBP12), which functions as a protein folding chaperone for proteins containing proline residues. Another example is trapoxin, a microbially derived cyclotetrapeptide that inhibits histone deacetylation in vivo and causes mammalian cells to undergo cell cycle arrest.51 Because the epoxyketone side chain of trapoxin is indispensable for activity, Schreiber et al. chose to replace one of the phenylalanine residues of trapoxin’s cyclic core with a lysine that could then be covalently linked to a solid support. The matrix was incubated with nuclear proteins from bovine thymus, and the bound polypeptides were eluted by boiling the matrix in 1% SDS buffer. Six major polypeptides with apparent molecular sizes between 45 and 50 kDa were detected by SDS-PAGE and silver staining. In addition, the authors also employed trapoxin to competitively inhibit the binding between the polypeptides and the matrix to validate the specificity of the results (Fig. 2b). Other examples are illustrated in Fig. 2c.52–55
Fig. 2.
Workflow of target identification with immobilized probes of natural medicines. a Identification of FK506 protein targets with an immobilized FK506 probe. b Identification of trapoxin protein targets with K-trap. c Structures of other reported immobilized probes of natural medicines
Although immobilized probes are easily synthesized and widely employed, one of their limitations, immobilization-induced steric hindrance, remains unsolved. The immobilization of the probes, which are always employed in CCCP, might influence the binding between the true targets and the reactive group, potentially leading to false-positive protein targets or the loss of potentially important protein targets. This could prove costly if an unsuitable protein target was selected for further evaluation.
Activity-based probe
To overcome immobilization-induced activity impairment, activity-based probes (ABPs) were developed for target identification in chemical proteomics. In the design of such probes, the first factor to consider is the activity of the drug molecule;31 in other words, the incorporation of the reporter group and the linker should not influence the bioactivity of the active molecule. Therefore, the SAR of the molecule should be studied or consulted before the start of the synthesis, and the probe’s pharmacological activity should be determined. Unlike immobilized probes, ABPs can interact with proteins in the active proteome before enrichment and even pass through the cell membrane to bind target proteins in living cells, potentially reflecting the true drug–target interactions under physiological conditions in cells.
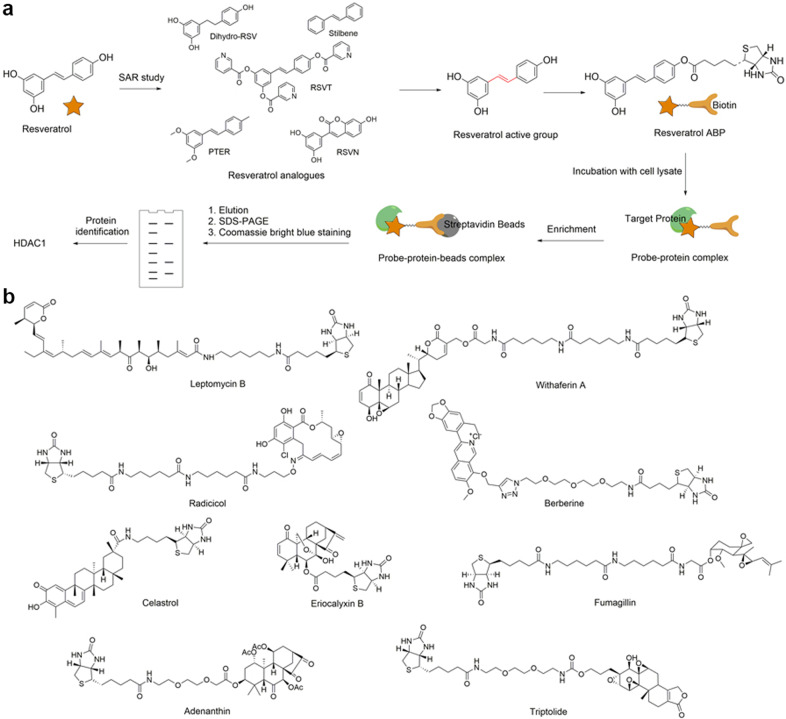
However, non-immobilization raises an obvious question: How does one enrich or detect probes and target proteins is the probe is not immobilized? In the structure of ABP, a reporter group is present to overcome this problem (Fig. 1b). Among the diverse reporter groups, biotin is most frequently utilized due to its strong affinity with avidin, allowing its enrichment using either avidin or streptavidin beads. For instance, to elucidate the mechanism of resveratrol, a natural product exhibiting anticancer activity in mouse melanoma cells, our group synthesized a probe by connecting a biotin tag to resveratrol based on an SAR study and validated the probe activity with in vitro biochemical experiments (Fig. 3a). Subsequently, the probe was incubated with lysates of melanoma cells and then enriched with streptavidin beads. As a result, we identified histone deacetylase I (HDAC1) as the protein target of resveratrol in mouse melanoma cells and revealed an epigenetic regulation pathway of focal adhesion kinase.41 Other excellent examples are listed in Fig. 3b.56–64
Fig. 3.
Schematic of target identification with activity-based probes of natural medicines. a Target identification of resveratrol with its activity-based probe. b Structures of some previously reported activity-based probes of natural medicines
In addition to the biotin tag, fluorescent tags are also widely used as the reporter group for target identification. Fluorescent-modified probes allow. the efficient and rapid detection of target proteins, but it cannot be enriched like biotin tags.65 However, in some cases, due to the large size of biotin, biotin can interfere with the original activity of the bioactive drug molecule. In addition, endogenous biotinylated proteins in the active proteome can interfere with identification by generating false-positive protein targets.66,67
Click chemistry probe
With advancements in bioorthogonal chemical reactions, especially the development of the click reaction, the limitations of biotinylated probes have been largely alleviated. In fact, the incorporation of an orthogonally reactive group in the structure of natural products has been one of the most widely used strategies for target identification in the last decade. With the orthogonal reaction group, such probes can undergo bioorthogonal click reactions with their complements (e.g., azide to alkyne,68 strained alkene to tetrazine,69 tetrazine to cyclopropane,70 etc. and vice versa), thereby covalently connecting probes to affinity tags (biotin-azide, biotin-cyclopropane, etc.) or fluorescent tags (rhodamine-azide, FITC-azide, etc.) for subsequent enrichment and target identification (Fig. 1b). Due to their relatively small sizes, these orthogonally reactive groups have little or no influence on the intrinsic pharmacological activity of the natural products, and the probes can easily reach the cytoplasm to bind target proteins in situ before the click reaction and enrichment.
