Abstract
Qinhuangdao coastal area is an important mariculture area in North China. Microbial communities play an important role in driving biogeochemical cycle and energy flow. It is necessary to identify the microbial communities and their functions in the coastal mariculture area of Qinhuangdao. In this study, the microbial community compositions and their metabolic potentials in the sediments and their surrounding seawaters of Qinhuangdao mariculture area were uncovered by the 16S rRNA gene amplicon sequencing and metagenomic shotgun sequencing approaches. The results of amplicon sequencing showed that Gammaproteobacteria and Alphaproteobacteria were predominant classes. Our datasets showed a clear shift in microbial taxonomic groups and the metabolic pathways in the sediments and surrounding seawaters. Metagenomic analysis showed that purine metabolism, ABC transporters, and pyrimidine metabolism were the most abundant pathways. Genes related to two-component system, TCA cycle and nitrogen metabolism exhibited higher abundance in sediments compared with those in seawaters. The presence of cadmium-resistant genes and ABC transporters suggested the ability of microorganisms to resist the toxicity of cadmium. In summary, this study provides comprehensive and significant differential signatures in the microbial community and metabolic pathways in Qinhuangdao mariculture area, and can develop effective microbial indicators to monitor mariculture area in the future.
Introduction
The microbial communities are widely distributed, and they play a vital role in marine ecosystems. The functions of microbial communities in marine environment balance various important biogeochemical cycles, including degradation of pollutants, nutrient cycles, and organic mineralization [1,2]. To date, ecosystem monitoring technologies are based on the monitoring of total biomass of microorganisms [3], algae [4,5], and macrozoobenthic species [6]. Studies have shown that the microbial diversity and metabolisms depend on the local environment, and are rapid and sensitive to environmental changes [7–9]. Therefore, responses of microbial communities associated with environmental changes can be used as microbial “indicators”. The identification of the core microbiome of a given environment is critical to define the health status of the environment, and to predict the response of microbial communities to anthropogenic changes [10,11]. However, functions of microbial communities are largely ignored in biological monitoring methods [12]. A recent study recommended that the structural or functional prokaryotic variables (biodiversity, abundance and metabolism) should be included in future implementations of the EU Marine Strategy Framewok Directive 2008/56/EC (MSFD) [13].
Metagenomics has been successfully applied to investigate microbial diversity, adaptation, evolution, and function [14]. The profiling of microbial communities can be revealed by high-throughput sequencing of targeted PCR amplification [15]. For example, through 16S rRNA genes sequencing, the microbial community variation can provide important baseline understandings of the microbial ecology and health assessment of the marine ecosystems [16,17]. Metabolism of microbial communities can be investigated by metagenomics without any prior knowledge. The metagenomic results are important evidences to measure some specific ecological processes. For instance, the metagenomic analysis revealed the metabolic versatility of microorganisms and their roles in biogeochemical cycles including nitrogen, carbon, and sulfur cycles at the Yap Trench of the western Pacific [18]. Metagenomics was also applied to study the upper and core regions of oxygen minimum zones in Arabian Sea, and confirmed the genomic potentials of active nitrogen cycle [19]. The metagenomic results revealed that the metabolic capacity of microorganisms might be weaken by contamination in mangrove sediments and the increase of greenhouse gas emission might be induced [20]. Recently, combining of 16S rRNA amplicon sequencing and metagenomic shotgun analysis revealed that chemical pollutants severely affected microbial structures and functions in sediments [21]. Therefore, high-throughput sequencing of amplicons and metagenomic shotgun sequencing can provide new perspective of complexity of microbial communities and their functional trait [22].
Bohai Sea is located in Northeast China and has been influenced by anthropogenic factors in recent years [23,24]. Previous studies revealed that the pollution in Bohai Sea is serious in coastal areas and that situation can be worse if no protection procedures are implemented [25]. The mariculture activities of Bay scallops (Argopecten irradians) have been conducted for over 30 years, and is a major industry that supports the economy of Qinhuangdao, which is a coastal city of Bohai Sea [17]. Hydrargyrum, Cadmium, and Plumbum are the predominant metal pollutants commonly found in the Bohai Sea, and their concentration in some areas were 12 to 150 times higher than the background concentrations [25]. In addition, Cadmium in fish species from Qinhuangdao had higher concentrations than that from the Pearl River Estuary, Zhejiang and Haikou of China [26].
Mitigating the threat of human activities to marine environment is a big challenge, thus it is necessary to develop new methods to monitor and assess the impact of human activities on marine sediments and the surrounding seawater columns in the mariculture areas, which could help us understand the restoration potentials of the marine ecosystems[27]. Previous studies have characterized the bacterioplankton communities of natural seawaters in Bohai Sea, and investigated the archaeal diversity and community structures in this area [16,17]. However, the lack of understanding of the metabolic capability of microorganisms in the sediments and surrounding seawaters of the Qinhuangdao mariculture area hinders the prediction of the capability of the microorganisms in responding to changes of environmental conditions. To our knowledge, this would be the first study to characterize the microbial communities with their metabolic potentials in Qinhuangdao mariculature area.
In this study, we collected two surface sediments samples and two surrounding seawater samples in Qinhuangdao mariculture area. Sequencing of 16S rRNA gene amplicons was used to identify the microbial communities. Metagenomic analysis was carried out to reveal the metabolic potentials. The goal of this research was to: (1) investigate the compositions of microbial communities in the sediments and surrounding seawaters; (2) provide the yet unexplored representative characteristic of metabolic potentials in Qinhuangdao aquaculture area; (3) identify the differences of metabolic potentials and key genes associated microbial communities in sediments and seawaters. The results of this study could help us understand the microbial communities and their metabolic patterns in Qinhuangdao mariculture area.
Materials and methods
Samples collection and environmental parameters description
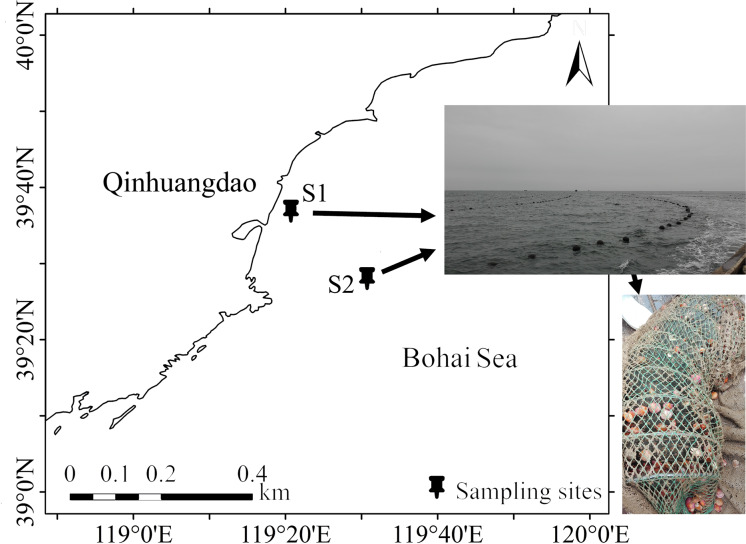
The field site access was approved by the departments of fishery administration. The seawaters were sampled on July 26, 2017 in two stations, S1 (39˚36’53” N, 119˚20’43” E) and S2 (39˚34’43” N, 119˚25’32”E) in Qinhuangdao mariculture area that farmed A. irradians, from 8 m and 10 m depth, respectively. (Fig 1). Seawaters were sampled in triplicate at each station, then homogeneously mixed prior to filtration, five litres of the pooled water were filtered through a 0.2 μm filter membrane (Millipore, Billerica, USA). Five grams of surface sediments were concomitantly collected at the S1 and S2 locations using a grab sampler made of stainless steel, and the sediment samples were named as S1S and S2S respectively. The sampled sediments were stored in sterile 5 ml storage tubes. All the filter membranes and sediments were stored at -80˚C for further analysis. Physicochemical properties of the seawater and sediment samples were determined. In detail, the values of parameters including temperature, pH, dissolved oxygen (DO), salinity, turbidity, electrical conductivity (EC), total dissolved solids (TDS), oxidation-reduction potential (ORP), total nitrogen (TN), total phosphorus (TP), total organic carbon (TOC) and total sulfur (TS) content in the seawater or sediments were determined (S1 Table).
