Abstract
Ralstonia solanacearum releases a set of effectors into plant cells that modify the host defence reaction. The role of the effector protein RipI during infection has not been elucidated. In this study, we demonstrated that transient overexpression of RipI induces the hypersensitive response (HR), up‐regulating the HR marker gene hin1, in Nicotiana benthamiana. Deletion of R. solanacearum ripI led to increased virulence in tomato (Solanum lycopersicum) plants. Through yeast two‐hybrid and pull‐down assays, we identified an interaction between the N. benthamiana transcription factor bHLH93 and RipI, both of which could be localized in the nucleus of Arabidopsis protoplasts. Silencing of bHLH93 markedly attenuated the RipI‐induced HR and induced expression of the PDF1.2 defence gene. These data demonstrate that the R. solanacearum effector RipI induces a host defence reaction by interacting with the bHLH93 transcription factor.
Keywords: bHLH93, defence reaction, Ralstonia solanacearum, RipI, transcription factor
Ralstonia solanacearum RipI is recognized by bHLH93 to induce a defence reaction in tobacco plants.
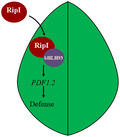
1.
Ralstonia solanacearum causes bacterial wilt in a wide range of host plants, including Nicotiana benthamiana. This pathogen is a species complex, a heterogeneous group of related but genetically distinct strains (Genin and Denny, 2012). The available genome sequences demonstrate that R. solanacearum possesses over 110 effector candidates that vary among isolates (Lonjon et al., 2018), secreted by the type III secretion system (T3SS).
The first complete genomic sequence of R. solanacearum was characterized from the strain GMI1000 isolated from tomato (Solanum lycopersicum) (Salanoubat et al., 2002). GMI1000 is pathogenic on the model plant Arabidopsis thaliana and nonpathogenic on N. benthamiana. The GMI1000 genome encodes around 72 effectors, one‐third of which are conserved in all R. solanacearum strains (Peeters et al., 2013). The RipP2 effector interacts with the Arabidopsis NB‐LRR protein RRS1‐R, leading to avirulence on A. thaliana Nd‐1 (Deslandes et al., 2003). RipP1 and RipAA act jointly to specify an incompatible interaction within Nicotiana plants (Poueymiro et al., 2009). RipAY and RipAK target plant redox regulators and host catalases, respectively, suppressing the immune response (Sun et al., 2017; Sang et al., 2018). Furthermore, a few effectors induce hypersensitive response (HR)‐like responses in different plant species. For example, RipAX2 induces an HR in eggplant (Solanum torvum) via an essential zinc‐protease motif (Nahar et al., 2014), and elicits the resistance of eggplant AG91‐25 (Morel et al., 2018).
A 47‐kDa effector protein, RipI, composed of 430 amino acids is encoded by RSc0041 in R. solanacearum GMI1000 (GenBank no. NC_003295.1) and contains two predicted motifs: an integrase motif from N54 to N174 and a DNA polymerase III subunit γ motif from N143 to N378 at the N‐terminus of RipI, respectively (Figure S1). Ectopic expression of RipI in Saccharomyces cerevisiae results in cell apoptosis; the integrase motif is essential for the apoptosis (Deng et al., 2016). To examine plant responses to RipI, we transiently expressed RipI in N. benthamiana leaves using Agrobacterium‐mediated transformation. An HR was evident 2 days post‐agroinfiltration, coupled with cell death and hydrogen peroxide accumulation (Figure 1a,b). Furthermore, the HR marker gene hin1 (Gopalan et al., 1996) was induced 17‐fold relative to control leaves agroinfiltrated with empty binary vector (Figure 1c).
FIGURE 1.
