Abstract
Superbugs’ resistivity against available natural products has become an alarming global threat, causing a rapid deterioration in public health and claiming tens of thousands of lives yearly. Although the rapid discovery of small molecules from plant and microbial origin with enhanced bioactivity has provided us with some hope, a rapid hike in the resistivity of superbugs has proven to be the biggest therapeutic hurdle of all times. Moreover, several distinct mechanisms endowed by these notorious superbugs make them immune to these antibiotics subsequently causing our antibiotic wardrobe to be obsolete. In this unfortunate situation, though the time frame for discovering novel “hit molecules” down the line remains largely unknown, our small hope and untiring efforts injected in hunting novel chemical scaffolds with unique molecular targets using high-throughput technologies may safeguard us against these life-threatening challenges to some extent. Amid this crisis, the current comprehensive review highlights the present status of knowledge, our search for bacteria Achilles’ heel, distinct molecular signaling that an opportunistic pathogen bestows to trespass the toxicity of antibiotics, and facile strategies and appealing therapeutic targets of novel drugs. Herein, we also discuss multidimensional strategies to combat antimicrobial resistance.
Keywords: antibiotic therapy, antibiotic-binding proteins, antimicrobial resistance, drug delivery systems, drug repurposing, pharmacodynamics, receptors
1. Introduction
1.1. Superbugs
The modern era has observed a worldwide explosion and tremendous upsurge in notorious, opportunistic, and recalcitrant superbugs that are highly resistant to antibiotics,1,2 and are the virulent cause of infections in humans and animals.3 Currently observed notorious hospital key pathogens are the ESKAPE group of organisms (Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter spp.).4 These pathogens endow intrinsic drug resistance that allows them to circumvent the effects of potent and bioactive chemical scaffolds, subsequently proving our antibiotics wardrobe to be obsolete. Bacteria are champions of evolution, and their frequently observed antibiotic resistance is due to the acquisition of resistant genes laterally from the environment,5 and incorporating them either in their plasmid or their genome. The environment harbors plenty of DNA from dead cadavers and these opportunistic pathogens are voracious grave eaters.6 This acquisition of foreign DNA mainly by conjugation enables them to activate their defensive manoeuvers and acquire immunity to different classes of compounds.6,7
Superbugs are eventually acquiring resistance to all classes of known compounds, either of natural or synthetic origin. Moreover, a glycopeptide antibiotic, vancomycin, with its target in preventing peptidoglycan synthesis of the cell envelope in Gram-positive bacteria, such as enterococci, streptococci, and staphylococci proved obsolete because of the presence of vancomycin resistant plasmids in these pathogens.2,8 However, the last resort antibiotic daptomycin possesses a special mechanism, which affects multiple functional aspects of bacterial cell membrane by inserting into the cell membrane in a phosphatidylglycerol-dependent fashion. This aggregation of daptomycin revamps membrane curvature creating a leakage from which ions ooze out. The leakage causes rapid depolarization subsequently causing the loss of membrane potential, which in turn leads to the inhibition of nucleotide and halting of protein synthesis. The inhibition of nucleotide synthesis eventually causes death of the bacterial cell.9 Therefore, the hunt for alternative and novel sources of chemical scaffolds with promising therapeutic leads, improved facile strategies of selecting potential drug candidates, novel break-through in searching magic bullets, and novel delivering mechanisms to the actual target is of crucial need.
1.2. Beneficial Microbes
A gram of soil harbors a 1 × 109 microbial population,10 and the origin of ca. > 80% of current antibiotics (either microbial synthesis or semisynthetic) are from soil-dwelling microbes.11 Thus, soil organisms (bacterium, fungus, and archaea) are prolific sources of hidden chemical diversity. Among the antibiotics producing organisms, Streptomyces are known to synthesize the highest number of antibiotics so far.12 These organisms can produce about 100 000 antibiotics; however, only a tiny fraction (<1%) of the antibiotics have been structurally elucidated.13−15 At the same time, each ecological niche is also a house and a major reservoir of resistant genes being accumulated over time by dead organisms. The pathogens residing in this atmosphere acquire resistivity against multidrugs by the acquisition of resistant genes through lateral transfer.16 Proof for this could be made through metagenomics data that provide ample evidence regarding a chunk of antibiotic-resistant genes housed in environmental soil, which we collectively refer to as “antibiotics resistome”.
Even though most microbes are classified as pathogens and have lethal properties, not all microbes currently known could be categorized as being harmful, which we refer to as “innocent bacteria”. Notable examples of these include the members belonging to the order Actinomycetales, which have profound tendencies to generate small-molecules of high therapeutic leads. These small-molecules (such as doxorubicin and daunorubicin from S. peucetius(17−19)) act as magic bullets with several clinical functions and have been widely utilized as blockbusters in sabotaging the virulence pathways of several notorious diseases. Moreover, teixobactin (a newly discovered antibiotic) is found effective against vancomycin-resistant enterococcus20 and Mycobacterium tuberculosis.21 Besides, with the advent of the “New Golden age” of antibiotics with the use of synthetic biology approaches that are integrated with in-silico simulations and combinatorial chemistry, we have subsequently gained our abilities to synthesize novel natural products with high therapeutic leads.22 Also, synthetic bioactive peptides have been recently formulated.23−26
1.3. Natural Products and Antibiotics and Their Classification
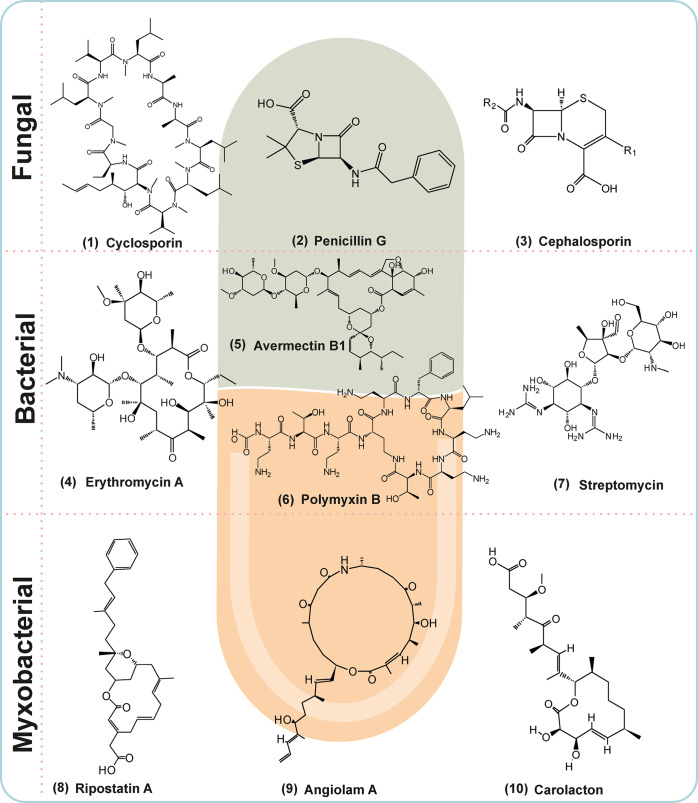
Natural products are evolutionarily optimized as drug-like molecules and are the best sources of high therapeutic leads.27 Fungi, bacteria, myxobacteria, and archaea are efficient producers of small molecules with enhanced bioactivity. Some frequently used fungal antibiotics include cephalosporin, cyclosporin, and penicillin G, while antibiotics of bacterial origin include avermectin B1, erythromycin A, polymyxin B, and streptomycin. Natural products from myxobacterial origin with antibacterial activity include angiolam A, carolacton, and ripostatin A (Figure 1). In addition, archaea are efficient producers of archaeocin antibiotics. For instance, halocins and sulfolobicins are produced by archaea belonging to the order Halobacteriales (Euryarchaeota) and Sulfolobales (Crenarchaeota), respectively.28−30
Figure 1.
Some notable examples of natural products synthesized by different micro-organisms: (1) Cyclosporin (immunosuppressant medication and natural product). (2 and 3) Penicillin G and cephalosporin (β-lactam antibiotics used in the treatment of the infections caused by staphylococci and streptococci). (4) Erythromycin A (a broad-spectrum macrolide antibiotic used in the treatment of bacterial infections; biosynthesized by Streptomyces erythreus). (5) Avermectin B1 (macrocyclic lactone derivatives with potent insecticidal and anthelmintic properties). (6) Polymyxin B (a polypeptide antibiotic biosynthesized by Paenibacillus polymyxa). (7) Streptomycin (an aminoglycoside antibiotic biosynthesized by Streptomyces griseus). (8) Ripostatin A (inhibitors of eubacterial RNA polymerase isolated from myxobacterium; Sorangium cellulosum). (9 and 10) Angiolam A and carolacton (isolated from myxobacteria Angiococcus disciformis and Sorangium cellulosum, respectively).
According to the mode of action and based on their chemical scaffolds, antibiotics have been broadly classified into several classes, namely, β-lactams, aminoglycosides, ansamycins, chloramphenicol, glycopeptides, lipopeptides, macrolides, quinolones, streptogramins, sulfonamides, and tetracyclines. Among these, chloramphenicol, macrolides, tetracyclines, and sulfonamides have bacteriostatic properties, while the remaining ones are bactericidal.31,32Figure 1 lists several key natural products and their source of origin.
1.4. Antibiotics Discovery and Developmental Pipeline
After an accidental discovery of antibiotic penicillin from a mold Penicillium notatum by Alexander Flemming in 1928, there was a growing interest in a search for “magic bullets” with huge therapeutic leads, and thus numerous metabolites with high bioactivity were identified thereafter. This allowed us to efficiently characterize most natural products of microbial origin during the 1950s and 1960s, which we refer to as the “Golden Age” of natural products drug discovery. The growing rate of finding new “hit molecules” severely declined thereafter (after the 1980s). Even though a large chunk of investment has been injected for discovering novel drugs, a highly repetitive rate of finding previously discovered molecules, the slow pace of novel drug discovery and development, and high attrition rates eventually caused global pharmaceutical industries to divert their attention.33 Instead, this led them to turn their interest in structure-based drug design (e.g., molecular docking), mechanism-based screening for bioassay-guided fractionation, and combinatorial chemistry.22 These approaches have now become the mainstay of high-throughput screening.34 However, with technological advancement and various new strategies being adopted, there is now a renaissance of interest in searching for the “hit molecules” (reviewed in ref (22)). Repurposing the old drugs to cure rare and common diseases has become an appealing therapeutic innovation lately because of the reduced costs, shorter-development timelines, and the use of de-risked compounds.33 This could also add a significant contribution to the “New Golden age” of natural product discovery.22 A notable example is the repurposing of a drug named ‘sildenafil citrate (Viagra)’ which is now used to cure erectile dysfunction; however, it was originally developed as an antihypertensive drug.35 Other examples of repurposed drugs include aspirin,36 raloxifene, and propranolol,37 and they have been targeted mostly to neurology, psychiatry, oncology, and infectious diseases. Thalidomide is a repurposed drug widely in use to cure leprosy and multiple myeloma.35 Some newly discovered small bioactive molecules are undergoing investigations using human cell cultures or animal models, but will take some time before they pass through phase III, and are channeled to the public markets for human consumption. Also, identification of a depsipeptide namely “teixobactin” from an uncultured environmental soil bacteria, ‘Eleftheria terrae’ using the bacterial iCHIP has been made very recently.20
Until last June 2019, ca. 42 potential antibiotics supposed to cure bacterial infections have been developed and are in the clinical development phase. Some of these drugs are xerava (eravacycline) and nuzyra (omadacycline) belonging to tetracycline class of compounds and target the 30S subunit of bacterial ribosomes especially against the ESKAPE pathogens. The U.S. Food and Drug Administration (FDA) approved these drugs in 2018. Other potential drugs currently in pending application are iclaprim (targets penicillin binding proteins; PBPs), cefiderocol (S-649266) (targets PBP), imipenem/cilastatin (targets PBP+ β-lactamase), relebactam (MK-7655A) (targets PBP + β-lactamase), lefamulin (BC-3781) (targets 50S subunit), and lascufloxacin (KRP-AM1977).38 According to the FDA, only 20% of Phase I drugs with possible roles to cure the bacterial infections will finally receive the FDA approval.
1.5. Antibiotics Developmental Horizon in the Last Two Decades
During the past two decades, more than 30 new antibiotics have been developed so far.39 Most of them are synthetic-derived (>50%), some natural product derived (40%), and the rest are new β-lactams/β-lactamase inhibitor combinations. Two-thirds of antibiotics derived from natural products are from an actinomycete, and the rest are from a fungal origin. Twenty percent of these antibiotics were classified as first-class antibiotics, such as bedaquiline (diarylquinoline), daptomycin (lipopeptide), fidaxomicin (tiacumicin), linezolid (oxazolidinone), and retapamulin (pleuromutilin). However, these antibiotics are active only against Gram-positive bacteria and have no antagonistic effects against Gram-negative pathogens. A single antibiotic namely diazabicyclooctane-type β-lactamase inhibitor (avibactam) was found to be effective against Gram-negative bacteria when used in combination with some β-lactams.39 Given the recalcitrant nature of multidrug resistant Gram-negative bacteria, a hunt for novel antibiotics that are highly active to these particular groups of bugs seems essential.
1.6. Pros and Cons of Previously Discovered Antibiotics
Some antibiotics such as chloramphenicol, lincosamides, macrolides, spectinomycin, sulfonamides, tetracyclines, and trimethoprim are bacteriostatic in nature, while some have bactericidal ((β-lactam antibiotics such as carbapenems, cephalosporins, monobactams, penicillin derivatives) and vancomycin) properties. These antibiotics have a very fast mode of action and are easy to administer. However, administrating higher doses of these antibiotics may cause the pathogens to build resistance and evolve gradually against the effect of these drugs. Moreover, the host immune system will suffer from negative consequences upon long exposure to antibiotics. Irrational use of antibiotics is a key driver of antibiotic resistance and is very detrimental, which includes prescription of incorrect doses, self-medication, and treatment of nonbacterial illness.40,41 In today’s context in which new and novel therapeutic agents are urgently required, a revisit to natural/herbal products that possess a novel mode of action for combating against the pathogens could prevent the spread of notorious pathogens.
Though we have discovered a lot of “hit molecules” with promising bioactivity against pathogens, unfortunately, all antibiotics currently available in the markets are not safe to use. Antibiotics, such as ketek (telithromycin), initially supposed to cure respiratory tract infections, have instigated the attention of the FDA, and have gained a boxed-warning because of its life-threatening side-effects (hepatotoxicity).42 Also, the negative consequences of using fluoroquinolone antibiotics that are likely to cause mental health problems along with low blood sugar disturbances have been brought to general notice by the FDA in 2018.43 Lately, antibiotics added to the soaps (such as triclosan and triclocarban), wipes, and cleaning supplies have reduced pathogens infection to human health.44 However, the FDA has banned the use of some other antibiotics (such as hexachlorophene) to be used as a soap supplement as it has negative consequences on human health (embryotoxic and produces some teratogenic effects).45 Honey added to the soaps act as a natural disinfectant, and is safe to use. Nevertheless, there has been a mixed perception of whether to consider these soap additives (active ingredient triclosan) as useful or are very harmful.44
1.7. Aspects and Challenges of Antibiotic Drug Development
A cocktail of antibiotics with other adjuvants can also facilitate to permeabilize the drugs across the cell membrane barrier and reach to the actual targets.46 This “Trojan Horse” strategy (achieving fast or facilitated antibiotics uptake) has been found effective in combining antibiotics with siderophores which facilitates the uptake of antibiotics by pathogens.47−49 Such strategy involves an efficient use of a specific Gram-positive antibiotic on Gram-negative bacteria, that eventually causes the bacterial suicide.50 A concoction of amoxicillin and clavulanic acid is being currently used as a front-line antibiotic.51 Moreover, a combination of different drugs has provided us with some hope as is observed in the treatment of tuberculosis and HIV; however, it may not be that effective in treating other ailments. Antibiotics currently in-use have more than one target, and unveiling actual targets and mode of action of new natural products will greatly facilitate resolving the crisis of antibiotics discovery.52 However, our severe negligence such as the wide use of broad-spectrum antibiotics, exposing pathogens to low-concentration of antibiotics below MIC, and their aberrant use even when not needed have forced pathogens to acquire mutations and evolve gradually. This strategy has allowed pathogens to circumvent the effects of traditional drugs.
2. Bacterial Membrane Architecture
The bacterial cell possesses a complex multilayered structure that helps them to thrive and afford protection against different hostile environments,53 and also limits antibiotic penetration. Owing to structural differences of their cell envelopes, they are further categorized into two major groups, namely, Gram-negative and Gram-positive bacteria. This differential membrane architecture affords varying permeability for the entry and/exit of drugs across the membrane. For instance, Gram-negative bacteria harbor an extra peptidoglycan layer compared to Gram-positive ones, which makes the former to be more recalcitrant to most antibiotics. Furthermore, an antibiotic highly lethal against a Gram-positive bacterium may find it difficult to trespass an outer membrane and peptidoglycan layer of Gram-negative bacterium, in turn with no effects of suppressing the growth of these bacteria. Thus, Gram-positive bacteria show considerable susceptibility to these antibiotics compared to their counterparts. This differential membrane architecture has further caused difficulties in disease diagnostics and their effective treatment. Some notorious Gram-positive organisms include glycopeptide-intermediate sensitivity S. aureus (GISA),54−56 methicillin-resistant Staphylococcus aureus (MRSA),57,58 penicillin-resistant Streptococcus pneumoniae, vancomycin-resistant Enterococcus (VRE) species, etc. The highly resistant Gram-negative bacteria include multidrug-resistant Acinetobacter baumannii, Pseudomonas aeruginosa, Stenotrophomonas maltophilia, etc. Besides, targeting the bacterial cell membrane by antibiotics is comparatively easier than the eukaryotic cells. This is because the anionic lipids in bacteria are exposed on the surface of the membranes, while in eukaryotes these anionic lipids are sequestered to the monolayer facing the interior of the cell or organelle. It is one of the crucial reason that most of the antimicrobial agents are cationic which facilitates greater selectivity for the bacterial membranes.59−61
2.1. Cell-Envelope of Gram-Negative Bacteria
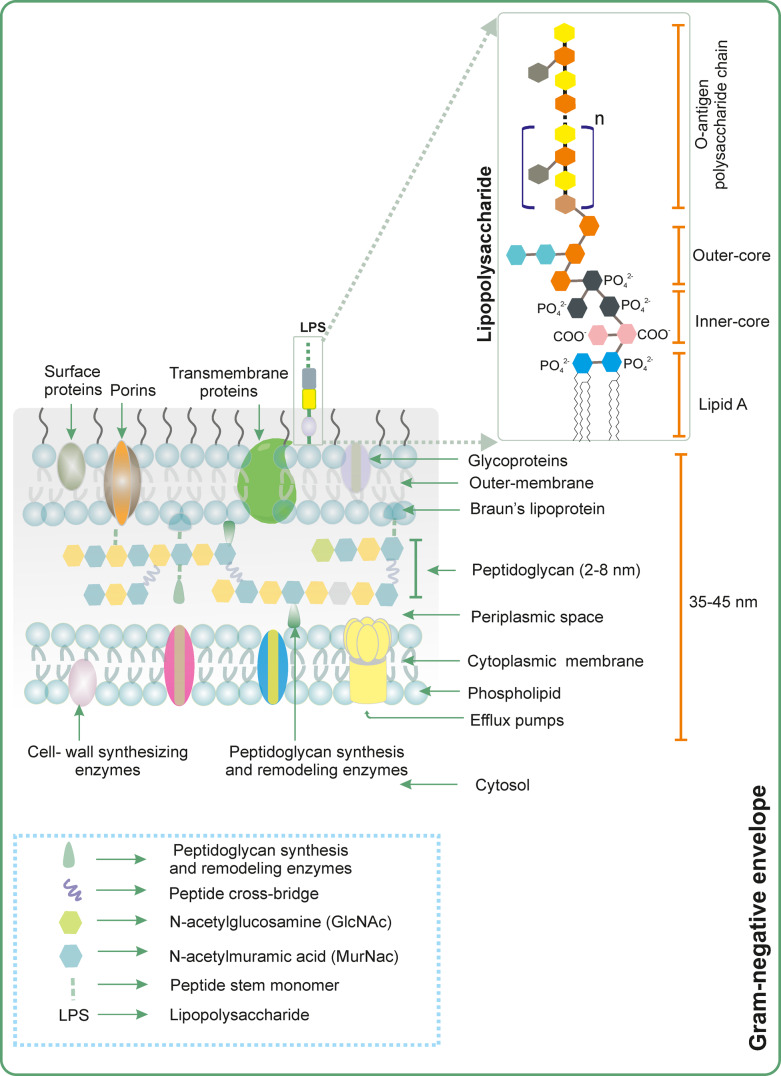
The cell envelope of Gram-negative bacteria comprises an asymmetric phospholipid layer, a thin peptidoglycan layer with the functionality of providing cell rigidity, and outer-membrane that is composed of inner phospholipid leaflet, and an outer leaflet of lipopolysaccharide.62−64 The lipid A comprises a bilayer formed by lipids, and inner-layer to this bilayer (called liposaccharide, LPS) is attached a set of core sugars where the first two of them are KDO [3-deoxy-d-manno-oct-2-ulosonic acid (also known as ketodeoxyoctulosonate)]. The link that attaches lipid A and KDO2 affords viability to the Gram-negative bacteria. To the inner side of the core is O-antigen oligosaccharide repeating units which for E. coli average 10–16 units, however for Shigella flexneri, they account for greater than 90.65 These long chains of carbohydrates are hydrated so that they form a barrier to the passage of hydrophobic molecules. These three moieties of LPS form a large amphipathic polymer66 (Figure 2).
Figure 2.
Gram-negative envelope of prokaryotes showing the extension of lipopolysaccharide (LPS) at its outer membrane. The arrangement of different segments of LPS: the O-antigen polysaccharide chain, outer core, inner-core, and lipid A is presented. In addition, different enzymes, proteins, and pumps that are localized in the Gram-negative bacteria are presented.
Smooth wild-type LPS as observed in Pseudomonas aeruginosa is less permeable than rough mutants lacking LPS core sugars as seen in E. coli and Salmonella Typhimurium.67 The outer membrane restricts the flow of both hydrophobic and hydrophilic molecules across the membrane. These are more resistant to amphiphilic and lipophilic inhibitors (antibiotics, dyes, detergents, free fatty acids and other chemotherapeutic agents) compared to Gram-positive bacteria.68 These LPSs also help to sustain the organism from a harsh environment and form biofilms in the human body eventually evading the human immune system.63 However, the porins which are water-filled β-barrel channels provide an easy access for the required nutrients and metabolic substrates through the lipid bilayer.69E. coli possesses trimeric porins, such as OmpF and OmpC that allow passage of small hydrophilic substances; however, P. aeruginosa is devoid of these trimeric porins. But instead, P. aeruginosa has monomeric porins such as OprF that allow slow permeation,70,71 while OprD is responsible for the uptake of basic amino acid, peptides plus carbapenems.72,73 Also, increased pump expression is caused by decreased porin expression as in nfxC mutants of P. aeruginosa where OprD downregulation causes impairment of carbapenem uptake, while MexEF-OprN is overexpressed, affecting the export of fluoroquinolone.74,75 In addition, the outer membrane of LPS is partially phosphorylated (the P-group confers net negative charge) and provides resistivity for the permeation of most antibiotics, causing pathogens such as P. aeruginosa to be recalcitrant against antibacterial agents.66,69 Moreover, an easy acquisition of resistant genes by Gram-negative bacteria through horizontal gene transfer and spontaneous mutations make them difficult to target and control.76
2.2. Cell-Envelope of Gram-Positive Bacteria
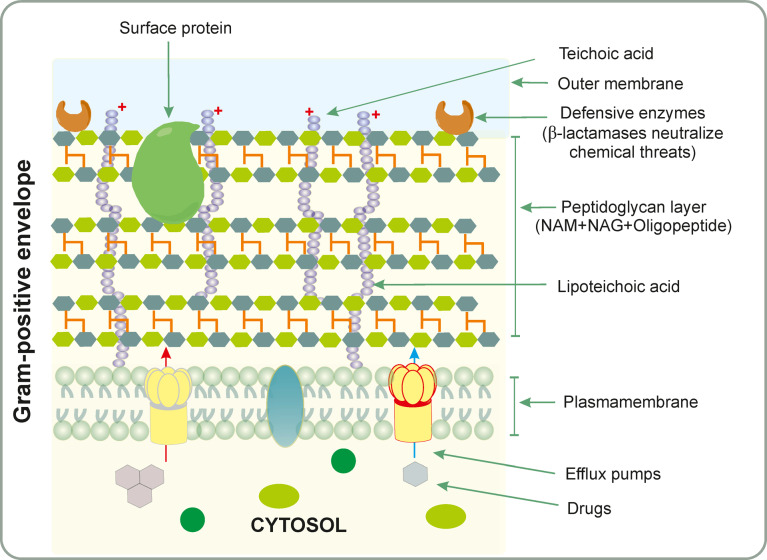
Gram-positive bacteria are devoid of an outer-membrane and have several layers of peptidoglycan unlike the thin layer in Gram-negative ones (Figure 3). Long anionic polymers of teichoic acids arise from these peptidoglycan layers.53 The rigid cell wall made up of the peptidoglycan layer in Gram-positive bacteria are structurally cross-linked, in which monomers of N-acetylglucosamine (NAG) and N-acetylmuramic acid (NAM) are inserted as a new building block of peptidoglycan giving rise to a chain of d-ala-d-ala.77 Antibiotics, such as vancomycin, target these building blocks (NAG and NAM), and prevent transpeptidase to act on these building blocks, in turn, preventing the cross-linking of the peptidoglycan layer. This causes the peptidoglycan layer to be more permeable for the influx of drugs, and the leaking out of bacterial cellular contents makes them prone to lysis.78 Recent identification has led to the findings that the complete saturation of the cell membrane with peptides also may lead to permeabilization and eventual membrane disruption.79 The modifications in the peptidoglycan cause an organism to evade innate immunity which involves amidation of d-Glu, and modification of l-ala by Gly and l-ornithine instead of meso-DAP) and glycan chains (O-acetylation of NAM, N-deacetylation of NAG and glycosylation of NAM) of PGN.80−82 This interconversion of phosphatidylglycerol from anionic to cationic or zwitterionic form by chemically utilizing bacterial enzymes gives them increased tolerance toward polycationic antimicrobial agents.83
Figure 3.