Fig 1. Map of study areas showing geographical locations, raft cultivations of A.irradians, and mesh cage used for cultivation.
ArcGIS 10.1 software (http://www.esri.com/software/arcgis) was used to develop the map.
DNA extraction, amplicon sequencing and analysis
The MoBio PowerSoil® DNA Isolation Kit (MO BIO Laboratories, USA) was used to extract the DNA from 0.2-μm filter membrane or 5 g of sediment. DNA concentration and quality were determined using NanoDrop spectrophotometer (ThermoScientific, Wilmington, DE, USA). The DNA extracts were used as the templates in PCR. The V4 variable region of the 16S rRNA genes was amplified by primers 16S V4-515F (5’-GTGCCAGCMGCCGCGGTAA-3’) and 16S V4-806R (5’-GGACTACHVGGGTWTCTAAT-3’) [28]. The 16S rRNA gene amplicons were prepared and subsequently sequenced with Illumina HiSeq high-throughput sequencing (2╳250 bp). Using the Qiime2 pipeline, effective tags were obtained through analyzing the sequencing output [29]. Operational Taxonomic Units (OTUs) were clustered by the effective tags at 97% sequence similarity. The analysis of taxonomic annotation was performed based on SILVA v132 reference database [30]. Relative abundance was represented by the percentage of each taxon in each sample to compare their microbial communities in phyla and class levels. The difference between the four communities was investigated by principal component analysis (PCA) using R software packages by considering the relative abundance of all microbial genera.
Metagenomic shotgun sequencing and analysis
The DNA extraction method was the same and described above. DNA was used for library preparation and detection. Qualified libraries were sequenced using Illumina with the paired-end length of 2╳150 bp, and the final raw data was used for bioinformatic analysis. The raw data was filtered by trimming adapter sequences and low quality reads (quality value less than 38 ≥40bp, and ≥10% N containing reads). The clean reads were de novo assembled for each DNA sample independently using MEGAHIT (—presets meta-large).The unassembled reads in each sample were merged together and assembled [31]. ORFs were predicted from the contigs of each sample as well as the contigs from the merged assembly using the MetaGeneMark software. Predicted genes were subjected for de-redundancy analysis using CD-HIT. Clean reads were mapped to representative genes by Bowtie 2, the number of reads mapped to each gene were used for the calculation of gene abundance [32,33]. Genes of which reads number in each sample was no more than 2 were filtered, thus the unigenes were obtained and used for subsequent analysis. The predicted protein sequences were searched by BLASTP with e-value≤1╳10−5 in the Kyoto Encyclopedia of Genes and Genomes (KEGG) database [34]. The annotated genes were assigned into KEGG pathways. Based on the functional abundance of level 3 KEGG pathways, PCA was used to cluster the samples. According to PCA clustering results, Metastat statistical analysis and metabolic pathway comparison analysis were carried out to distinguish functional composition differences.
Nucleotide sequence accession numbers
Metagenomic datasets are available online at NCBI, BioSample accessions: SAMN13744940, SAMN13744941, SAMN13744942, SAMN13744943 for Seawater1 (S1), Sediment1 (S1S), Seawater2 (S2), and Sediment2 (S2S) samples respectively. The 16S rRNA datasets are available online at NCBI, BioSample accessions: SAMN13752121, SAMN13752122, SAMN13752123, SAMN13752124 for (Seawater sample1) S1, (Sediment sample1) S1S, (Seawater sample2) S2, and (Sediment sample2) S2S samples respectively.
Results
Taxonomic characterization of the microbial communities
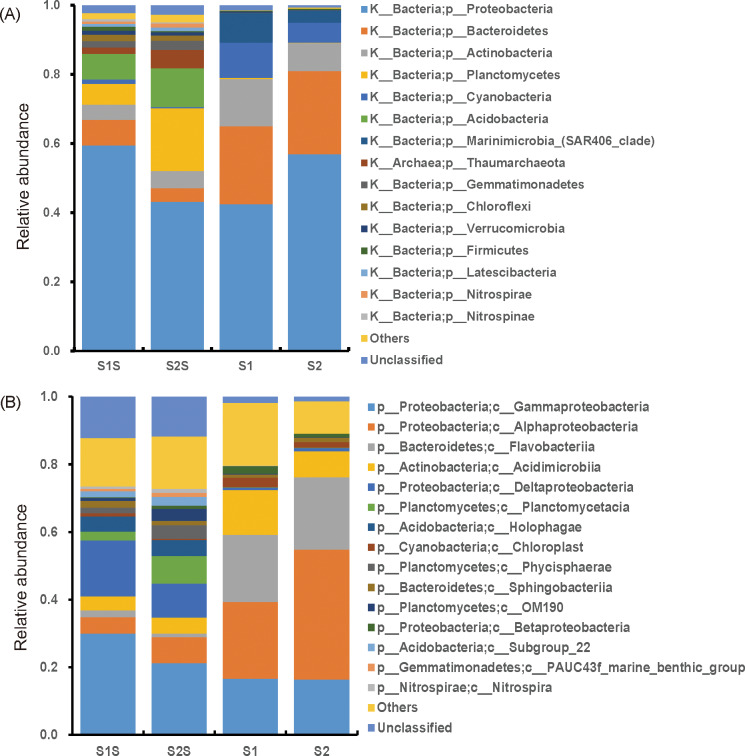
Based on the sequencing of 16S rRNA genes, 61,799, 88,645, 91,113 and 86,122 clean reads were obtained from the S1S, S2S, S1 and S2 samples, respectively. A total of 4,784 OTUs were found in the sediment samples while only 860 OTUs were in the seawater samples. Specifically, the numbers of OTUs detected in each sample were 3,530, 4,271, 709, and 617 in S1S, S2S, S1, and S2 samples, respectively. The relative abundance of the phyla and classes in different samples are shown in Fig 2. Proteobacteria, Bacteroidetes, and Actinobacteria are the three most dominant phyla, accounting for 49.69%, 15.04%, and 8.08%, respectively (Fig 2A). At class level, the top three dominant classes were Gammaproteobacteria (20.35%), Alphaproteobacteria (19.36%), and Flavobacteriia (11.85%). Specifically, Gammaproteobacteria and Alphaproteobacteria are both the sub-divisions of Proteobacteria. Gammaproteobacteria was the dominant class of in sediments (25.6%) while the dominant class in seawaters was Alphaproteobacteria (30.5%) (Fig 2B).
