Abstract
Electrochemical impedimetric biosensors (EIBs) have a simple structure and can be used to rapidly and sensitively detect and measure hazards in food. EIBs detect and measure target molecules by transducing biochemical reactions on their surface to electrical signal outputs responding to a sinusoidal electrical signal input. Due to their structural simplicity and analytical sensitivity, EIBs are regarded as the most potent method of food hazard monitoring that can be implemented in the food supply chain. This paper discusses the theoretical background, structure, and construction of EIB and its applications in food safety.
Keywords: Electrochemical impedance spectroscopy, Biosensor, Food safety, Pathogen, Mycotoxin
Introduction
Food safety is a key public health issue that begins with monitoring food hazards in including pathogens and chemical contaminants, and is achieved by eliminating or reducing food hazards to acceptable levels. As food hazards can enter the food supply chain at any point from farm to table, monitoring should be implemented at all points. Therefore, methods for monitoring food safety that can be easily implemented within the food supply chain are required. There have been marked advances in food safety monitoring technology over the past two decades, and various monitoring methods have been developed and are currently in use. In particular, electrochemical impedimetric biosensors (EIBs) have attracted a great deal of attention from food safety scientists and administrators. EIBs directly detect and measure target molecules with no sample preparation requirement, and can therefore be used for inline monitoring of hazards in the food supply chain. The sensitivity of EIBs for the detection and measurement of food hazards is comparable to or better than that of other biosensors and traditional methods (Ahmed et al., 2014; Bahadır and Sezgintürk, 2016; Malvano et al., 2019). EIBs can detect and measure food hazards in less than 1 h (Ahmed et al., 2014; Chai et al., 2010; Malvano et al., 2019). In combination with the Internet of Things (IoT), EIBs may evolve into real-time food safety monitoring systems (Durresi, 2016). Currently, the integration of EIBs into smart devices for food hazard detection has been intensively investigated (Huang et al., 2018; Rosati et al., 2019a). However, further research is required for incorporation of EIBs into the food supply chain. This review discusses the theoretical background, structure, and construction of EIBs, and their potential applications for food safety, to stimulate interest in their development for use in real-time inline food hazard monitoring.
Theoretical background of EIBs
EIBs probe their target molecules by measuring impedance, which is enhanced by the formation of antibody–antigen or ligand–receptor complexes on their surface. Electrochemical impedance is the amount of opposition that an electrochemical cell (e.g., the EIB) presents to the flow of an electrical current on application of a small-amplitude sinusoidal voltage. The sinusoidal voltage input (V) as a function of time (t) can be expressed using the maximum amplitude of voltage (V0) and radial frequency (ω; ω = 2πf, where f is the linear frequency, represented by the number of cycles per second), and is also expressed as a complex number in Eq. 1:
| 1 |
Current output (I) from EIB responding to the sinusoidal voltage input will be a sinusoid at the same ω, but shifted in phase (ϕ) and altered in terms of the maximum amplitude of the current (I0). Thus, I can be expressed as Eq. 2.
| 2 |
According to Ohm’s law, impedance (Z) as a function of ω is V divided by I and can be represented as a complex number. Based on Euler’s relationship, Z can be expressed as a polar and rectangular coordinate form of a complex number, as shown in Eq. 3. Z in rectangular coordinate form can be characterized as a real part (Zre) and imaginary part (Zim), referred to as resistance and reactance, respectively. Zim is enhanced due to ϕ and accounts for capacitance and inductance. However, the biological recognition elements and target molecules of EIBs, such as antibodies, antigens, receptors, DNAs, aptamers, etc., are not sufficiently electrochemically active to significantly alter the inductance (Rishpon and Buchner, 2005).
| 3 |
Electrolyte resistance (Rs), double-layer capacitance (Cdl), and charge-transfer resistance (Rct) at the electrode/electrolyte interface may be involved in the alteration of Z on application of a sample to EIBs. The electrolytes in a sample solution govern Rs. Rs is independent of the target molecules in the sample solution, and can be determined by measuring Zre of the sample solution at high f, from 0.1 to 10 MHz (Carminati et al., 2015; Itagaki et al., 2007; Manickam et al., 2012). Cdl depends on the thickness (d) of the electrical double layer (EDL) formed at the electrode/electrolyte interface, as well as the dielectric constant of the sample solution. The formation of antibody–antigen or ligand–receptor complexes on the EIB surface may alter the physicochemical characteristics of the interface between the EIB surface and sample solution, and may increase d in particular. If the effect of immunoreaction on the EIB surface on inductance is negligible, Cdl dominates Zim. Zim is linearly related to the inverse of Cdl (Eq. 4), and Cdl is inversely proportional to d (Eq. 5). Thus, the formation of antibody–antigen or ligand–receptor complexes on the EIB surface decreases Cdl with increasing d (Carminati et al., 2015; Prodromidis, 2010). Changes in Cdl that are specific to immunoreactions on the EIB surface can be identified by measuring Z at f from 10 to 1000 Hz (Carminati et al., 2015; Prodromidis, 2010).
| 4 |
| 5 |
where ε is the dielectric constant of the sample solution.
