Abstract
Large population-genomic sequencing studies can enable highly-powered analyses of sequence signatures of natural selection. Genome repositories now available for Saccharomyces yeast make it a premier model for studies of the molecular mechanisms of adaptation. We mined the genomes of hundreds of isolates of the sister species S. cerevisiae and S. paradoxus to identify sequence hallmarks of adaptive divergence between the two. From the top hits we focused on a set of genes encoding membrane proteins of the peroxisome, an organelle devoted to lipid breakdown and other specialized metabolic pathways. In-depth population- and comparative-genomic sequence analyses of these genes revealed striking divergence between S. cerevisiae and S. paradoxus. And from transcriptional profiles we detected non-neutral, directional cis-regulatory variation at the peroxisome membrane genes, with overall high expression in S. cerevisiae relative to S. paradoxus. Taken together, these data support a model in which yeast species have differentially tuned the expression of peroxisome components to boost their fitness in distinct niches.
Keywords: population genomics, budding yeast, molecular evolution, peroxisomes
A central goal of large-scale population genomics is to understand how organisms adapt to their environments, by enabling the search for signatures of natural selection in genome sequences. In humans, for example, the 1000 Genomes project has met with some satisfying successes, as candidate adaptive loci have emerged that had not been detected in smaller samples (Field et al. 2016). For organisms with compact genomes and short generation times, sequencing projects of similar scope may lead to an even deeper understanding of the molecular mechanisms of adaptation.
We set out to use budding yeasts as a testbed for large-scale molecular-evolution analysis of population genomes. Despite decades of work in the laboratory, the adaptive history of Saccharomyces in the wild remains poorly understood. Previous sequence surveys have reported relatively few signals of positive selection within or between Saccharomyces species (Kellis et al. 2003; Liti et al. 2009; Elyashiv et al. 2010; Vishnoi et al. 2011; Engle and Fay 2012) (although balancing selection may be more easily detected in this system (Jakobson et al. 2019)). To shed more light on the adaptive history of yeast in the wild, we harnessed a recently published compendium of >1000 S. cerevisiae strain genomes (Peter et al. 2018) to search for loci with marked divergence between this species and its close relative S. paradoxus. From among the hits in this scan, we chose a set of genes that function in the peroxisome membrane for a detailed analysis of divergence in sequence and expression, and an inference of the underlying selective forces.
Methods
Strains and sequences
Data sources for all analyses are collated in Table S3. For population-genomic analyses of coding regions, we collated sequences from 1011 S. cerevisiae strains from (Peter et al. 2018) and S. paradoxus population-genomic data as follows.
For analyses involving European S. paradoxus, we downloaded assemblies and annotation data for S. paradoxus from 12 strains from (Bergström et al. 2014). We used a custom Python script to extract S. paradoxus coding region nucleotide sequences from their annotated location in the genome. For each gene, we aligned the sequences from the strains from a given S. cerevisiae population defined in (Peter et al. 2018) and all 10 European S. paradoxus strains using MUSCLE (Edgar 2004) with the ‘–maxiters’ setting set to 2. We eliminated any sequence consisting of more than 5% gaps, and for the S. cerevisiae populations, we did not analyze any with fewer than 10 strains. We used the alignments as input into the DXY calculation as detailed below.
For analyses involving North American S. paradoxus, we downloaded sequence data for the North American A, B, and C subpopulations from (Durand et al. 2019). We aligned coding region sequences from each to those of vineyard S. cerevisiae from (Peter et al. 2018) using MUSCLE with the ‘–maxiters’ setting set to 2 and eliminated any sequence consisting of more than 5% gaps. We discarded any genes with fewer than 300 S. cerevisiae strain sequences or any genes with missing S. paradoxus sequences in each population, leaving pools of 4285, 3880, and 3868 genes for S. paradoxus subpopulations A, B, and C, respectively.
For population-genomic analyses of promoter regions, we downloaded sequence and annotation data for 10 strains of the wine/European S. cerevisiae promoter regions defined in (Liti et al. 2009). For a given gene we extracted the sequence 500 base pairs upstream of the start codon from each strain, which we defined as the promoter. We likewise extracted the promoter sequence for each gene from each of the 10 European S. paradoxus strains from (Bergström et al. 2014). We then aligned the complete set of promoter sequences from S. cerevisiae and S. paradoxus using MUSCLE with the ‘–maxiters’ setting set to 2 and eliminated any sequence consisting of more than 50% gaps or unspecified nucleotides noted as ‘N.’