Fig 2.
High-throughput sequencing of 16S rRNA gene revealing the relative abundance of phylum (A) and class (B) in the sediment (S1S and S2S) and seawater (S1 and S2) samples. The top 15 phyla or classes with the largest relative abundance were presented and the rest phyla or classes were set as others or unclassified.
Functional profiles of microbial communities from KEGG functional analysis
After removing low quality reads, a total of 51 Gbp clean data was generated for the four samples, with 12.9 Gbp for S1, 12.3 Gbp for S2, 12.3 Gbp for S1S, and 13.2 Gbp for S2S. After assembling of the clean reads, there were 435,438 contigs with the average length of 974 bp. Venn diagram showed the number of identified genes and their distribution in sediments and seawaters (S1 Fig). There were 280,552 genes common for sediments and seawaters, while there were 666,483 and 1,024,221 genes for sediments and seawaters, respectively (S1 Fig). Functional properties of these genes were investigated using KEGG Orthology (KO). Of the identified protein-coding genes, 45.9% were assigned against KO database. In detail, the number ratios of genes involving in organismal systems, metabolism, human diseases, genetic information processing, environmental information processing, and cellular processes were 1.65%, 31.26%, 3.19%, 5.86%, 4.25%, and 3.80% (S2 Fig). Besides, the relative abundances of reads assigned to level 1 and level 2 KEGG pathways in the four samples were showed in S3 Fig.
Differential analysis of microbial taxonomic profiles and metabolic pathways
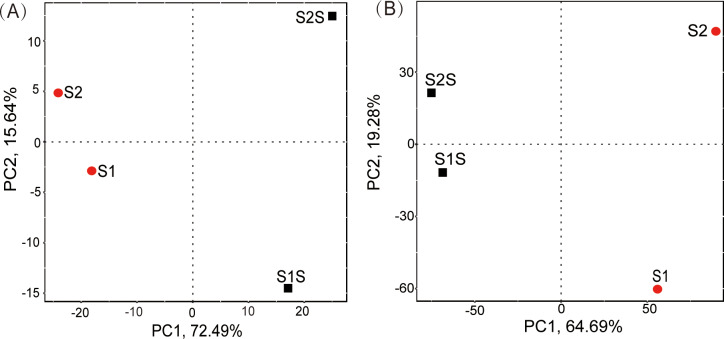
PCA was used to statistically explore and visualize the difference between the different microbial communities (Fig 3). Two seawater samples and two sediment samples were clustered together based on the microbial compositions at the genus level (Fig 3A). PCA was also performed to cluster the metagenomic data based on their KEGG annotations. Obvious differential distributions of KEGG pathways between the sediments and seawaters were observed (Fig 3B). Therefore, there are not only differences in microbial taxonomic profile but also significant variations in the potential metabolic pathways.
Fig 3.
PCA analysis results based on (A) genus abundance from 16S rRNA gene amplicon sequencing and (B) functional abundance of level 3 KEGG pathways. The red and black dots are sediment and seawater samples, respectively. The abscissa and the ordinate represent the first principal component (PC1) and the second principal component (PC2) respectively. The percentage represents the contribution of the principal components to the samples difference.
Top abundant pathways from KEGG functional analysis
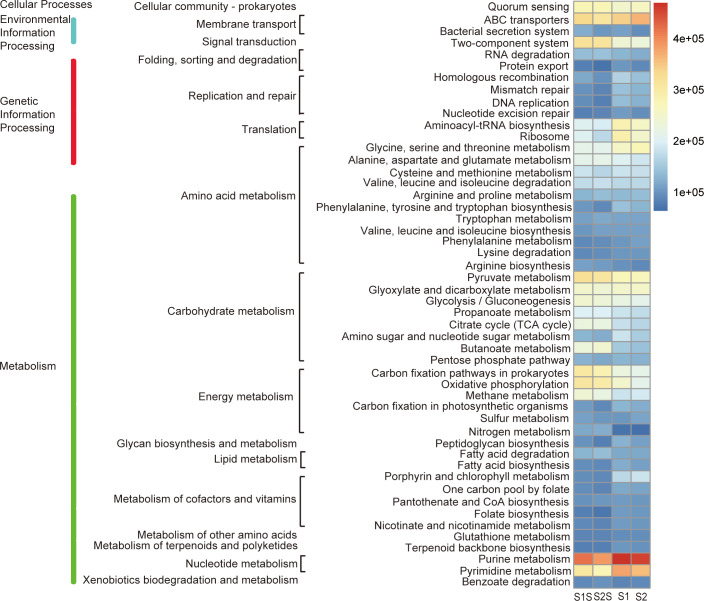
The top three abundant pathways were purine metabolism, ABC transporters and pyrimidine metabolism. Furthermore, the purine metabolism and pyrimidine metabolism belong to nucleotide metabolism. ABC transporters belong to membrane transport pathway (Fig 4).
Fig 4. Heatmap profile showing the relative abundance of top 50 most abundant metabolic pathways in the sediment and seawater samples.
The three columns of descriptions from left to right represent the level 1, level 2, and level 3 KEGG pathways.
Purine metabolism
A total of 208 genes were involved in purine metabolism in the sediment and seawater samples. The highest abundance gene was nrdAE that encoding ribonucleoside-diphosphate reductase alpha chain. In addition, rpoB, rpoC, PFAS, polA and dnaE that encoding DNA-directed RNA polymerase subunit beta, DNA-directed RNA polymerase subunit beta', phosphoribosylformylglycinamidine synthase, DNA polymerase I, and DNA polymerase III subunit alpha also exhibited high abundance.
ABC transporters
323 genes annotated as ABC transporters were retrieved in the four metagenomes. Of these genes, genes (livF, livG, livK, livM, and livH), which belonged to branched-chain amino acid transport system, showed the highest abundances.
Pyrimidine metabolism
A total of 143 genes involved in pyrimidine metabolism were identified. Similar to purine metabolism, nrdAE was also the highest abundant gene, followed by rpoC, rpoB, CPA2 (encoding carbamoyl-phosphate synthase large subunit), dnaE, and polA.