Rct accounts for the diffusion of electrolytes from the bulk solution to the EIB surface, which is expected, especially when redox reactions occur (Carminati et al., 2015; Prodromidis 2010). Redox reactions can be enhanced by introducing redox probes, such as ferricyanide, into the sample solution or coupling redox reporters, such as graphene oxide, gold nanoparticle, and titanium carbide, with EIBs (Carminati et al., 2015; Li et al., 2017; Liang et al., 2019; Lu et al., 2012). Redox reactions affect current flow and Rct (Carminati et al., 2015). With the formation of antibody–antigen, ligand–receptor, protein–aptamer, and DNA–DNA complexes on the EIB surface, d is increased and ions near the complexes are relocated, thereby altering Rct (Bard, 1980; Manickam et al., 2012; Prodromidis, 2010). In particular, Rct is altered more if the electrical potential of the EIB versus an additionally implemented reference electrode is maintained at a certain voltage (Bard, 1980; Park et al., 2018; Prodromidis. 2010). Rct is an electrical parameter consisting of Zre, and the changes therein caused by immunoreactions on the EIB surface are frequency-dependent. Rct can be characterized by Zre at f from 0.1 to 1.0 Hz (Carminati et al., 2015; Prodromidis, 2010). Consequently, electrical parameters, including Rs, Cdl, and Rct, can be characterized by electrochemical impedance spectroscopy (EIS) of an EIB over a wide f from 0.1 Hz to 10 MHz (Maalouf et al., 2007a; Radhakrishnan et al., 2014).
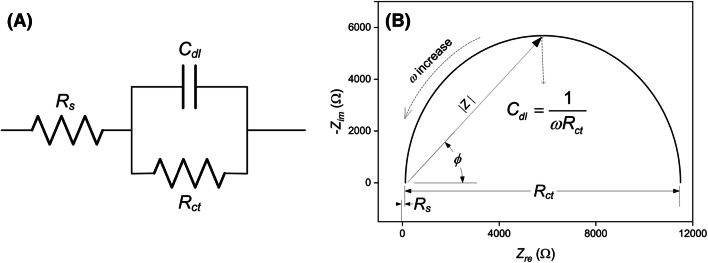
The electrochemical impedance spectrum can be presented using Nyquist plots (− Zim versus Zre) and Bode plots (Z, ϕ, Zre, and Zim versus f). The electrical parameters of a circuit model equivalent to the EIB system can be characterized using a Nyquist plot. A classical circuit model with an electrode/electrolyte interface is presented in Fig. 1A. The Nyquist plot of the equivalent circuit in Fig. 1A is presented in Fig. 1B. On the Nyquist plot, Z is presented as a vector of length |Z|. The angle between the Z vector and the axis of Zre is ϕ (Fig. 1B). The Z of the equivalent circuit in Fig. 1A can be expressed using Rs, Rct, and Cdl, and follows Eq. 6. With Eq. 6, the Zre and Zim can be expressed as Eqs. 7 and 8.
| 6 |
| 7 |
| 8 |
Fig. 1.
(A) A classical circuit model of the electrode/electrolyte interface. (B) Nyquist plot of the classical circuit model
As ω → 0 and ∞, limited forms of Zre can be obtained as shown in Eq. 9. Thus, Rct can be obtained by subtracting the minimum value of Zre (Zremin) from the maximum value of Zre (Zremax), as shown in Eq. 10.
| 9 |
| 10 |
The Nyquist plot of the equivalent circuit produces a semicircle with a radius of half Rct (Bard, 1980). Hence, the maximum value of − Zim (− Zimmax) is centered at Zre = Rs + Rct/2. Using Eq. 7, Cdl can be obtained with Eqs. 11 and 12.
| 11 |
| 12 |
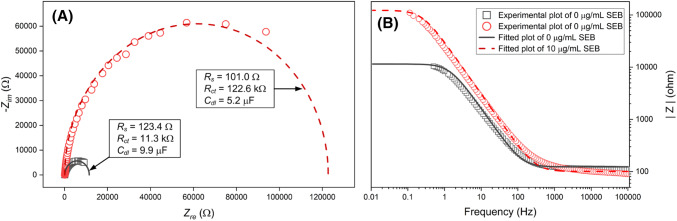
However, experimentally obtained Zre and − Zim often do not produce a complete semicircle in a Nyquist plot due to the nonuniform current distribution on the electrode surface (Cheng and Chen, 2013). The Nyquist plot obtained by EIS measurement of EIBs must frequently be fitted. Figure 2A shows Nyquist plots obtained experimentally from EIS of the EIB for Staphylococcus enterotoxin B (SEB), and Nyquist plots fitted mathematically using EIS Spectrum Analyzer software v1.0 (Bondarenko and Ragoisha, 2005). The Rs, Rct, and Cdl derived from the EIB for SEB were calculated based on the equivalent circuit presented in Fig. 1A. It is obvious that Rct and Cdl derived from the EIB for SEB increased and decreased with complexation of SEB with anti-SEB antibodies immobilized on the EIB surface (Fig. 1C). As the Cdl decreased, Zim also increased. Although a Nyquist plot is critical to characterize electrical parameters, Z, derived from an EIB, it is difficult to determine the dependence of the electrical parameters on the frequency. Bode plots provide frequency information, and are useful to determine the frequency range needed to obtain stable values of electrical parameters.
Fig. 2.