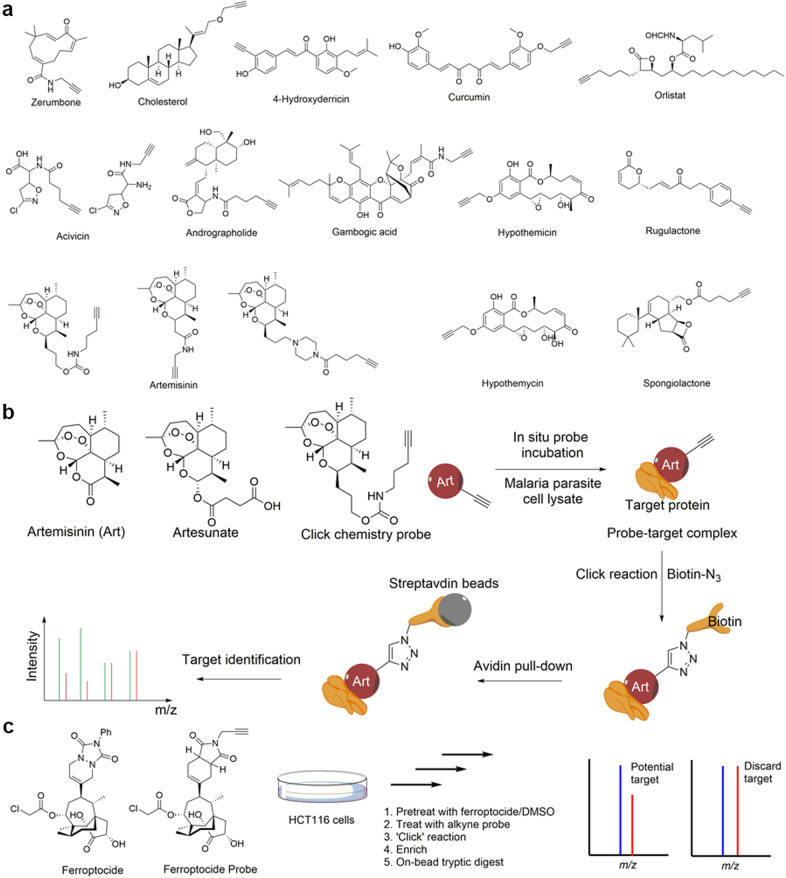
Many research groups, such as Tate’s group at Imperial College, Sieber’s group at the Technical University of Munich and Lin’s group at the National University of Singapore, have made great achievements in the target identification of natural products with click chemistry probes, including acivicin,71 curcumin,67 andrographolide,72 artemisinin,73 zerumbone74 and cholesterol.75 (Fig. 4a). Taking artemisinin as an example (Fig. 4b), Wang et al. utilized a click chemistry probe of artemisinin to identify its protein targets in Plasmodium falciparum and made two important findings: (i) heme, rather than free ferrous iron, is predominantly responsible for artemisinin activation; and (ii) artemisinin may kill the parasite through a promiscuous targeting mechanism. Because modifying artemisinin’s structure without influencing its activity is quite difficult, the authors synthesized a click chemistry probe derived from artesunate, an analog of artemisinin that also exhibits antimalarial activity. After activity validation, the probe was incubated with malaria parasites to fully bind the target proteins. Then, the target–probe complex was modified with a biotin tag through a click reaction and enriched with streptavidin beads. Finally, a total of 124 parasite proteins were identified, of which 33 proteins had previously been reported. Moreover, taking OAT (ornithine metabolism, arginine and proline metabolism) as a representative target, the activation mechanism of artemisinin was studied. It was observed that the probe itself did not bind to OAT, and its binding required the addition of hemin and was further enhanced in the presence of vitamin C, Na2S2O4 or glutathione, which reduce hemin to heme. By contrast, the addition of ferrous iron had no detectable effect on probe-OAT binding, revealing that heme, rather than free ferrous iron, has a predominant role in artemisinin activation.
Fig. 4.
Schematic of target identification with click chemistry probes of natural medicines. a Structures of some reported click chemistry probes of natural medicines. b Identification of artemisinin protein targets with its click chemistry probe. c Identification of ferroptocide protein targets with its click chemistry probe
Another example is ferroptocide,76 a small molecule chemically derived from the diterpene natural product pleuromutilin that rapidly and robustly induces ferroptotic death of cancer cells. After biological evaluation, Llabani et al. synthesized a click chemistry probe of ferroptocide to identify the protein targets in HCT 116 cells (Fig. 4c). The cells were pretreated with 20 μM ferroptocide or DMSO for 60 min and then incubated with 20 μM ferroptocide probe for 30 min; a click reaction with biotin-azide and enrichment with streptavidin magnetic beads followed. On-bead trypsin digestion coupled to LC/LC–MS/MS analysis provided a list of over 300 targets. With subsequent CRISPR knockout studies, the authors found that ferroptocide is an inhibitor of thioredoxin, a key component of the antioxidant system in the cell, and positively modulates the immune system in a murine model of breast cancer.
Photoaffinity probe
All the click chemistry probes mentioned above possess active groups that can covalently modify the amino acid residues in target proteins, leading to steady binding between the probes and the targets during the click reaction and enrichment. However, some natural products, such as resveratrol, interact with their protein targets through noncovalent secondary bonds, including hydrogen bonds, ionic bonds and hydrophobic interactions. For these bioactive molecules, simple click chemistry probes are unsuitable for target identification because the binding between the active molecule and the target protein can be disrupted during the click and enrich processes due to their noncovalent interactions. For such cases, the photoaffinity labeling technique (PAL) was developed.77–79
Photoaffinity probes generally consist of a click chemistry probe skeleton for target binding and enrichment and a photoaffinity group for fixing the binding between the probe and the targets (Fig. 1b). After incubation with the active proteome, the photoaffinity probe generates highly active free radical intermediates to covalently bind the target protein under certain wavelengths of light, and this is followed by click chemistry and target enrichment. Frequently used photoaffinity groups include benzophenone, aryl azide, and diazirine, and of these, benzophenone is the most widely applied due to its stability and ease of synthesis.80,81 In recent years, diazirine-based photoaffinity probe synthesis has attracted much interest due to its small size and high efficiency. Unfortunately, photoaffinity groups can intrinsically bind some nonspecific proteins (e.g., diazirine to voltage-dependent anion channels),82,83 which may affect the accuracy of the results.
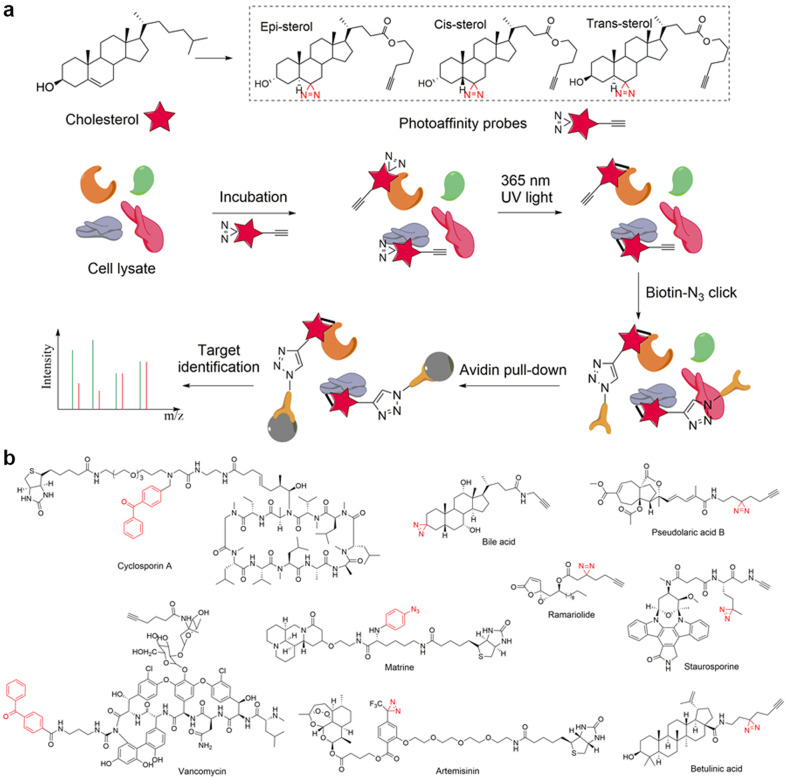
Cravatt’s group has utilized photoaffinity probes directly in living mammalian cells to globally map the binding proteins of cholesterol, an essential structural component of cellular membranes that serves as a precursor of several classes of signaling molecules.84 Based on cholesterol’s structure, they first designed and synthesized a set of sterol probes (Fig. 5a), each of which contained a photoactivatable diazirine group at the 6 position of the steroid core, which is a modification that has previously been shown to minimally perturb the biophysical properties of cholesterol. Then, living human cells were incubated with the probes and irradiated with 365 nm UV light to covalently cross-link the probe with the targets. After biotin modification through a click reaction and enrichment with streptavidin beads, the target proteins were identified with a quantitative proteomics approach. Over 250 cholesterol-binding proteins, including receptors, channels and enzymes involved in many established and previously unreported interactions, were identified. Other examples of photoaffinity probes are illustrated in Fig. 5b.85–93
Fig. 5.
Schematic of target identification with photoaffinity probes of natural medicines. a Cholesterol target identification with its photoaffinity probe. b Structures of some reported photoaffinity probes of natural medicines. The photoaffinity groups are indicated in red
Nonlabeling approach
In all the types of probes described above, the addition of an exogenous group could interfere with the pharmacological activity of the natural product. Moreover, for some natural products, their structures have no active sites suitable for modification, thereby limiting the application of these probe-based methods. For this reason, some nonlabeling chemical proteomics approaches, such as drug affinity responsive target stability (DARTS),94,95 stability of proteins from rates of oxidation (SPROX),96,97 cellular thermal shift assay (CETSA)98,99 and thermal proteome profiling (TPP),100 were developed for target identification. DARTS identifies targets by detecting enzymolysis changes based on the fact that the binding of the drug molecule stabilizes target proteins to trypsin-induced hydrolysis. For instance, Piazza et al.101 utilized a DARTS-based approach to identify the protein targets of three metabolites: adenosine 5′-triphosphate (ATP), l-phenylalanine (l-Phe) and phosphoenolpyruvate (PEP). In the study (Fig. 6a), proteomes were extracted under conditions that preserve the structures of native proteins, and the extracts were exposed to the small molecule of interest. Samples were subjected to limited proteolysis with the broad-specificity protease proteinase K to generate structure-specific protein fragments. The fragments were then digested with the sequence-specific protease trypsin to generate peptide mixtures amenable to bottom-up proteomic analysis. The peptides were analyzed with MS, and the targets were identified by comparing the peptides in the presence and absence of the small molecule. A total of 231 targets were observed for ATP, and 129 and 41 protein targets were identified for PEP and l-Phe, respectively.