Differentially abundant pathways from KEGG functional analysis
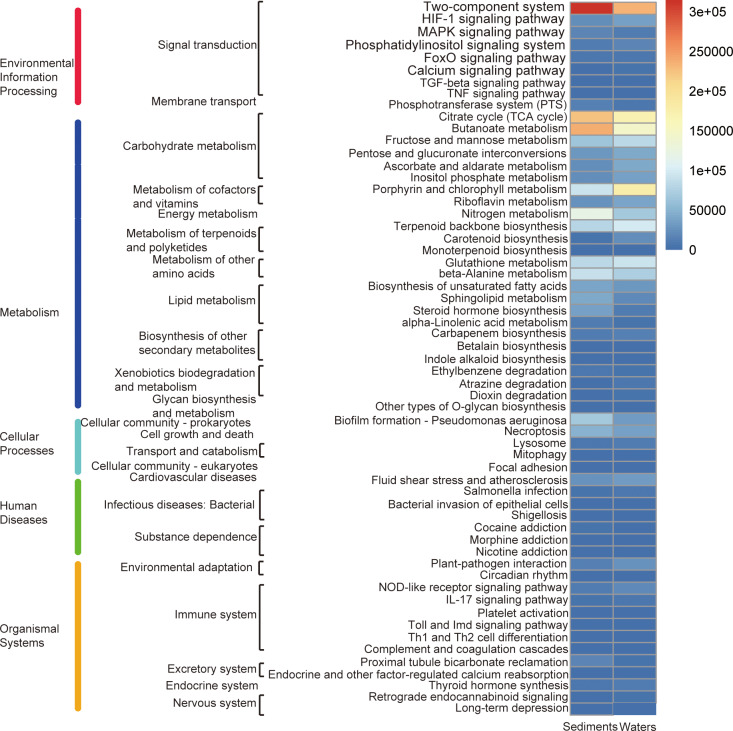
Further analysis showed that two-component system, citrate cycle (TCA cycle), and nitrogen metabolism were identified as differentially abundant pathways (Fig 5).
Fig 5. The relative abundance of top 50 most differentially abundant metabolic pathways in the sediment and seawater samples.
The three columns of descriptions from left to right represent the level 1, level 2, and level 3 KEGG pathways.
Two-component system
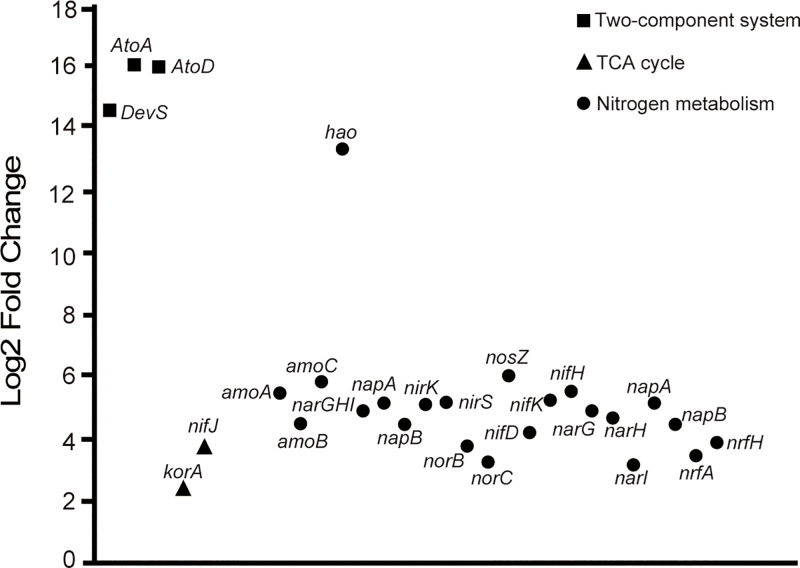
A total of 295 genes were present in the two-component system. Of these 295 genes, 182 genes exhibited higher abundance in sediments than in seawaters, while 113 genes exhibited higher abundance in seawaters than in sediments. Notably, genes including DevS, AtoA, and AtoD that encode sensor histidine kinase DevS, acetoacetate CoA-transferase beta subunit, and acetoacetate CoA-transferase alpha subunit were only identified in sediment samples (Fig 6).
Fig 6. Fold changes of selected genes that were more abundant in sediments than in seawaters.
Genes were selected from two-component system, TCA cycle, and nitrogen metabolism.
TCA cycle
A total of 59 genes involved in TCA cycle were identified for all the samples. 36 genes exhibited higher abundance in sediments than in seawaters, while 23 genes exhibited higher abundance in seawaters than in sediments. Overall, TCA cycle exhibited high abundance in sediments compared with that in seawaters. The genes nifJ and korA that encoding pyruvate-ferredoxin/flavodoxin oxidoreductase and 2-oxoglutarate/2-oxoacid ferredoxin oxidoreductase subunit alpha were more abundant in the sediments significantly (P<0.05) (Fig 6).
Nitrogen metabolism
Microbial communities in both sediments and seawaters possessed diverse metabolic genes involved in nitrogen metabolism (Fig 4, Fig 5). 36 genes were more abundant in sediments than in seawaters. The predicted nitrogen metabolism pathway was summarized in S4 Fig. For instance, genes ammonia monooxygenase (amoA, amoB, and amoC) were more abundant in the sediments than in the seawaters. The sequences annotated as “hao” were only found in sediment metagenomes. The marker genes including narGHI or napAB, norBC, nirK or nirS, nosZ encode the denitrification-related enzymes including nitrate reductase, nitric oxide reductase, nitrite reductase, and nitrous oxide reductase. These genes were all more abundant in sediments than in seawaters. Moreover, there were more abundant genes involved in nitrogen fixation (nifDKH) and in dissimilatory nitrate reduction to ammonium (DNRA) (narGHI, napAB, nirBD, and nrfAH) in sediments than in seawaters (Fig 6).
Identification of cadmium-resistant genes
In particular, the metagenomics revealed several cadmium-resistant genes. These genes include czcA, czcB, czcC, czcD, cadC, and zipB coding for czcA (cobalt-zinc-cadmium resistance protein), czcB (membrane fusion protein), czcC (outer membrane protein), czcD (cobalt-zinc-cadmium efflux system protein), cadC (lead/cadmium/zinc/bismuth-responsive transcriptional repressor), and zipB (zinc and cadmium transporter). The relative abundances of cadmium-resistant genes in sediments and seawaters were summarized in S3 Table.
Discussion
The present study is based on the comparative metagenomic evaluation of sediments and surrounding seawaters in Qinghuangdao mariculture area. The identification of microbial community compositions and their metabolic pathways could provide valuable insights into the biogeochemical process in Qinhuangdao coastal mariculture area.
According to the sequencing of 16S rRNA genes, high abundance of Proteobacteria is a common feature in both sediments and seawaters, and this phylum has been characterized to actively participate in biogeochemical processes of lake or marine ecosystems [35,36]. Previous study had revealed that Proteobacteria showed the highest relative abundance in sediments of the Bohai Sea, Yellow Sea and South China Sea [37]. Bacteroidetes and Actinobacteria were also the major phyla in cooccurrence with Proteobacteria, which are the typical heterotrophic bacteria playing roles in degrading marine organic nitrogen [37]. Similar to our results, many researches have reported that Proteobacteria (especially Alphaproteobacteria and Gammaproteobacteria), Bacteroidetes and Actinobacteria are the most important components of the offshore microbiome [38–42]. On class level, Gammaproteobacteria and Alphaproteobacteria were the most predominant class in sediments and seawaters, respectively. The result is consistent with previous findings that Gammaproteobacteria is an important class that has universal distribution in marine sediments [43] while Alphaproteobacteria dominates in the temperate coastal marine waters [44].
