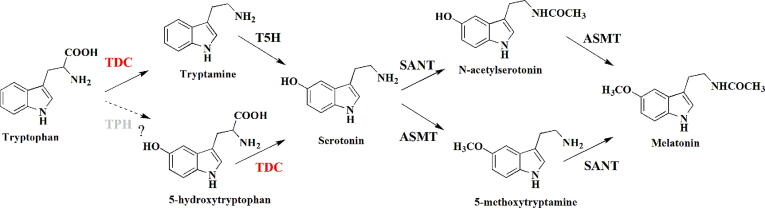
Graphical abstract
Keywords: Crystal structure, Tryptophan decarboxylase, Melatonin biosynthesis, Substrate specificity
Abstract
Plant tryptophan decarboxylase (TDC) is a type II Pyridoxal-5′-phosphate-dependent decarboxylase (PLP_DC) that could be used as a target to genetically improve crops. However, lack of accurate structural information on plant TDC hampers the understanding of its decarboxylation mechanisms. In the present study, the crystal structures of Oryza sativa TDC (OsTDC) in its complexes with pyridoxal-5′-phosphate, tryptamine and serotonin were determined. The structures provide detailed interaction information between TDC and its substrates. The Y359 residue from the loop gate is a proton donor and forms a Lewis acid-base pair with serotonin/tryptamine, which is associated with product release. The H214 residue is responsible for PLP binding and proton transfer, and its proper interaction with Y359 is essential for OsTDC enzyme activity. The extra hydrogen bonds formed between the 5-hydroxyl group of serotonin and the backbone carboxyl groups of F104 and P105 explain the discrepancy between the catalytic activity of TDC in tryptophan and in 5-hydroxytryptophan. In addition, an evolutionary analysis revealed that type II PLP_DC originated from glutamic acid decarboxylase, potentially as an adaptive evolution of mechanism in organisms in extreme environments. This study is, to our knowledge, the first to present a detailed analysis of the crystal structure of OsTDC in these complexes. The information regarding the catalytic mechanism described here could facilitate the development of protocols to regulate melatonin levels and thereby contribute to crop improvement efforts to improve food security worldwide.
Introduction
Melatonin (N-acetyl-5-methoxytryptamine) was previously considered as a promoter produced mainly in animals by the pineal gland [1], [2]. However, recently, it has been reported to be involved in numerous physiological processes, with anti-aging, immune system enhancement, and reproductive activity regulation properties [3], [4]. In addition, melatonin has potential applications in human disease prevention and treatment, and this hormone could limit cancer progression by inhibiting cell growth and migration [5], [6], [7]. Besides, studies have revealed that all organisms, including plants produce melatonin, potentially as a defense agent against various abiotic stresses factors [8]. Melatonin is present in different parts of plants, including leaves, stems, roots, fruits, and seeds, and shares structural features with indoleacetic acid (IAA) [9]. Numerous studies have focused on the physiological functions of melatonin in plants and its benefits, when compared with IAA, as an excellent modulation target for crop improvement [10]. For instance, an increase in melatonin in transgenic rice or exogenous melatonin application could regulate seed germination and root elongation under abiotic stresses factors such as cold and osmotic stress [11], [12], [13].
The classic melatonin biosynthetic pathway in vertebrates has been studied extensively and demonstrated in other organisms, including plants [14], [15]. The biosynthesis of melatonin in plants appears to be much more complicated with four pathways [16], [17]. The conserved biosynthetic pathway in plants consists of four enzymatic reactions, and tryptophan decarboxylase (TDC) initiates the pathway by synthesizing tryptamine (TSS) from tryptophan. Subsequently, P450 tryptamine 5-hydroxylase (T5H) produces serotonin (SRO), followed by two concurrent methylation/acylation steps that generate melatonin (Fig. 1, Fig. S1) [16], [18]. Most of the enzymes involved in the pathway have been identified and studied extensively in plants, excluding tryptophan hydroxylase (TPH). However, several studies have demonstrated that the hydroxylation of tryptophan could also initiate melatonin biosynthesis in plants. For example, the suppression of O. sativa T5H leads to an unexpected increase in tryptophan, tryptamine, 5-hydroxytryptophan and melatonin concentrations, with corresponding increases in transcriptional levels of OsTDC [19]. TDC could catalyze both tryptophan and 5-hydroxytryptophan [20], and an alternative melatonin biosynthetic pathway similar to that in animals could be triggered in plants by some biotic or abiotic stresses factors [16], [21]. Therefore, TDC could regulate melatonin synthesis as a bi-functional enzyme via a dual catalysis pathway.
Fig. 1.
TDC mediates multiple melatonin biosynthesis pathways in rice. At least four different pathways are involved in melatonin biosynthesis in rice. TPH has not yet been identified in rice and is indicated in gray.
TDC is an aromatic-L-amino-acid decarboxylase (AADC; EC4.1.1.28) that has been classified evolutionarily as a type II Pyridoxal-5′-phosphate-dependent decarboxylase (PLP_DC) (Fig. S2) [22]. Generally, AADC utilizes Pyridoxal-5′-phosphate (PLP) as a co-factor to catalyze the decarboxylation of a few aromatic L-amino acids and their α-methylated derivatives, including tryptophan, tyrosine, and L-DOPA [23], [24], [25]. TDC has been isolated from various plant species [26], [27], and has been reported to exhibit strict substrate specificity for tryptophan, and not for phenolic L-amino acids (phenylalanine and tyrosine) and their derivatives, which are specifically recognized by L-DOPA decarboxylase (DDC) and tyrosine decarboxylase (TyDC) [28], [29]. The structures of several type II PLP_DCs have already been resolved [30], [31], [32], [33], and the elucidation of such structures facilitates the understanding of the catalytic mechanisms of such types of enzymes. PLP is bound covalently to the Ɛ-amino group of a conserved lysine residue. Although the architectures of the PLP_DCs are conserved, they exhibit strict substrate specificity (Fig. S3, Table S1). Consequently, the specific mechanisms of substrate binding are diverse. The only TDC whose structure that has been resolved, which originated from gut microbiota (PDB ID: 4OBV), displays major differences in evolutionary and sequence identity with plant TDC. Therefore, it is not a very useful source of information in attempts to clarify the decarboxylation dynamics in plant TDCs. Additionally, without accurate structural information on plant TDCs, it is impossible to describe how TDCs distinguish tryptophan from 5-hydroxytryptophan based solely on primary sequence data.
In the present study, we investigated the potential roles of TDC in melatonin biosynthesis from a structural perspective. The crystal structure of a plant TDC from O. sativa and the complexes it forms with PLP, TSS, and SRO were examined. In addition, a comparative analysis and systematic phylogenetic analysis of the type II PLP_DC family were performed. Considering the importance of melatonin in rice and other plants, the catalytic mechanism of OsTDC described in the present study could facilitate the development of protocols for the regulation of the melatonin levels in plants as a crop improvement tool [16], [34].
Materials and methods
Chemicals and reagents
Chemicals and reagents such as tryptophan and tryptamine that were used in this study were purchased from Sigma-Aldrich (St. Louis, MO, USA) or Aladdin (Shanghai, China). Kanamycin was purchased from Melonepharm (Dalian, China) and yeast extract and tryptone were from OXOID (Basingstoke, UK). KOD-Plus-Neo was purchased from TOYOBO (Shanghai, China).