Gram-positive envelope of prokaryotes showing the teichoic acid which provides a net positive charge to the cell. The arrangement of different pumps and surface proteins has been revealed.
3. Antibiotics Targets and Resistance by Bacteria
3.1. Different Mechanism of Actions and Their Targets
Several possible mechanisms of drug targets for the generation of the next novel antibiotics can be envisaged and are discussed in detail as follows.
3.1.1. Cell-Wall Targeting Antibiotics
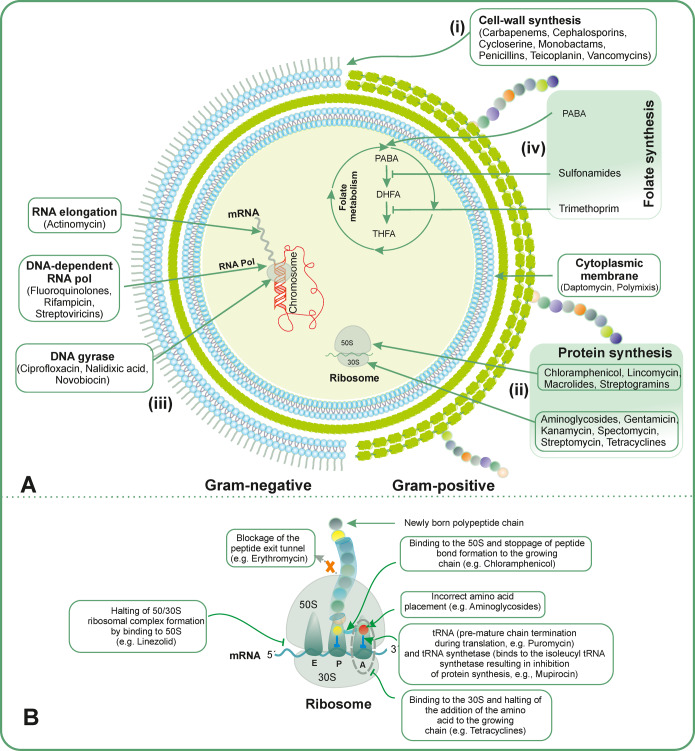
Some antibiotics tend to weaken or disrupt the bacterial plasma-membrane/cell, making the cell wall more prone to cell lysis.84 Antibiotics such as ampicillin target the peptidoglycan layer and inhibit the synthesis of bacterial cell walls. Other notable examples of this class of antibiotics include bacitracin, carbapenems, cephalosporins, penicillins, and vancomycin. Penicillin G- a β-lactam- targets the biosynthesis of the bacterial cell wall.85 The synthesis of other derivatives of this compound with additional scaffolds such as carbapenems, cephalosporins, and monobactams has also been performed.86 Glycopeptides that include vancomycin and teicoplanin also target the cell wall of Gram-positive infections. These glycopeptides bind to the d-Ala-d-Ala termini of membrane-linked cell wall precursors, blocking further incorporation to the growing peptidoglycan chain, and thus preventing the formation of cross-links87−89 (Figure 4A(i)).
Figure 4.
Antibiotics target in both Gram-positive and Gram-negative pathogens. (A) Different antibiotics with their major targets are shown. (B) Antibiotics with their major target in Ribosomal subunits that cause a halt in protein synthesis are depicted.
Some compounds, such as daptomycin, defensins, and polymyxin B have cationic properties and tend to disrupt the bacterial cell walls by forming the pores in the cell membrane.85 Their cationic properties help attract them to the negative charges of lipopolysaccharide of Gram-negative bacteria and thus disrupt the membrane.90 Thus, these antibiotics, mainly polymyxin, are in use for the treatment of respiratory pathogens, such as Klebsiella pneumoniae and P. aeruginosa. An antibiotic namely d-cycloserine blocks the interconversion of an amino-acid alanine to d-alanyl-d-alanine in the bacterial cell wall synthesis.91
3.1.2. Protein-Synthesis Targeting Antibiotics
Some antibiotics can hinder the synthesis of ribosomal proteins.85 These distinct groups of antibiotics can target different ribosomal subunits, either the 50S or 30S. Notable examples of these antibiotics include aminoglycosides, chloramphenicol, erythromycin, macrolides, oxazolidinones, streptomycin, and tetracyclines. Antibiotics such as chloramphenicol, macrolides, and oxazolidinones target to the 50S subunit, while aminoglycosides and tetracyclines target to ribosomal 30S subunit.92 Similarly, kanamycin binds to the bacterial 30S ribosomal subunit, thereby preventing the translocation of peptidyl-tRNA from the A-site to P-site. This causes the misread of mRNA and inhibits protein synthesis. Similarly, macrolides such as erythromycin cause a blockade of the peptide exit tunnel of a ribosomal large subunit (50S) (Figure 4A(ii),B).93,94
Bacteria have now developed several new and unique strategies to cope with antibiotics that target the protein-synthesizing machinery of the pathogens. Streptomycin, an aminoglycoside antibiotic produced by the soil bacterium Streptomyces griseus,95 and the first drug used for the treatment of tuberculosis96 has the properties that can mutate a 30S ribosomal subunit in bacteria by causing a point mutation in the S12 protein of the small unit, eventually hampering the protein synthesis. Thus, streptomycin-resistant bacteria have now developed new strategies, whereby an alteration in 30S protein S12 causes the failure of drugs to bind to their ribosomal protein and bypass the effects of antibiotics. Recently identified aminoglycoside-acetylating genes aac(2′) and eis2 are known to confer resistance against streptomycin in Mycobacterium abscessus.96 Notable examples of this include studies in Bacillus subtilis, Mycobacterium tuberculosis, and Salmonella Typhimurium.97 Streptomycin resistant Mycobacterium tuberculosis has a mutation in the rpsL gene that encodes ribosomal protein S12.98 Some bacteria are also immune to spectinomycin antibiotics that can mutate protein S5 of the 30S subunit. Antibiotics, for instance, actinomycin, cause halting of the RNA elongation. Antibiotics such as chloramphenicol prevent the formation of the peptide bond to the growing chain, causing its premature termination. Similarly, some other antibiotics, such as tetracyclines, also prevent the addition of amino acid to the growing chain in the 50S subunit of the ribosome. However, aminoglycoside antibiotics function in a very different way, placing an incorrect amino acid to the growing peptide chain. The exit of the nascent peptide through an exit tunnel can be blocked by erythromycin, preventing the extension of the peptide chain. Also, some antibiotics such as puromycin and mupirocin act on tRNA and tRNA synthetase that stimulate the premature termination of chain and binding to the isoleucyl tRNA, respectively (Figure 4A,B).99
3.1.3. Nucleotide (DNA/RNA) Synthesis Hampers of the Bug
Enzymes such as DNA gyrases, also called topoisomerase II, precede before DNA helicase, and negatively unwound the supercoiled double-stranded DNA making way for the DNA helicase (Figure 4A(iii)). These enzymes introduce negative supercoiling to DNA to relieve torsional stress arising from the transcription and replication complexes,100,101 which subsequently cause an alteration in DNA topology and is responsible for the illegitimate recombination. These enzymes are ubiquitous in prokaryotes, while they are scarcely traced in eukaryotes. The differences in nucleotide sequences and similarity provide varying affinity for different molecules, proving them as an ideal candidate for antibiotics discovery.
Inhibitors of gyrase fall in two different classes, which are aminocoumarins (includes novobiocin) and quinolones (includes ciprofloxacin, pefloxacin, ofloxacin, and nalidixic acid). Gyrl (gyrases inhibitory proteins) is an inhibitor of DNA gyrases in E. coli.102 The aminocoumarins work through competitive inhibition by binding to the ATP-binding pocket of Gyrase B enzyme, which makes them appealing targets for developing broad-spectrum inhibitors. However, quinolones along with these enzymes prevent the replicating DNA from decatenating.103,104 Nevertheless, the bugs that are resistant to the binding of quinoline possess mutated topoisomerases.105
The binding of two subunits A (GyrA) and B (GyrB) of gyrase together to DNA causes ATP hydrolysis introducing the negative supercoils. Each subunit has further two subunits that assemble to form a complex of A2B2. The GyrA causes nicks in DNA, while the GyrB introduces negative supercoils, followed by resealing by GyrA. Remarkably, the ATP hydrolysis provides the energy necessary for the reaction and is catalyzed by the GyrB subunit. The quinoline binding to GyrA inhibits the cutting of DNA and its resealing function. Antibiotics such as nalidixic acid and oxalinic acid cause selective inhibition of GyrA, while GyrB is inhibited by coumermycin A and novobiocin. This inhibition in subunits has a negative role that prevents the blockage of super twisting activity thus avoiding the transcription. Moreover, the induction of heat-shock proteins such as LetD encoded by F-factor inhibits DNA gyrase activity,106 which proves that these enzymes are good candidates for developing both anticancer therapy and as antibacterial agents.107
3.1.4. Inhibitors of Folates Biosynthesis
In most organisms, folates provide special cofactors required for the biosynthesis of vital cellular components.108 Humans lack de novo synthesis of folates and are auxotrophic to folates. Thus, external supplement is needed for humans survival (either in the form of vitamin B9 or folic acid). Many microbes such as Bifidobacteria and plants rely on de novo folate biosynthesis. This, however, makes these microbes much more vulnerable to antibiotics. Interestingly, the presence of folate biosynthetic pathways in microbes but not in humans makes them ideal targets for the development of antimicrobial drugs.109 Available shreds of evidence reveal that the disruption of folate biosynthesis in prokaryotes makes them more vulnerable to antibiotics.110
Antibiotics, such as sulfonamides and trimethoprim cause impairment of folate metabolism and inhibit folate biosynthesis in microorganisms (Figure 4A (iv)). The para-aminobenzoic acid (PABA), an intermediate in folate biosynthesis, is generated from chorismate by the synergistic effects of 4-amino-4-deoxychorismate synthase (EC 6.2.1.85) and 4-amino-4-deoxychorismate lyase (EC 4.1.3.38).111 The PABA and pteridine form dihydropteroic acid and then to dihydrofolic acid (DHFA) by the activity of pteridine synthetase and dihydrofolate synthetase (EC 6.3.2.12), respectively. This DHFA formation process can be halted by sulfonamides. The subsequent conversion of DHFA to tetrahydrofolic acid (THFA) is carried out by an enzyme dihydrofolate reductase (EC 1.5.1.3), while this is arrested by tetramethoprim.112 Further conversion of THFA gives rise to thymidine, purines, and methionine. Thus, the use of these antibiotics sulfonamides and trimethoprim can largely control the biosynthesis of nucleotides and amino acids in microorganisms.
3.1.5. Inhibition of Mycolic Acid Biosynthesis
Mycolic acids are specific lipid key components of a mycobacterial cell wall,113 and these are formed from three covalently linked macromolecules (i.e., peptidoglycan, arabinogalactan, and mycolic acid) also called mycolyl–arabinogalactan–peptidoglycan complex (mAGP).114 The peptidoglycan, which is a polysaccharide, encapsulates the cytoplasmic membrane and is common in bacteria. Nevertheless, the peptidoglycan of Mycobacterium (causative agent of both tuberculosis and leprosy)113,114 differs from others in (i) N-glycosylation of muramic acid in Mycobacterium(115) and Nocardia, which in other bacteria is N-acetylated, and (ii) the cross-linking of peptides includes a proportion of bonds between two residues of meso-diaminopimelic acid in the peptidoglycan of Mycobacterium, while in other bacteria it is meso-DAP-d-alanine linkages.114
The mycolic acid helps bacterium to grow inside the macrophages so that they can bypass the host tissue physiology and host defense system.116,117 Three major types of mycolic acids, namely, alpha-, methoxy-, and keto- are synthesized by Mycobacterium tuberculosis. The biosynthesis of mycolic acids occurs through five distinct stages. First, the synthesis of C20–C26 straight carbon saturated fatty acids provides an alkyl branch, followed by the synthesis of meromycolic acid chain that offers a carbon skeleton and further modifications to introduce another functional group in the growing chain. This follows the Claisen-type condensation step with an ultimate reduction. The resulting scaffold is then acted upon by various mycolyltransferases which ultimately process cellular lipids.114,118
As M. tuberculosis mutants show increased resistance to antibiotics and macrophages,119 understanding the detailed molecular mechanism on the emergence of total drug resistance of these pathogens proves a major determinant in controlling these pathogens. Their resistivity against antibiotics can occur naturally, irrespective of their exposure to drugs such as rifampicin, isoniazid, isoxyl, thiacetazone, and streptomycin.120 The mechanisms of action of these drugs, however, differ from each other.120 The higher resistance of pathogens to these antibiotics is because of the monotherapy or low-concentration (below the MIC level), or improperly administered antibiotics whereby the persisters tend to evolve gradually and gain resistance to these antibiotics. Some drugs known to inhibit the biosynthesis of mycolic acid are ethionamide, isoniazid, isoxyl, thiolactomycin, and triclosan.114 Owing to the low permeability of the cell wall, mycobacteria show intrinsic resistance to these antibiotics, and these are the mycobacterial porin that limit the transverse of hydrophilic antibiotics and biocides across the membranes.121 Moreover, these tuberculosis pathogens negatively affect and cause down-regulation of critical innate effector immune responses of machrophages,122 for instance, the fusion of lysosome and phagosome, autophagy, antigen presentation and inhibition of reaction oxygen species. In addition, bacteria causing multidrug-resistant tuberculosis (MDR-TB) shows resistance against isoniazid and rifampin. The treatment of MDR-TB, however, can comprise the long use of fluoroquinolones and aminoglycosides,123 which are considered as the second line drugs.
The protein synthesis inhibitors in mycolate biosynthesis and the mycolic acids have crucial roles in the biosynthesis of the mycobacterial cell wall. All the key enzymes involved in the mycolic acid biosynthetic pathway, including fatty acid synthesis (KasA, KasB, MabA, InhA, HadABC), mycolic acid modifying enzyme such as SAM-dependent methyltransferases, fatty acid activating and condensing enzymes (FadD32, Acc, Pks13), transporters such as MmpL3 and transferases (Antigen 85A-C), are potential therapeutic targets.124
3.2. Multidrug Resistance in Bacteria
The growth of opportunistic pathogenic bacteria can be circumvented by elucidating their sensing and invading strategies. For this, hunting clinically relevant new chemical entities that endow the capacity to suppress the growth and spread of these bugs is of dire need. Many fungal endophytes, soil-dwelling, and marine actinomycetes are well-known primary and prolific sources of clinically relevant antibiotics and may help provide novel drugs with a unique mode of action against pathogens invasion.
Methicillin-resistant Staphylococcus aureus (MRSA) builds resistance not only to methicillin (which was initially developed to fight against penicillinase-producing S. aureus) but also to aminoglycosides, chloramphenicol, lincosamides, macrolides, and tetracycline.125,126 An antibiotic, namely vancomycin, was used for the treatment of MRSA infections; however, Enterococcus displays resistance to vancomycin, thus this species also gained the title of MRSA in 2002.127 Some untreatable variants are newly emerging that are observed to circumvent vancomycin effects in a hospital-acquired pathogen, namely Staphylococcus aureus. However, notoriously pathogenic bacteria resistant to vancomycin are Enterococcus faecium, and E. faecalis, which are β-lactam resistant and are known to be cured either with daptomycin, dalfopristin, or linezolid antibiotics.128
Microbes harbor several mechanisms to build resistance against multidrugs, such as the biochemical mechanism of resistance,129−131 receptors inactivation at the cell membrane,131,132 reduced membrane permeability,68,69,76,133−136 and the increased expression of genes that encode multidrug efflux pumps.125 These resistivity mechanisms can also occur via several mechanisms, such as accumulation of mutations, antimicrobial modifications (for instance, synthesis of β-lactamase enzymes), acquiring mobile genetic elements (transposons) through lateral gene transfer, cell-wall modification that includes LPS modification, reduced expression of cell-wall porins that prevent drugs influx, and overexpression of efflux pumps. Resistance, biosynthetic, and transporter encoding genes are often clustered together within the biosynthetic cluster, which provides a window of opportunity to sequester newly synthesized cargo extracellularly before a toxic threshold is reached. Some widely used resistance mechanisms endowed by pathogens are presented in the following sections:
3.2.1. Biochemical Mechanisms of Resistance
3.2.1.1. Mutational Alteration or Overproduction of the Target Protein
Macrolides and lincosamides make alterations in a ribosome, while β-lactams, quinolones, rifampin, etc. are responsible for enzymatic alterations in penicillin-binding proteins (high-affinity target sites of all β-lactam antibiotics). The overproduction of the target protein is done by sulfa, TMP, glycopeptides, etc. The penicillin-resistant mutants of Bacillus subtilis have an alteration in one of five penicillin-binding components, which allows them to resist the effect of penicillin.137 Thus, considering the worldwide explosion of antimicrobial resistance, drugs that possess the enhanced capability to tolerate the hydrolysis of β-lactamase are urgently needed.138
3.2.1.2. Enzymatic Inactivation of the Drug
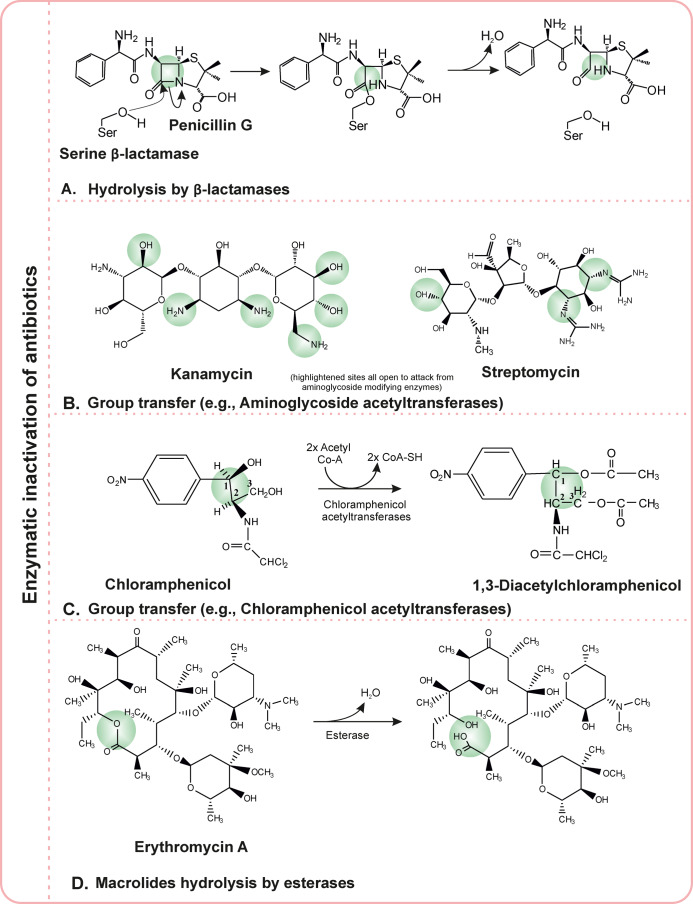
Pathogens generate enzymes (e.g., β-lactamase) that inactivate or neutralize the effect of antibiotics by several mechanisms (Figure 5A). Several processes for this mechanism include hydrolysis (β-lactam antibiotics), group transfer (modifications by acetyl transfer, phosphorylation, glycosylation, nucleotidylation, ribosylation and thiol transfer19,139), and the redox mechanisms.139
Figure 5.
Different microbial enzymes that inactivate the effect of antibiotics: (A) hydrolysis by B-lactamases; (B) group transfer (aminoglycoside acetyltransferases as an example); (C) group transfer (chloramphenicol acetyltransferases as an example); (D) macrolides hydrolysis by esterases (erythromycin A as an example).
Three distinct enzymatic inactivation mechanisms are observed in pathogens: (i) by β-lactamases (such as penicillinases) and cephalosporinases,140 (ii) by modification of aminoglycosides,141 and (iii) by esterase that hydrolyzes macrolides.142 The β-lactamases inactivate the β-lactam ring of penicillins,140 cephalosporins, and other related drugs making them nonfunctional. Some notable pathogens that can withstand the effect of β-lactams are Enterobacteriaceae, Haemophilus influenza, and S. aureus. The inactivation by modification of aminoglycosides141 or chloramphenicol by acetylation or phosphorylation by acetyltransferases and phosphotransferase; however, do not affect the pathogens.139,143 The acetyltransferase catalyzes the acylation of hydroxyl groups (Figure 5B,C). Pathogens such as S. aureus, Streptococcus, Pseudomonas, and Enterobacteriaceae are resistant against aminoglycosides and chloramphenicol by enzymatically acetylating these antibiotics. The inactivation done by esterase hydrolyzes macrolides142 causing an intracellular modification of the drugs making them ineffective.