The sediment samples clustered distinctively from seawater samples and displayed a higher microbial diversity than the seawater samples. The metagenomic data also suggested that the sediment samples obtained from two different locations were more similar, while the seawater samples from two different locations clustered together. Therefore, the habitat substrates are likely the main factors shaping the microbial community characteristics not only at taxonomic level, but also at metabolic level. Studies have found that sediment contains more nutrients than seawater column because of the reservoirs of absorbed nutrients and high-molecular-weight organic nitrogen, and protects microorganisms from sunlight and predation. These advantages lead to sediments contain greater microbial diversity and higher microbial abundance than in the seawater columns [45,46]. Rich nutrient sediments in aquaculture have the necessary conditions for the proliferation of entire microbial communities and may be a source of new genes [47].
The microbial communities can mediate diverse metabolic processes that might be generally underrepresented. It is noteworthy that purine metabolism and pyrimidine metabolism were the most active pathways in all the communities. Ribonucleoside diphosphate reductase has been considered to be a protein that is rigorously required for elongation of DNA replication, and has a key role in regulating DNA synthesis [48]. The dnaE gene encodes the catalytic subunit of DNA polymerase III that is the main DNA replicative polymerase. The polA gene encodes DNA polymerase I that plays a role in excision-repair processes to process Okazaki fragments and fill gaps [49]. The genes rpoB and rpoC encode the major catalytic subunits of RNA polymerase in bacteria [50]. Phosphoribosyl formylglycinamidine synthase is involved in the purine biosynthetic pathway, which is necessary for nucleic acid synthesis and formation of ATP. The large subunit of arginine-specific carbamoyl phosphate synthetase was encoded by CPA2 gene and is involved in arginine biosynthesis [51]. All the highly abundant genes reported here suggested their important roles in synthesis of nucleic acid and amino acid. Result also indicated that ATP-binding cassette (ABC) transporters of microbial communities were active in this mariculture area, which couple ATP hydrolysis to actively transport of a wide variety of nutrients (e.g., some sugars, amino acids, and vitamins) [52]. The most conspicuous signature of the ABC transporters is the very high abundance of livF, livG, livK, livM, and livH genes that encode branched-chain amino acid transport ABC transport system. The function of this system is absorbing and transporting exogenous branched-chain amino acids by cells [53]. As one of an organism’s primary interfaces with the environment, the microorganisms probably heavily invested in the ABC transporters to efficiently sense and response to the environmental changes [54].
Two-component systems play important roles in adapting changing environmental conditions for microorganisms. Through two-component systems, microorganisms can modify cellular physiological activities including catalyzing reactions, initiating gene expression, or modifying protein-protein interactions [55]. DevS has been found to be a sensor of hypoxia, CO, and NO, and catalyze a reaction of DevR with ATP to produce phosphor-DevR [56]. The genes involved in metabolism of short-chain fatty acid constitute ato system. AtoA and AtoD supported the pathway of degradation of short-chain fatty acid in the sediments and eventually produced ATP by TCA cycle [57]. The high abundance of genes in TCA cycle in sediments compared to in seawaters might be indicative of the standard pathway for heterotrophic microorganisms to synthesize ATP and carry out a wider metabolic network contributing to other aspects of metabolism [58,59]. Notably, the gene coding for PFO (nifJ) as a part required for nitrogenase expression is apparently present in the TCA cycle [60]. Nitrogen cycle is critical for removing excess nutrients from marine ecosystem species. Research has revealed that the microbial communities in the upper Mississippi River sediment had large genomic potential for nitrogen cycling pathway regardless of mussel assemblages [61]. As key enzymes for nitrification, ammonia monooxygenase (amoA, amoB, and amoC) and hydroxylamine dehydrogenase (hao) co-catalyze the oxidation of ammonia to nitrite. The presence of amoA, amoB, amoC and hao in the sediments suggested that the process hydroxylamine might be an important intermediate. In that process, ammonia was used as an energy source and then the hydroxylamine was catalyzed to nitrite [62]. Enzymes including nitrate reductase, nitrite reductase, nitric oxide reductase, and nitrous oxide reductase are encoded by narGHI or napAB, nirK or nirS, norBC, and nosZ respectively. All the enzymes participate in the denitrification processes including the reduction of NO3- to NO2-, NO, N2O, and N2. The more abundance of these marker genes in the sediments suggests the elevated capacity of denitrification in sediments.
Cadmium is the most common heavy metal and is easily detected in shellfish farming environments worldwide. The existence of some genes with specific roles has important ecological significance. The presence of cadmium-resistant genes suggests the ability of microorganisms in this environment to resist the toxicity of cadmium. Proteins including czcA, czcB, czcC, and czcD are encoded by czc operon [63] which can pump Zn (Ⅱ), Cd (Ⅱ), and Co (Ⅱ) from cells, and constitute the best understood genetic mechanism of bacterial metal resistance [64]. The cadC gene encodes Cd(Ⅱ)/Pb(Ⅱ)-specific ATPase efflux pump and regulates cad cadmium resistance operon expression [65]. ZIPB protein, the Zrt-, Irt-like protein (ZIP) homolog which can transport a broad spectrum of metal ions including Cd2+, Zn2+, Cu2+, and Mn2+ [66]. All the genes above related to cadmium-resistance help assessing the functional potential of cadmium tolerance or resistance associated with the microbial community. In addition, ABC transporters can also have an vital role in detoxification of heavy metal, and resistance to the contamination of cadmium [67,68].
As far as we know, this is the first study to report the molecular inner operation mechanism of sediments and surrounding seawaters in the Qinghuangdao mariculture area. This study is a starting point to monitor the microbial communities and their metabolisms in Qinhuangdao mariculture area, especially those impacted by aquaculture activities. Although data provided here only reveals the metabolic potential of microbial community, it provides the basis for determining the actual metabolic activities of the microorganisms in the future.
Conclusions
In summary, this study provides insights into molecular characteristics of the microbial community compositions and their metabolic potentials. From the 16S rRNA gene amplicon dataset, various levels of taxonomic diversity in the microbial communities could be outlined. Metagenomic data showed the key metabolic pathways together with key genes. This study leads to the findings as the follows: (1) Proteobacteria was the overwhelming phylum in both sediments and waters, and Gammaproteobacteria and Alphaproteobacteria were the predominate classes. (2) Habitat type (i.e., sediments and seawaters) not the locations might be the reason for the variation in the microbiome in the Qinhuangdao mariculture area. (3) The microbial communities in Qinhuangdao mariculture area were mainly predicted to be active on the purine metabolism, ABC transporters and pyrimidine metabolism. (4) Two component system, TCA cycle, and nitrogen metabolism were the more abundant pathways in the sediments than in the seawaters. (5) cadmium-resistant genes and ABC transporters might play roles in resisting the toxicity of cadmium. Together, these results evidenced a broad microbial diversity with versatile metabolic pathways for the microbial communities in Qinhuangdao mariculture area.
Supporting information
(TIF)
The number on the bar chart represents the number of genes on the annotation. Another axis is the functional annotation information in KEGG.
(TIF)
The number on the bar chart represents the percentage of reads annotated to each KEGG pathway.
(TIF)
Enzyme commission number and name of gene product are shown in the boxes. Genes only found in sediments are labeled with asterisk. Enzyme commission number is shown in the boxes.
(TIF)
(PDF)
(PDF)
(PDF)
Acknowledgments
We thank to Dr. Qinglin Wang (Hebei Normal University of Science and Technology) for assistance in sampling.
Data Availability
Metagenomic datasets are available online at NCBI, BioSample accessions: SAMN13744940, SAMN13744941, SAMN13744942, SAMN13744943 for Seawater1 (S1), Sediment1 (S1S), Seawater2 (S2), and Sediment2 (S2S) samples respectively. The 16S rRNA datasets are available online at NCBI, BioSample accessions: SAMN13752121, SAMN13752122, SAMN13752123, SAMN13752124 for (Seawater sample1) S1, (Sediment sample1) S1S, (Seawater sample2) S2, and (Sediment sample2) S2S samples respectively.
Funding Statement
This work was supported by the Fundamental Research Funds for Central Public Welfare Scientific Research Institutes of China [grant number 2019YSKY-007](SW).
References
- 1.Petro C, Starnawski P, Schramm A, Kjeldsen KU. Microbial community assembly in marine sediments. Aquat Microb Ecol. 2017; 79: 177–195. [Google Scholar]
- 2.Yang X, Tsibart A, Nam H, Hur J, Elnaggar A, Tack F, et al. Effect of gasification biochar application on soil quality: Trace metal behavior, microbial community, and soil dissolved organic matter. J Hazard Mater. 2019; 365: 684–694. 10.1016/j.jhazmat.2018.11.042 [DOI] [PubMed] [Google Scholar]
- 3.Dequiedt S, Saby NPA, Lelievre M, Jolivet C, Thioulouse J, Toutain B, et al. Biogeographical patterns of soil molecular microbial biomass as influenced by soil characteristics and management. Global Ecol Biogeogr. 2011; 20: 641–652. [Google Scholar]
- 4.Reavie ED, Jicha TM, Angradi TR, Bolgrien DW, Hill BH. Algal assemblages for large river monitoring: Comparison among biovolume, absolute and relative abundance metrics. Ecol Indic. 2010; 10: 167–177. [Google Scholar]
- 5.Zhang Y, Lin X, Shi X, Lin L, Luo H, Li L, et al. Metatranscriptomic signatures associated with phytoplankton regime shift from diatom dominance to a dinoflagellate bloom. Front Microbiol. 2019; 10: 590 10.3389/fmicb.2019.00590 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Magurran AE, Baillie SR, Buckland ST, Dick JM, Elston DA, Scott EM, et al. Long-term datasets in biodiversity research and monitoring: assessing change in ecological communities through time. Trends Ecol Evol. 2010; 25: 574–582. 10.1016/j.tree.2010.06.016 [DOI] [PubMed] [Google Scholar]
- 7.Dafforn KA, Baird DJ, Chariton AA, Sun MY, Brown MV, Simpson SL, et al. Faster, Higher and Stronger? The Pros and Cons of Molecular Faunal Data for Assessing Ecosystem Condition. Adv Ecol Res. 2014; 51: 1–40. [Google Scholar]
- 8.Kearns PJ, Angell JH, Feinman SG, Bowen JL. Long-term nutrient addition differentially alters community composition and diversity of genes that control nitrous oxide flux from salt marsh sediments. Estuar Coast Shelf S. 2015; 154: 39–47. [Google Scholar]
- 9.Chariton AA, Sun MY, Gibson JF, Webb JA, Leung KMY, Hickey CW, et al. Emergent technologies and analytical approaches for understanding the effects of multiple stressors in aquatic environments. Mar Freshwater Res. 2016; 67: 414–428. [Google Scholar]
- 10.Shade A, Handelsman J. Beyond the Venn diagram: the hunt for a core microbiome. Environ Microbiol. 2012; 14: 4–12. 10.1111/j.1462-2920.2011.02585.x [DOI] [PubMed] [Google Scholar]
- 11.Tavares TCL, Normando LRO, De Vasconcelos ATR, Gerber AL, Agnezlima LF, Melo VMM. Metagenomic analysis of sediments under seaports influence in the Equatorial Atlantic Ocean. Sci Total Environ. 2016; 557: 888–900. 10.1016/j.scitotenv.2016.03.141 [DOI] [PubMed] [Google Scholar]
- 12.Birrer SC, Dafforn KA, Sun MY, Williams RBH, Potts J, Scanes P, et al. Using meta-omics of contaminated sediments to monitor changes in pathways relevant to climate regulation. Environ Microbiol. 2018; 21: 389–401. 10.1111/1462-2920.14470 [DOI] [PubMed] [Google Scholar]
- 13.Caruso G, La Ferla R, Azzaro M, Zoppini A, Marino G, Petochi T, et al. Microbial assemblages for environmental quality assessment: Knowledge, gaps and usefulness in the European Marine Strategy Framework Directive. Crit Rev Microbiol. 2016; 42: 883–904. 10.3109/1040841X.2015.1087380 [DOI] [PubMed] [Google Scholar]
- 14.Zhang S, Hu Z, Wang H. Metagenomic analysis exhibited the co-metabolism of polycyclic aromatic hydrocarbons by bacterial community from estuarine sediment. Environ Int. 2019; 129: 308–319. 10.1016/j.envint.2019.05.028 [DOI] [PubMed] [Google Scholar]
- 15.Callahan BJ, Wong J, Heiner C, Oh S, Theriot CM, Gulati AS, et al. High-throughput amplicon sequencing of the full-length 16S rRNA gene with single-nucleotide resolution. Nucleic Acids Res. 2018; 47: e103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Wang S, Zhang Y, He J, Jia X, Lin J, Li M, et al. Molecular analyses of bacterioplankton communities with highly abundant Vibrio clades: a case study in Bohai Sea coastal waters. J Oceanol Limnol. 2019; 37: 1638–1648. [Google Scholar]
- 17.Wang S, Zheng X, Xia H, Shi D, Fan J, Wang P, et al. Archaeal community variation in the Qinhuangdao coastal aquaculture zone revealed by high-throughput sequencing. PLOS ONE. 2019; 14: e0218611 10.1371/journal.pone.0218611 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zhang X, Xu W, Liu Y, Cai M, Luo Z, Li M. Metagenomics reveals microbial diversity and metabolic potentials of seawater and surface sediment from a Hadal biosphere at the Yap trench. Front Microbiol. 2018; 9:2402 10.3389/fmicb.2018.02402 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Luke C, Speth DR, Kox MAR, Villanueva L, Jetten MSM. Metagenomic analysis of nitrogen and methane cycling in the Arabian Sea oxygen minimum zone. PeerJ. 2016; 4: 28–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Li Y, Zheng L, Zhang Y, Liu H, Jing H. Comparative metagenomics study reveals Pollution induced changes of microbial genes in mangrove sediments. Sci Rep. 2019; 9: 5739 10.1038/s41598-019-42260-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lu X, Chen C, Zheng T. Metagenomic insights into effects of chemical pollutants on microbial community composition and function in estuarine sediments receiving polluted river water. Microb Ecol. 2017; 73: 791–800. 10.1007/s00248-016-0868-8 [DOI] [PubMed] [Google Scholar]