(A) Nyquist plots for an EIB for SEB, and mathematically fitted Nyquist plots. (B) Bode plots of |Z| versus f obtained from the EIB for SEB. EIB for SEB was developed using an anodic aluminum substrate and APTES. An anodic aluminum substrate with pores approximately 30 nm in diameter was treated with APTES. Anti-SEB was covalently immobilized on APES-SAMs deposited on the anodic aluminum substrate using glutaraldehyde. EIS of the EIB for SEB was performed at a biased potential of 0.1 V (vs. an Ag/AgCl reference electrode), in the absence or presence of 10 mg/mL SEB in 0.3% NaCl solution
Structure and construction of EIBs
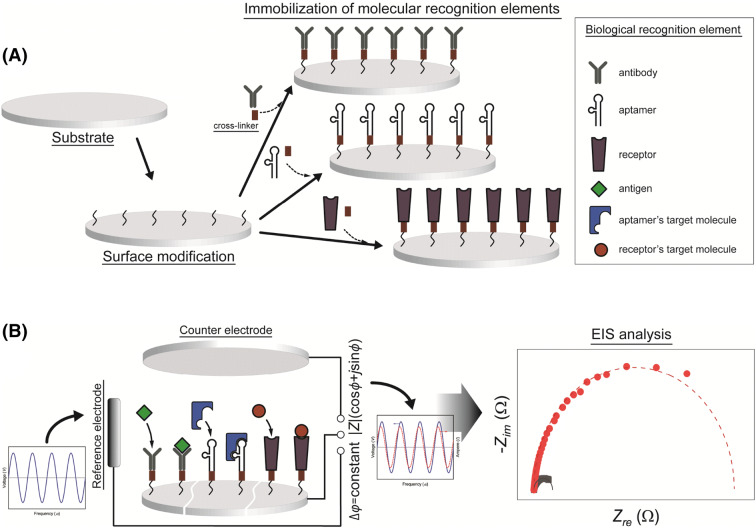
An EIB consists of a signal transducer, an electrically conductive electrode substrate, and biological recognition elements (Fig. 3A) (Leca-Bouvier and Blum, 2005). For EIBs, an EIS analyzer acts as a signal transducer. An electrode substrate mediates biological recognition (Fig. 3B). Mercury, platinum, graphite, gold, stainless steel, silicon, and aluminum are the most frequently used materials for the electrode substrate (Săndulescu et al., 2015). There have been a number of studies on the use of nanoporous metal oxides and orderly structured carbon composites as the electrode substrate, to increase the surface area and sensitivity of EIBs (Ali et al., 2014; Ania et al., 2018; Bonanni et al., 2012; Chai and Takhistov, 2012). Metal nanoparticles have been used to intensify the electrochemical signal outputs from EIBs (Derkus et al., 2014; Lin et al., 2019; Peng et al., 2006).
Fig. 3.
Schematic illustration of (A) the structure and construction, and (B) detection and measurement process, of EIB
Biological recognition elements include antibodies, aptamers, and receptors (Fig. 3A). To obtain impedimetric signal outputs specific for biological recognition at the EIB/sample solution interface, biological recognition elements should be immobilized intimately on the electrode substrate (Vashist et al., 2014). The electrode substrate may need to be chemically functionalized, and biological recognition elements can be chemically immobilized on the substrate by crosslinkers (Fig. 3A) (Nicosia and Huskens 2014; Vashist et al., 2014). Gold is reactive to thiols (Ron and Rubinstein, 1998). Thiol-based polymers, including peptides, proteins, and alkanethiols, can covalently bind to the gold surface and form self-assembled monolayers (SAMs) (Abad et al., 2006; Nicosia and Huskens 2014; Niu et al., 2012). Biological recognition elements can be immobilized by covalent binding with the SAMs of thiol-based polymers through the use of crosslinkers, such as protein A, protein G, and bifunctional amide compounds (Abad et al., 2006; Icoz et al., 2018). Metal oxides, such as silicon oxide and aluminum oxide, have hydroxyl groups on their surface and are reactive to silane compounds (Plueddemann, 1991). 3-Aminopropyltriethoxysilane (APTES) has been widely used to functionalize metal oxide surfaces (Chai et al., 2012a, b; Huy et al., 2011; Plueddemann 1991; Vashist et al., 2014). APTES bind electrostatically to metal oxides and form SAMs (Plueddemann, 1991). Antibodies and receptors can bind covalently to APTES with crosslinkers, such as glutaraldehyde (Fig. 3A) (Chai et al., 2012a, b; Vashist et al., 2014). APTES is also useful for functionalization of the surface of carbon composites (Luong et al., 2004; Zheng et al., 2013). Unlike the case of immobilization of proteins on the electrode substrate, DNA and aptamers must be conjugated with the thiol or amine group at the 3′ or 5′ end for immobilization on the electrode substrate (Lu et al., 2007; Paniel et al., 2013). Depending on the conjugated groups, DNA and aptamers can be immobilized directly on the gold surface or crosslinked with APTES-SAMs deposited on the electrode substrate (Keighley et al., 2008; Sauthier et al., 2002; Tam et al., 2009; Walsh et al., 2001; Wang et al., 2013).
EIBs for detection of food hazards
Food hazard detection methods should not only be simple and easy to operate, thus allowing onsite monitoring, but also sensitive and reliable to prevent the consumption of contaminated and deteriorated foods. EIBs can identify biochemical reactions of biological recognition elements with their target molecules at the EIB/sample interface. Furthermore, EIBs do not require additional sample preparation steps, and are therefore among the most useful analytical methods for onsite detection of food hazards. This article discusses research regarding the use of EIBs for the detection of major food poisoning bacteria and mycotoxins.