Pathogens maintain a sufficient level of endogenous esterases144 that belong to a hydrolase class of enzymes and are involved in catalyzing the formation and breakdown of ester bonds through inter- and intramolecular transesterification reactions (Figure 5D). Mostly, these enzymes act on macrolides in vivo, causing the intracellular modifications and making them nonfunctional. However, esterases of Bacillus cereus (Bcr135 and Bcr136) do not have functional properties to inactivate the macrolides.145 The enzymes responsible for resistance to macrolides, rifamycins, β-lactams, and capreomycin are Erm41, Arr_Mab, BlaMab, and Eis2, respectively. Similarly, enzymes responsible for resistance to aminoglycosides, such as kanamycin B [AAC(2′) and Eis2] have also been identified.146−149
3.2.1.3. Acquisition of Resistant Genes
Next-generation sequencing has provided massive information on genome sequences in an unprecedented way. This approach allows us to rapidily map the horizontally acquired genes in different species in a fraction of time. Pathogens endow unparalleled tendencies of obtaining a significant amount of foreign DNA from the environment and incorporate them into their genome. This allows them to acquire resistance to different classes of compounds and causes replacement in the target molecules. For instance, Neisseria meningitides and Streptococcus pneumoniae have resistance to penicillin because of incorporating the production of mosaic proteins that were acquired horizontally.150 Similarly, genes (30–60 kb) encoding methicillin-resistant penicillin-binding proteins induced by methicillin and other β-lactams were acquired from other species in S. aureus.127,151 Moreover, Yersiniabactin gene cluster (ca. 40 kb) encoding the virulence factor lying in a high-pathogenicity island of Yersinia pestis was also transferred to E. coli.152
3.2.1.4. Bypassing the Target
Vancomycin, the last resort of antibiotics until a few years ago showed broad bioactivity to notorious pathogens by a distinct mechanism. This antibiotic binds to a substrate, the lipid-linked disaccharide pentapeptide, a precursor of cell-wall peptidoglycan, and makes pathogens unable to easily withstand the effects of these antibiotics. However, pathogens such as enterococci have now developed resistive mechanisms to circumvent the effect of these antibiotics. These pathogens are inhabitants of the human intestinal tract; however, lately, they have also developed mechanisms to curtail the effects of β-lactams, aminoglycosides, macrolides, and tetracycline.153 Vancomycin tends to bind to the d-Ala-d-Ala of the pentapeptide.154 However, the resistant mutant has a modified structure of this pentapeptide by an ester structure, d-Ala-D-lactic acid. This bypasses the identification and binding of vancomycin to this modified structure of the pentapeptide.155 Nevertheless, the resistance to these antibiotics requires the participation of resistance genes that need to be acquired from other organisms by the HGL method.125
Currently, daptomycin (a nonribosomal peptide isolated from Streptomyces roseosporus) is known to be the last resort antibiotics that binds to ribosomal protein S19 in humans.156 It has a unique mode of action against Gram-negative bacteria,157 the mechanism of which is however not fully elucidated. Besides, presumptions have been made that the underlying mechanism should be very unique and possibly involve a dependency on calcium.158 In 2003, the FDA approved the use of this drug for treatment of skin infections caused by Gram-positive bacteria, and in 2006 for treatment of bacteremia and endocarditis caused by Staphylococcus aureus, including MRSA.159
3.2.1.5. Resistance Mechanism of Biofilms against Antibiotics
Microorganisms tend to form an extracellular matrix called biofilms (matrix-enclosed bacterial population) that consist of a highly hydrated polymeric matrix surrounded by some interstitial voids and offer restricted diffusion of antibiotics. These biofilms are proven to be a key virulence factor for a wide range of microbial consortia160 and are highly resistant to antibiotics, antimicrobial agents, and also can efficiently overcome host innate immunity.161−163 An extracellular polymeric substance released by microbes helps them adhere to and form a matrix embedding the cells giving rise to a biofilm. A notable example of biofilm formation includes the colonization of the lungs’ surface of patients suffering from cystic fibrosis by P. aeruginosa. Also, Gram-positive bacteria such as S. epidermis, S. aureus, and enterococci form a resistant biofilm that is difficult to address by currently available antibiotic therapies, and these have a high-level of natural resistance to antimicrobial compounds.164−166
The anaerobic condition or nutrients depletion restricts bacterial growth whereby they enter into the stationary phase, and they are recalcitrant to growth-dependent antimicrobial killing. Two different hypotheses exist regarding the antibiotic tolerance by biofilms, namely, (i) slow or incomplete penetration of antibiotics,167−170 and (ii) changes in the chemical environment.171
Due to the high resistivity of biofilm infection, the “old” last-antibiotics such as colistin have been revisited.172−175 A cocktail of chemicals such as streptokinase and streptodornase inhibit biofilm growth of P. aeruginosa and S. aureus. Besides, enzymes such as oxidoreductases and polysaccharide–hydrolyzing enzymes are bactericidal in action and help remove biofilms.176 Similarly, alginate lyase causes diffusion of aminoglycosides and suppresses the growth of P. aeruginosa.177 A phytochemical, namely, allicin from garlic origin showed higher activity against the biofilm formation of C. albicans and S. epidermis.178 Some plant-derived biofilm inhibitors are ellagic acid, ginkgoneolic acid, hyperforin, tannic acid, (R)-norbgugaine, ginkgolic acid, and taxol.179 Plants such as garlic, gingko, Fritillaria delavayi, taxus, and some lichens (Cetraria nepalensis, Everniastrum nepalense, Parmelia milghenensis, P. minarum, Usnea longifolia) are enriched with antimicrobial compounds that have potential to suppress the formation of biofilms.180,181
Biofilms formed by an alginate-overproducing strain of Pseudomonas aeruginosa have a complex structural architecture that allows them to more resist tobramycin than biofilms formed by an isogenic nonmucoid strain. This suggests that an altered structure of biofilms cause increased resistance of pathogens against antibiotics.182 Quorum sensing molecules such as acyl-homoserine lactones induce biofilm formation by altering the patterns of gene expression such as in quorum sensing deficient mutants of P. aeruginosa.183 Also, the upregulation of a gene algC helps synthesize exopolysaccharide alginate in P. aeruginosa.184,185 Higher gene expression was observed for the attached cells during biofilm formation as seen in P. aeruginosa (algC, algD, algU, rpoS, and genes controlling polyphosphokinase synthesis)185 and Staphylococcus aureus.186
Interestingly, well-known bacterial resistivity mechanisms such as efflux pumps, modifying enzymes, and target mutations are not responsible for the formation and protection of the biofilms. Once the biofilm is formed, organisms can resist the toxicity of high doses of antibiotics compared to their earlier ones.187,188 For instance, the MIC of ampicillin in aqueous solution is 2 μg/mL for Klebsiella pneumoniae. However, the resistivity upsurges up to 5000 μg/mL ampicillin once the organism forms the biofilm.189
The formation of a bacterial biofilm can also take place by the subinhibitory concentrations of some antibiotics and contributes the bacterial persistence in chronic infections.190,191 Some notable examples of this biofilm formation include inducible resistance of P. aeruginosa and E. coli using subinhibitory concentrations (0.3 μg/mL; 0.3× the MIC) of aminoglycoside antibiotics such as tobramycin.192 The aminoglycoside response regulator (arr) encoding an inner membrane phosphodiesterase seems highly responsible for forming biofilm-specific aminoglycoside resistance in P. aeruginosa.193 The MIC of the bacteria is very less in the planktonic state compared to that of biofilms as is observed in P. aeruginosa.194 The MIC of Staphylococcus was found three folds higher (2048 μg/mL) in a biofilm state than in the planktonic state.195 The treatment of a biofilm also depends upon the time of its formation. For instance, a decline in the effect of vancomycin was observed when the biofilm aged from 6 h to 2 days.196
3.2.1.6. Preventing Drug Access to Targets
The functions of influx and efflux pumps depend on the type of transporters, species, and the substrates obtained. In bacteria, the influx of drugs occurs possibly by three different mechanisms, for instance, porin channels (OmpF of E. coli and OprF of P. aeruginosa), specific-protein channels (CarO of A. baumannii and OprD of P. aeruginosa for carbapenems), and a lipid bilayer region (asymmetric LPS). The diffusion of antibiotics takes place after its entry via the outer membrane to the inner membrane; however, they are prone to immediate extrusion by utilizing various efflux transporters. Interestingly, the impermeability of the drugs is enhanced by imipenem, while the pathogens active efflux pump is activated at once that immediately shuttles the substrates extracellularly.68,197,198
3.2.2. Multidrug Efflux Pumps and Their Regulation
These multidrug pumps are ubiquitous and are an important class of resistance determinants in pathogens that help to shuttle the substrates across the biological membrane.199 They recognize varying ranges of antibacterial substrates differing in both valency and structures and are mediated by integral membrane proteins. Antibiotics such as aminoglycosides, fluoroquinolones, tetracyclines, β-lactams, and macrolides are prone to immediate extracellular efflux.200−202
In Gram-positive bacteria, efflux pumps such as QacA, NorA (Bmr), and Smr pump out solute molecules across a single cytoplasmic membrane layer.68 The NorA (Bmr) pumps out basic dyes which are lipophilic cations and includes puromycin and fluoroquinolones. In addition, QacA and Smr both export quaternary ammonium compounds and basic dyes. In Gram-negative pathogens, the solutes need to trespass several layers en route to an inner periplasmic membrane.68 However, efflux pumps from the peptidoglycan or periplasm layer promptly channel substrates extracellularly that enter through outer-membrane. The overexpression of multidrug exporter pumps in Gram-negative bacteria causes resistance to different antibiotics, such as Smr (resistance to tetraphenylphosphonium and methylviologen) and EmrE (tetracycline, erythromycin and sulfadiazine).203
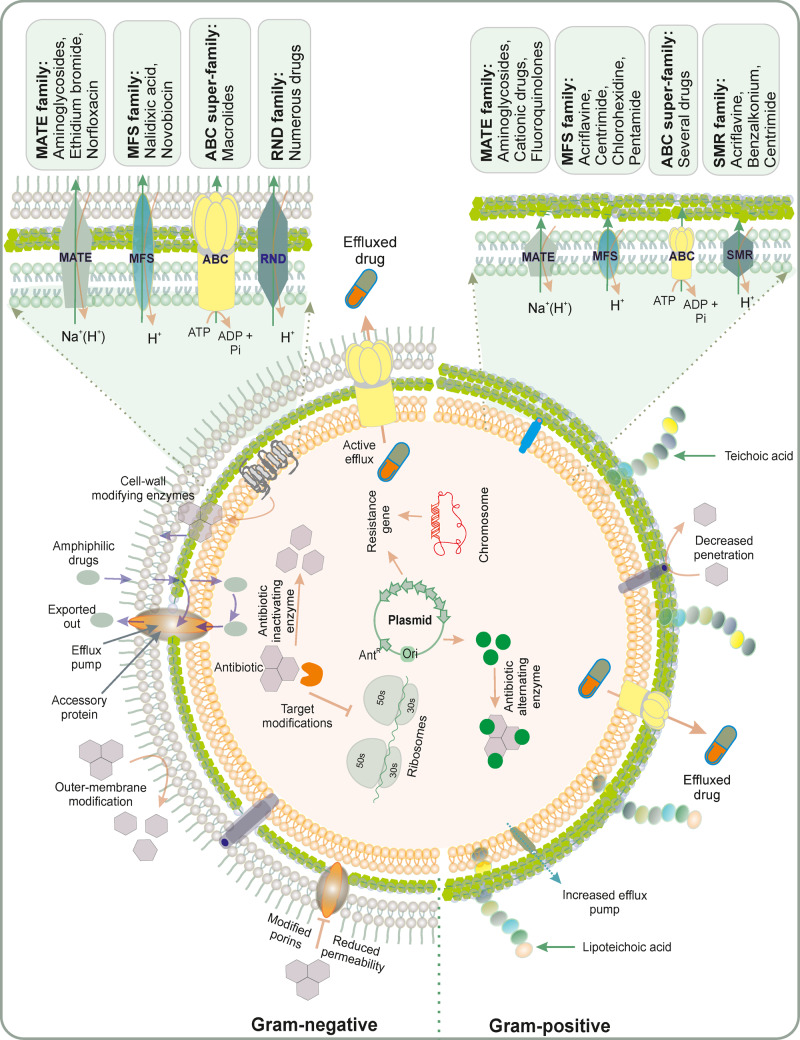
Bacteria endow different classes of efflux transporters and vary between different bacterial species. The efflux pump in Gram-negative bacteria needs to cross a double membrane system, such as the AcrAB system of E. coli and the MexAB-OprM system.204,205 Major classes of transporters in bacteria are primary transporters that include ABC (ATP-binding cassette) superfamilies, and secondary transporters that include MATE (multidrug and toxic compound extrusion), MFS (major facilitator superfamily), RND, and SMR (small multidrug resistance).206 Apart from ABC transporters, all other known transporters are secondary transporters as they are dependent on proton motive force. These transporters have different structural complexity and distinction can be made based on their domain organization and functions involved.206 The characterization of efflux systems into six major families has been performed based on sequence and functional similarities,207 as provided below (Figure 6).
Figure 6.
Family of different pumps and transporters in Gram-positive and Gram-negative bacteria.
3.2.2.1. ATP-Binding Cassette (ABC) Superfamily Transporters
ABC transporters are primary and ubiquitous membrane transport systems found in both prokaryotes and eukaryotes.206 ABC transporters are broadly grouped into two different types, importers (Type I and Type II) and exporters (10 different families based on membrane topology, viz., ABC(A-H), ABC-I (Caf16p-like), and YDR061W-like).206 These membrane-bound proteins possess two hydrophobic transmembrane domains and two cytoplasmic domains that recognize and confer resistance to antibiotics, vitamins, xenobiotics, amino acids, and sugars. The domain comprises two short peptide motifs, a Walker A (glycine-rich) and a Walker B motif (hydrophobic).206 The ABC signature is a consensus sequence and is unique in ABC domains. These transporters utilize the energy by the hydrolysis of ATP.206 The well-characterized members of this superfamily are MalEFGK, MDR, and CFTR.
Some notable examples of ABC-type transporters are MacAB in Neisseria gonorrheae, MsbA and MAcAB in E. coli, SmdAB in Serratia marcescens, and VcaM in Vibrio cholera.208 The SmrA in E. coli provides resistance to doxorubicin, fluoroquinolones, and tetracycline.209 MacABCsm provides intrinsic resistance to macrolides, aminoglycosides, and polymyxins plus tolerance to oxidative stress and biofilm formation. The ABC transporter such as CaCDR1 in C. albicans, a fungal pathogen, is overexpressed in fluconazole-resistant isolates. In E. coli, the MacA–MacB–TolC ABC types tripartite multidrug efflux pump actively extrudes substrate, including macrolide antibiotics and polypeptide virulence factors.210 The energy required for the transport is supplied by the ATPase MacB.210 This MacAB–TolC assembly may also transport heat-stable enterotoxin II, across the inner membrane to the periplasm by the Sec machinery, where it undergoes maturation.211 In S. aureus, ABC pumps recognize and shuttle tetracycline. Thus, understanding the structure–function relationships of these efflux pumps that underlie their transport mechanism and drug selectivity could enrich our nascent knowledge and help combat the challenges posed by notorious pathogens.
3.2.2.2. Multidrug and Toxic Compound Extrusion (MATE) Superfamily Transporters
MATE transporters are mainly Na+/drug antiporters,212 and they utilize the electrochemical potential of sodium ions as the driving force across the membranes, rather than the proton gradient used by the RND, MFS, and RND families.213 The expression of these efflux proteins in different species affords resistance toward a cocktail of drugs. For instance, overexpression of MepA from Staphylococcus aureus rendered them to build resistance to tigecycline (a glycylcycline antibiotic), which was originally considered to have antimicrobial activity toward MRSA and VRSA strains.214
NorM is the first MATE transporter (a Na+/drug antiporter) to be characterized by Vibrio parahemolyticus in 1998.215,216 This can recognize fluoroquinolones (such as norfloxacin) as transport substrates. Most MATE-family transporters are Na+-coupled transporters and transport Na+; however, some are H+/drug antiporters, such as PmpM (from Pseudomonas aeruginosa).213,217 Prokaryotic MATE transporters are classified into DinF and NorM, while eukaryotic MATE belongs to a single-family.212,218 Some notable examples of MATE transporters in microbes are AbeM (from Acinetobacter baumannii), BexA (from Bacteroides thetaiotaomicron), CdeA (from Clostridium difficile), ClbM (from E. coli),219 DinF (from Ralstonia solanacearum), HmrM (from Haemophilus influenza), Mdtk (from Salmonella enterica serovar Typhimurium), NorM (from Erwinia amylovora and Neisseria meningitides), NorMI (from Brucella melitensis), PmpM (from Pseudomonas aeruginosa), VcmA, VcrM, VcrB, VcrD, VcrH, and VcrN (from Vibrio cholera), MepA (from Staphylococcus aureus), VmrA (from Vibrio parahemolyticus), and YdhE (from E. coli; a homologue of NorM in Vibrio parahemolyticus). They have varying substrates, such as ciprofloxacin, norfloxacin, Na+, TPP+, EtBr, kanamycin, streptomycin, etc. Several numbers of MATE transporters have been traced in organisms such as Clostridium perfringens and Vibrio parahemolyticus (strain RIMD2210633); each harbor 11 and 9 numbers of MATE transporters, respectively.220
Identification of MATE transporters have been made in other domains of life. For instance, a single characterized Erc1MATE-type transporter of yeast Saccharomyces cerevisiae confers resistance to ethionine and has a functional role in the vesicular storage of S-adenosylmethionine.221 MATE proteins in mammals including humans are expressed predominantly in the liver and kidneys222 and are involved in the export of xenobiotics and cationic clinical drugs.223
3.2.2.3. Major Facilitator Superfamily (MFS) Transporters
MFS transporters are substrate-specific porters that are ubiquitous and catalyze the transport of sugars, drugs, and metabolic intermediates in organisms of all domains of life.224 These are the largest group of secondary transporters and show a high degree of structural conformation.225 The MDR transporters belonging to this MFS family have diverse substrate specificities and are most prevalent in microbial genomes.226 The structural architecture of these transporters comprises two domains, each having six helices, and the domains are linked by a long cytoplasmic loop.227−230 These MFS transporters form tripartite pumps with other partner proteins in Gram-negative pathogens, which direct the active shuttle of substrate extracellularly from the inner membrane and cytoplasm.231 The substrates coupling in these transporters involve alternate binding and releasing of the substrate.232 However, the conformational changes of the MFS proteins determine the transport mechanism.233 The mutations in local repressor and global regulatory genes, mutations in the promoter of definite transporters, and insertional elements inserted upstream of transporters encoding genes cause overexpression of these efflux pumps.234,235
These MFS encoding proteins include antiporters, symporters, and uniporters. The substrate recognition for this class of porters includes amino acids, cations, drugs, chromophores, enzyme cofactors, iron chelates, organic and inorganic acids, monosaccharides, nucleobases, nucleosides, nucleotides, oligosaccharides, peptides, and vitamins.233,236,237 The notable members of this class include Bmr protein in Bacillus subtilis,238 EmrB protein in E. coli,239 and QacA protein in Staphylococcus aureus.240,241 Some well-characterized members of this family include LacY, Glut1, and TetB. Moreover, AmpG of E. coli transports peptidoglycan degradation products and penicillin, Jen1 of Saccharomyces cerevisiae are lactate/pyruvate porters, NanT of E. coli transports sialic acid, NrtP is nitrate/nitrite porters of Prochlorococcus, QepA of E. coli clinical isolates efflux fluoroquinolone, and Qnr provides resistance against fluoroquinolone.242 EmrD, a close homologue with Bmr of Bacillus subtilis, LmrP of Lactococcus lactis, and NorA of Staphylococcus aureus is a proton-dependent multidrug transporter of E. coli and shuttles amphipathic compounds across the biological membrane.243 The crystal structures of EmrD243 and YajR244 revealed tripartite assemblies that span the bacterial cell envelope and provide some insights into this complex machinery. The CraA MFS pump in A. baumannii is a homologue of the MdfA pump in E. coli and confers resistance to chloramphenicol.245 The EmrB efflux pump of E. coli has, however, wider substrate recognition and confers resistance to a range of hydrophobic toxins (carbonyl cyanide m-chlorophenyl hydrazine).239 The EmrB along with EmrA (a membrane fusion protein) plus TolC (outer membrane protein) forms a tripartite system allowing substrates to trespass across the membranes246 (Figure 6).
3.2.2.4. Small Multidrug Resistance (SMR) Protein Family Transporters
This class of transporter is involved in the efflux of lipophilic cationic drugs in bacteria.247,248 Two distinct types of classes within this family are found, namely, small multidrug efflux systems and Sug proteins.249,250 The membrane topology of this family proteins reveals a polypeptide chain spanning the membrane four times (four α-helical transmembrane segments) and are the smallest drug efflux proteins.250 Typical and well-characterized members of this family include multidrug efflux proteins such as QacC, Smr, or Ebr from Staphylococcus sp.,249,251 and QacE from Klebsiella aerogenes.252 The multidrug Ebr efflux proteins for translocating lipophilic cations found in E. coli are also designated as MvrC and EmrE.207,253 These family proteins are found to be conserved in different prokaryotes,248 such as in E. coli. In this family of transporters, MDR pumps are found only in one of the two families.248 The EmrE of E. coli that functions as an inverted homodimer belongs to the SMR family proteins.254
3.2.2.5. Resistance-Nodulation-Cell-Division (RND) Family Transporters
These resistance-nodulation-cell-division (RND) family transporters are efflux pumps that translocate drugs and toxic cations extracellularly and are being driven by the proton motive force.255,256 They recognize the hydrophobicity of a substrate and are the ideal target candidates for developing the antibiotic adjuvants.257 Even though they are restricted in the inner membrane, they have intrinsic interaction with the periplasmic membrane (by the periplasmic adaptor protein) and outer membrane, in turn forming a tripartite complex. Notable examples of this complex in Gram-negatives can be viewed as AcrAB-TolC and MexAB-OprM in E. coli and P. aeruginosa, respectively.258,259 Gram-negative prokaryotes have this kind of pump.248
Inhibition of the drug efflux pump could also act as antivirulence drugs.257P. aeruginosa endows 12 efflux pumps of the RND family.234,260−266 The outer membrane component, namely TolC, works in conjunction with RND, ABC, and MFS efflux pumps,267 and the lack of any of these components causes them to be dysfunctional.268 The division of the RND family into two subfamilies—(i) hydrophobic and amphiphilic efflux RND (HAE-RND) and (ii) heavy metal efflux RND (HAE-RND) family—has been made.269 This classification is based on the recognition of their substrates and their translocation. Among the RND family, four RND efflux pumps such as MexAB-OprM, MexCD-OprJ, MexEF-OprN, and MexXY-OprM contribute the most significant and intrinsic MDR in P. aeruginosa.257,261,270 This MexB recognizes and transports various compounds such as aminoglycosides, cadmium, cerulenin, erythromycin, fluoroquinolones, macrolides, sulfonamides, tetracyclines, thiolactomycin, triclosan, trimethoprim, vanadium, and zinc.271 RND pumps also shuttle cobalt and nickel ions. Similarly, MexF and MexY also transport different compounds as substrates.272 Some other known RND members in P. aeruginosa are CzrAB-OpmN, MexGHI-OpmD, MexJK, and MexXY. The master regulators for these members are CzrRS (sensor for CzrAB-OpmN),273 MexL (represses MexJK),274 MexR (represses MexAB-OprM),275,276 MexZ (represses MexXY),277 and NfxB (represses MexCD-OprJ).278 These RND pumps are vital in pathogenic clinical bacterial strains effluxing out most antibiotics. Most of the work about these pumps has been concentrated only in E. coli and P. aeruginosa. Experiments on knocking-out some crucial efflux pumps, such as MexAB-OprM, MexEF-OprN, and MexGHI-OpmD have revealed an altered level of extracellular acylated homoserine lactones.279,280 An observance of different phenotypes for these mutated strains maybe because of quorum sensing.281
Moreover, in light of the antibiotics efflux by the pathogens, the way to circumvent this is by the development of chemical entities which the organisms’ efflux pump does not recognize, fails to shuttle out, or partially channels out.282 Together with this, an antibiotic that endows the capacity to inhibit the efflux pump can also work in most scenarios. However, pathogens have specially designed efflux pumps that get rid of the effects of the antibiotic, and efficiently translocate the antibacterial drugs out of the membrane. An efflux inhibitor in pathogens such as P. aeruginosa is phenylalanine–arginine β-naphthylamide (PAβN) that is active against MexAB-OprM, MexEF-OprN, MexCD-OprJ, and MexXY-OprM.283,284 The substrates of an RNP pump, namely minocycline and trimethoprim act as an efflux inhibitor of nitrocefin H33342 by AcrB in E. coli and S. enterica serovar Typhimurium.285−289 Even though RND multidrug efflux pumps of pathogen can shuttle a wide range of antimicrobial compounds, they do not tend to translocate the aminoglycosides.272
3.2.2.6. Proteobacterial Antimicrobial Compound Efflux (PACE) Family Transporters
This family of transporters has recently been characterized as a new class of resistance determinants in some Gram-negative human pathogens. They are known to efflux cationic biocides,266,290 and have the ability to confer resistance to a wide-range of biocides (such as antiseptics and disinfectants).291 It has been hypothesized that these transporters are likely to be powered by the H+ gradient.290 These transporters are found to be conserved in some γ-Proteobacterial species (Acinetobacter baumannii, Pseudomonas aeruginosa, and Vibrio parahemolyticus) and β-Proteobacterial species (Burkholderia cenocepacia) and Firmicutes (Veillonella parvula).290 They are thought to be acquired through lateral transfer from other species as the genome of most E. coli do not encode the PACE efflux proteins;199,290 however, some strains of E. coli encode one of the four PACE protein variants.290
3.2.3. Membrane Permeability
Many pathogens, for instance, P. aeruginosa possess intrinsic resistance to available natural products owing to their low permeability of an outer-membrane that works in conjunction with multidrug efflux transporters.292,293 The fate and efficacy of antibiotics are determined by their capacity to reach the target and have either bactericidal or bacteriostatic effects. For this, antibiotics need to trespass the barrier afforded by the cell membrane and access the intracellular targets of the pathogen. Moreover, the distinct membrane barrier for Gram-negatives and Gram-positives is considered as the major determinant for antimicrobial resistance in these pathogens,294 as could be exemplified by the low permeability of macrolides.295 Besides, Gram-negatives are also known to have acquired genes from other species that convey resistance to β-lactam antibiotics and cause low permeability of antibiotics to reach the target.296,297 Even sometimes, the permeability of the outer membrane works in conjunction with efflux pumps, to have enhanced resistance,298 which could be exemplified in biofilm-forming pathogens.299
3.2.4. Resistance Mutation
Pathogens, when subjected to lower doses of antibiotics (below MIC), evolve drastically300 as is observed in Salmonella enterica upon exposure to sub-MIC levels of streptomycin. The weak selection pressure may cause pathogens to gradually build higher tolerance capacity to elevated concentrations of antibiotics. However, the evolutionary mechanisms for the development of resistance against antibiotics remain enigmatic. Evidence suggests that the alteration of the rRNA target, reduction in aminoglycoside uptake, and induction of aminoglycoside-modifying enzyme AadA are three different mechanisms that allow pathogens to tolerate a high amount of antibiotics. Besides, disinfectants and heavy metals can also contribute to resistance evolution.301−303
Streptomycin in higher concentrations causes mutations in the gene rpsL that encodes ribosomal protein S12304,305 and confers resistance to an elevated level of this antibiotic (>1024 mg/L).306−308 Similarly, a mutation in the gene trkH (a potassium symporter) renders an organism to gain resistance against streptomycin, which before mutation was known to provide resistance against aminoglycoside in E. coli.309,310 Besides, mutation in the genes gidB (encoding a 16S rRNA methyltransferase), trkH (a potassium symporter), nuoG (encoding subunits of NADH dehydrogenase), cyoB (encoding bo3 ubiquinol terminal oxidase subunit 1), and znuA (encoding proteins involved in zinc uptake) generated high-level (>46 fold higher concentration) of streptomycin resistance.300 Moreover, mutations in the genes cyoB, nuoG, and trkH cause reduction in streptomycin uptake,311−314 and also cause impairment of the electron transport system.310,315 Thus, the bacterial population which has gained resistance mutation before exposure to antibiotics is unaffected by antibiotics even in higher doses, and the pathogens still tend to grow.