Fig. 6.
a Schematic of target identification of ATP, PEP and l-Phe with DARTS. b Workflow of target fishing and MS identification. c Workflow of chemical proteomics combined with protein microarray
Unlike DARTS, SPROX detects the oxidation level of methionine in proteins to identify targets due to changes in antioxidant ability following the binding of the molecule.96 CETSA covers a wider range of applications than DARTS and SPROX, such as target identification in living cells, cell lysates and tissues. It is based on the thermodynamic stability alterations induced by the drug molecule’s binding. To overcome the challenges of low sensitivity and throughput in CETSA, TPP was developed.100 It is derived from CETSA but also allows the identification of off-targets and biomarkers. For these nonlabeling methods, the natural products need not be modified or attached to exogenous groups, allowing complete retention of their intrinsic bioactivities. Although nonlabeling chemical proteomics approaches have been widely applied in the target identification of natural products and medicines,97,102 they suffer from a few drawbacks, such as the tedious condition groping process and insufficient target sensitivity against nonlabeling molecules.
Target identification
After probe synthesis and subsequent target enrichment, the target proteins are identified with proteomic analysis by mass spectrometry, which is a valuable tool.103,104 In the early days of research, mass spectrometry was applied to identify specific bands in SDS electrophoretic gels or spots in two-dimensional electrophoretic gels.105,106 Because the targets are confirmed by comparing the gray values of proteins in different groups (probe vs control), this method may lead to low-abundance target loss and nonspecific results. To overcome these challenges, quantitative proteomics approaches for measuring abundance changes of many proteins in multiple samples have been developed.107–109 In addition, with the development of chip technology, protein microarrays have also been employed for target identification in chemical proteomics approaches.110,111 In this section, we will describe several different protein identification methods in detail, including their workflows, features and practical applications.
Gel separation and band identification
Briefly, the target proteins in the active proteome are enriched with molecule-derived probes and subsequently separated through SDS-PAGE or two-dimensional electrophoresis. Coomassie brilliant blue staining or silver staining can be used to visualize the separated proteins. Then, the gels containing the enriched proteins from different groups (always probe vs control) were aligned to identify the distinct bands or spots, followed by gel excision and in-gel digestion. Finally, the target proteins were identified by identifying the postdigested peptides with mass spectrometry (Fig. 6b). In the example mentioned above, our group identified resveratrol’s targets in mouse melanoma cells with this method (Fig. 3a). After probe synthesis and target enrichment, the binding proteins were eluted with SDS loading buffer. Then, the targets were separated by SDS-PAGE, followed by Coomassie brilliant blue staining, and we discovered two distinct bands compared with the control lane (DMSO). The two bands were excised and identified as acetyl-CoA acetyltransferase 1 (ACAT1) and HDAC1. With in vivo and in vitro experiments, we confirmed that resveratrol inhibits focal adhesion kinase (FAK) expression by interacting with HDAC1.41 Although the method has been widely used in protein identification,112–114 it has two disadvantages: (i) some low-abundance but vital target proteins are still invisible after Coomassie brilliant blue staining or even silver staining, resulting in target loss; and (ii) some distinct bands or spots, especially bands, contain more than one protein, so the nonspecific binding in these bands or spots may also be identified.
Quantitative proteomics
To overcome the deficiencies in-gel separation and band identification, quantitative methods have been incorporated into chemical proteomics.40,115–118 By comparing the relative abundance of proteins between different samples or against appropriate negative controls, proteins with a higher abundance ratio (>1.5 or 2, different values have been used) than a threshold are identified as specific targets, thus avoiding nonspecific binders. Due to the precision of the method, low-abundance targets can also be identified.119 Quantitative proteomics approaches mainly include stable isotope labeling by amino acids in cell culture (SILAC), chemical labeling approaches and label-free approaches.
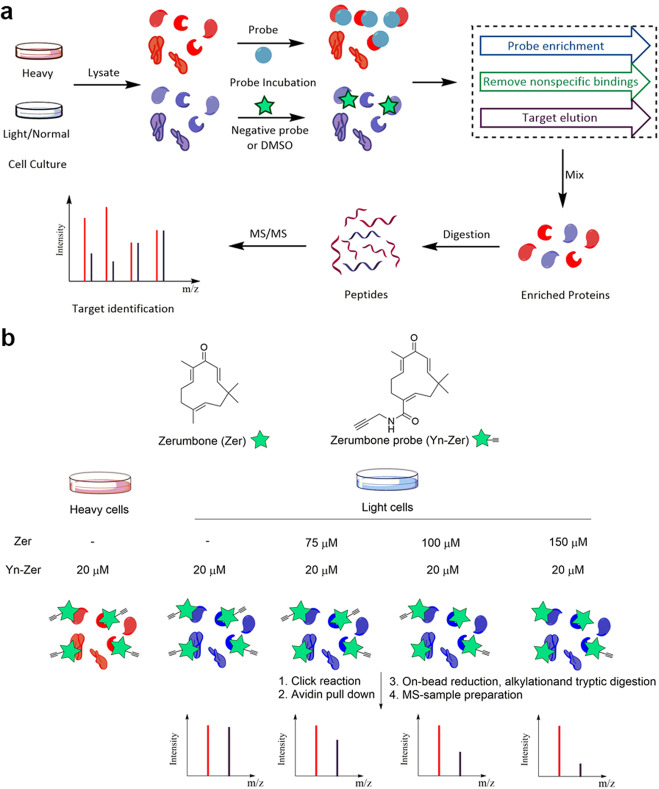
SILAC has become the most frequently used quantitative proteomics approach in target identification since its development in 2002.120 It uses isotopically labeled amino acids to stably label proteins during cell culture and determines the relative quantity of peptides by comparing the molecular weight shifts after MS. As the initial step of a typical SILAC experiment, two parallel cell populations are cultured in different media (one contains normal amino acids and another contains isotopically labeled amino acids, namely, “heavy” amino acids121) resulting in molecular weight differences in the newly synthesized proteins between the two populations after several generations of cell culture (Fig. 7a). Next, the synthesized probe and negative control (usually DMSO) are incubated with the heavy cells and normal cells, respectively. After “target fishing”, the enriched proteins from the two groups are pooled for subsequent identification to avoid measurement error. According to the shift in the molecular weight of the proteins from the two groups, the specific target proteins can be easily identified by comparing the relative protein abundances between the two groups.
Fig. 7.
Schematic of protein identification with SILAC. a General workflow of chemical proteomics combined with SILAC. b Target identification of zerumbone through a chemical proteomics approach coupled with SILAC
To date, many studies have applied chemical proteomics approaches along with SILAC to identify protein targets of various natural products as well as natural medicines. For instance, Kalesh et al.74 applied a “spike-in” SILAC method to identify the cellular targets of zerumbone (Zer), a phytochemical with diverse biological activities ranging from anti-inflammatory to anticancer properties. As shown in Fig. 7b, the authors initially synthesized a click chemistry probe of zerumbone (Yn-Zer), and the probe’s activity was confirmed to be similar to that of zerumbone. In parallel, lysates of HeLa cells labeled with 15N413C6-arginine and 15N213C6-lysine (heavy cells) were treated with 20 μM Yn-Zer, whereas lysates from cells cultured in normal medium (light cells) were treated with 20 μM Yn-Zer, 20 μM Yn-Zer combined with 75 μM Zer, 20 μM Yn-Zer combined with 100 μM Zer, and 20 μM Yn-Zer combined with 150 μM Zer. The addition of Zer competitively inhibited the binding between the target proteins and Yn-Zer. With SILAC, the relative amounts of the fished proteins (compared to the heavy group) were determined, and proteins with lower relative amounts with increasing Zer concentration were identified as specific targets. Finally, a total of 62 proteins that are involved in vital biological processes showed statistically significant enrichments, with many of these proteins having key roles in regulating apoptosis and cell survival.
