Abstract
N6-methyladenosine (m6A), the most abundant modification in eukaryotic cells, regulates RNA transcription, processing, splicing, degradation, and translation. Circular RNA (circRNA) is a class of covalently closed RNA molecules characterized by universality, diversity, stability and conservatism of evolution. Accumulating evidence shows that both m6A modification and circRNAs participate in the pathogenesis of multiple diseases, such as cancers, neurological diseases, autoimmune diseases, and infertility. Recently, m6A modification has been identified for its enrichment and vital biological functions in regulating circRNAs. In this review, we summarize the role of m6A modification in the regulation and function of circRNAs. Moreover, we discuss the potential applications and possible future directions in the field.
Keywords: M6A, CircRNA, M6A modified circRNA, Innate immunity, Tumour
Background
Circular RNA (circRNA) is a class of single-stranded covalently closed RNA molecules that was first discovered in pathogens by Sanger et al. in 1976 [1]. It is now generally accepted that circRNA is generated by a process named back-splicing [2], and increasing studies have demonstrated that circRNA plays important roles in the occurrence, development and prognosis of various diseases, including tumorigenesis [3–5], neurodevelopmental processes [6] autoimmune responses [7], and infertility [8]. However, studies on how circRNA is regulated before exerting specific biological functions are still limited [9].
To date, over 160 types of chemical modifications have been identified in RNA molecules, of which methylation is the most common type [10]. The methods of methylation modifications of RNA include N6-methyladenosine (m6A), 5-methylcytosine (m5C), N1-methyladenosine (m1A), 5-hydroxymethylcytosine (5hmC), N6, 2′-O-dimethyladenosine (m6Am), 7-methylguanine (m7G), etc. [11], of which m6A modification is the most abundant type in eukaryotic cells [12]. Previous studies have shown that m6A modification is a dynamic and reversible process and regulates RNA transcription, processing, splicing, degradation, and translation [13–17]. The occurrence and development of many diseases, such as tumours [18], obesity [19], infertility [20], autoimmune disease [21] and neurological disease [22], are closely related to alteration of m6A modification.
Although research on the regulatory mechanism of m6A modification of mRNA has made great progress [23], for some non-coding RNAs, especially circRNAs, the regulatory network of m6A has not been fully elucidated [24]. In this review, we summarize the role of m6A modification in circRNA regulation and function. Furthermore, we discuss the potential applications and possible future directions in this field.
M6A writers, erasers, and readers
The regulation function of m6A is mainly accomplished by three homologous factors referred to as “writers”, “erasers” and “readers”. M6A “writers” are proteins involved in the formation of the methyltransferase complex, including methyltransferase-like 3 and 14 proteins (METTL3 and METTL14) and their cofactors WT1 associated protein (WTAP), RNA-binding motif protein 15/15B (RBM15/15B), Vir-like m6A methyltransferase associated (VIRMA), and zinc finger CCCH-type containing 13 (ZC3H13); METTL3, as the earliest identified and most well-known component [25], is an S-adenosylmethionine (SAM) binding protein and is highly conserved in various eukaryotic species [26, 27]. Notably, except for the above readers that function in a form of complexes, a homologue of METTL3 (METTL16) has been identified as a novel independent RNA methyltransferase that regulates cellular SAM levels and methylates U6 small nuclear RNA [28].
The dynamic and reversible m6A process (Fig. 1) also relies on some demethylases (erasers). Fat mass and obesity-associated protein (FTO), the first protein identified to catalyse m6A demethylation [29], works together with a homologue of itself (ALKBH5, [30] to maintain the balance of m6A levels in the transcriptome [31]. ALKBH3 is a recently discovered demethylase that prefers to perform its demethylation function on tRNA rather than on mRNA or rRNA [32]. In addition, ALKBH3 is also a generally accepted DNA repair enzyme and has the potential to be a molecular marker for tumours [33]. M6A-modified RNA requires a class of variable RNA-binding proteins (readers) to perform specific biological functions. Proteins of the YT521-B homology (YTH) domain family, including YTHDC1, YTHDC2, YTHDF1, YTHDF2 and YTHDF3 [34], were the first five characterized m6A readers in humans that have a conserved m6A-binding domain. The heterogeneous nuclear ribonucleoprotein (HNRNP) family is another group of RNA-binding proteins (RBPs) that serves as m6A readers. Heterogeneous nuclear ribonucleoprotein A2/B1 (HNRNPA2B1) specifically recognizes m6A-modified RNA and acts as a mediator in m6A-dependent nuclear RNA processing [35]. In contrast, HNRNPC and HNRNPG cannot directly bind to the m6A site, but they can mediate the selective splicing process of transcripts containing m6A modification by identifying and binding to the m6A-dependent structural switches [36]. Translation initiation factor 3 (eIF3) initiates the translation procedure by binding to the m6A site in the 5′-UTR of mRNA, while the family of insulin-like growth factor 2 mRNA-binding proteins (IGF2BPs, including IGF2BP1/2/3) makes the target gene and the corresponding translation more stable [37]. Moreover, proline rich coiled-coil 2 A (Prrc2a) is a novel m6A reader that stabilizes mRNA expression by binding to a consensus GGACU motif in the coding sequence (CDS) in an m6A-dependent manner [38].
Fig. 1.
Dynamic and reversible m6A process. The installation, removal and identification of m6A are conducted by writers, readers, and erasers, respectively. Writers refer to the m6A complex, including METTL3, METTL14, WTAP, RBM15/15B, VIRMA and ZC3H13. Besides, METTL16 is a novel independent RNA methyltransferase. Erasers are proteins that own demethylases activity, including FTO, ALKBH5, ALKBH3. Readers are proteins that recognize the m6A modification and perform multiple functions in RNA metabolism, some of which identified so far are YTH family, HNRNP family, eIF3, IGF2BPs and Prrc2a
The dynamic reversibility of m6A modification is closely associated with the normal physiological activities of the organism. Studies have revealed that m6A-modified mRNA or non-coding RNA (mainly miRNA and lncRNA) plays crucial roles in spermatogenesis [39], T cell homeostasis [40], Drosophila sex determination [41], heat shock responses [42], reprogramming and pluripotency [43], as well as other processes. Considering the significance of m6A modification in the regulation of gene expression and various biological functions, dysregulation of m6A levels contributes to diverse diseases, especially for some cancers. Recent studies have indicated that both aberrant m6A modification and abnormal expression of m6A regulatory proteins can both be detected in acute myeloid leukaemia (AML) [44], hepatocellular carcinoma (HCC) [45], glioblastoma stem cells (GSCs) [46], breast cancer [47], obesity [19], infertility [20], autoimmune disease [21] and neurological disease [22].
Characteristics, regulatory mechanisms and biological functions of circRNA
According to their origin, circRNAs can be classified into four broad categories, exonic circRNAs (ecircRNAs), intronic circRNAs (ciRNAs), exon-intron circRNAs (EIciRNAs) and others, ranging from virus, tRNA, rRNA, snRNA [48]. In general, circRNAs can be detected in most organisms, including archaea [49], plants [50], parasites [51], and most mammals [52]. Previous studies have shown that there are more than 25,000 different RNAs that generate corresponding circRNAs in human fibroblasts [53]. Different circRNAs can also be produced by the same gene through alternative circularization [54], which causes the diversity of circRNAs. Another important characteristic of circRNAs is that they cannot be degraded by exonucleases and are therefore more stable than linear circRNAs [55]. Homology studies between different species have shown that circRNAs are highly conserved in evolution between species. The level of homology of circRNA in mice and humans reaches 20% or more [56], while that in pigs and mice is between 15 and 20% [57]. The last but most practical characteristic of circRNAs is that their expression levels vary according to different tissues and different growth stages, which is an essential characteristic for an ideal disease biomarker. Expression profiles of different tissues in humans and mice show that nerve tissue (especially brain tissue) contains more circRNA than other tissues [58], and the expression level of circRNA is gradually upregulated with the development of the brain.