Food poisoning bacteria are the most dangerous food hazards, and pose a major threat to human health. A large number of studies on the detection of food poisoning bacteria in foods have been conducted (Bridier, 2018; Hoorfar, 2011). Various antibodies that bind directly to food poisoning bacteria, such as pathogenic Escherichia coli and Salmonella spp., are commercially available, and EIBs can serve as a universal platform for these pathogens. An EIB with anti-E. coli O157:H7 on a gold-coated electrode detected the presence of 7 CFU/mL of E. coli O157:H7 in a ferrous solution (Joung et al., 2012). An increase in Rct was observed as E. coli from a sample bound to anti-E. coli on the EIB, in proportion to the concentration of E. coli included in the sample (Joung et al., 2012; Maalouf et al., 2007b). The sensitivity of the EIB for E. coli O157:H7 was improved by attaching electron transferring mediators; the limit of detection (LOD) of the EIB was 3 CFU/mL (Malvano et al., 2018). Similar to the results of the EIB for E. coli, the binding of Salmonella spp. with anti-Salmonella immobilized on a gold electrode caused an increase in Rct (Mantzila et al., 2008; Pournaras et al., 2008). The EIB for Salmonella spp., constructed on a gold electrode using tyramine as a surface modifier, exhibited a LOD of 20 CFU/mL Salmonella spp. (Liu et al., 2018). Aptamers, as biological recognition elements of EIBs, have been investigated due to their high binding specificity and affinity to their target bacteria (Teng et al., 2016). Aptamer-based EIBs for E. coli O157:H7 and E. coli O111 showed a LOD at the level of 100 CFU/mL (Brosel-Oliu et al., 2018; Luo et al., 2012). An aptamer-based EIB for Salmonella Typhimurium, constructed on a gold electrode functionalized by conductive polymer, could detect the presence of this bacterium at 3 CFU/mL (Sheikhzadeh et al., 2016). Viable S. Typhimurium could be selectively measured with an EIB developed using aptamers with high affinity to viable S. Typhimurium but poor affinity to dead S. Typhimurium (Labib et al., 2012). The EIB for viable S. Typhimurium had a LOD of 600 CFU/mL S. Typhimurium (Labib et al., 2012).
Mycotoxins are poisonous substances produced by fungi (Omotayo et al., 2019) that can cause disease and death in humans, and are therefore under strict governmental regulation (European Commission, 2010; KFDA, 2020; US FDA, 2016). The development of EIBs for mycotoxins has focused on the detection of ochratoxin A and aflatoxins, due to their prevalence and toxicity (Malvano et al., 2019; Omotayo et al., 2019). An EIB with anti-ochratoxin A immobilized on an indium oxide electrode showed a linear response, in terms of Rct, to ochratoxin concentrations ranging from 1 to 10 ng/mL (Khan and Dhayal 2009). An EIB for ochratoxin A built on a gold electrode showed similar results to one based on an indium oxide electrode (Radi et al., 2009). The acceptable limit established for ochratoxin A in food products is 5 ng/g (Codex STAN 1995). The sensitivity of the EIBs described above was not sufficient to meet existing regulations established for ochratoxin A. An EIB that could measure ochratoxin A at concentrations in food products below 0.5 ng/g was reported (Tang et al., 2016). That EIB, based on competitive immunoreaction, had a reference ochratoxin A-immobilized carbon electrode and signal tags (anti-ochratoxin A-immobilized and manganese oxide-adsorbed graphene oxide nanosheets), and measured impedance; signal tags bound to the electrode could detect the presence of 0.055 pg/mL ochratoxin A (Tang et al., 2016).
Aflatoxins are a family of mycotoxins mainly produced by Aspergillus species (Dutton et al., 1985). Four major types of aflatoxins are found in food: aflatoxin B1, B2, G1, and G2 (Bennett and Klich, 2003). Aflatoxin B1 is the most common and toxic aflatoxin in food, but all aflatoxins are toxic and carcinogenic (Wakenell, 2016). Aflatoxin regulations are often based on the sum of aflatoxin B1, B2, G1, and G2, and the maximum permissible level of total aflatoxins in food established by the CODEX Alimentarius Commission is 15 ng/g (Codex STAN 1995). Antibodies specific to aflatoxin B1 are commercially available, and many antibodies developed using aflatoxin B1 show good cross-affinity to aflatoxin B2, G1, and G2 (Ertekin et al., 2016; Gathumbi et al., 2001). An EIB with anti-aflatoxin B1 immobilized on the carbon electrode, where carbon nanotubes were physically adsorbed, showed a linear increase in Rct with increasing level of aflatoxin B1 from 0.1 to 10 ng/mL (Yu et al., 2015). A highly sensitive EIB that could directly measure aflatoxin B1 was developed by immobilizing anti-aflatoxin B1 on carbon nanotubes covalently anchored on the gold electrode. The carbon nanotubes were covalently anchored on the surface of a gold electrode via formation of cysteine SAMs on the gold surface and subsequent activation of cysteine SAMs using 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC) and N-hydroxysuccinimide (NHS). The EIB showed a linear response, in terms of Rct, to aflatoxin B1 concentrations ranging from 0.1 to 20 pg/mL (Costa et al., 2017). The sensitivity of the EIB for aflatoxin B1 was improved by graphene oxide and conductive polymer (Wang et al., 2015). Graphene oxide was deposited on a carbon electrode and anti-aflatoxin B1 was cross-linked to the graphene oxide with conductive polymer. The EIB for aflatoxin B1 developed using graphene oxide and conductive polymer exhibited a significant increase in Rct even in the presence of 10 fg/mL aflatoxin B1. The EIB also showed a linear increase in Rct with increasing aflatoxin B1 concentration from 10 fg/mL to 10 pg/mL (Wang et al., 2015). A cost-effective, disposable but highly sensitive EIB for aflatoxin B1 was developed using a gold CD-trode (the gold layer used for recordable compact discs) (Foguel et al., 2016). Anti-aflatoxin B1 was immobilized covalently on a gold CD-trode by surface functionalization, using lipoic acid and subsequent EDC/NHS activation. The Rct from the EIB increased in proportion to the increase aflatoxin B1 concentration, from 1.56 to 31.2 ng/mL, and had a LOD of 0.11 ng/mL.