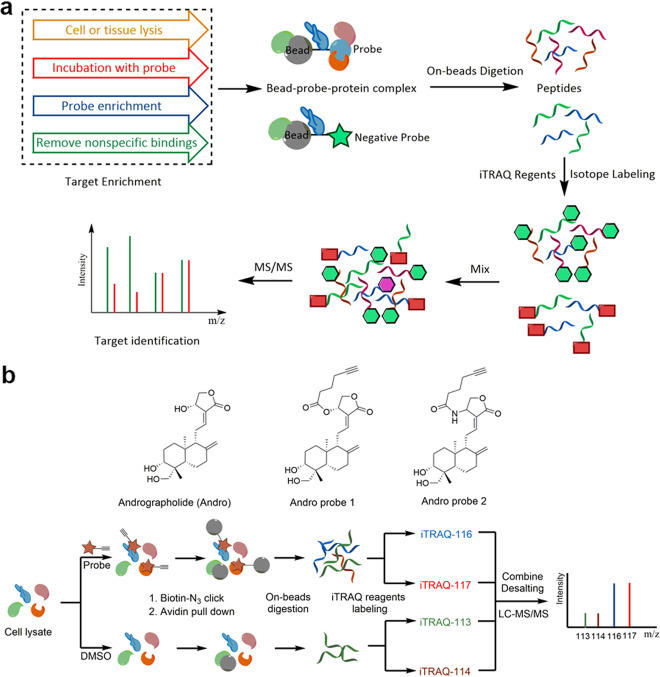
Compared with SILAC, isobaric tags for relative and absolute quantification (iTRAQ), a typical chemical labeling approach, can be used to perform stable isotope labeling of peptides digested from proteins and utilizes labeling reagents to quantify reporter ions fragmented by MS/MS,119 affording many advantages.122,123 For example, in some applications, such as identifying targets in natural microbial communities or primary tissue samples, SILAC is not suitable due to its complicated labeling process during cell culture, whereas iTRAQ is postapplicative.124 Moreover, in a SILAC experiment, at most three samples can be determined at one time, whereas iTRAQ can simultaneously analyze up to eight samples.125 The general workflow for target identification with iTRAQ is illustrated in Fig. 8a, and it only differs from SILAC in the labeling process. In iTRAQ, the peptides digested from proteins of different groups are incubated with different iTRAQ regents for isotope labeling. Due to the mass difference of reporter ions in different iTRAQ reagents, the relative protein abundance in different groups can be calculated. iTRAQ has also been widely applied in the identification of targets of natural medicines.126,127
Fig. 8.
Schematic of protein identification with iTRAQ. a Workflow of chemical proteomics combined with iTRAQ. b Andrographolide target identification with chemical proteomics combined with iTRAQ
For example, Wang et al.72 utilized a clickable activity-based probe derived from andrographolide, a natural product with known anti-inflammatory and anticancer effects, to enrich protein targets in live cancer cells (Fig. 8b). In the assay, the cells were first incubated with the clickable probe or with DMSO as a negative control. Following sequential probe binding, biotin modification through a click reaction, avidin pull-down and thorough washing, the target proteins were digested with trypsin. The resulting peptides were labeled with their respective iTRAQ regents (control group with 113 and 114, whereas probe group with 116 and 117) and pooled for further identification and quantification via LC–MS/MS. The results were also validated through cell migration and invasion assays, revealing that andrographolide has a potential novel application as a tumor metastasis inhibitor.
For the label-free quantitative proteomics approach, protein abundance is calculated by detecting the MS signal densities of peptides digested from the specific protein or the number of MS/MS signals corresponding to peptides and proteins.128 Due to the missing labeling process, this method is much simpler and more cost-efficient.129 However, it has disadvantages in accuracy and throughput, especially in some promiscuous cases, such as samples with heavy backgrounds, compared to quantitative labeling proteomics approaches.130,131 Moreover, in the label-free approach, only one MS run can be done per sample, so samples of different groups have to be examined in separate MS runs, which might lead to operating error, leading to reduced accuracy in the results.
Protein microarray
In addition to MS-based approaches to identify the protein targets of drug molecules of interest, protein microarrays are another approach that has been utilized frequently. The main function of protein microarray technology is investigating the functional properties of immobilized proteins, the interaction of proteins, and their enzyme activity, and it has also been utilized to study the interactions between proteins and natural products or medicines for decades.132,133 Combined with chemical proteomics, this method can also be applied for target identification, providing a platform for analyzing the interactions between small molecules and thousands of proteins.134 In a typical protein microarray approach (Fig. 6c), diverse proteins are first immobilized on a miniature high-density array, followed by the labeling of the molecule to be tested with an affinity tag, such as biotin, a fluorophore, a photoaffinity group or a radioactive isotope, to allow the molecule-linked proteins to be easily traced. Notably, this method is high-throughput, allowing the identification of target and off-target proteins in the whole proteome on the microarray in a single run.135,136 However, it also possesses certain disadvantages. For example, active molecules need to be modified with tags that might influence their intrinsic activities. Along with the development of mass spectrometry techniques, protein microarrays are always combined with mass spectrometry to overcome modification-induced activity alterations.137
Summary and outlook
In the development of new natural medicines, target identification facilitates the determination of the MOA and side effects, accelerating this process from discovery to market. Along with the development of chemical biology and proteomics, the chemical proteomics approach has become a popular method in target identification of small active molecules, especially natural products, providing an important theoretical basis for novel natural medicine research and development. In most cases, natural products need to be modified with reporter tags to facilitate enrichment or detection, which might influence their intrinsic pharmacological activities, thereby leading to a biased target result. Moreover, some nonlabeling chemical proteomics approaches for target identification are not well qualified in promiscuous cases, and their low accuracy and throughput limit their broad application. Therefore, developing a highly accurate nonlabeling chemical proteomics approach with high-throughput is imperative. Although some studies have simultaneously applied two or more different strategies for target identification to avoid nonspecific binding and narrow target collection and obtained a more accurate target list, the MOA predicted from the targets must be validated by biochemical methods. Collectively, the use of chemical proteomics will continue to be a key tool to drive the discovery of new therapeutic compounds of natural origin.
Acknowledgements
We gratefully acknowledge financial support from the National Natural Science Foundation of China (81903588, 81803456 and 81841001); the Major National Science and Technology Program of China for Innovative Drug (2017ZX09101002-001-001-05); the Fundamental Research Funds for the Central Public Welfare Research Institutes (ZXKT18003); the Natural Science Foundation of Jiangsu Province (BK20190799, China); the Science and Technology Foundation of Shenzhen (JCYJ20180305164128430); the International Cooperation Foundation of Shenzhen (GJHZ20180928171602104); the Shenzhen Economic and Information Committee “Innovation Chain and Industry Chain” integration special support plan project (20180225112449943); and the Shenzhen Public Service Platform on Tumor Precision Medicine and Molecular Diagnosis.
Competing interests
The authors declare no competing interests.
Footnotes
These authors contributed equally: Xiao Chen, Yutong Wang, Nan Ma
Contributor Information
Zhen Liang, Email: Liang.zhen@szhospital.com.
Chang Zou, Email: zou.chang@szhospital.com.
Jigang Wang, Email: jgwang@icmm.ac.cn.