Protein expression and purification
O. sativa TDC was cloned into the pET-28a expression vector with a C-terminal hexahistidine tag. The plasmid was confirmed and transformed into Escherichia coli BL21 (DE3) for expression. The purification procedure was performed according to previous study with minor modifications [35]. After induction for 18 h with 0.1 mM IPTG at 20 °C, the cells were harvested by centrifugation, and the pellets were resuspended in lysis buffer (50 mM Tris-HCl pH 8.0, 400 mM NaCl, 10% glycerol, 1 mM PMSF). The cells were then lysed by sonication, and the debris was removed by ultracentrifugation. The supernatant was mixed with Ni-NTA agarose beads (Qiagen) and rocked for 1 h at 4 °C before elution with 250 mM imidazole. The proteins were further purified on Hitrap Q (GE Healthcare, USA). After a final step of gel-filtration chromatography on a Superdex 200 column (GE Healthcare, USA) equilibrated with 25 mM Tris-HCl pH 8.0, 150 mM NaCl, the purified proteins were concentrated to 15 mg/mL and stored at −80 °C.
Crystallization
All Crystals of TDC related were grown by sitting-drop vapor diffusion at 4 °C. For the TDC-PLP complex, the TDC protein at a concentration of 15 mg/mL was premixed with 10 mM PLP and then mixed with an equal volume of reservoir solution containing 20% (w/v) polyethylene glycol 1000, 100 mM imidazole pH 8.0 and 200 mM calcium acetate. For the TDC-PLP-SRO complex and the TDC-PLP-TSS complex, except premixed the PLP, supplied with 10 mM tryptophan and 10 mM 5-hydroxyltryptophan, respectively. The crystals suitable for X-ray diffracting appeared within 3 to 5 days. The crystals were gradually transferred into a harvesting solution containing 22% (w/v) polyethylene glycol 1000, 100 mM imidazole pH 8.0 and 200 mM calcium acetate, 25% (v/v) glycerol, followed by flash-freezing in liquid nitrogen for storage [35].
Data collection and structure determination
Data sets were collected under cryogenic conditions (100 K) at Shanghai Synchrotron Radiation Facility (SSRF) beamlines BL18U1 and BL19U1. All data were indexed, integrated and scaled by HKL3000 [36]. Further data processing was carried out using CCP4 suit [37]. The first TDC structure was solved by molecular replacement with the homologous human HDC (PDB ID: 4E1O) as the search model using the program PHASER, the others used TDC itself as the search model [38]. All the structures were iteratively built with COOT and refined with PHENIX program [39], [40]. Data collection and structure refinement statistics are summarized in Table 1. All figures were generated with PyMOL [41].
Table 1.
Data collection and refinement statistics for TDC.
| 6KHO (TDC + PLP) | 6KHP (TDC + PLP + TSS) | 6KHN (TDC + PLP + SRO) | |
|---|---|---|---|
| Data collection | |||
| Wavelength(Å) | 0.97915 | 0.91905 | 0.97915 |
| Space group | P3121 | P3121 | P3121 |
| Cell dimensions | |||
| a, b, c(Å) | 155.7,155.7, 102.2 | 155.8, 155.8, 102.1 | 156.3, 156.3, 102.4 |
| α, β, γ (°) | 90, 90, 120 | 90, 90, 120 | 90, 90, 120 |
| Resolution(Å) | 1.97 | 2.30 | 2.30 |
| Rmerge (%) | 8.3(62.5)* | 11.6(71.7)* | 10.2(71.2)* |
| I/σI | 39.6(2.5)* | 15.3(2.3)* | 30.3(3.5)* |
| Completeness (%) | 97.5(93.9)* | 99.8(99.6)* | 99.9(1 0 0)* |
| Redundancy | 9.3(8.1)* | 15.9(13.3)* | 8.0(7.5)* |
| Refinement | |||
| Resolution(Å) | 42.72–1.97 | 28.13–2.29 | 47.88–2.29 |
| No. of reflections | 98,051 | 63,528 | 64,580 |
| Rwork/Rfree (%) | 16.8/19.4 | 16.3/19.7 | 16.2/20.1 |
| No. of atoms | |||
| Protein | 7398 | 7372 | 7372 |
| Ligand | 30 | 42 | 43 |
| PEG | 14 | 14 | 14 |
| ACT | 16 | 8 | 8 |
| Ca | 4 | 4 | 4 |
| Water | 508 | 409 | 334 |
| B-factors(Å2) | |||
| Protein | 36.7 | 35.6 | 39.1 |
| Ligand | 37.6 | 47.1 | 47.5 |
| PEG | 40.7 | 38.5 | 54.9 |
| ACT | 43.1 | 43.1 | 48.8 |
| Ca | 49.9 | 54.2 | 59.3 |
| Water | 42.4 | 37.2 | 38.1 |
| RMS deviations | |||
| Bond lengths (Å) | 0.007 | 0.007 | 0.007 |
| Bond angles (°) | 0.829 | 0.813 | 0.838 |
| Ramanchandran plot | |||
| Favored region | 97.07% | 97.18% | 96.97% |
| Allowed region | 100% | 100% | 100% |
| Outlier region | 0.00% | 0.00% | 0.00% |
Highest resolution shell is shown in parenthesis.
Enzymatic activity assays and HPLC/electrospray-ionization quadrupole time-of-flight mass spectrometry (Q-TOF MS) analysis
For protein activity analysis, 2.5 μg protein with 1 mM PLP and 0.5 mM different substrates were mixed in 0.2 M potassium phosphate buffer with a total volume of 200 μL (PBS, pH 7.5). The reaction was started with the addition of PLP and ended with the addition of 200 μL 0.1 M hydrochloric acid (HCl). Reaction was conducted in a temperature control instrument with a constant temperature of 37 °C. The reaction product was centrifuged in 12000 rpm for 10 min and the supernatant was subjected to HPLC for analysis. HPLC and Q-TOF MS analysis were conducted according to the previous reports with some minor modification [42]. The solvent gradient conditions A (methanol): B (H2O with 0.1% formic acid) (v/v) for tryptophan and tryptamine detection are as follows: 0 min, 5:95; 5 min, 60:40; 15 min, 95:5. While, for 5-hydroxytryptophan and serotonin detection, the gradient conditions are as follows: 0 min, 5:95; 5 min, 20:80; 10 min, 30:70; 15 min, 100:0. For Q-TOF MS analysis, the HPLC system was connected to Q-TOF MS spectrometer (Agilent Technologies, Santa Clara, CA, USA) equipped with an electrospray interface to identify the products. The conditions of the ESI source were as follows: drying gas (N2) flow rate, 8.0 l/min; drying gas temperature, 300 °C; nebulizer, 241 kPa (35 psig); capillary voltage, 4000 V; fragmentor voltage, 150 V; collision energy, 30 eV; skimmer voltage, 60 V, and octopole radio frequency, 250 V [42].