In conclusion, EIBs have a number of advantages over conventional and optical biosensors. Unlike optical-based biosensors, EIBs do not require excitation sources, filters, or lenses. EIBs can directly qualify and quantify their target molecules in food, and have comparable or better sensitivity than optical biosensors. The EIB is a versatile platform that can be modified to measure different food hazards through replacement of biological recognition elements. EIBs can be manufactured using consumer-grade inkjet printers (Rosati et al., 2019a, b). The EIB for a bacteriophage produced using an inkjet printer showed better sensitivity than a traditional method for bacteriophage detection (Rosati et al., 2019a). EIBs appear to be a sensitive and cost-effective means of food hazard detection suitable for mass production. With advances in mobile phone technology, there have been a number of studies concerned with integration of EIBs into smartphones (Huang et al., 2018; Rosati et al., 2019b). In the near future, EIBs are expected to be implemented throughout the food supply chain as for inline and real-time monitoring of food hazards.
Acknowledgements
This study was supported by a grant from the National Research Foundation of Korea (NRF-2019R1I1A3A01052210), funded by the Korean government.
List of symbols
- V
The sinusoidal voltage input
- V0
The maximum amplitude of V
- I
The current output
- I0
The maximum amplitude of I
- f
The linear frequency
- t
Time
- ω
The radial frequency
- ϕ
The phase shift of I
- Z
The impedance
- |Z|
The absolute value of Z
- Zre
The real part of Z
- Zim
The imaginary part of Z
- Zminre
The minimum value of Zre
- Zmaxre
The maximum value of Zre
- Rs
The electrolyte resistance
- Cdl
The double-layer capacitance
- Rct
The charge-transfer resistance
- ε
The dielectric constant
- d
The thickness of the electrical double layer
Footnotes
The original version of this article was revised due to a retrospective Open Access order.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Change history
9/5/2020
The article “Electrochemical impedimetric biosensors for food safety”, written by Changhoon Chai and Se-Wook Oh, was originally published Online First without Open Access.
References
- Abad JM, Pita M, Fernández VM. Immobilization of proteins on gold surfaces. In: Guisan JM (ed). Immobilization of Enzymes and Cells. Totowa: Humana Press, pp. 229-238 (2006)
- Ahmed A, Rushworth JV, Hirst NA, Millner PA. Biosensors for whole-cell bacterial detection. Clin. Microbiol. Rev. 2014;27:631–646. doi: 10.1128/CMR.00120-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ali MA, Srivastava S, Mondal K, Chavhan PM, Agrawal VV, John R, Sharma A, Malhotra BD. A surface functionalized nanoporous titania integrated microfluidic biochip. Nanoscale. 2014;6:13958–13969. doi: 10.1039/c4nr03791j. [DOI] [PubMed] [Google Scholar]
- Ania CO, Gomis-Berenguer A, Dentzer J, Vix-Guterl C. Nanoconfinement of glucose oxidase on mesoporous carbon electrodes with tunable pore sizes. J. Electroanal. Chem. 2018;808:372–379. [Google Scholar]
- Bahadır EB, Sezgintürk MK. A review on impedimetric biosensors. Artif. Cells Nanomed. Biotechnol. 2016;44:248–262. doi: 10.3109/21691401.2014.942456. [DOI] [PubMed] [Google Scholar]
- Bard AJ. Electrochemical methods : Fundamentals and applications. New York: Wiley; 1980. [Google Scholar]
- Bennett JW, Klich M. Mycotoxins. Clin. Microbiol. Rev. 2003;16:497–516. doi: 10.1128/CMR.16.3.497-516.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonanni A, Loo AH, Pumera M. Graphene for impedimetric biosensing. TrAC Trends Anal. Chem. 37: 12-21 (2012)
- Bondarenko A, Ragoisha G. Inverse problem in potentiodynamic electrochemical impedance. In: Pomerantsev AL (ed). Progress in Chemometrics Research. Nova Science Publishers: New York, pp. 89-102 (2005)
- Bridier A. Foodborne bacterial pathogens: Methods and protocols. New York: Springer; 2018. [Google Scholar]
- Brosel-Oliu S, Ferreira R, Uria N, Abramova N, Gargallo R, Muñoz-Pascual F-X, Bratov A. Novel impedimetric aptasensor for label-free detection of Escherichia coli O157:H7. Sensor. Actuat. B: Chem. 2018;255:2988–2995. [Google Scholar]
- Carminati M, Ferrari G, Bianchi D, Sampietro M. Impedance spectroscopy for biosensing: Circuits and applications. In: Sawan M (ed). Handbook of biochips: Integrated circuits and systems for biology and medicine. New York: Springer, pp. 1–24 (2015)
- Chai C, Lee J, Takhistov P. Direct detection of the biological toxin in acidic environment by electrochemical impedimetric immunosensor. Sensors. 2010;10:11414–11427. doi: 10.3390/s101211414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chai C, Lee J, Park J, Takhistov P. Antibody immobilization on a nanoporous aluminum surface for immunosensor development. Appl. Surf. Sci. 2012;263:195–201. [Google Scholar]