References
- 1.Newman DJ, Cragg GM. Natural products as sources of new drugs over the last 25 years. J. Nat. Prod. 2007;70:461–477. doi: 10.1021/np068054v. [DOI] [PubMed] [Google Scholar]
- 2.Guo Z. The modification of natural products for medical use. Acta Pharm. Sin. B. 2017;7:119–136. doi: 10.1016/j.apsb.2016.06.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Carlson EE. Natural products as chemical probes. ACS Chem. Biol. 2010;5:639–653. doi: 10.1021/cb100105c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Rodrigues T, Reker D, Schneider P, Schneider G. Counting on natural products for drug design. Nat. Chem. 2016;8:531. doi: 10.1038/nchem.2479. [DOI] [PubMed] [Google Scholar]
- 5.Gouyette A. Synthesis of deuterium-labelled elliptinium and its use in metabolic studies. Biomed. Environ. Mass Spectrom. 1988;15:243–247. doi: 10.1002/bms.1200150502. [DOI] [PubMed] [Google Scholar]
- 6.Caille P, et al. Phase II trial of elliptinium in advanced renal cell carcinoma. Cancer Treat. Rep. 1985;69:901–902. [PubMed] [Google Scholar]
- 7.Jones RN, Fritsche TR, Sader HS, Ross JE. Activity of retapamulin (SB-275833), a novel pleuromutilin, against selected resistant gram-positive cocci. Antimicrob. Agents Chemother. 2006;50:2583–2586. doi: 10.1128/AAC.01432-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Berman AY, Motechin RA, Wiesenfeld MY, Holz MK. The therapeutic potential of resveratrol: a review of clinical trials. NPJ Precis. Oncol. 2017;1:35. doi: 10.1038/s41698-017-0038-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bonnefont-Rousselot D. Resveratrol and cardiovascular diseases. Nutrients. 2016;8:250. doi: 10.3390/nu8050250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Xia N, Daiber A, Förstermann U, Li H. Antioxidant effects of resveratrol in the cardiovascular system. Br. J. Pharmacol. 2017;174:1633–1646. doi: 10.1111/bph.13492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kunnumakkara AB, et al. Curcumin, the golden nutraceutical: multitargeting for multiple chronic diseases. Br. J. Pharm. 2017;174:1325–1348. doi: 10.1111/bph.13621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chen, Y. et al. Platinum complexes of curcumin delivered by dual-responsive polymeric nanoparticles improve chemotherapeutic efficacy based on the enhanced anti-metastasis activity and reduce side effects. Acta Pharm. Sin. B (2019). 10.1016/j.apsb.2019.10.011. [DOI] [PMC free article] [PubMed]
- 13.Zhang T, et al. Inhalation treatment of primary lung cancer using liposomal curcumin dry powder inhalers. Acta Pharm. Sin. B. 2018;8:440–448. doi: 10.1016/j.apsb.2018.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ding Y, et al. Discovery and development of natural product oridonin-inspired anticancer agents. Eur. J. Med. Chem. 2016;122:102–117. doi: 10.1016/j.ejmech.2016.06.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Zhu L, Li M, Liu X, Du L, Jin Y. Inhalable oridonin-loaded poly (lactic-co-glycolic) acid large porous microparticles for in situ treatment of primary non-small cell lung cancer. Acta Pharm. Sin. B. 2017;7:80–90. doi: 10.1016/j.apsb.2016.09.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Chai X-Y, et al. Six insecticidal isoryanodane diterpenoids from the bark and twigs of Itoa orientalis. Tetrahedron. 2008;64:5743–5747. [Google Scholar]
- 17.Liu L, Li A-L, Zhao M-B, Tu P-F. Tetralones and flavonoids from Pyrola calliantha. Chem. Biodivers. 2007;4:2932–2937. doi: 10.1002/cbdv.200790242. [DOI] [PubMed] [Google Scholar]
- 18.Jin W, et al. Simultaneous analysis of multiple bioactive constituents in Rheum tanguticum Maxim. ex Balf. by high-performance liquid chromatography coupled to tandem mass spectrometry. Rapid Commun. Mass Spectrom. 2007;21:2351–2360. doi: 10.1002/rcm.3086. [DOI] [PubMed] [Google Scholar]
- 19.Chen X, et al. Target identification with quantitative activity based protein profiling (ABPP) Proteomics. 2017;17:1600212. doi: 10.1002/pmic.201600212. [DOI] [PubMed] [Google Scholar]
- 20.Comess KM, et al. Emerging approaches for the identification of protein targets of small molecules-a practitioners’ perspective. J. Med. Chem. 2018;61:8504–8535. doi: 10.1021/acs.jmedchem.7b01921. [DOI] [PubMed] [Google Scholar]
- 21.Schenone M, Dančik V, Wagner BK, Clemons PA. Target identification and mechanism of action in chemical biology and drug discovery. Nat. Chem. Biol. 2013;9:232. doi: 10.1038/nchembio.1199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Zhong, C. et al. Identification of bioactive anti-angiogenic components targeting tumor endothelial cells in Shenmai injection using multidimensional pharmacokinetics. Acta Pharm. Sin. B (2019). 10.1016/j.apsb.2019.12.011. [DOI] [PMC free article] [PubMed]
- 23.Rix U, Superti-Furga G. Target profiling of small molecules by chemical proteomics. Nat. Chem. Biol. 2009;5:616–624. doi: 10.1038/nchembio.216. [DOI] [PubMed] [Google Scholar]
- 24.Majumder A, Biswal MR, Prakash MK. One drug multiple targets: An approach to predict drug efficacies on bacterial strains differing in membrane composition. ACS Omega. 2019;4:4977–4983. [Google Scholar]
- 25.Klessig DF, Tian M, Choi HW. Multiple targets of salicylic acid and its derivatives in plants and animals. Front. Immunol. 2016;7:206. doi: 10.3389/fimmu.2016.00206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zon LI, Peterson RT. In vivo drug discovery in the zebrafish. Nat. Rev. Drug Discov. 2005;4:35. doi: 10.1038/nrd1606. [DOI] [PubMed] [Google Scholar]
- 27.Lamb J, et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science. 2006;313:1929–1935. doi: 10.1126/science.1132939. [DOI] [PubMed] [Google Scholar]
- 28.Hillenmeyer ME, et al. The chemical genomic portrait of yeast: uncovering a phenotype for all genes. Science. 2008;320:362–365. doi: 10.1126/science.1150021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Caligiuri M, et al. MASPIT: three-hybrid trap for quantitative proteome fingerprinting of small molecule-protein interactions in mammalian cells. Chem. Biol. 2006;13:711–722. doi: 10.1016/j.chembiol.2006.05.008. [DOI] [PubMed] [Google Scholar]
- 30.Cohen AA, et al. Dynamic proteomics of individual cancer cells in response to a drug. Science. 2008;322:1511–1516. doi: 10.1126/science.1160165. [DOI] [PubMed] [Google Scholar]
- 31.Wang, S. et al. Advanced activity-based protein profiling application strategies for drug development. Front. Pharmacol.9, 1–9 (2018). [DOI] [PMC free article] [PubMed]
- 32.Lum KM, et al. Mapping protein targets of bioactive small molecules using lipid-based chemical proteomics. ACS Chem. Biol. 2017;12:2671–2681. doi: 10.1021/acschembio.7b00581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Bantscheff M, Scholten A, Heck AJR. Revealing promiscuous drug-target interactions by chemical proteomics. Drug Discov. Today. 2009;14:1021–1029. doi: 10.1016/j.drudis.2009.07.001. [DOI] [PubMed] [Google Scholar]
- 34.Yang, Y., Fonović, M. & Verhelst, S. H. L. Cleavable linkers in chemical proteomics applications. Methods Mol Biol.1191, 185–203 (2017). [DOI] [PubMed]
- 35.Itzhak Daniel N., Sacco Francesca, Nagaraj Nagarjuna, Tyanova Stefka, Mann Matthias, Murgia Marta. SILAC-based quantitative proteomics using mass spectrometry quantifies endoplasmic reticulum stress in whole HeLa cells. Disease Models & Mechanisms. 2019;12(11):dmm040741. doi: 10.1242/dmm.040741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Jing Y, Wan J, Angelidaki I, Zhang S, Luo G. iTRAQ quantitative proteomic analysis reveals the pathways for methanation of propionate facilitated by magnetite. Water Res. 2017;108:212–221. doi: 10.1016/j.watres.2016.10.077. [DOI] [PubMed] [Google Scholar]