3.3. Honorable Suicide of Unfortunate Bacterial Pathogens
Organisms tend to alter their behavior and display negative interactions in detrimental conditions.316 The limiting factors for their growth are the depletion of resources, production of toxic byproducts, or the addition of antibiotics/natural products in the culture media. This behavior causes oscillatory dynamics that have a significant role in the ecology and evolution of microbes. “Quorum sensing” involves teamwork and constant chemical communication via which bacteria respond to the neighboring bacterial population. An exchange of signals occurs between these organisms in response to different stressors, such as availability of the nutrients or during infection, which allows the bacterial population to thrive in a varying environment. However, when the challenging condition persists, bacterium undergoes its mass destruction, which we refer to as ‘programmed cell death (PCD)/ apoptosis’. During this event, the bacterial cells put themselves to death.
Apoptosis in E. coli is caused by two different genes, mazE and MazF, which are located on the same operon. The gene mazF encodes a killer toxin, while mazE encodes antitoxin. The damage in mazE gene or the protein encoded by it is inhibited, whereby mazF unfolds leading the cells to die.317 Besides, various toxins secreted by bacteria in challenging conditions cause them to undergo suicide, while sparing uninfected neighbors. Bacteria such as E. coli undergo PCD when viruses infect them. This strategy causes the virus to get entrapped within bacterial cells, avoiding their escape to the external milieu, which in turn prevents their population from becoming severely damaged. However, the surviving noninfected bacterial population immediately self-destructs once the virus releases and the population is infected.
A similar strategy applies to the addition of antibiotics to the growing bacterial culture. The supplement of antibiotics causes bacteria to sequestrate these toxic compounds within their cell-envelope until the threshold is reached and subject themselves to die before the toxic compounds are expelled out to the environment. This prevents bacterial neighbors from being mutated with the toxic compound, eventually preventing their mass destruction. As for other strategies, bacteria such as Paenibacillus sp. modify their growth conditions to acidic pH which leads to their mass destruction.316 Such conditions lead to the exponential growth of these organisms followed by their exponential death. Moreover, a family of proteins that encode zeta toxins (PezT) causes cells of Streptococcus pneumoniae to undergo severe stress triggering them further to commit suicide.318,319 Thus, whatever the stresses may be, bacterial pathogens are always willing to devote themselves to the cost of their life to save their neighboring population.
4. Key Possibilities to Combat against the Rising Superbugs
In the scenario of rising superbugs, different sources for the origin of natural products have been identified. The plant origin and microbial origin need to be rigorously examined which may provide a new horizon in developing new strategies to combat against antibiotic resistance. Some key possibilities are discussed below.
4.1. Altering the Chemical Scaffolds of Old Antibiotics
Drug repurposing represents a manifestation of the shift from a single to multitarget paradigm in drug discovery. Changing the molecular scaffold of old antibiotics by the utilization of synthetic biology tools may make them no longer recognizable by efflux pumps of pathogens, and can prevent them from expelling out. This provides substantial benefit to pharma-industries because of the use of de-risked compounds, low developmental costs, utilization of compounds with known pharmacological traits, and short developmental plans. The repurposed drugs may find new targets and unveil the novel pathways that may be of substantial importance.33 Some highly valued repurposed drugs are auranofin, bithionol, celecoxib, cotrimazole, dapoxetine, duloxetine, ketoconazole, minoxidil, raloxifene, sildenafil, and thalidomide. However, altering the scaffold of the molecule may cause the antibiotic to be less effective to act against its intended target. For instance, human P-glycoprotein (P-gp; an ABC efflux transporter) recognizes the substrate taxol. Nonetheless, the chemical modification of taxol prevents the P-gp from recognizing the modified taxol, and thus it is allowed to cross the blood-brain barrier before being transported out.320
Three different approaches are being widely used in drug repurposing, such as, candidate identification, mechanistic assessment of the drug effect, and efficacy evaluation.33 These approaches fall under two major categories: (i) computational approaches and (ii) experimental approaches. The compound identification approach, which take almost two years, involves the search for the targets and novel insights for screening specialized metabolites and their platforms. The next step involves the acquisition of a compound that involves licensing of the compounds, which almost takes two more years.321 Once a novel compound has been discovered, it needs to pass through different phases (I, II, and III) before it is channeled to the clinical trials (to be used for human consumption). The overall drug development process takes at least 12 years to release to the market.321 Currently, some pharmaceutical companies are stepping up their efforts in developing a compound named ETX1317, which binds and inhibits the activity of β-lactamase. The inhibition of this enzyme will allow β-lactam antibiotics to work very efficiently, and as this compound needs to be injected intravenously, it will prove very useful in treating multidrug-resistant infections.
4.2. Probiotics Therapy
Probiotics alter the metabolic or immunological activity of the indigenous gut microbiota.322 They target the surface of epithelium by specific adhesion found mediated by specific organelles, for instance, fimbriae. The use of probiotics is an appealing therapy for curing different diseases, such as chronic diarrhea, allergies, and eczema.323 Some notable examples of gut probiotics are Lactobacillus rhamnosus, L. plantarum, L. reuteri, and L. agilis,324 bifidobacteria, propionibacteria, and enterococci. The first two of these examples are being perceived as “protective” intestinal bacteria325 because of their ability to tolerate acidic conditions rather than other bacteria.326
The human colon comprises 1 × 1011 microorganisms per gram. The gut microflora produces essential mucosal nutrients, such as short-chain fatty acids, arginine, cysteine, and glutamine.325 These probiotics also secrete antimicrobial substances, overgrow the pathogens, and stimulate the intestinal immune defense system. Many gastrointestinal dysfunctions are based on the disturbances or imbalances of intestinal microflora. The gut microbiota suffer from serious threats, whereby a tough competition exists for binding to the walls through the adhesion receptors and colonization of the gut. The gut mucosal barriers are the first line in host defense, and probiotics colonize the mucus in a very efficient manner to prevent it from infection. For instance, L. plantarum endows mannose-specific adhesion327 similar to E. coli, Klebsiella, and Shigella, making them appealing species for further clinical research.
Pathogens provide severe clinical complications associated with a significant economic burden. One promising way for treating them is an engraftment of microbiota from a healthy donor into a recipient to restore the normal gut microbial community structure.322 A commonly used microbial source for the transplantation is fecal microbiota from infants’ colons, dominated by bifidobacteria and staphylococci, suggesting that the diversity of organisms depends on the regular food intake. This alteration of gastrointestinal microflora is the most effective, inexpensive, and easy technique and has been widely used for treating recalcitrant Clostridium difficile infection in clinical therapies.322,328 In addition, microbiota-related dysfunction can be cured by this process.322 The common route of administration for these strains is en route nasogastric/nasojejunal tube, retention enemas, and colonoscopy. Probiotics also stimulate host immunity by balancing the intestinal microflora. An oral administration of fecal enemas from healthy adults can cure Pseudomembranous colitis that is caused by the pathogenic strain of C. difficile.328−331 Interestingly, a nonpathogenic strain of C. difficile can occupy the gut surface that prevents pathogenic strains from infection. The oral administration of probiotics has several functions, such as immunomodulation of local and systemic immune response, lowering of serum cholesterol and blood pressure, reduction in the occurrence of bladder cancer, shortening rotavirus diarrhea, and modulation of endogenous microflora.332−334 The use of probiotics may be a good strategy in the area of gut infections; however, it may not be very useful for the treatment of cellulitis or bacteremia.
Some organisms designated by the FDA as GRAS (Generally Recognized as Safe) have found applications to certain probiotics when they are supplemented with food. Species such as Aspergillus oryzae, Bifidobacterium lactis, Carnobacterium maltaromaticum, Fusarium venenatum, Oenococcus oeni, Saccharomyces cerevisiae, Schizosaccharomyces pombe, and Streptococcus thermophiles are some GRAS species recognized by the FDA.
4.3. Overexpression of Efflux Pump Inhibitors
Multidrug resistance efflux pumps play a vital role in antibiotics resistance in both Gram-negative and Gram-positive bacteria.260 Currently, the failure in targeting the use of antibiotics toward preventing the spread of infectious diseases have been observed because of the overexpression of active efflux pumps nestled in the bacterial cell wall and have the ability to shuttle out cytotoxic drugs (either old or the new drugs), or other unwanted molecules. Thus, the efflux pumps remain a major determinant for knowing the efficacy of metabolites with different scaffolds, and a promising strategy in combating against pathogenic organisms.335 Some well-characterized efflux pump inhibitors (EPI) are peptidomimetics and pyridopyrimidines traced in several pathogens.336−338
The EPI first to be characterized was phenyl-arginine β-naphthylamide (PAβN) with its role to inhibit the expression of efflux systems of P. aeruginosa, that is, MexAB-OprM, MexCD-OprJ, MexEF-OprN, and MexXY-OprM.339 This inhibitor also represses the expression of different pumps in different organisms as well, for instance, AcrAB-TolC of E. coli, K. pneumoniae, and S.Typhimurium.339 Recently, four different small molecules that inhibit the expression of efflux pumps in E. coli and interact with AcrAB-TolC efflux pump component AcrA were identified.340 These small molecules namely chlorobiocin and SLU-258 are likely to be involved in binding to a site between the lipoyl and β-barrel domains of AcrA, which affects the expression of efflux pump.340 In addition, thiophene derivatives are known to promote the inhibition of the NorA efflux pump, making the antibiotic ciprofloxacin active against the S. aureus SA-119B strain that constitutively overexpressed the NorA efflux pump.260,341,342 Another best characterized EPI for Gram-positive bacteria is NMP (1-(1-naphthylmethyl)-piperazine).343,344 However, the toxicity of EPIs to human cells at higher concentrations has proven to be a downside and has remained a controversy in clinical development.344 Some other compounds that act as EPIs are globomycin, carbonyl cyanide m-chlorophenylhydrazone, various quinolines, and aryl piperazine derivatives.345 Some of the established strategies which prevent the efflux of drug from the cells are the disruption of pump assembly, altered drug structure that are no longer identified by EPs, collapse of efflux energy (proton motive force), and application of competitive/noncompetitive inhibitor with drug and blocking inner/outer membrane proteins.346−348 Some synthetic EPIs such as carbonylcyanide m-chlorophenylhydrazone, chlorpromazine, and PAβN have been known to prevent biofilm formation in E. coli, P. aeruginosa, and S. aureus.349
4.4. Essential Oils in Conjunction with Antibiotics to Treat Recalcitrant Bacterial Pathogens
Essential oils (EOs) are concentrated volatile and aromatic natural products of plant origin (stored by plants in secretory cells, cavities, glandular trichomes, or epidermic cells). These secondary metabolites (SMs) are a cocktail of primary terpenoids (monoterpenes and sesquiterpenes) along with other various molecules such as acids, alcohols, aldehydes, aliphatic hydrocarbons, acyclic esters, or lactones. They also comprise carvacrol, cinnamaldehyde, cinnamic acid, eugenol, and thymol. These are known to have some synergistic effects in combination with antibiotics,350 and have enormous clinical implementations.
Owing to the hydrophobicity of EOs, a thick peptidoglycan layer of Gram-positive bacteria facilitates free passage of these molecules through their cellular envelope. The Gram-negatives though are recalcitrant to these molecules; however, the passage is not checked completely, allowing some to filter in through the porins.351 In micro-organisms, EOs can cause degradation of the cell wall,352 cytoplasmic coagulation,353 damage to membrane proteins,354 puncturing of the cell-wall causing leakage of cellular contents,355 damage to cytoplasmic membrane,356 and decrease in inner ATP pool by the reduction of ATP synthesis.356 They reduce membrane potential and proton motive force.357 The loss in microbial viability may be caused by the destabilization of the phospholipid layer, malfunction, and permeability of the plasma membrane (by destroying the electron transport system),358 and inactivation of the enzymatic reactions and cellular proteins.359
Currently, more than 3000 of EOs are known; however, only <10% of those are in daily use, and mostly as fragrances. Some widely studied EOs are oregano (from Origanum vulgare) and thyme (from Thymus vulgaris) that comprises mostly carvacrol, thymol, g-terpinene, r-cymene,360,361 and Cinnamic acid and trans-cinnamaldehyde from Cinnamomum zeylanicum.362 Given the increasing clinical resistance, myriads of synergistic alternatives have been developed as a replacement to antibiotics, or as a development of adjuvants.363 This includes combining nonantibiotic drugs with antibiotics364 or combining environmental antimicrobials with antibiotics.363 In addition, EOs in conjunction with antibiotics (basically β-lactam) and nanoparticles365 help reduce the emergence of resistant variants and suppress the growth of plasmid-conferred multidrug-resistant pathogens.366,367 Similarly, amoxicillin and oxacillin proved effective against MRSA and Acinetobacter baumannii when used in conjunction with EOs of Pistacia lentiscus, Pituranthos chloranthus, and Teucruim ramosissimum.367 Besides, the synergistic effect of eugenol with ampicillin revealed bioactivity against Streptococcus criceti and S. gordonii, while with gentamicin showed bioactivity against S. sanguinis and Porphyromonas gingivalis.368 Moreover, Carvacrol showed a synergistic effect along with ampicillin and nitrofurantoin by inhibiting the growth of Klebsiella oxytoca.369 EOs from Momordica charantia,370Punica granatum,371 and Rosmarinus officinalis(372) inhibit the MDR efflux pump of MRSA; however, some notorious pathogens have intrinsic resistance to these EOs, as revealed by Pseudomonas spp. (overexpression of MexAB-oprM efflux pump373). Thus, use of EOs in conjunction with other antimicrobials could help eradicate the burgeoning clinical resistivity to some extent.
5. Concluding Thoughts and Future Perspectives
In the current scenario, in which an increased resistance of superbugs against the cocktail of antibiotics prevails along with recurring and emerging new infectious diseases, the search for novel “hit molecules” that govern the power to suppress the spread of bacterial pathogens is urgently needed. Discovering new antibacterial leads from novel validated targets may help us to combat the existing challenges of antimicrobial resistance. Even though tremendous efforts are being injected in discovering new bioactive molecules, high throughput strategies still need to be developed to trace new bioactive entities, which can act as a magic bullet against obnoxious superbugs. As such, the recent identification of a compound namely teixobactin from Eleftheria terrae, an uncultivable or poorly cultivable microbe, and darobactin which is active against Gram-negative pathogens have equally instigated researchers’ attention. Nevertheless, some very poorly studied places or virgin habitat for research, such as marine species or the Himalayan soil-dwelling microbes and the symbiotic organisms (symbionts from the guts of termites, spiders, or other arthropods) are some of the enticing possibilities for discovering novel metabolites-synthesizing strains. In addition, anticipating the next move of pathogens beforehand by predicting their possible evolution and identifying their actual targets may control the spread of these bugs. Moreover, performing genomics zoom-in and trying to activate the cryptic, silent, or orphan biosynthetic pathways and the use of recently discovered strategies in finding new chemical entities might unveil some successful stories about new chemical scaffolds. Although gaps remain in our knowledge to find new “hit molecules”, new strategies injected in this field are likely to discover ideal candidates with enhanced bioactivity in the next few years.
The authors declare no competing financial interest.
References
- WHO . (2014) Antimicrobial resistance: global report on surveillance, World Health Organization, France. [Google Scholar]
- Walsh C. T.; Fisher S. L.; Park I. S.; Prahalad M.; Wu Z. (1996) Bacterial resistance to vancomycin: Five genes and one missing hydrogen bond tell the story. Chem. Biol. 3, 21–28. 10.1016/S1074-5521(96)90079-4. [DOI] [PubMed] [Google Scholar]
- Moir D. T.; Di M.; Opperman T.; Schweizer H. P.; Bowlin T. L. (2007) A high-throughput, homogeneous, bioluminescent assay for Pseudomonas aeruginosa gyrase inhibitors and other DNA-damaging agents. J. Biomol. Screening 12, 855–864. 10.1177/1087057107304729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rice L. B. B. (2008) Federal funding for the study of antimicrobial resistance in nosocomial pathogens: No ESKAPE. J. Infect. Dis. 197, 1079–1081. 10.1086/533452. [DOI] [PubMed] [Google Scholar]
- de la Cruz F.; Davies J. (2000) Horizontal gene transfer and the origin of species: lessons from bacteria. Trends Microbiol. 8, 128–133. 10.1016/S0966-842X(00)01703-0. [DOI] [PubMed] [Google Scholar]
- Chen I.; Dubnau D. (2004) DNA uptake during bacterial transformation. Nat. Rev. Microbiol. 2, 241–249. 10.1038/nrmicro844. [DOI] [PubMed] [Google Scholar]
- Jain R.; Rivera M. C.; Lake J. A. (1999) Horizontal gene transfer among genomes: the complexity hypothesis. Proc. Natl. Acad. Sci. U. S. A. 96, 3801–3806. 10.1073/pnas.96.7.3801. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Werner G.; Strommenger B.; Witte W. (2008) Acquired vancomycin resistance in clinically relevant pathogens. Future Microbiol. 3, 547–562. 10.2217/17460913.3.5.547. [DOI] [PubMed] [Google Scholar]
- Pogliano J.; Pogliano N.; Silverman J. A. (2012) Daptomycin-mediated reorganization of membrane architecture causes mislocalization of essential cell division proteins. J. Bacteriol. 194, 4494–4504. 10.1128/JB.00011-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whitman W. B.; Coleman D. C.; Wiebe W. J. (1998) Prokaryotes: The unseen majority. Proc. Natl. Acad. Sci. U. S. A. 95, 6578–6583. 10.1073/pnas.95.12.6578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kieser T, Bibb M. J., Buttner M. J., and Chater K. F. (2000) Practical Streptomyces Genetics, 1st ed., The John Innes Foundation, Norwich. [Google Scholar]
- de Lima Procópio R. E.; da Silva I. R.; Martins M. K.; de Azevedo J. L.; de Araújo J. M. (2012) Antibiotics produced by Streptomyces. Braz. J. Infect. Dis. 16, 466–471. 10.1016/j.bjid.2012.08.014. [DOI] [PubMed] [Google Scholar]
- Watve M. G.; Tickoo R.; Jog M. M.; Bhole B. D. (2001) How many antibiotics are produced by the genus Streptomyces?. Arch. Microbiol. 176, 386–390. 10.1007/s002030100345. [DOI] [PubMed] [Google Scholar]
- Watve M.; Shejval V.; Sonawane C.; Rahalkar M. (2000) The ’ K ’ selected oligophilic bacteria: A key to uncultured diversity?. Curr. Sci. 78, 1535–1542. [Google Scholar]
- Ward D. M.; Weller R.; Bateson M. M. (1990) 16S rRNA sequences reveal numerous uncultured microorganisms in a natural community. Nature 345, 63–65. 10.1038/345063a0. [DOI] [PubMed] [Google Scholar]
- Benveniste R.; Davies J. (1973) Aminoglycoside antibiotic-inactivating enzymes in actinomycetes similar to those present in clinical isolates of antibiotic-resistant bacteria. Proc. Natl. Acad. Sci. U. S. A. 70, 2276–2280. 10.1073/pnas.70.8.2276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lomovskaya N.; Otten S. L.; Doi-Katayama Y.; Fonstein L.; Liu X. C.; Takatsu T.; Inventi-Solari A.; Filippini S.; Torti F.; Colombo A. L.; Hutchinson C. R. (1999) Doxorubicin overproduction in Streptomyces peucetius: Cloning and characterization of the dnrU ketoreductase and dnrV genes and the doxA cytochrome P-450 hydroxylase gene. J. Bacteriol. 181, 305–318. 10.1128/JB.181.1.305-318.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Otten S. L.; Ferguson J.; Hutchinson C. R. (1995) Regulation of daunorubicin production in Streptomyces peucetius by the dnrR2 locus. J. Bacteriol. 177, 1216–1224. 10.1128/JB.177.5.1216-1224.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Padmini N.; Ajilda A. A. K.; Sivakumar N.; Selvakumar G. (2017) Extended spectrum β-lactamase producing Escherichia coli and Klebsiella pneumoniae: critical tools for antibiotic resistance pattern. J. Basic Microbiol. 57, 460–470. 10.1002/jobm.201700008. [DOI] [PubMed] [Google Scholar]
- Ling L. L.; Schneider T.; Peoples A. J.; Spoering A. L.; Engels I.; Conlon B. P.; Mueller A.; Hughes D. E.; Epstein S.; Jones M.; Lazarides L.; Steadman V. A.; Cohen D. R.; Felix C. R.; Fetterman K. A.; Millett W. P.; Nitti A. G.; Zullo A. M.; Chen C.; Lewis K. (2015) A new antibiotic kills pathogens without detectable resistance. Nature 517, 455–459. 10.1038/nature14098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zong Y.; Fang F.; Meyer K. J.; Wang L.; Ni Z.; Gao H.; Lewis K.; Zhang J.; Rao Y. (2019) Gram-scale total synthesis of teixobactin promoting binding mode study and discovery of more potent antibiotics. Nat. Commun. 10, 1–11. 10.1038/s41467-019-11211-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baral B.; Akhgari A.; Metsä-Ketelä M. (2018) Activation of microbial secondary metabolic pathways: Avenues and challenges. Synth. Syst. Biotechnol. 3, 163–178. 10.1016/j.synbio.2018.09.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blaskovich M. A. T.; Zuegg J.; Elliott A. G.; Cooper M. A. (2015) Helping chemists discover new antibiotics. ACS Infect. Dis. 1, 285–287. 10.1021/acsinfecdis.5b00044. [DOI] [PubMed] [Google Scholar]
- Möhler J. S.; Sim W.; Blaskovich M. A. T.; Cooper M. A.; Ziora Z. M. (2018) Silver bullets: A new lustre on an old antimicrobial agent. Biotechnol. Adv. 36, 1391–1411. 10.1016/j.biotechadv.2018.05.004. [DOI] [PubMed] [Google Scholar]
- Yin L. M.; Edwards M. A.; Li J.; Yip C. M.; Deber C. M. (2012) Roles of hydrophobicity and charge distribution of cationic antimicrobial peptides in peptide-membrane interactions. J. Biol. Chem. 287, 7738–7745. 10.1074/jbc.M111.303602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li J.; Koh J. J.; Liu S.; Lakshminarayanan R.; Verma C. S.; Beuerman R. W. (2017) Membrane active antimicrobial peptides: Translating mechanistic insights to design. Front. Neurosci. 11, 73. 10.3389/fnins.2017.00073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Newman D. J.; Cragg G. M. (2016) Natural products as sources of new drugs from 1981 to 2014. J. Nat. Prod. 79, 629–661. 10.1021/acs.jnatprod.5b01055. [DOI] [PubMed] [Google Scholar]
- Besse A.; Peduzzi J.; Rebuffat S.; Carré-Mlouka A. (2015) Antimicrobial peptides and proteins in the face of extremes: Lessons from archaeocins. Biochimie 118, 344–355. 10.1016/j.biochi.2015.06.004. [DOI] [PubMed] [Google Scholar]
- Kumar V.; Tiwari S. K. (2017) Halocin HA1: An archaeocin produced by the haloarchaeon Haloferax larsenii HA1. Process Biochem. 61, 202–208. 10.1016/j.procbio.2017.06.010. [DOI] [Google Scholar]
- Shand R. F., and Leyva K. J. (2007) Peptide and Protein Antibiotics from the Domain Archaea: Halocins and Sulfolobicins, in Bacteriocins, pp 93–109, Springer Berlin Heidelberg, Berlin, Heidelberg. [Google Scholar]
- Van Hoek A. H. A. M.; Mevius D.; Guerra B.; Mullany P.; Roberts A. P.; Aarts H. J. M. (2011) Acquired antibiotic resistance genes: An overview. Front. Microbiol. 2, 1–27. 10.3389/fmicb.2011.00203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adzitey F. (2016) Antibiotic classes and antibiotic susceptibility of bacterial isolates from selected Poultry; A mini review. Worlds Vet. J. 6, 36–41. 10.5455/wvj.20150853. [DOI] [Google Scholar]
- Pushpakom S.; Iorio F.; Eyers P. A.; Escott K. J.; Hopper S.; Wells A.; Doig A.; Guilliams T.; Latimer J.; McNamee C.; Norris A.; Sanseau P.; Cavalla D.; Pirmohamed M. (2019) Drug repurposing: progress, challenges and recommendations. Nat. Rev. Drug Discovery 18, 41–58. 10.1038/nrd.2018.168. [DOI] [PubMed] [Google Scholar]
- Butler M. S. (2004) The role of natural product chemistry in drug discovery. J. Nat. Prod. 67, 2141–2153. 10.1021/np040106y. [DOI] [PubMed] [Google Scholar]
- Ashburn T. T.; Thor K. B. (2004) Drug repositioning: Identifying and developing new uses for existing drugs. Nat. Rev. Drug Discovery 3, 673–683. 10.1038/nrd1468. [DOI] [PubMed] [Google Scholar]
- Dehmer S. P., Maciosek M. V., and Flottemesch T. (2015) Aspirin use to prevent cardiovascular disease and cancer: A decision analysis, Agency for Healthcare Research and Quality, Rockville, MD. [PubMed] [Google Scholar]
- Cavalla D.; Singal C. (2012) Retrospective clinical analysis for drug rescue: For new indications or stratified patient groups. Drug Discovery Today 17, 104–109. 10.1016/j.drudis.2011.09.019. [DOI] [PubMed] [Google Scholar]
- Pucci M. J.; Bush K. (2013) Investigational antimicrobial agents of 2013. Clin. Microbiol. Rev. 26, 792–821. 10.1128/CMR.00033-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Butler M. S.; Blaskovich M. A.; Cooper M. A. (2017) Antibiotics in the clinical pipeline at the end of 2015. J. Antibiot. 70, 3–24. 10.1038/ja.2016.72. [DOI] [PubMed] [Google Scholar]
- Ilić K.; Jakovljević E.; Škodrić-Trifunović V. (2012) Social-economic factors and irrational antibiotic use as reasons for antibiotic resistance of bacteria causing common childhood infections in primary healthcare. Eur. J. Pediatr. 171, 767–777. 10.1007/s00431-011-1592-5. [DOI] [PubMed] [Google Scholar]
- Mboya E. A.; Sanga L. A.; Ngocho J. S. (2018) Irrational use of antibiotics in the Moshi Municipality Northern Tanzania: a cross sectional study. Pan Afr. Med. J. 31, 165. 10.11604/pamj.2018.31.165.15991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ross D. B. (2007) The FDA and the Case of Ketek. N. Engl. J. Med. 356, 1601–1604. 10.1056/NEJMp078032. [DOI] [PubMed] [Google Scholar]
- USFDA (2018) FDA updates warnings for fluoroquinolone antibiotics on risks of mental health and low blood sugar adverse reactions, U.S. Food Drug Admin..