For coding-region analyses and, separately, promoter analyses using a given population, we discarded any genes for which we had sequence data for fewer than 80% of the respective strains, yielding a total genomic set numbering between 3430 and 3521 genes. We used the alignments as input into the DXY calculation as detailed below.
For Gene Ontology (GO) enrichment tests as described below, we mapped each gene to its gene ontology groups based on data from geneontology.com (Ashburner et al. 2000). We created a pool of genes from which to resample, eliminating dubious ORFs and those mapped only to the broadest GO terms (Molecular Function, Biological Process, or Cellular Component). Genes were annotated as essential or nonessential based on observations in the S. cerevisiae type strain (Winzeler et al. 1999).
Sequence analyses
Divergence:
We used the alignments of ORF or promoter sequences from a given S. cerevisiae and S. paradoxus population as input into custom Python scripts to run the DXY test (Noor and Bennett 2009), using the formula
where nx is the number of S. cerevisiae strains, ny is the number of S. paradoxus strains, and k is the number of sites with different nucleotides in the same position for each pair of sequences.
To test GO:0005778 for enriched DXY, for a given S. cerevisiae population and S. paradoxus population, we calculated mtrue, the median DXY across all g genes in the GO term. Next, from the complete set of genes in the genome with alignments and GO annotation (see above), we randomly sampled g ORFs or promoters, ensuring the same proportion of essential genes as in the focal term, and calculated the median DXY of this sample. We repeated this procedure 10,000 times and tabulated the proportion of these resampled groups that yielded a median DXY greater than or equal to mtrue, which we used as a one-sided p value assessing the significance of the enrichment of high DXY in the focal term.
Polymorphism:
We used the sequences from the vineyard S. cerevisiae population from (Peter et al. 2018) and, separately, European S. paradoxus from (Bergström et al. 2014), as input into a custom Python script to measure , nucleotide diversity, using the formula
where xi and xj are the frequencies of sequences i and j respectively and is the number of nucleotide differences per site in the sequences. The code iterates through each sequence, pairing it with all other sequences and calculates the number of sites with different alleles divided by the total number of sites, returning the sum across all pairs of sequences. As above, we used a resampling test to calculate a one-sided p value for elevated median nucleotide diversity in the peroxisome membrane genes compared to resampled cohorts.
Phylogenetics:
For phylogenetic analysis of Saccharomyces sensu stricto, we downloaded sequence data for S. cerevisiae strain S288C, S. paradoxus strain CBS432, S. mikatae strain IFO1815, and S. uvarum strain CBS7001 (Christie et al. 2004; Liti et al. 2009; Scannell et al. 2011). We aligned each open reading frame using PRANK (Löytynoja 2014). For phylogenetic analysis including more Saccharomycetaceae species, we downloaded DNA sequences of the 11 closest relatives of S. cerevisiae and S. paradoxus from (Shen et al. 2018) and aligned them using MUSCLE (Edgar 2004) with the ‘–maxiters’ setting set to 2. We used the CodeML module from the PAML package (Yang 2007) to infer branch length for each gene for which we also had data in our population-genomic analyses (see above). The model used an unrooted star tree for analysis of Saccharomyces sensu stricto and the tree from (Shen et al. 2018) for analysis of the larger species set; in each case we assumed a single protein evolutionary rate. Then, for each species set, for each branch in turn we used the nucleotide branch length as input into a resampling approach as above, and we carried out a one-sided significance test for long branch length in the genes of GO:0005778. The ETE toolkit phylogenetic tree viewer was used to create the trees in Figure S1 (Huerta-Cepas et al. 2016).
