Abstract
The sudden outbreak of novel coronavirus has caused a global concern due to its infection rate and mortality. Despite extensive research, there are still no specific drugs or vaccines to combat SARS-CoV-2 infection. Hence, this study was designed to evaluate some plant-based active compounds for drug candidacy against SARS-CoV-2 by using virtual screening methods and various computational analyses. A total of 27 plant metabolites were screened against SARS-CoV-2 main protease proteins (MPP), Nsp9 RNA binding protein, spike receptor binding domain, spike ecto-domain and HR2 domain using a molecular docking approach. Four metabolites, i.e., asiatic acid, avicularin, guajaverin, and withaferin showed maximum binding affinity with all key proteins in terms of lowest global binding energy. The crucial binding sites and drug surface hotspots were unravelled for each viral protein. The top candidates were further employed for ADME (absorption, distribution, metabolism, and excretion) analysis to investigate their drug profiles. Results suggest that none of the compounds render any undesirable consequences that could reduce their drug likeness properties. The analysis of toxicity pattern revealed no significant tumorigenic, mutagenic, irritating, or reproductive effects by the compounds. However, withaferin was comparatively toxic among the top four candidates with considerable cytotoxicity and immunotoxicity. Most of the target class by top drug candidates belonged to enzyme groups (e.g. oxidoreductases hydrolases, phosphatases). Moreover, results of drug similarity prediction revealed two approved structural analogs of Asiatic acid i.e. Hydrocortisone (DB00741) (previously used for SARS-CoV-1 and MERS) and Dinoprost-tromethamine (DB01160) from DrugBank. In addition, two other biologically active compounds, Mupirocin (DB00410) and Simvastatin (DB00641) could be an option for the treatment of viral infections. The study may pave the way to develop effective medications and preventive measure against SARS-CoV-2. Due to the encouraging results, we highly recommend further in vivo trials for the experimental validation of our findings.
Keywords: SARS-CoV-2, Plant metabolites, Main protease proteins, Molecular docking, ADME analysis, Drug target
Graphical abstract
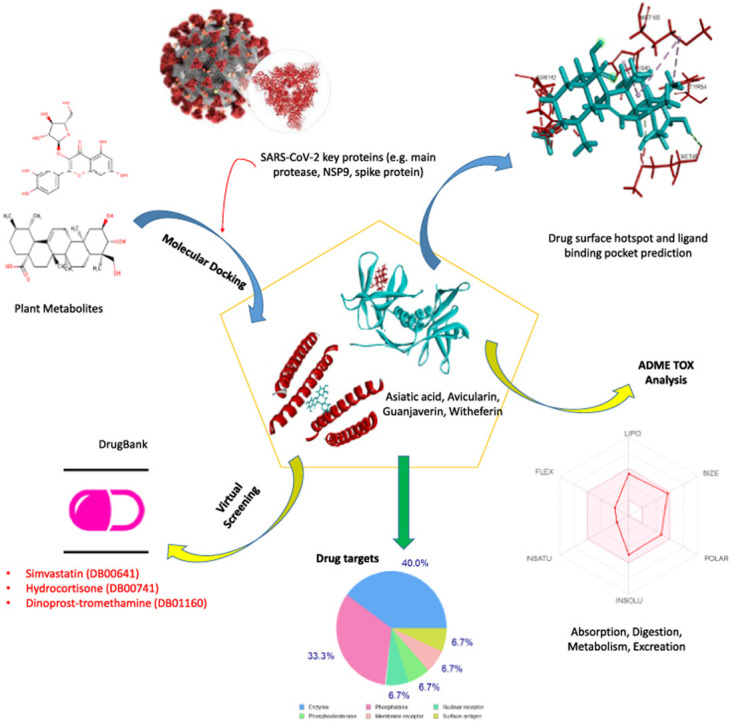
Highlights
-
•
Asiatic acid, Avicularin, Guajaverin & Withaferin showed maximum binding affinity.
-
•
The crucial binding sites & drug surface hotspots were unravelled for key viral protein.
-
•
ADME analysis revealed that top candidates do not show any undesirable consequences.
-
•
Witheferin was comparatively toxic with considerable cytotoxicity and immunotoxicity.
-
•
Hydrocortisone and Simvastatin could be an option to treat SARS-CoV-2 infections.
1. Introduction
The sudden outbreak of novel coronavirus (SARS-CoV-2) infection emanated from Wuhan, China and spread throughout the world excepting a few countries to date [1]. The virus is responsible for causing novel disease, which WHO officially called COVID-19. As of April 23, 2020, World Health Organization (WHO) estimated that new coronavirus touched 213 countries, areas or territories [2,3]. The infection rate is increasing. However, the fatality rate of SARS-CoV-2 (3.4%) estimated by WHO is lower than previous fatal diseases SARS and MERS, which had 9.6% and 35% death rates, respectively [4,5].
Coronaviruses are enveloped, positive single-stranded RNA viruses with large genome size ranging from 26 kb to 32 kb. These viruses are representative of four subfamilies, which include alpha-, beta-, gamma- and delta-coronaviruses. SARS-CoV-2 showed more sequence similarity with SARS-CoV than MERS-CoV when genome sequences of these mentioned viruses were compared [6]. But they also exhibit dissimilarities that can influence their process of pathogenesis [7,8]. SARS-CoV-2 infects humans through the same entry point of the ACE receptor which is expressed in the respiratory tract [9,10]. However, among various proteins, four proteins are commonly found in the structure of all coronaviruses representing spike (S), envelope (E), membrane (M), and nucleocapsid (N) [8]. The initial and important stage of viral entry into host cell is receptor recognition [11]. The assembly of viral particle involves M protein and E protein, while virus binding and entrance into host cell take place by S protein with the assistance of SARS-CoV-2 angiotensin-converting enzyme [10,12].
The coronavirus (SARS-CoV-2) belongs to the family of Beta-coronaviruses, which are responsible for causing severe human respiratory syndrome [3,13]. The virus is spread mainly through community transmission, while SARS and MERS affect people via nosocomial spread [14]. It can transmit from one individual to other by respiratory droplets. SARS-CoV-2 infected patients have general signs and symptoms, suffering initially from common flu-like fever, sputum production, dyspnoea, headache, sore throat/pharyngalgia, and diarrhoea, which may further lead to express life-threatening symptoms including fatal pneumonia [15]. COVID-19 affected patients, either symptomatic or asymptomatic, were detected with the nose area containing a higher viral load than in throat [16]. A critically ill patient has a series of complexities with progression of disease.
The efficacy and safety of antivirals require evaluation by clinical trial [3]. There is no efficient, safe, and specific potential therapeutic to be approved for rapid remedy of this new respiratory syndrome to date [17,18]. Clinical trials of some drugs have been started, yet till now, only a few candidates have shown some efficacy in in vitro studies [19]. Not many have progressed to randomized animal or human trials; hence they may have limited use to counter infection [19]. Many countries and some pharmaceutical companies announced their headway and programs to develop vaccines (e.g. subunit, mRNA, DNA, live-vector vaccine) against the virus. But the developmental process of making human vaccine from concept to licensure may require years [20]. As the epidemic is still spreading, medicinal plants may be alternatively used in making drugs as early as possible. Several scientific researchers reported the helpfulness of plants due to their medicinal value and therapeutic uses as drugs from the ancient times [21]. Plant-derived active compounds of different plant parts are useful for treating diseases including diarrhoea, headache, and inflammation, and bacterial and fungal infections. From prehistoric times, traditional people utilized these plants for the remedial purposes of health deteriorating diseases because of the existence of numerous phytochemicals [22]. Various limitations are associated with modern treatment options including drug-resistance, severe side effects, adverse toxicity profiles, complicated medication etc. Natural products have the potential to form the basis of holistic health care [23]. The properties of antioxidants render medicinal plants to be effective in treating life-threatening diseases such as cancer, Alzheimer's disease, diabetes, malaria, and cardiac diseases (Table 1 ) while minimizing drug toxicity [24].
Table 1.
List of plant metabolites used in the study with respective source and activities.
| Metabolites | PubChem CID | Class | Source | Activities | References |
|---|---|---|---|---|---|
| Allicin | 65036 | S-containing compound | Allium sativum | Antimicrorial, antiviral Antioxidant, anti-cancer activity | [25] |
| Andrographolide | 5318517 | Diterpenoid labdane | Andrographis paniculata | antioxidant, anti-inflammatory, and anti-cancer | [26] |
| Apigenin | 5280443 | Flavonoid | Vegetable and fruit | Effective in cancer, depression, diabetes & Alzheimer's disease, | [27] |
| Asiatic acid | 119034 | Aglycone type pentacyclic triterpenoids | Centella asiatica | Antioxidant, cardioprotective, anti-inflammatory, antitumor, neuroprotective, antimicrobial | [28] |
| Avicularin | 5490064 | quercetin-3-a-L arabinofuranoside (flavonoid) | Psidium guyava, Lespedeza cuneata | anti-inflammatory, anti-oxidant, hepatoprotective activity | [29] |
| Capsaicin | 1548943 | Alkaloid | Capsicum genus | Pruritis, pain relief, non-steroidal anti-inflammatory drug induced gastritis | [30] |
| Chavibetol | 596375 | Phenylpropanoid | Piper betle | immunomodulatory, radical scavenging | [31] |
| Cinnamic acid | 444539 | Aromatic carboxylic acids | Cinnamomum species | Antibacterial, antifungal, antimalarial, antitubercular | [32] |
| Curcumin | 969516 | Polyphenolic compound | Curcuma longa | antibacterial, anti-inflammatory antiviral, antioxidant, anti-arthritis & anti-cancer activity | [33] |
| Eugenol | 3314 | Phenylpropanoid | Ocimum tenuiflorum, Eugenia caryophyllata | antimicrobial, anti-inflammatory, analgesic and antioxidant | [34] |
| Arjunone | 14034821 | Flavonoids | Terminalia arjuna | Arjunone and other compounds have role in antioxidant, antiatherogenic, anti-inflammatory, anti-carcinogenic activity | [35] |
| Galangin | 5281616 | Flavonol | Honey, Alpinia officinarum, propolis | Anti-cancer, anti-mutagenic, anti-oxidative, radical scavenging etc. | [36] |
| Gentisic acid | 3469 | Phenolic acid |
Gentiana, Citrus, H. rosa-sinensis, O. europaea, S. indicum |
Antioxidant, neuroprotective, antiinflammatory, hepatoprotective, antimicrobial activities | [37] |
| Guajaverin | 5481224 | Flavonoid | Psidium guyava | Anti-plaque activity | [38] |
| Kaempferol | 5280863 | Flavonoid aglycone | Vegetable and fruit | Anti-inflammatory, antioxidant, antimicrobial, antitumor, cardioprotective, and antidiabetic activities | [39] |
| Luteolin | 5280445 | Flavonoid | Carrots, celery peppers, olive peppermint | Anticancer, antioxidant, antimicrobial, anti-inflammatory, and activities | [40] |
| m-Coumaric acid | 637541 | Phenolic acid | Solanum nigrum | Role in pharmacological activities | [41] |
| Piperic acid | 5370536 | Alkaloid | Piper nigrum | No known function | [42] |
| Piperine | 638024 | Alkaloid | Piper spp. | Anticancer, antimicrobial, antimalarial | [42] |
| Quercetine | 5280343 | Flavonoid | Diverse plant species | Antioxidant, cardiovascular, antiviral, anti-inflammatory, anticancer, antimicrobial | [43] |
| Swertiamarin | 442435 | Secoiridoid glycoside | Swertia chirata | Anti-arthritic, anti-diabetic Cardio-protective, Anticancer, Anti-hepatitis, Antibacterial, anti-atherosclerotic | [44] |
| Swertinin | 5491517 | Secoiridoid glycoside | Swertia chirata | Role in pharmacological activities | [45] |
| Thymoquinone | 10281 | Monoterpene | Nigella sativa | Anti-oxidant and anti-inflammatory properties, Anti-microbial, Anti-arthritic, anti-cancer efficacy | [46] |
| Vincamine | 15376 | Alkaloid | Catharanthus roseus, Vinca minor | Cerebral disorders, antiulcer activity, cerebrovascular insufficiencies | [47] |
| Vitexin | 5280441 | Apigenin flavone glucoside | Crataegus species | Anti-inflammatory effects, anti-oxidant effects, anti-carcinogenic effects, anti-viral effects | [48] |
| Withaferin | 265237 | Steroidal lactone | Withania somnifera | Anti-cancer, adaptogenic, anti-stress, immunomodulatory, anti-inflammatory, anti-tumor, cardioprotective, and neuroprotective activities. | [49] |
| Zingiberene | 92776 | Isoprenoids | Zingiber Officinale | Anti-ulcer, antibacterial, cytoxic effect | [50] |
The expansion of natural product as new medicine or drug to resist the emerging virus SARS-CoV-2 could be done to bypass the side effects of synthetic drugs. Therefore, the study aimed at evaluating some plant-based active compound for drug candidacy against SARS-CoV-2 through virtual screening methods and various computational investigations.