- Chai C, Takhistov P. Control of the lateral interactions of immobilized proteins using surface nanoporous-patterning. Appl. Surf. Sci. 2012;263:104–110. [Google Scholar]
- Cheng Q, Chen Z. The cause analysis of the incomplete semi-circle observed in high frequency region of EIS obtained from TEL-covered pure copper. Int. J. Electrochem. Sci. 2013;8:8282–8290. [Google Scholar]
- Codex STAN. STAN 193-1995. Codex General Standard for Contaminants and Toxins in Food and Feed, Codex Alimentarius Commission (1995)
- Costa MP, Frías IAM, Andrade CAS, Oliveira MDL. Impedimetric immunoassay for aflatoxin B1 using a cysteine modified gold electrode with covalently immobilized carbon nanotubes. Microchim. Acta. 2017;184:3205–3213. [Google Scholar]
- Derkus B, Cebesoy Emregul K, Mazi H, Emregul E, Yumak T, Sinag A. Protein A immunosensor for the detection of immunoglobulin G by impedance spectroscopy. Bioprocess Biosystems Eng. 2014;37:965–976. doi: 10.1007/s00449-013-1068-2. [DOI] [PubMed] [Google Scholar]
- Durresi M. (Bio)Sensor Integration with ICT tools for supplying chain management and traceability in agriculture. In: Scognamiglio V, Rea G, Arduini F, Palleschi G (eds). Comprehensive Analytical Chemistry. Elsevier, pp. 389-413 (2016)
- Dutton MF, Ehrlich K, Bennett JW. Biosynthetic relationship among aflatoxins B1, B2, M1, and M2. Appl. Environ. Microbiol. 1985;49:1392–1395. doi: 10.1128/aem.49.6.1392-1395.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ertekin Ö, Pirinçci ŞŞ, Öztürk S. Monoclonal IgA antibodies for aflatoxin immunoassays. Toxins. 2016;8:148. doi: 10.3390/toxins8050148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- European Commission. Commission Regulation (EC) No 165/2010 of 26 February 2010 amending Regulation (EC) No 1881/2006 setting maximum levels for certain contaminants in foodstuffs as regards aflatoxins. Off. J. Eur. Union: 8–12 (2010)
- Foguel MV, Furlan Giordano G, De Sylos CM, Carlos IZ, Pupim Ferreira AA, Benedetti AV, Yamanaka H. A low-cost label-free AFB1 impedimetric immunosensor based on functionalized CD-trodes. Chemosensors. 2016;4:17. [Google Scholar]
- Gathumbi JK, Usleber E, Märtlbauer E. Production of ultrasensitive antibodies against aflatoxin B1. Lett. Appl. Microbiol. 2001;32:349–351. doi: 10.1046/j.1472-765x.2001.00914.x. [DOI] [PubMed] [Google Scholar]
- Hoorfar J. Rapid detection, characterization, and enumeration of foodborne pathogens. American Society of Microbiology (2011) [DOI] [PubMed]
- Huang X, Xu D, Chen J, Liu J, Li Y, Song J, Ma X, Guo J. Smartphone-based analytical biosensors. Analyst. 2018;143:5339–5351. doi: 10.1039/c8an01269e. [DOI] [PubMed] [Google Scholar]
- Huy TQ, Hanh NTH, Van Chung P, Anh DD, Nga PT, Tuan MA. Characterization of immobilization methods of antiviral antibodies in serum for electrochemical biosensors. Appl. Surf. Sci. 2011;257:7090–7095. [Google Scholar]
- Icoz K, Soylu MC, Canikara Z, Unal E. Quartz-crystal microbalance measurements of CD19 antibody immobilization on gold surface and capturing B lymphoblast cells: Effect of surface functionalization. Electroanalysis. 2018;30:834–841. [Google Scholar]
- Itagaki M, Suzuki S, Shitanda I, Watanabe K. Electrochemical impedance and complex capacitance to interpret electrochemical capacitor. Electrochemistry. 2007;75:649–655. [Google Scholar]
- Joung C-K, Kim H-N, Im H-C, Kim H-Y, Oh M-H, Kim Y-R. Ultra-sensitive detection of pathogenic microorganism using surface-engineered impedimetric immunosensor. Sensor. Actuat. B: Chem. 2012;161:824–831. [Google Scholar]
- Keighley SD, Li P, Estrela P, Migliorato P. Optimization of DNA immobilization on gold electrodes for label-free detection by electrochemical impedance spectroscopy. Biosensors Bioelectron. 2008;23:1291–1297. doi: 10.1016/j.bios.2007.11.012. [DOI] [PubMed] [Google Scholar]
- KFDA. Chapter 5. Standards and specifications for each food products. Food standard Codex (2020)
- Khan R, Dhayal M. Chitosan/polyaniline hybrid conducting biopolymer base impedimetric immunosensor to detect Ochratoxin-A. Biosensors Bioelectron. 2009;24:1700–1705. doi: 10.1016/j.bios.2008.08.046. [DOI] [PubMed] [Google Scholar]
- Labib M, Zamay AS, Kolovskaya OS, Reshetneva IT, Zamay GS, Kibbee RJ, Sattar SA, Zamay TN, Berezovski MV. Aptamer-based viability impedimetric sensor for bacteria. Anal. Chem. 2012;84:8966–8969. doi: 10.1021/ac302902s. [DOI] [PubMed] [Google Scholar]
- Leca-Bouvier B, Blum LJ. Biosensors for protein detection: A review. Anal. Lett. 2005;38:1491–1517. [Google Scholar]
- Li X, Huang Y, Chen M, Tong Y, Zhang C. A label-free electrochemical bisphenol A immunosensor based on chlorogenic acid as a redox probe. Anal. Methods. 2017;9:2183–2188. [Google Scholar]
- Liang Y, Zhao X, Wang N, Wang J, Chen H, Bai L, Wang W. A label-free immunosensor based on PHEMA/graphene oxide nanocomposite for simultaneous electrochemical determination of alpha fetoprotein. RSC Adv. 2019;9:17187–17193. doi: 10.1039/c9ra02565k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin D, Pillai RG, Lee WE, Jemere AB. An impedimetric biosensor for E. coli O157:H7 based on the use of self-assembled gold nanoparticles and protein G. Microchim. Acta 186: 169 (2019) [DOI] [PubMed]
- Liu J, Jasim I, Abdullah A, Shen Z, Zhao L, El-Dweik M, Zhang S, Almasri M. An integrated impedance biosensor platform for detection of pathogens in poultry products. Sci. Rep. 2018;8:16109. doi: 10.1038/s41598-018-33972-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu Z, Li CM, Zhou Q, Bao Q-L, Cui X. Covalently linked DNA/protein multilayered film for controlled DNA release. J. Colloid Interface Sci. 2007;314:80–88. doi: 10.1016/j.jcis.2007.05.018. [DOI] [PubMed] [Google Scholar]
- Lu L, Liu B, Zhao Z, Ma C, Luo P, Liu C, Xie G. Ultrasensitive electrochemical immunosensor for HE4 based on rolling circle amplification. Biosensors Bioelectron. 2012;33:216–221. doi: 10.1016/j.bios.2012.01.004. [DOI] [PubMed] [Google Scholar]
- Luo C, Lei Y, Yan L, Yu T, Li Q, Zhang D, Ding S, Ju H. A rapid and sensitive aptamer-based electrochemical biosensor for direct detection of Escherichia coli O111. Electroanalysis. 2012;24:1186–1191. [Google Scholar]
- Luong JHT, Hrapovic S, Wang D, Bensebaa F, Simard B. Solubilization of multiwall carbon nanotubes by 3-aminopropyltriethoxysilane towards the fabrication of electrochemical biosensors with promoted electron transfer. Electroanalysis. 2004;16:132–139. [Google Scholar]
- Maalouf R, Chebib H, Saïkali Y, Vittori O, Sigaud M, Jaffrezic-Renault N. Amperometric and impedimetric characterization of a glutamate biosensor based on Nafion® and a methyl viologen modified glassy carbon electrode. Biosensors Bioelectron. 2007;22:2682–2688. doi: 10.1016/j.bios.2006.11.003. [DOI] [PubMed] [Google Scholar]
- Maalouf R, Fournier-Wirth C, Coste J, Chebib H, Saïkali Y, Vittori O, Errachid A, Cloarec J-P, Martelet C, Jaffrezic-Renault N. Label-free detection of bacteria by electrochemical impedance spectroscopy: Comparison to surface plasmon resonance. Anal. Chem. 2007;79:4879–4886. doi: 10.1021/ac070085n. [DOI] [PubMed] [Google Scholar]
- Malvano F, Pilloton R, Albanese D. Sensitive detection of escherichia coli O157:H7 in food products by impedimetric immunosensors. Sensors. 2018;18:2168. doi: 10.3390/s18072168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malvano F, Pilloton R, Albanese D. Label-free impedimetric biosensors for the control of food safety – A review. Int. J. Environ. Anal. Chem. 2019;1:1–24. [Google Scholar]
- Manickam A, Johnson CA, Kavusi S, Hassibi A. Interface design for CMOS-integrated electrochemical impedance spectroscopy (EIS) biosensors. Sensors. 2012;12:14467–14488. doi: 10.3390/s121114467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mantzila AG, Maipa V, Prodromidis MI. Development of a faradic impedimetric immunosensor for the detection of Salmonella typhimurium in milk. Anal. Chem. 2008;80:1169–1175. doi: 10.1021/ac071570l. [DOI] [PubMed] [Google Scholar]
- Nicosia C, Huskens J. Reactive self-assembled monolayers: from surface functionalization to gradient formation. Mater. Horiz. 2014;1:32–45. [Google Scholar]
- Niu Y, Matos AI, Abrantes LM, Viana AS, Jin G. Antibody oriented immobilization on gold using the reaction between carbon disulfide and amine groups and its application in immunosensing. Langmuir. 2012;28:17718–17725. doi: 10.1021/la303032f. [DOI] [PubMed] [Google Scholar]
- Omotayo OP, Omotayo AO, Mwanza M, Babalola OO. Prevalence of mycotoxins and their consequences on human health. Toxicol. Res. 2019;35:1–7. doi: 10.5487/TR.2019.35.1.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paniel N, Baudart J, Hayat A, Barthelmebs L. Aptasensor and genosensor methods for detection of microbes in real world samples. Methods. 2013;64:229–240. doi: 10.1016/j.ymeth.2013.07.001. [DOI] [PubMed] [Google Scholar]
- Park JS, Kim HJ, Lee J-H, Park JH, Kim J, Hwang KS, Lee BC. Amyloid beta detection by Faradaic electrochemical impedance spectroscopy using interdigitated microelectrodes. Sensors. 2018;18:426. doi: 10.3390/s18020426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peng H, Soeller C, Cannell MB, Bowmaker GA, Cooney RP, Travas-Sejdic J. Electrochemical detection of DNA hybridization amplified by nanoparticles. Biosensors Bioelectron. 2006;21:1727–1736. doi: 10.1016/j.bios.2005.08.011. [DOI] [PubMed] [Google Scholar]
- Plueddemann EP. Nature of adhesion through silane coupling agents. pp. 115–152. In: Silane Coupling Agents. Springer US, Boston, MA (1991)
- Pournaras AV, Koraki T, Prodromidis MI. Development of an impedimetric immunosensor based on electropolymerized polytyramine films for the direct detection of Salmonella typhimurium in pure cultures of type strains and inoculated real samples. Anal. Chim. Acta. 2008;624:301–307. doi: 10.1016/j.aca.2008.06.043. [DOI] [PubMed] [Google Scholar]