Gene expression analysis
To analyze differential expression between S. cerevisiae and S. paradoxus, we downloaded RNA-seq data sets from (Schraiber et al. 2013; Artieri and Fraser 2014), which each profiled S. cerevisiae strain S288C, S. paradoxus strain CBS438, and their hybrid after growth in rich glucose medium. In the case of (Artieri and Fraser 2014) for each gene we averaged the reported expression from two replicates for each strain in turn. In each data set, for a given gene, we refer to the total differential expression between the species as the log2 of expression in purebred S. cerevisiae, relative to the analogous quantity in purebred S. paradoxus; cis-regulatory variation is reported as the log2 ratio of allele-specific expression from the two species’ alleles in the hybrid. For each data set, we assessed whether GO:0005778 was enriched for dramatic cis-regulatory variation as follows. We used the measurement of cis-regulatory variation, for each gene for which we also had data in our population-genomic analyses (see above), as input into a one-sided resampling test as above; we then doubled the resulting p-value to yield the result of a two-sided test. We carried out an analogous test for the measurement of total differential expression between the parent species when cultured separately, but we used a one-sided test for elevated expression in S. cerevisiae, under the expectation that the direction of differential expression between the species would conform to that of cis-regulatory divergence. For concision, only the data from (Schraiber et al. 2013) are visualized in Figure 2. For analysis of ribosomal profiling data, we accessed allele-specific ribosomal occupancy in the hybrid and differential ribosomal occupancy between the species when grown separately from (Artieri and Fraser 2014), and analyzed them as above.
In addition, for the analysis of expression profiles of S. cerevisiae and other species in Figure S2, we used measurements of cis-regulatory variation and total differential expression from (Schraiber et al. 2013).
Data availability
Analysis scripts are available at https://github.com/clairedubin/peroxisome_evolution. Supplemental material available at figshare: https://doi.org/10.25387/g3.12124485.
Results
A genome-wide scan for divergence between S. cerevisiae and S. paradoxus:
To identify candidate cases of adaptation in budding yeasts, we first aligned coding sequences from 1011 S. cerevisiae strains (Peter et al. 2018) and 12 strains of S. paradoxus (Bergström et al. 2014). We eliminated gap-rich alignments from further analysis, retaining 3664 genes well-suited for molecular-evolution testing. Next, we reasoned that vineyard/wine S. cerevisiae, as the most deeply sequenced well-defined population of this species currently identified (Peter et al. 2018), would be ideal for exploratory population-genomic analyses. Likewise, the majority of the S. paradoxus isolates in our set (10 of 12) were members of a well-defined European population (Bergström et al. 2014), suitable for population genomics. For each gene, we tabulated variants between all pairs of vineyard S. cerevisiae and European S. paradoxus, and we used these counts to calculate the species divergence metric DXY, a suggestive hallmark of positive selection (Noor and Bennett 2009) (Table S1A).
Among the top-scoring genes in this DXY scan, we noted PEX18, a peroxisome membrane-associated protein importer, and PEX27, a peroxisome biogenesis factor also resident in the membrane (Table S1A). The potential for evolutionary change between budding yeasts in peroxisome factors was of particular interest given the history of divergence in the function of this organelle across fungi more broadly (Gabaldón 2010). We suspected that other peroxisome factors might also have been subject to divergence between S. cerevisiae and S. paradoxus, and as such we examined the complete set of annotated peroxisome membrane genes in yeast (the Gene Ontology term GO:0005778). Indeed, this group was enriched for high DXY in our analysis of vineyard S. cerevisiae and European S. paradoxus (resampling P = 0.0227 with all genes beside PEX18 and PEX27, P = 0.0069 including all genes; Figure 1). On the strength of this signal, we chose the peroxisome membrane gene cohort for further study of molecular change between the species.
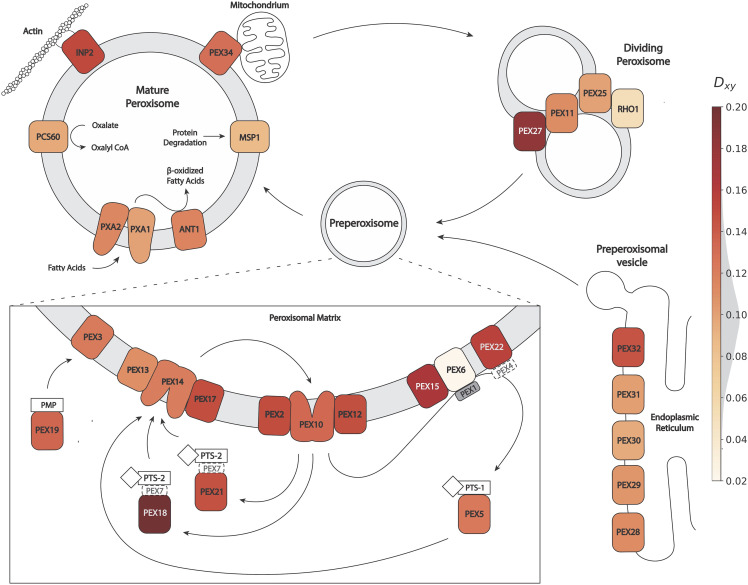
Figure 1.