2. Material and methods
2.1. Retrieval of SARS-CoV-2 proteins/protein-domains and plant metabolites
The 3D structures of SARS-CoV-2 main proteases (6W63, 6LU7), Nsp9 (Non-structural protein-9) RNA binding protein (6W4B), spike receptor binding domain (6M0J), spike ecto-domain (6VYB), and HR2 Domain (6LVN) were retrieved from the RCSB Protein Data Bank [51]. A total 27 plant metabolites belonging to different classes were extracted from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) in SDS (3D) format (Table 1) [52]. The structures were further converted into the PDB format by Open Babel v2.3 [53].
2.2. Screening of plant metabolites against SARS-CoV-2 proteins/protein-domains
Molecular docking is an effective approach for screening suitable therapeutics against specific drug target of deadly pathogens [54]. This powerful tool is used to model the interaction between small ligands and macromolecules, thereby paving the way for drug discovery [55]. The binding affinity of 27 plant metabolites with different SARS-CoV-2 proteins/protein domains (drug targets/macromolecules) were determined by using the PatchDock server [56]. Recently, alpha-ketoamide (CID 6482451) has been suggested as a SARS-CoV-2 MPP inhibitor by experimental study [57]. The ligand was used as a positive control for the present study, and employed for docking analysis against all six macromolecules. The docked complexes were further refined via the FireDock refinement tool [58]. The ligand bond complexes were visualized by Discovery Studio v3.1 and PyMOL v2.0 [59,60].
2.3. Analysis of drug surface hotspot and ligand binding pocket prediction
The drug surface hotspot of SARS-CoV-2 proteins was analysed by investigating the docked complexes with the top metabolites using LigPlot+, Discovery Studio and PyMOL v.2.0 software [59,60]. Binding patterns of asiatic acid, avicularin, guajaverin, and withaferin with six macromolecules were allowed for comparative structural analysis. Moreover, interaction of Alpha-ketoamide with the studied proteins were also investigated.
2.4. Drug profile analysis of top metabolites
Absorption, distribution, metabolism, and excretion (ADME) are four major criteria that influence the drug levels and kinetics of drug exposure to the tissues within an organism. The pharmacological activity and performance of a drug is largely controlled by these parameters [61]. The SwissADME server was used to assess the ADME properties of top four metabolites [62]. The BOILED-Egg model was employed to calculate the blood-brain barrier (BBB) in the studied compounds [63]. The relative toxicity of top drug candidates were analysed via the ProToxII server [64]. This popular webserver efficiently predicts various toxicity endpoints by incorporating molecular similarity, fragment tendency and fragment similarity methods. The server also predicted the oral toxicity based on the analysis of two-dimensional (2D) similarity to compounds with a known median lethal doses (LD50). The set used for the prediction consists of approximately 38,000 unique compounds with known oral LD50 values measured in rodents [65]. Additionally, OSIRIS Property Explorer were employed to investigate the undesired effects of these compounds [66].
2.5. Prediction of drug targets and available drug molecules from DrugBank
SwissTargetPrediction was utilized to estimate the possible macromolecular targets of predicted drug candidates [67]. The server predicts based on a combination of 2D and 3D similarity with a library of 370000 known bioactive compounds on approximately 3000 proteins. Moreover, the SwissSimilarity web tools were used to identify potential drug molecules against SARS-CoV-2 based on homology screening of predicted top drug candidates. The server allowed ligand-based virtual screening of several libraries of small molecules to find approved, experimental, or commercially available drugs from DrugBank using different approaches including FP2 fingerprints, electroshape, spectrophores, and align-IT [68].
3. Results
3.1. Screening of plant metabolites against SARS-CoV-2
All of the retrieved structures of SARS-CoV-2 proteins/protein-domains (macromolecules) and plant metabolites (ligands) were optimized and employed for molecular docking to predict the affinity between the above-mentioned ligands and the macromolecules. The metabolites were ranked based on global binding energy and the results depict that the top four scorers (metabolites) were the same for each of the macromolecules in terms of minimum binding energy (Table 2 and Supplementary File 1). In each case, asiatic acid, avicularin, guajaverin, and withaferin showed the best binding interactions with six studied macromolecules (Fig. 1 and Table 2). Moreover, asiatic acid exhibited the highest binding affinity with SARS-CoV-2 main protease (−53.05 kcal/mol), Nsp9 RNA binding protein (−50.04 kcal/mol), and spike ecto-domain (60.68 kcal/mol) (Fig. 2 and Table 2), while guajaverin bound with the spike receptor binding domain and HR2 Domain with a binding energy of −47.34 kcal/mol and −28.73 kcal/mol, respectively (Fig. 3 and Table 2).
Table 2.
Analysis of global binding energy and interaction sites of the screened top 4 metabolites.
| Macromolecules | Ligands | Global Energy | ACE | Score | Area | Ligand binding residues |
|---|---|---|---|---|---|---|
| 6W63 | α-ketoamide (Control) | −56.92 | −16.84 | 4560 | 526.40 | Asp197, Leu272, Gly275, Leu286, Leu287, Asp289 |
| Asiatic acid | −53.05 | −15.26 | 4916 | 577.10 | His41, Met49, Tyr54, Asn142, Met165 | |
| Avicularin | −48.62 | −18.50 | 4694 | 532.10 | Thr25, Thr26, His41, Cys44, Ser46, Met49, Gly143, Cys145 | |
| Guajaverin | −48.48 | −15.12 | 4450 | 497.50 | Thr25, His41, Cys44, Met49, Asn142, Cys145, Met165, Asp187, Arg188 | |
| Withaferin | −48.46 | −14.08 | 4984 | 597.40 | His41, Met49, Met165, Pro168, Ala191 | |
| 6W4B | α-ketoamide (Control) | −48.60 | −16.39 | 4458 | 504.60 | Phe41, Trp54, Ile66, Thr68, Glu69 |
| Asiatic acid | −50.04 | −16.37 | 4998 | 564.20 | Met13, Gly39, Arg40, Phe41, Val42, Phe57, Pro58, Ile66 | |
| Withaferin | −47.95 | −13.30 | 4896 | 570.40 | Arg40, Val42, Phe57, Pro58,Lys59, Ser60, Ile66 | |
| Guajaverin | −42.72 | −10.63 | 4548 | 641.40 | Asn1, Asn2, Glu3, Gln50, Pro72, Pro73 | |
| Avicularin | −39.83 | −23.80 | 4556 | 514.50 | Met49, Met165, Glu166, Thr190 | |
| 6VYB | α-ketoamide (Control) | −63.94 | −17.32 | 5728 | 705.10 | Thr547, Gly548, Thr549,Asp745, Val976 |
| Asiatic acid | −60.68 | −22.33 | 6276 | 771.50 | Phe338, Ala363, Tyr365, Leu368, Cys379, Pro384, Leu387, Leu390, Phe392, Val395, Cys432, Ile434, Leu513, Val524 | |
| Withaferin | −60.19 | −20.49 | 5760 | 793.10 | Ile410, Pro412, Leu425, Pro426, Cys432, Val433, Phe464, Val512, Leu513 | |
| Guajaverin | −55.24 | −17.51 | 5208 | 659.20 | Ile410, Pro412, Lys424, Gly431, Cys432, Val433, Val512 | |
| Avicularin | −52.93 | −17.15 | 5474 | 683.30 | Ala411, Pro412, Leu425, Cys432, Val433, Val512 | |
| 6LVN | α-ketoamide (Control) | −25.52 | −2.71 | 4318 | 564.20 | Ile16, Asn20, Lys24, Asn27, Glu28 |
| Guajaverin | −28.73 | −2.13 | 3696 | 443.50 | Asp17, Arg18, Glu21, Lys24 | |
| Withaferin | −28.11 | −1.24 | 4376 | 507.70 | Lys14, Lys24, Arg18 | |
| Asiatic acid | −27.58 | −1.12 | 4366 | 500.30 | Lys14, Lys24, Asp17, Arg18, Glu21 | |
| Avicularin | −26.48 | −1.22 | 3986 | 465.10 | Asp17, Arg18, Glu21, Asn20 | |
| 6M0J | α-ketoamide (Control) | −60.50 | −9.34 | 5374 | 655.40 | Lys94, Tyr196, Asp206, Glu208, Val209, Asn210 |
| Guajaverin | −47.34 | −11.22 | 4554 | 575.60 | Leu95, Gln98, Ala99, Glu208, Asn210, Ala396, Lys562, Trp566 | |
| Withaferin | −46.84 | −11.13 | 5598 | 640.50 | Leu95, Gln98, His195, Tyr196, Lys562 | |
| Asiatic acid | −45.69 | −13.09 | 5978 | 691.70 | Leu95, Gln98, Ala99, Tyr202, Asp206, Glu208, Val209, Ala396, Lys562, Pro565, Trp566 | |
| Avicularin | −43.13 | −11.09 | 5232 | 604.20 | Lys94, Leu95, Tyr196, Val209, Asn210, Ala396, Lys562 | |
| 6LU7 | α-ketoamide (Control) | −56.13 | −15.07 | 4578 | 492.00 | Asp197, Lys236, Tyr237, Leu272 |
| Avicularin | −54.04 | −14.77 | 4584 | 520.60 | His41, Met49, His164, Met165, Glu166, Pro168, Thr190, Asp187, Arg188, Gln189 | |
| Guajaverin | −51.69 | −12.92 | 4182 | 515.50 | Met49, Phe140, His163, His164, Met165, Glu166, Arg188, Gln192 | |
| Withaferin | −47.08 | −14.06 | 4708 | 560.60 | Thr24, Met49, His41, Cys145, Met165, Arg188 | |
| Asiatic acid | −43.52 | −13.90 | 5050 | 562.20 | Met49, Leu141, Asn142, Ser144, Cys145, His163, Glu166 |
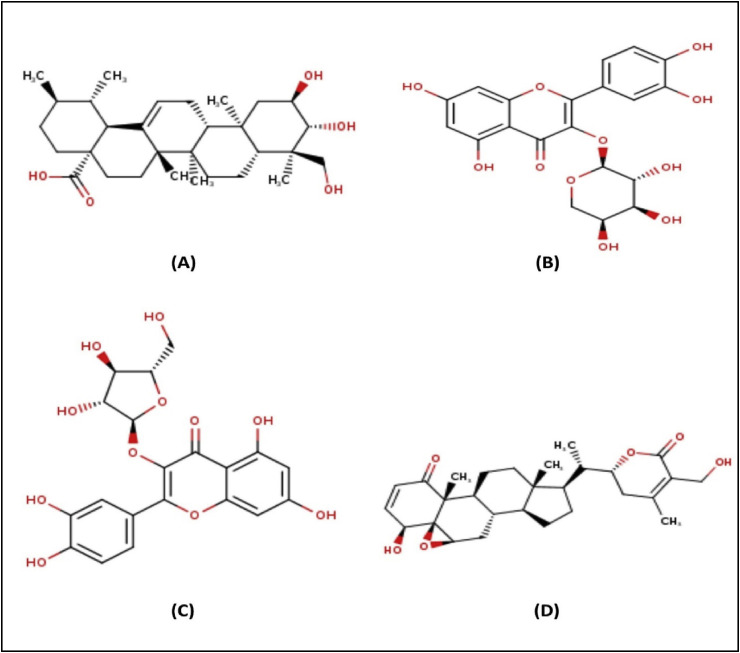
Fig. 1.