Bioinformatics and evolutionary trails analysis
Multiple sequence alignment and protein structure alignment was performed using DNAMAN (Lynnon Corp., Quebec, Canada) and PyMOL, respectively. A neighbor-joining phylogenetic analysis of OsTDC and other 66 kinds of PLP_DCs in different species was created in MEGA [43]. The tree was constructed with 1000 bootstrap replicates and was generated by online tool iTOL [44]. For evolutionary trails analysis, more than 1000 decarboxylases were retrieved from 76 identified species according to the studies by Kumar et al [45], [46]. The retrieved protein sequences were then checked for the typical PLP_DC domain architecture by using Pfam and SMART tools [47], [48]. The protein sequences lacking the characteristics PLP_DC domain were discarded and not included in further analyses. According to APG classification system [49], the 76 identified species were classified and their evolutionary relationship was also estimated [50]. In addition, the estimated divergence times of the identified species shown relative to the geological time scale according to Zeng et al [50].
Site-directed mutagenesis of TDC
Site-directed mutagenesis experiment was conducted using a PCR method with KOD-plus-neo and the primers used in this experiment are listed in Table S2 [51]. After PCR, the PCR products were digested with the DpnI restriction endonuclease. The mutants were then transformed into chemically competent E. coli DH5α cells and the plasmids from positive strains were extracted for sequencing. Positive plasmids were subsequently transferred to E. coli BL21 (DE3) for protein expression, purification and analyzing the enzymatic activity.
Preparation of graphs
Unless special comments, the results were present as triplicate experiments ± standard deviation (SD). Unless produced with the specific software (for example, MEGA and PyMOL), the original graphs were generated using Microsoft Office PowerPoint 2010, OriginPro 8 (OriginLab Corporation, Northampton, MA, USA) or GraphPad Prism software. When it is necessary, the graphs were merged with Adobe PhotoShop CS6 (V13.0.1.1).
Results
Overall structure of OsTDC
The crystal structures of the TDC-PLP binary complex, as well as the TDC-PLP-TSS and TDC-PLP-SRO ternary complexes, were determined at resolutions of 1.97 Å, 2.30 Å and 2.30 Å, respectively (Table 1). The overall structures revealed that OsTDC is a homodimer and the two subunits are symmetric with almost identical structures (Fig. 2a, b). Each monomeric unit is composed of three domains: the N-terminal small domain, the middle large domain and the C-terminal small domain (Fig. S4). The N-terminal small domain contains three parallel α helices that pack against the other monomer and is linked to the middle large domain by a long loop (Fig. S4). The first 21 residues in the N-terminal domain are invisible in our electron density maps, even with several rounds of refinement. The middle large domain is composed of a typical seven-stranded β-sheet surrounded by α-helices (Fig. S4). In addition, the C-terminal small domain contains a four-strand anti-parallel β-sheet surrounded by three α helices (Fig. S4). The active cavity lies at the border of the two monomers, with PLP and substrates almost docked in one monomer, while the other one remains essential for TDC activity, particularly a loop from residue 350 to 367 (Fig. 2a, b and Fig. S4). The loop, which is formed by the other subunit, covers on the surface of the active cavity and forms a closed state, which facilitates the completion of the catalytic reaction (Fig. S5). The Y359 residue from the loop region is vital for catalysis and is relatively conserved in all PLP_DCs, despite great variation in amino acid residues in the region (Fig. 3).
Fig. 2.
Overall structure of OsTDC in complexes with PLP-TSS/SRO. (a, b) The structure of OsTDC in complex with PLP-TSS/SRO. One subunit is colored with salmon and the other subunit is separated by domains and labeled with green, slate blue and yellow, respectively. The loop gate is colored in magenta. The PLP (cyan), TSS (green) and SRO (green) are shown in stick-ball and labeled. (c, d) Electron density maps of PLP-K330, TSS-Y359 and SRO-Y359. Contours are drawn at the 1.0 σ level; the residues are shown as sticks. The K330, Y359, PLP, and TSS/SRO are colored in salmon, magenta, blue and green, respectively.
Fig. 3.
Mutation analysis of OsTDC and sequence alignment of PLP_DCs. (a) Relative activities of wild type OsTDC and its variants against tryptophan (red) and 5-hydroxytryptophan (blue). The activities of the OsTDC against tryptophan and 5-hydroxytryptophan are set to 100%. Each bar represents the mean value from triplicate experiments ± SD (n = 3). (b) Excerption of key amino acid residues from the multiple sequence alignment results of different PLP_DCs. Accession numbers are listed on the left, residues of OsTDC are labeled with numbers, and the highly conserved residues are marked with red color. The alignment was created by ClustalW and generated by online tool Espript 3.0. Different PLP_DCs and their substrate structures are marked in the right with vertical bars with different colors.
An initial attempt to crystallize the apo form of TDC failed, and crystallization was only possible after the PLP co-factor was supplied, which indicates a conformational transition from the TDC apo form to its PLP binding form. Although tryptophan and 5-hydroxytryptophan were included for co-crystallization with TDC, the only electron density features observed were for the TSS moiety of tryptophan and the SRO moiety of 5-hydroxytryptophan, potentially because of decarboxylation (Fig. 2). Therefore, we obtained two ternary complexes containing their products TSS (TDC-PLP-TSS) and SRO (TDC-PLP-SRO) (Fig. 2).
Co-factor binding site
PLP co-factor is functionally conserved in all PLP_DCs and it always binds to a conserved lysine to form one Schiff base [22], [32]. In our structures, continuous electron density distribution supports the Schiff base linkage between PLP and K330, and K330A mutation leads to complete inactivation (Fig. 2c, d and Fig. 3a). The result indicates that K330 plays a fundamental role in its decarboxylation catalysis. The decarboxylation reaction involves the cleavage of the Schiff base bond and the formation of a new quinoid transition Schiff base bond between the substrate and PLP (Fig. S6). The two Schiff bases are nearly perpendicular to each other, which we following by superimposition of OsTDC-PLP-SRO onto pig DDC (1JS3) and hHDC (4E1O) (Fig. S7). In addition, the methyl group of A300 and the imidazole ring of H214 anchor the pyridine ring of PLP to form a sandwich structure (Fig. S6j and S8). Correspondingly, mutants, such as H214A, H214F, H214Q and H214Y, drastically decrease OsTDC decarboxylation activity, which suggests that H214 has a key role in the maintenance of OsTDC catalytic activity (Fig. 3a). The protonated pyridine nitrogen of PLP forms a pair of salt bridges with the carboxyl group of D298, and the O3 atom of the pyridine ring of PLP forms a hydrogen bond with the hydroxyl group of T273 (Fig. S6j and S8). Although the residues are highly conserved in the known PLP_DCs [30], [33], the motif residues involved in stabilizing the PLP phosphate group are variable in different PLP_DCs (Fig. 3b). In the OsTDC, the O1P atom is hydrogen bonded to the backbone amino group of T175 (Fig. S8); the O2P atom is hydrogen bonded to the hydroxyl group of S176; and the O2P atom can also form hydrogen bonds with the backbone amino group of S176 and V380 from the other monomer (Fig. S8); and the O3P atom forms hydrogen bonds with the backbone amino group of G381 from the other monomer (Fig. S8).