Based on adequate studies on the characteristics of circRNA, an increasing number of studies have focused on its regulatory function [59, 60] (Table 1). The most classical network in which circRNA exerts a specific function occurs through acting as competing endogenous RNA (ceRNA). CircRNAs with a miRNA response element (MRE) can bind specific miRNAs to negatively regulate their activity, so circRNAs can also be considered “miRNA sponges”. The first circRNA defined as an “miRNA sponge” was ciRS-7, and it was first identified in human and mouse brains by Thomas B et al. in 2013 [72]. In addition, circRNAs can also perform specific physiological functions by interacting with some RBPs. In most cases, these circRNAs act as a “separant” to inhibit the function or transport of RBPs. CircEIF3J and circPIAP2, which are predominantly detected in the nucleus, can interact with U1 snRNP and promote transcription of their parental genes [73]. Interestingly, some circRNAs located in the cytoplasm have similar protein binding abilities. CircFoxo3 interacts with inhibitor of DNA binding 1 (ID-1), E2F transcription factor 1 (E2F1), focal adhesion kinase (FAK), and hypoxia inducible factor 1 subunit α (HIF1-α) so that these components are retained together in the cytoplasm [74]. Moreover, recent studies have shown that some circRNAs could be translated into proteins [75, 76]. In the absence of a dissociative 5′ end, the translation of circRNAs cannot be initiated by traditional cap-dependent regulatory elements and therefore requires an internal ribosome entry site (IRES) or other elements to activate a cap-independent pathway. To support this claim, Wang et al. engineered an IRES in a circRNA and then corresponding protein translated by this circRNA was detected in 293 T cells [77]. Recently, another study found that m6A modification was abundant in many circRNAs, and this kind of methylation modification could drive circRNA translation in a manner similar to IRES [78].
Table 1.
Roles of circRNA in different cancers
| Functions | CircRNA | Cancer | Dysregulation | References |
|---|---|---|---|---|
| MiRNA sponge | circ_0026134 | Lung cancer | Up | [61] |
| circ_0005963 | Colorectal cancer | Up | [62] | |
| circ_000684 | Gastric cancer | Up | [63] | |
| circ_0051443 | Hepatocellular cancer | Down | [64] | |
| Binding to protein | circ-Amotl1 | Breast cancer | Up | [65] |
| circ-Foxo3 | Breast cancer | Down | [66] | |
| circ-ZKSCAN1 | Hepatocellular cancer | Down | [67] | |
| Translation template | circ-FBXW7 | Glioblastoma | Down | [68] |
| circ-SHPRH | Glioblastoma | Down | [69] | |
| circ-PPP1R12A | Colon cancer | Up | [70] | |
| circ-β-catenin | Liver cancer | Up | [71] |
Although still in its infancy, circRNAs have been found to be closely related to the occurrence, development and prognosis of various diseases (Fig. 2). Recent studies have demonstrated that the dysregulation of circRNAs exists in different cancers, neuropsychological diseases, autoimmune diseases, infertility, diabetes, nephropathy, arthritis, etc., but few of these circRNAs have been verified to have biological functions. Some studies considered that it might be related to the epigenetic modification of circRNA [79, 80], and m6A modification is the first role that comes into sight.
Fig. 2.
Role of circRNA and m6A modification in various diseases. Three major biological functions of circRNAs are shown on the left. Three homologous factors involved in the regulatory function of m6A are listed on the right
Role of m6A methylation in the regulation of circRNAs
Current studies have identified that dysregulation of m6A modification contributes to various diseases, especially for some cancers. Generally, m6A functions as a double-edged sword. In most cases, aberrant m6A modification contributes to tumorigenesis and tumour progression. However, recent studies revealed that abnormal m6A level can also cause tumour suppression [81]. Since m6A functions via affecting RNA metabolism primarily, researchers have focused their attention on m6A-modified mRNA in recent years. Currently, m6A-modified ncRNAs, especially m6A-modified circRNAs, remain to be further explored. Here, we summarize the role of m6A modification in circRNA regulation and function.
M6A modification regulates circRNA translation
Recent studies have shown that some circRNAs have protein-coding potential [75, 82], and the translation process can be driven by m6A [78]. In general, the translation of RNA in eukaryotic cells requires a eukaryotic translation initiation factor 4F (eIF4F) complex, which is composed of three initiation factors, eIF4A (a helicase protein), eIF4E (a m7G reader) and eIF4G (a scaffold protein) [83]. On mRNA, these transcription initiation elements are located on the cap structure of the 5′ end, so here we define it as a cap-dependent pathway [84]. However, this traditional cap-dependent pathway does not work in a closed circular transcript in the absence of a dissociative 5′ end. Therefore, some cap-independent translation initiation mechanisms, such as the IRES-dependent pathway and m6A-dependent pathway (Fig. 3), have been proposed to explain the protein-coding ability of some circRNAs. IRESs are sequences that mediate the binding between ribosomes and RNA, thus initiating translation. The reported protein-coding circRNAs driven by IRES include circZNF609 in myogenesis [75], circMbL in fly head extracts [82], circSHPRH and circFBXW7 in glioma tumorigenesis [68, 69], and circβ-catenin in liver cancer growth [71].
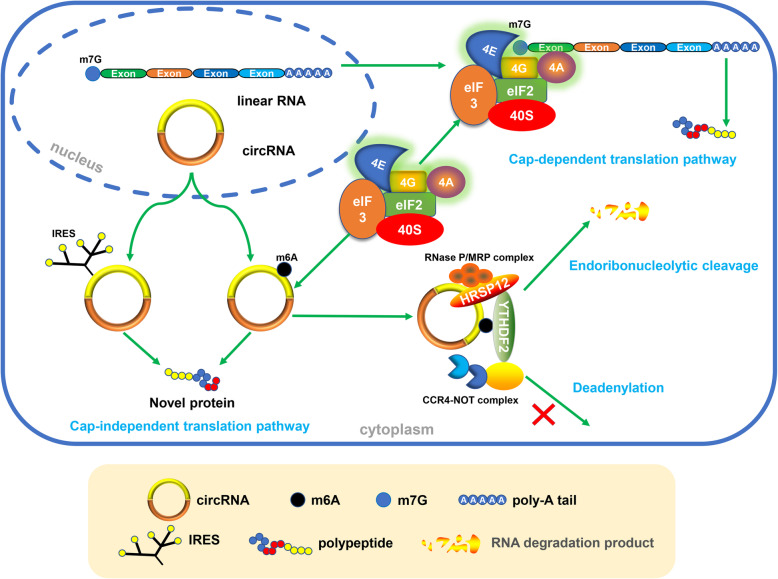
Fig. 3.