- Prodromidis MI. Impedimetric immunosensors—A review. Electrochim. Acta. 2010;55:4227–4233. [Google Scholar]
- Radhakrishnan R, Suni II, Bever CS, Hammock BD. Impedance biosensors: Applications to sustainability and remaining technical challenges. ACS Sustain. Chem. Eng. 2014;2:1649–1655. doi: 10.1021/sc500106y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Radi A-E, Muñoz-Berbel X, Lates V, Marty J-L. Label-free impedimetric immunosensor for sensitive detection of ochratoxin A. Biosensors Bioelectron. 2009;24:1888–1892. doi: 10.1016/j.bios.2008.09.021. [DOI] [PubMed] [Google Scholar]
- Rishpon J, Buchner V. Electrochemical antibody-based sensors. pp. 329–373. In: Comprehensive Analytical Chemistry. Gorton L (ed). Elsevier, Amsterdam (2005)
- Ron H, Rubinstein I. Self-assembled monolayers on oxidized metals. 3. Alkylthiol and dialkyl disulfide assembly on gold under electrochemical conditions. J. Am. Chem. Soc. 120: 13444–13452 (1998)
- Rosati G, Cunego A, Fracchetti F, Del Casale A, Scaramuzza M, De Toni A, Torriani S, Paccagnella A. Inkjet printed interdigitated biosensor for easy and rapid detection of bacteriophage contamination: A preliminary study for milk processing control applications. Chemosensors. 2019;7:8. [Google Scholar]
- Rosati G, Ravarotto M, Sanavia M, Scaramuzza M, De Toni A, Paccagnella A. Inkjet sensors produced by consumer printers with smartphone impedance readout. Sens. Biosensing Res. 2019;26:100308. [Google Scholar]
- Săndulescu R, Tertis M, Cristea C, Bodoki E. New materials for the construction of electrochemical biosensors. pp. 1–36. In: Biosensors-Micro and Nanoscale Applications. InTech (2015)
- Sauthier ML, Carroll RL, Gorman CB, Franzen S. Nanoparticle layers assembled through DNA hybridization: Characterization and optimization. Langmuir. 2002;18:1825–1830. [Google Scholar]
- Sheikhzadeh E, Chamsaz M, Turner APF, Jager EWH, Beni V. Label-free impedimetric biosensor for Salmonella Typhimurium detection based on poly [pyrrole-co-3-carboxyl-pyrrole] copolymer supported aptamer. Biosensors Bioelectron. 2016;80:194–200. doi: 10.1016/j.bios.2016.01.057. [DOI] [PubMed] [Google Scholar]
- Tam PD, Van Hieu N, Chien ND, Le A-T, Anh Tuan M. DNA sensor development based on multi-wall carbon nanotubes for label-free influenza virus (type A) detection. J. Immunol. Methods. 2009;350:118–124. doi: 10.1016/j.jim.2009.08.002. [DOI] [PubMed] [Google Scholar]
- Tang J, Huang Y, Zhang C, Liu H, Tang D. Amplified impedimetric immunosensor based on instant catalyst for sensitive determination of ochratoxin A. Biosensors Bioelectron. 2016;86:386–392. doi: 10.1016/j.bios.2016.06.080. [DOI] [PubMed] [Google Scholar]
- Teng J, Yuan F, Ye Y, Zheng L, Yao L, Xue F, Chen W, Li B. Aptamer-based technologies in foodborne pathogen detection. Front. Microbiol. 7 (2016) [DOI] [PMC free article] [PubMed]
- US FDA. Chemical contaminants, metals, natural toxins and pesticides guidance documents and regulations. U.S. Food and Drug Administration (2016)
- Vashist SK, Lam E, Hrapovic S, Male KB, Luong JHT. Immobilization of antibodies and enzymes on 3-aminopropyltriethoxysilane-functionalized bioanalytical platforms for biosensors and diagnostics. Chem. Rev. 2014;114:11083–11130. doi: 10.1021/cr5000943. [DOI] [PubMed] [Google Scholar]
- Wakenell P. CHAPTER 15 - Management and medicine of backyard poultry. pp. 550–565. In: Current Therapy in Avian Medicine and Surgery. Speer BL (ed). W.B. Saunders (2016)
- Walsh MK, Wang X, Weimer BC. Optimizing the immobilization of single-stranded DNA onto glass beads. J. Biochem. Biophys. Methods. 2001;47:221–231. doi: 10.1016/s0165-022x(00)00146-9. [DOI] [PubMed] [Google Scholar]
- Wang D, Hu W, Xiong Y, Xu Y, Ming Li C. Multifunctionalized reduced graphene oxide-doped polypyrrole/pyrrolepropylic acid nanocomposite impedimetric immunosensor to ultra-sensitively detect small molecular aflatoxin B1. Biosensors Bioelectron. 2015;63:185–189. doi: 10.1016/j.bios.2014.06.070. [DOI] [PubMed] [Google Scholar]
- Wang W, Wang Y, Tu L, Klein T, Feng Y, Wang J. Surface modification for protein and DNA immobilization onto GMR biosensor. IEEE Trans. Magn. 2013;49:296–299. [Google Scholar]
- Yu L, Zhang Y, Hu C, Wu H, Yang Y, Huang C, Jia N. Highly sensitive electrochemical impedance spectroscopy immunosensor for the detection of AFB1 in olive oil. Food Chem. 2015;176:22–26. doi: 10.1016/j.foodchem.2014.12.030. [DOI] [PubMed] [Google Scholar]
- Zheng D, Vashist SK, Dykas MM, Saha S, Al-Rubeaan K, Lam E, Luong JHT, Sheu F-S. Graphene versus multi-walled carbon nanotubes for electrochemical glucose biosensing. Materials. 2013;6:1011–1027. doi: 10.3390/ma6031011. [DOI] [PMC free article] [PubMed] [Google Scholar]