- 37.Moulder R, Bhosale SD, Goodlett DR, Lahesmaa R. Analysis of the plasma proteome using iTRAQ and TMT-based Isobaric labeling. Mass Spectrom. Rev. 2018;37:583–606. doi: 10.1002/mas.21550. [DOI] [PubMed] [Google Scholar]
- 38.West GM, et al. Mass spectrometry-based thermal shift assay for protein- ligand binding analysis. Anal. Chem. 2010;82:5573–5581. doi: 10.1021/ac100465a. [DOI] [PubMed] [Google Scholar]
- 39.Akter S, et al. Chemical proteomics reveals new targets of cysteine sulfinic acid reductase. Nat. Chem. Biol. 2018;14:995–1004. doi: 10.1038/s41589-018-0116-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.van Rooden EJ, et al. Mapping in vivo target interaction profiles of covalent inhibitors using chemical proteomics with label-free quantification. Nat. Protoc. 2018;13:752. doi: 10.1038/nprot.2017.159. [DOI] [PubMed] [Google Scholar]
- 41.Chen X, et al. Comparative profiling of analog targets: A case study on resveratrol for mouse melanoma metastasis suppression. Theranostics. 2018;8:3504–3516. doi: 10.7150/thno.24336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Bar-Peled L, et al. Chemical proteomics identifies druggable vulnerabilities in a genetically defined cancer. Cell. 2017;171:696–709. doi: 10.1016/j.cell.2017.08.051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Hu L, Fawcett JP, Gu J. Protein target discovery of drug and its reactive intermediate metabolite by using proteomic strategy. Acta Pharm. Sin. B. 2012;2:126–136. [Google Scholar]
- 44.Speers AE, Cravatt BF. Activity-based protein profiling (ABPP) and click chemistry (CC)—ABPP by MudPIT mass spectrometry. Curr. Protoc. Chem. Biol. 2009;1:29–41. doi: 10.1002/9780470559277.ch090138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Cuatrecasas P, Wilchek M, Anfinsen CB. Selective enzyme purification by affinity chromatography. Proc. Natl Acad. Sci. USA. 1968;61:636. doi: 10.1073/pnas.61.2.636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Harding MW, Galat A, Uehling DE, Schreiber SL. A receptor for the immuno-suppressant FK506 is a cis–trans peptidyl-prolyl isomerase. Nature. 1989;341:758. doi: 10.1038/341758a0. [DOI] [PubMed] [Google Scholar]
- 47.Crews CM, Collins JL, Lane WS, Snapper ML, Schreiber SL. GTP-dependent binding of the antiproliferative agent didemnin to elongation factor 1 alpha. J. Biol. Chem. 1994;269:15411–15414. [PubMed] [Google Scholar]
- 48.Knockaert M, et al. Intracellular targets of cyclin-dependent kinase inhibitors: identification by affinity chromatography using immobilised inhibitors. Chem. Biol. 2000;7:411–422. doi: 10.1016/s1074-5521(00)00124-1. [DOI] [PubMed] [Google Scholar]
- 49.Sieber, S. A. & Cravatt, B. F. Analytical platforms for activity-based protein profiling–exploiting the versatility of chemistry for functional proteomics. Chem. Commun.22, 2311–2319 (2006). [DOI] [PubMed]
- 50.Yang P, Liu K. Activity-based protein profiling: recent advances in probe development and applications. ChemBioChem. 2015;16:712–724. doi: 10.1002/cbic.201402582. [DOI] [PubMed] [Google Scholar]
- 51.Taunton J, Hassig CA, Schreiber SL. A mammalian histone deacetylase related to the yeast transcriptional regulator Rpd3p. Science. 1996;272:408–411. doi: 10.1126/science.272.5260.408. [DOI] [PubMed] [Google Scholar]
- 52.Li J, et al. Artemisinins target GABAA receptor signaling and impair α cell identity. Cell. 2017;168:86–100. doi: 10.1016/j.cell.2016.11.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Capolupo A, et al. Determination of gymnemic acid I as a protein biosynthesis inhibitor using chemical proteomics. J. Nat. Prod. 2017;80:909–915. doi: 10.1021/acs.jnatprod.6b00793. [DOI] [PubMed] [Google Scholar]
- 54.Cassiano C, et al. In cell scalaradial interactome profiling using a bio-orthogonal clickable probe. Chem. Commun. 2014;50:6043–6045. doi: 10.1039/c4cc00989d. [DOI] [PubMed] [Google Scholar]
- 55.Margarucci L, et al. Chemical proteomics-driven discovery of oleocanthal as an Hsp90 inhibitor. Chem. Commun. 2013;49:5844–5846. doi: 10.1039/c3cc41858h. [DOI] [PubMed] [Google Scholar]
- 56.Zhao Q, et al. Natural products triptolide, celastrol, and withaferin A inhibit the chaperone activity of peroxiredoxin I. Chem. Sci. 2015;6:4124–4130. doi: 10.1039/c5sc00633c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Dong T, et al. Ainsliadimer A selectively inhibits IKKα/β by covalently binding a conserved cysteine. Nat. Commun. 2015;6:6522. doi: 10.1038/ncomms7522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Sin N, et al. The anti-angiogenic agent fumagillin covalently binds and inhibits the methionine aminopeptidase, MetAP-2. Proc. Natl Acad. Sci. USA. 1997;94:6099–6103. doi: 10.1073/pnas.94.12.6099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Kong L-M, et al. Identification and validation of p50 as the cellular target of eriocalyxin B. Oncotarget. 2014;5:11354. doi: 10.18632/oncotarget.2461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Klaić L, Morimoto RI, Silverman RB. Celastrol analogues as inducers of the heat shock response. Design and synthesis of affinity probes for the identification of protein targets. ACS Chem. Biol. 2012;7:928–937. doi: 10.1021/cb200539u. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.WonáKim S, et al. Identification of actin as a direct proteomic target of berberine using an affinity-based chemical probe and elucidation of its modulatory role in actin assembly. Chem. Commun. 2017;53:7045–7047. doi: 10.1039/c7cc02789c. [DOI] [PubMed] [Google Scholar]
- 62.Ki SW, et al. Radicicol binds and inhibits mammalian ATP citrate lyase. J. Biol. Chem. 2000;275:39231–39236. doi: 10.1074/jbc.M006192200. [DOI] [PubMed] [Google Scholar]
- 63.Bargagna-Mohan P, et al. The tumor inhibitor and antiangiogenic agent withaferin A targets the intermediate filament protein vimentin. Chem. Biol. 2007;14:623–634. doi: 10.1016/j.chembiol.2007.04.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Kudo N, et al. Leptomycin B inactivates CRM1/exportin 1 by covalent modification at a cysteine residue in the central conserved region. Proc. Natl Acad. Sci. USA. 1999;96:9112–9117. doi: 10.1073/pnas.96.16.9112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Charron G, et al. Robust fluorescent detection of protein fatty-acylation with chemical reporters. J. Am. Chem. Soc. 2009;131:4967–4975. doi: 10.1021/ja810122f. [DOI] [PubMed] [Google Scholar]
- 66.Wright MH, Sieber SA. Chemical proteomics approaches for identifying the cellular targets of natural products. Nat. Prod. Rep. 2016;00:1–28. doi: 10.1039/c6np00001k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Wang J, et al. In situ proteomic profiling of curcumin targets in HCT116 colon cancer cell line. Sci. Rep. 2016;6:22146. doi: 10.1038/srep22146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Baskin JM, et al. Copper-free click chemistry for dynamic in vivo imaging. Proc. Natl Acad. Sci. USA. 2007;104:16793–16797. doi: 10.1073/pnas.0707090104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Devaraj NK, Upadhyay R, Haun JB, Hilderbrand SA, Weissleder R. Fast and sensitive pretargeted labeling of cancer cells through a tetrazine/trans-cyclooctene cycloaddition. Angew. Chem. Int. Ed. 2009;48:7013–7016. doi: 10.1002/anie.200903233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Patterson DM, Nazarova LA, Xie B, Kamber DN, Prescher JA. Functionalized cyclopropenes as bioorthogonal chemical reporters. J. Am. Chem. Soc. 2012;134:18638–18643. doi: 10.1021/ja3060436. [DOI] [PubMed] [Google Scholar]