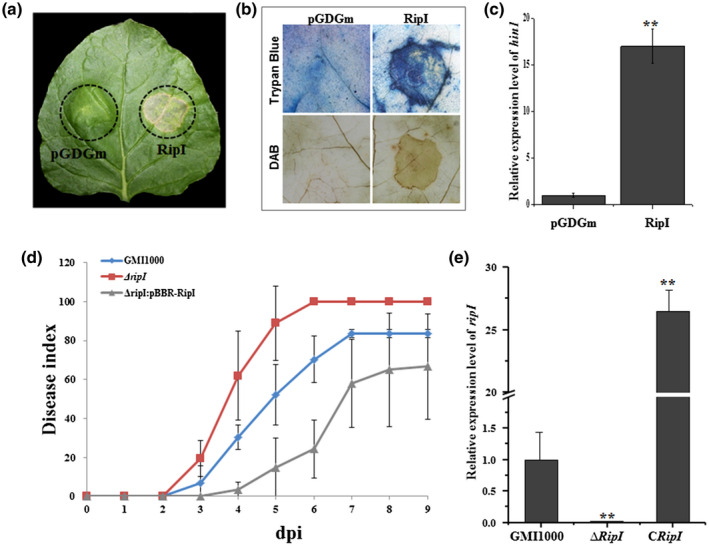
RipI induces a defence reaction in host plants. (a) Hypersensitive response (HR) induced by transient heterologous expression of RipI in Nicotiana benthamiana leaves. Dotted circles indicate inoculated areas. Agrobacterium tumefaciens GV3101 harbouring empty vector control pGDGm or pGDGm‐RipI was prepared to OD600 = 1.0 and infiltrated into N. benthamiana leaves. Photographs were taken at 2 days post‐agroinfiltration. (b) Detection of cell death and hydrogen peroxide accumulation by tissue staining. N. benthamiana leaves were harvested 36 hr post‐agroinfiltration and stained with trypan blue and 3,3′‐diaminobenzidine (DAB). (c) Assessment of hin1 transcript level by quantitative reverse transcription PCR. hin1 transcript level induced by RipI was compared with that induced by an empty vector pGDGm control (set as 1) at 36 hr post‐agroinfiltration. Error bars represent the standard deviation from three replicates. Differences were evaluated using Student's t test (**p < .01). (d) Progression of bacterial wilt on tomato plants inoculated with the wild type (GMI1000), mutant ΔripI, or the complemented strain ΔripI:pBBR‐RipI. Disease severity was rated for 9 days after stem inoculation (dpi). Each time point represents the mean disease severity of six inoculated plants per treatment. Error bars represent the standard deviation from three independent experiments. (e) Transcription of the ripI gene in the wild type GMI1000, mutant ΔripI, and the complemented strain ΔripI:pBBR‐RipI (CRipI). RNAs were extracted from cells cultured in minimal medium M63. Error bars represent the standard deviation from three replicates. Expression level in GMI1000 was set to 1. Differences were evaluated using Student's t test (**p < .01). All experiments were replicated three times with similar results and representative results are shown
To investigate the role of the R. solanacearum RipI effector in virulence, we constructed the deletion mutant ΔripI in strain GMI1000. Although wilt symptoms initially appeared at 3 days post‐inoculation (dpi), tomato plants inoculated with ΔripI displayed more rapid disease development than those inoculated with wild‐type GMI1000. The disease index of ΔripI was 20.1, while that of the wild type was only 8.3 at 3 dpi. By 6 dpi, all ΔripI‐infected plants had fully wilted, and the disease index of GMI1000 was 82.7 (Figure 1d). We observed wilt symptoms in the tomato plants inoculated with the ΔripI strain complemented with ripI (ΔripI:pBBR‐RipI) at 4 dpi, 1 day later than observed in the wild type and mutant ΔripI; however, the disease index of the complemented strain was lower than that of the wild type and ΔripI (Figure 1d). We examined the transcript levels of ripI in the three strains by quantitative reverse transcription‐PCR (RT‐qPCR). A higher transcript level was observed in the complemented strain than in the wild type and no transcript was detected in the ΔripI mutant (Figure 1e).