- Aiello A. E.; Larson E. L.; Levy S. B. (2007) Consumer antibacterial soaps: effective or just risky?. Clin. Infect. Dis. 45, S137–S147. 10.1086/519255. [DOI] [PubMed] [Google Scholar]
- (1971) FDA warns on hexachlorophene bath. New York Times Dec, 49. [Google Scholar]
- Gill E. E.; Franco O. L.; Hancock R. E. W. (2015) Antibiotic adjuvants: Diverse strategies for controlling drug-resistant pathogens. Chem. Biol. Drug Des. 85, 56–78. 10.1111/cbdd.12478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Möllmann U.; Heinisch L.; Bauernfeind A.; Köhler T.; Ankel-Fuchs D. (2009) Siderophores as drug delivery agents: Application of the “trojan Horse” strategy. BioMetals 22, 615–624. 10.1007/s10534-009-9219-2. [DOI] [PubMed] [Google Scholar]
- Górska A.; Sloderbach A.; Marszałł M. P. (2014) Siderophore-drug complexes: potential medicinal applications of the ‘Trojan horse’ strategy. Trends Pharmacol. Sci. 35, 442–449. 10.1016/j.tips.2014.06.007. [DOI] [PubMed] [Google Scholar]
- Mislin G. L. A.; Schalk I. J. (2014) Siderophore-dependent iron uptake systems as gates for antibiotic Trojan horse strategies against Pseudomonas aeruginosa. Metallomics 6, 408–420. 10.1039/C3MT00359K. [DOI] [PubMed] [Google Scholar]
- Schalk I. J. (2018) A trojan-horse strategy including a bacterial suicide action for the efficient use of a specific Gram-positive antibiotic on Gram-negative bacteria. J. Med. Chem. 61, 3842–3844. 10.1021/acs.jmedchem.8b00522. [DOI] [PubMed] [Google Scholar]
- Tobert J. A. (2003) Lovastatin and beyond: The history of the HMG-CoA reductase inhibitors. Nat. Rev. Drug Discovery 2, 517–526. 10.1038/nrd1112. [DOI] [PubMed] [Google Scholar]
- Wohlleben W.; Mast Y.; Stegmann E.; Ziemert N. (2016) Antibiotic drug discovery. Microb. Biotechnol. 9, 541–548. 10.1111/1751-7915.12388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Silhavy T. J.; Kahne D.; Walker S. (2010) The bacterial cell envelope. Cold Spring Harbor Perspect. Biol. 2, 1–16. 10.1101/cshperspect.a000414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tenover F. C.; Lancaster M. V.; Hill B. C.; Steward C. D.; Stocker S. A.; Hancock G. A.; O’Hara C. M.; Clark N. C.; Hiramatsu K. (1998) Characterization of Staphylococci with reduced susceptibilities to vancomycin and other glycopeptides. J. Clin. Microbiol. 36, 1020–1027. 10.1128/JCM.36.4.1020-1027.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liñares J. (2001) The VISA/GISA problem: therapeutic implications. Clin. Microbiol. Infect. 7 (Suppl 4), 8–15. 10.1046/j.1469-0691.2001.00054.x. [DOI] [PubMed] [Google Scholar]
- Rybak M. J.; Akins R. L. (2001) Emergence of methicillin-resistant Staphylococcus aureus with intermediate glycopeptide resistance: Clinical significance and treatment options. Drugs 61, 1–7. 10.2165/00003495-200161010-00001. [DOI] [PubMed] [Google Scholar]
- Péchère J. C. (1999) Current and future management of infections due to methicillin-resistant staphylococci infections: the role of quinupristin/dalfopristin. J. Antimicrob. Chemother. 44 (Suppl A), 11–8. 10.1093/jac/44.suppl_1.11. [DOI] [PubMed] [Google Scholar]
- Enright M. C.; Robinson D. A.; Randle G.; Feil E. J.; Grundmann H.; Spratt B. G. (2002) The evolutionary history of methicillin-resistant Staphylococcus aureus (MRSA). Proc. Natl. Acad. Sci. U. S. A. 99, 7687–7692. 10.1073/pnas.122108599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alves D.; Olívia Pereira M. (2014) Mini-review: Antimicrobial peptides and enzymes as promising candidates to functionalize biomaterial surfaces. Biofouling 30, 483–499. 10.1080/08927014.2014.889120. [DOI] [PubMed] [Google Scholar]
- Fjell C. D.; Hiss J. A.; Hancock R. E. W.; Schneider G. (2012) Designing antimicrobial peptides: Form follows function. Nat. Rev. Drug Discovery 11, 37–51. 10.1038/nrd3591. [DOI] [PubMed] [Google Scholar]
- Epand R. M.; Vogel H. J. (1999) Diversity of antimicrobial peptides and their mechanisms of action. Biochim. Biophys. Acta, Biomembr. 1462, 11–28. 10.1016/S0005-2736(99)00198-4. [DOI] [PubMed] [Google Scholar]
- Silver L. L. (2016) A Gestalt approach to Gram-negative entry. Bioorg. Med. Chem. 24, 6379–6389. 10.1016/j.bmc.2016.06.044. [DOI] [PubMed] [Google Scholar]
- Whitfield C.; Trent M. S. (2014) Biosynthesis and export of bacterial lipopolysaccharides. Annu. Rev. Biochem. 83, 99–128. 10.1146/annurev-biochem-060713-035600. [DOI] [PubMed] [Google Scholar]
- Ruiz N.; Kahne D.; Silhavy T. J. (2009) Transport of lipopolysaccharide across the cell envelope: The long road of discovery. Nat. Rev. Microbiol. 7, 677–683. 10.1038/nrmicro2184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morona R.; Daniels C.; Van Den Bosch L. (2003) Genetic modulation of Shigella flexneri 2a lipopolysaccharide O antigen modal chain length reveals that it has been optimized for virulence. Microbiology 149, 925–939. 10.1099/mic.0.26141-0. [DOI] [PubMed] [Google Scholar]
- Gu Y.; Stansfeld P. J.; Zeng Y.; Dong H.; Wang W.; Dong C. (2015) Lipopolysaccharide is inserted into the outer membrane through an intramembrane hole, a lumen gate, and the lateral opening of lptd. Structure 23, 496–504. 10.1016/j.str.2015.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nikaido H.; Varra M. (1985) Molecular basis of bacterial outer membrane permeability. Microbiol Mol. Biol. Rev. 67, 593–656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nikaido H. (1996) Multidrug efflux pumps of gram-negative bacteria. J. Bacteriol. 178, 5853–5859. 10.1128/JB.178.20.5853-5859.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Delcour A. H. (2009) Outer membrane permeability and antibiotic resistance. Biochim. Biophys. Acta, Proteins Proteomics 1794, 808–816. 10.1016/j.bbapap.2008.11.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sugawara E.; Nagano K.; Nikaido H. (2010) Factors affecting the folding of Pseudomonas aeruginosa OprF porin into the one-domain open conformer. mBio 1, 1–9. 10.1128/mBio.00228-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sugawara E.; Nestorovich E. M.; Bezrukov S. M.; Nikaido H. (2006) Pseudomonas aeruginosa porin OprF exists in two different conformations. J. Biol. Chem. 281, 16220–16229. 10.1074/jbc.M600680200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trias J.; Nikaido H. (1990) Protein D2 channel of the Pseudomonas aeruginosa outer membrane has a binding site for basic amino acids and peptides. J. Biol. Chem. 265, 15680–15684. [PubMed] [Google Scholar]
- Nikaido H. (2003) Molecular basis of bacterial outer membrane permeability revisited. Microbiol. Mol. Biol. Rev. 67, 593–656. 10.1128/MMBR.67.4.593-656.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishino K.; Nikaido E.; Yamaguchi A. (2009) Regulation and physiological function of multidrug efflux pumps in Escherichia coli and Salmonella. Biochim. Biophys. Acta, Proteins Proteomics 1794, 834–843. 10.1016/j.bbapap.2009.02.002. [DOI] [PubMed] [Google Scholar]
- Castanheira M.; Deshpande L. M.; Costello A.; Davies T. A.; Jones R. N. (2014) Epidemiology and carbapenem resistance mechanisms of carbapenem-non-susceptible Pseudomonas aeruginosa collected during 2009–11 in 14 European and Mediterranean countries. J. Antimicrob. Chemother. 69, 1804–1814. 10.1093/jac/dku048. [DOI] [PubMed] [Google Scholar]
- Cox G.; Wright G. D. (2013) Intrinsic antibiotic resistance: Mechanisms, origins, challenges and solutions. Int. J. Med. Microbiol. 303, 287–292. 10.1016/j.ijmm.2013.02.009. [DOI] [PubMed] [Google Scholar]
- Hiramatsu K. (2001) Vancomycin-resistant Staphylococcus aureus: a new model of antibiotic resistance. Lancet Infect. Dis. 1, 147–155. 10.1016/S1473-3099(01)00091-3. [DOI] [PubMed] [Google Scholar]
- Gardete S.; Tomasz A. (2014) Mechanisms of vancomycin resistance in Staphylococcus aureus. J. Clin. Invest. 124, 2836–2840. 10.1172/JCI68834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malanovic N.; Lohner K. (2016) Gram-positive bacterial cell envelopes: The impact on the activity of antimicrobial peptides. Biochim. Biophys. Acta, Biomembr. 1858, 936–946. 10.1016/j.bbamem.2015.11.004. [DOI] [PubMed] [Google Scholar]
- Moynihan P. J.; Sychantha D.; Clarke A. J. (2014) Chemical biology of peptidoglycan acetylation and deacetylation. Bioorg. Chem. 54, 44–50. 10.1016/j.bioorg.2014.03.010. [DOI] [PubMed] [Google Scholar]
- Sukhithasri V.; Nisha N.; Biswas L.; Anil Kumar V.; Biswas R. (2013) Innate immune recognition of microbial cell wall components and microbial strategies to evade such recognitions. Microbiol. Res. 168, 396–406. 10.1016/j.micres.2013.02.005. [DOI] [PubMed] [Google Scholar]
- Münch D.; Sahl H.-G. (2015) Structural variations of the cell wall precursor lipid II in Gram-positive bacteria — Impact on binding and efficacy of antimicrobial peptides. Biochim. Biophys. Acta, Biomembr. 1848, 3062–3071. 10.1016/j.bbamem.2015.04.014. [DOI] [PubMed] [Google Scholar]
- Beceiro A.; Tomas M.; Bou G. (2013) Antimicrobial resistance and virulence: a successful or deleterious association in the bacterial world?. Clin. Microbiol. Rev. 26, 185–230. 10.1128/CMR.00059-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bush K.; Bradford P. A. (2016) β-Lactams and β-Lactamase Inhibitors: An Overview. Cold Spring Harbor Perspect. Med. 6, a025247 10.1101/cshperspect.a025247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCarter Y. S.; Wencewicz T. (2017) Antibiotics: Challenges, mechanisms, opportunities. Lab. Med. 48, e42. [Google Scholar]
- Guitor A. K.; Wright G. D. (2018) Antimicrobial resistance and respiratory infections. Chest 154, 1202–1212. 10.1016/j.chest.2018.06.019. [DOI] [PubMed] [Google Scholar]
- Ge M.; Onishi H. R.; Kohler J.; Silver L. L.; Kerns R.; Fukuzawa S.; Thompson C.; Kahne D. (1999) Vancomycin derivatives that inhibit peptidoglycan biosynthesis without binding D-Ala-D-Ala. Science (Washington, DC, U. S.) 284, 507–511. 10.1126/science.284.5413.507. [DOI] [PubMed] [Google Scholar]
- Rao J. (1998) A trivalent system from vancomycin·D-Ala-D-Ala with higher affinity than avidin·biotin. Science (Washington, DC, U. S.) 280, 708–711. 10.1126/science.280.5364.708. [DOI] [PubMed] [Google Scholar]
- Healy V. L.; Lessard I. A.; Roper D. I.; Knox J. R.; Walsh C. T. (2000) Vancomycin resistance in enterococci: reprogramming of the D-ala-D-Ala ligases in bacterial peptidoglycan biosynthesis. Chem. Biol. 7, R109–19. 10.1016/S1074-5521(00)00116-2. [DOI] [PubMed] [Google Scholar]
- Velkov T.; Roberts K. D.; Nation R. L.; Wang J.; Thompson P. E.; Li J. (2014) Teaching “old” polymyxins new tricks: New-generation lipopeptides targeting gram-negative “superbugs.. ACS Chem. Biol. 9, 1172–1177. 10.1021/cb500080r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wright G. D.; Walsh C. T. (2005) D-Alanyl-D-alanine ligases and the molecular mechanism of vancomycin resistance. Acc. Chem. Res. 25, 468–473. 10.1021/ar00022a006. [DOI] [Google Scholar]
- Brown D. (2015) Antibiotic resistance breakers: Can repurposed drugs fill the antibiotic discovery void?. Nat. Rev. Drug Discovery 14, 821–832. 10.1038/nrd4675. [DOI] [PubMed] [Google Scholar]
- Ban N.; Beckmann R.; Cate J. H. D.; Dinman J. D.; Dragon F.; Ellis S. R.; Lafontaine D. L. J.; Lindahl L.; Liljas A.; Lipton J. M.; McAlear M. A.; Moore P. B.; Noller H. F.; Ortega J.; Panse V. G.; Ramakrishnan V.; Spahn C. M. T.; Steitz T. A.; Tchorzewski M.; Tollervey D.; Warren A. J.; Williamson J. R.; Wilson D.; Yonath A.; Yusupov M. (2014) A new system for naming ribosomal proteins. Curr. Opin. Struct. Biol. 24, 165–169. 10.1016/j.sbi.2014.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yonath A.; Bashan A. (2004) Ribosomal crystallography: initiation, peptide bond formation, and amino acid polymerization are hampered by antibiotics. Annu. Rev. Microbiol. 58, 233–251. 10.1146/annurev.micro.58.030603.123822. [DOI] [PubMed] [Google Scholar]
- Schatz A.; Bugle E.; Waksman S. A. (1944) Streptomycin, a substance exhibiting antibiotic activity against Gram-positive and Gram-negative bacteria. Exp. Biol. Med. 55, 66–69. 10.3181/00379727-55-14461. [DOI] [PubMed] [Google Scholar]
- Dal Molin M.; Gut M.; Rominski A.; Haldimann K.; Becker K.; Sander P. (2018) Molecular mechanisms of intrinsic Streptomycin resistance in Mycobacterium abscessus. Antimicrob. Agents Chemother. 62, 1–11. 10.1128/AAC.01427-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Finken M.; Kirschner P.; Meier A.; Wrede A.; Böttger E. C. (1993) Molecular basis of streptomycin resistance in Mycobacterium tuberculosis: alterations of the ribosomal protein S12 gene and point mutations within a functional 16S ribosomal RNA pseudoknot. Mol. Microbiol. 9, 1239–1246. 10.1111/j.1365-2958.1993.tb01253.x. [DOI] [PubMed] [Google Scholar]
- Honore N.; Cole S. T. (1994) Streptomycin resistance in mycobacteria. Antimicrob. Agents Chemother. 38, 238–242. 10.1128/AAC.38.2.238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodman C. D.; Pasaje C. F. A.; Kennedy K.; McFadden G. I.; Ralph S. A. (2016) Targeting protein translation in organelles of the Apicomplexa. Trends Parasitol. 32, 953–965. 10.1016/j.pt.2016.09.011. [DOI] [PubMed] [Google Scholar]
- Liu L. F.; Wang J. C. (1987) Supercoiling of the DNA template during transcription. Proc. Natl. Acad. Sci. U. S. A. 84, 7024–7027. 10.1073/pnas.84.20.7024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxwell A.; Gellert M. (1986) Mechanistic aspects of DNA topoisomerases. Adv. Protein Chem. 38, 69–107. 10.1016/S0065-3233(08)60526-4. [DOI] [PubMed] [Google Scholar]
- Nakanishi A.; Oshida T.; Matsushita T.; Imajoh-Ohmi S.; Ohnuki T. (1998) Identification of DNA gyrase inhibitor (GyrI) in Escherichia coli. J. Biol. Chem. 273, 1933–1938. 10.1074/jbc.273.4.1933. [DOI] [PubMed] [Google Scholar]
- Zechiedrich E. L.; Khodursky A. B.; Cozzarelli N. R. (1997) Topoisomerase IV, not gyrase, decatenates products of site-specific recombination in Escherichia coli. Genes Dev. 11, 2580–2592. 10.1101/gad.11.19.2580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zechiedrich E. L.; Cozzarelli N. R. (1995) Roles of topoisomerase IV and DNA gyrase in DNA unlinking during replication in Escherichia coli. Genes Dev. 9, 2859–2869. 10.1101/gad.9.22.2859. [DOI] [PubMed] [Google Scholar]
- Fitzgibbon J. E.; John J. F.; Delucia J. L.; Dubin D. T. (1998) Topoisomerase mutations in trovafloxacin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 42, 2122–2124. 10.1128/AAC.42.8.2122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maki S.; Takiguchi S.; Miki T.; Horiuchi T. (1992) Modulation of DNA supercoiling activity of Escherichia coli DNA gyrase by F plasmid proteins. J. Biol. Chem. 267, 12244–12251. [PubMed] [Google Scholar]
- Al-Nadaf A. H.; Salah S. A.; Taha M. O. (2018) Discovery of new Gyrase β inhibitors via structure based modeling. Comput. Biol. Chem. 74, 263–272. 10.1016/j.compbiolchem.2018.03.020. [DOI] [PubMed] [Google Scholar]
- Andlid T. A., D’Aimmo M. R., and Jastrebova J. (2018) Folate and Bifidobacteria, in The Bifidobacteria and Related Organisms, pp 195–212, Elsevier. [Google Scholar]
- Dawadi S.; Kordus S. L.; Baughn A. D.; Aldrich C. C. (2017) Synthesis and analysis of bacterial folate metabolism intermediates and antifolates. Org. Lett. 19, 5220–5223. 10.1021/acs.orglett.7b02487. [DOI] [PubMed] [Google Scholar]
- Morgan J.; Smith M.; Mc Auley M. T.; Enrique Salcedo-Sora J. (2018) Disrupting folate metabolism reduces the capacity of bacteria in exponential growth to develop persisters to antibiotics. Microbiology 164, 1432–1445. 10.1099/mic.0.000722. [DOI] [PubMed] [Google Scholar]
- O’Rourke P. E. F.; Eadsforth T. C.; Fyfe P. K.; Shepherd S. M.; Hunter W. N. (2011) Pseudomonas aeruginosa 4-Amino-4-Deoxychorismate Lyase: Spatial conservation of an active site tyrosine and classification of two types of enzyme. PLoS One 6, e24158 10.1371/journal.pone.0024158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kahn S. B.; Fein S. A.; Brodsky I. (1968) Effects of trimethoprim on folate metabolism in man. Clin. Pharmacol. Ther. 9, 550–560. 10.1002/cpt196895550. [DOI] [PubMed] [Google Scholar]
- Marrakchi H.; Lanéelle M. A.; Daffé M. (2014) Mycolic acids: Structures, biosynthesis, and beyond. Chem. Biol. 21, 67–85. 10.1016/j.chembiol.2013.11.011. [DOI] [PubMed] [Google Scholar]
- Schroeder E. K.; de Souza N.; Santos D. S.; Blanchard J. S.; Basso L. A. (2002) Drugs that inhibit mycolic acid biosynthesis in Mycobacterium tuberculosis. Curr. Pharm. Biotechnol. 3, 197–225. 10.2174/1389201023378328. [DOI] [PubMed] [Google Scholar]
- Ngadjeua F.; Braud E.; Saidjalolov S.; Iannazzo L.; Schnappinger D.; Ehrt S.; Hugonnet J.-E.; Mengin-Lecreulx D.; Patin D.; Ethève-Quelquejeu M.; Fonvielle M.; Arthur M. (2018) Critical impact of peptidoglycan precursor amidation on the activity of L, D-transpeptidases from Enterococcus faecium and Mycobacterium tuberculosis. Chem. - Eur. J. 24, 5743–5747. 10.1002/chem.201706082. [DOI] [PubMed] [Google Scholar]
- Raman K.; Rajagopalan P.; Chandra N. (2005) Flux balance analysis of mycolic acid pathway: Targets for anti-tubercular drugs. PLoS Comput. Biol. 1, 0349–0358. 10.1371/journal.pcbi.0010046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gengenbacher M.; Kaufmann S. H. E. (2012) Mycobacterium tuberculosis: Success through dormancy. FEMS Microbiol. Rev. 36, 514–532. 10.1111/j.1574-6976.2012.00331.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhatt A.; Molle V.; Besra G. S.; Jacobs W. R.; Kremer L. (2007) The Mycobacterium tuberculosis FAS-II condensing enzymes: their role in mycolic acid biosynthesis, acid-fastness, pathogenesis and in future drug development. Mol. Microbiol. 64, 1442–1454. 10.1111/j.1365-2958.2007.05761.x. [DOI] [PubMed] [Google Scholar]
- Queval C. J.; Brosch R.; Simeone R. (2017) The Macrophage: A disputed fortress in the battle against Mycobacterium tuberculosis. Front. Microbiol. 8, 2284. 10.3389/fmicb.2017.02284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alahari A.; Trivelli X.; Guérardel Y.; Dover L. G.; Besra G. S.; Sacchettini J. C.; Reynolds R. C.; Coxon G. D.; Kremer L. (2007) Thiacetazone, an antitubercular drug that inhibits cyclopropanation of cell wall mycolic acids in Mycobacteria. PLoS One 2, e1343 10.1371/journal.pone.0001343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lambert P. A. (2002) Cellular impermeability and uptake of biocides and antibiotics in Gram-positive bacteria and mycobacteria. J. Appl. Microbiol. 92, 46S–54S. 10.1046/j.1365-2672.92.5s1.7.x. [DOI] [PubMed] [Google Scholar]
- Bhat K. H., and Yaseen I. (2018) Mycobacterium tuberculosis: Macrophage takeover and modulation of innate effector responses, in Mycobacterium - Research and Development, In Tech, 10.5772/intechopen.75003 [DOI] [Google Scholar]
- Millard J.; Ugarte-Gil C.; Moore D. A. J. (2015) Multidrug resistant tuberculosis. BMJ. 350, h882 10.1136/bmj.h882. [DOI] [PubMed] [Google Scholar]
- North E. J.; Jackson M.; Lee R. E. (2013) New approaches to target the mycolic acid biosynthesis pathway for the development of tuberculosis therapeutics. Curr. Pharm. Des. 20, 4357–4378. 10.2174/1381612819666131118203641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nikaido H. (2009) Multidrug resistance in bacteria. Annu. Rev. Biochem. 78, 119–146. 10.1146/annurev.biochem.78.082907.145923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neu H. C. (1992) The Crisis in Antibiotic Resistance. Science (Washington, DC, U. S.) 257, 1064–1073. 10.1126/science.257.5073.1064. [DOI] [PubMed] [Google Scholar]
- de Lencastre H.; Oliveira D.; Tomasz A. (2007) Antibiotic resistant Staphylococcus aureus: a paradigm of adaptive power. Curr. Opin. Microbiol. 10, 428–435. 10.1016/j.mib.2007.08.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arias C. A.; Contreras G. A.; Murray B. E. (2010) Management of multidrug-resistant enterococcal infections. Clin. Microbiol. Infect. 16, 555–562. 10.1111/j.1469-0691.2010.03214.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bell A. A. (1981) Biochemical mechanisms of disease resistance. Annu. Rev. Plant Physiol. 32, 21–81. 10.1146/annurev.pp.32.060181.000321. [DOI] [Google Scholar]
- Mazel D.; Davies J. (1999) Antibiotic resistance in microbes. Cell. Mol. Life Sci. 56, 742–754. 10.1007/s000180050021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pallasch T. J. (2003) Antibiotic resistance. Dent. Clin. North Am. 47, 623–39. 10.1016/S0011-8532(03)00039-9. [DOI] [PubMed] [Google Scholar]
- Nizet V. (2006) Antimicrobial peptide resistance mechanisms of human bacterial pathogens. Curr. Issues Mol. Biol. 8, 11–26. [PubMed] [Google Scholar]
- Nikaido H. (1998) Multiple antibiotic resistance and efflux. Curr. Opin. Microbiol. 1, 516–523. 10.1016/S1369-5274(98)80083-0. [DOI] [PubMed] [Google Scholar]