M6A modification regulates circRNA translation and degradation. The translation of circRNAs requires m6A modification or IRES, which is different from the traditional cap-dependent pathway of linear RNAs. M6A-modified circRNAs are endoribonuclease-cleaved via the YTHDF2-HRSP12-RNase P/MRP axis
However, a recent study conducted by Yang et al. broadens our horizons on the coding landscape of the human transcriptome. An m6A-driven translation pathway was proposed and verified in cellular responses to environmental stress [78]. In this study, circRNAs containing m6A motifs were detected to be translated, and the efficiency of translation was validated to be modulated by the m6A level. Mechanistically, this m6A-driven translation was initiated by factor eIF4G2 and m6A reader YTHDF3, enhanced by methyltransferase METTL3/14, and inhibited by demethylase FTO. Moreover, the m6A level of some endogenous circRNAs was tested, and the results showed that the m6A motif was abundant in circRNAs. In terms of the whole human transcriptome, m6A-modified circRNAs with coding potential are not rare [85, 86]. Finally, 33 endogenous peptides encoded by the back-splice junctions of circRNAs were chosen for functional analysis. However, regrettably, no functional enrichment was detected despite the translation of these circRNAs being indeed elevated when facing cellular stress.
Notably, these two cap-independent translation pathways might not function independently. Legnini et al. reanalysed m6A-Seq and immunoprecipitation data [15] and combined the data with other m6A immunoprecipitation (IP) results in myoblasts alone [75]. The results showed that a high m6A methylation level was detected in the IRES-activated protein-coding circRNA circZNF609, suggesting a possible connection between these two cap-independent pathways.
M6A modification facilitates circRNA degradation
Due to their closed circular structure, circRNAs are naturally more stable than their parental linear RNAs, as they are not the primary targets of foreign chemicals or exonucleases. This has been validated by many studies related to the characterization of circRNAs [59, 79]. CircRNAs are rarely degraded prior to the corresponding parental linear circRNAs in Actinomycin D and RNase R treatment. However, how circRNA is degraded and what factors contribute to the surveillance pathway remain largely unknown.
A previous study reported that circRNAs with near perfect complementary miRNA target sites could be degraded in an Ago2-slicer-dependent manner, but for those circRNAs without miRNA sponge function or specific microRNA target sites, this method does not work [87]. Another study found that the depletion of GW182 (a key component of the P-body and RNAi machine) resulted in the accumulation of endogenous circular transcripts. However, the depletion of other P-body components or RNAi complex factors did not have similar effects, indicating that GW182, not the P-body or RNAi machine, affected the degradation of circRNAs [88]. Regrettably, GW182 shows little effect on the nuclear export of circRNAs, and its functions in the cytoplasm has not been fully elucidated, so other studies are needed to explain the degradation of circRNA.
The endoribonucleolytic cleavage pathway is one of the pathways by which m6A-modified RNAs are degraded. As a new star in the field of non-coding RNA research, m6A-modified circRNAs were also found to be endoribonuclease-cleaved via a YTHDF2-HRSP12-RNase P/MRP axis [89] (Fig. 3). HRSP12 is an adaptor protein that bridges YTHDF2 (m6A reader protein) and RNase P/MRP (endoribonucleases) to form a YTHDF2-HRSP12-RNase P/MRP complex, for which YTHDF2 is the guide. When an m6A-modified circRNA is recognized by YTHDF2, regardless of whether it occupies an HRSP12-binding site, RNase P/MRP always performs its endonuclease function. The only difference is that the existence of the HRSP12 binding site greatly improves the efficiency of endoribonucleolytic cleavage. Subsequently, the m6A-modified circRNA is selectively downregulated. What follows is a change in the biological function of circRNAs. Thus, we can conclude that one of the ways that m6A modification regulates the biological function of circRNAs is to affect their degradation.
M6A modified circRNA in innate immunity
Innate immunity (also named non-specific immunity) is the natural immune defence function formed by the body in the process of development and evolution. It plays a decisive role in controlling and resolving the inflammatory response to tissue damage [90]. A recent study found that innate immunity can be activated differently by exogenous and endogenous RNAs [91].
All transcripts directly generated by RNA polymerase II bear an m7G cap, and RIG-I (also known as DDX58) senses a triphosphate at the 5′ end [92]; these are essential elements for immune monitoring. Due to the closed circular structure, circRNAs are supposed to be able to escape from the end monitoring system. However, recent studies showed that the invasion of some exogenous circRNAs still leads to potent induction of innate immunity genes and confers protection against viral infection [93], while endogenous circRNAs form some 16–26 bp imperfect RNA duplexes to resist the double-stranded RNA (dsRNA)-activated protein kinase (PKR) in innate immunity [94] (Fig. 4). One of the explanations was found to describe how the immune system defined endogenous versus foreign circRNA as m6A modification.
Fig. 4.
M6A-modified circRNAs in innate immunity and tumours. M6A modification defines endogenous versus foreign circRNA in innate immunity. M6A modification of circNSUN2 promotes the liver metastasis of colorectal cancer by facilitating cytoplasmic export and forming a circNSUN2/IGF2BP2/HMGA2 RNA-protein ternary complex to stabilize HMGA2 mRNA
A study conducted by Y. Grace et al. found that a circRNA generated by ZKSCAN1 introns (circSELF), but not autocatalytic splicing (circFOREIGN), is associated with WTAP and KIAA1429 (m6A writers) as well as YTHDF2 and HNRNPC (m6A readers) [80]. Further research found that different levels of m6A modification were detected in these two circRNAs, and m6A modification marked circRNA as “SELF”. CircSELF can escape innate immunological surveillance via YTHDF2-mediated suppression, which is consistent with a recent study showing that m6A-modified RNAs could be recruited by YTHDF proteins and induced into phase-separated condensates via their N-terminal disordered domains [95]. These results suggest that human circRNAs may be marked by the covalent m6A modification, which is essential for the recognition function of innate immunity.
M6A-modified circRNA in tumours
Since m6A and circRNAs are both closely related to tumours, it is natural to speculate that m6A modification might regulate the function of circRNAs in various tumours. Herein, we briefly review recent studies of m6A-modified circRNAs associated with tumours.
As the third most prevalent and the second most deadly malignancy worldwide, colorectal cancer is still a major threat to human health, especially in China [96]. Clinically, the liver metastasis of colorectal cancer is the most common organ metastasis and leads to poor prognosis beyond 5 years [97]. Recently, Chen et al. found that m6A modification of circNSUN2 promotes the liver metastasis of colorectal cancer by facilitating cytoplasmic export and forming a circNSUN2/IGF2BP2/HMGA2 RNA-protein ternary complex to stabilize HMGA2 mRNA [79] (Fig. 4). HMGA2, a high mobility group AT-hook 2, is already widely believed to be related to the progression of colorectal cancer [98, 99]. These results illuminate how m6A modification affects the interaction between circRNA and RBP.
Cervical cancer is a prevalent gynaecological cancer with a relatively poor prognosis [100], and almost all cervical cancers are caused by oncogenic types of human papillomavirus (HPV) [101]. CircE7 is an oncoprotein-encoding circRNA generated by HPV that is closely related to the growth of CaSki cervical carcinoma cells both in vitro and in vivo. Interestingly, m6A modification is detected and verified to be an essential motif for the protein-coding ability of circE7 [102], which is consistent with the ideas mentioned above that m6A modification facilitates circRNA translation and helps foreign circRNAs escape immune monitoring. Moreover, circE7 is not a special case that is specifically expressed or modified by m6A. Another study identified more than 1 thousand m6A-modified circRNAs in human embryonic stem cells (hESCs) and showed that m6A circRNAs are also abundant in HeLa cells [103], which expands our understanding of the breadth and regulatory aspects of m6A modification.
In addition to modifying circRNA directly, m6A can also affect the function of circRNA via changing the methylation state of downstream molecules. As one of the main response factors downstream of the Hippo pathway, YAP is closely related to the occurrence and development of various tumours [104, 105]. In hepatocellular cancer, circ_104075 can absorb miR-582-3p to stimulate tumorigenesis via YAP [106]. M6A modification in the 3′-UTR of YAP induces the interaction with miR-382-5p and subsequently leads to the inhibition of YAP. Then, the promoting effect of circRNA_104075 on hepatocellular cancer is inhibited. In addition, a combinative bioinformatics prediction of m6A level, IRES and open reading frame (ORF) could indicate the protein-coding potential of circPVRL3 in gastric cancer [107].