A yeast two‐hybrid screen using RipI as bait showed that the N. benthamiana transcription factor bHLH93 (Niben101Scf05329g05016.1) interacts with RipI (https://solgenomics.net/organism/Nicotiana_benthamiana/genome). To confirm this interaction, recovered pGADT7 plasmid was retransformed into AH109 together with RipI bait. As expected, the transformant was able to grow on SD –His –Leu –Trp –Ade medium supplemented with 20 μg/ml X‐α‐galactosidase (Figure 2a). Furthermore, a glutathione‐S‐transferase (GST) pull‐down assay confirmed that RipI physically interacts with bHLH93 (Figure 2b).
FIGURE 2.
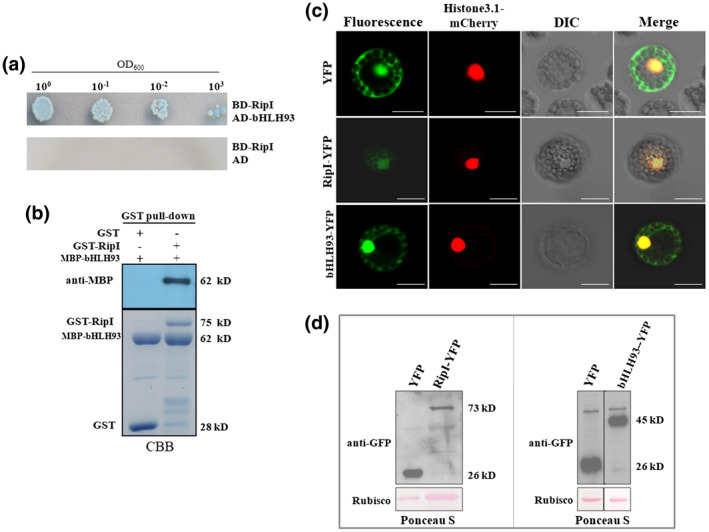
RipI interacts with the bHLH93 transcription factor in Nicotiana benthamiana. (a) Yeast two‐hybrid assays showing the interaction between RipI and bHLH93. The transformants were prepared to a cell density of OD600 = 1.0 and diluted in a 10‐fold series. For each concentration, 2 μl was spotted and incubated on synthetic defined SD −Ade −Leu −Trp −His plates supplemented with 20 μg/ml X‐α‐galactosidase (X‐α‐gal) for 4 days at 30 °C. The transformant containing BD‐RipI and empty vector pGADT7 served as a negative control. (b) RipI interacts with bHLH93 in vitro in glutathione‐S‐transferase (GST) pull‐down assays. Recombinant GST‐RipI and maltose binding protein (MBP)‐bHLH93 fusions were subjected to GST pull‐down analysis. GST tag and MBP‐bHLH93 fusions were used as the negative control. Gel stained with Coomassie brilliant blue (CBB) is shown. Interacting proteins were identified by immunoblotting using anti‐MBP antibodies. The experiment was repeated three times with similar results. (c) Subcellular localization of RipI and bHLH93 in Arabidopsis protoplasts. Arabidopsis protoplasts were transfected for coexpression of mCherry‐histone3.1 (serving as a nuclear localization marker) and either RipI or bHLH93 fused with yellow fluorescent protein (YFP). Images were recorded at 8 hr post‐transfection for visualization of YFP fluorescence at 488 nm and mCherry fluorescence at 580 nm. The colour of YFP fluorescence was set to green to facilitate detection of the fluorescence merged with mCherry fluorescence (green merged with red is shown as yellow, while yellow merged with red is shown as orange). Differential interference contrast (DIC) images were also photographed. Bars in all images represent 20 μm. (d) Immunoblot analysis of RipI expression in Arabidopsis protoplasts using green fluorescent protein (GFP) polyclonal antiserum (anti‐GFP). Total proteins were extracted from Arabidopsis protoplasts. Arabidopsis protoplasts transfected with empty vector expressing YFP were used as the control. The loading control was RuBisCO stained with Ponceau S. All experiments were replicated three times with similar results and representative results are shown
Next, we examined the subcellular localization of bHLH93 and RipI using Arabidopsis protoplasts. Although some protoplasts transfected with RipI‐yellow fluorescent protein (YFP) perished, we identified RipI in the nucleus of a number of protoplasts (Figure 2c). This was consistent with the nuclear localization of RipI reported in yeast protoplasts (Deng et al., 2016). bHLH93‐YFP was located in the nucleus, membrane, and cytoplasm, and its fluorescence was much brighter than that of RipI‐YFP (Figure 2c). Transfection with bHLH93‐YFP was more successful than transfection with RipI‐YFP, resulting in many more fluorescent protoplasts. Immunoblot analysis using anti‐GFP antibodies confirmed the stronger signals from protoplasts transfected with bHLH93‐YFP (Figure 2d). No fluorescence was detected from protoplasts co‐transfected with RipI and bHLH93.