- 71.Battenberg OA, Yang Y, Verhelst SHL, Sieber SA. Target profiling of 4-hydroxyderricin in S. aureus reveals seryl-tRNA synthetase binding and inhibition by covalent modification. Mol. Biosyst. 2013;9:343–351. doi: 10.1039/c2mb25446h. [DOI] [PubMed] [Google Scholar]
- 72.Wang J, et al. A quantitative chemical proteomics approach to profile the specific cellular targets of andrographolide, a promising anticancer agent that suppresses tumor metastasis. Mol. Cell. Proteom. 2014;13:876–886. doi: 10.1074/mcp.M113.029793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Wang J, et al. Haem-activated promiscuous targeting of artemisinin in Plasmodium falciparum. Nat. Commun. 2015;6:10111. doi: 10.1038/ncomms10111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Kalesh KA, Clulow JA, Tate EW. Target profiling of zerumbone using a novel cell-permeable clickable probe and quantitative chemical proteomics. Chem. Commun. 2015;51:5497–5500. doi: 10.1039/c4cc09527h. [DOI] [PubMed] [Google Scholar]
- 75.Ciepla P, et al. New chemical probes targeting cholesterylation of Sonic Hedgehog in human cells and zebrafish. Chem. Sci. 2014;5:4249–4259. doi: 10.1039/c4sc01600a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Llabani E, et al. Diverse compounds from pleuromutilin lead to a thioredoxin inhibitor and inducer of ferroptosis. Nat. Chem. 2019;11:521–532. doi: 10.1038/s41557-019-0261-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Smith E, Collins I. Photoaffinity labeling in target-and binding-site identification. Future Med. Chem. 2015;7:159–183. doi: 10.4155/fmc.14.152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Xiao F, Zhang X, Lei X. Recent developments and applications of photoconjugation chemistry. Chim. Int. J. Chem. 2018;72:782–790. doi: 10.2533/chimia.2018.782. [DOI] [PubMed] [Google Scholar]
- 79.Liu X, Dong T, Zhou Y, Huang N, Lei X. Exploring the binding proteins of glycolipids with bifunctional chemical probes. Angew. Chem. Int. Ed. 2016;55:14330–14334. doi: 10.1002/anie.201608827. [DOI] [PubMed] [Google Scholar]
- 80.Lee B, et al. Design, synthesis and biological evaluation of photoaffinity probes of antiangiogenic homoisoflavonoids. Bioorg. Med. Chem. Lett. 2016;26:4277–4281. doi: 10.1016/j.bmcl.2016.07.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Wu Y, et al. Development of a multifunctional benzophenone linker for peptide stapling and photoaffinity labelling. ChemBioChem. 2016;17:689–692. doi: 10.1002/cbic.201500648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Budelier MM, et al. Photoaffinity labeling with cholesterol analogues precisely maps a cholesterol-binding site in voltage-dependent anion channel-1. J. Biol. Chem. 2017;292:9294–9304. doi: 10.1074/jbc.M116.773069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Cheng WWL, et al. Multiple neurosteroid and cholesterol binding sites in voltage-dependent anion channel-1 determined by photo-affinity labeling. Biochim. Biophys. Acta. 2019;1864:1269–1279. doi: 10.1016/j.bbalip.2019.06.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Hulce JJ, Cognetta AB, Niphakis MJ, Tully SE, Cravatt BF. Proteome-wide mapping of cholesterol-interacting proteins in mammalian cells. Nat. Methods. 2013;10:259–264. doi: 10.1038/nmeth.2368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Wang D, et al. Identification of Annexin A2 as a target protein for plant alkaloid matrine. Chem. Commun. 2017;53:5020–5023. doi: 10.1039/c7cc02227a. [DOI] [PubMed] [Google Scholar]
- 86.Guo H, Xu J, Hao P, Ding K, Li Z. Competitive affinity-based proteome profiling and imaging to reveal potential cellular targets of betulinic acid. Chem. Commun. 2017;53:9620–9623. doi: 10.1039/c7cc04190j. [DOI] [PubMed] [Google Scholar]
- 87.Konziase B. Biotinylated probes of artemisinin with labeling affinity toward Trypanosoma brucei brucei target proteins. Anal. Biochem. 2015;482:25–31. doi: 10.1016/j.ab.2015.04.020. [DOI] [PubMed] [Google Scholar]
- 88.Eirich J, Orth R, Sieber SA. Unraveling the protein targets of vancomycin in living S. aureus and E. faecalis cells. J. Am. Chem. Soc. 2011;133:12144–12153. doi: 10.1021/ja2039979. [DOI] [PubMed] [Google Scholar]
- 89.Zhou Y, et al. Chemical proteomics reveal CD147 as a functional target of pseudolaric acid B in human cancer cells. Chem. Commun. 2017;53:8671–8674. doi: 10.1039/c7cc04345g. [DOI] [PubMed] [Google Scholar]
- 90.Lehmann J, Richers J, Pöthig A, Sieber SA. Synthesis of ramariolide natural products and discovery of their targets in mycobacteria. Chem. Commun. 2017;53:107–110. doi: 10.1039/c6cc08365j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Zhuang S, Li Q, Cai L, Wang C, Lei X. Chemoproteomic profiling of bile acid interacting proteins. ACS Cent. Sci. 2017;3:501–509. doi: 10.1021/acscentsci.7b00134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Shi H, Cheng X, Sze SK, Yao SQ. Proteome profiling reveals potential cellular targets of staurosporine using a clickable cell-permeable probe. Chem. Commun. 2011;47:11306–11308. doi: 10.1039/c1cc14824a. [DOI] [PubMed] [Google Scholar]
- 93.Lamos SM, et al. Mixed isotope photoaffinity reagents for identification of small-molecule targets by mass spectrometry. Angew. Chem. Int. Ed. 2006;45:4329–4333. doi: 10.1002/anie.200600743. [DOI] [PubMed] [Google Scholar]
- 94.Lomenick B, et al. Target identification using drug affinity responsive target stability (DARTS) Proc. Natl Acad. Sci. USA. 2009;106:21984–21989. doi: 10.1073/pnas.0910040106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Lomenick B, Jung G, Wohlschlegel JA, Huang J. Target identification using drug affinity responsive target stability (DARTS) Curr. Protoc. Chem. Biol. 2011;3:163–180. doi: 10.1002/9780470559277.ch110180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Strickland EC, et al. Thermodynamic analysis of protein-ligand binding interactions in complex biological mixtures using the stability of proteins from rates of oxidation. Nat. Protoc. 2013;8:148. doi: 10.1038/nprot.2012.146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.DeArmond PD, Xu Y, Strickland EC, Daniels KG, Fitzgerald MC. Thermodynamic analysis of protein–ligand interactions in complex biological mixtures using a shotgun proteomics approach. J. Proteome Res. 2011;10:4948–4958. doi: 10.1021/pr200403c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Molina DM, et al. Monitoring drug target engagement in cells and tissues using the cellular thermal shift assay. Science. 2013;341:84–87. doi: 10.1126/science.1233606. [DOI] [PubMed] [Google Scholar]
- 99.Jafari R, et al. The cellular thermal shift assay for evaluating drug target interactions in cells. Nat. Protoc. 2014;9:2100. doi: 10.1038/nprot.2014.138. [DOI] [PubMed] [Google Scholar]
- 100.Savitski MM, et al. Tracking cancer drugs in living cells by thermal profiling of the proteome. Science. 2014;346:1255784. doi: 10.1126/science.1255784. [DOI] [PubMed] [Google Scholar]
- 101.Piazza I, et al. A map of protein-metabolite interactions reveals principles of chemical communication. Cell. 2018;172:358–372.e23. doi: 10.1016/j.cell.2017.12.006. [DOI] [PubMed] [Google Scholar]
- 102.Kost GC, et al. A novel anti-cancer agent, 1-(3, 5-dimethoxyphenyl)-4-[(6-fluoro-2-methoxyquinoxalin-3-yl) aminocarbonyl] piperazine (RX-5902), interfereswith β-catenin function through Y593 phospho-p68 RNA helicase. J. Cell Biochem. 2015;116:1595–1601. doi: 10.1002/jcb.25113. [DOI] [PubMed] [Google Scholar]