Chemical structures of asiatic acid (A), guajaverin (B), avicularin (C) and withaferin (D).
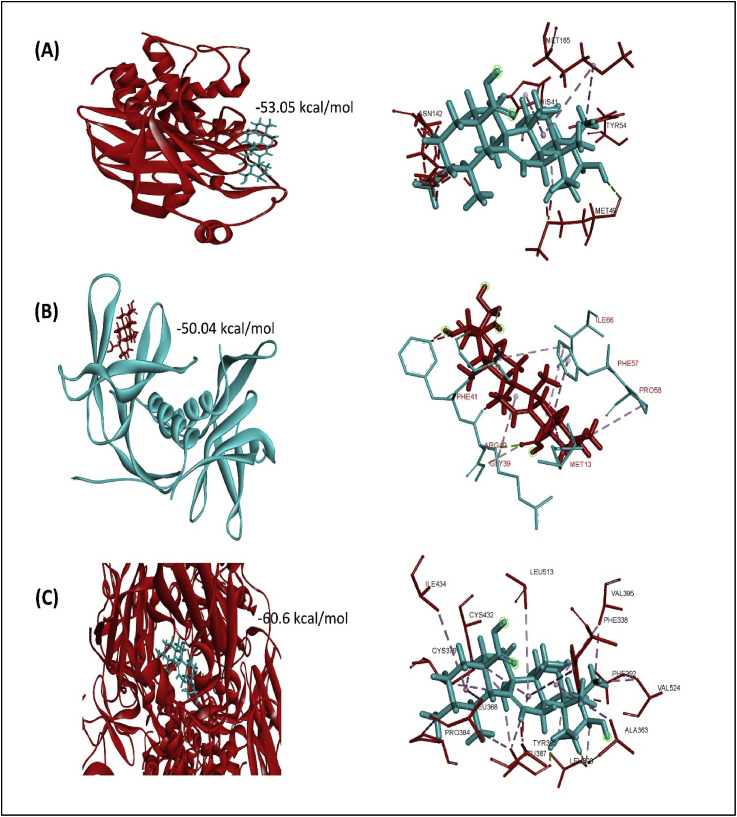
Fig. 2.
Molecular interaction of asiatic acid with SARS-CoV-2 main protease (A), Nsp9 RNA binding protein (B), and spike ecto-domain (C).
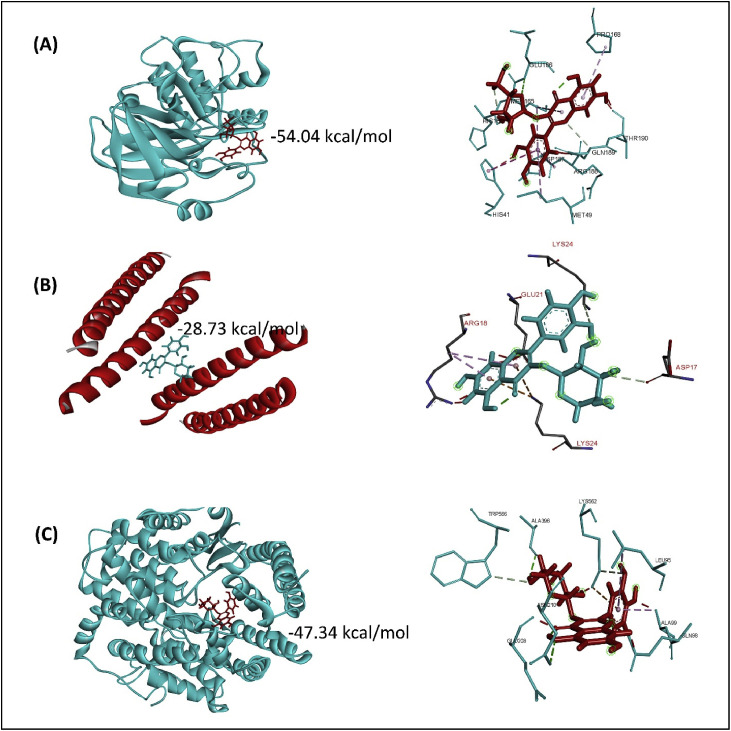
Fig. 3.
Molecular interaction of SARS-CoV-2 main protease with avicularin (A), HR2 domain with guajaverin (B), and spike receptor-binding domain with guajaverin (C).
3.2. Analysis of drug surface hotspot and ligand binding pocket prediction
The structural conformation of the docked complex was analysed to unravel the drug surface hotspot of studied SARS-CoV-2 proteins. The ligand binding pattern and interacting residues with their respective positions were investigated (Table 2). Results revealed that the amino acids from 41 to 54 and 142–190 positions were crucial for the binding interactions of SARS-CoV-2 main protease protein (6W63). Moreover, His41, Cys44, Met49, Asn142, Cys145, Met165 were involved in maximum cases to form the docked complexes. The ligands showed highest binding affinity for 39–73 and 142–166 regions of Nsp9 RNA binding protein (6W4B). Again, the residues from 94 to 99 and 563–566 regions were identified as top surface hotspots for spike receptor binding domain (6M0J) where the position Lys94, Leu95, Tyr196, Lys562, Trp566 were most dominant (Table 2).
3.3. ADME analysis of top drug candidates
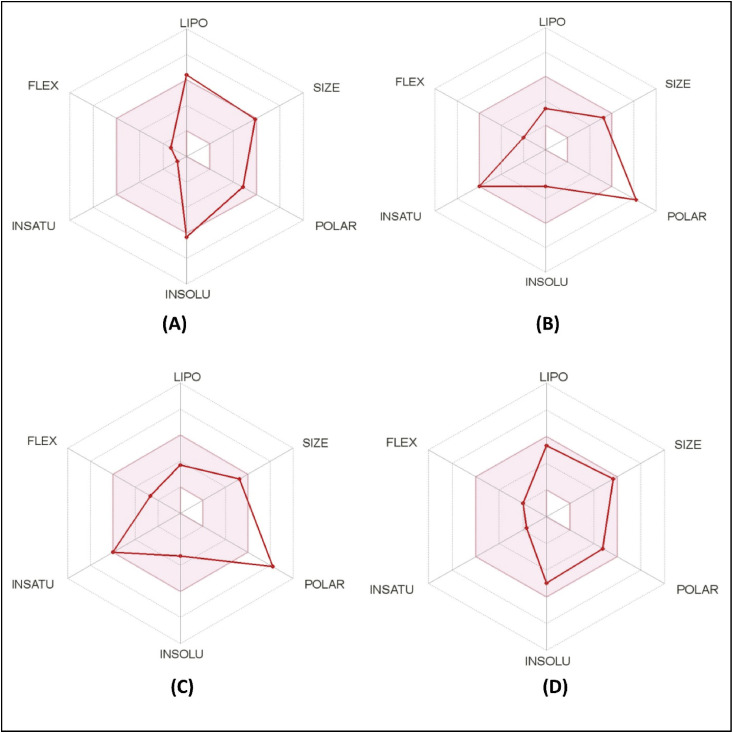
Different ADME properties, i.e., physicochemical parameters, pharmacokinetics, lipophilicity, water solubility, medicinal chemistry of top drug candidates were estimated to evaluate their drug profiles (Fig. 4 and Table 3 ). Analysis of inhibition effects with different CYP isoforms (CYP1A2, CYP2D6, CYP2C9, CYP2C19, CYP3A4) revealed that none of the candidates had such an interaction possibility with any cytochromes P450 isoforms. GI absorption was found higher for asiatic acid and withaferin, while lower for guajaverin and avicularin. Moreover, blood-brain barrier (BBB) permeation was calculated by the BOILED-Egg model, which revealed no BBB permeant among the studied top drug candidates. Each candidate was water soluble from a moderate to high level, while guajaverin and avicularin showed maximum solubility (Table 3).
Fig. 4.
ADME analysis of top four metabolites; A: Asiatic acid, B: Guajaverin, C: Avicularin, and D: Withaferin.
Table 3.
Drug profile and ADME analysis of the top four metabolites.