We investigated the requirement for bHLH93 in HR induction by RipI using tobacco rattle virus (TRV)‐induced gene silencing. A 529‐bp fragment including coding and 3′ untranslated region (UTR) sequence of bHLH93 was cloned into pTRV‐RNA2, generating pTRV2‐bHLH93. pTRV2‐gfp carrying the gfp (green fluorescent protein) gene fragment served as a negative control and pTRV2‐PDS (phytoene desaturase) was used to determine silencing efficiency. At 13 dpi, when photobleaching symptoms were observed on the upper leaves of PDS‐silenced plants, RipI was transiently expressed in bHLH93‐silenced plants to examine HR induction. RipI agroinfiltrated at three concentrations (OD600 = 0.01, 0.1, and 1.0) induced a weaker HR in bHLH93‐silenced plants than in control plants transformed with pTRV1 and pTRV2‐gfp (TRV‐gfp) (Figure 3a). RT‐qPCR confirmed the silencing effect of bHLH93 (Figure 3b).
FIGURE 3.
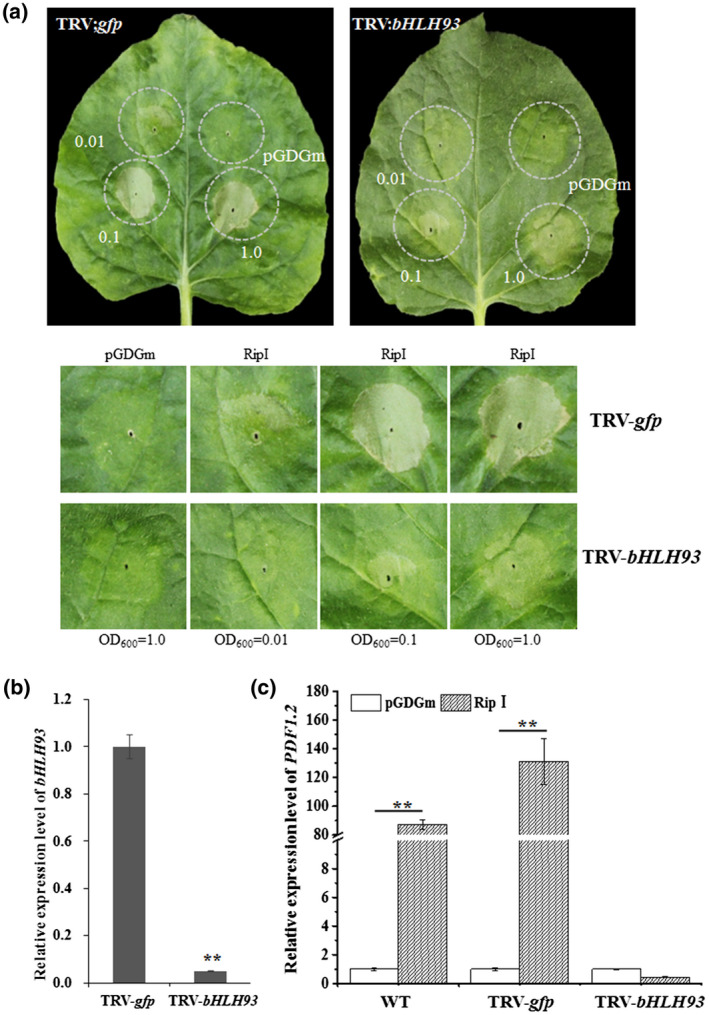
Requirement of bHLH93 for defence induction by RipI. (a) Silencing of bHLH93 attenuated the hypersensitive response (HR) induced by RipI. Procedures were similar to those shown in Figure 1a. Leaves were inoculated with three concentrations (OD600 = 0.01, 0.1, and 1.0) of the pGDGm‐RipI construct. The lower images show enlargements of areas exhibiting HR, whereas the whole leaves are shown above. (b) Quantitative reverse transcription PCR analysis of bHLH93 transcript level in bHLH93‐silenced plants. RNAs were isolated from the upper new leaves when the photobleaching phenotype was observed in phytoene desaturase (PDS)‐silenced plants (positive control). The transcript level in plants transformed with tobacco rattle virus (TRV)‐gfp was used as a control to monitor expression change. Error bars represent the standard deviation from three replicates. Differences were evaluated using Student's t tests (**p < .01). (c) bHLH93 is required for PDF1.2 induction by RipI. PDF1.2 transcript was quantified in wild type (WT) and