- Nikaido H. (1994) Prevention of drug access to bacterial targets: Permeability barriers and active efflux. Science (Washington, DC, U. S.) 264, 382–389. 10.1126/science.8153625. [DOI] [PubMed] [Google Scholar]
- Vila J.; Martí S.; Sánchez-Céspedes J. (2007) Porins, efflux pumps and multidrug resistance in Acinetobacter baumannii. J. Antimicrob. Chemother. 59, 1210–1215. 10.1093/jac/dkl509. [DOI] [PubMed] [Google Scholar]
- Bolla J.-M.; Alibert-Franco S.; Handzlik J.; Chevalier J.; Mahamoud A.; Boyer G.; Kieć-Kononowicz K.; Pagès J.-M. (2011) Strategies for bypassing the membrane barrier in multidrug resistant Gram-negative bacteria. FEBS Lett. 585, 1682–1690. 10.1016/j.febslet.2011.04.054. [DOI] [PubMed] [Google Scholar]
- Buchanan C. E.; Strominger J. L. (1976) Altered penicillin binding components in penicillin resistant mutants of Bacillus subtilis. Proc. Natl. Acad. Sci. U. S. A. 73, 1816–1820. 10.1073/pnas.73.6.1816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levy N.; Bruneau J. M.; Le Rouzic E.; Bonnard D.; Le Strat F.; Caravano A.; Chevreuil F.; Barbion J.; Chasset S.; Ledoussal B.; Moreau F.; Ruff M. (2019) Structural basis for E. coli penicillin binding protein (PBP)2 inhibition, a platform for drug design. J. Med. Chem. 62, 4742–4754. 10.1021/acs.jmedchem.9b00338. [DOI] [PubMed] [Google Scholar]
- Wright G. D. (2005) Bacterial resistance to antibiotics: Enzymatic degradation and modification. Adv. Drug Delivery Rev. 57, 1451–1470. 10.1016/j.addr.2005.04.002. [DOI] [PubMed] [Google Scholar]
- Thomson K. S.; Smith Moland E. (2000) Version 2000: The new β-lactamases of Gram-negative bacteria at the dawn of the new millennium. Microbes Infect. 2, 1225–1235. 10.1016/S1286-4579(00)01276-4. [DOI] [PubMed] [Google Scholar]
- Xu Z.; Stogios P. J.; Quaile A. T.; Forsberg K. J.; Patel S.; Skarina T.; Houliston S.; Arrowsmith C.; Dantas G.; Savchenko A. (2017) Structural and functional survey of environmental aminoglycoside acetyltransferases reveals functionality of resistance enzymes. ACS Infect. Dis. 3, 653–665. 10.1021/acsinfecdis.7b00068. [DOI] [PubMed] [Google Scholar]
- Magnet S.; Blanchard J. S. (2005) Molecular insights into aminoglycoside action and resistance. Chem. Rev. 105, 477–98. 10.1021/cr0301088. [DOI] [PubMed] [Google Scholar]
- Davies J. (1994) Inactivation of antibiotics and the dissemination of resistance genes. Science (Washington, DC, U. S.) 264, 375–382. 10.1126/science.8153624. [DOI] [PubMed] [Google Scholar]
- Antonczak A. K.; Simova Z.; Tippmann E. M. (2009) A critical examination of Escherichia coli esterase activity. J. Biol. Chem. 284, 28795–28800. 10.1074/jbc.M109.027409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morar M.; Pengelly K.; Koteva K.; Wright G. D. (2012) Mechanism and diversity of the erythromycin esterase family of enzymes. Biochemistry 51, 1740–1751. 10.1021/bi201790u. [DOI] [PubMed] [Google Scholar]
- Nash K. A.; Brown-Elliott A. B.; Wallace R. J. (2009) A Novel gene, erm(41), confers inducible macrolide resistance to clinical isolates of Mycobacterium abscessus but is absent from Mycobacterium chelonae. Antimicrob. Agents Chemother. 53, 1367–1376. 10.1128/AAC.01275-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rominski A.; Roditscheff A.; Selchow P.; Böttger E. C.; Sander P. (2017) Intrinsic rifamycin resistance of Mycobacterium abscessus is mediated by ADP-ribosyltransferase MAB_0591. J. Antimicrob. Chemother. 72, 376–384. 10.1093/jac/dkw466. [DOI] [PubMed] [Google Scholar]
- Dubee V.; Soroka D.; Cortes M.; Lefebvre A.-L.; Gutmann L.; Hugonnet J.-E.; Arthur M.; Mainardi J.-L. (2015) Impact of β-lactamase inhibition on the activity of ceftaroline against Mycobacterium tuberculosis and Mycobacterium abscessus. Antimicrob. Agents Chemother. 59, 2938–2941. 10.1128/AAC.05080-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rominski A.; Selchow P.; Becker K.; Brülle J. K.; Dal Molin M.; Sander P. (2017) Elucidation of Mycobacterium abscessus aminoglycoside and capreomycin resistance by targeted deletion of three putative resistance genes. J. Antimicrob. Chemother. 72, 2191–2200. 10.1093/jac/dkx125. [DOI] [PubMed] [Google Scholar]
- Spratt B. G. (1994) Resistance to antibiotics mediated by target alterations. Science (Washington, DC, U. S.) 264, 388–393. 10.1126/science.8153626. [DOI] [PubMed] [Google Scholar]
- Kuroda M.; Ohta T.; Uchiyama I.; Baba T.; Yuzawa H.; Kobayashi I.; Cui L.; Oguchi A.; Aoki K.; Nagai Y.; Lian J.; Ito T.; Kanamori M.; Matsumaru H.; Maruyama A.; Murakami H.; Hosoyama A.; Mizutani-Ui Y.; Takahashi N. K.; Sawano T.; Inoue R.; Kaito C.; Sekimizu K.; Hirakawa H.; Kuhara S.; Goto S.; Yabuzaki J.; Kanehisa M.; Yamashita A.; Oshima K.; Furuya K.; Yoshino C.; Shiba T.; Hattori M.; Ogasawara N.; Hayashi H.; Hiramatsu K. (2001) Whole genome sequencing of meticillin-resistant Staphylococcus aureus. Lancet 357, 1225–1240. 10.1016/S0140-6736(00)04403-2. [DOI] [PubMed] [Google Scholar]
- Carniel E. (2002) Plasmids and pathogenicity islands of Yersinia. Curr. Top. Microbiol. Immunol. 264, 89–108. 10.1007/978-3-642-56031-6_6. [DOI] [PubMed] [Google Scholar]
- Tenover F. C. (2006) Mechanisms of antimicrobial resistance in bacteria. Am. J. Infect. Control 34, S3–S10. 10.1016/j.ajic.2006.05.219. [DOI] [PubMed] [Google Scholar]
- Kim S. J.; Matsuoka S.; Patti G. J.; Schaefer J. (2008) Vancomycin derivative with damaged D-Ala-D-Ala binding cleft binds to cross-linked peptidoglycan in the cell wall of Staphylococcus aureus. Biochemistry 47, 3822–3831. 10.1021/bi702232a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Courvalin P. (2006) Vancomycin resistance in Gram-positive cocci. Clin. Infect. Dis. 42, S25–S34. 10.1086/491711. [DOI] [PubMed] [Google Scholar]
- Gotsbacher M. P.; Cho S.; Kwon H. J.; Karuso P. (2016) Daptomycin, a last-resort antibiotic, binds ribosomal protein S19 in humans. Proteome Sci. 15, 16. 10.1186/s12953-017-0124-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imai Y.; Meyer K. J.; Iinishi A.; Favre-Godal Q.; Green R.; Manuse S.; Caboni M.; Mori M.; Niles S.; Ghiglieri M.; Honrao C.; Ma X.; Guo J. J.; Makriyannis A.; Linares-Otoya L.; Böhringer N.; Wuisan Z. G.; Kaur H.; Wu R.; Mateus A.; Typas A.; Savitski M. M.; Espinoza J. L.; O’Rourke A.; Nelson K. E.; Hiller S.; Noinaj N.; Schäberle T. F.; D’Onofrio A.; Lewis K. (2019) A new antibiotic selectively kills Gram-negative pathogens. Nature 576, 459–464. 10.1038/s41586-019-1791-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller A.; Wenzel M.; Strahl H.; Grein F.; Saaki T. N. V.; Kohl B.; Siersma T.; Bandow J. E.; Sahl H. G.; Schneider T.; Hamoen L. W. (2016) Daptomycin inhibits cell envelope synthesis by interfering with fluid membrane microdomains. Proc. Natl. Acad. Sci. U. S. A. 113, E7077–E7086. 10.1073/pnas.1611173113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sauermann R.; Rothenburger M.; Graninger W.; Joukhadar C. (2008) Daptomycin: A review 4 years after first approval. Pharmacology 81, 79–91. 10.1159/000109868. [DOI] [PubMed] [Google Scholar]
- Koo H.; Allan R. N.; Howlin R. P.; Stoodley P.; Hall-Stoodley L. (2017) Targeting microbial biofilms: Current and prospective therapeutic strategies. Nat. Rev. Microbiol. 15, 740–755. 10.1038/nrmicro.2017.99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flemming H. C.; Wingender J.; Szewzyk U.; Steinberg P.; Rice S. A.; Kjelleberg S. (2016) Biofilms: An emergent form of bacterial life. Nat. Rev. Microbiol. 14, 563–575. 10.1038/nrmicro.2016.94. [DOI] [PubMed] [Google Scholar]
- Van Acker H.; Van Dijck P.; Coenye T. (2014) Molecular mechanisms of antimicrobial tolerance and resistance in bacterial and fungal biofilms. Trends Microbiol. 22, 326–333. 10.1016/j.tim.2014.02.001. [DOI] [PubMed] [Google Scholar]
- Lebeaux D.; Ghigo J.-M.; Beloin C. (2014) Biofilm-related infections: Bridging the gap between clinical management and fundamental aspects of recalcitrance toward antibiotics. Microbiol. Mol. Biol. Rev. 78, 510–543. 10.1128/MMBR.00013-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cunningham R.; Cheesbrough J. (1992) Comparative activity of glycopeptide antibiotics against coagulase-negative stapfaylococci embedded in fibrin clots. J. Antimicrob. Chemother. 30, 321–326. 10.1093/jac/30.3.321. [DOI] [PubMed] [Google Scholar]
- Higashi J. M.; Wang I. W.; Shlaes D. M.; Anderson J. M.; Marchant R. E. (1998) Adhesion of Staphylococcus epidermidis and transposon mutant strains to hydrophobic polyethylene. J. Biomed. Mater. Res. 39, 341–350. 10.1002/(SICI)1097-4636(19980305)39:3<341::AID-JBM1>3.0.CO;2-J. [DOI] [PubMed] [Google Scholar]
- Raad I.; Darouiche R.; Hachem R.; Sacilowski M.; Bodey G. P. (1995) Antibiotics and prevention of microbial colonization of catheters. Antimicrob. Agents Chemother. 39, 2397–2400. 10.1128/AAC.39.11.2397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stewart P. S. (2002) Mechanisms of antibiotic resistance in bacterial biofilms. Int. J. Med. Microbiol. 292, 107–113. 10.1078/1438-4221-00196. [DOI] [PubMed] [Google Scholar]
- Hall M. R.; McGillicuddy E.; Kaplan L. J. (2014) Biofilm: Basic principles, pathophysiology, and implications for clinicians. Surg. Infect. (Larchmt). 15, 1–7. 10.1089/sur.2012.129. [DOI] [PubMed] [Google Scholar]
- Nishimura S.; Tsurumoto T.; Yonekura A.; Adachi K.; Shindo H. (2006) Antimicrobial susceptibility of Staphylococcus aureus and Staphylococcus epidermidis biofilms isolated from infected total hip arthroplasty cases. J. Orthop. Sci. 11, 46–50. 10.1007/s00776-005-0968-7. [DOI] [PubMed] [Google Scholar]
- Stewart P. S.; Costerton J. W. (2001) Antibiotic resistance of bacteria in biofilms. Lancet 358, 135–138. 10.1016/S0140-6736(01)05321-1. [DOI] [PubMed] [Google Scholar]
- Sutherland I. W. (2001) The biofilm matrix - An immobilized but dynamic microbial environment. Trends Microbiol. 9, 222–227. 10.1016/S0966-842X(01)02012-1. [DOI] [PubMed] [Google Scholar]
- Velkov T.; Roberts K. D.; Li J. (2017) Rediscovering the octapeptins. Nat. Prod. Rep. 34, 295–309. 10.1039/C6NP00113K. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li J.; Milne R. W.; Nation R. L.; Turnidge J. D.; Smeaton T. C.; Coulthard K. (2004) Pharmacokinetics of colistin methanesulphonate and colistin in rats following an intravenous dose of colistin methanesulphonate. J. Antimicrob. Chemother. 53, 837–840. 10.1093/jac/dkh167. [DOI] [PubMed] [Google Scholar]
- Li J.; Rayner C. R.; Nation R. L.; Owen R. J.; Spelman D.; Tan K. E.; Liolios L. (2006) Heteroresistance to colistin in multidrug-resistant Acinetobacter baumannii. Antimicrob. Agents Chemother. 50, 2946–2950. 10.1128/AAC.00103-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li J.; Nation R. L.; Turnidge J. D.; Milne R. W.; Coulthard K.; Rayner C. R.; Paterson D. L. (2006) Colistin: the re-emerging antibiotic for multidrug-resistant Gram-negative bacterial infections. Lancet Infect. Dis. 6, 589–601. 10.1016/S1473-3099(06)70580-1. [DOI] [PubMed] [Google Scholar]
- Johansen C.; Falholt P.; Gram L. (1997) Enzymatic removal and disinfection of bacterial biofilms. Appl. Environ. Microbiol. 63, 3724–3728. 10.1128/AEM.63.9.3724-3728.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hatch R. A.; Schiller N. L. (1998) Alginate lyase promotes diffusion of aminoglycosides through the extracellular polysaccharide of mucoid Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 42, 974–977. 10.1128/AAC.42.4.974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shuford J. A.; Steckelberg J. M.; Patel R. (2005) Effects of fresh garlic extract on Candida albicans biofilms. Antimicrob. Agents Chemother. 49, 473–473. 10.1128/AAC.49.1.473.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rabin N.; Zheng Y.; Opoku-Temeng C.; Du Y.; Bonsu E.; Sintim H. O. (2015) Agents that inhibit bacterial biofilm formation. Future Med. Chem. 7, 647–671. 10.4155/fmc.15.7. [DOI] [PubMed] [Google Scholar]
- Baral B.; Maharjan B. L. (2011) Assessment of antimicrobial and phytochemical potentials of high altitudinal Nepalese lichens. J. Microbiol. Biotechnol. Food Sci. 1, 98–112. [Google Scholar]
- Maharjan B. L.; Mainali S.; Baral B. (2011) Phytochemical screening and antimicrobial assay of some Nepalese medicinal plants. Sci. World 9, 90–92. 10.3126/sw.v9i9.5526. [DOI] [Google Scholar]
- Hentzer M.; Teitzel G. M.; Balzer G. J.; Heydorn A.; Molin S.; Givskov M.; Parsek M. R. (2001) Alginate overproduction affects Pseudomonas aeruginosa biofilm structure and function. J. Bacteriol. 183, 5395–5401. 10.1128/JB.183.18.5395-5401.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shih P.; Huang C. (2002) Effects of quorum-sensing deficiency on Pseudomonas aeruginosa. J. Antimicrob. Chemother. 49, 309–314. 10.1093/jac/49.2.309. [DOI] [PubMed] [Google Scholar]
- Davies D. G.; Chakrabarty A. M.; Geesey G. G. (1993) Exopolysaccharide production in biofilms: Substratum activation of alginate gene expression by Pseudomonas aeruginosa. Appl. Environ. Microbiol. 59, 1181–1186. 10.1128/AEM.59.4.1181-1186.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davies D. G.; Geesey G. G. (1995) Regulation of the alginate biosynthesis gene algC in Pseudomonas aeruginosa during biofilm development in continuous culture. Appl. Environ. Microbiol. 61, 860–867. 10.1128/AEM.61.3.860-867.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Becker P.; Hufnagle W.; Peters G.; Herrmann M. (2001) Detection of differential gene expression in biofilm-forming versus planktonic populations of Staphylococcus aureus using micro-representational-difference analysis. Appl. Environ. Microbiol. 67, 2958–2965. 10.1128/AEM.67.7.2958-2965.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anwar H.; Van Biesen T.; Dasgupta M.; Lam K.; Costerton J. W. (1989) Interaction of biofilm bacteria with antibiotics in a novel in vitro chemostat system. Antimicrob. Agents Chemother. 33, 1824–1826. 10.1128/AAC.33.10.1824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams I.; Venables W. A.; Lloyd D.; Paul F.; Critchley I. (1997) The effects of adherence to silicone surfaces on antibiotic susceptibility in Staphylococcus aureus. Microbiology 143, 2407–2413. 10.1099/00221287-143-7-2407. [DOI] [PubMed] [Google Scholar]
- Anderl J. N.; Franklin M. J.; Stewart P. S. (2000) Role of antibiotic penetration limitation in Klebsiella pneumoniae biofilm resistance to ampicillin and ciprofloxacin. Antimicrob. Agents Chemother. 44, 1818–1824. 10.1128/AAC.44.7.1818-1824.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whiteley M.; Bangera M. G.; Bumgarner R. E.; Parsek M. R.; Teitzel G. M.; Lory S.; Greenberg E. P. (2001) Gene expression in Pseudomonas aeruginosa biofilms. Nature 413, 860–864. 10.1038/35101627. [DOI] [PubMed] [Google Scholar]
- Mah T.-F.; Pitts B.; Pellock B.; Walker G. C.; Stewart P. S.; O'Toole G. A. (2003) A genetic basis for Pseudomonas aeruginosa biofilm antibiotic resistance. Nature 426, 1–5. 10.1038/nature02122. [DOI] [PubMed] [Google Scholar]
- Hoffman L. R.; D’Argenio D. A.; MacCoss M. J.; Zhang Z.; Jones R. A.; Miller S. I. (2005) Aminoglycoside antibiotics induce bacterial biofilm formation. Nature 436, 1171–1175. 10.1038/nature03912. [DOI] [PubMed] [Google Scholar]
- D’Argenio D. A.; Miller S. I. (2004) Cyclic di-GMP as a bacterial second messenger. Microbiology 150, 2497–2502. 10.1099/mic.0.27099-0. [DOI] [PubMed] [Google Scholar]
- Brooun A.; Liu S.; Lewis K. (2000) A dose-response study of antibiotic resistance in Pseudomonas aeruginosa biofilms. Antimicrob. Agents Chemother. 44, 640–646. 10.1128/AAC.44.3.640-646.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khardori N.; Yassien M.; Wilson K. (1995) Tolerance of Staphylococcus epidermidis grown from indwelling vascular catheters to antimicrobial agents. J. Ind. Microbiol. 15, 148–151. 10.1007/BF01569818. [DOI] [PubMed] [Google Scholar]
- Monzón M.; Oteiza C.; Leiva J.; Lamata M.; Amorena B. (2002) Biofilm testing of Staphylococcus epidermidis clinical isolates: Low performance of vancomycin in relation to other antibiotics. Diagn. Microbiol. Infect. Dis. 44, 319–324. 10.1016/S0732-8893(02)00464-9. [DOI] [PubMed] [Google Scholar]
- Davin-Regli A.; Bolla J.-M.; James C. E.; Lavigne J.-P.; Chevalier J.; Garnotel E.; Molitor A.; Pagès J.-M. (2008) Membrane permeability and regulation of drug “influx and efflux” in enterobacterial pathogens. Curr. Drug Targets 9, 750–759. 10.2174/138945008785747824. [DOI] [PubMed] [Google Scholar]
- Mazzariol A.; Cornaglia G.; Nikaido H. (2000) Contributions of the AmpC beta-lactamase and the AcrAB multidrug efflux system in intrinsic resistance of Escherichia coli K-12 to Beta-lactams. Antimicrob. Agents Chemother. 44, 1387–1390. 10.1128/AAC.44.5.1387-1390.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hassan K. A.; Liu Q.; Henderson P. J. F.; Paulsen I. T. (2015) Homologs of the Acinetobacter baumannii acei transporter represent a new family of bacterial multidrug efflux systems. mBio 6, 1–5. 10.1128/mBio.01982-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mahamoud A.; Chevalier J.; Alibert-Franco S.; Kern W. V.; Pagès J. M. (2007) Antibiotic efflux pumps in Gram-negative bacteria: The inhibitor response strategy. J. Antimicrob. Chemother. 59, 1223–1229. 10.1093/jac/dkl493. [DOI] [PubMed] [Google Scholar]
- Van Bambeke F.; Balzi E.; Tulkens P. M (2000) Antibiotic efflux pumps. Biochem. Pharmacol. 60, 1–24. 10.1016/S0006-2952(00)00291-4. [DOI] [PubMed] [Google Scholar]
- Stavri M.; Piddock L. J. V.; Gibbons S. (2007) Bacterial efflux pump inhibitors from natural sources. J. Antimicrob. Chemother. 59, 1247–1260. 10.1093/jac/dkl460. [DOI] [PubMed] [Google Scholar]
- Yerushalmi H.; Lebendiker M.; Schuldiner S. (1995) EmrE, an Escherichia coli 12-kDa multidrug transporter, exchanges toxic cations and H+ and is soluble in organic solvents. J. Biol. Chem. 270, 6856. 10.1074/jbc.270.12.6856. [DOI] [PubMed] [Google Scholar]
- Wandersman C. (1992) Secretion across the bacterial outer membrane. Trends Genet. 8, 317–322. 10.1016/0168-9525(92)90264-5. [DOI] [PubMed] [Google Scholar]
- Wandersman C.; Delepelaire P. (1990) TolC, an Escherichia coli outer membrane protein required for hemolysin secretion. Proc. Natl. Acad. Sci. U. S. A. 87, 4776–4780. 10.1073/pnas.87.12.4776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baral B. (2017) Evolutionary trajectories of entomopathogenic fungi ABC transporters. Adv. Genet. 98, 117–154. 10.1016/bs.adgen.2017.07.002. [DOI] [PubMed] [Google Scholar]
- Lewis K. (1994) Multidrug resistance pumps in bacteria: variations on a theme. Trends Biochem. Sci. 19, 119–123. 10.1016/0968-0004(94)90204-6. [DOI] [PubMed] [Google Scholar]
- Lin Y. T.; Huang Y. W.; Liou R. S.; Chang Y. C.; Yang T. C. (2014) MacABCsm, an ABC-type tripartite efflux pump of Stenotrophomonas maltophilia involved in drug resistance, oxidative and envelope stress tolerances and biofilm formation. J. Antimicrob. Chemother. 69, 3221–3226. 10.1093/jac/dku317. [DOI] [PubMed] [Google Scholar]
- Al-Hamad A.; Upton M.; Burnie J. (2009) Molecular cloning and characterization of SmrA, a novel ABC multidrug efflux pump from Stenotrophomonas maltophilia. J. Antimicrob. Chemother. 64, 731–734. 10.1093/jac/dkp271. [DOI] [PubMed] [Google Scholar]
- Fitzpatrick A. W. P.; Llabrés S.; Neuberger A.; Blaza J. N.; Bai X.-C.; Okada U.; Murakami S.; van Veen H. W.; Zachariae U.; Scheres S. H. W.; Luisi B. F.; Du D. (2017) Structure of the MacAB-TolC ABC-type tripartite multidrug efflux pump. Nat. Microbiol. 2, 17070. 10.1038/nmicrobiol.2017.70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamanaka H.; Kobayashi H.; Takahashi E.; Okamoto K. (2008) MacAB is involved in the secretion of Escherichia coli heat-stable enterotoxin II. J. Bacteriol. 190, 7693–7698. 10.1128/JB.00853-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown M. H.; Paulsen I. T.; Skurray R. A. (1999) The multidrug efflux protein NorM is a prototype of a new family of transporters. Mol. Microbiol. 31, 393–395. 10.1046/j.1365-2958.1999.01162.x. [DOI] [PubMed] [Google Scholar]