- 103.Ryan DJ, Spraggins JM, Caprioli RM. Protein identification strategies in MALDI imaging mass spectrometry: a brief review. Curr. Opin. Chem. Biol. 2019;48:64–72. doi: 10.1016/j.cbpa.2018.10.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Brunet, M. A. & Roucou, X. Mass spectrometry-based proteomics analyses using the OpenProt database to unveil novel proteins translated from non-canonical open reading frames. JoVE. 146, e59589 (2019). [DOI] [PubMed]
- 105.Gharahdaghi F, Weinberg CR, Meagher DA, Imai BS, Mische SM. Mass spectrometric identification of proteins from silver-stained polyacrylamide gel: a method for the removal of silver ions to enhance sensitivity. Electrophor. Int. J. 1999;20:601–605. doi: 10.1002/(SICI)1522-2683(19990301)20:3<601::AID-ELPS601>3.0.CO;2-6. [DOI] [PubMed] [Google Scholar]
- 106.Raikos V, Hansen R, Campbell L, Euston SR. Separation and identification of hen egg protein isoforms using SDS–PAGE and 2D gel electrophoresis with MALDI-TOF mass spectrometry. Food Chem. 2006;99:702–710. [Google Scholar]
- 107.Ong S-E, Foster LJ, Mann M. Mass spectrometric-based approaches in quantitative proteomics. Methods. 2003;29:124–130. doi: 10.1016/s1046-2023(02)00303-1. [DOI] [PubMed] [Google Scholar]
- 108.Mann M. Functional and quantitative proteomics using SILAC. Nat. Rev. Mol. Cell Biol. 2006;7:952–958. doi: 10.1038/nrm2067. [DOI] [PubMed] [Google Scholar]
- 109.Zhu, W., Smith, J. W. & Huang, C.-M. Mass spectrometry-based label-free quantitative proteomics. Biomed Res. Int. 2010, 840518 (2009). [DOI] [PMC free article] [PubMed]
- 110.Templin MF, et al. Protein microarray technology. Drug Discov. Today. 2002;7:815–822. doi: 10.1016/s1359-6446(00)01910-2. [DOI] [PubMed] [Google Scholar]
- 111.Wilson DS, Nock S. Recent developments in protein microarray technology. Angew. Chem. Int. Ed. 2003;42:494–500. doi: 10.1002/anie.200390150. [DOI] [PubMed] [Google Scholar]
- 112.Wang D-Y, et al. Target identification of kinase inhibitor alisertib (MLN8237) by using DNA-programmed affinity labeling. Chem. Eur. J. 2017;23:10906–10914. doi: 10.1002/chem.201702033. [DOI] [PubMed] [Google Scholar]
- 113.Rodriguez-Furlan C, Zhang C, Raikhel N, Hicks GR. Drug affinity responsive target stability (DARTS) to resolve protein–small molecule interaction in arabidopsis. Curr. Protoc. Plant Biol. 2017;2:370–378. doi: 10.1002/cppb.20062. [DOI] [PubMed] [Google Scholar]
- 114.Ehe BK, et al. Identification of a DYRK1A-mediated phosphorylation site within the nuclear localization sequence of the hedgehog transcription factor GLI1. Biochem. Biophys. Res. Commun. 2017;491:767–772. doi: 10.1016/j.bbrc.2017.07.107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Kleiner RE, Hang LE, Molloy KR, Chait BT, Kapoor TM. A chemical proteomics approach to reveal direct protein-protein interactions in living cells. Cell Chem. Biol. 2018;25:110–120. doi: 10.1016/j.chembiol.2017.10.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Yang F, Gao J, Che J, Jia G, Wang C. A dimethyl-labeling-based strategy for site-specifically quantitative chemical proteomics. Anal. Chem. 2018;90:9576–9582. doi: 10.1021/acs.analchem.8b02426. [DOI] [PubMed] [Google Scholar]
- 117.Chen Y, Liu Y, Hou X, Ye Z, Wang C. Quantitative and site-specific chemoproteomic profiling of targets of acrolein. Chem. Res. Toxicol. 2019;32:467–473. doi: 10.1021/acs.chemrestox.8b00343. [DOI] [PubMed] [Google Scholar]
- 118.Ye Z, et al. Chemoproteomic profiling reveals ethacrynic acid targets adenine nucleotide translocases to impair mitochondrial function. Mol. Pharm. 2018;15:2413–2422. doi: 10.1021/acs.molpharmaceut.8b00250. [DOI] [PubMed] [Google Scholar]
- 119.Wang J, et al. Target identification of natural and traditional medicines with quantitative chemical proteomics approaches. Pharmacol. Ther. 2016;162:10–22. doi: 10.1016/j.pharmthera.2016.01.010. [DOI] [PubMed] [Google Scholar]
- 120.Ong S-E, et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteomics. 2002;1:376–386. doi: 10.1074/mcp.m200025-mcp200. [DOI] [PubMed] [Google Scholar]
- 121.Abarca ML, et al. Functional and quantitative proteomics using SILAC in cancer research. Junta Dir. 1994;60:31. [Google Scholar]
- 122.Wiese S, Reidegeld KA, Meyer HE, Warscheid B. Protein labeling by iTRAQ: a new tool for quantitative mass spectrometry in proteome research. Proteomics. 2007;7:340–350. doi: 10.1002/pmic.200600422. [DOI] [PubMed] [Google Scholar]
- 123.Deng F, et al. iTRAQ-based quantitative proteomic analysis of esophageal squamous cell carcinoma. Tumor Biol. 2016;37:1909–1918. doi: 10.1007/s13277-015-3840-1. [DOI] [PubMed] [Google Scholar]
- 124.Lau H-T, Suh HW, Golkowski M, Ong S-E. Comparing SILAC-and stable isotope dimethyl-labeling approaches for quantitative proteomics. J. Proteome Res. 2014;13:4164–4174. doi: 10.1021/pr500630a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Ow SY, et al. Quantitative shotgun proteomics of enriched heterocysts from Nostoc sp. PCC 7120 using 8-plex isobaric peptide tags. J. Proteome Res. 2008;7:1615–1628. doi: 10.1021/pr700604v. [DOI] [PubMed] [Google Scholar]
- 126.Wang, J. et al. Drug target identification using an iTRAQ-based quantitative chemical proteomics approach—based on a target profiling study of andrographolide. Methods Enzymol.586, 291–309. [DOI] [PubMed]
- 127.Li Y, et al. iTRAQ-based pharmacoproteomics reveals potential targets of berberine, a promising therapy for ulcerative colitis. Eur. J. Pharmacol. 2019;850:167–179. doi: 10.1016/j.ejphar.2019.02.021. [DOI] [PubMed] [Google Scholar]
- 128.Zybailov B, et al. Statistical analysis of membrane proteome expression changes in Saccharomyces c erevisiae. J. Proteome Res. 2006;5:2339–2347. doi: 10.1021/pr060161n. [DOI] [PubMed] [Google Scholar]
- 129.Vowinckel J, et al. Cost-effective generation of precise label-free quantitative proteomes in high-throughput by microLC and data-independent acquisition. Sci. Rep. 2018;8:1–10. doi: 10.1038/s41598-018-22610-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Asara JM, Christofk HR, Freimark LM, Cantley LC. A label-free quantification method by MS/MS TIC compared to SILAC and spectral counting in a proteomics screen. Proteomics. 2008;8:994–999. doi: 10.1002/pmic.200700426. [DOI] [PubMed] [Google Scholar]
- 131.Megger DA, Bracht T, Meyer HE, Sitek B. Label-free quantification in clinical proteomics. Biochim. Biophys. Acta. 2013;1834:1581–1590. doi: 10.1016/j.bbapap.2013.04.001. [DOI] [PubMed] [Google Scholar]
- 132.Lee Y, et al. ProteoChip: a highly sensitive protein microarray prepared by a novel method of protein immobilization for application of protein-protein interaction studies. Proteomics. 2003;3:2289–2304. doi: 10.1002/pmic.200300541. [DOI] [PubMed] [Google Scholar]
- 133.Haab BB. Advances in protein microarray technology for protein expression and interaction profiling. Curr. Opin. Drug Discov. Dev. 2001;4:116–123. [PubMed] [Google Scholar]
- 134.Xu Q, Lam KS. Protein and chemical microarrays—powerful tools for proteomics. Biomed. Res. Int. 2003;2003:257–266. doi: 10.1155/S1110724303209220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Aguilar-Mahecha A, Hassan S, Ferrario C, Basik M. Microarrays as validation strategies in clinical samples: tissue and protein microarrays. Omics J. Integr. Biol. 2006;10:311–326. doi: 10.1089/omi.2006.10.311. [DOI] [PubMed] [Google Scholar]
- 136.Hett EC, et al. Selectivity determination of a small molecule chemical probe using protein microarray and affinity capture techniques. ACS Comb. Sci. 2016;18:611–615. doi: 10.1021/acscombsci.6b00089. [DOI] [PubMed] [Google Scholar]
- 137.Zhou Y, Liu Z, Rothschild KJ, Lim MJ. Proteome-wide drug screening using mass spectrometric imaging of bead-arrays. Sci. Rep. 2016;6:26125. doi: 10.1038/srep26125. [DOI] [PMC free article] [PubMed] [Google Scholar]