in bHLH93‐silenced plants transiently expressing RipI. The transcript level in plants transformed with TRV‐gfp served as a negative control. RNAs were isolated 36 hr post‐agroinfiltration with RipI. Error bars represent the standard deviation from three replicates. Differences were evaluated using Student's t test (**p < .01). All experiments were replicated three times with similar results and representative results are shown
Salicylic acid (SA), jasmonic acid (JA), and ethylene (ET) induce diverse but cross‐linked defence signalling pathways against microbial pathogens in plants (Zhang et al., 2017; Nakano and Mukaihara, 2018). The elicitation of an HR and attenuation of virulence prompted us to investigate whether RipI was associated with any defence signalling pathway. PDF1.2, a defensin gene involved in the JA and ET signalling responses (Robert‐Seilaniantz et al., 2007; Vos et al., 2015), was induced by RipI (Figure 3c). However, PDF1.2 transcript in bHLH93‐silenced plants was dramatically reduced in response to transient expression of RipI (Figure 3c). The silencing effect of bHLH93 in bHLH93‐silenced plants was confirmed by RT‐qPCR (Figure S3).
Although RipI induces the defence response, it does not function as an avirulence protein specifying an incompatible interaction with N. benthamiana in the GMI1000 genetic background. The ripI mutant of GMI1000 retained the ability to induce a HR in N. benthamiana, which was jointly determined by RipAA and RipA1 effectors or RipB from strain RS1000 (Poueymiro et al., 2009; Nakano and Mukaihara, 2019). The cell death induced by RipI observed here was consistent with the previous report that ectopic expression of RipI leads to apoptosis in yeast (Deng et al., 2016). As the arginine at position 117 within the integrase motif is required for its inhibition of yeast growth (Deng et al., 2016), the involvement of the integrase motif in inducing cell death in N. benthamiana should be clarified.