- 22.Li J, Jin X, Zhang X, Chen L, Liu J, Zhang H, et al. Comparative metagenomics of two distinct biological soil crusts in the Tengger Desert, China. Soil Biol Biochem. 2020; 140: 107637. [Google Scholar]
- 23.Gao Y, Yang Q, Li H, Wang X, Zhan A. Anthropogenic pollutant-driven geographical distribution of mesozooplankton communities in estuarine areas of the Bohai Sea, China. Sci Rep. 2019; 9: 9668 10.1038/s41598-019-46047-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Gao X, Chen C-TA. Heavy metal pollution status in surface sediments of the coastal Bohai Bay. Water Res. 2012; 46: 1901–1911. 10.1016/j.watres.2012.01.007 [DOI] [PubMed] [Google Scholar]
- 25.Li Y, Guo L, Feng H. Status and trends of sediment metal pollution in Bohai Sea, China. Curr Pollut Rep. 2015; 1: 191–202. [Google Scholar]
- 26.Zhang W, Wang W-X. Large-scale spatial and interspecies differences in trace elements and stable isotopes in marine wild fish from Chinese waters. J Hazard Mater. 2012; 215–216: 65–74. 10.1016/j.jhazmat.2012.02.032 [DOI] [PubMed] [Google Scholar]
- 27.Shahidul Islam M, Tanaka M. Impacts of pollution on coastal and marine ecosystems including coastal and marine fisheries and approach for management: a review and synthesis. Mar Pollut Bull. 2004; 48: 624–649. 10.1016/j.marpolbul.2003.12.004 [DOI] [PubMed] [Google Scholar]
- 28.Caporaso JG, Lauber CL, Walters WA, Berglyons D, Lozupone CA, Turnbaugh PJ, et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA. 2011; 108: 4516–4522. 10.1073/pnas.1000080107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010; 7: 335–336. 10.1038/nmeth.f.303 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Yilmaz P, Parfrey LW, Yarza P, Gerken J, Pruesse E, Quast C, et al. The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acids Res. 2014; 42: 643–648. 10.1093/nar/gkt888 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature. 2010; 464: 59–65. 10.1038/nature08821 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Cotillard A, Kennedy SP, Kong LC, Prifti E, Pons N, Le Chatelier E, et al. Dietary intervention impact on gut microbial gene richness. Nature. 2013; 500: 585–588. 10.1038/nature12480 [DOI] [PubMed] [Google Scholar]
- 33.Le Chatelier E, Nielsen T, Qin J, Prifti E, Hildebrand F, Falony G, et al. Richness of human gut microbiome correlates with metabolic markers. Nature. 2013; 500: 541–546. 10.1038/nature12506 [DOI] [PubMed] [Google Scholar]
- 34.Altermann E, Klaenhammer TR. PathwayVoyager: pathway mapping using the Kyoto Encyclopedia of Genes and Genomes (KEGG) database. BMC Genomics. 2005; 6: 60–60. 10.1186/1471-2164-6-60 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Zhang J, Yang Y, Zhao L, Li Y, Xie S, Liu Y. Distribution of sediment bacterial and archaeal communities in plateau freshwater lakes. Appl Microbiol Biot. 2015; 99: 3291–3302. [DOI] [PubMed] [Google Scholar]
- 36.Saleem F, Azim MK, Mustafa A, Kori JA, Hussain MS. Metagenomic profiling of fresh water lakes at different altitudes in Pakistan. Ecol Inform. 2019; 51: 73–81. [Google Scholar]
- 37.Zhang J, Chen M, Huang J, Guo X, Zhang Y, Liu D, et al. Diversity of the microbial community and cultivable protease-producing bacteria in the sediments of the Bohai Sea, Yellow Sea and South China Sea. PLOS ONE. 2019; 14: [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Ye Q, Wu Y, Zhu Z, Wang X, Li Z, Zhang J. Bacterial diversity in the surface sediments of the hypoxic zone near the Changjiang Estuary and in the East China Sea. Microbiologyopen. 2016; 5: 323–339. 10.1002/mbo3.330 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Duncan KR, Haltli B, Gill KA, Kerr RG. Bioprospecting from marine sediments of New Brunswick, Canada: exploring the relationship between total bacterial diversity and actinobacteria diversity. Mar Drugs. 2014; 12: 899–925. 10.3390/md12020899 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Wang J, Tao S, Yu K, Jiang R, Liu M, Liu X, et al. Bacterial diversity in the surface layer of sediments from the East China Sea. Evol Ecol Res. 2016; 17: 721–736. [Google Scholar]
- 41.Hoffmann K, Hassenruck C, Salmancarvalho V, Holtappels M, Bienhold C. Response of bacterial communities to different detritus compositions in arctic deep-sea sediments. Front Microbiol. 2017; 8:266 10.3389/fmicb.2017.00266 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Li Y, Wu C, Zhou M, Wang ET, Zhang Z, Liu W, et al. Diversity of Cultivable Protease-Producing Bacteria in Laizhou Bay Sediments, Bohai Sea, China. Front Microbiol. 2017; 8: 405 10.3389/fmicb.2017.00405 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Hunter EM, Mills HJ, Kostka JE. Microbial community diversity associated with carbon and nitrogen cycling in permeable shelf sediments. Appl Environ Microb. 2006; 72: 5689–5701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Dang H, Li T, Chen M, Huang G. Cross-Ocean Distribution of Rhodobacterales Bacteria as Primary Surface Colonizers in Temperate Coastal Marine Waters. Appl Environ Microb. 2008; 74: 52–60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Kobiyama A, Ikeo K, Reza MS, Rashid J, Yamada Y, Ikeda Y, et al. Metagenome-based diversity analyses suggest a strong locality signal for bacterial communities associated with oyster aquaculture farms in Ofunato Bay. Gene. 2018; 665: 149–154. 10.1016/j.gene.2018.04.073 [DOI] [PubMed] [Google Scholar]
- 46.Jorgensen SL, Hannisdal B, Lanzen A, Baumberger T, Flesland K, Fonseca RG, et al. Correlating microbial community profiles with geochemical data in highly stratified sediments from the Arctic Mid-Ocean Ridge. Pro Natl Acad Sci USA. 2012; 109: 16764–16765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Perkins TL, Clements K, Baas JH, Jago CF, Jones DL, Malham SK, et al. Sediment composition influences spatial variation in the abundance of human pathogen indicator bacteria within an estuarine environment. PLOS ONE. 2014; 9: e112951 10.1371/journal.pone.0112951 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Martinezporchas M, Vargasalbores F. Microbial metagenomics in aquaculture: a potential tool for a deeper insight into the activity. Rev Aqua. 2017; 9: 42–56. [Google Scholar]
- 49.Guzman EC, Caballero JL, Jimenezsanchez A. Ribonucleoside diphosphate reductase is a component of the replication hyperstructure in Escherichia coli. Mol Microbiol. 2002; 43: 487–495. 10.1046/j.1365-2958.2002.02761.x [DOI] [PubMed] [Google Scholar]
- 50.Hastings PJ, Hersh MN, Thornton PC, Fonville NC, Slack A, Frisch RL, et al. Competition of Escherichia coli DNA Polymerases I, II and III with DNA Pol IV in Stressed Cells. PLOS ONE. 2010; 5: e10862 10.1371/journal.pone.0010862 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Lee YH, Nam KH, Helmann JD. A Mutation of the RNA Polymerase β′ Subunit (rpoC) Confers Cephalosporin Resistance in Bacillus subtilis. Antimicrob Agents Ch. 2013; 57: 56–65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Liu X, Cai Y, Zhang X, Zhang H, Zheng X, Zhang Z. Carbamoyl phosphate synthetase subunit MoCpa2 affects development and pathogenicity by modulating arginine biosynthesis in magnaporthe oryzae. Front Microbiol. 2016; 7: 2023–2023. 10.3389/fmicb.2016.02023 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Cuthbertson L, Kos V, Whitfield C. ABC transporters involved in export of cell surface glycoconjugates. Microbiol. Mol Biol R. 2010; 74: 341–362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Hosie AHF, Poole PS. Bacterial ABC transporters of amino acids. Res Microbiol. 2001; 152: 259–270. 10.1016/s0923-2508(01)01197-4 [DOI] [PubMed] [Google Scholar]
- 55.Giuliani SE, Frank AM, Corgliano DM, Seifert C, Hauser L, Collart F. Environment sensing and response mediated by ABC transporters. BMC Genomics 2011; 12: 1–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Skerker JM, Prasol MS, Perchuk BS, Biondi EG, Laub MTJPB. Two-component signal transduction pathways regulating growth and cell cycle progression in a bacterium: a system-level analysis. PloS Biol. 2005; 3: e334 10.1371/journal.pbio.0030334 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Sousa EHS, Gonzalez G, Gillesgonzalez MA. Target DNA stabilizes Mycobacterium tuberculosis DevR/DosR phosphorylation by the full‐length oxygen sensors DevS/DosS and DosT. FEBS J. 2017; 284: 3954–3967. 10.1111/febs.14284 [DOI] [PubMed] [Google Scholar]
- 58.Jenkins LS, Nunn WD. Genetic and molecular characterization of the genes involved in short-chain fatty acid degradation in Escherichia coli: the ato system. J Bacteriol. 1987; 169: 42–52. 10.1128/jb.169.1.42-52.1987 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Glockner FO, Kube M, Bauer M, Teeling H, Lombardot T, Ludwig W, et al. Complete genome sequence of the marine planctomycete Pirellula sp strain 1. Proc Natl Acad Sci USA. 2003; 100: 8298–8303. 10.1073/pnas.1431443100 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Araujo WL, Nunesnesi A, Nikoloski Z, Sweetlove LJ, Fernie AR. Metabolic control and regulation of the tricarboxylic acid cycle in photosynthetic and heterotrophic plant tissues. Plant Cell Environ. 2012; 35: 1–21. 10.1111/j.1365-3040.2011.02332.x [DOI] [PubMed] [Google Scholar]
- 61.Schmitz O, Kentemich T, Zimmer W, Hundeshagen B, Bothe H. Identification of the nifJ gene coding for pyruvate: ferredoxin oxidoreductase in dinitrogen-fixing cyanobacteria. Arch Microbiol. 1993; 160: 62–67. [PubMed] [Google Scholar]
- 62.Black EM. Assessing the impacts of native freshwater mussels on nitrogen cycling microbial communities using metagenomics. Ph.D. thesis, The University of Iowa. 2018.
- 63.Cerqueira T, Barroso C, Froufe H, Egas C, Bettencourt R. Metagenomic signatures of microbial communities in deep-sea hydrothermal sediments of Azores vent fields. Microb Ecol. 2018; 76: 387–403. 10.1007/s00248-018-1144-x [DOI] [PubMed] [Google Scholar]
- 64.Grosse C, Anton A, Hoffmann T, Franke S, Schleuder G, Nies DH. Identification of a regulatory pathway that controls the heavy-metal resistance system Czc via promoter czcNp in Ralstonia metallidurans. Arch Microbiol. 2004; 182: 109–118. 10.1007/s00203-004-0670-8 [DOI] [PubMed] [Google Scholar]
- 65.Intorne AC, De Oliveira MVV, Pereira LDM, Filho GADS. Essential role of the czc determinant for cadmium, cobalt and zinc resistance in Gluconacetobacter diazotrophicus PAl 5. Int Microbiol. 2012; 15: 69–78. 10.2436/20.1501.01.160 [DOI] [PubMed] [Google Scholar]
- 66.Busenlehner LS, Pennella MA, Giedroc DP. The SmtB/ArsR family of metalloregulatory transcriptional repressors: structural insights into prokaryotic metal resistance. Fems Microbiol Rev. 2003; 27: 131–143. 10.1016/S0168-6445(03)00054-8 [DOI] [PubMed] [Google Scholar]
- 67.Lin W, Chai J, Love J, Fu D. Selective Electrodiffusion of Zinc Ions in a Zrt-, Irt-like Protein, ZIPB. J Biol Chem. 2010; 285: 39013–39020. 10.1074/jbc.M110.180620 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Feng G, Xie T, Wang X, Bai J, Tang L, Zhao H, et al. Metagenomic analysis of microbial community and function involved in cd-contaminated soil. BMC Microbiol. 2018; 18: 11–11. 10.1186/s12866-018-1152-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
(TIF)
The number on the bar chart represents the number of genes on the annotation. Another axis is the functional annotation information in KEGG.
(TIF)
The number on the bar chart represents the percentage of reads annotated to each KEGG pathway.
(TIF)
Enzyme commission number and name of gene product are shown in the boxes. Genes only found in sediments are labeled with asterisk. Enzyme commission number is shown in the boxes.
(TIF)
(PDF)
(PDF)
(PDF)
Data Availability Statement
Metagenomic datasets are available online at NCBI, BioSample accessions: SAMN13744940, SAMN13744941, SAMN13744942, SAMN13744943 for Seawater1 (S1), Sediment1 (S1S), Seawater2 (S2), and Sediment2 (S2S) samples respectively. The 16S rRNA datasets are available online at NCBI, BioSample accessions: SAMN13752121, SAMN13752122, SAMN13752123, SAMN13752124 for (Seawater sample1) S1, (Sediment sample1) S1S, (Seawater sample2) S2, and (Sediment sample2) S2S samples respectively.