Peroxisome membrane genes are enriched for high DXY, a population-based metric of sequence divergence, between S. cerevisiae vineyard strains and European S. paradoxus. Each colored element reports DXY at one gene of GO:0005778 (peroxisomal membrane), overlaid on a cartoon localization of the encoded protein during peroxisome fission (top right); de novo peroxisome biogenesis from the endoplasmic reticulum (bottom right); and function of the mature organelle (left). The inset shows machinery for protein recruitment and import into peroxisomes during maturation. Client proteins are shown in white, as diamonds (with peroxisome targeting signals PTS-1 or PTS-2) or a rectangle (a peroxisomal matrix protein, PMP). Symbols with dashed outlines represent proteins not in GO:0005778. Pex1 (gray) is a member of GO:0005778 but did not have sequence data meeting our quality criteria. Hmg1 and Hmg2 also fall into GO:0005778 but are not shown for clarity (DXY = 0.087 and 0.092 respectively). The genome-wide distribution of DXY values is shown superimposed on the color bar legend.
Species-wide signals of divergence at peroxisome membrane genes
We hypothesized that sequence divergence from S. paradoxus at peroxisome membrane genes might not be particular to vineyard S. cerevisiae, but instead could reflect a species-wide trend. To test this, we repeated our analysis of DXY on the peroxisome membrane group using the isolates from each in turn of 23 S. cerevisiae populations defined in (Peter, et al. 2018), as a comparison against European S. paradoxus (Table S1A). We detected elevated DXY in peroxisome membrane genes in each case (Table 1). Next, by a similar logic, we asked whether the trend for divergence from S. cerevisiae would be upheld in analyses of S. paradoxus from localities beside Europe. Using three North American S. paradoxus populations (Durand et al. 2019), we again found striking divergence in the peroxisome membrane gene group (Table S1B). We conclude that the sequences of peroxisome membrane genes are strikingly distinct between the two species, regardless of which isolates and populations we analyze.
Table 1. DXY enrichment among peroxisomal membrane genes.
| POPULATION | MEDIAN DXY FOR GO:0005778 | pa |
|---|---|---|
| French Guiana, human | 0.1104 | 0.0118 |
| Ale beer | 0.1106 | 0.0021 |
| West African cocoa | 0.1114 | 0.0043 |
| African palm wine | 0.1112 | 0.0068 |
| European wine | 0.1108 | 0.0069 |
| European wine subclade 1 | 0.1107 | 0.0073 |
| European wine subclade 2 | 0.1107 | 0.0083 |
| European wine subclade 3 | 0.1107 | 0.0089 |
| European wine subclade 4 | 0.1107 | 0.0089 |
| Ecuadorean | 0.1097 | 0.0145 |
| North American oak | 0.1096 | 0.0116 |
| Asian islands | 0.1116 | 0.0060 |
| Sake | 0.1112 | 0.0068 |
| Asian fermentation | 0.1113 | 0.0071 |
| Alpechin | 0.1125 | 0.0040 |
| Brazilian bioethanol | 0.1107 | 0.0044 |
| French dairy | 0.1109 | 0.0047 |
| African beer | 0.1119 | 0.0031 |
| Mosaic beer | 0.1107 | 0.0064 |
| Mixed origin | 0.1096 | 0.0066 |
| Mosaic region 1 | 0.1101 | 0.0083 |
| Mosaic region 2 | 0.1107 | 0.0051 |
| Mosaic region 3 | 0.1105 | 0.0071 |
Each row reports the enrichment among peroxisomal membrane genes (GO:0005778) for elevated DXY, a population-based metric of sequence divergence, in a comparison of the indicated S. cerevisiae population and European S. paradoxus. In a given population, the median DXY across all genes in the genome is between 0.0971 and 0.0988.
Resampling-based significance from a test for elevated DXY using strains of the indicated S. cerevisiae population.