Applications and future directions
Considering the stability and conserved nature of their structure, the potential of circRNAs as diagnostic biomarkers and therapeutic targets is unquestionable and is supported by the growing number of circRNA-related studies in recent years [108]. However, the relationship between epigenetic modification and circRNA functions is still largely unknown. As one of the most abundant RNA modifications, m6A provides us with an intermediate mechanism by which circRNAs are regulated by upstream molecules and allows us to predict and interfere with disease progression caused by the dysregulation of circRNAs. There is no doubt that it would greatly expand our understanding of circRNA and drive its applications.
Notably, no specific biological functions have been detected in the majority of already discovered circRNAs, which is also one of the reasons that circRNAs were regarded as by-products of splicing when first discovered [109]. Considering the ubiquitous m6A modification in annotated functional circRNAs, we speculate that it might be related to the tissue and developmental stage specificity of circRNA. That is, specific circRNAs present differential expression only if they have been activated by specific molecular mechanisms, such as m6A, in specific tissues, developmental stages and subcellular locations. To test this conjecture, a combination analysis of the m6A Hi-Res chip and RNA-seq would be helpful for our future research on the biological function and clinical application of m6A-modified circRNAs.
Conclusions
With the broad application of high-throughput sequencing technology and bioinformatics analysis in scientific research, increasing numbers of m6A-modified circRNAs will be found and tested. By then, our understanding of how m6A modification regulates circRNA will not be confined to the four limited aspects of translation, degradation, immunity, and tumours. Other effects of m6A on circRNA, such as processing or splicing effects, and the biological functions of m6A-modified circRNAs in other non-neoplastic diseases could be further investigated.
Since the current understanding of m6A-modified circRNAs is only at the tip of the iceberg, there is still a long way to go to reveal its further regulatory mechanisms and subsequent biological functions in diseases. At this stage, we propose that more m6A regulated circRNAs could be developed to diagnostic biomarkers and therapeutic targets in the future. With the existing technical advancements, it is no longer a technical problem to identify the characterization, localization, transport and degradation of circRNAs in living cells. We anticipate that methods for simplifying the detection of m6A levels of specific circRNAs and for effectively extracting circRNAs with low abundance in limited samples, such as exosomal circRNAs, will progress in the field.
Acknowledgements
Not applicable.
Abbreviations
- 5hmC
5-hydroxymethylcytosine
- AML
Acute myeloid leukaemia
- CDS
Coding sequence
- ceRNA
Competing endogenous RNA
- CircRNA
Circular RNA
- ciRNAs
Intronic circRNAs
- E2F1
E2F transcription factor 1
- ecircRNAs
Exonic circRNAs
- EIciRNAs
Exon-intron circRNAs
- eIF3
Translation initiation factor 3
- FAK
Focal adhesion kinase
- FTO
Fat mass and obesity-associated protein
- GSCs
Glioblastoma stem cells
- HCC
Hepatocellular carcinoma
- hESCs
Human embryonic stem cells
- HIF1-α
Hypoxia inducible factor 1 subunit α
- HNRNP
Heterogeneous nuclear ribonucleoprotein
- HPV
Human papillomavirus
- ID-1
Inhibitor of DNA binding 1
- IGF2BPs
Insulin-like growth factor 2 mRNA-binding proteins
- IP
Immunoprecipitation
- IRES
Internal ribosome entry site
- M1A
N1-methyladenosine
- M5C
5-methylcytosine
- M6A
N6-methyladenosine
- M6Am
N6, 2′-O-dimethyladenosine
- M7G
7-methylguanine
- METTL14
Methyltransferase-like 14 protein
- METTL3
Methyltransferase-like 3 protein
- MRE
miRNA response element
- ORF
Open reading frame
- PRCC2A
Proline rich coiled-coil 2 A
- RBM15/15B
RNA-binding motif protein 15/15B
- RBPs
RNA-binding proteins
- SAM
S-adenosylmethionine
- VIRMA
Vir-like m6A methyltransferase associated
- WTAP
WT1 associated protein
- YTH
YT521-B homology
- ZC3H13
Zinc finger CCCH-type containing 13
Authors’ contributions
LLZ and CFH wrote the manuscript and created the figures. DTY, JBL and ZQS provided direction and guidance throughout the preparation of this manuscript. CC, YXG reviewed and made significant revisions to the manuscript. LLZ collected and prepared the related papers. All authors read and approved the final manuscript.
Funding
This study was supported by The National Natural Science Foundation of China (81972663, 81560385), Key Scientific Research Projects of Institutions of Higher Education in Henan Province (19A310024), The Medical Scientific and Technological Research Project of Henan Province (201702027), The China Postdoctoral Science Foundation (2019 T120648, 2017 M610462), The National Natural Science Foundation of Henan Province (182300410342), The Health Commission Technology Talents Overseas Training Project of Henan Province (2018140) and The Key Scientific Research Project of Henan Higher Education Institutions (20A310024).
Availability of data and materials
Not applicable.
Ethics approval and consent to participate
Not applicable.
Consent for publication
All of the authors are aware of and agree to the content of the paper and their being listed as a co-author of the paper.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Lele Zhang and Chaofeng Hou contributed equally to this work.
Contributor Information
Detao Yin, Email: detaoyin@zzu.edu.cn.
Jinbo Liu, Email: 1999liujb@163.com.
Zhenqiang Sun, Email: zqsun82@csu.edu.cn.