Substrate binding and specificity
The substrate-binding pocket in TDC is formed by several helices and loops. Hydrophobic residues, W95, F103, F104 and L336, from one subunit, and V125, F127, L360 and V380, from an adjacent subunit, provide a hydrophobic environment for the substrate molecules (Fig. S8). Mutation of the residues into alanine decreases or eliminates activity (Fig. 3a). The loop region (residue 350 to 367 in TDC), considered to control the entry and release of substrates and products, is disordered in most of the resolved PLP_DCs [30], [33]. Nevertheless, it could be well built in the electron density of one subunit in our structures (Fig. S5c). We observed that the amino groups of SRO and TSS were linked with the hydroxyl group of Y359 in a Lewis acid-base form (Fig. 2). Mutants Y359A, Y359F, and Y359H completely eliminated its activity; however, E358A and K361A mutations did not influence its activity (Fig. 3a). The results indicated that Y359 is crucial for substrate recognition and acts as a proton donor during catalysis. The imidazole nitrogen of H214 is proximal to the hydroxyl group of Y359, at a distance of 2.9 Å, which suggests the existence of proton transfer chains between these residues (Fig. 4 and Fig S7). Such a phenomenon is not observed in hHDC structure and gut microbiota TDC considering the corresponding tyrosine residue is located relatively far from the histidine residue and the substrate (Fig. S7b).
Fig. 4.
Superimposition of OsTDC Structures. (a) The TDC-PLP, TDC-PLP-TSS and TDC-PLP-SRO structures are colored with green, slate blue and salmon, respectively. The gate loop region is colored with magenta. PLP, TSS, and SRO are shown as sticks. (b) A view of the active pocket of OsTDC-PLP-TSS. (c) A view of active pocket of OsTDC-PLP-SRO. For (b) and (c), the residues are shown in sticks with rainbow colors, the hydrogen bonds are indicated with red dotted lines.
TDC has been reported to exhibit varying catalytic efficiency against tryptophan and 5-hydroxytryptophan [18], [52]. To investigate how TDC distinguishes tryptophan from 5-hydroxytryptophan, we superimposed the crystal structures of TDC-PLP-TSS, TDC-PLP-SRO and TDC-PLP. According to the results, there is no conformational transition; the TSS and SRO molecule occupied similar positions (Fig. 4). In addition, the substitution of the substrate did not alter the active site architecture of TDC considerably, and the binding details were almost identical in each complex, with the aromatic ring of TSS and SRO burying deeply within the active site cleft and penetrating the cofactor ring plane (Fig. 4). A remarkable feature of the 5-hydroxytryptophan structure, which distinguishes it from the tryptophan structure, is that 5-hydroxytryptophan has an additional hydroxyl group on the indole ring. Two notable hydrogen bonds may form between the 5-hydroxyl group of SRO and the backbone carboxyl group of F104 and P105 (Fig. 4c).
Previous studies have demonstrated that the specificity for different aromatic amino acid substrates could be driven by the hydrophobic nature of the residues around the active pocket, in addition to the size of the active pocket [33], [53]. The single active site amino acid residue that could define plant AADC indole and benzene substrate selectivity corresponded to G381 in TDC. In addition, an in vitro enzyme assay revealed that OsTDC could not catalyze tyrosine and L-DOPA (Fig. S9). The distance between G381 and TSS is greater than 3.5 Å, and could be smaller than 3.0 Å if glycine mutates into serine (Fig. S10). The reduced steric hindrance and higher hydrophobicity are potential reasons why the binding pocket of TDC is capable of accommodating tryptophan, which, as a substrate, is configurationally larger than histidine and tyrosine.
Comparison with other type II PLP_DCs
The crystal structures of several type II PLP_DCs from different species have been resolved [30], [31], [32], [33], [54]. Structural comparison revealed that TDC exhibits a structural resemblance to the resolved PLP_DCs considering they adopt similar fold and dimeric configurations (Fig. S3). All of them consist of three characteristic domains and have similar active centers and essential conserved residues, such as the base-stacking histidine residue, which is vital for PLP binding [55]. Besides, the location and binding characteristics of PLP are almost identical in OsTDC and the PLP_DCs (Fig. S7). A common feature of catalytic mechanisms among all the PLP_DCs is the presence of a lysine residue, observed in the active sites of all the PLP enzyme structures (e.g., K303 in pig kidney DDC), which forms a Schiff base with the co-factor in the absence of a substrate. The lysine is involved in external aldimine formation and hydrolysis, and in product release. Another conserved catalytic residue is a tyrosine (e.g., Y332 in pig kidney DDC) located in the middle of the conserved mobile loop, and it interacts with the substrate by forming a hydrogen bond. The corresponding residues observed in OsTDC are H214, K330 and Y359. The consistency among the structures could be linked to their functional similarity. However, multiple sequence alignment revealed that despite their conserved structures, the PLP_DCs varied in length and exhibited very low levels of sequence identity/similarity, particularly in the substrate binding pocket, which may explain why individual enzymes could generate specific products with unique physiological functions. Comparisons were also performed among the TDCs identified from various plant species. As expected, they shared high sequence identity (48–87%), particularly in the region around the active pocket (Fig. S11), which is consistent with their evolutionary relationship or catalytic similarity, even among different plant lineages (Fig. S2).
To clarify the relationship between TDC and other PLP_DCs, we conducted a systematic phylogenetic analysis (Fig. 5). Among the four types of decarboxylases, type II PLP_DCs form a large and important group and are phylogenetically present in almost all organisms ranging from primitive bacteria to humans. The evolutionary relationships among the selected type II PLP_DCs were classified and evaluated and the results indicated that all the type II PLP_DCs originated from glutamic acid decarboxylase (GAD). The primitive bacterium Escherichia coli only had GADs, without any histidine decarboxylase (HDC) or AADC enzymes. Green algae such as Chlamydomonas reinhardtii and Coccomyxa subellipsoidea C-169 had a few more PLP_DCs including HDC and AADC. Their genes exhibited a notable expansion in land plants and the gene number of different PLP_DCs in different species, with obvious differences in evolution times. Our findings are consistent with previously identified PLP_DC expansion trends in land plants [45]. Generally, more GADs and AADCs were identified in monocots than in dicots, while more HDCs were identified in dicots. However, there were exceptions, and a few dicot species such as Glycine max had more GADs and HDCs over AADC genes, while Brassica napus had more GADs and AADCs.
Fig. 5.
Evolutionary trails of PLP_DC. More than 1000 decarboxylases were retrieved from 76 identified species and then the retrieved protein sequences were checked for the typical PLP_DC domain architecture using Pfam and SMART tools. The PLP_DCs were classified and analyzed statistically. Different colors indicate different PLP_DCs and lengths of the bars represent the number of different PLP_DCs in different species. The evolutionary relationships among the 76 identified species were classified and estimated according to the APG classification system. The estimated divergence times are also illustrated.
Discussion
Melatonin plays various roles in plant growth and development, and is an effective target molecule for increasing crop yield, and nutraceutical value, and resistance to diseases [56], [57]. Melatonin-rich transgenic species, such as rice and tomato, have been studied extensively by overexpressing vertebrates’ genes that are involved melatonin biosynthesis in them [11], [58], [59], [60], [61]. The melatonin biosynthetic process in plants involves at least four enzymes, including TDC, T5H, SNAT, and ASMT, whose functional and biochemical features have been clarified [16], [34]. For example, studies have revealed that TDC could catalyze the decarboxylation of 5-hydroxytryptophan as well; therefore, 5-hydroxytryptophan could also initiate melatonin biosynthesis [16]. However, lack of structural information on the enzymes has limited their application in crop genetic improvement. In the present study, the crystal structure of OsTDC and its complexes with different ligands, TSS and SRO, together with the in vitro enzymatic assays revealed that it exhibits catalytic activity both in tryptophan and 5-hydroxytryptophan. In addition, the structures observed reinforce the hypothesis proposing the existence of a TDC-mediated dual melatonin biosynthesis pathway [16]. The identification of TPH in plants that convert tryptophan into 5-hydroxytryptophan could further confirm the hypothesis directly.
AADCs enzymes often form a homodimer with similar architectures to execute their biological functions [30], [62]. The lysine residue plays a key role in their activity by linking with the PLP co-factor to form a Schiff bases and it is conserved [62]. In addition, histidine or tyrosine residues around the substrate often act as proton donors to facilitate the enzyme activity [32], [54], [63], [64], [65]. The tyrosine residue is often located at a loop region and it is difficult to visualize in numerous crystal structures [64], [66]. Nevertheless, we tracked the electron density signals of the loop regions (residues 350–367) and substrates (Fig. 2 and Fig. S5c). In addition to the K330-PLP, which exhibits a continuous electron density, we observed that the hydroxyl group of Y359 was very close to the amine group of tryptamine/serotonin (Fig. 2). Site-directed mutagenesis results indicated that Y359 is critical for substrate recognition and could act as a proton donor during catalysis, which is consistent with the results of Williams and Burkhard et al [33], [64]. Notably, another conserved residue, H214, is in a hydrogen bond distance with Y359. Mutation of either of them to other amino acids eliminated enzyme activity (Fig. 3a). Considering histidine could often act as a general base [67], [68], [69], we concluded that H214 could act together with Y359 to facilitate proton transfer. However, in LbTyDC, the corresponding histidine residue, H241 has different conformations in apo and PLP-bound structures (Fig. S7), indicating that histidine residue is vital for PLP binding [32].
AADCs often exhibit strict substrate selectivity, which is puzzling, considering they share high amino acid identities [29]. In the current study, we confirmed that OsTDC maintains activity in tryptophan and 5-hydroxytryptophan, but exhibits no activity in tyrosine and L-DOPA (Fig. 3a, Fig. S1 and S9). However, DDC has the capacity to convert both tyrosine and L-DOPA, which indicated that such substrate heterozygosity may exist in most AADCs [30], [64]. Our structures included two ligand products, TSS and SRO, which revealed that the substrates tryptophan and 5-hydroxytryptophan bind to OsTDC with similar configurations (Fig. 4). The substrates binding pocket and active cavities are composed mainly of W95, F103, F104 and L336, hydrophobic residues from one subunit, and V125, F127, L360 and V380, from the adjacent subunit (Fig. S8). The residues are relatively variable across different AADCs based on our multiple sequence alignment results (Fig. 3b) and other reported structures that may confer substrate specificity [30], [31], [32], [33], [54], [64].
Compared to TSS, SRO has an additional hydroxyl group and form hydrogen bonds with the backbone carboxyl group of F104 and P105, which could limit the release of SRO, in turn, imposing greater selectivity for tryptophan than for 5-hydroxytryptophan (Fig. 4c). The impaired release of SRO may explain why suppression of rice T5H induces the upregulation of TDC transcription to maintain melatonin levels [19]. The change in catalytic efficiency and substrate affinity of the S354G mutant in the hHDC and S586A mutant in LbTDC strongly suggests that steric hindrance and hydrophobicity influence substrate specificity [31], [32]. However, there are also studies of a CrTDC G370S mutant and a CsG351S/L mutant conferring the TDC decarboxylation ability to L-DOPA and tyrosine [29], [70]. Combined with the fact that 5-hydroxytryptophan is configurationally larger than tryptophan, we concluded that the size of the substrate and the active site, together with the hydrogen bonds formed between enzymes and substrates, determine how TDC interacts with tryptophan and 5-hydroxytryptophan.
The overall structures of the selected PLP_DCs seem like with OsTDC (Fig. S3), however, there are considerable differences in activity [30], [31], [32], [33]. Consequently, their unique functions evolved require investigation. Therefore, 76 PLP_DCs in four types were analyzed and the results indicated that type II PLP_DCs could have originated from GAD and diverged in the course of evolution (Fig. 5). HDC and AADC emerged almost at the same time in early algae and achieved high gene-level amplification from terrestrial plants (Selaginella moellendorffii) approximately 300 Mya in the Permian (Fig. 5). Selective pressures in plants are unique, and therefore, may be driven by evolutionary trajectories of the metabolic enzymes in a lineage-specific manner [71]. It is estimated that atmospheric oxygen concentrations began to rise 2.5 billion years ago, which may have forced the earliest organisms to incorporate photosynthesis into their metabolic machinery to adapt to the environments [72]. Considering the transition from anaerobic metabolism to aerobic metabolism at the time and the reactive oxygen species that would be produced inevitably, the gene-level amplification of AADC in different plant species could be due to the need for different secondary metabolites to facilitate adaptation to the environment, as well as the maintenance of species specificity [71]. This finding could also reveal that L-aromatic amino acids can be used as precursors to multiple metabolic pathways for producing different secondary metabolites.
Conclusions
In summary, we have presented the crystal structure of a plant TDC from O. sativa and its complexes formed with different ligands, which provide structural-based evidence of a TDC-mediated dual melatonin biosynthesis pathway. The Y359 residue from the loop gate is a proton donor and forms a Lewis acid-base pair with SRO/TSS, which is associated with product release. The conserved H214 acts jointly with Y359 to facilitate proton transfer. Additionally, the 5-hydroxyl group of SRO form hydrogen bonds with the backbone carboxyl group of F104 and P105, which could limit its release. The PLP_DC originated from GAD could be a form of adaptive evolution in organisms to tolerate stressful environments. Structural analyses combined with evolutionary analyses provide an accurate characterization of the conserved TDC. Furthermore, the high sequence identity/similarity observed between OsTDC and other identified plant TDC makes it possible to generate more accurate structures of other plant TDC via protein modeling.
Protein data bank accession numbers
The atomic coordinates and structural factors for the structures of TDC have been deposited to the RCSB Protein Data Bank with the PDB ID 6KHO, 6KHP and 6KHN, respectively.
Compliance with Ethics Requirements
All Institutional and National Guidelines for the care and use of animals (fisheries) were followed.
Declaration of Competing Interest
The authors have no conflict of interest to declare.
Acknowledgements
This Project was funded by National Natural Science Foundation of China (31970596, 81703637) and the Open Project of State Key Laboratory of Natural Medicines (SKLNMKF201708). This research was also supported by the Natural Science Fund in Jiangsu Province (BK20170736). We thank the staffs from BL18U1/BL19U1 beamlines of National Facility for Protein Science in Shanghai (NFPS) at Shanghai Synchrotron Radiation Facility for assistance during data collection.
Footnotes
Peer review under responsibility of Cairo University.
Supplementary data to this article can be found online at https://doi.org/10.1016/j.jare.2020.06.004.
Contributor Information
Yucheng Zhao, Email: zhaoyucheng1986@126.com.
Zhixiong Zeng, Email: zengzx@mail.hzau.edu.cn.
Appendix A. Supplementary material
The following are the Supplementary data to this article:
References
- 1.Masana M.I., Sumaya I.C., Becker-Andre M., Dubocovich M.L. Behavioral characterization and modulation of circadian rhythms by light and melatonin in C3H/HeN mice homozygous for the RORbeta knockout. Am J Physiol Regul Integr Comp Physiol. 2007;292:2357–2367. doi: 10.1152/ajpregu.00687.2006. [DOI] [PubMed] [Google Scholar]
- 2.Paredes S.D., Sanchez S., Parvez H., Rodriguez A.B., Barriga C. Altered circadian rhythms of corticosterone, melatonin, and phagocytic activity in response to stress in rats. Neuro Endocrinol Lett. 2007;28:489–495. [PubMed] [Google Scholar]
- 3.Slominski A.T., Hardeland R., Zmijewski M.A., Slominski R.M., Reiter R.J., Paus R. Melatonin: A cutaneous perspective on its production, metabolism, and functions. J Invest Dermatol. 2018;138:490–499. doi: 10.1016/j.jid.2017.10.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Liu Y., Zhang L., Zhang H., Liu B., Wu Z., Zhao W. Exogenous melatonin modulates apoptosis in the mouse brain induced by high-LET carbon ion irradiation. J Pineal Res. 2012;52:47–56. doi: 10.1111/j.1600-079X.2011.00917.x. [DOI] [PubMed] [Google Scholar]
- 5.Dallaspezia S., Benedetti F. Melatonin, circadian rhythms, and the clock genes in bipolar disorder. Curr Psychiatry Rep. 2009;11:488–493. doi: 10.1007/s11920-009-0074-1. [DOI] [PubMed] [Google Scholar]
- 6.Zhang J.J., Meng X., Li Y., Zhou Y., Xu D.P., Li S. Effects of melatonin on liver injuries and diseases. Int J Mol Sci. 2017;18:673. doi: 10.3390/ijms18040673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Zhang S., Qi Y., Zhang H., He W., Zhou Q., Gui S. Melatonin inhibits cell growth and migration, but promotes apoptosis in gastric cancer cell line, SGC7901. Biotech Histochem. 2013;88:281–289. doi: 10.3109/10520295.2013.769633. [DOI] [PubMed] [Google Scholar]
- 8.Kaur H., Mukherjee S., Baluska F., Bhatla S.C. Regulatory roles of serotonin and melatonin in abiotic stress tolerance in plants. Plant Signal Behav. 2015;10 doi: 10.1080/15592324.2015.1049788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Arnao M.B., Hernandez-Ruiz J. Functions of melatonin in plants: a review. J Pineal Res. 2015;59:133–150. doi: 10.1111/jpi.12253. [DOI] [PubMed] [Google Scholar]
- 10.Arnao M.B., Hernandez-Ruiz J. Melatonin and its relationship to plant hormones. Ann Bot. 2018;121:195–207. doi: 10.1093/aob/mcx114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Park S., Back K. Melatonin promotes seminal root elongation and root growth in transgenic rice after germination. J Pineal Res. 2012;53:385–389. doi: 10.1111/j.1600-079X.2012.01008.x. [DOI] [PubMed] [Google Scholar]
- 12.Byeon Y., Park S., Lee H.Y., Kim Y.S., Back K. Elevated production of melatonin in transgenic rice seeds expressing rice tryptophan decarboxylase. J Pineal Res. 2014;56:275–282. doi: 10.1111/jpi.12120. [DOI] [PubMed] [Google Scholar]
- 13.Zhang N., Zhao B., Zhang H.J., Weeda S., Yang C., Yang Z.C. Melatonin promotes water-stress tolerance, lateral root formation, and seed germination in cucumber (Cucumis sativus L.) J Pineal Res. 2013;54:15–23. doi: 10.1111/j.1600-079X.2012.01015.x. [DOI] [PubMed] [Google Scholar]
- 14.Weissbach H., Redfield B.G., Axelrod J. Biosynthesis of melatonin: enzymic conversion of serotonin to N-acetylserotonin. Biochim Biophys Acta. 1960;43:352–353. doi: 10.1016/0006-3002(60)90453-4. [DOI] [PubMed] [Google Scholar]
- 15.Liu T., Borjigin J. N-acetyltransferase is not the rate-limiting enzyme of melatonin synthesis at night. J Pineal Res. 2005;39:91–96. doi: 10.1111/j.1600-079X.2005.00223.x. [DOI] [PubMed] [Google Scholar]
- 16.Back K., Tan D.X., Reiter R.J. Melatonin biosynthesis in plants: multiple pathways catalyze tryptophan to melatonin in the cytoplasm or chloroplasts. J Pineal Res. 2016;61:426–437. doi: 10.1111/jpi.12364. [DOI] [PubMed] [Google Scholar]
- 17.Tan D.X., Hardeland R., Back K., Manchester L.C., Alatorre-Jimenez M.A., Reiter R.J. On the significance of an alternate pathway of melatonin synthesis via 5-methoxytryptamine: comparisons across species. J Pineal Res. 2016;61:27–40. doi: 10.1111/jpi.12336. [DOI] [PubMed] [Google Scholar]
- 18.Murch S.J., KrishnaRaj S., Saxena P.K. Tryptophan is a precursor for melatonin and serotonin biosynthesis in in vitro regenerated St. John's wort (Hypericum perforatum L. cv. Anthos) plants. Plant Cell Rep. 2000;19:698–704. doi: 10.1007/s002990000206. [DOI] [PubMed] [Google Scholar]
- 19.Park S., Byeon Y., Back K. Transcriptional suppression of tryptamine 5-hydroxylase, a terminal serotonin biosynthetic gene, induces melatonin biosynthesis in rice (Oryza sativa L.) J Pineal Res. 2013;55:131–137. doi: 10.1111/jpi.12053. [DOI] [PubMed] [Google Scholar]
- 20.Park M., Kang K., Park S., Back K. Conversion of 5-hydroxytryptophan into serotonin by tryptophan decarboxylase in plants, Escherichia coli, and yeast. Biosci Biotechnol Biochem. 2008;72:2456–2458. doi: 10.1271/bbb.80220. [DOI] [PubMed] [Google Scholar]
- 21.Reiter R.J., Tan D.X., Zhou Z., Cruz M.H., Fuentes-Broto L., Galano A. Phytomelatonin: assisting plants to survive and thrive. Molecules. 2015;20:7396–7437. doi: 10.3390/molecules20047396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Sandmeier E., Hale T.I., Christen P. Multiple evolutionary origin of pyridoxal-5'-phosphate-dependent amino acid decarboxylases. Eur J Biochem. 1994;221:997–1002. doi: 10.1111/j.1432-1033.1994.tb18816.x. [DOI] [PubMed] [Google Scholar]
- 23.Okazaki Y., Shizuri Y. Identification of the aromatic L-amino acid decarboxylase (AADC) gene and its expression in the attachment and metamorphosis of the barnacle, Balanus amphitrite. Dev Growth Differ. 2001;43:33–41. doi: 10.1046/j.1440-169x.2001.00552.x. [DOI] [PubMed] [Google Scholar]
- 24.Zhu M.Y., Juorio A.V. Aromatic L-amino acid decarboxylase: biological characterization and functional role. Gen Pharmacol. 1995;26:681–696. doi: 10.1016/0306-3623(94)00223-a. [DOI] [PubMed] [Google Scholar]
- 25.Kubovcakova L., Krizanova O., Kvetnansky R. Identification of the aromatic L-amino acid decarboxylase gene expression in various mice tissues and its modulation by immobilization stress in stellate ganglia. Neuroscience. 2004;126:375–380. doi: 10.1016/j.neuroscience.2004.04.005. [DOI] [PubMed] [Google Scholar]
- 26.De Luca V., Marineau C., Brisson N. Molecular cloning and analysis of cDNA encoding a plant tryptophan decarboxylase: comparison with animal DOPA decarboxylases. Proc Natl Acad Sci U S A. 1989;86:2582–2586. doi: 10.1073/pnas.86.8.2582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Pang X., Wei Y., Cheng Y., Pan L., Ye Q., Wang R. The tryptophan decarboxylase in Solanum lycopersicum. Molecules. 2018;23:998. doi: 10.3390/molecules23050998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Facchini P.J., Huber-Allanach K.L., Tari L.W. Plant aromatic L-amino acid decarboxylases: evolution, biochemistry, regulation, and metabolic engineering applications. Phytochemistry. 2000;54:121–138. doi: 10.1016/s0031-9422(00)00050-9. [DOI] [PubMed] [Google Scholar]
- 29.Torrens-Spence M.P., Lazear M., von Guggenberg R., Ding H., Li J. Investigation of a substrate-specifying residue within Papaver somniferum and Catharanthus roseus aromatic amino acid decarboxylases. Phytochemistry. 2014;106:37–43. doi: 10.1016/j.phytochem.2014.07.007. [DOI] [PubMed] [Google Scholar]
- 30.Giardina G., Montioli R., Gianni S., Cellini B., Paiardini A., Voltattorni C.B. Open conformation of human DOPA decarboxylase reveals the mechanism of PLP addition to Group II decarboxylases. Proc Natl Acad Sci U S A. 2011;108:20514–20519. doi: 10.1073/pnas.1111456108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Komori H., Nitta Y., Ueno H., Higuchi Y. Structural study reveals that Ser-354 determines substrate specificity on human histidine decarboxylase. J Biol Chem. 2012;287:29175–29183. doi: 10.1074/jbc.M112.381897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zhu H., Xu G., Zhang K., Kong X., Han R., Zhou J. Crystal structure of tyrosine decarboxylase and identification of key residues involved in conformational swing and substrate binding. Sci Rep. 2016;6:27779. doi: 10.1038/srep27779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Williams B.B., Van Benschoten A.H., Cimermancic P., Donia M.S., Zimmermann M., Taketani M. Discovery and characterization of gut microbiota decarboxylases that can produce the neurotransmitter tryptamine. Cell Host Microbe. 2014;16:495–503. doi: 10.1016/j.chom.2014.09.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Fan J., Xie Y., Zhang Z., Chen L. Melatonin: a multifunctional factor in plants. Int J Mol Sci. 2018;19:1528. doi: 10.3390/ijms19051528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Xue J., Chen H., Wu J., Takeuchi M., Inoue H., Liu Y. Structure of the fission yeast S. pombe telomeric Tpz1-Poz1-Rap1 complex. Cell Res. 2017;27:1503–1520. doi: 10.1038/cr.2017.145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Minor W., Cymborowski M., Otwinowski Z., Chruszcz M. HKL-3000: the integration of data reduction and structure solution–from diffraction images to an initial model in minutes. Acta Crystallogr D Biol Crystallogr. 2006;62:859–866. doi: 10.1107/S0907444906019949. [DOI] [PubMed] [Google Scholar]
- 37.Winn M.D., Ballard C.C., Cowtan K.D., Dodson E.J., Emsley P., Evans P.R. Overview of the CCP4 suite and current developments. Acta Crystallogr D Biol Crystallogr. 2011;67:235–242. doi: 10.1107/S0907444910045749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.McCoy A.J., Grosse-Kunstleve R.W., Adams P.D., Winn M.D., Storoni L.C., Read R.J. Phaser crystallographic software. J Appl Crystallogr. 2007;40:658–674. doi: 10.1107/S0021889807021206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Emsley P., Cowtan K. Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr. 2004;60:2126–2132. doi: 10.1107/S0907444904019158. [DOI] [PubMed] [Google Scholar]
- 40.Adams P.D., Afonine P.V., Bunkoczi G., Chen V.B., Davis I.W., Echols N. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D Biol Crystallogr. 2010;66:213–221. doi: 10.1107/S0907444909052925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Schrodinger L. The PyMOL molecular graphics system, Version 2.3., https://www.pymol.org.
- 42.Zhao Y., Wang N., Zeng Z., Xu S., Huang C., Wang W. Cloning, functional characterization, and catalytic mechanism of a bergaptol O-methyltransferase from Peucedanum praeruptorum Dunn. Front Plant Sci. 2016;7:722. doi: 10.3389/fpls.2016.00722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Kumar S., Nei M., Dudley J., Tamura K. MEGA: a biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief Bioinform. 2008;9:299–306. doi: 10.1093/bib/bbn017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Letunic I., Bork P. Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Res. 2019;47:256–259. doi: 10.1093/nar/gkz239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Kumar R. Evolutionary trails of plant Group II Pyridoxal phosphate-dependent decarboxylase genes. Front Plant Sci. 2016;7:1268. doi: 10.3389/fpls.2016.01268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Kumar R., Jiwani G., Pareek A., SravanKumar T., Khurana A., Sharma A.K. Evolutionary profiling of Group II Pyridoxal-phosphate-dependent decarboxylases suggests expansion and functional diversification of histidine decarboxylases in Tomato. Plant Genome. 2016;9:1–15. doi: 10.3835/plantgenome2015.07.0057. [DOI] [PubMed] [Google Scholar]
- 47.El-Gebali S., Mistry J., Bateman A., Eddy S.R., Luciani A., Potter S.C. The Pfam protein families database in 2019. Nucleic Acids Res. 2019;47:427–432. doi: 10.1093/nar/gky995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Schultz J., Copley R.R., Doerks T., Ponting C.P., Bork P. SMART: a web-based tool for the study of genetically mobile domains. Nucleic Acids Res. 2000;28:231–234. doi: 10.1093/nar/28.1.231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Bremer B., Bremer K., Chase M.W., Fay M.F., Reveal J.L., Soltis D.E. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG III. Bot J Linn Soc. 2009;161:105–121. [Google Scholar]
- 50.Zeng L., Zhang Q., Sun R., Kong H., Zhang N., Ma H. Resolution of deep angiosperm phylogeny using conserved nuclear genes and estimates of early divergence times. Nat Commun. 2014;5:4956. doi: 10.1038/ncomms5956. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Wang W., Malcolm B.A. Two-stage PCR protocol allowing introduction of multiple mutations, deletions and insertions using QuikChange site-directed mutagenesis. Biotechniques. 1999;26:680–682. doi: 10.2144/99264st03. [DOI] [PubMed] [Google Scholar]
- 52.Kang S., Kang K., Lee K., Back K. Characterization of rice tryptophan decarboxylases and their direct involvement in serotonin biosynthesis in transgenic rice. Planta. 2007;227:263–272. doi: 10.1007/s00425-007-0614-z. [DOI] [PubMed] [Google Scholar]
- 53.Coon S.L., Roseboom P.H., Baler R., Weller J.L., Namboodiri M.A., Koonin E.V. Pineal serotonin N-acetyltransferase: expression cloning and molecular analysis. Science. 1995;270:1681–1683. doi: 10.1126/science.270.5242.1681. [DOI] [PubMed] [Google Scholar]
- 54.Han Q., Ding H., Robinson H., Christensen B.M., Li J. Crystal structure and substrate specificity of Drosophila 3,4-dihydroxyphenylalanine decarboxylase. PLoS ONE. 2010;5:8826. doi: 10.1371/journal.pone.0008826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Ishii S., Mizuguchi H., Nishino J., Hayashi H., Kagamiyama H. Functionally important residues of aromatic L-amino acid decarboxylase probed by sequence alignment and site-directed mutagenesis. J Biochem. 1996;120:369–376. doi: 10.1093/oxfordjournals.jbchem.a021422. [DOI] [PubMed] [Google Scholar]
- 56.Tan D.X., Hardeland R., Manchester L.C., Korkmaz A., Ma S., Rosales-Corral S. Functional roles of melatonin in plants, and perspectives in nutritional and agricultural science. J Exp Bot. 2012;63:577–597. doi: 10.1093/jxb/err256. [DOI] [PubMed] [Google Scholar]
- 57.Nawaz M.A., Huang Y., Bie Z., Ahmed W., Reiter R.J., Niu M. Melatonin: current status and future perspectives in plant science. Front Plant Sci. 2015;6:1230. doi: 10.3389/fpls.2015.01230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Byeon Y., Back K. An increase in melatonin in transgenic rice causes pleiotropic phenotypes, including enhanced seedling growth, delayed flowering, and low grain yield. J Pineal Res. 2014;56:408–414. doi: 10.1111/jpi.12129. [DOI] [PubMed] [Google Scholar]
- 59.Kang K., Lee K., Park S., Kim Y.S., Back K. Enhanced production of melatonin by ectopic overexpression of human serotonin N-acetyltransferase plays a role in cold resistance in transgenic rice seedlings. J Pineal Res. 2010;49:176–182. doi: 10.1111/j.1600-079X.2010.00783.x. [DOI] [PubMed] [Google Scholar]
- 60.Park S., Lee D.E., Jang H., Byeon Y., Kim Y.S., Back K. Melatonin-rich transgenic rice plants exhibit resistance to herbicide-induced oxidative stress. J Pineal Res. 2013;54:258–263. doi: 10.1111/j.1600-079X.2012.01029.x. [DOI] [PubMed] [Google Scholar]
- 61.Wang L., Zhao Y., Reiter R.J., He C.J., Liu G.S., Lei Q. Changes in melatonin levels in transgenic 'Micro-Tom' tomato overexpressing ovine AANAT and ovine HIOMT genes. J Pineal Res. 2014;56:134–142. doi: 10.1111/jpi.12105. [DOI] [PubMed] [Google Scholar]
- 62.Schneider G., Kack H., Lindqvist Y. The manifold of vitamin B-6 dependent enzymes. Structure. 2000;8:1–6. doi: 10.1016/s0969-2126(00)00085-x. [DOI] [PubMed] [Google Scholar]
- 63.Bertoldi M., Castellani S., Bori Voltattorni C. Mutation of residues in the coenzyme binding pocket of DOPA decarboxylase, Effects on catalytic properties. Eur J Biochem. 2001;268:2975–2981. doi: 10.1046/j.1432-1327.2001.02187.x. [DOI] [PubMed] [Google Scholar]
- 64.Burkhard P., Dominici P., Borri-Voltattorni C., Jansonius J.N., Malashkevich V.N. Structural insight into Parkinson's disease treatment from drug-inhibited DOPA decarboxylase. Nat Struct Biol. 2001;8:963–967. doi: 10.1038/nsb1101-963. [DOI] [PubMed] [Google Scholar]
- 65.Bertoldi M., Gonsalvi M., Contestabile R., Voltattorni C.B. Mutation of tyrosine 332 to phenylalanine converts DOPA decarboxylase into a decarboxylation-dependent oxidative deaminase. J Biol Chem. 2002;277:36357–36362. doi: 10.1074/jbc.M204867200. [DOI] [PubMed] [Google Scholar]
- 66.Fenalti G., Law R.H., Buckle A.M., Langendorf C., Tuck K., Rosado C.J. GABA production by glutamic acid decarboxylase is regulated by a dynamic catalytic loop. Nat Struct Mol Biol. 2007;14:280–286. doi: 10.1038/nsmb1228. [DOI] [PubMed] [Google Scholar]
- 67.Newmister S.A., Romminger S., Schmidt J.J., Williams R.M., Smith J.L., Berlinck R.G.S. Unveiling sequential late-stage methyltransferase reactions in the meleagrin/oxaline biosynthetic pathway. Org Biomol Chem. 2018;16:6450–6459. doi: 10.1039/c8ob01565a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Offen W., Martinez-Fleites C., Yang M., Kiat-Lim E., Davis B.G., Tarling C.A. Structure of a flavonoid glucosyltransferase reveals the basis for plant natural product modification. EMBO J. 2006;25:1396–1405. doi: 10.1038/sj.emboj.7600970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Tramonti A., De Biase D., Giartosio A., Bossa F., John R.A. The roles of His-167 and His-275 in the reaction catalyzed by glutamate decarboxylase from Escherichia coli. J Biol Chem. 1998;273:1939–1945. doi: 10.1074/jbc.273.4.1939. [DOI] [PubMed] [Google Scholar]
- 70.Kalb D., Gressler J., Hoffmeister D. Active-site engineering expands the substrate profile of the basidiomycete L-Tryptophan decarboxylase CsTDC. ChemBioChem. 2016;17:132–136. doi: 10.1002/cbic.201500438. [DOI] [PubMed] [Google Scholar]
- 71.Torrens-Spence MP, Chiang YC, Smith T, Vicent MA, Wang Y, Weng JK. Structural basis for independent origins of new catalytic machineries in plant AAAD proteins. bioRxiv 2018:404970. [DOI] [PMC free article] [PubMed]
- 72.Manchester L.C., Coto-Montes A., Boga J.A., Andersen L.P.H., Zhou Z., Galano A. Melatonin: an ancient molecule that makes oxygen metabolically tolerable. J Pineal Res. 2015;59:403–419. doi: 10.1111/jpi.12267. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.