Directional cis-regulatory divergence at peroxisome membrane genes
We next aimed to investigate the possible molecular mechanisms of peroxisome membrane gene evolution. We found no detectable signal in tests of non-neutral amino acid substitution rates (data not shown). As such, we instead hypothesized that evolution could have tuned the regulation of this gene cohort differently between S. cerevisiae and S. paradoxus. To evaluate this, we made use of our laboratory’s test for cis-regulatory variants that drive expression in the same direction across unlinked genes of a pathway, a pattern unlikely under neutrality (Bullard et al. 2010). We applied this scheme to two independently collected profiles of cis-regulatory divergence between S. cerevisiae and S. paradoxus in rich glucose medium, each measured from allele-specific expression in the interspecies hybrid (Schraiber et al. 2013; Artieri and Fraser 2014). The results revealed significant directional cis-regulatory divergence in the peroxisomal membrane group (resampling P = 0.0062 and 0.0320, respectively, from the two data sources), with the S. cerevisiae allele of each gene tending to drive higher expression relative to the S. paradoxus allele (Figure 2).
Figure 2.
S. cerevisiae expresses peroxisome membrane genes more highly than does S. paradoxus, as a product in part of cis-regulatory changes. Each cell reports expression, as a ratio between S. cerevisiae and S. paradoxus, of the indicated peroxisome membrane gene from GO:0005778. Total, expression measured in purebred species; cis, expression from the indicated species’ allele in a diploid interspecific hybrid, reflecting effects of cis-regulatory divergence.
We also inspected profiles of total expression divergence between the species—the product of DNA sequence variants acting both in cis and in trans, as measured from expression in purebreds. Using this quantity we detected directional change in peroxisome membrane genes with higher expression in S. cerevisiae, as was true in our analysis of cis-regulatory variation (Figure 2; resampling P = 0.0035 and 0.0275, respectively, from the two data sources). Furthermore, in ribosomal profiling data (Artieri and Fraser 2014), we detected higher ribosomal occupancy of peroxisomal membrane genes in cultures of S. cerevisiae relative to S. paradoxus, attesting to the relevance of the mRNA expression difference between the species in terms of protein abundance (though no such imbalance could be detected between the alleles in the hybrid; resampling P = 0.0171 and 0.7754, respectively). We conclude that, with growth conditions held constant, S. cerevisiae and S. paradoxus are hard-wired to express peroxisome membrane genes differently, as a function in part of a suite of cis-regulatory changes at the unlinked gene loci.
We reasoned that many of the elements underlying cis-acting expression variation between the species in our gene cohort would likely fall in promoters. Yet the initial evidence of sequence divergence that we had noted for the peroxisome membrane genes was in our DXY test applied to coding regions (Table 1 and Table S2). We expected that, if such a signal were the consequence of linked selection on adaptive variants in promoters, the latter would mirror what we had detected in open reading frames, and exhibit significant divergence. Consistent with this prediction, we detected high DXY in the promoters of the peroxisome membrane genes, in a comparison of the European S. cerevisiae and European S. paradoxus populations (resampling P = 0.0019). Thus at a sizeable window centered on a given gene in the cohort, encompassing both the upstream region and the open reading frame, strains of the two species are robustly and recurrently different from one another in terms of sequence. Together, this trend and our expression-based test support a model in which the regulation of the peroxisome membrane genes has been subject to different selective pressures in S. cerevisiae and S. paradoxus.
Inferring a history of evolutionary volatility at peroxisome membrane genes
In principle, divergence in peroxisome membrane genes could reflect a history in which an ancestral state was established before the divergence of S. cerevisiae and S. paradoxus, followed by stabilizing selection in one species and directional selection in the other. Alternatively, the peroxisome membrane gene group may have been evolutionarily more volatile, undergoing changes in multiple Saccharomycete lineages. To investigate this, we pursued phylogenetic analyses of DNA sequence across species, using a small set of Saccharomyces sensu stricto species (Christie et al. 2004; Liti et al. 2009; Scannell et al. 2011) and, separately, a larger set of Saccharomycetaceae (Shen et al. 2018). These tests revealed an excess of sequence variation in the peroxisome membrane genes along each branch of the phylogeny, significantly so in most cases (Figure S1). Likewise, expression profiles in rich glucose medium (Schraiber et al. 2013) revealed a regulatory program in the distant relative S. uvarum that was distinct from the expression levels of either S. cerevisiae or S. paradoxus (Figure S2). The latter suggests an evolutionary event in the S. uvarum lineage unrelated to the changes between our two focal species. These data preclude the straightforward inference of a state of the peroxisome membrane group ancestral to Saccharomyces sensu stricto or those species farther out, and reflect a likely history of evolutionary volatility over millions of years.