References
- 1.Sanger HL, Klotz G, Riesner D, Gross HJ, Kleinschmidt AK. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc Natl Acad Sci U S A. 1976;73:3852–3856. doi: 10.1073/pnas.73.11.3852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Chen LL, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015;12:381–388. doi: 10.1080/15476286.2015.1020271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Dong W, Bi J, Liu H, Yan D, He Q, Zhou Q, Wang Q, Xie R, Su Y, Yang M, et al. Circular RNA ACVR2A suppresses bladder cancer cells proliferation and metastasis through miR-626/EYA4 axis. Mol Cancer. 2019;18:95. doi: 10.1186/s12943-019-1025-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wang S, Zhang Y, Cai Q, Ma M, Jin LY, Weng M, Zhou D, Tang Z, Wang JD, Quan Z. Circular RNA FOXP1 promotes tumor progression and Warburg effect in gallbladder cancer by regulating PKLR expression. Mol Cancer. 2019;18:145. doi: 10.1186/s12943-019-1078-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Zhu Z, Rong Z, Luo Z, Yu Z, Zhang J, Qiu Z, Huang C. Circular RNA circNHSL1 promotes gastric cancer progression through the miR-1306-3p/SIX1/vimentin axis. Mol Cancer. 2019;18:126. doi: 10.1186/s12943-019-1054-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Dube U, Del-Aguila JL, Li Z, Budde JP, Jiang S, Hsu S, Ibanez L, Fernandez MV, Farias F, Norton J, et al. An atlas of cortical circular RNA expression in Alzheimer disease brains demonstrates clinical and pathological associations. Nat Neurosci. 2019;22:1903–1912. doi: 10.1038/s41593-019-0501-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cardamone G, Paraboschi EM, Rimoldi V, Duga S, Solda G, Asselta R. The characterization of GSDMB splicing and Backsplicing profiles identifies novel isoforms and a circular RNA that are Dysregulated in multiple sclerosis. Int J Mol Sci. 2017;18:576. doi: 10.3390/ijms18030576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Shen J, Chen L, Cheng J, Jin X, Mu Y, Li Q, Xia L, Gao Y, Xia Y. Circular RNA sequencing reveals the molecular mechanism of the effects of acupuncture and moxibustion on endometrial receptivity in patients undergoing infertility treatment. Mol Med Rep. 2019;20:1959–1965. doi: 10.3892/mmr.2019.10386. [DOI] [PubMed] [Google Scholar]
- 9.Meng J, Chen S, Han JX, Qian B, Wang XR, Zhong WL, Qin Y, Zhang H, Gao WF, Lei YY, et al. Twist1 regulates Vimentin through Cul2 circular RNA to promote EMT in hepatocellular carcinoma. Cancer Res. 2018;78:4150–4162. doi: 10.1158/0008-5472.CAN-17-3009. [DOI] [PubMed] [Google Scholar]
- 10.Boccaletto P, Machnicka MA, Purta E, Piatkowski P, Baginski B, Wirecki TK, de Crecy-Lagard V, Ross R, Limbach PA, Kotter A, et al. MODOMICS: a database of RNA modification pathways. 2017 update. Nucleic Acids Res. 2018;46:D303–D307. doi: 10.1093/nar/gkx1030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Li J, Yang X, Qi Z, Sang Y, Liu Y, Xu B, Liu W, Xu Z, Deng Y. The role of mRNA m (6) a methylation in the nervous system. Cell Biosci. 2019;9:66. doi: 10.1186/s13578-019-0330-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wang X, Wu R, Liu Y, Zhao Y, Bi Z, Yao Y, Liu Q, Shi H, Wang F, Wang Y. m (6) a mRNA methylation controls autophagy and adipogenesis by targeting Atg5 and Atg7. Autophagy. 2019:1–15. [DOI] [PMC free article] [PubMed]
- 13.Barbieri I, Tzelepis K, Pandolfini L, Shi J, Millan-Zambrano G, Robson SC, Aspris D, Migliori V, Bannister AJ, Han N, et al. Promoter-bound METTL3 maintains myeloid leukaemia by m (6) A-dependent translation control. Nature. 2017;552:126–131. doi: 10.1038/nature24678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Bartosovic M, Molares HC, Gregorova P, Hrossova D, Kudla G, Vanacova S. N6-methyladenosine demethylase FTO targets pre-mRNAs and regulates alternative splicing and 3′-end processing. Nucleic Acids Res. 2017;45:11356–11370. doi: 10.1093/nar/gkx778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Zhao X, Yang Y, Sun BF, Shi Y, Yang X, Xiao W, Hao YJ, Ping XL, Chen YS, Wang WJ, et al. FTO-dependent demethylation of N6-methyladenosine regulates mRNA splicing and is required for adipogenesis. Cell Res. 2014;24:1403–1419. doi: 10.1038/cr.2014.151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Du H, Zhao Y, He J, Zhang Y, Xi H, Liu M, Ma J, Wu L. YTHDF2 destabilizes m (6) A-containing RNA through direct recruitment of the CCR4-NOT deadenylase complex. Nat Commun. 2016;7:12626. doi: 10.1038/ncomms12626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Cheng M, Sheng L, Gao Q, Xiong Q, Zhang H, Wu M, Liang Y, Zhu F, Zhang Y, Zhang X, et al. The m (6) a methyltransferase METTL3 promotes bladder cancer progression via AFF4/NF-kappaB/MYC signaling network. Oncogene. 2019;38:3667–3680. doi: 10.1038/s41388-019-0683-z. [DOI] [PubMed] [Google Scholar]
- 18.Lin X, Chai G, Wu Y, Li J, Chen F, Liu J, Luo G, Tauler J, Du J, Lin S, et al. RNA m (6) a methylation regulates the epithelial mesenchymal transition of cancer cells and translation of snail. Nat Commun. 2019;10:2065. doi: 10.1038/s41467-019-09865-9. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 19.Chen J, Du B. Novel positioning from obesity to cancer: FTO, an m (6) a RNA demethylase, regulates tumour progression. J Cancer Res Clin Oncol. 2019;145:19–29. doi: 10.1007/s00432-018-2796-0. [DOI] [PubMed] [Google Scholar]
- 20.Ivanova I, Much C, Di Giacomo M, Azzi C, Morgan M, Moreira PN, Monahan J, Carrieri C, Enright AJ, O'Carroll D. The RNA m (6) a reader YTHDF2 is essential for the post-transcriptional regulation of the maternal Transcriptome and oocyte competence. Mol Cell. 2017;67:1059–1067. doi: 10.1016/j.molcel.2017.08.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Winkler R, Gillis E, Lasman L, Safra M, Geula S, Soyris C, Nachshon A, Tai-Schmiedel J, Friedman N, VTK L-T, et al. m (6) a modification controls the innate immune response to infection by targeting type I interferons. Nat Immunol. 2019;20:173–182. doi: 10.1038/s41590-018-0275-z. [DOI] [PubMed] [Google Scholar]
- 22.Weng YL, Wang X, An R, Cassin J, Vissers C, Liu Y, Liu Y, Xu T, Wang X, Wong SZH, et al. Epitranscriptomic m (6) a regulation of axon regeneration in the adult mammalian nervous system. Neuron. 2018;97:313–325. doi: 10.1016/j.neuron.2017.12.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Jia R, Chai P, Wang S, Sun B, Xu Y, Yang Y, Ge S, Jia R, Yang YG, Fan X. m (6) a modification suppresses ocular melanoma through modulating HINT2 mRNA translation. Mol Cancer. 2019;18:161. doi: 10.1186/s12943-019-1088-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Fazi F, Fatica A. Interplay between N (6)-Methyladenosine (m (6) a) and non-coding RNAs in cell development and Cancer. Front Cell Dev Biol. 2019;7:116. doi: 10.3389/fcell.2019.00116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Liu J, Yue Y, Han D, Wang X, Fu Y, Zhang L, Jia G, Yu M, Lu Z, Deng X, et al. A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat Chem Biol. 2014;10:93–95. doi: 10.1038/nchembio.1432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Li F, Yi Y, Miao Y, Long W, Long T, Chen S, Cheng W, Zou C, Zheng Y, Wu X, et al. N (6)-Methyladenosine modulates nonsense-mediated mRNA decay in human Glioblastoma. Cancer Res. 2019;79:5785–5798. doi: 10.1158/0008-5472.CAN-18-2868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Knuckles P, Carl SH, Musheev M, Niehrs C, Wenger A, Buhler M. RNA fate determination through cotranscriptional adenosine methylation and microprocessor binding. Nat Struct Mol Biol. 2017;24:561–569. doi: 10.1038/nsmb.3419. [DOI] [PubMed] [Google Scholar]