- He G.-X.; Kuroda T.; Mima T.; Morita Y.; Mizushima T.; Tsuchiya T. (2004) An H+-coupled multidrug efflux pump, PmpM, a member of the MATE family of transporters, from Pseudomonas aeruginosa. J. Bacteriol. 186, 262–265. 10.1128/JB.186.1.262-265.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McAleese F.; Petersen P.; Ruzin A.; Dunman P. M.; Murphy E.; Projan S. J.; Bradford P. A. (2005) A novel MATE family efflux pump contributes to the reduced susceptibility of laboratory-derived Staphylococcus aureus mutants to tigecycline. Antimicrob. Agents Chemother. 49, 1865–1871. 10.1128/AAC.49.5.1865-1871.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morita Y.; Kodama K.; Shiota S.; Mine T.; Kataoka A.; Mizushima T.; Tsuchiya T. (1998) NorM, a putative multidrug efflux protein, of Vibrio parahaemolyticus and its homolog in Escherichia coli. Antimicrob. Agents Chemother. 42, 1778–1782. 10.1128/AAC.42.7.1778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morita Y.; Kataoka A.; Shiota S.; Mizushima T.; Tsuchiya T. (2000) NorM of Vibrio parahaemolyticus is an Na+-driven multidrug efflux pump. J. Bacteriol. 182, 6694–6697. 10.1128/JB.182.23.6694-6697.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Su X.-Z.; Chen J.; Mizushima T.; Kuroda T.; Tsuchiya T. (2005) AbeM, an H+-coupled Acinetobacter baumannii multidrug efflux pump belonging to the MATE family of transporters. Antimicrob. Agents Chemother. 49, 4362–4364. 10.1128/AAC.49.10.4362-4364.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Omote H.; Hiasa M.; Matsumoto T.; Otsuka M.; Moriyama Y. (2006) The MATE proteins as fundamental transporters of metabolic and xenobiotic organic cations. Trends Pharmacol. Sci. 27, 587–593. 10.1016/j.tips.2006.09.001. [DOI] [PubMed] [Google Scholar]
- Mousa J. J.; Yang Y.; Tomkovich S.; Shima A.; Newsome R. C.; Tripathi P.; Oswald E.; Bruner S. D.; Jobin C. (2016) MATE transport of the E. coli-derived genotoxin colibactin. Nat. Microbiol. 1, 15009. 10.1038/nmicrobiol.2015.9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuroda T.; Tsuchiya T. (2009) Multidrug efflux transporters in the MATE family. Biochim. Biophys. Acta, Proteins Proteomics 1794, 763–768. 10.1016/j.bbapap.2008.11.012. [DOI] [PubMed] [Google Scholar]
- Shiomi N.; Fukuda H.; Fukuda Y.; Murata K.; Kimura A. (1991) Nucleotide sequence and characterization of a gene conferring resistance to ethionine in yeast Saccharomyces cerevisiae. J. Ferment. Bioeng. 71, 211–215. 10.1016/0922-338X(91)90269-M. [DOI] [Google Scholar]
- Otsuka M.; Matsumoto T.; Morimoto R.; Arioka S.; Omote H.; Moriyama Y. (2005) A human transporter protein that mediates the final excretion step for toxic organic cations. Proc. Natl. Acad. Sci. U. S. A. 102, 17923–17928. 10.1073/pnas.0506483102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyauchi H.; Moriyama S.; Kusakizako T.; Kumazaki K.; Nakane T.; Yamashita K.; Hirata K.; Dohmae N.; Nishizawa T.; Ito K.; Miyaji T.; Moriyama Y.; Ishitani R.; Nureki O. (2017) Structural basis for xenobiotic extrusion by eukaryotic MATE transporter. Nat. Commun. 8, 1633. 10.1038/s41467-017-01541-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Griffith J. K.; Baker M. E.; Rouch D. A.; Page M. G. P.; Skurray R. A.; Paulsen I. T.; Chater K. F.; Baldwin S. A.; Henderson P. J. F. (1992) Membrane transport proteins: implications of sequence comparisons. Curr. Opin. Cell Biol. 4, 684–695. 10.1016/0955-0674(92)90090-Y. [DOI] [PubMed] [Google Scholar]
- Du D.; van Veen H. W.; Murakami S.; Pos K. M.; Luisi B. F. (2015) Structure, mechanism and cooperation of bacterial multidrug transporters. Curr. Opin. Struct. Biol. 33, 76–91. 10.1016/j.sbi.2015.07.015. [DOI] [PubMed] [Google Scholar]
- Putman M.; van Veen H. W.; Konings W. N. (2000) Molecular properties of bacterial multidrug transporters. Microbiol. Mol. Biol. Rev. 64, 672–693. 10.1128/MMBR.64.4.672-693.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abramson J. (2003) Structure and mechanism of the lactose permease of Escherichia coli. Science (Washington, DC, U. S.) 301, 610–615. 10.1126/science.1088196. [DOI] [PubMed] [Google Scholar]
- Wisedchaisri G.; Park M.-S.; Iadanza M. G.; Zheng H.; Gonen T. (2014) Proton-coupled sugar transport in the prototypical major facilitator superfamily protein XylE. Nat. Commun. 5, 4521. 10.1038/ncomms5521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun J.; Bankston J. R.; Payandeh J.; Hinds T. R.; Zagotta W. N.; Zheng N. (2014) Crystal structure of the plant dual-affinity nitrate transporter NRT1.1. Nature 507, 73–77. 10.1038/nature13074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deng D.; Xu C.; Sun P.; Wu J.; Yan C.; Hu M.; Yan N. (2014) Crystal structure of the human glucose transporter GLUT1. Nature 510, 121–125. 10.1038/nature13306. [DOI] [PubMed] [Google Scholar]
- Huang Y.-W.; Hu R.-M.; Chu F.-Y.; Lin H.-R.; Yang T.-C. (2013) Characterization of a major facilitator superfamily (MFS) tripartite efflux pump EmrCABsm from Stenotrophomonas maltophilia. J. Antimicrob. Chemother. 68, 2498–2505. 10.1093/jac/dkt250. [DOI] [PubMed] [Google Scholar]
- Fluman N.; Ryan C. M.; Whitelegge J. P.; Bibi E. (2012) Dissection of mechanistic principles of a secondary multidrug efflux protein. Mol. Cell 47, 777–787. 10.1016/j.molcel.2012.06.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen D. E.; Podell S.; Sauer J. D.; Swanson M. S.; Saier M. H. (2008) The phagosomal nutrient transporter (Pht) family. Microbiology 154, 42–53. 10.1099/mic.0.2007/010611-0. [DOI] [PubMed] [Google Scholar]
- Piddock L. J. V. (2006) Multidrug-resistance efflux pumps- not just for resistance. Nat. Rev. Microbiol. 4, 629–636. 10.1038/nrmicro1464. [DOI] [PubMed] [Google Scholar]
- Piddock L. J. V. (2006) Clinically relevant chromosomally encoded multidrug resistance efflux pumps in bacteria. Clin. Microbiol. Rev. 19, 382–402. 10.1128/CMR.19.2.382-402.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lorca G. L.; Barabote R. D.; Zlotopolski V.; Tran C.; Winnen B.; Hvorup R. N.; Stonestrom A. J.; Nguyen E.; Huang L.-W.; Kim D. S.; Saier M. H. (2007) Transport capabilities of eleven gram-positive bacteria: Comparative genomic analyses. Biochim. Biophys. Acta, Biomembr. 1768, 1342–1366. 10.1016/j.bbamem.2007.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saier M. H. Jr; Paulsen I. T. (2001) Phylogeny of multidrug transporters. Semin. Cell Dev. Biol. 12, 205–213. 10.1006/scdb.2000.0246. [DOI] [PubMed] [Google Scholar]
- Neyfakh A. A.; Bidnenko V. E.; Chen L. B. (1991) Efflux-mediated multidrug resistance in Bacillus subtilis: similarities and dissimilarities with the mammalian system. Proc. Natl. Acad. Sci. U. S. A. 88, 4781–4785. 10.1073/pnas.88.11.4781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lomovskaya O.; Lewis K. (1992) Emr, an Escherichia coli locus for multidrug resistance. Proc. Natl. Acad. Sci. U. S. A. 89, 8938–8942. 10.1073/pnas.89.19.8938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tennent J. M.; Lyon B. R.; Midgley M.; Jones G.; Purewal A. S.; Skurray R. A. (1989) Physical and biochemical characterization of the qmA gene encoding antiseptic and disinfectant resistance in Staphylococcus aureus. Microbiology 135, 1–10. 10.1099/00221287-135-1-1. [DOI] [PubMed] [Google Scholar]
- Rouch D.; Cram D.; DiBerardino D.; Littlejohn T.; Skurray R. (1990) Efflux-mediated antiseptic resistance gene qacA from Staphylococcus aureus: commonancestry with tetracycline- and sugar-transport proteins. Mol. Microbiol. 4, 2051. 10.1111/j.1365-2958.1990.tb00565.x. [DOI] [PubMed] [Google Scholar]
- Yamane K.; Wachino J.-I.; Suzuki S.; Kimura K.; Shibata N.; Kato H.; Shibayama K.; Konda T.; Arakawa Y. (2007) New plasmid-mediated fluoroquinolone efflux pump, QepA, found in an Escherichia coli clinical isolate. Antimicrob. Agents Chemother. 51, 3354–3360. 10.1128/AAC.00339-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yin Y. (2006) Structure of the multidrug transporter EmrD from Escherichia coli. Science (Washington, DC, U. S.) 312, 741–744. 10.1126/science.1125629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang D.; Zhao Y.; Wang X.; Fan J.; Heng J.; Liu X.; Feng W.; Kang X.; Huang B.; Liu J.; Zhang X. C. (2013) Structure of the YajR transporter suggests a transport mechanism based on the conserved motif A. Proc. Natl. Acad. Sci. U. S. A. 110, 14664–14669. 10.1073/pnas.1308127110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roca I.; Marti S.; Espinal P.; Martinez P.; Gibert I.; Vila J. (2009) CraA, a aajor facilitator superfamily efflux pump associated with chloramphenicol resistance inAcinetobacter baumannii. Antimicrob. Agents Chemother. 53, 4013–4014. 10.1128/AAC.00584-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanabe M.; Szakonyi G.; Brown K. A.; Henderson P. J. F.; Nield J.; Byrne B. (2009) The multidrug resistance efflux complex, EmrAB from Escherichia coli forms a dimer in vitro. Biochem. Biophys. Res. Commun. 380, 338–342. 10.1016/j.bbrc.2009.01.081. [DOI] [PubMed] [Google Scholar]
- Chung Y. J.; Saier M. H. (2001) SMR-type multidrug resistance pumps. Curr. Opin. Drug Discovery Devel. 4, 237–245. [PubMed] [Google Scholar]
- Saier M. H.; Paulsen I. T.; Sliwinski M. K.; Pao S. S.; Skurray R. A.; Nikaido H. (1998) Evolutionary origins of multidrug and drug-specific efflux pumps in bacteria. FASEB J. 12, 265–274. 10.1096/fasebj.12.3.265. [DOI] [PubMed] [Google Scholar]
- Grinius L.; Dreguniene G.; Goldberg E. B.; Liao C. H.; Projan S. J. (1992) A staphylococcal multidrug resistance gene product is a member of a new protein family. Plasmid 27, 119–129. 10.1016/0147-619X(92)90012-Y. [DOI] [PubMed] [Google Scholar]
- Paulsen I. T.; Skurray R. A.; Tam R.; Saier M. H.; Turner R. J.; Weiner J. H.; Goldberg E. B.; Grinius L. L. (1996) The SMR family: A novel family of multidrug efflux proteins involved with the efflux of lipophilic drugs. Mol. Microbiol. 19, 1167–1175. 10.1111/j.1365-2958.1996.tb02462.x. [DOI] [PubMed] [Google Scholar]
- Littlejohn T. G.; DiBerardino D.; Messerotti L. J.; Spiers S. J.; Skurray R. A. (1991) Structure and evolution of a family of genes encoding antiseptic and disinfectant resistance in Staphylococcus aureus. Gene 101, 59–66. 10.1016/0378-1119(91)90224-Y. [DOI] [PubMed] [Google Scholar]
- Paulsen I. T.; Skurray R. A. (1993) Topology, structure and evolution of two families of proteins involved in antibiotic and antiseptic resistance in eukaryotes and prokaryotes - an analysis. Gene 124, 1–11. 10.1016/0378-1119(93)90755-R. [DOI] [PubMed] [Google Scholar]
- Morimyo M.; Hongo E.; Hama-Inaba H.; Machida I. (1992) Cloning and characterization of the mvrC gene of Escherichia coli K-12 which confers resistance against methyl viologen toxicity. Nucleic Acids Res. 20, 3159–65. 10.1093/nar/20.12.3159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y.-J.; Pornillos O.; Lieu S.; Ma C.; Chen A. P.; Chang G. (2007) X-ray structure of EmrE supports dual topology model. Proc. Natl. Acad. Sci. U. S. A. 104, 18999–19004. 10.1073/pnas.0709387104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nikaido H.; Basina M.; Nguyen V.; Rosenberg E. Y. (1998) Multidrug efflux pump AcrAB of Salmonella typhimurium excretes only those β -Lactam antibiotics containing lipophilic side chains. J. Bacteriol. 180, 4686–4692. 10.1128/JB.180.17.4686-4692.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferreira M. M. C.; Kiralj R. (2004) QSAR study of β-lactam antibiotic efflux by the bacterial multidrug resistance pump AcrB. J. Chemom. 18, 242–252. 10.1002/cem.867. [DOI] [Google Scholar]
- Rampioni G.; Pillai C. R.; Longo F.; Bondì R.; Baldelli V.; Messina M.; Imperi F.; Visca P.; Leoni L. (2017) Effect of efflux pump inhibition on Pseudomonas aeruginosa transcriptome and virulence. Sci. Rep. 7, 11392. 10.1038/s41598-017-11892-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seeger M. A.; Eicher T. (2011) Structural asymmetry of AcrB. Science (80-.). 1295, 1295–1299. [DOI] [PubMed] [Google Scholar]
- Husain F.; Bikhchandani M.; Nikaido H. (2011) Vestibules are part of the substrate path in the multidrug efflux transporter AcrB of Escherichia coli. J. Bacteriol. 193, 5847–5849. 10.1128/JB.05759-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zechini B.; Versace I. (2009) Inhibitors of multidrug resistant efflux systems in bacteria. Recent Pat. Anti-Infect. Drug Discovery 4, 37–50. 10.2174/157489109787236256. [DOI] [PubMed] [Google Scholar]
- Fernández L.; Hancock R. E. W. (2012) Adaptive and mutational resistance: Role of porins and efflux pumps in drug resistance. Clin. Microbiol. Rev. 25, 661–681. 10.1128/CMR.00043-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nikaido H.; Pagès J. M. (2012) Broad-specificity efflux pumps and their role in multidrug resistance of Gram-negative bacteria. FEMS Microbiol. Rev. 36, 340–363. 10.1111/j.1574-6976.2011.00290.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blair J. M. A.; Webber M. A.; Baylay A. J.; Ogbolu D. O.; Piddock L. J. V. (2015) Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 13, 42–51. 10.1038/nrmicro3380. [DOI] [PubMed] [Google Scholar]
- Blair J. M. A.; Webber M. A.; Baylay A. J.; Ogbolu D. O.; Piddock L. J. V. (2015) Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 13, 42–51. 10.1038/nrmicro3380. [DOI] [PubMed] [Google Scholar]
- Delmar J. A.; Su C.-C.; Yu E. W. (2014) Bacterial multidrug efflux transporters. Annu. Rev. Biophys. 43, 93–117. 10.1146/annurev-biophys-051013-022855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun J.; Deng Z.; Yan A. (2014) Bacterial multidrug efflux pumps: Mechanisms, physiology and pharmacological exploitations. Biochem. Biophys. Res. Commun. 453, 254–267. 10.1016/j.bbrc.2014.05.090. [DOI] [PubMed] [Google Scholar]
- Turlin E.; Heuck G.; Simões Brandão M. I.; Szili N.; Mellin J. R.; Lange N.; Wandersman C. (2014) Protoporphyrin (PPIX) efflux by the MacAB-TolC pump in Escherichia coli. MicrobiologyOpen 3, 849–859. 10.1002/mbo3.203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tal N.; Schuldiner S. (2009) A coordinated network of transporters with overlapping specificities provides a robust survival strategy. Proc. Natl. Acad. Sci. U. S. A. 106, 9051–9056. 10.1073/pnas.0902400106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nies D. H. (2003) Efflux-mediated heavy metal resistance in prokaryotes. FEMS Microbiol. Rev. 27, 313–339. 10.1016/S0168-6445(03)00048-2. [DOI] [PubMed] [Google Scholar]
- Dreier J.; Ruggerone P. (2015) Interaction of antibacterial compounds with RND efflux pumps in Pseudomonas aeruginosa. Front. Microbiol. 6, 660. 10.3389/fmicb.2015.00660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lister P. D.; Wolter D. J.; Hanson N. D. (2009) Antibacterial-resistant Pseudomonas aeruginosa: Clinical impact and complex regulation of chromosomally encoded resistance mechanisms. Clin. Microbiol. Rev. 22, 582–610. 10.1128/CMR.00040-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morita Y.; Tomida J.; Kawamura Y. (2012) MexXY multidrug efflux system of Pseudomonas aeruginosa. Front. Microbiol. 3, 408. 10.3389/fmicb.2012.00408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hassan M.-T.; van der Lelie D.; Springael D.; Römling U.; Ahmed N.; Mergeay M. (1999) Identification of a gene cluster, czr, involved in cadmium and zinc resistance in Pseudomonas aeruginosa. Gene 238, 417–425. 10.1016/S0378-1119(99)00349-2. [DOI] [PubMed] [Google Scholar]
- Chuanchuen R.; Narasaki C. T.; Schweizer H. P. (2002) The MexJK efflux pump of Pseudomonas aeruginosa requires OprM for antibiotic efflux but not for efflux of Triclosan. J. Bacteriol. 184, 5036–5044. 10.1128/JB.184.18.5036-5044.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saito K.; Eda S.; Maseda H.; Nakae T. (2001) Molecular mechanism of MexR-mediated regulation of MexAB-OprM efflux pump expression in Pseudomonas aeruginosa. FEMS Microbiol. Lett. 195, 23–28. 10.1111/j.1574-6968.2001.tb10492.x. [DOI] [PubMed] [Google Scholar]
- Lim D.; Poole K.; Strynadka N. C. J. (2002) Crystal structure of the MexR repressor of the mexRAB-oprM multidrug efflux operon of Pseudomonas aeruginosa. J. Biol. Chem. 277, 29253–29259. 10.1074/jbc.M111381200. [DOI] [PubMed] [Google Scholar]
- Westbrock-Wadman S.; Sherman D. R.; Hickey M. J.; Coulter S. N.; Zhu Y. Q.; Warrener P.; Nguyen L. Y.; Shawar R. M.; Folger K. R.; Stover C. K. (1999) Characterization of a Pseudomonas aeruginosa efflux pump contributing to aminoglycoside impermeability. Antimicrob. Agents Chemother. 43, 2975–2983. 10.1128/AAC.43.12.2975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okazaki T.; Hirai K. (1992) Cloning and nucleotide sequence of the Pseudomonas aeruginosa nfxB gene, conferring resistance to new quinolones. FEMS Microbiol. Lett. 76, 197–202. 10.1111/j.1574-6968.1992.tb05462.x. [DOI] [PubMed] [Google Scholar]
- Kohler T.; van Delden C.; Curty L. K.; Hamzehpour M. M.; Pechere J.-C. (2001) Overexpression of the MexEF-OprN multidrug efflux system affects cell-to-cell signaling in Pseudomonas aeruginosa. J. Bacteriol. 183, 5213–5222. 10.1128/JB.183.18.5213-5222.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aendekerk S.; Ghysels B.; Cornelis P.; Baysse C. (2002) Characterization of a new efflux pump, MexGHI-OpmD, from Pseudomonas aeruginosa that confers resistance to vanadium. Microbiology 148, 2371–2381. 10.1099/00221287-148-8-2371. [DOI] [PubMed] [Google Scholar]
- Evans K.; Passador L.; Srikumar R.; Tsang E.; Nezezon J.; Poole K. (1998) Influence of the MexAB-OprM multidrug efflux system on quorum sensing in Pseudomonas aeruginosa. J. Bacteriol. 180, 5443–5447. 10.1128/JB.180.20.5443-5447.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayashi K.; Fukushima A.; Hayashi-Nishino M.; Nishino K. (2014) Effect of methylglyoxal on multidrug-resistant Pseudomonas aeruginosa. Front. Microbiol. 5, 1–6. 10.3389/fmicb.2014.00180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Watkins W. J.; Landaverry Y.; Léger R.; Litman R.; Renau T. E.; Williams N.; Yen R.; Zhang J. Z.; Chamberland S.; Madsen D.; Griffith D.; Tembe V.; Huie K.; Dudley M. N. (2003) The relationship between physicochemical properties, in vitro activity and pharmacokinetic profiles of analogues of diamine-containing efflux pump inhibitors. Bioorg. Med. Chem. Lett. 13, 4241–4244. 10.1016/j.bmcl.2003.07.030. [DOI] [PubMed] [Google Scholar]
- Kriengkauykiat J.; Porter E.; Lomovskaya O.; Wong-Beringer A. (2005) Use of an efflux pump inhibitor to determine the prevalence of efflux pump-mediated fluoroquinolone resistance and multidrug resistance in Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 49, 565–570. 10.1128/AAC.49.2.565-570.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takatsuka Y.; Chen C.; Nikaido H. (2010) Mechanism of recognition of compounds of diverse structures by the multidrug efflux pump AcrB of Escherichia coli. Proc. Natl. Acad. Sci. U. S. A. 107, 6559–6565. 10.1073/pnas.1001460107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coyne S.; Rosenfeld N.; Lambert T.; Courvalin P.; Périchon B. (2010) Overexpression of resistance-nodulation-cell division pump AdeFGH confers multidrug resistance in Acinetobacter baumannii. Antimicrob. Agents Chemother. 54, 4389–4393. 10.1128/AAC.00155-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piddock L. J. V., Garvey M. I., Rahman M. M., and Gibbons S. (2010) Natural and synthetic compounds such as trimethoprim behave as inhibitors of efflux in Gram-negative bacteria J. Antimicrob. Chemother. 65, 1215–1223. [DOI] [PubMed] [Google Scholar]
- Amin I.; Richmond G. E.; Sen P.; Koh T.; Piddock L. J.; Chua K. (2013) A method for generating marker-less gene deletions in multidrug-resistant Acinetobacter baumannii. BMC Microbiol. 13, 158. 10.1186/1471-2180-13-158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Podnecky N. L.; Wuthiekanun V.; Peacock S. J.; Schweizer H. P. (2013) The BpeEF-OprC efflux pump is responsible for widespread trimethoprim resistance in clinical and environmental Burkholderia pseudomallei isolates. Antimicrob. Agents Chemother. 57, 4381–4386. 10.1128/AAC.00660-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hassan K. A.; Elbourne L. D. H.; Li L.; Gamage H. K. A. H.; Liu Q.; Jackson S. M.; Sharples D.; Kolstø A. B.; Henderson P. J. F.; Paulsen I. T. (2015) An ace up their sleeve: A transcriptomic approach exposes the AceI efflux protein of Acinetobacter baumannii and reveals the drug efflux potential hidden in many microbial pathogens. Front. Microbiol. 6, 1–6. 10.3389/fmicb.2015.00333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hassan K. A.; Liu Q.; Elbourne L. D. H.; Ahmad I.; Sharples D.; Naidu V.; Chan C. L.; Li L.; Harborne S. P. D.; Pokhrel A.; Postis V. L. G.; Goldman A.; Henderson P. J. F.; Paulsen I. T. (2018) Pacing across the membrane: the novel PACE family of efflux pumps is widespread in Gram-negative pathogens. Res. Microbiol. 169, 450–454. 10.1016/j.resmic.2018.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hancock R. E. W. (1998) Resistance mechanisms in Pseudomonas aeruginosa and other nonfermentative Gram-negative bacteria. Clin. Infect. Dis. 27, S93–S99. 10.1086/514909. [DOI] [PubMed] [Google Scholar]