Discussion
In this work, we have harnessed the wealth of population-genomic data available for S. cerevisiae (Peter et al. 2018) in tests of divergence between this species and S. paradoxus. To date, hits from such scans have been at a premium, which may reflect the limited statistical power associated with smaller yeast genome cohorts (Kellis et al. 2003; Elyashiv et al. 2010; Vishnoi et al. 2011; Engle and Fay 2012; Bergström et al. 2014). Indeed, most studies of adaptive loci in Saccharomyces have found their gene candidates by methods other than molecular-evolution surveys (Will et al. 2010; Engle and Fay 2012; Martin et al. 2012; Fraser et al. 2012; Roop et al. 2016; Weiss et al. 2018; Duan et al. 2019). By contrast, the strong molecular-evolution signal we report here attests to the power of the deep population-sampling approach.
Our analysis has centered on a pattern of sequence divergence from S. paradoxus across hundreds of S. cerevisiae isolates in peroxisome membrane genes. Given that we have used the absolute divergence measure DXY rather than metrics that normalize divergence by within-species polymorphism, our conclusions are not contingent on any effects of the latter (and the peroxisome membrane gene group was not an outlier with respect to polymorphism in any case; resampling P = 0.68 and P = 0.25 for vineyard S. cerevisiae and European S. paradoxus, respectively). Rather, we interpret the elevated DXY signal as an indicator of a history of positive selection on the gene cohort in one or both species, although strictly speaking it could also be consistent with balancing selection in the ancestor (Guerrero and Hahn 2017).
Our findings dovetail with the widespread divergence in peroxisomal function across the fungal kingdom in general, which has perhaps been facilitated by the ease of gains and losses of peroxisomal localization signals in protein sequences (Yanagida et al. 2015). Apart from its conserved role in fatty acid oxidation, the peroxisome has evolved in particular fungi to carry out methanol catabolism; antibiotic, siderophore, and biotin biosynthesis; and a wound-healing function in filamentous species (Gabaldón 2010; Maruyama and Kitamoto 2013; Lim and Keller 2014). However, our data do not afford any insight into possible biochemical changes in the function of this organelle between S. cerevisiae and S. paradoxus. Instead, the strongest mechanistic inference we can make derives from our analysis of gene expression: S. cerevisiae and S. paradoxus have differentially tuned when and how peroxisome membrane genes come on, in a pattern inconsistent with neutrality as a driving force. It is tempting to speculate that the high expression by S. cerevisiae could reflect a historical need to boost fatty acid metabolism in its niche. As no conclusions about niche or ecology can be drawn from our genomics approach, future ecological and phenotypic analyses relevant to the peroxisome will be of prime interest.
A key conclusion from our findings is that variants in many peroxisome membrane genes—with regulatory changes at promoters likely of particular importance—came together to build an evolutionary innovation as S. cerevisiae and S. paradoxus diverged. Assuming that these changes represent an adaptation to an environmental challenge, we can infer that no single Mendelian locus was sufficient to solve the ecological problem at hand. This complexity would be consistent with the deleterious effects of overexpressing individual peroxisome membrane genes on their own in yeast (Elgersma et al. 1997; Sopko et al. 2006). As population sequencing data sets grow, such instances of polygenic adaptation (Pritchard and Di Rienzo 2010) may prove to be the norm, in systems from microbes to mammals.
Acknowledgments
The authors thank Faisal AlZaben for assistance with PAML; John Dueber, Parbir Grewal, Jennifer Samson, and members of the Brem lab for helpful discussions; and Adam Arkin for his generosity with space and resources. This work was funded by NIH R01 GM120430-A1 to R.B.B.
Footnotes
Supplemental material available at figshare: https://doi.org/10.25387/g3.12124485.
Communicating editor: A. Rokas
Literature Cited
- Artieri C. G., and Fraser H. B., 2014. Evolution at two levels of gene expression in yeast. Genome Res. 24: 411–421. 10.1101/gr.165522.113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ashburner M., Ball C. A., Blake J. A., Botstein D., Butler H. et al. , 2000. Gene Ontology: tool for the unification of biology. Nat. Genet. 25: 25–29. 10.1038/75556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bergström A., Simpson J. T., Salinas F., Barré B., Parts L. et al. , 2014. A high-definition view of functional genetic variation from natural yeast genomes. Mol. Biol. Evol. 31: 872–888. 10.1093/molbev/msu037 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bullard J. H., Mostovoy Y., Dudoit S., and Brem R. B., 2010. Polygenic and directional regulatory evolution across pathways in Saccharomyces. Proc. Natl. Acad. Sci. USA 107: 5058–5063. 10.1073/pnas.0912959107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Christie K. R., Weng S., Balakrishnan R., Costanzo M. C., Dolinski K. et al. , 2004. Saccharomyces Genome Database (SGD) provides tools to identify and analyze sequences from Saccharomyces cerevisiae and related sequences from other organisms. Nucleic Acids Res. 32: D311–D314. 10.1093/nar/gkh033 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duan S.-F., Shi J.-Y., Yin Q., Zhang R.-P., Han P.-J. et al. , 2019. Reverse Evolution of a Classic Gene Network in Yeast Offers a Competitive Advantage. Curr. Biol. 29: 1126–1136.e5. 10.1016/j.cub.2019.02.038 [DOI] [PubMed] [Google Scholar]
- Durand É., Gagnon-Arsenault I., Hallin J., Hatin I., Dubé A. K. et al. , 2019. Turnover of ribosome-associated transcripts from de novo ORFs produces gene-like characteristics available for de novo gene emergence in wild yeast populations. Genome Res. 29: 932–943. 10.1101/gr.239822.118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edgar R. C., 2004. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32: 1792–1797. 10.1093/nar/gkh340 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elgersma Y., Kwast L., van den Berg M., Snyder W. B., Distel B. et al. , 1997. Overexpression of Pex15p, a phosphorylated peroxisomal integral membrane protein required for peroxisome assembly in S.cerevisiae, causes proliferation of the endoplasmic reticulum membrane. EMBO J. 16: 7326–7341. 10.1093/emboj/16.24.7326 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elyashiv E., Bullaughey K., Sattath S., Rinott Y., Przeworski M. et al. , 2010. Shifts in the intensity of purifying selection: an analysis of genome-wide polymorphism data from two closely related yeast species. Genome Res. 20: 1558–1573. 10.1101/gr.108993.110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engle E. K., and Fay J. C., 2012. Divergence of the Yeast Transcription Factor FZF1 Affects Sulfite Resistance. PLoS Genet 8: e1002763 10.1371/journal.pgen.1002763 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Field Y., Boyle E. A., Telis N., Gao Z., Gaulton K. J. et al. , 2016. Detection of human adaptation during the past 2000 years. Science 354: 760–764. 10.1126/science.aag0776 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fraser H. B., Levy S., Chavan A., Shah H. B., Perez J. C. et al. , 2012. Polygenic cis-regulatory adaptation in the evolution of yeast pathogenicity. Genome Res. 22: 1930–1939. 10.1101/gr.134080.111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gabaldón T., 2010. Peroxisome diversity and evolution. Philos. Trans. R. Soc. Lond. B Biol. Sci. 365: 765–773. 10.1098/rstb.2009.0240 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guerrero R. F., and Hahn M. W., 2017. Speciation as a sieve for ancestral polymorphism. Mol. Ecol. 26: 5362–5368. 10.1111/mec.14290 [DOI] [PubMed] [Google Scholar]
- Huerta-Cepas J., Serra F., and Bork P., 2016. ETE 3: Reconstruction, Analysis, and Visualization of Phylogenomic Data. Mol. Biol. Evol. 33: 1635–1638. 10.1093/molbev/msw046 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jakobson C. M., She R., and Jarosz D. F., 2019. Pervasive function and evidence for selection across standing genetic variation in S. cerevisiae. Nat. Commun. 10: 1222 10.1038/s41467-019-09166-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kellis M., Patterson N., Endrizzi M., Birren B., and Lander E. S., 2003. Sequencing and comparison of yeast species to identify genes and regulatory elements. Nature 423: 241–254. 10.1038/nature01644 [DOI] [PubMed] [Google Scholar]
- Lim F. Y., and Keller N. P., 2014. Spatial and temporal control of fungal natural product synthesis. Nat. Prod. Rep. 31: 1277–1286. 10.1039/C4NP00083H [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liti G., Carter D. M., Moses A. M., Warringer J., Parts L. et al. , 2009. Population genomics of domestic and wild yeasts. Nature 458: 337–341. 10.1038/nature07743 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Löytynoja A., 2014. Phylogeny-aware alignment with PRANK. Methods Mol. Biol. 1079: 155–170. 10.1007/978-1-62703-646-7_10 [DOI] [PubMed] [Google Scholar]
- Martin H. C., Roop J. I., Schraiber J. G., Hsu T. Y., and Brem R. B., 2012. Evolution of a membrane protein regulon in Saccharomyces. Mol. Biol. Evol. 29: 1747–1756. 10.1093/molbev/mss017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maruyama J.-I., and Kitamoto K., 2013. Expanding functional repertoires of fungal peroxisomes: contribution to growth and survival processes. Front. Physiol. 4: 177 10.3389/fphys.2013.00177 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noor M. F., and Bennett S. M., 2009. Islands of speciation or mirages in the desert? Examining the role of restricted recombination in maintaining species. Heredity 103: 439–444. 10.1038/hdy.2009.151 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peter J., De Chiara M., Friedrich A., Yue J.-X., Pflieger D. et al. , 2018. Genome evolution across 1,011 Saccharomyces cerevisiae isolates. Nature 556: 339–344. 10.1038/s41586-018-0030-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pritchard J. K., and Di Rienzo A., 2010. Adaptation - not by sweeps alone. Nat. Rev. Genet. 11: 665–667. 10.1038/nrg2880 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roop J. I., Chang K. C., and Brem R. B., 2016. Polygenic evolution of a sugar specialization trade-off in yeast. Nature 530: 336–339. 10.1038/nature16938 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scannell D. R., Zill O. A., Rokas A., Payen C., Dunham M. J., et al. , 2011. The Awesome Power of Yeast Evolutionary Genetics: New Genome Sequences and Strain Resources for the Saccharomyces sensu stricto Genus. G3 (Bethesda) 1: 11–25. 10.1534/g3.111.000273 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schraiber J. G., Mostovoy Y., Hsu T. Y., and Brem R. B., 2013. Inferring evolutionary histories of pathway regulation from transcriptional profiling data. PLOS Comput. Biol. 9: e1003255 10.1371/journal.pcbi.1003255 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen X.-X., Opulente D. A., Kominek J., Zhou X., Steenwyk J. L. et al. , 2018. Tempo and Mode of Genome Evolution in the Budding Yeast Subphylum. Cell 175: 1533–1545.e20. 10.1016/j.cell.2018.10.023 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sopko R., Huang D., Preston N., Chua G., Papp B. et al. , 2006. Mapping pathways and phenotypes by systematic gene overexpression. Mol. Cell 21: 319–330. 10.1016/j.molcel.2005.12.011 [DOI] [PubMed] [Google Scholar]
- Vishnoi A., Sethupathy P., Simola D., Plotkin J. B., and Hannenhalli S., 2011. Genome-wide survey of natural selection on functional, structural, and network properties of polymorphic sites in Saccharomyces paradoxus. Mol. Biol. Evol. 28: 2615–2627. 10.1093/molbev/msr085 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss C. V., Roop J. I., Hackley R. K., Chuong J. N., Grigoriev I. V. et al. , 2018. Genetic dissection of interspecific differences in yeast thermotolerance. Nat. Genet. 50: 1501–1504. 10.1038/s41588-018-0243-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Will J. L., Kim H. S., Clarke J., Painter J. C., Fay J. C. et al. , 2010. Incipient balancing selection through adaptive loss of aquaporins in natural Saccharomyces cerevisiae populations. PLoS Genet. 6: e1000893 10.1371/journal.pgen.1000893 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winzeler E. A., Shoemaker D. D., Astromoff A., Liang H., Anderson K. et al. , 1999. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285: 901–906. 10.1126/science.285.5429.901 [DOI] [PubMed] [Google Scholar]
- Yanagida H., Gispan A., Kadouri N., Rozen S., Sharon M. et al. , 2015. The Evolutionary Potential of Phenotypic Mutations. PLoS Genet. 11: e1005445 10.1371/journal.pgen.1005445 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang Z., 2007. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24: 1586–1591. 10.1093/molbev/msm088 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Analysis scripts are available at https://github.com/clairedubin/peroxisome_evolution. Supplemental material available at figshare: https://doi.org/10.25387/g3.12124485.