- 28.Pendleton KE, Chen B, Liu K, Hunter OV, Xie Y, Tu BP, Conrad NK. The U6 snRNA m (6) a methyltransferase METTL16 regulates SAM Synthetase intron retention. Cell. 2017;169:824–835. doi: 10.1016/j.cell.2017.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Jia G, Fu Y, Zhao X, Dai Q, Zheng G, Yang Y, Yi C, Lindahl T, Pan T, Yang YG, He C. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol. 2011;7:885–887. doi: 10.1038/nchembio.687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Zheng G, Dahl JA, Niu Y, Fedorcsak P, Huang CM, Li CJ, Vagbo CB, Shi Y, Wang WL, Song SH, et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell. 2013;49:18–29. doi: 10.1016/j.molcel.2012.10.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Huang Y, Yan J, Li Q, Li J, Gong S, Zhou H, Gan J, Jiang H, Jia GF, Luo C, Yang CG. Meclofenamic acid selectively inhibits FTO demethylation of m6A over ALKBH5. Nucleic Acids Res. 2015;43:373–384. doi: 10.1093/nar/gku1276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ueda Y, Ooshio I, Fusamae Y, Kitae K, Kawaguchi M, Jingushi K, Hase H, Harada K, Hirata K, Tsujikawa K. AlkB homolog 3-mediated tRNA demethylation promotes protein synthesis in cancer cells. Sci Rep. 2017;7:42271. doi: 10.1038/srep42271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Beharry AA, Lacoste S, O'Connor TR, Kool ET. Fluorescence monitoring of the oxidative repair of DNA alkylation damage by ALKBH3, a prostate Cancer marker. J Am Chem Soc. 2016;138:3647–3650. doi: 10.1021/jacs.6b00986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Liu N, Dai Q, Zheng G, He C, Parisien M, Pan T. N (6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature. 2015;518:560–564. doi: 10.1038/nature14234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Alarcon CR, Goodarzi H, Lee H, Liu X, Tavazoie S, Tavazoie SF. HNRNPA2B1 is a mediator of m (6) A-dependent nuclear RNA processing events. Cell. 2015;162:1299–1308. doi: 10.1016/j.cell.2015.08.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Liu N, Zhou KI, Parisien M, Dai Q, Diatchenko L, Pan T. N6-methyladenosine alters RNA structure to regulate binding of a low-complexity protein. Nucleic Acids Res. 2017;45:6051–6063. doi: 10.1093/nar/gkx141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, Zhao BS, Mesquita A, Liu C, Yuan CL, et al. Recognition of RNA N (6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018;20:285–295. doi: 10.1038/s41556-018-0045-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Wu R, Li A, Sun B, Sun JG, Zhang J, Zhang T, Chen Y, Xiao Y, Gao Y, Zhang Q, et al. A novel m (6) a reader Prrc2a controls oligodendroglial specification and myelination. Cell Res. 2019;29:23–41. doi: 10.1038/s41422-018-0113-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wang S, Chai P, Jia R, Jia R. Novel insights on m (6) a RNA methylation in tumorigenesis: a double-edged sword. Mol Cancer. 2018;17:101. doi: 10.1186/s12943-018-0847-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Takada K, Jameson SC. Naive T cell homeostasis: from awareness of space to a sense of place. Nat Rev Immunol. 2009;9:823–832. doi: 10.1038/nri2657. [DOI] [PubMed] [Google Scholar]
- 41.Haussmann IU, Bodi Z, Sanchez-Moran E, Mongan NP, Archer N, Fray RG, Soller M. m (6) a potentiates Sxl alternative pre-mRNA splicing for robust Drosophila sex determination. Nature. 2016;540:301–304. doi: 10.1038/nature20577. [DOI] [PubMed] [Google Scholar]
- 42.Zhou J, Wan J, Gao X, Zhang X, Jaffrey SR, Qian SB. Dynamic m (6) a mRNA methylation directs translational control of heat shock response. Nature. 2015;526:591–594. doi: 10.1038/nature15377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Aguilo F, Zhang F, Sancho A, Fidalgo M, Di Cecilia S, Vashisht A, Lee DF, Chen CH, Rengasamy M, Andino B, et al. Coordination of m (6) a mRNA methylation and gene transcription by ZFP217 regulates Pluripotency and reprogramming. Cell Stem Cell. 2015;17:689–704. doi: 10.1016/j.stem.2015.09.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Li Z, Weng H, Su R, Weng X, Zuo Z, Li C, Huang H, Nachtergaele S, Dong L, Hu C, et al. FTO plays an oncogenic role in acute myeloid leukemia as a N (6)-Methyladenosine RNA Demethylase. Cancer Cell. 2017;31:127–141. doi: 10.1016/j.ccell.2016.11.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Chen M, Wei L, Law CT, Tsang FH, Shen J, Cheng CL, Tsang LH, Ho DW, Chiu DK, Lee JM, et al. RNA N6-methyladenosine methyltransferase-like 3 promotes liver cancer progression through YTHDF2-dependent posttranscriptional silencing of SOCS2. Hepatology. 2018;67:2254–2270. doi: 10.1002/hep.29683. [DOI] [PubMed] [Google Scholar]
- 46.Cui Q, Shi H, Ye P, Li L, Qu Q, Sun G, Sun G, Lu Z, Huang Y, Yang CG, et al. m (6) a RNA methylation regulates the Self-renewal and tumorigenesis of Glioblastoma stem cells. Cell Rep. 2017;18:2622–2634. doi: 10.1016/j.celrep.2017.02.059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Cai X, Wang X, Cao C, Gao Y, Zhang S, Yang Z, Liu Y, Zhang X, Zhang W, Ye L. HBXIP-elevated methyltransferase METTL3 promotes the progression of breast cancer via inhibiting tumor suppressor let-7g. Cancer Lett. 2018;415:11–19. doi: 10.1016/j.canlet.2017.11.018. [DOI] [PubMed] [Google Scholar]
- 48.Chen I, Chen CY, Chuang TJ. Biogenesis, identification, and function of exonic circular RNAs. Wiley Interdiscip Rev RNA. 2015;6:563–579. doi: 10.1002/wrna.1294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Danan M, Schwartz S, Edelheit S, Sorek R. Transcriptome-wide discovery of circular RNAs in Archaea. Nucleic Acids Res. 2012;40:3131–3142. doi: 10.1093/nar/gkr1009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Wang PL, Bao Y, Yee MC, Barrett SP, Hogan GJ, Olsen MN, Dinneny JR, Brown PO, Salzman J. Circular RNA is expressed across the eukaryotic tree of life. PLoS One. 2014;9:e90859. doi: 10.1371/journal.pone.0090859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Broadbent KM, Broadbent JC, Ribacke U, Wirth D, Rinn JL, Sabeti PC. Strand-specific RNA sequencing in plasmodium falciparum malaria identifies developmentally regulated long non-coding RNA and circular RNA. BMC Genomics. 2015;16:454. doi: 10.1186/s12864-015-1603-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Ji P, Wu W, Chen S, Zheng Y, Zhou L, Zhang J, Cheng H, Yan J, Zhang S, Yang P, Zhao F. Expanded expression landscape and prioritization of circular RNAs in mammals. Cell Rep. 2019;26:3444–3460. doi: 10.1016/j.celrep.2019.02.078. [DOI] [PubMed] [Google Scholar]