| Parameter | Top Main Protease Protein Inhibitors of SARS-CoV-2 |
||||
|---|---|---|---|---|---|
| Asiatic acid | Guajaverin | Avicularin | Withaferin | ||
| Physicochemical parameters | Formula | C30H48O5 | C20H18O11 | C20H18O11 | C28H38O6 |
| Molecular weight | 488.70 g/mol | 434.35 g/mol | 434.35 g/mol | 470.60 g/mol | |
| No. H-bond acceptor | 5 | 11 | 11 | 6 | |
| No. H-bond donors | 4 | 7 | 7 | 2 | |
| Molar Refractivity | 139.24 | 104.19 | 104.19 | 127.49 | |
| TPSA | 97.99 Å2 | 190.28 Å2 | 190.28 Å2 | 96.36 Å2 | |
| Lipophilicity | Log Po/w (iLOGP) | 2.95 | 1.77 | 1.86 | 3.24 |
| Log Po/w (XLOGP3) | 5.70 | 0.43 | 0.98 | 3.83 | |
| Log Po/w (WLOGP) | 5.03 | 0.10 | 0.10 | 3.35 | |
| Log Po/w (MLOGP) | 4.14 | −2.06 | −2.06 | 2.75 | |
| Log Po/w (SILICOS-IT) | 3.96 | −0.10 | 0.06 | 3.93 | |
| Consensus Log Po/w | 4.36 | 0.03 | 0.19 | 3.42 | |
| Pharmacokinetics | GI absorption | High | Low | Low | High |
| BBB permeant | No | No | No | No | |
| P-gp substrate | Yes | No | No | Yes | |
| CYP1A2 inhibitor | No | No | No | No | |
| CYP2C19 inhibitor | No | No | No | No | |
| CYP2C9 inhibitor | No | No | No | No | |
| CYP2D6 inhibitor | No | No | No | No | |
| CYP3A4 inhibitor | No | No | No | No | |
| Log Kp (skin permeation) | - 5.23 cm/s | −8.64 cm/s | −8.25 cm/s | −6.45 cm/s | |
| Water Solubility | Log S (ESOL) | −6.33 | −2.99 | −3.27 | −4.97 |
| Solubility | 2.29e-4 mg/ml; 4.69e-7 mol/l |
4.47e-01 mg/ml; 1.03e-03 mol/l | 2.34e-01 mg/ml; 5.39e-04 mol/l | 5.01e-03 mg/ml; 1.07e-05 mol/l | |
| Class | Poorly soluble | Soluble | Soluble | Moderately soluble | |
| Log S (SILICOS-IT) | −4.28 | −1.94 | −2.07 | −3.79 | |
| Solubility | 2.59e-2 mg/ml; 5.31e-05 mol/l | 4.96e+00 mg/ml; 1.14e-02 mol/l | 3.71e+0 mg/ml; 8.55e-3 mol/l | 7.54e-02 mg/ml; 1.60e-04 mol/l | |
| Class | Moderately soluble | Soluble | Soluble | Soluble | |
| Medicinal Chemistry | Leadlikeness | No; 2 violations: MW > 350, XLOGP3>3.5 | No; 1 violation: MW > 350 | No; 1 violation: MW > 350 | No; 2 violations: MW > 350, XLOGP3>3.5 |
| Bioavailability Score | 0.56 | 0.17 | 0.17 | 0.55 | |
| PAINS | 0 alert | 1 alert: catechol_A | 1 alert: catechol_A | 0 alert | |
| Synthetic accessibility | 6.56 | 5.05 | 5.04 | 6.83 | |
3.4. Toxicity pattern analysis of top drug candidates
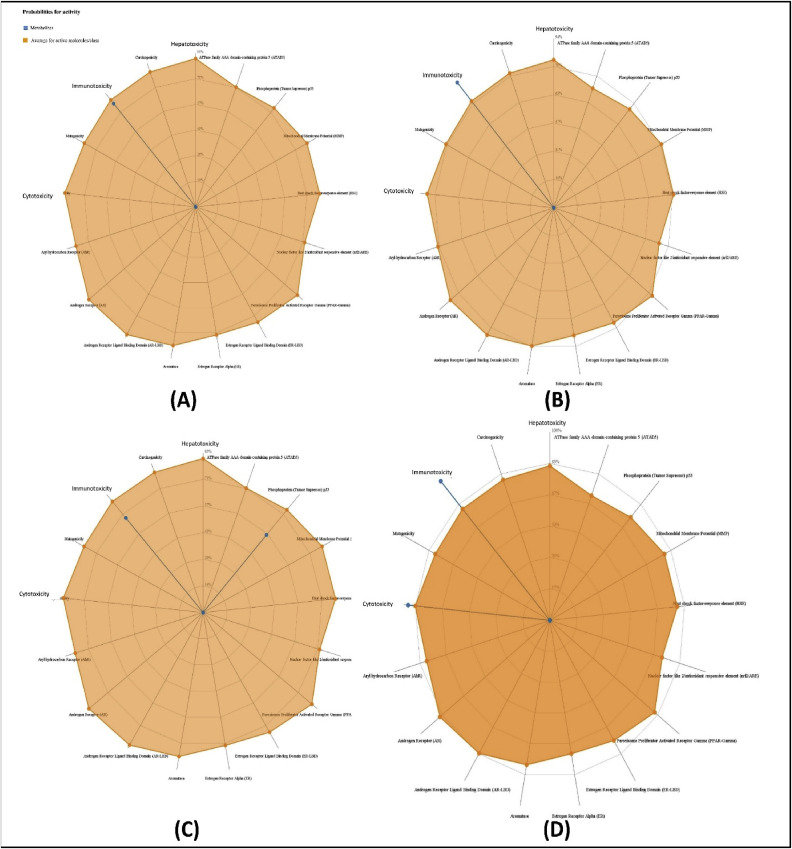
Prediction of various toxicity endpoints such as acute toxicity, hepatotoxicity, cytotoxicity, carcinogenicity, mutagenicity, immunotoxicity, adverse outcomes (Tox21) pathways and toxicity targets were analysed (Table 4 ). Results revealed that guajaverin and avicularin fell in the category of toxicity class 5, while the predicted toxicity group for Asiatic acid and withaferin were 4 and 2 respectively (the lower the class the higher the toxicity). Estimated LD50 for asiatic acid, avicularin, guajaverin and withaferin were 2000, 5000, 5000 and 7 mg/kg respectively. The toxicity radar in Fig. 5 illustrates the confidence of positive toxicity results compared to the average of its class. None of the compounds showed any undesired effects such as tumorigenicity, mutagenicity, irritating, or reproductive effects. Withaferin, however, was found to be relatively toxic among the four candidates, with considerable cytotoxicity and immunotoxicity (Fig. 5).
Table 4.
Toxicity model reports of the top four drug candidates.
| Classification | Target | Prediction and Probability |
|||
|---|---|---|---|---|---|
| Asiatic Acid | Aviculerin | Guajaverin | Withaferin | ||
| Organ toxicity | Hepatotoxicity | Inactive (0.91) | Inactive (0.80) | Inactive (0.80) | Inactive (0.93) |
| Toxicity end points | Carcinogenicity | Inactive (0.70) | Inactive (0.79) | Inactive (0.79) | Inactive (0.55) |
| Toxicity end points | Immunotoxicity | Active (0.77) | Active (0.68) | Active (0.93) | Active (0.99) |
| Toxicity end points | Mutagenicity | Inactive (0.81) | Inactive (0.73) | Inactive (0.79) | Inactive (0.79) |
| Toxicity end points | Cytotoxicity | Inactive (0.73) | Inactive (0.72) | Inactive (0.69) | Active (0.87) |
| Tox21-Nuclear receptor signalling pathways | Aryl hydrocarbon Receptor (AhR) | Inactive (0.99) | Inactive (0.85) | Inactive (0.90) | Inactive (0.98) |
| Tox21-Nuclear receptor signalling pathways | Androgen Receptor (AR) | Inactive (0.59) | Inactive (0.92) | Inactive (0.96) | Inactive (0.63) |
| Tox21-Nuclear receptor signalling pathways | Androgen Receptor Ligand Binding Domain (AR-LBD) | Inactive (0.51) | Inactive (0.98) | Inactive (0.97) | Inactive (0.54) |
| Tox21-Nuclear receptor signalling pathways | Aromatase | Inactive (0.91) | Inactive (0.98) | Inactive (0.97) | Inactive (0.80) |
| Tox21-Nuclear receptor signalling pathways | Estrogen Receptor Alpha (ER) | Inactive (0.73) | Inactive (0.85) | Inactive (0.92) | Inactive (0.60) |
| Tox21-Nuclear receptor signalling pathways | Estrogen Receptor Ligand Binding Domain (ER-LBD) | Inactive (0.97) | Inactive (0.99) | Inactive (0.99) | Inactive (0.98) |
| Tox21-Nuclear receptor signalling pathways | Peroxisome Proliferator Activated Receptor Gamma (PPAR-γ) | Inactive (0.97) | Inactive (0.93) | Inactive (0.94) | Inactive (0.91) |
| Tox21-Stress response pathways | Nuclear factor (erythroid-derived 2)-like 2/antioxidant responsive element | Inactive (0.89) | Inactive (0.91) | Inactive (0.94) | Inactive (0.86) |
| Tox21-Stress response pathways | Heat shock factor response element (HSE) | Inactive (0.89) | Inactive (0.91) | Inactive (0.94) | Inactive (0.86) |
| Tox21-Stress response pathways | Mitochondrial Membrane Potential (MMP) | Inactive (0.85) | Inactive (0.89) | Inactive (0.89) | Inactive (0.80) |
| Tox21-Stress response pathways | Phosphoprotein (Tumor Supressor) p53 | Inactive (0.93) | Active (0.55) | Inactive (0.72) | Inactive (0.75) |
| Tox21-Stress response pathways | ATPase family AAA domain-containing protein 5 (ATAD5) | Inactive (0.96) | Inactive (0.96) | Inactive (0.96) | Inactive (0.94) |
Fig. 5.
Toxicity patterns of the top four drug candidates; A: Asiatic acid, B: Guajaverin, C: Avicularin, and D: Withaferin.
3.5. Prediction of drug targets and available drug molecules from DrugBank
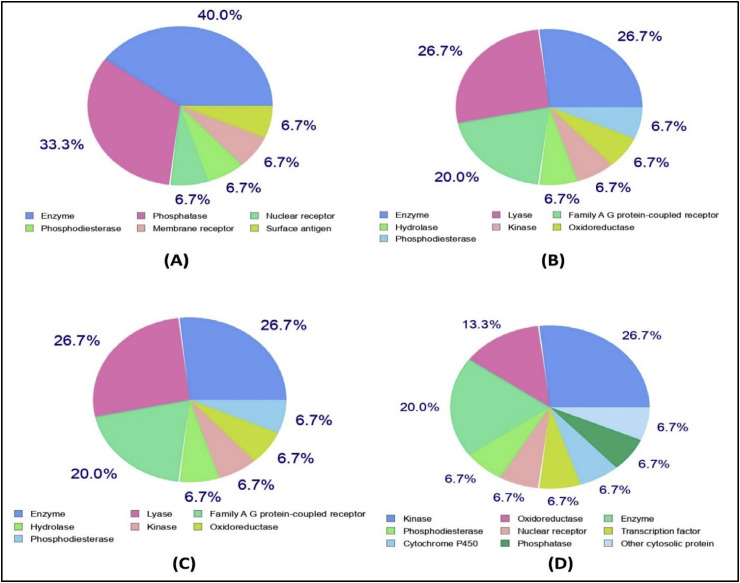
Most of the target class belonged to enzymes, kinase proteins, oxidoreductases (i.e. aldose reductase, aldo-keto reductase), phosphatases and lyases (i.e. carbonic anhydrase) (Fig. 6 and Table 5 ). Ligand-based virtual screening was performed to predict biologically active small compounds against SARS-CoV-2 from DrugBank. Two approved drugs, Hydrocortisone (DB00741) and Dinoprost-tromethamine (DB01160) were found analogous to asiatic acid with prediction scores of 50.52 and 50.53, respectively. Moreover, results revealed the similarity of Mupirocin (DB00410) and Simvastatin (DB00641) with withaferin, with a high prediction score (Table 6 ). The findings suggest that these could be potential drug candidates against SARS-CoV-2, thus requiring further experimental trials.
Fig. 6.
Prediction of drug targets for asiatic acid (A), guajaverin (B), avicularin (C), and withaferin (D).
Table 5.