- Fischbach M. A.; Walsh C. T. (2009) Antibiotics For emerging pathogens. Science 325, 1089–1093. 10.1126/science.1176667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nikaido H. (1994) Prevention of drug acces to bacterial target: Permeability barriers and active efflux. Science 264, 382–388. 10.1126/science.8153625. [DOI] [PubMed] [Google Scholar]
- Vaara M. (1993) Outer membrane permeability barrier to azithromycin, clarithromycin, and roxithromycin in gram-negative enteric bacteria. Antimicrob. Agents Chemother. 37, 354–356. 10.1128/AAC.37.2.354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Charrel R. N.; Pages J. M.; De Micco P.; Mallea M. (1996) Prevalence of outer membrane porin alteration in B-lactam-antibiotic- resistant Enterobacter aerogenes. Antimicrob. Agents Chemother. 40, 2854–2858. 10.1128/AAC.40.12.2854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martínez-Martínez L.; Hernández-Allés S.; Albertí S.; Tomás J. M.; Benedi V. J.; Jacoby G. A. (1996) In vivo selection of porin-deficient mutants of Klebsiella pneumoniae with increased resistance to cefoxitin and expanded-spectrum cephalosporins. Antimicrob. Agents Chemother. 40, 342–348. 10.1128/AAC.40.2.342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hancock R. E. W. (1997) The bacterial outer membrane as a drug barrier. Trends Microbiol. 5, 37–42. 10.1016/S0966-842X(97)81773-8. [DOI] [PubMed] [Google Scholar]
- Donlan R. M. (2000) Role of Biofilms in Antimicrobial Resistance. ASAIO J. 46, S47–S52. 10.1097/00002480-200011000-00037. [DOI] [PubMed] [Google Scholar]
- Wistrand-Yuen E.; Knopp M.; Hjort K.; Koskiniemi S.; Berg O. G.; Andersson D. I. (2018) Evolution of high-level resistance during low-level antibiotic exposure. Nat. Commun. 9, 1599. 10.1038/s41467-018-04059-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gullberg E.; Cao S.; Berg O. G.; Ilbäck C.; Sandegren L.; Hughes D.; Andersson D. I.; Lipsitch M. (2011) Selection of resistant bacteria at very low antibiotic concentrations. PLoS Pathog. 7, e1002158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffman R. M.; Becket E.; Lee C.; Yang H.; Yuan J.; Miller J. H.; Liu A.; Maiz M.; Tamae C.; Fong A.; Wahba C.; Tran K. P.; Salih A.; Medrano L.; Lee K. (2011) Selective advantage of resistant strains at trace levels of antibiotics: a simple and ultrasensitive color test for detection of antibiotics and genotoxic agents. Antimicrob. Agents Chemother. 55, 1204–1210. 10.1128/AAC.01182-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gullberg E.; Albrecht L. M.; Karlsson C.; Sandegren L.; Andersson D. I. (2014) Selection of a multidrug resistance plasmid by sublethal levels of antibiotics and heavy metals. mBio 5, e01918 10.1128/mBio.01918-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ozaki M.; Mizushima S.; Nomura M. (1969) Identification and functional characterization of the protein controlled by the streptomycin-resistant locus in E. coli. Nature 222, 333–339. 10.1038/222333a0. [DOI] [PubMed] [Google Scholar]
- Funatsu G.; Wittmann H. G. (1972) Ribosomal proteins. Location of amino-acid replacements in protein S12 isolated from Escherichia coli mutants resistant to streptomycin. J. Mol. Biol. 68, 547–550. 10.1016/0022-2836(72)90108-8. [DOI] [PubMed] [Google Scholar]
- Gill A. E.; Amyes S. G. B. (2004) The contribution of a novel ribosomal S12 mutation to aminoglycoside resistance of Escherichia coli mutants. J. Chemother. 16, 347–349. 10.1179/joc.2004.16.4.347. [DOI] [PubMed] [Google Scholar]
- Spagnolo F.; Rinaldi C.; Sajorda D. R.; Dykhuizen D. E. (2016) Evolution of resistance to continuously increasing streptomycin concentrations in populations of Escherichia coli. Antimicrob. Agents Chemother. 60, 1336–42. 10.1128/AAC.01359-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olkkola S.; Juntunen P.; Heiska H.; Hyytiäinen H.; Hänninen M. L. (2010) Mutations in the rpsL gene are involved in streptomycin resistance in Campylobacter coli. Microb. Drug Resist. 16, 105–110. 10.1089/mdr.2009.0128. [DOI] [PubMed] [Google Scholar]
- Oz T.; Guvenek A.; Yildiz S.; Karaboga E.; Tamer Y. T.; Mumcuyan N.; Ozan V. B.; Senturk G. H.; Cokol M.; Yeh P.; Toprak E. (2014) Strength of selection pressure is an important parameter contributing to the complexity of antibiotic resistance evolution. Mol. Biol. Evol. 31, 2387–2401. 10.1093/molbev/msu191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lazar V.; Pal Singh G.; Spohn R.; Nagy I.; Horvath B.; Hrtyan M.; Busa-Fekete R.; Bogos B.; Mehi O.; Csorgo B.; Posfai G.; Fekete G.; Szappanos B.; Kegl B.; Papp B.; Pal C. (2013) Bacterial evolution of antibiotic hypersensitivity. Mol. Syst. Biol. 9, 700. 10.1038/msb.2013.57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bryan L. E.; Van Den Elzen H. M. (1976) Streptomycin accumulation in susceptible and resistant strains of Escherichia coli and Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 9, 928–938. 10.1128/AAC.9.6.928. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Campbell B. D.; Kadner R. J. (1980) Relation of aerobiosis and ionic strength to the uptake of dihydrostreptomycin in Escherichia coli. Biochim. Biophys. Acta, Bioenerg. 593, 1–10. 10.1016/0005-2728(80)90002-X. [DOI] [PubMed] [Google Scholar]
- Bryan L. E.; Kwan S. (1983) Roles of ribosomal binding, membrane potential, and electron transport in bacterial upake of streptomycin and gentamicin. Antimicrob. Agents Chemother. 23, 835–845. 10.1128/AAC.23.6.835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taber H. W.; Mueller J. P.; Miller P. F.; Arrow A. S. (1987) Bacterial uptake of aminoglycoside antibiotics. Microbiol. Rev. 51, 439–457. 10.1128/MMBR.51.4.439-457.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rhoads D. B.; Epstein W. (1977) Energy coupling to Net K+ transport in Escherichia coli K-12. J. Biol. Chem. 252, 1394–1401. [PubMed] [Google Scholar]
- Ratzke C.; Denk J.; Gore J. (2018) Ecological suicide in microbes. Nat. Ecol. Evol. 2, 867–872. 10.1038/s41559-018-0535-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis K. (2003) Programmed death in bacteria. Microbiol. Mol. Biol. Rev. 64, 503–514. 10.1128/MMBR.64.3.503-514.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mutschler H.; Gebhardt M.; Shoeman R. L.; Meinhart A. (2011) A novel mechanism of programmed cell death in bacteria by toxin-antitoxin systems corrupts peptidoglycan synthesis. PLoS Biol. 9, 1001036. 10.1371/journal.pbio.1001033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sedwick C. (2011) PezT: A Bacterial Suicide Gene. PLoS Biol. 9, e1001036 10.1371/journal.pbio.1001036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rice A.; Liu Y.; Michaelis M. L.; Himes R. H.; Georg G. I.; Audus K. L. (2005) Chemical modification of paclitaxel (Taxol) reduces P-Glycoprotein interactions and increases permeation across the blood-brain barrier in vitro and in situ. J. Med. Chem. 48, 832–838. 10.1021/jm040114b. [DOI] [PubMed] [Google Scholar]
- Ashburn T. T.; Thor K. B. (2004) Drug repositioning: Identifying and developing new uses for existing drugs. Nat. Rev. Drug Discovery 3, 673–683. 10.1038/nrd1468. [DOI] [PubMed] [Google Scholar]
- Borody T. J.; Khoruts A. (2012) Fecal microbiota transplantation and emerging applications. Nat. Rev. Gastroenterol. Hepatol. 9, 88–96. 10.1038/nrgastro.2011.244. [DOI] [PubMed] [Google Scholar]
- Floch M. H. (2012) The Power of Poop. J. Clin. Gastroenterol. 46, 625–626. 10.1097/MCG.0b013e3182667a93. [DOI] [PubMed] [Google Scholar]
- Molin G.; Jeppsson B.; Johansson M.-L.; Ahrné S.; Nobaek S.; Ståhl M.; Bengmark S. (1993) Numerical taxonomy of Lactobacillus spp. associated with healthy and diseased mucosa of the human intestines. J. Appl. Bacteriol. 74, 314–323. 10.1111/j.1365-2672.1993.tb03031.x. [DOI] [PubMed] [Google Scholar]
- Bengmark S. (1996) Econutrition and health maintenance - A new concept to prevent GI inflammation, ulceration and sepsis. Clin. Nutr. 15, 1–10. 10.1016/S0261-5614(96)80253-6. [DOI] [PubMed] [Google Scholar]
- Bengmark S. (2000) Colonic food: pre- and probiotics. Am. J. Gastroenterol. 95, S5–S7. 10.1016/S0002-9270(99)00807-2. [DOI] [PubMed] [Google Scholar]
- Adlerberth I; Ahrne S; Johansson M L; Molin G; Hanson L A; Wold A E (1996) A mannose-specific adherence mechanism in Lactobacillus plantarum conferring binding to the human colonic cell line HT-29. Appl. Environ. Microbiol. 62, 2244–2251. 10.1128/AEM.62.7.2244-2251.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kassam Z.; Lee C. H.; Yuan Y.; Hunt R. H. (2013) Fecal microbiota transplantation for Clostridium difficile infection: Systematic review and meta-analysis. Am. J. Gastroenterol. 108, 500–508. 10.1038/ajg.2013.59. [DOI] [PubMed] [Google Scholar]
- Bakken J. S.; Borody T.; Brandt L. J.; Brill J. V.; Demarco D. C.; Franzos M. A.; Kelly C.; Khoruts A.; Louie T.; Martinelli L. P.; Moore T. A.; Russell G.; Surawicz C. (2011) Treating clostridium difficile infection with fecal microbiota transplantation. Clin. Gastroenterol. Hepatol. 9, 1044–1049. 10.1016/j.cgh.2011.08.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamada N.; Chen G. Y.; Inohara N.; Núñez G. (2013) Control of pathogens and pathobionts by the gut microbiota. Nat. Immunol. 14, 685–690. 10.1038/ni.2608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ananthakrishnan A. N. (2011) Clostridium difficile infection: Epidemiology, risk factors and management. Nat. Rev. Gastroenterol. Hepatol. 8, 17–26. 10.1038/nrgastro.2010.190. [DOI] [PubMed] [Google Scholar]
- Salminen S.; Ouwehand A.; Benno Y.; Lee Y. K. (1999) Probiotics: how should they be defined?. Trends Food Sci. Technol. 10, 107–110. 10.1016/S0924-2244(99)00027-8. [DOI] [Google Scholar]
- Ouwehand A. C., Salminen S., and Isolauri E. (2002) Probiotics: an overview of beneficial effects, in Lactic Acid Bacteria: Genetics, Metabolism and Applications, pp 279–289, Springer Netherlands, Dordrecht. [PubMed] [Google Scholar]
- Isolauri E.; Salminen S.; Ouwehand A. C. (2004) Probiotics. Best Pract. Res. Clin. Gastroenterol. 18, 299–313. 10.1016/j.bpg.2003.10.006. [DOI] [PubMed] [Google Scholar]
- Alibert S.; Pagès J. (2007) Efflux pump inhibitors in bacteria. Expert Opin. Ther. Pat. 17, 883–888. 10.1517/13543776.17.8.883. [DOI] [Google Scholar]
- Askoura M.; Mattawa W.; Abujamel T.; Taher I. (2011) Efflux pump inhibitors (EPIs) as new antimicrobial agents against Pseudomonas aeruginosa. Libyan J. Med. 6, 5870. 10.3402/ljm.v6i0.5870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoshida K.; Nakayama K.; Kuru N.; Kobayashi S.; Ohtsuka M.; Takemura M.; Hoshino K.; Kanda H.; Zhang J. Z.; Lee V. J.; Watkins W. J. (2006) MexAB-OprM specific efflux pump inhibitors in Pseudomonas aeruginosa. Part 5: Carbon-substituted analogues at the C-2 position. Bioorg. Med. Chem. 14, 1993–2004. 10.1016/j.bmc.2005.10.043. [DOI] [PubMed] [Google Scholar]
- Lomovskaya O.; Bostian K. A. (2006) Practical applications and feasibility of efflux pump inhibitors in the clinic—A vision for applied use. Biochem. Pharmacol. 71, 910–918. 10.1016/j.bcp.2005.12.008. [DOI] [PubMed] [Google Scholar]
- Kourtesi C. (2013) Microbial efflux systems and inhibitors: Approaches to drug discovery and the challenge of clinical implementation. Open Microbiol. J. 7, 34–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Darzynkiewicz Z. M.; Green A. T.; Abdali N.; Hazel A.; Fulton R. L.; Kimball J.; Gryczynski Z.; Gumbart J. C.; Parks J. M.; Smith J. C.; Zgurskaya H. I. (2019) Identification of binding sites for efflux pump inhibitors of the AcrAB-TolC component AcrA. Biophys. J. 116, 648–658. 10.1016/j.bpj.2019.01.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cruz R. M. D.; Zelli R.; Benshain S.; Cruz R. M. D.; Siqueira-Junior J. P.; Decout J.-L.; Mingeot-Leclercq M.-P.; Mendonca-Junior F. J. B. (2020) Synthesis and evaluation of new 2-aminothiophene derivatives as Staphylococcus aureus efflux pump inhibitors. ChemMedChem 15, 716. 10.1002/cmdc.201900688. [DOI] [PubMed] [Google Scholar]
- Fournier dit Chabert J.; Marquez B.; Neville L.; Joucla L.; Broussous S.; Bouhours P.; David E.; Pellet-Rostaing S.; Marquet B.; Moreau N.; Lemaire M. (2007) Synthesis and evaluation of new arylbenzo[b]thiophene and diarylthiophene derivatives as inhibitors of the NorA multidrug transporter of Staphylococcus aureus. Bioorg. Med. Chem. 15, 4482–4497. 10.1016/j.bmc.2007.04.023. [DOI] [PubMed] [Google Scholar]
- Lomovskaya O.; Warren M. S.; Lee A.; Galazzo J.; Fronko R.; Lee M.; Blais J.; Cho D.; Chamberland S.; Renau T.; Leger R.; Hecker S.; Watkins W.; Hoshino K.; Ishida H.; Lee V. J. (2001) Identification and characterization of inhibitors of multidrug resistance efflux pumps in Pseudomonas aeruginosa: Novel agents for combination therapy. Antimicrob. Agents Chemother. 45, 105–116. 10.1128/AAC.45.1.105-116.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li X. Z.; Plésiat P.; Nikaido H. (2015) The challenge of efflux-mediated antibiotic resistance in Gram-negative bacteria. Clin. Microbiol. Rev. 28, 337–418. 10.1128/CMR.00117-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pagès J.-M.; Masi M.; Barbe J. (2005) Inhibitors of efflux pumps in Gram-negative bacteria. Trends Mol. Med. 11, 382–389. 10.1016/j.molmed.2005.06.006. [DOI] [PubMed] [Google Scholar]
- Shriram V.; Khare T.; Bhagwat R.; Shukla R.; Kumar V. (2018) Inhibiting bacterial drug efflux pumps via phyto-therapeutics to combat threatening antimicrobial resistance. Front. Microbiol. 9, 2990. 10.3389/fmicb.2018.02990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wong K.; Ma J.; Rothnie A.; Biggin P. C.; Kerr I. D. (2014) Towards understanding promiscuity in multidrug efflux pumps. Trends Biochem. Sci. 39, 8–16. 10.1016/j.tibs.2013.11.002. [DOI] [PubMed] [Google Scholar]
- Pagès J. M.; Amaral L. (2009) Mechanisms of drug efflux and strategies to combat them: Challenging the efflux pump of Gram-negative bacteria. Biochim. Biophys. Acta, Proteins Proteomics 1794, 826–833. 10.1016/j.bbapap.2008.12.011. [DOI] [PubMed] [Google Scholar]
- Baugh S.; Phillips C. R.; Ekanayaka A. S.; Piddock L. J. V.; Webber M. A. (2014) Inhibition of multidrug efflux as a strategy to prevent biofilm formation. J. Antimicrob. Chemother. 69, 673–681. 10.1093/jac/dkt420. [DOI] [PubMed] [Google Scholar]
- Langeveld W. T.; Veldhuizen E. J. A.; Burt S. A. (2014) Synergy between essential oil components and antibiotics: a review. Crit. Rev. Microbiol. 40, 76–94. 10.3109/1040841X.2013.763219. [DOI] [PubMed] [Google Scholar]
- Plésiat P.; Nikaido H. (1992) Outer membranes of Gram-negative bacteria are permeable to steroid probes. Mol. Microbiol. 6, 1323–1333. 10.1111/j.1365-2958.1992.tb00853.x. [DOI] [PubMed] [Google Scholar]
- Gill A. O.; Holley R. A. (2006) Disruption of Escherichia coli, Listeria monocytogenes and Lactobacillus sakei cellular membranes by plant oil aromatics. Int. J. Food Microbiol. 108, 1–9. 10.1016/j.ijfoodmicro.2005.10.009. [DOI] [PubMed] [Google Scholar]
- Gustafson L.; Chew M.; Bell W. (1998) Effects of tea tree oil on Escherichia coli. Lett. Appl. Microbiol. 26, 194–198. 10.1046/j.1472-765X.1998.00317.x. [DOI] [PubMed] [Google Scholar]
- Juven B. J.; Kanner J.; Schved F.; Weisslowicz H. (1994) Factors that interact with the antibacterial action of thyme essential oil and its active constituents. J. Appl. Bacteriol. 76, 626–631. 10.1111/j.1365-2672.1994.tb01661.x. [DOI] [PubMed] [Google Scholar]
- Lambert R. J. W.; Skandamis P. N.; Coote P. J.; Nychas G.-J. E. (2001) A study of the minimum inhibitory concentration and mode of action of oregano essential oil, thymol and carvacrol. J. Appl. Microbiol. 91, 453–462. 10.1046/j.1365-2672.2001.01428.x. [DOI] [PubMed] [Google Scholar]
- Burt S. (2004) Essential oils: their antibacterial properties and potential applications in foods-a review. Int. J. Food Microbiol. 94, 223–253. 10.1016/j.ijfoodmicro.2004.03.022. [DOI] [PubMed] [Google Scholar]
- Ultee A.; Smid E. J. (2001) Influence of carvacrol on growth and toxin production by Bacillus cereus. Int. J. Food Microbiol. 64, 373–378. 10.1016/S0168-1605(00)00480-3. [DOI] [PubMed] [Google Scholar]
- Tassou C.; Koutsoumanis K.; Nychas G.-J. E. (2000) Inhibition of Salmonella enteritidis and Staphylococcus aureus in nutrient broth by mint essential oil. Food Res. Int. 33, 273–280. 10.1016/S0963-9969(00)00047-8. [DOI] [Google Scholar]
- Burt S. A.; Reinders R. D. (2003) Antibacterial activity of selected plant essential oils against Escherichia coli O157:H7. Lett. Appl. Microbiol. 36, 162–167. 10.1046/j.1472-765X.2003.01285.x. [DOI] [PubMed] [Google Scholar]
- Burt S. A.; Vlielander R.; Haagsman H. P.; Veldhuizen E. J. A. (2005) Increase in activity of essential oil components carvacrol and thymol against Escherichia coli O157:H7 by addition of food stabilizers. J. Food Prot. 68, 919–926. 10.4315/0362-028X-68.5.919. [DOI] [PubMed] [Google Scholar]
- Burt S. (2004) Essential oils: their antibacterial properties and potential applications in foods—a review. Int. J. Food Microbiol. 94, 223–253. 10.1016/j.ijfoodmicro.2004.03.022. [DOI] [PubMed] [Google Scholar]
- Woehrlin F.; Fry H.; Abraham K.; Preiss-Weigert A. (2010) Quantification of flavoring constituents in Cinnamon: High variation of Coumarin in Cassia Bark from the German retail market and in authentic samples from Indonesia. J. Agric. Food Chem. 58, 10568–10575. 10.1021/jf102112p. [DOI] [PubMed] [Google Scholar]
- Bush K.; Courvalin P.; Dantas G.; Davies J.; Eisenstein B.; Huovinen P.; Jacoby G. A.; Kishony R.; Kreiswirth B. N.; Kutter E.; Lerner S. A.; Levy S.; Lewis K.; Lomovskaya O.; Miller J. H.; Mobashery S.; Piddock L. J. V.; Projan S.; Thomas C. M.; Tomasz A.; Tulkens P. M.; Walsh T. R.; Watson J. D.; Witkowski J.; Witte W.; Wright G.; Yeh P.; Zgurskaya H. I. (2011) Tackling antibiotic resistance. Nat. Rev. Microbiol. 9, 894–896. 10.1038/nrmicro2693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ejim L.; Farha M. A.; Falconer S. B.; Wildenhain J.; Coombes B. K.; Tyers M.; Brown E. D.; Wright G. D. (2011) Combinations of antibiotics and nonantibiotic drugs enhance antimicrobial efficacy. Nat. Chem. Biol. 7, 348–350. 10.1038/nchembio.559. [DOI] [PubMed] [Google Scholar]
- Rai M.; Paralikar P.; Jogee P.; Agarkar G.; Ingle A. P.; Derita M.; Zacchino S. (2017) Synergistic antimicrobial potential of essential oils in combination with nanoparticles: Emerging trends and future perspectives. Int. J. Pharm. 519, 67–78. 10.1016/j.ijpharm.2017.01.013. [DOI] [PubMed] [Google Scholar]
- Yap P. S. X.; Lim S. H. E.; Hu C. P.; Yiap B. C. (2013) Combination of essential oils and antibiotics reduce antibiotic resistance in plasmid-conferred multidrug resistant bacteria. Phytomedicine 20, 710–713. 10.1016/j.phymed.2013.02.013. [DOI] [PubMed] [Google Scholar]
- Lahmar A.; Bedoui A.; Mokdad-Bzeouich I.; Dhaouifi Z.; Kalboussi Z.; Cheraif I.; Ghedira K.; Chekir-Ghedira L. (2017) Reversal of resistance in bacteria underlies synergistic effect of essential oils with conventional antibiotics. Microb. Pathog. 106, 50–59. 10.1016/j.micpath.2016.10.018. [DOI] [PubMed] [Google Scholar]
- Moon S.-E.; Kim H.-Y.; Cha J.-D. (2011) Synergistic effect between clove oil and its major compounds and antibiotics against oral bacteria. Arch. Oral Biol. 56, 907–916. 10.1016/j.archoralbio.2011.02.005. [DOI] [PubMed] [Google Scholar]
- Zhang D.; Hu H.; Rao Q.; Zhao Z. (2011) Synergistic effects and physiological responses of selected bacterial isolates from animal feed to four natural antimicrobials and two antibiotics. Foodborne Pathog. Dis. 8, 1055–1062. 10.1089/fpd.2010.0817. [DOI] [PubMed] [Google Scholar]
- Coutinho H. D. M.; Costa J. G. M.; Falcão-Silva V. S.; Siqueira-Júnior J. P.; Lima E. O. (2010) Effect of Momordica charantia L. in the resistance to aminoglycosides in methicilin-resistant Staphylococcus aureus. Comp. Immunol. Microbiol. Infect. Dis. 33, 467–471. 10.1016/j.cimid.2009.08.001. [DOI] [PubMed] [Google Scholar]
- Braga L. C.; Leite A. A.; Xavier K. G.; Takahashi J. A.; Bemquerer M. P.; Chartone-Souza E.; Nascimento A. M. (2005) Synergic interaction between pomegranate extract and antibiotics against Staphylococcus aureus. Can. J. Microbiol. 51, 541–547. 10.1139/w05-022. [DOI] [PubMed] [Google Scholar]
- Oluwatuyi M.; Kaatz G. W.; Gibbons S. (2004) Antibacterial and resistance modifying activity of Rosmarinus officinalis. Phytochemistry 65, 3249–3254. 10.1016/j.phytochem.2004.10.009. [DOI] [PubMed] [Google Scholar]
- Papadopoulos C. J.; Carson C. F.; Chang B. J.; Riley T. V. (2008) Role of the MexAB-OprM efflux pump of Pseudomonas aeruginosa in tolerance to Tea tree (Melaleuca alternifolia) oil and its monoterpene components Terpinen-4-ol, 1,8-Cineole, and -Terpineol. Appl. Environ. Microbiol. 74, 1932–1935. 10.1128/AEM.02334-07. [DOI] [PMC free article] [PubMed] [Google Scholar]