bHLH proteins belong to a superfamily of transcription factors with redundant members in plants (Ledent and Vervoort, 2001). For example, 133 genes encode bHLHs in A. thaliana (Heim et al., 2003). These proteins play multiple roles in cell and tissue development and regulate hormone metabolism, improving tolerance to drought stress (Heim et al., 2003; Bruex et al., 2012). While 324 bHLH family genes in N. benthamiana have been identified (Yu et al., 2019), the biological roles of most NbbHLH genes remain unknown and only 278 NbbHLHs sequences were released. Though all the NbbHLHs contain bHLH domains, the identities of either full‐length sequence or bHLH domain sequence among 278 NbbHLHs are low (Figure S2). The RipI‐interacting protein Niben101Scf05329g05016.1 is annotated as bHLH93 in the N. benthamiana genome database. In this study, we established that bHLH93 was targeted by the R. solanacearum effector RipI and induced a defence reaction in N. benthamiana. Both RipI and bHLH93 were located in the nucleus; however, no fluorescence was detected in Arabidopsis protoplasts co‐transfected with RipI and bHLH93, suggesting that the simultaneous expression of RipI and bHLH93 may accelerate cell death progression, causing a rapid degradation of the nucleus.
JA signalling is a known target of plant bacterial T3SS effectors. The RipAL effector contributes to R. solanacearum virulence in pepper (Capsicum annuum) plants by inducing JA production (Nakano and Mukaihara, 2018). The Pseudomonas syringae effector HopX1 functions as a cysteine protease that activates gene expression in the JA response pathway through degrading jasmonate ZIM domain (JAZ) proteins independently of the Skip‐Cullin‐F‐box (SCF) complex (Gimenez‐Ibanez et al., 2014). HopZ1a relies on its acetyltransferase activity to acetylate JAZ proteins in advance of proteasome‐dependent degradation (Lewis et al., 2013). AvrB directly targets a pathway consisting of RAR1, HSP90, MPK4, and RIN4, inducing JA responses and increasing plant susceptibility to P. syringae (He et al., 2004; Cui et al., 2010). Not only that, the gene of N. benthamiana related with JA response also was induced by type III secreted effector(s) of R. solanacearum , such as the DS1 gene. DS1‐silenced plants infected by R. solanacearum displayed hyperinduction of PR4 expression (Nakano et al., 2013). The elevated expression of N. benthamiana PDF1.2 in response to RipI suggests that the JA and ET signalling pathways had been activated. Furthermore, RipI could not induce PDF1.2 expression in bHLH93‐silenced plants, implying that bHLH93 functions in signalling pathways mediated by RipI. Further studies are needed to clarify how RipI induces PDF1.2 expression and whether bHLH93 directly regulates JA and ET signalling.
In conclusion, the R. solanacearum effector RipI plays a role in host defence induction, as revealed by the HR induced by transient heterologous expression of RipI in N. benthamiana. Mutagenesis of ripI in GMI1000 increased its virulence on tomato plants. Furthermore, the bHLH93 transcription factor interacted with RipI and was involved in the RipI‐induced defence response in N. benthamiana. These findings provide insight into the role of RipI in virulent R. solanacearum strains in host plants.
Supporting information
ACKNOWLEDGEMENTS
This work was supported by the National Natural Science Foundation of China (31671988, 31872919, 31701752, and 31801696), the Science Foundation of Fujian Provincial Science & Technology Department (2017J01434), and the Scientific Project of Fujian Provincial Education Department (JA15152). We thank Yule Liu (School of Life Sciences, Tsinghua University, Beijing) for providing the pTRV1, pTRV2, and pTRV2‐PDS plasmids.
Zhuo T, Wang X, Chen Z, et al. The Ralstonia solanacearum effector RipI induces a defence reaction by interacting with the bHLH93 transcription factor in Nicotiana benthamiana . Molecular Plant Pathology. 2020;21:999–1004. 10.1111/mpp.12937
Tao Zhuo and Xue Wang contributed equally to this work.
DATA AVAILABILITY STATEMENT
The data that support the findings of this study are available from the corresponding author upon reasonable request.
REFERENCES
- Bruex, A. , Kainkaryam, R.M. , Wieckowski, Y. , Kang, Y.H. , Bernhardt, C. et al (2012) A gene regulatory network for root epidermis cell differentiation in Arabidopsis . PLoS Genetics, 8, e1002446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cui, H. , Wang, Y. , Xue, L. , Chu, J. , Yan, C. , Fu, J. et al (2010) Pseudomonas syringae effector protein AvrB perturbs Arabidopsis hormone signaling by activating MAP kinase 4. Cell Host & Microbe, 7, 164–175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deng, M. , Sun, Y. , Li, P. , Fu, B. , Shen, D. and Lu, Y. (2016) The phytopathogenic virulent effector protein RipI induces apoptosis in budding yeast Saccharomyces cerevisiae . Toxicon, 121, 109–118. [DOI] [PubMed] [Google Scholar]
- Deslandes, L. , Olivier, J. , Peeters, N. , Feng, D.X. , Khounlotham, M. , Boucher, C. et al (2003) Physical interaction between RRS1‐R, a protein conferring resistance to bacterial wilt, and PopP2, a type III effector targeted to the plant nucleus. Proceedings of the National Academy of Sciences of the United States of America, 100, 8024–8029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Genin, S. and Denny, T.P. (2012) Pathogenomics of the Ralstonia solanacearum species complex. Annual Review of Phytopathology, 50, 67–89. [DOI] [PubMed] [Google Scholar]
- Gimenez‐Ibanez, S. , Boter, M. , Fernandez‐Barbero, G. , Chini, A. , Rathjen, J.P. and Solano, R. (2014) The bacterial effector HopX1 targets JAZ transcriptional repressors to activate jasmonate signaling and promote infection in Arabidopsis . PLoS Biology, 12, e1001792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gopalan, S. , Wei, W. and He, S.Y. (1996) hrp gene‐dependent induction of hin1: a plant gene activated rapidly by both harpins and the avrPto gene‐mediated signal. The Plant Journal, 10, 591–600. [DOI] [PubMed] [Google Scholar]
- He, P. , Chintamanani, S. , Chen, Z. , Zhu, L. , Kunkel, B.N. , Alfano, J.R. et al (2004) Activation of a COI1‐dependent pathway in Arabidopsis by Pseudomonas syringae type III effectors and coronatine. The Plant Journal, 37, 589–602. [DOI] [PubMed] [Google Scholar]
- Heim, M.A. , Jakoby, M. , Werber, M. , Martin, C. , Weisshaar, B. and Bailey, P.C. (2003) The basic helix‐loop‐helix transcription factor family in plants: a genome‐wide study of protein structure and functional diversity. Molecular Biology and Evolution, 20, 735–747. [DOI] [PubMed] [Google Scholar]
- Ledent, V. and Vervoort, M. (2001) The basic helix‐loop‐helix protein family: comparative genomics and phylogenetic analysis. Genome Research, 11, 754–770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis, J.D. , Lee, A.H. , Hassan, J.A. , Wan, J. , Hurley, B. , Jhingree, J.R. et al (2013) The Arabidopsis ZED1 pseudokinase is required for ZAR1‐mediated immunity induced by the Pseudomonas syringae type III effector HopZ1a. Proceedings of the National Academy of Sciences of the United States of America, 110, 18722–18727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lonjon, F. , Peeters, N. , Genin, S. and Vailleau, F. (2018) In vitro and in vivo secretion/translocation assays to identify novel Ralstonia solanacearum type 3 effectors In: Medina C. and López‐Baena F. (Eds.) Host–Pathogen Interactions. New York: Humana Press, pp. 209–222. [Google Scholar]
- Morel, A. , Guinard, J. , Lonjon, F. , Sujeeun, L. , Barberis, P. , Genin, S. et al (2018) The eggplant AG91‐25 recognizes the type III‐secreted effector RipAX2 to trigger resistance to bacterial wilt (Ralstonia solanacearum species complex). Molecular Plant Pathology, 19, 2459–2472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nahar, K. , Matsumoto, I. , Taguchi, F. , Inagaki, Y. , Yamamoto, M. , Toyoda, K. et al (2014) Ralstonia solanacearum type III secretion system effector Rip36 induces a hypersensitive response in the nonhost wild eggplant Solanum torvum . Molecular Plant Pathology, 15, 297–303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakano, M. and Mukaihara, T. (2018) Ralstonia solanacearum type III effector RipAL targets chloroplasts and induces jasmonic acid production to suppress salicylic acid‐mediated defense responses in plants. Plant Cell Physiology, 59, 2576–2589. [DOI] [PubMed] [Google Scholar]
- Nakano, M. and Mukaihara, T. (2019) The type III effector RipB from Ralstonia solanacearum RS1000 acts as a major avirulence factor in Nicotiana benthamiana and other Nicotiana species. Molecular Plant Pathology, 20, 1237–1251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakano, M. , Nishihara, M. , Yoshioka, H. , Takahashi, H. , Sawasaki, T. , Ohnishi, K. et al (2013) Suppression of DS1 phosphatidic acid phosphatase confirms resistance to Ralstonia solanacearum in Nicotiana benthamiana . PLoS ONE, 8, e75124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peeters, N. , Carrère, S. , Anisimova, M. , Plener, L. , Cazalé, A.C. and Genin, S. (2013) Repertoire, unified nomenclature and evolution of the type III effector gene set in the Ralstonia solanacearum species complex. BMC Genomics, 14, 859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poueymiro, M. , Cunnac, S. , Barberis, P. , Deslandes, L. , Peeters, N. , Cazale‐Noel, A.C. et al (2009) Two type III secretion system effectors from Ralstonia solanacearum GMI1000 determine host‐range specificity on tobacco. Molecular Plant‐Microbe Interactions, 22, 538–550. [DOI] [PubMed] [Google Scholar]
- Robert‐Seilaniantz, A. , Navarro, L. , Bari, R. and Jones, J.D. (2007) Pathological hormone imbalances. Current Opinion in Plant Biology, 10, 372–379. [DOI] [PubMed] [Google Scholar]
- Salanoubat, M. , Genin, S. , Artiguenave, F. , Gouzy, J. , Mangenot, S. , Arlat, M. et al (2002) Genome sequence of the plant pathogen Ralstonia solanacearum . Nature, 415, 497–502. [DOI] [PubMed] [Google Scholar]
- Sang, Y. , Wang, Y. , Ni, H. , Cazalé, A.C. , She, Y.M. , Peeters, N. et al (2018) The Ralstonia solanacearum type III effector RipAY targets plant redox regulators to suppress immune responses. Molecular Plant Pathology, 19, 129–142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun, Y. , Li, P. , Deng, M. , Shen, D. , Dai, G. , Yao, N. et al (2017) The Ralstonia solanacearum effector RipAK suppresses plant hypersensitive response by inhibiting the activity of host catalases. Cellular Microbiology, 19, e12736. [DOI] [PubMed] [Google Scholar]
- Yu, J. , Ai, G. , Shen, D. , Chai, C. , Jia, Y. , Liu, W. et al (2019) Bioinformatical analysis and prediction of Nicotiana benthamiana bHLH transcription factors in Phytophthora parasitica resistance. Genomics, 111, 473–482. [DOI] [PubMed] [Google Scholar]
- Vos, I.A. , Moritz, L. , Pieterse, C.M. and Van Wees, S. (2015) Impact of hormonal crosstalk on plant resistance and fitness under multi‐attacker conditions. Frontiers in Plant Science, 6, 639. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang, L. , Zhang, F. , Melotto, M. , Yao, J. and He, S.Y. (2017) Jasmonate signaling and manipulation by pathogens and insects. Journal of Experimental Botany, 68, 1371–1385. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.