- 53.Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, Marzluff WF, Sharpless NE. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19:141–157. doi: 10.1261/rna.035667.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One. 2012;7:e30733. doi: 10.1371/journal.pone.0030733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495:333–338. doi: 10.1038/nature11928. [DOI] [PubMed] [Google Scholar]
- 56.Guo JU, Agarwal V, Guo H, Bartel DP. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 2014;15:409. doi: 10.1186/s13059-014-0409-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Veno MT, Hansen TB, Veno ST, Clausen BH, Grebing M, Finsen B, Holm IE, Kjems J. Spatio-temporal regulation of circular RNA expression during porcine embryonic brain development. Genome Biol. 2015;16:245. doi: 10.1186/s13059-015-0801-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.You X, Vlatkovic I, Babic A, Will T, Epstein I, Tushev G, Akbalik G, Wang M, Glock C, Quedenau C, et al. Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat Neurosci. 2015;18:603–610. doi: 10.1038/nn.3975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Huang X, He M, Huang S, Lin R, Zhan M, Yang D, Shen H, Xu S, Cheng W, Yu J, et al. Circular RNA circERBB2 promotes gallbladder cancer progression by regulating PA2G4-dependent rDNA transcription. Mol Cancer. 2019;18:166. doi: 10.1186/s12943-019-1098-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Xu L, Feng X, Hao X, Wang P, Zhang Y, Zheng X, Li L, Ren S, Zhang M, Xu M. CircSETD3 (Hsa_circ_0000567) acts as a sponge for microRNA-421 inhibiting hepatocellular carcinoma growth. J Exp Clin Cancer Res. 2019;38:98. doi: 10.1186/s13046-019-1041-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Chang H, Qu J, Wang J, Liang X, Sun W. Circular RNA circ_0026134 regulates non-small cell lung cancer cell proliferation and invasion via sponging miR-1256 and miR-1287. Biomed Pharmacother. 2019;112:108743. doi: 10.1016/j.biopha.2019.108743. [DOI] [PubMed] [Google Scholar]
- 62.Wang X, Zhang H, Yang H, Bai M, Ning T, Deng T, Liu R, Fan Q, Zhu K, Li J, et al. Exosome-delivered circRNA promotes glycolysis to induce chemoresistance through the miR-122-PKM2 axis in colorectal cancer. Mol Oncol. 2020;14:539–555. doi: 10.1002/1878-0261.12629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Lin S, Song S, Sun R, Zhang M, Du Y, Zhang D, Xu W, Wang H. Oncogenic circular RNA Hsa-circ-000684 interacts with microRNA-186 to upregulate ZEB1 in gastric cancer. FASEB J. 2020. [DOI] [PubMed]
- 64.Chen W, Quan Y, Fan S, Wang H, Liang J, Huang L, Chen L, Liu Q, He P, Ye Y. Exosome-transmitted circular RNA hsa_circ_0051443 suppresses hepatocellular carcinoma progression. Cancer Lett. 2020;475:119–128. doi: 10.1016/j.canlet.2020.01.022. [DOI] [PubMed] [Google Scholar]
- 65.Yang Q, Du WW, Wu N, Yang W, Awan FM, Fang L, Ma J, Li X, Zeng Y, Yang Z, et al. A circular RNA promotes tumorigenesis by inducing c-myc nuclear translocation. Cell Death Differ. 2017;24:1609–1620. doi: 10.1038/cdd.2017.86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Du WW, Yang W, Liu E, Yang Z, Dhaliwal P, Yang BB. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 2016;44:2846–2858. doi: 10.1093/nar/gkw027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Zhu YJ, Zheng B, Luo GJ, Ma XK, Lu XY, Lin XM, Yang S, Zhao Q, Wu T, Li ZX, et al. Circular RNAs negatively regulate cancer stem cells by physically binding FMRP against CCAR1 complex in hepatocellular carcinoma. Theranostics. 2019;9:3526–3540. doi: 10.7150/thno.32796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Yang Y, Gao X, Zhang M, Yan S, Sun C, Xiao F, Huang N, Yang X, Zhao K, Zhou H, et al. Novel role of FBXW7 circular RNA in repressing Glioma tumorigenesis. J Natl Cancer Inst. 2018;110. [DOI] [PMC free article] [PubMed]
- 69.Zhang M, Huang N, Yang X, Luo J, Yan S, Xiao F, Chen W, Gao X, Zhao K, Zhou H, et al. A novel protein encoded by the circular form of the SHPRH gene suppresses glioma tumorigenesis. Oncogene. 2018;37:1805–1814. doi: 10.1038/s41388-017-0019-9. [DOI] [PubMed] [Google Scholar]
- 70.Zheng X, Chen L, Zhou Y, Wang Q, Zheng Z, Xu B, Wu C, Zhou Q, Hu W, Wu C, Jiang J. A novel protein encoded by a circular RNA circPPP1R12A promotes tumor pathogenesis and metastasis of colon cancer via hippo-YAP signaling. Mol Cancer. 2019;18:47. doi: 10.1186/s12943-019-1010-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Liang WC, Wong CW, Liang PP, Shi M, Cao Y, Rao ST, Tsui SK, Waye MM, Zhang Q, Fu WM, Zhang JF. Translation of the circular RNA circbeta-catenin promotes liver cancer cell growth through activation of the Wnt pathway. Genome Biol. 2019;20:84. doi: 10.1186/s13059-019-1685-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, Kjems J. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495:384–388. doi: 10.1038/nature11993. [DOI] [PubMed] [Google Scholar]
- 73.Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, Zhong G, Yu B, Hu W, Dai L, et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015;22:256–264. doi: 10.1038/nsmb.2959. [DOI] [PubMed] [Google Scholar]
- 74.Du WW, Yang W, Chen Y, Wu ZK, Foster FS, Yang Z, Li X, Yang BB. Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur Heart J. 2017;38:1402–1412. doi: 10.1093/eurheartj/ehw001. [DOI] [PubMed] [Google Scholar]
- 75.Legnini I, Di Timoteo G, Rossi F, Morlando M, Briganti F, Sthandier O, Fatica A, Santini T, Andronache A, Wade M, et al. Circ-ZNF609 is a circular RNA that can be translated and functions in Myogenesis. Mol Cell. 2017;66:22–37. doi: 10.1016/j.molcel.2017.02.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Begum S, Yiu A, Stebbing J, Castellano L. Novel tumour suppressive protein encoded by circular RNA, circ-SHPRH, in glioblastomas. Oncogene. 2018;37:4055–4057. doi: 10.1038/s41388-018-0230-3. [DOI] [PubMed] [Google Scholar]
- 77.Wang Y, Wang Z. Efficient backsplicing produces translatable circular mRNAs. RNA. 2015;21:172–179. doi: 10.1261/rna.048272.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Yang Y, Fan X, Mao M, Song X, Wu P, Zhang Y, Jin Y, Yang Y, Chen LL, Wang Y, et al. Extensive translation of circular RNAs driven by N(6)-methyladenosine. Cell Res. 2017;27:626–641. doi: 10.1038/cr.2017.31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Chen RX, Chen X, Xia LP, Zhang JX, Pan ZZ, Ma XD, Han K, Chen JW, Judde JG, Deas O, et al. N(6)-methyladenosine modification of circNSUN2 facilitates cytoplasmic export and stabilizes HMGA2 to promote colorectal liver metastasis. Nat Commun. 2019;10:4695. doi: 10.1038/s41467-019-12651-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Chen YG, Chen R, Ahmad S, Verma R, Kasturi SP, Amaya L, Broughton JP, Kim J, Cadena C, Pulendran B, et al. N6-Methyladenosine modification controls circular RNA immunity. Mol Cell. 2019;76:96–109. doi: 10.1016/j.molcel.2019.07.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Ma JZ, Yang F, Zhou CC, Liu F, Yuan JH, Wang F, Wang TT, Xu QG, Zhou WP, Sun SH. METTL14 suppresses the metastatic potential of hepatocellular carcinoma by modulating N(6) -methyladenosine-dependent primary MicroRNA processing. Hepatology. 2017;65:529–543. doi: 10.1002/hep.28885. [DOI] [PubMed] [Google Scholar]