Predicted drug targets for asiatic acid, guajaverin, aviculerin, and withaferin.
| Metab-olites | Drug Targets | Common Name | Uniprot ID | ChEMBL ID | Target Class | Probability |
|---|---|---|---|---|---|---|
| Asiatic Acid | Aldo-keto reductase family 1 member B10 | AKR1B10 | O60218 | CHEMBL5983 | Enzyme | |
| Protein-tyrosine phosphatase 1B | PTPN1 | P18031 | CHEMBL335 | Phosphatase | ||
| 11-β-hydroxysteroid dehydrogenase 1 | HSD11B1 | P28845 | CHEMBL4235 | Enzyme |  |
|
| DNA polymerase beta | POLB | P06746 | CHEMBL2392 | Enzyme |  |
|
| T-cell protein-tyrosine phosphatase | PTPN2 | P17706 | CHEMBL3807 | Phosphatase |  |
|
| Phospholipase A2 group 1B | PLA2G1B | P04054 | CHEMBL4426 | Enzyme |  |
|
| Guajaverin & Aviculerin | Aldose reductase | AKR1B1 | P15121 | CHEMBL1900 | Enzyme | |
| Carbonic anhydrase II | CA2 | P00918 | CHEMBL205 | Lyase | ||
| Carbonic anhydrase VII | CA7 | P43166 | CHEMBL2326 | Lyase | ||
| Carbonic anhydrase XII | CA12 | O43570 | CHEMBL3242 | Lyase | ||
| Carbonic anhydrase IV | CA4 | P22748 | CHEMBL3729 | Lyase | ||
| NADPH oxidase 4 | NOX4 | Q9NPH5 | CHEMBL1250375 | Enzyme | ||
| Adrenergic receptor alpha-2 | ADRA2C | P18825 | CHEMBL1916 | Family A G protein-coupled-receptor | ||
| Acetylcholinesterase | ACHE | P22303 | CHEMBL220 | Hydrolase | ||
| Quinone reductase 2 | NQO2 | P16083 | CHEMBL3959 | Enzyme | ||
| Ribosomal protein S6 kinase alpha 3 | RPS6KA3 | P51812 | CHEMBL2345 | Kinase |  |
|
| Neuromedin-U receptor 2 | NMUR2 | Q9GZQ4 | CHEMBL1075144 | Family A G protein-coupled receptor |  |
|
| Withaferin | Protein kinase C alpha | PRKCA | P17252 | CHEMBL299 | Kinase |  |
| Cyclooxygenase-2 | PTGS2 | P35354 | CHEMBL230 | Oxidoreductase |  |
|
| Isoleucyl-tRNA synthetase | IARS | P41252 | CHEMBL3235 | Enzyme |  |
|
| Protein kinase C delta | PRKCD | Q05655 | CHEMBL2996 | Kinase |  |
|
| HMG-CoA reductase | HMGCR | P04035 | CHEMBL402 | Oxidoreductase |  |
|
| Phosphodiesterase 4D | PDE4D | Q08499 | CHEMBL288 | Phosphodiesterase |  |
|
| Telomerase reverse transcriptase | TERT | O14746 | CHEMBL2916 | Enzyme |  |
|
| Androgen Receptor | AR | P10275 | CHEMBL1871 | Nuclear receptor |  |
|
| Protein kinase C epsilon | PRKCE | Q02156 | CHEMBL3582 | Kinase |  |
|
| Proto-oncogene c-JUN | JUN | P05412 | CHEMBL4977 | Transcription factor |  |
|
| Protein-tyrosine phosphatase 1B | PTPN1 | P18031 | CHEMBL335 | Phosphatase |  |
Table 6.
Predicted bioactive molecules from DrugBank.
| Metabolites | Drug bank id | Name | Score | Status |
|---|---|---|---|---|
| Asiatic acid | DB00741 | Hydrocortisone | 0.539 | Approved |
| DB01160 | Dinoprost Tromethamine | 0.529 | Approved | |
| DB07886 | (11alpha,14beta)-11,17,21-trihydroxypregn-4-ene-3,20-dione | 0.539 | Experimental | |
| DB07209 | (8R,9Z,12Z)-8-hydroxy-6-oxooctadeca-9,12-dienoic acid | 0.510 | Experimental | |
| Guajaverin | DB08995 | Diosmin | 0.280 | Approved |
| DB02375 | Myricetin | 0.236 | Experimental | |
| Withaferin | DB00410 | Mupirocin | 0.481 | Approved |
| DB00641 | Simvastatin | 0.447 | Approved | |
| DB08224 | hexahydro-7-methyl-8-[2-[(2r,4r)-tetrahydro-4-hydroxy-6-oxo-2h-pyran-2-yl]ethyl]-1-naphthalenol | 0.501 | Experimental | |
| DB04775 | Reidispongiolide C | 0.479 | Experimental | |
| Avicularin | DB08995 | Diosmin | 0.249 | Approved |
| DB02375 | Myricetin | 0.210 | Experimental |
4. Discussion
Excessive infection rates and mortality of SARS-CoV-2 led the researchers to concentrate immensely on developing strategies for combating infections caused by the pathogen [[69], [70], [71]]. Regardless of this praiseworthy initiative, there are still no specific drugs or approved vaccines that could treat SARS-CoV-2 infected patients [72,73]. Though some candidates are in the investigational stages, many of them raise controversial issues [74,75]. Plant-derived natural products play a significant role by being a lead molecule in the development of drug candidates [76]. Hence, in the present study, attempts were taken to evaluate some plant-derived metabolites as inhibitory agents of SARS-CoV-2 based on their binding affinities to the key proteins of the pathogen.
The contribution of computational biology has accelerated the pace of drug discovery [77]. It is now used in the biopharmaceutical industry to discover and develop new lead compounds against many infectious pathogens [77,78]. By this route, one can visualize the possibilities of binding of potential small molecules as ligands/inhibitors [76]. Phytomolecules like Baicalein, Luteolin, Quercetin, and Kaempferol are potential antiviral agents against a wide range of important viruses including Dengue, HIV, H5N1 influenza A virus, Coxsackie virus, CHIKV, and Japanese encephalitis virus [79]. Recent studies have focused on MPP inhibitors of SARS-CoV-2 i.e. alpha-ketoamide, Hydroxy, Remdesivir, Chloroquine and Favipiravir to evaluate their potency as drugs [80,81]. Several in silico strategies were also adopted to screen putative drug candidates against SARS-CoV-2 [82,83]. However, all these experiments used either main protease proteins or RNA-dependent RNA polymerase of SARS-CoV-2 as probable drug targets. In this study, we screened some natural metabolites against SARS-CoV-2 main proteases (6W63, 6LU7), Nsp9 (Non-structural protein-9) RNA binding protein (6W4B), spike receptor binding domain (6M0J), spike ecto-domain (6VYB), and HR2 domain (6LVN) using a molecular docking approach [[84], [85], [86]]. The polyproteins of coronavirus are cleaved and transformed in mature non-structural proteins (Nsp) by proteases [87]. As a putative component in the replication complex, Nsp9 may possibly have an RNA binding activity. Viral replication complexes are frequently membrane associated and Nsp9 helps in this case. The entry of coronavirus into host cells, on the contrary, is mediated by the transmembrane spike glycoprotein that forms homotrimers protruding from the viral surface. S protein comprises two functional subunits responsible for binding to the host cell receptor (S1) and fusion of the viral and cellular membranes (S). After the attachment of the receptor-binding subunit to the receptor, the HR1 and HR2 domains in the membrane fusion subunit interact with each other and form a six-helix bundle, and this conformational change results in a close apposition of the fusion peptide, leading to virus-cell membrane fusion [88]. Thus, all of these proteins represent an attractive pharmacological target for SARS-CoV-2.
Results revealed that asiatic acid had highest binding affinity with SARS-CoV-2 main protease (−53.05 kcal/mol), Nsp9 RNA binding protein (−50.04 kcal/mol) and spike ecto-domain (60.68 kcal/mol) (Fig. 2 and Table 2). Remarkably, four metabolites i.e. asiatic acid, avicularin, guajaverin and withaferin scored best for each six macromolecules and bound with minimum global binding energy (Table 2 and Supplementary File 1). The scores of top candidates were either close or in some instances lower than alpha ketoamide, a positive control used in the present study (Table 2). Asiatic acid, a triterpenoid derivative from Centella asiatica, displayed antioxidative, anti-inflammatory, and protective properties against neurotoxicity induced by glutamate- or b-amyloid previously [89]. Bian et al. also reported the inhibitory activities of asiatic acid and effectivity against fibroproliferative disorders (Keloids) through blocking the TGF-β/Smad pathway [90]. Withanolides are nature-derived secondary metabolites produced in Withania somnifera via oxidation of steroids, which have medicinal value like anti-inflammation, anti-cancer, adaptogenic and anti-oxidant effects [91]. Withaferin, a steroidal lactone from this group, suppresses HIV-1 LTR transcription and viral replication and also has a vital function to inhibit herpes simplex virus [92,93]. It has anti-inflammatory properties and also shows neuro-protective activity against Aβ neurotoxicity [94,95]. A molecular docking and simulation study revealed the vital function of withaferin to attenuate the neuraminidase of H1N1 influenza virus [96].
Guajaverin (Quercetin 3-arabinopyranoside) and avicularin (quercetin- 3-O-α-L-arabinofuranoside) are the main bioactive components of guava leaves with hypoglycemic properties and inhibitory capacity against free fatty acid release [97]. Previously, anti-plaque activity of guajaverine was attributed to its microbicidial activity against the growth of Strep [38]. Avicularin, a flavonoid of plants, displayed diverse pharmacological properties such as anti-inflammatory effects and anti-infectious effects against pathogens [98,99]. Lee et al. reported the effective anti-oxidant potentiality of Avicularin from Lespedeza cuneata [100]. Researchers also identified hepatoprotective activity of avicularin extracted from the aerial parts of Lespedeza cuneata against lesions caused by t‐BHP in HepG2 cells [101]. It has also been suggested to inhibit activation of ERK signaling pathways through LPS-stimulated overproduction of pro-inflammatory mediators and cytokine [98]. Avicularin may suppress the inflammatory response, and causes apoptosis in human RA synovial cells through obstructing the activation of the MEK/NF-κB pathway, thus preventing rheumatoid arthritis (RA) in vitro [102].
In the present study, we revealed the molecular interactions of top drug candidates with SARS-CoV-2 key proteins (Fig. 2 and 3 and Table 2). The binding sites for each ligand occupied the catalytic domain of SARS-CoV-2 main protease protein [103]. Among the common binding residues, His41 and Cys145 form the catalytic dyad and act as a substrate recognition site [103,104]. The top candidates were well fitted into the active pocket of MPP where several hydrophobic amino acid residues including Met49, Gly143, Cys145, Met165, Pro168, Ala191 compose a relatively hydrophobic environment, which may help to stabilize its conformation [104]. The crucial binding sites of Nsp9 protein (39–73 region) are characterized by positively charged, glycine rich β-loops, which were proposed to be involved in RNA binding [105]. Moreover, we targeted three distinct domains of SARS-CoV-2 spike protein, all of which play essential roles in the mechanism of viral entry into the host cell [106]. The investigation may be useful to unravel the main drug target hotspot and medicinal chemistry of the investigational drugs currently under trials against SARS-CoV-2. ADME data, whether experimentally measured or computationally predicted, provide key insights into how a drug will ultimately be treated or accepted by the body. Hence, while a drug lead may exhibit phenomenal efficacy in vitro, poor ADME results often invariably terminate its development [107]. Computational methods play a key role in anticipating potential ADME and toxicity problems and reducing the number of experiments that involve animal testing. Therefore, the topmost drug candidates were employed for ADME analysis to investigate their drug profiles. None of the metabolites, however, showed any undesirable consequences that could reduce their drug likeness properties. SARS-CoV-2 appears as a severe acute respiratory disease not a neuro disease [108]. Thus, there is no need to permeate the blood brain barrier (BBB) for being an effective molecule against SARS-CoV-2. However, no BBB permeants were found among the top drug candidates. Most of the target class for the top drug candidates belonged to the categories of enzymes (e.g. oxidoreductases, hydrolase, phosphatases, lyases (Table 5). The major protease proteins (protein hydrolase) of SARS-CoV-2 thus can be a specific target for these natural metabolites. Guajaverin and avicularin are two isomers and derivatives of quercetin with a glycoside substituent in their chemical structure [109,110]. The significant similarity between these two polar compounds in terms of structure, chemical formula, molar mass, and other physicochemical parameters (Table 3) may be responsible for covering the same targets by these two flavonoids (Table 5). The toxicity of drug impurities is closely related to their structure. Structure-activity relationships (SARs) have been widely used in Europe and the United States to predict toxicity by computer [111]. The toxicity prediction results in the present study revealed negligible tumorigenic, mutagenic, irritating, or reproductive effects by the drug candidates, though withaferin was found to be comparatively toxic among the top four compounds. Despite having medicinal importance [112,113], the ability of withaferin to inhibit cell growth and induce apoptosis in in vitro and in vivo models were reported [114,115]. Moreover, dose dependent toxicity and other adverse effects such as elevation of liver enzymes, skin rash, fever etc. were observed by researchers [115,116]. Our study also revealed the hepatotoxic and cytotoxic nature of withaferin through computational investigations.
However, drug similarity prediction identified two approved structural analogs of withaferin, Mupirocin (DB00410) and Simvastatin (DB00641) which could be alternative choices, and therefore require further in vivo investigations. Ligand-based virtual screening using asiatic acid predicted two other biologically active compounds, Hydrocortisone (DB00741) and Dinoprost-tromethamine (DB01160) from DrugBank. Interestingly, Hydrocortisone, a cortisone based drug, was previously used during the SARS-CoV-1 and MERS outbreak [117]. Diosmin, on the contrary, is used as a supplementary drug and is found in various natural plants [118]. Myricetin showed the potential to inhibit reverse transcriptase of the RLV and HIV viruses, while characterized as having antioxidative and prooxidative properties. It is also a potent anticarcinogen and antimutagen [119]. The most significant finding of this study is Simvastatin, which can block downstream molecules (key factors in virus infectivity) and can control severe influenza and pneumonia through prevention of excess cytokine release [120]. The results suggest that all these compounds could be potential drug candidates against SARS-CoV-2. The study may pave the way to develop effective medications and preventive measures against SARS-CoV-2 in the future.
5. Conclusion
The results suggest that asiatic acid, avicularin, and guajaverin could be options to treat SARS-CoV-2 associated infections. Furthermore, two biologically active structural analogs from DrugBank i.e. Hydrocortisone and Simvastain may be effective and show potency against the viral pathogen. However, all the investigational drugs of SARS-CoV-2 are under strict regulation by the World Health Organization. Due to the encouraging results, we highly recommend further in vivo trials for the experimental validation of our findings.
Declaration of competing interest
The authors declare that they have no conflict of interests.
Acknowledgements
The authors would like to acknowledge the Department of Microbial Biotechnology, Department of Pharmaceuticals and Industrial Biotechnology, and the Department of Plant and Environmental Biotechnology of Sylhet Agricultural University, Sylhet-3100, Bangladesh for the technical support provided for this research work.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.imu.2020.100367.
Funding information
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Appendix A. Supplementary data
The following is the Supplementary data to this article:
References
- 1.Yao T.T., Qian J.D., Zhu W.Y., Wang Y., Wang G.Q.A. Systematic review of lopinavir therapy for SARS coronavirus and MERS coronavirus-A possible reference for coronavirus disease-19 treatment option. J Med Virol. 2020 doi: 10.1002/jmv.25729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.World Health Organization Coronavirus disease (COVID-19) pandemic 2020. https://www.who.int/emergencies/diseases/novel-coronavirus-2019
- 3.Zhou M., Zhang X., Qu J. Coronavirus disease 2019 (COVID-19): a clinical update. Front Med. 2020;2:1–10. doi: 10.1007/s11684-020-0767-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Guo Y.R., Cao Q.D., Hong Z.S., Tan Y.Y., Chen S.D., Jin H.J. The origin, transmission and clinical therapies on coronavirus disease 2019 (COVID-19) outbreak - an update on the status. Mil Med Res. 2020;7(1):1–10. doi: 10.1186/s40779-020-00240-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.De Wit E., Van Doremalen N., Falzarano D., Munster V.J. SARS and MERS: recent insights into emerging coronaviruses. Nat Rev Microbiol. 2016;14(8):523–534. doi: 10.1038/nrmicro.2016.81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Chan J.F., Kok K.H., Zhu Z., Chu H., To K.K., Yuan S. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg Microb Infect. 2020;9(1):221–236. doi: 10.1080/22221751.2020.1719902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kannan S., Shaik Syed Ali P., Sheeza A., Hemalatha K. COVID-19 (Novel Coronavirus 2019) - recent trends. Eur Rev Med Pharmacol Sci. 2020;24(4):2006–2011. doi: 10.26355/eurrev_202002_20378. [DOI] [PubMed] [Google Scholar]
- 8.Mousavizadeh L., Ghasemi S. Genotype and phenotype of COVID-19: their roles in pathogenesis. J Microbiol Immunol Infect. 2020;S1684–1182(20):30082–30087. doi: 10.1016/j.jmii.2020.03.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zhou P., Yang X.L., Wang X.G., Hu B., Zhang L., Zhang W. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579(7798):270–273. doi: 10.1038/s41586-020-2012-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Li F. Receptor recognition mechanisms of coronaviruses: a decade of structural studies. J Virol. 2015;89:1954–1964. doi: 10.1128/JVI.02615-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Structure Li F. Function, and evolution of coronavirus spike proteins. Annu Rev Virol. 2016;3(1):237–261. doi: 10.1146/annurev-virology-110615-042301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wan Y., Shang J., Graham R., Baric R.S., Li F. Receptor recognition by novel coronavirus from Wuhan: an analysis based on decade-long structural studies of SARS. J Virol. 2020;94(7) doi: 10.1128/JVI.00127-20. e127-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Velavan T.P., Meyer C.G. The COVID-19 epidemic. Trop Med Int Health. 2020;25(3):278–280. doi: 10.1111/tmi.13383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Munster V.J., Koopmans M., van Doremalen N., van Riel D., de Wit E. A novel coronavirus emerging in China - key questions for impact assessment. N Engl J Med. 2020;382(8):692–694. doi: 10.1056/NEJMp2000929. [DOI] [PubMed] [Google Scholar]
- 15.Zheng J. SARS-CoV-2: an emerging coronavirus that causes a global threat. Int J Biol Sci. 2020;16(10):1678–1685. doi: 10.7150/ijbs.45053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zou L., Ruan F., Huang M., Liang L., Huang H., Hong Z. SARS-CoV-2 viral load in upper respiratory specimens of infected patients. N Engl J Med. 2020;10:1056. doi: 10.1056/NEJMc2001737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lu H. Drug treatment options for the 2019-new coronavirus (2019-nCoV) Biosci Trends. 2020;14(1) doi: 10.5582/bst.2020.01020. 69-7. [DOI] [PubMed] [Google Scholar]
- 18.Li G., De Clercq E. Therapeutic options for the 2019 novel coronavirus (2019-nCoV) Nat Rev Drug Discov. 2020;19(3):149–150. doi: 10.1038/d41573-020-00016-0. [DOI] [PubMed] [Google Scholar]
- 19.Dhama K., Sharun K., Tiwari R., Dadar M., Malik Y.S., Singh K.P. COVID-19, an emerging coronavirus infection: advances and prospects in designing and developing vaccines, immunotherapeutics, and therapeutics. Hum Vaccines Immunother. 2020;18:1–7. doi: 10.1080/21645515.2020.1735227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bregu M., Draper S.J., Hill A.V., Greenwood B.M. Accelerating vaccine development and deployment: report of a Royal Society satellite meeting. Philos Trans R Soc Lond B Biol Sci. 2011;366(1579):2841–2849. doi: 10.1098/rstb.2011.0100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Koc Suheda, Belgin S., Isgor, Yasemin G., Isgor, Moghaddam N.S. The potential medicinal value of plants from Asteraceae family with antioxidant defense enzymes as biological targets. Pharmaceut Biol. 2015;53(5):746–751. doi: 10.3109/13880209.2014.942788. [DOI] [PubMed] [Google Scholar]
- 22.Kumar V., Suri S., Prasad R., Gat Y., Sangma C., Jakhu H., Sharma M. Bioactive compounds, health benefits and utilization of Rhododendron: a comprehensive review. Agric Food Secur. 2019;8:6. [Google Scholar]
- 23.Cheuka P.M., Mayoka G., Mutai P., Chibale K. The role of natural products in drug discovery and development against neglected tropical diseases. Molecules. 2016;22(1):58. doi: 10.3390/molecules22010058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Karimi A., Majlesi M., Rafieian-Kopaei M. Herbal versus synthetic drugs; beliefs and facts. J Nephropharmacol. 2015;4(1):27–30. [PMC free article] [PubMed] [Google Scholar]
- 25.El-Saber Batiha G., Magdy Beshbishy A., Wasef L., Elewa Y.H., A Al-Sagan A., El-Hack A. Chemical constituents and pharmacological activities of garlic (allium sativum L.): a review. Nutrients. 2020;12(3):872. doi: 10.3390/nu12030872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Mussard E., Cesaro A., Lespessailles E., Legrain B., Berteina-Raboin S., Toumi H. Andrographolide, a natural antioxidant: an update. Antioxidants. 2019;8(12):571. doi: 10.3390/antiox8120571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Salehi B., Venditti A., Sharifi-Rad M., Kręgiel D., Sharifi-Rad J., Durazzo A. The therapeutic potential of apigenin. Int J Mol Sci. 2019;20(6):1305. doi: 10.3390/ijms20061305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Nagoor Meeran M.F., Goyal S.N., Suchal K., Sharma C., Patil C.R., Ojha S.K. Pharmacological properties, molecular mechanisms, and pharmaceutical development of asiatic acid: a pentacyclic triterpenoid of therapeutic promise. Front Pharmacol. 2018;9:892. doi: 10.3389/fphar.2018.00892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Wang Z., Li F., Quan Y., Shen J. Avicularin ameliorates human hepatocellular carcinoma via the regulation of NF-κB/COX-2/PPAR-γ activities. Mol Med Rep. 2019;19(6):5417–5423. doi: 10.3892/mmr.2019.10198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Hayman M., Kam P.C.A. Capsaicin: a review of its pharmacology and clinical applications. Curr Anaesth Crit Care. 2019;19:338–343. [Google Scholar]
- 31.Bhalerao S.A., Verma D.R., Gavankar R.V., Teli N.C., Rane Y.Y., Didwana V.S. Phytochemistry, pharmacological profile and therapeutic uses of piper betle linn.– an overview. J Pharmacogn Phytochem. 2013;1(2):10–19. [Google Scholar]
- 32.Guzman J.D. Natural cinnamic acids, synthetic derivatives and hybrids with antimicrobial activity. Molecules. 2014;19(12) doi: 10.3390/molecules191219292. 19292‐19349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Al-Samydai A., Jaber N. Pharmacological aspects of curcumin: review article. Int J Pharm. 2018;5(6):313–326. [Google Scholar]
- 34.Nejad S.M., Özgüneş H., Başaran N. Pharmacological and toxicological properties of eugenol. The Turkish J Pharm Sci. 2017;14(2):201–206. doi: 10.4274/tjps.62207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Amalraj A., Gopi S. Medicinal properties of Terminalia arjuna (roxb.) wight & arn.: a review. J Tradit Compl Med. 2016;7(1):65‐78. doi: 10.1016/j.jtcme.2016.02.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Patel D.K., Patel K., Gadewar M., Tahilyani V. Pharmacological and bioanalytical aspects of galangin-a concise report. Asian Pacific J Trop Biomed. 2012;2(1):S449–S455. doi: 10.1016/S2221-1691(12)60239-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Abedi F., Razavi B.M., Hosseinzadeh H.A. Review on gentisic acid as a plant derived phenolic acid and metabolite of aspirin: comprehensive pharmacology, toxicology, and some pharmaceutical aspects. Phytother Res. 2019;34(4):729–741. doi: 10.1002/ptr.6573. [DOI] [PubMed] [Google Scholar]
- 38.Prabu G.R., Gnanamani A., Sadulla S. Guaijaverin–a plant flavonoid as potential antiplaque agent against Streptococcus mutans. J Appl Microbiol. 2006;101(2):487–495. doi: 10.1111/j.1365-2672.2006.02912.x. [DOI] [PubMed] [Google Scholar]
- 39.Imran M., Salehi B., Sharifi-Rad J., Aslam Gondal T., Saeed F., Kaempferol Imran A. A key emphasis to its anticancer potential. Molecules. 2019;24(12):2277. doi: 10.3390/molecules24122277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Lopez-Lazaro M. Distribution and biological activities of the flavonoid Luteolin. Med Chem. 2009;9(1):31–59. doi: 10.2174/138955709787001712. [DOI] [PubMed] [Google Scholar]
- 41.Ohnishi R., Ito H., Iguchi A., Shinomiya K., Kamei C., Hatano T. Effects of chlorogenic acid and its metabolites on spontaneous locomotor activity in mice. Biosci Biotechnol Biochem. 2006;70(10):2560–2563. doi: 10.1271/bbb.60243. [DOI] [PubMed] [Google Scholar]
- 42.Maalik A., Khan F.A., Mumtaz A., Mehmood A., Azhar S., Atif M. Pharmacological applications of quercetin and its derivatives: a short review. Trop J Pharmaceut Res. 2014;13(9):1561–1566. [Google Scholar]
- 43.Mgbeahuruike E.E., Yrjönen T., Vuorela H., Holm Y. Bioactive compounds from medicinal plants: focus on Piper species. South Afr J Bot. 2017;112:54–69. [Google Scholar]
- 44.Kumar V., Van Staden J.A. Review of swertia chirayita (gentianaceae) as a traditional medicinal plant. Front Pharmacol. 2016;6:308. doi: 10.3389/fphar.2015.00308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Singh R.L., Singh P., Agarwal A. Chemical constituents and bio-pharmacological activities of Swertia chirata: a review. NPAIJ. 2012;8(6):238–247. [Google Scholar]
- 46.Ahmad A., Mishra R.K., Vyawahare A., Kumar A., Rehman M.U., Qamar W. Thymoquinone (2-Isopropyl-5-methyl-1, 4-benzoquinone) as a chemopreventive/anticancer agent: chemistry and biological effects. Saudi Pharmaceut J. 2019;27(8):1113–1126. doi: 10.1016/j.jsps.2019.09.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Barrales-Cureño H.J. Pharmacological applications and in vitro biotechnological production of anticancer alkaloids of Catharanthus roseus. Biotecnol Apl. 2015;32(1):1101–1110. [Google Scholar]
- 48.He M., Min J.W., Kong W.L., He X.H., Li J.X., Peng B.W. A review on the pharmacological effects of vitexin and isovitexin. Fitoterapia. 2016;115:74–85. doi: 10.1016/j.fitote.2016.09.011. [DOI] [PubMed] [Google Scholar]
- 49.Patel K., Singh R.B., Patel D.K. Pharmacological and analytical aspects of withaferin A: a concise report of current scientific literature. Asian Pac J Reprod. 2013;2(3):238–243. [Google Scholar]
- 50.Johji Y., Michihiko M., Rong H.Q., Hisashi M., Hajime F. The anti-ulcer effect in rats of ginger constituents. J Ethnopharmacol. 1988;23(2–3):299–304. doi: 10.1016/0378-8741(88)90009-8. [DOI] [PubMed] [Google Scholar]
- 51.Rose P.W., Prlić A., Altunkaya A., Bi C., Bradley A.R., Christie C.H. The RCSB protein data bank: integrative view of protein, gene and 3D structural information. Nucleic Acids Res. 2017;45:D271–D281. doi: 10.1093/nar/gkw1000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Kim S., Thiessen P.A., Bolton E.E., Chen J., Fu G., Gindulyte A. PubChem substance and compound databases. Nucleic Acids Res. 2016;44(D1):D1202–D1213. doi: 10.1093/nar/gkv951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.O'Boyle N.M., Banck M., James C.A., Morley C., Vandermeersch T., Hutchison G.R. Open Babel: an open chemical toolbox. J Cheminf. 2011;3(1):33. doi: 10.1186/1758-2946-3-33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Meng X.Y., Zhang H.X., Mezei M., Cui M. Molecular docking: a powerful approach for structure-based drug discovery. Curr Comput Aided Drug Des. 2011;7(2):146–157. doi: 10.2174/157340911795677602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Kitchen D.B., Decornez H., Furr J.R., Bajorath J. Docking and scoring in virtual screening for drug discovery: methods and applications. Nat Rev Drug Discov. 2004;3(11):935–949. doi: 10.1038/nrd1549. [DOI] [PubMed] [Google Scholar]
- 56.Schneidman-Duhovny D., Inbar Y., Nussinov R., Wolfson H.J. PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res. 2005;33(suppl_2):W363–W367. doi: 10.1093/nar/gki481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Zhang L., Lin D., Sun X., Curth U., Drosten C., Sauerhering L. Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors. Science. 2020;368(6489):409–412. doi: 10.1126/science.abb3405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Mashiach E., Schneidman-Duhovny D., Andrusier N., Nussinov R., Wolfson H.J. FireDock: a web server for fast interaction refinement in molecular docking. Nucleic Acids Res. 2008;36(suppl_2):W229–W232. doi: 10.1093/nar/gkn186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Wang Q., He J., Wu D., Wang J., Yan J., Li H. Interaction of α-cyperone with human serum albumin: determination of the binding site by using Discovery Studio and via spectroscopic methods. J Lumin. 2015;164:81–85. [Google Scholar]
- 60.DeLano W.L. Pymol: an open-source molecular graphics tool. CCP4 Newslett Protein Crystallogr. 2002;40(1):82–92. [Google Scholar]
- 61.Balani S.K., Miwa G.T., Gan L.S., Wu J.T., Lee F.W. Strategy of utilizing in vitro and in vivo ADME tools for lead optimization and drug candidate selection. Curr Top Med Chem. 2005;5(11):1033–1038. doi: 10.2174/156802605774297038. [DOI] [PubMed] [Google Scholar]
- 62.Daina A., Michielin O., Zoete V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep. 2017;7:42717. doi: 10.1038/srep42717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Daina A., Zoete V. A boiled‐egg to predict gastrointestinal absorption and brain penetration of small molecules. ChemMedChem. 2016;11(11):1117–1121. doi: 10.1002/cmdc.201600182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Banerjee P., Eckert A.O., Schrey A.K., Preissner R. ProTox-II: a webserver for the prediction of toxicity of chemicals. Nucleic Acids Res. 2018;46(W1):W257–W263. doi: 10.1093/nar/gky318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Drwal M.N., Banerjee P., Dunkel M., Wettig M.R., Preissner R. ProTox: a web server for the in silico prediction of rodent oral toxicity. Nucleic Acids Res. 2014;42(W1):W53–W58. doi: 10.1093/nar/gku401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Sander T. OSIRIS property explorer. Org Chem Portal. 2001 https://www.organicchemistry.org/prog/peo/ [Google Scholar]
- 67.Daina A., Michielin O., Zoete V. Swiss Target Prediction: updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res. 2019;47(W1):W357–W364. doi: 10.1093/nar/gkz382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Zoete V., Daina A., Bovigny C., Michielin O. Swiss similarity: a web tool for low to ultra high throughput ligand-based virtual screening. J Chem Inf Model. 2016;56(8):1399–1404. doi: 10.1021/acs.jcim.6b00174. [DOI] [PubMed] [Google Scholar]
- 69.Wilder-Smith A., Chiew C.J., Lee V.J. Can we contain the COVID-19 outbreak with the same measures as for SARS? Lancet Infect Dis. 2020;20:e102–e107. doi: 10.1016/S1473-3099(20)30129-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Yuen K.S., Ye Z.W., Fung S.Y., Chan C.P., Jin D.Y. SARS-CoV-2 and COVID-19: the most important research questions. Cell Biosci. 2020;10(1):1–5. doi: 10.1186/s13578-020-00404-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Lake M.A. What we know so far: COVID-19 current clinical knowledge and research. Clin Med. 2020;20(2):124. doi: 10.7861/clinmed.2019-coron. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Fang Y., Zhang H., Xie J., Lin M., Ying L., Pang P., Ji W. Sensitivity of chest CT for COVID-19: comparison to RT-PCR. Radiology. 2020;19:200432. doi: 10.1148/radiol.2020200432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Prompetchara E., Ketloy C., Palaga T. Immune responses in COVID-19 and potential vaccines: lessons learned from SARS and MERS epidemic. Asian Pac J Allergy Immunol. 2020;38(1):1–9. doi: 10.12932/AP-200220-0772. [DOI] [PubMed] [Google Scholar]
- 74.Zhou D., Dai S.M., Tong Q. COVID-19: a recommendation to examine the effect of hydroxychloroquine in preventing infection and progression. J Antimicrob Chemother. 2020 doi: 10.1093/jac/dkaa114. pii:dkaa114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Dong L., Hu S., Gao J. Discovering drugs to treat coronavirus disease 2019 (COVID-19) Drug Discov Therapeut. 2020;14(1):58–60. doi: 10.5582/ddt.2020.01012. [DOI] [PubMed] [Google Scholar]
- 76.Joseph J., Bhaskaran R., Kaliraj M., Muthuswamy M., Suresh A., Monodon P. Molecular Docking of Phytoligands to the viral protein receptor. Rab7 Bioinformation. 2017;13(4):116. doi: 10.6026/97320630013116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Hirono S. An introduction to the computer-aided structure-based drug design--applications of bioinformatics to drug discovery. Rinsho byori. The Japanese journal of clinical pathology. 2002;50(1):45–51. [PubMed] [Google Scholar]
- 78.Ivanov A.S., Veselovsky A.V., Dubanov A.V., Skvortsov V.S. Bioinformatics platform development: from gene to lead compound. Methods Mol Biol. 2006;316:389–431. doi: 10.1385/1-59259-964-8:389. [DOI] [PubMed] [Google Scholar]
- 79.Habbu PV, Mahadevan KM, Shastry RA, Manjunatha H. Antimicrobial activity of flavanoid sulphates and other fractions of Argyreiaspeciosa (Burm. [PubMed]; f) Boj Indian J Exp Biol. 2009;47(2):121–128. [PubMed] [Google Scholar]
- 80.Al-Tawfiq J.A., Al-Homoud A.H., Memish Z.A. Travel Med Infect Dis; 2020. Remdesivir as a possible therapeutic option for the COVID-19; p. 101615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Colson P., Rolain J.M., Lagier J.C., Brouqui P., Raoult D. Chloroquine and hydroxychloroquine as available weapons to fight COVID-19. Int J Antimicrob Agents. 2020:105932. doi: 10.1016/j.ijantimicag.2020.105932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Parvez M., Alam S., Karim M., Hasan M., Jaman J., Karim Z. 2020. Prediction of potential inhibitors for RNA-dependent RNA polymerase of SARS-CoV-2 using comprehensive drug repurposing and molecular docking approach. arXiv:2004.07086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Hasan M., Parvez M.S., Azim K.F., Imran A.S., Raihan T., Gulshan A. Main protease inhibitors and drug surface hotspot for the treatment of COVID-19: drug repurposing and molecular docking approach. ChemRxiv (Preprint) 2020 doi: 10.1016/j.biopha.2021.111742. https://doi: 10.26434/chemrxiv.12118857 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Chang M.W., Ayeni C., Breuer S., Torbett B.E. Virtual screening for HIV protease inhibitors: a comparison of AutoDock 4 and Vina. PloS One. 2010;5(8) doi: 10.1371/journal.pone.0011955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Hasan M., Ghosh P.P., Azim K.F., Mukta S., Abir R.A., Nahar J. Reverse vaccinology approach to design a novel multi-epitope subunit vaccine against avian influenza A (H7N9) virus. Microb Pathog. 2019;130:19–37. doi: 10.1016/j.micpath.2019.02.023. [DOI] [PubMed] [Google Scholar]
- 86.Azim K.F., Lasker T., Akter R., Hia M.M., Bhuiyan O.F., Hasan M. 2019. Conglomeration of highly antigenic nucleoproteins to inaugurate a heterosubtypic next generation vaccine candidate against Arenaviridae family. bioRxiv (Preprint) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Hilgenfeld R. From SARS to MERS: crystallographic studies on coronaviral proteases enable antiviral drug design. FEBS J. 2014;281:4085–4096. doi: 10.1111/febs.12936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Moore J.P., Doms R.W. The entry of entry inhibitors: a fusion of science and medicine. Proc Natl Acad Sci USA. 2003;100:10598–10602. doi: 10.1073/pnas.1932511100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Krishnamurthy R.G., Senut M., Zemke D., Min J., Frenkel M.B., Greenberg E.J. Asiatic acid, a pentacyclic triterpene from centellaasiatica, is neuroprotective in a mouse model of focal cerebral ischemia. J Neurosci Res. 2009;87:2541–2550. doi: 10.1002/jnr.22071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Bian D., Zhang J., Wu X., Dou Y., Yang Y., Tan Q. Asiatic acid isolated from Centella asiatica inhibits TGF-β1-induced collagen expression in human keloid fibroblasts via PPAR-γ activation. Int J Biosci. 2013;9(10):1032–1042. doi: 10.7150/ijbs.7273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Vaishnavi K., Saxena N., Shah N. Differential activities of the two closely related withanolides, Withaferin A and Withanone: bioinformatics and experimental evidences. PLoS One. 2012;7(9) doi: 10.1371/journal.pone.0044419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Shi T., Wilhelm E., Bell B., Dumais N. Nf-κb-Dependent inhibition of HIV-1 transcription by withaferin A. HIV Curr Res. 2017;2:1. [Google Scholar]
- 93.Grover A., Agrawal V., Shandilya A., Bisaria V.S., Sundar D. Non-nucleosidic inhibition of Herpes simplex virus DNA polymerase: mechanistic insights into the anti-herpetic mode of action of herbal drug withaferin A. BMC Bioinf. 2011;12(Suppl 13):S22. doi: 10.1186/1471-2105-12-S13-S22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.White P.T., Subramanian C., Motiwala H.F., Cohen M.S. Natural withanolides in the treatment of chronic diseases. Adv Exp Med Biol. 2016;928:329–373. doi: 10.1007/978-3-319-41334-1_14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Tiwari S., Atluri V.S.R., Yndart Arias A. Withaferin A suppresses beta amyloid in app expressing cells: studies for tat and cocaine associated neurological dysfunctions. Front Aging Neurosci. 2018;10:291. doi: 10.3389/fnagi.2018.00291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Cai Z., Zhang G., Tang B., Liu Y., Fu X., Zhang X. Promising anti-influenza properties of active constituent of withaniasomnifera ayurvedic herb in targeting neuraminidase of H1N1 influenza: computational study. Cell Biochem Biophys. 2015;72:727–739. doi: 10.1007/s12013-015-0524-9. [DOI] [PubMed] [Google Scholar]
- 97.Wen O., Xiaoai Z., Lei S., Xuexiang C., Shumin Y., Yong C. Hypoglycemic activity of avicularin and guaijaverin in guava leaves [J] Food Sci (N Y) 2016;37(7) 168-17. [Google Scholar]
- 98.Vo V.A., Lee J.W., Chang J.E., Kim J.Y., Kim N.H., Lee H.J. Avicularin inhibits lipopolysaccharide-induced inflammatory response by suppressing ERK phosphorylation in RAW macrophages. Biomol Ther. 2012;20(6):532–537. doi: 10.4062/biomolther.2012.20.6.532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Shen Z., Xu Y., Jiang X., Wang Z., Guo Y., Pan W., Hou J. Avicularin relieves depressive-like behaviors induced by chronic unpredictable mild stress in mice. Med Sci Mon Int Med J Exp Clin Res: Int Med J Exp Clin Res. 2019;25:2777–2784. doi: 10.12659/MSM.912401. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 100.Lee J.S., Lee A.Y., Quilantang N.G., Geraldino P.J.L., Cho E.J., Lee S. Anti-oxidant activity of avicularin and isovitexin from Lespedeza cuneata. J Appl Biol Chem. 2019;62(2):143–147. [Google Scholar]
- 101.Kim S.M., Kang K., Jho E.H., Jung Y.J., Nho C.W., Um B.H. Hepatoprotective effect of flavonoid glycosides from Lespedeza cuneata against oxidative stress induced by tert-butyl hyperoxide. Phytother Res. 2011;25(7):1011–1017. doi: 10.1002/ptr.3387. [DOI] [PubMed] [Google Scholar]
- 102.Wang W., Zheng H., Zheng M., Liu X., Yu J. Protective effect of avicularin on rheumatoid arthritis and its associated mechanisms. Exp Ther Med. 2018;16(6):5343–5349. doi: 10.3892/etm.2018.6872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Yang H., Yang M., Ding Y., Liu Y., Lou Z., Zhou Z. The crystal structures of severe acute respiratory syndrome virus main protease and its complex with an inhibitor. Proc Natl Acad Sci Unit States Am. 2003;100(23):13190–13195. doi: 10.1073/pnas.1835675100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Wu C., Liu Y., Yang Y., Zhang P., Zhong W., Wang Y. Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharm Sin B. 2020;10(5):766–788. doi: 10.1016/j.apsb.2020.02.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Littler D., Gully B., Colson R.N., Rossjohn J. 2020. Crystal structure of the SARS-CoV-2 non-structural protein 9, Nsp9. BioRxiv (preprint) 013920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Shang J., Wan Y., Luo C., Ye G., Geng Q., Auerbach A. Cell entry mechanisms of SARS-CoV-2. Proc Natl Acad Sci Unit States Am. 2020;117(21):11727–11734. doi: 10.1073/pnas.2003138117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Wishart D.S. Improving early drug discovery through ADME modelling. Drugs R. 2007;8(6):349–362. doi: 10.2165/00126839-200708060-00003. [DOI] [PubMed] [Google Scholar]
- 108.Astuti Y. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): an overview of viral structure and host response. Diabetes Metab. Syndrome. 2020;14(4):407–412. doi: 10.1016/j.dsx.2020.04.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Díaz-de-Cerio E., Verardo V., Gómez-Caravaca A.M., Fernández-Gutiérrez A., Segura-Carretero A. Determination of polar compounds in guava leaves infusions and ultrasound aqueous extract by HPLC-ESI-MS. J Chem. 2015;2015:1–9. [Google Scholar]
- 110.Ortega J.T., Estrada O., Serrano M.L., Contreras W., Orsini G., Pujola F.H. Glycosylated flavonoids from psidium guineense as major inhibitors of HIV-1 replication in vitro. Nat Prod Commun. 2017;12(7):1049–1052. [Google Scholar]
- 111.Guan L., Yang H., Cai Y., Sun L., Di P., Li W., Liu G. ADMET-score–a comprehensive scoring function for evaluation of chemical drug-likeness. Med Chem Comm. 2019;10(1):148–157. doi: 10.1039/c8md00472b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Bhattacharya S.K., Satyan K.S., Ghosal S. Antioxidant activity of glycowithanolides from Withania somnifera Indian. J Exp Biol. 1997;35:236–239. [PubMed] [Google Scholar]
- 113.Misra L., Mishra P., Pandey A., Sangwan R.S., Sangwan N.S., Tuli R. Withanolides from Withania somnifera roots. Phytochemistry. 2008;69(4):1000–1004. doi: 10.1016/j.phytochem.2007.10.024. [DOI] [PubMed] [Google Scholar]
- 114.Stan S.D., Hahm E.R., Warin R., Singh S.V. Withaferin A causes FOXO3a-and Bim-dependent apoptosis and inhibits growth of human breast cancer cells in vivo. Canc Res. 2008;68(18):7661–7669. doi: 10.1158/0008-5472.CAN-08-1510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Safder T., Person E., Cohen M., Mukerji R., Samadi A., Grogan P. Vitro and in vivo cytotoxic effects of withaferin A on retinal epithelial pigmented cells. Invest Ophthalmol Vis Sci. 2012;53(14):5364. [Google Scholar]
- 116.Pires N., Gota V., Gulia A., Hingorani L., Agarwal M., Puri A. Safety and Pharmacokinetics of Withaferin-A in advanced stage high grade Osteosarcoma: A phase I trial. J Ayurveda Integr Med. 2019;11(1):68–72. doi: 10.1016/j.jaim.2018.12.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Barzilai D., Plavnick J., Hazani A., Einath R., Kleinhaus N., Kanter Y. Use of hydrocortisone in the treatment of acute myocardial infarction: summary of a clinical trial in 446 patients. Chest. 1972;61(5):488–491. doi: 10.1378/chest.61.5.488. [DOI] [PubMed] [Google Scholar]
- 118.Moldovan Z., Bunaciu A.A., Al-Omar M.A., Aboul-Enein H.Y.A. Spectrophotometric method for diosmin determination. Open Chem Biomed Methods J. 2010;3:123–127. [Google Scholar]
- 119.Ong K.C., Khoo H.E. Biological effects of myricetin. Gen Pharmacol Vasc Syst. 1997;29(2):121–126. doi: 10.1016/s0306-3623(96)00421-1. [DOI] [PubMed] [Google Scholar]
- 120.Jung K., Wang Q., Kim Y., Scheuer K., Zhang Z., Shen Q. The effect of simvastatin or interferon-α on infectivity of human norovirus using a gnotobiotic pig model for the study of antivirals. PloS One. 2012;7(7) doi: 10.1371/journal.pone.0041619. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.