- 82.Pamudurti NR, Bartok O, Jens M, Ashwal-Fluss R, Stottmeister C, Ruhe L, Hanan M, Wyler E, Perez-Hernandez D, Ramberger E, et al. Translation of CircRNAs. Mol Cell. 2017;66:9–21. doi: 10.1016/j.molcel.2017.02.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Pelletier J, Sonenberg N. The organizing principles of eukaryotic ribosome recruitment. Annu Rev Biochem. 2019;88:307–335. doi: 10.1146/annurev-biochem-013118-111042. [DOI] [PubMed] [Google Scholar]
- 84.Haimov O, Sinvani H, Dikstein R. Cap-dependent, scanning-free translation initiation mechanisms. Biochim Biophys Acta. 1849;2015:1313–1318. doi: 10.1016/j.bbagrm.2015.09.006. [DOI] [PubMed] [Google Scholar]
- 85.Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell. 2012;149:1635–1646. doi: 10.1016/j.cell.2012.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, Cesarkas K, Jacob-Hirsch J, Amariglio N, Kupiec M, et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature. 2012;485:201–206. doi: 10.1038/nature11112. [DOI] [PubMed] [Google Scholar]
- 87.Hansen TB, Wiklund ED, Bramsen JB, Villadsen SB, Statham AL, Clark SJ, Kjems J. miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO J. 2011;30:4414–4422. doi: 10.1038/emboj.2011.359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Jia R, Xiao MS, Li Z, Shan G, Huang C. Defining an evolutionarily conserved role of GW182 in circular RNA degradation. Cell Discov. 2019;5:45. doi: 10.1038/s41421-019-0113-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Park OH, Ha H, Lee Y, Boo SH, Kwon DH, Song HK, Kim YK. Endoribonucleolytic cleavage of m(6)A-containing RNAs by RNase P/MRP complex. Mol Cell. 2019;74:494–507. doi: 10.1016/j.molcel.2019.02.034. [DOI] [PubMed] [Google Scholar]
- 90.Kany S, Janicova A, Relja B. Innate immunity and alcohol. J Clin Med. 2019;8. [DOI] [PMC free article] [PubMed]
- 91.Streicher F, Jouvenet N. Stimulation of innate immunity by host and viral RNAs. Trends Immunol. 2019;40:1134–1148. doi: 10.1016/j.it.2019.10.009. [DOI] [PubMed] [Google Scholar]
- 92.Dickey TH, Song B, Pyle AM. RNA binding activates RIG-I by releasing an autorepressed signaling domain. Sci Adv. 2019;5:eaax3641. doi: 10.1126/sciadv.aax3641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Chen YG, Kim MV, Chen X, Batista PJ, Aoyama S, Wilusz JE, Iwasaki A, Chang HY. Sensing Self and foreign circular RNAs by intron identity. Mol Cell. 2017;67:228–238. doi: 10.1016/j.molcel.2017.05.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Liu CX, Li X, Nan F, Jiang S, Gao X, Guo SK, Xue W, Cui Y, Dong K, Ding H, et al. Structure and degradation of circular RNAs regulate PKR activation in innate immunity. Cell. 2019;177:865–880. doi: 10.1016/j.cell.2019.03.046. [DOI] [PubMed] [Google Scholar]
- 95.Ries RJ, Zaccara S, Klein P, Olarerin-George A, Namkoong S, Pickering BF, Patil DP, Kwak H, Lee JH, Jaffrey SR. m(6) a enhances the phase separation potential of mRNA. Nature. 2019;571:424–428. doi: 10.1038/s41586-019-1374-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424. doi: 10.3322/caac.21492. [DOI] [PubMed] [Google Scholar]
- 97.Grossman JG, Nywening TM, Belt BA, Panni RZ, Krasnick BA, DeNardo DG, Hawkins WG, Goedegebuure SP, Linehan DC, Fields RC. Recruitment of CCR2(+) tumor associated macrophage to sites of liver metastasis confers a poor prognosis in human colorectal cancer. Oncoimmunology. 2018;7:e1470729. doi: 10.1080/2162402X.2018.1470729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Wang X, Liu X, Li AY, Chen L, Lai L, Lin HH, Hu S, Yao L, Peng J, Loera S, et al. Overexpression of HMGA2 promotes metastasis and impacts survival of colorectal cancers. Clin Cancer Res. 2011;17:2570–2580. doi: 10.1158/1078-0432.CCR-10-2542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Chang HY, Ye SP, Pan SL, Kuo TT, Liu BC, Chen YL, Huang TC. Overexpression of miR-194 reverses HMGA2-driven signatures in colorectal Cancer. Theranostics. 2017;7:3889–3900. doi: 10.7150/thno.20041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Liu R, Shuai Y, Luo J, Zhang Z. SEMA3C promotes cervical Cancer growth and is associated with poor prognosis. Front Oncol. 2019;9:1035. doi: 10.3389/fonc.2019.01035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Chibwesha CJ, Stringer JSA. Cervical Cancer as a global concern: contributions of the dual epidemics of HPV and HIV. JAMA. 2019;322:1558–1560. doi: 10.1001/jama.2019.16176. [DOI] [PubMed] [Google Scholar]
- 102.Zhao J, Lee EE, Kim J, Yang R, Chamseddin B, Ni C, Gusho E, Xie Y, Chiang CM, Buszczak M, et al. Transforming activity of an oncoprotein-encoding circular RNA from human papillomavirus. Nat Commun. 2019;10:2300. doi: 10.1038/s41467-019-10246-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Zhou C, Molinie B, Daneshvar K, Pondick JV, Wang J, Van Wittenberghe N, Xing Y, Giallourakis CC, Mullen AC. Genome-wide maps of m6A circRNAs identify widespread and cell-type-specific methylation patterns that are distinct from mRNAs. Cell Rep. 2017;20:2262–2276. doi: 10.1016/j.celrep.2017.08.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.White SM, Avantaggiati ML, Nemazanyy I, Di Poto C, Yang Y, Pende M, Gibney GT, Ressom HW, Field J, Atkins MB, Yi C. YAP/TAZ inhibition induces metabolic and signaling rewiring resulting in targetable vulnerabilities in NF2-deficient tumor cells. Dev Cell. 2019;49:425–443. doi: 10.1016/j.devcel.2019.04.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Wang L, Wang C, Tao Z, Zhao L, Zhu Z, Wu W, He Y, Chen H, Zheng B, Huang X, et al. Curcumin derivative WZ35 inhibits tumor cell growth via ROS-YAP-JNK signaling pathway in breast cancer. J Exp Clin Cancer Res. 2019;38:460. doi: 10.1186/s13046-019-1424-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Zhang X, Xu Y, Qian Z, Zheng W, Wu Q, Chen Y, Zhu G, Liu Y, Bian Z, Xu W, et al. circRNA_104075 stimulates YAP-dependent tumorigenesis through the regulation of HNF4a and may serve as a diagnostic marker in hepatocellular carcinoma. Cell Death Dis. 2018;9:1091. doi: 10.1038/s41419-018-1132-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Sun HD, Xu ZP, Sun ZQ, Zhu B, Wang Q, Zhou J, Jin H, Zhao A, Tang WW, Cao XF. Down-regulation of circPVRL3 promotes the proliferation and migration of gastric cancer cells. Sci Rep. 2018;8:10111. doi: 10.1038/s41598-018-27837-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Wang Y, Wu C, Zhang Y, Yang Y, Ren Z, Lammi MJ, Guo X. Screening for differentially expressed circRNA between Kashin-Beck disease and osteoarthritis patients based on circRNA chips. Clin Chim Acta. 2020;501:92–101. doi: 10.1016/j.cca.2019.10.026. [DOI] [PubMed] [Google Scholar]
- 109.Nigro JM, Cho KR, Fearon ER, Kern SE, Ruppert JM, Oliner JD, Kinzler KW, Vogelstein B. Scrambled exons. Cell. 1991;64:607–613. doi: 10.1016/0092-8674(91)90244-s. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable.