Abstract
Alveolar macrophages are the first line of host defence against mycobacteria, but an insufficient host response allows survival of bacteria within macrophages. We aimed to investigate the role of Toll-like receptor 9 (TLR9) activation in macrophage defence against mycobacteria. Human in vitrodifferentiated macrophages as well as human and mouse alveolar macrophages showed TLR9 mRNA and protein expression. The cells were markedly activated by DNA isolated from attenuated mycobacterial strains (H37Ra and Mycobacterium bovis BCG) as assessed by measuring cytokine expression by real-time PCR, whereas synthetic phosphorothioate-modified oligonucleotides had a much lower potency to activate human macrophages. Intracellular replication of H37Ra was higher in macrophages isolated from TLR9-deficient mice than in macrophages from wild-type mice, whereas H37Rv showed equal survival in cells from wild-type or mutant mice. Increased bacterial survival in mouse macrophages was accompanied by altered cytokine production as determined by Luminex bead assays. In vivo infection experiments also showed differential cytokine production in TLR9-deficient mice compared to wild-type animals. Both human monocyte-derived macrophages as well as human alveolar macrophages showed reduced activation upon treatment with DNA isolated from bacteria from virulent (M. bovis and H37Rv) compared to attenuated mycobacteria. We suggest attenuated TLR9 activation contributes to the insufficient host response against virulent mycobacteria.
Key Words: Pathogenesis; Oligonucleotide; Pathogen-associated molecular patterns; Toll-like receptor; Virulence factors; Tuberculosis; Host defence; Inflammation, lung; Macrophages; Mycobacteria
Introduction
Myobacterium tuberculosis is an intracellular pathogen that can persist and propagate within human macrophages. Control and resolution of mycobacterial infections has been shown to depend on a coordinated innate and adaptive immune response, primarily involving macrophages and CD4+ T cells, respectively [1]. Over the last several years, the Toll-like receptors (TLR) have emerged as important transducers of the innate immune response [2]. Although DNA isolated from mycobacteria was the first DNA to be shown to induce immunostimulatory action in mammalian cells more than 20 years ago [3, 4], the relevance of TLR activation by mycobacterial DNA during tuberculosis infection has only recently been a matter of interest [5].
Bacterial DNA is now recognized as the natural ligand for TLR9 [6]. What distinguishes bacterial from mammalian DNA is its highly increased content of under-methylated 5’-CG-3’ sequences, usually referred to as CpG. The frequency of the base sequence CG is suppressed in mammals.
In addition to these differences in the frequency of CpG sequences between bacterial and mammalian DNA, there are also differences in the methylation state of the DNA. Whereas most CpG sequences in mammalian DNA are methylated, practically all bacterial CpG sequences are non-methylated. Non-methylated CpG sequences within defined motifs have been shown to bind to and signal via intracellular TLR9 [7, 8].
Whether human monocytes or human macrophages express functional TLR9 and therefore can be activated by bacterial DNA or synthetic CpG oligonucleotides has been controversial [9, 10, 11, 12]. Evidence that these mononuclear phagocytes do not express TLR9 was supported by a recent study showing lack of responsiveness of primary monocyte-derived macrophages and THP-1 cells towards CpG oligonucleotides [13]. While activation of human monocytes by CpG had been described [14], this activation was suggested to depend on a contamination of the primary monocyte preparation with plasmacytoid dendritic cells [15]. Other recent data, however, argue in favour of the presence of TLR9 in cells of the human monocytic lineage [16, 17, 18, 19] and reveal that bacterial DNA might represent a much better stimulus for TLR9 activation in macrophages than CpG oligonucleotides [20]. Nevertheless, there remains significant controversy regarding whether human macrophages express functional TLR9.
Because TLR9 activation by mycobacterial DNA could influence the innate response of macrophages to infection, we sought to clarify the expression of TLR9 on human and murine macrophages as well as the impact of mycobacterial DNA on macrophage function. Our data revealed a high sensitivity of macrophages to mycobacterial DNA compared to synthetic immunostimulatory oligonucleotides. Most interestingly, DNA from virulent mycobacterial strains showed a markedly reduced potency to activate macrophages than DNA from attenuated mycobacterial strains. These observations might help explain the attenuated macrophage activation upon infection with virulent mycobacteria.
Materials and Methods
Materials
Mouse IgG2a isotype (clone HOPC-1) was obtained from Southern Biotechnology, human IgG, biotinylated goat anti-mouse F(ab’) fragment and Cy5- or PE-labelled streptavidin were from Jackson Immunolab and FITC-labelled anti-CD14, PE-labelled anti-human TLR9 (eB72-1665), FITC-IgG2a and PE-IgG2a κ isotype controls from eBioscience. Anti NF-κB/p65 was purchased from Santa Cruz Biotechnology. IRDyeTM 800 conjugated anti-mouse IgG and anti-β-actin were purchased from Rockland. IRDye 680 conjugated anti-rabbit IgG was from LI-COR Biosciences. Chloroquine (CQ, diphosphate salt; Sigma) was dissolved in water at a stock concentration of 100 mM, syringe filtered (0.2 µM), aliquoted and stored at −20° until use. CQ did not impair macrophage viability as assessed by the MTT test performed as described previously (data not shown) [21].
Bacterial Culture
Mycobacteria (Mycobacterium bovis BCG Pasteur ATCC 27289, wild-type M. bovis ATCC19210, H37Rv ATCC 27294 and H37Ra ATCC 25177) were grown in Middlebrook 7H9 broth containing 10% ADC, 0.2% glycerol and 0.05% Tween 80 (7H9-ADCT) or on Middlebrook 7H10 agar containing OADC (Becton Dickinson), 0.5% glycerol and antifungal cycloheximide (100 µg ml–1; Sigma-Aldrich). Antibiotics included hygromycin (50 µg ml–1) and kanamycin (25 µg ml–1).
Cell Culture
M-CSF-Differentiated Macrophages
Peripheral blood was obtained after informed consent from normal volunteers at the General Clinical Research Center at The Scripps Research Institute. Alternatively, buffy coats from the blood bank Saarbrücken were used. Mononuclear cells were isolated from the blood as previously described [22] and then differentiated into macrophages by treatment with 50 ng/ml M-CSF (Promocell) in RPMI-1640 containing 1% FCS or in RPMI-1640 containing 20% FCS for 5–7 days. Cells were seeded into 12- or 24-well plates and treated with DNA in triplicates, followed by RNA isolation and real-time PCR analysis. Three independent cell isolations were performed for each treatment.
THP-1 Cells
THP-1 cells were cultivated in RPMI-1640 with 10% FCS and differentiated by treatment with 100 nM PMA for 48 h.
Human Alveolar Macrophages
Human alveolar macrophages were obtained by bronchoalveolar lavage (BAL) after informed consent from normal volunteers at the General Clinical Research Center at The Scripps Research Institute or from surgical samples from the SHG Kliniken Völklingen. Four successive 60-ml lavages were performed with 37°C sterile normal saline solution. BAL fluid was filtered through a sterile 100-µm mesh filter (Falcon) in wash buffer containing DNase. BAL cells were cultured in RPMI-1640 supplemented with 50% FCS, penicillin/streptomycin and amphotericin for 2 h at 37°C before removing non-adherent cells. Cells were cultured in 12- or 24-well plates overnight and treated with DNA in triplicates followed by RNA isolation and real-time PCR analysis or by immunostaining. Five independent cell isolations were performed.
Mouse Alveolar Macrophages
C57BL/6J mice were obtained from the Jackson Laboratory and housed under specific pathogen-free conditions in the animal resource centre at the La Jolla Institute for Allergy and Immunology. TLR9CpG1/CpG1 mice were maintained under pathogen-free conditions in The Scripps Research Institute animal care facility. All mice used in the experiments were 8–10 weeks of age. All experimental procedures were conducted in accordance with the institutional guidelines for animal care and use. Alveolar macrophages were obtained by BAL via the trachea. Briefly, mice were anaesthetized, the trachea was exposed and a disposable plastic cannula with a 1.0-ml syringe attached was inserted through an incision immediately posterior to the larynx. The respiratory tract was then washed out in a standard, reproducible manner with 1 ml PBS + 5% FCS which was infused and withdrawn several times. Erythrocytes were lysed with Sigma lysis solution and viable cell counts were determined by Trypan blue exclusion. Alveolar macrophages from 4 mice per strain were pooled and cultured at a concentration of 60,000 cells/well in LabTek chambers in RPMI-1640 containing 10% FCS and antibiotics. After 2 h of adherence, media were replaced and the cells were incubated overnight. The next day, macrophages were treated with 1 µg/ml LPS or 5 µg/ml BCG DNA, stained, then analysed. Two independent cell isolations were performed for each strain.
Mouse Bone Marrow Macrophage Cultivation and Mycobacterial Infection
Macrophages were prepared essentially as described by Kiemer and Vollmar [23], whereby cells were differentiated by adding 10 ng/ml recombinant mouse M-CSF (R&D Systems) to the media for 6 days. Cells were suspended at a density of 6 × 105/ml in DMEM supplemented with 10% FCS, 10 mM HEPES, 0.06 g/l l-glutamine, 1 mM pyruvate and penicillin/streptomycin, and were plated into 24-well plates at 500 µl/well. Antibiotics were taken off the cells 3 days before the experiments. Differentiated cells were activated with 1 × 106 units/ml recombinant interferon-γ (IFN-γ [R&D Systems) for 24 h and mycobacteria (virulent H37Rv or attenuated H37Ra) were added at an MOI of 2:1. After 4 h of infection, macrophages were washed 3 times with DMEM to remove extracellular bacteria. Mock-infected (PBS) macrophages served as negative controls for each experiment. The culture supernatant of macrophages was collected after 24 h and 7 days, filtered through a 0.2-µM pore membrane, and stored at −80°C until analysis. Tumour necrosis factor-α (TNF-α) and MCP-1 levels in the supernatants were measured by a Luminex bead assay (Upstate Beadlyte) and nitrite concentrations by the Griess Reagent System (Promega). Macrophages were pooled from 6–9 mice per strain. Intracellular uptake and replication of the bacteria were determined by lysing macrophages with 200 µl of PBS/0.5% Triton X-100 and plating serial dilutions on 7H11 agar to enumerate CFU. This was performed at 4 h (day 0), day 3 and day 7. CFU were enumerated after 3–4 weeks of plate incubation.
Mouse Infection
Mice were obtained and housed as described above. Eight- to ten-week-old TLR9CpG1/CpG1 and C57BL/6 mice were infected with either M. tuberculosis H37Rv or M. tuberculosis H37Ra via the inhalation route using the Inhalation Exposure System (Glas-col). Twenty-four hours after the infection, the right lungs of 3 mice (per each infection per mouse strain) were harvested, homogenized, and the whole homogenate was plated on 7H11-OADC plates to determine the dose of infection. Average M. tuberculosis H37Rv CFU in TLR9CpG1/CpG1 or C57BL/6 lungs were 128 and 181, respectively. Average M. tuberculosis H37Ra CFU in TLR9CpG1/CpG1 or C57BL/6 lungs were 201 and 211, respectively. In our previous experiments with similar doses, bacteria were not found in liver and lungs 24 h after the infection. Therefore, CFU numbers per right lung × 2 is an estimation of bacterial load per mouse. In this regard, these bacterial loads per mouse are somewhat lower than the high-dose inoculom infection described in a recent study [5].
CFU in lung, spleen and liver at later time points were determined by plating appropriately diluted organ homogenates. CFU on 7H11-OADC were counted 3–4 weeks after plating.
Loss of weight accompanied by failure to groom, ruffled fur and lethargy were used to determine the endpoint of morbidity, in addition (in some instances) to the recommendation of the veterinary staff of the North Animal Facility of UC Berkeley. Health of the mice was monitored daily by the veterinary staff. Statistical analysis of survival curve was performed by Kaplan-Meier plot. Differences were considered significant at p < 0.05.
Homogenates from day 0 and lungs at 18 days after infection were filtered through a 0.2-µM pore membrane and stored at −80°C until analysis. TNF-α, interleukin-1β (IL-1β), IFN-γ and IL-6 levels in the supernatants were measured by a Luminex bead assay (Upstate Beadlyte).
Monoclonal Antibody 2-1E2 Production
For immunization with TLR9 protein, a fusion protein consisting of the extracellular domain of human TLR9 (amino acids 1–815) and human IgG1Fc was constructed, stably expressed in HEK293 cells and purified from cell lysates by protein A affinity chromatography. C57BL/6 mice were injected 3 times with 20 µ g human TLR9 protein mixed with 10 nmol of CpG-ODN 1668 and fusion was performed according to standard protocols. Screening and cloning of hybridomas led to the generation of antibodies reacting against TLR9. The antibody 2–1E2 is of the IgG2a isotype and recognizes human TLR9. Specificity of the antibody was tested by transfecting COS-1 or CHO-K1 cells with either a pcDNA3.1 vector encoding human TLR9, with a vector encoding TLR9 attached to a C-terminal YFP tag [8] or an empty control vector followed by Western blot or FACS analysis 24 h after transfection. COS-1 cells were transfected using the Nucleofector II device and the Cell Line Nucleofector Kit V (Amaxa) and CHO-K1 cells were transfected using the Fugene HD Transfection Reagent (Roche) according to the manufacturers’ instructions.
Western Blotting
Cells were lysed in modified RIPA buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1 mM EDTA, 1% EDTA, 1% Triton X-100, 0.2% sodium deoxycholate, 0.1% SDS, 5 µg/ml aprotinin and 5 µg/ml leupeptin) and stored at −80°C. For Western blot analysis, the samples were boiled with sample buffer, separated by SDS-PAGE on 8% gels, and then transferred to Immobilon FL-PVDF membrane (Millipore). The membranes were blocked in blocking buffer for near-infraded Western blotting (Rockland) overnight, incubated for 3 h with mouse anti-human TLR9 (clone 2-1E2), washed 3 times, and incubated with IRDye 800 conjugated anti-mouse IgG (Rockland) for 1.5 h. After washing, blots were scanned with an Odyssey Infrared Imaging System (LI-COR Biosciences). Actin served as loading control.
RNA Isolation and Reverse Transcription
Total RNA was extracted using either RNA Stat-60 (Iso-Tex Diagnostics Inc.) or RNeasy mini or micro kit columns (Qiagen). DNA was digested during the RNA isolation procedure using the RNase-free DNase 1 treatment kit (Qiagen). One microgram of RNA was denatured at 65°C for 5 min, placed on ice, and then reverse transcribed in a total volume of 20 µl using random hexamers, dNTPs, RNase inhibitor and 1 U of Superscript II reverse transcriptase (Life Technologies) for 1 h at 42°C.
RT-PCR and Real-Time Quantitative PCR
The iCycler iQ (Bio-Rad) was used for real-time quantitative PCR. The primers used are given in the online suppl. table 1 (www.karger.com/doi/10.1159/000142731). Normalization was done on β-actin and GAPDH. Dual-labelled probes were obtained from BioSearch Technologies (Novato). Standards, from 10 to 0.0001 amol of the PCR product cloned into pGEM-T Easy (Promega), were run alongside the samples to generate a standard curve. All samples and standards were analysed in triplicate. The PCR reaction mixture consisted of 1.5 mM Tris-HCl, 5 mM KCl, 2 mM dNTPs, 3 or 4 mM Mg2+, 100 ng of sense and antisense primers, either 2.5 (TNF-α, IL-10, IL-12) or 1.5 pmol (GAPDH, β-actin, IP10, TLR2, TLR4, TLR9) of the respective dual-labelled probe, and 1 U of AmpliTaq gold (Applied Biosystems) in a total volume of 25 µl. The reaction conditions were 95°C for 10 min followed by 40 cycles of 30 s at 94°C, 30 s at 60°C and 30 s at 72°C. The starting amount of cDNA in each sample was calculated using the iCycler iQ software package (Bio-Rad). Qualitative RT-PCR was performed under the same conditions in a Bio-Rad iCycler thermal cycler. Reaction products were run on agarose gel and stained with ethidium bromide.
DNA Isolation, Digestion and Methylation
Before DNA isolation, bacteria were centrifuged and boiled for 10 min. DNA was isolated according to a previously published method [24]. Briefly, bacteria from cultures at an OD600 of 1 were pelleted and resuspended in TE buffer. TE-saturated phenol pH 8 and glass beads were added and the suspension was vortexed for 3 min. The phenol phase was re-extracted with TE, followed by the addition of 0.2 vol. 5% sodium deoxycholate and incubation at 56°C for 90 min. Protein was removed by at least 3 cycles of extraction with phenol:chloroform:isoamyl. DNA was precipitated by the addition of sodium acetate/ethanol, washed in 70% ethanol and resuspended in Tris buffer. Any residual RNA was digested with Qiagen RNase and residual protein was removed by chloroform:isoamyl extraction. DNA was again precipitated, washed and resuspended in Tris buffer. The isolation was performed under sterile conditions in order to avoid bacterial contamination from the surrounding area. Additional precipitation and washing steps including purification with Triton X-114 were included to assure purity of the DNA [25]. We checked all DNA preparations with a commercially available limulus amoebocyte lysate assay (Bio Whittaker) in order to exclude LPS contaminations. Three different DNA preparations were used for macrophage activation studies. For further control experiments, DNA was either digested with DNase or cytosine residues (C5) were methylated by the SssI methyltransferase [26].
DNA was digested with DNase (500 µg/ml; Sigma) for 2 h at 37°C under sterile conditions. After digestion, DNase was heat inactivated and the required volume of the reaction correlative with the amount of DNA digested was added to a culture of monocyte-derived macrophages. In addition, mock-treated H37Ra-DNA, which was homologously treated in the absence of DNase, was added to cells. After 2 h of incubation, RNA was isolated and real-time PCR analysis was performed.
Cytosine residues of BCG DNA were methylated by SssI methyltransferase (New England Biolabs) under sterile conditions according to the manufacturer's instructions. In order to determine whether methylation in fact occurred, digestion was performed with HpaII as a restriction enzyme sensitive to CG methylation. After methylation, SssI was heat inactivated and the required volume of the reaction correlative with the amount of DNA methylated was added to a culture of monocyte-derived macrophages. In addition, BCG DNA, which was homologously treated in the absence of SssI, was added to cells. After 2 h of incubation, RNA was isolated and real-time PCR analysis was performed.
Cell Staining
Cells were fixed in 4% formaldehyde in PBS for 15 min at room temperature and permeabilized in FACS buffer containing 0.5% saponin and 0.5% BSA. Unspecific binding was blocked by addition of human IgG, followed by treatment with mouse anti-human TLR9, biotin-goat anti-mouse F(ab’)2, streptavidin-PE or streptavidin-Cy5, and FITC-mouse anti-human CD14 or the respective isotypes or by the use of PE-labelled anti-TLR9 antibody. For FACS analysis, cells were detached from the plates in TEN buffer (40 mM Tris, 1 mM EDTA, 150 mM NaCl, macrophages) or with trypsin-EDTA (COS-1) before staining and analysed in a Becton Dickinson FACS Calibur flow cytometer after staining. NF-κB (p65) staining was done in 0.2% Triton X-100-permeabilized cells counterstained by Toto-3 in PBS [27]. For confocal microscopy and laser scanning, cytometry cells were grown in LabTek chambers (Nunc) and mounted in Fluoromount-G/propidium iodide followed by analysis with a Bio-Rad 1024 confocal microscope and a CompuCyte LSC laser scanning cytometer. The microscope was a Bio-Rad MRC1024 laser scanning confocal microscope, equipped with a krypton/argon mixed gas laser with excitation wavelengths of 488, 568 and 647 nm. It was attached to a Zeiss Axiovert S100TV inverted microscope with infinity-corrected optics. A C-Apo 40× objective was used to analyse labelled cells. Images were collected using LaserSharp (version 3.2) software (Bio-Rad) and processed with Photoshop software (Adobe). A CompuCyte laser scanning cytometer equipped with a 20-mW argon-ion air-cooled laser and a 5-mW HeNe laser equipped with a DP11 digital camera (Olympus) were used to measure Cy5 fluorescence. Scanning was done using the 20× objective. Propidium iodide fluorescence was used as the contouring parameter.
Results
Human Macrophages Express TLR9 mRNA and Protein and Are Responsive to Bacterial DNA
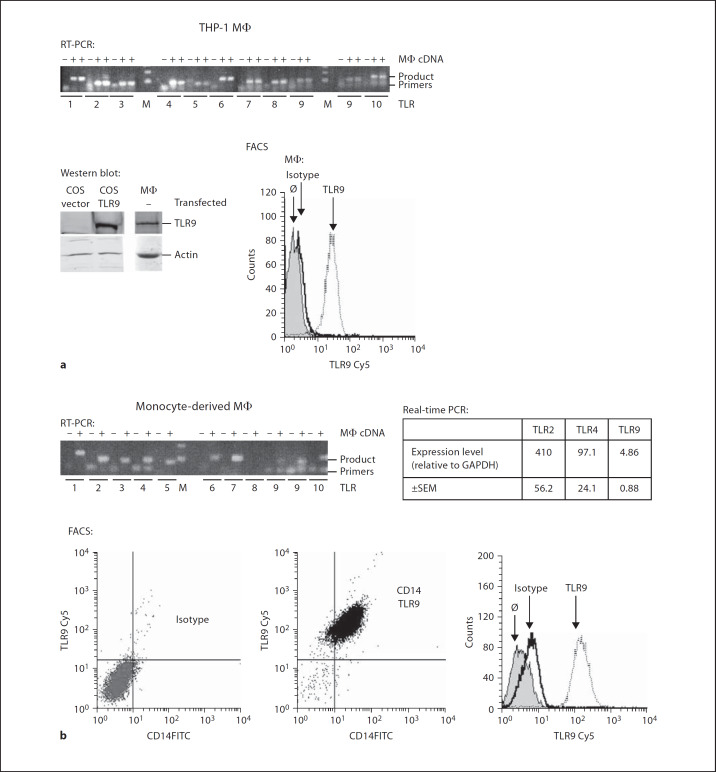
mRNA expression of all human TLRs (TLR1–10) was detected by conventional RT-PCR in PMA-differentiated THP-1 macrophages (fig. 1a) [28]. Expression of TLR9 protein was then investigated by FACS analysis and Western blotting, employing a custom antibody against human TLR9. To demonstrate antibody specificity, COS-1 or CHO cells were transfected with a plasmid for untagged or YFP-linked TLR9 or an empty vector, then analysed for TLR9 expression. TLR9 expression was detected by FACS analysis in TLR9-transfected but not in empty vector-transfected cells (data not shown). Similarly, immunoblotting for TLR9 showed a band only in TLR9-transfected COS-1 cells (fig. 1a). In contrast, THP-1 macrophages showed evidence of TLR9 expression by Western blot and FACS analysis (fig. 1a).
Fig. 1.
Human monocyte-derived macrophages express TLR9. THP-1 cells were differentiated by PMA (a) and primary monocytes were differentiated into macrophages by M-CSF treatment (b). RT-PCR and real-time PCR: RNA isolation, reverse transcription and qualitative RT-PCR for TLRs1–10 as well as quantitative real-time PCR for TLR2, TLR4 and TLR9 were performed as described in Materials and Methods. Qualitative RT-PCR was performed in the presence (+) or absence (–) of cDNA. Reaction products of qualitative RT-PCR reactions and a size marker (M) were electrophoresed in agarose and stained with ethidium bromide. Expression levels of TLR2, TLR4 and TLR9 are expressed as mRNA copies of the respective gene per 1,000 copies of GAPDH. Quantification was performed by comparing starting amounts of cDNAs with a standard curve of amplicon cloned into pGEM-T Easy. Data represent means ± SEM of cells from 5–7 donors. Western blot: COS-1 cells were transfected with a pcDNA3.1 vector encoding TLR9 or a vector control. Transfected cells as well as human macrophages were lysed. TLR9 was detected by Western blot by a custom antibody and visualized by an IRDye 800 conjugated anti-mouse IgG. Actin served as a loading control. FACS: Primary macrophages and differentiated THP-1 cells were stained with a custom antibody against TLR9 followed by detection with a secondary antibody via biotin/streptavidin/Cy5 and an FITC-labelled commercial antibody against CD14 as described in Materials and Methods. Cells were analysed by flow cytometry. Data show 1 representative of 3 independent experiments with cells from different donors.
We then analysed expression of TLR9 on human monocyte-derived macrophages. The primary macrophages displayed mRNA expression for all human TLRs (TLR1–10) as detected by conventional RT-PCR (fig. 1b). In order to estimate the expression level of TLR9 in primary macrophages, we performed quantitative real-time PCR analysis using a standard curve of the amplicon cloned into pGEM-T Easy. The assay was linear over 7 orders of magnitude in a range between 100 and 1 × 10–4 amol cDNA/reaction. Normalizing TLR expression to the housekeeping gene GAPDH demonstrated marked differences in the amount of mRNA expressed, with TLR2 showing the highest and TLR9 the lowest expression levels (fig. 1b). TLR9 mRNA was detected by quantitative RT-PCR in all macrophage preparations tested (7 separate experiments). Since CD14 has been shown to be expressed by monocyte-derived human macrophages but not by dendritic cells [29], primary macrophages were co-stained with anti-TLR9 and anti-CD14 for FACS analysis. M-CSF-differentiated CD14+ macrophages showed a marked expression of TLR9 as assessed by FACS staining for TLR9 (fig. 1b). CD14+ monocytes, obtained from peripheral blood, also showed staining for TLR9 (data not shown). A commercially available antibody (eBioscience) gave similar results (data not shown).
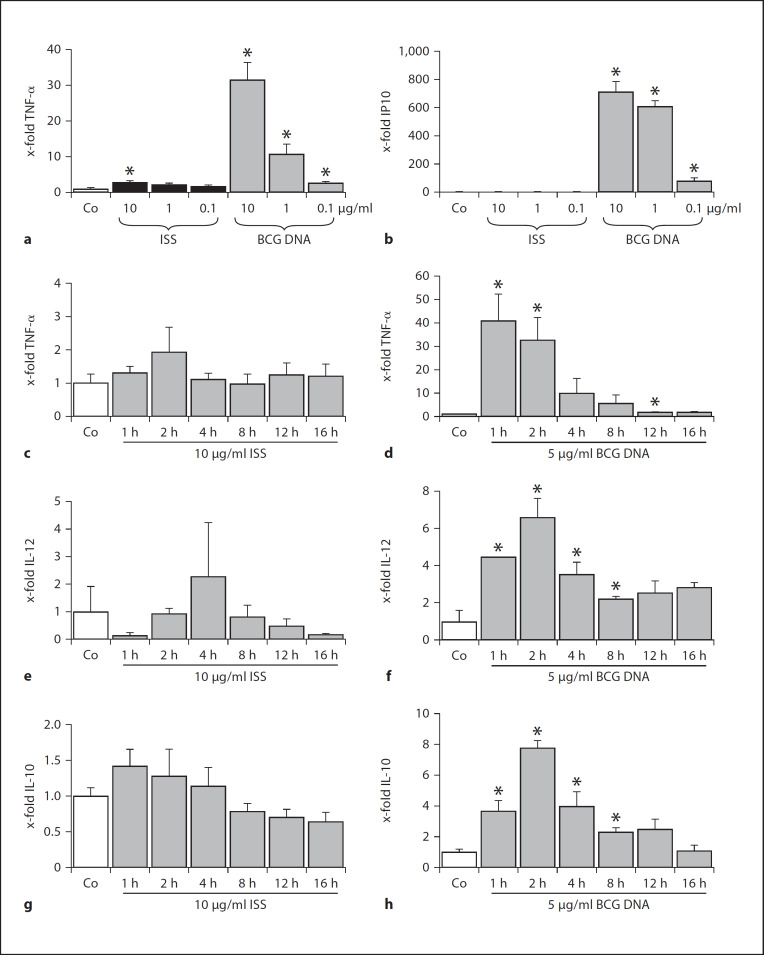
We next tested whether TLR9 in human macrophages was functional by determining whether macrophages were responsive to stimulation with TLR9 ligands. Monocyte-derived human macrophages were treated with different concentrations of a CpG-containing oligonucleotide [phosphorothioate-modified immunostimulatory sequence (ISS) 1018, 5’-TGACTGTGAACGTTCGAGATGA-3’], then induction of cytokine mRNA was measured by quantitative RT-PCR. While we observed a dose-dependent increase in TNF-α mRNA (fig. 2a), the extent of cytokine induction observed was relatively weak compared to the increase in cytokine induction in other TLR9 ligand-responsive cells [30]. Employment of a different ISS (ODN 2006, 5’-tcg tcg ttt tgt cgt ttt gtc gtt-3’) also resulted in a low but significant induction of TNF-α at a concentration of 10 µg/ml, whereas the respective GC control (sequence: 5’-tgc tgc ttt tgt gct ttt gtg ctt-3’; both purchased from Invivogen) did not induce TNF-α in the highest concentration tested (data not shown). This suggests that the activatory potential of ISS is not due to an unspecific effect of phosphorothioate groups. Since bacterial DNA was recently reported to be a much stronger inducer of TLR9-dependent responses in macrophages compared to CpG-containing oligonucleotides [20], we assessed the impact of genomic DNA (isolated from the attenuated M. bovis BCG strain) as an alternative activator of TLR9 [4]. Interestingly, we saw a markedly higher TNF-α response in human macrophages treated with BCG DNA compared to treatment with ISS (fig. 2a). We also confirmed that stimulation of macrophages with BCG DNA caused a marked induction of other cytokine mRNAs, such as IP10 as well as IL-12 and IL-10 (fig. 2b, e-h). Indeed, the induction of IP10 mRNA by BCG DNA was even greater than that observed for TNF-α (fig. 2b). The differences in the extent of activation seen between BCG DNA and ISS DNA were not due to differences in the kinetics of the cytokine mRNA induction, since induction time patterns were comparable for ISS and BCG DNA (fig. 2c, d). Although we did observe substantial inter-experiment variability in the extent of macrophage activation depending on the blood donor, the intra-experiment variability was low. Thus, we always stimulated macrophages with ISS and mycobacterial DNA on the same day using the same macrophage preparation in order to allow direct comparison (fig. 2). In general, all experiments show means from 3 cell preparations, each performed in triplicate.
Fig. 2.
Activation of human monocyte-derived macrophages by mycobacterial DNA. Primary monocyte-derived macrophages were left untreated (Co) or treated with ISS (ISS 1018, phosphorothioate-modified oligonucleotide 5’-TGACTGTGAACGTTCGAGATGA-3’) or genomic DNA from M. bovis BCG. a, b Cells were treated for 3 h in the indicated concentrations (0.1–10 µg/ml). c, d Cells were treated for the indicated times (1–16 h) in the indicated concentrations (5 or 10 µg/ml). After treatment, RNA was isolated followed by real-time PCR analysis for TNF-α (a, c, d), IP10 (b), IL-12 (e, f) or IL-10 (g, h). Values for cytokines were normalized to the housekeeping gene GAPDH. Data are expressed as x-fold TNF-α or IP10 induction with expression levels of untreated cells set as 1. Data show means ± SEM of 3 experiments performed in triplicates. * p < 0.05 significantly different from Co (t test).
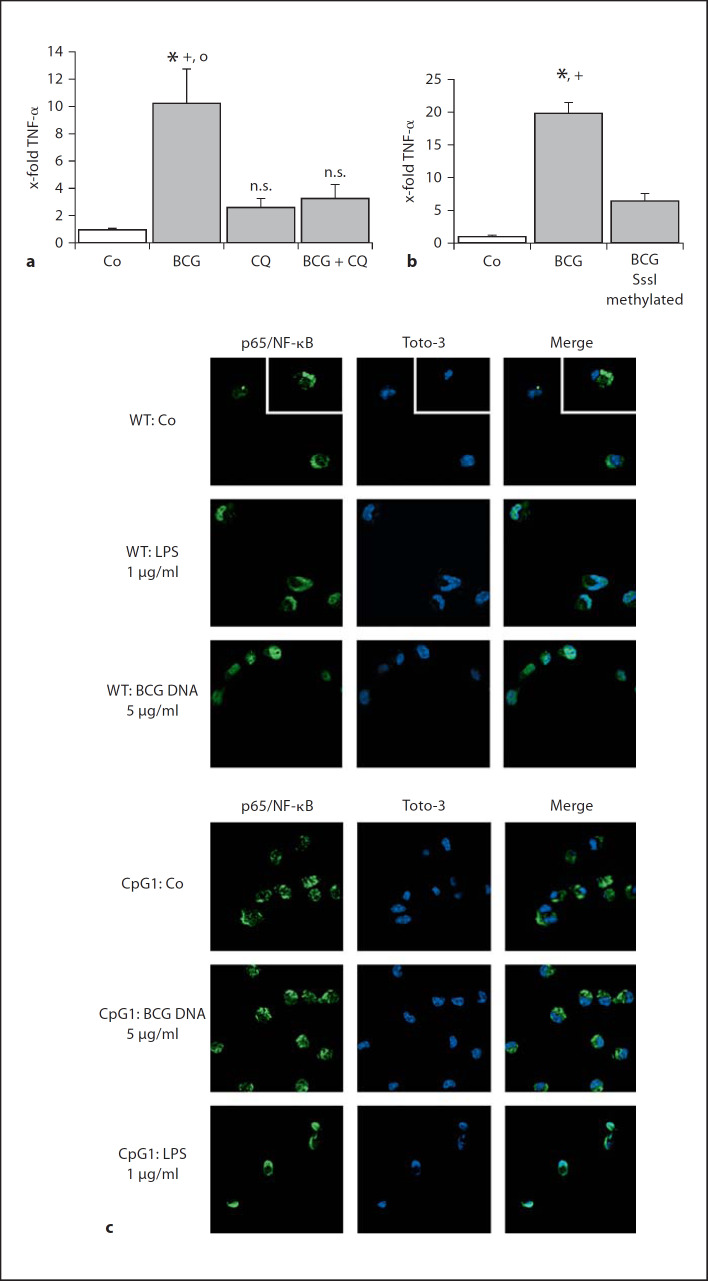
We then performed several experiments to demonstrate that the activation of human macrophages by BCG DNA was due to DNA and not evoked by contaminants. Potential contamination of the DNA preparations with endotoxin was assessed using a commercially available endotoxin assay (limulus amoebocyte lysate) with a detection sensitivity of 0.03 EU/ml. None of our DNA preparations showed any reactivity using this sensitive assay. Additional controls were performed in order to exclude effects from other potential contaminants, most likely TLR2 agonists as mycobacterial cell wall components. First, we co-treated cells with the TLR9 inhibitor CQ [31, 32], which does not interfere with TLR2 activation [33], together with BCG DNA. CQ completely abrogated TNF-α induction induced by BCG DNA treatment, whereas it had no effect when given to the cells alone (fig. 3a). As a complementary control, we methylated cytosine residues (C5) on BCG DNA with SssI methylase. As expected, methylation of the cytosine residues largely abrogated TNF-α mRNA induction by our DNA preparation (fig. 3b). Taken together, these experiments strongly support the interpretation that it is the DNA itself that mediates the profound effect of mycobacterial DNA preparations on macrophage activation, and that this effect is transduced through its interaction with TLR9.
Fig. 3.
Activation by DNA preparations is specific for non-methylated DNA. a, b DNA was isolated from M. bovis BCG mycobacteria and added to primary monocyte-derived macrophages for 3 h followed by RNA isolation and real-time PCR analysis for TNF-α. a Cells were either left untreated (Co), or treated with CQ (10 µM), BCG DNA (5 µg/ml) or a combination of both, whereby CQ was added to the cells 1 h before BCG DNA. * p < 0.05 significantly different from Co; + p < 0.05 significantly different from CQ; ° p < 0.05 significantly different from BCG + CQ. n.s. = Not significantly different from Co (t test). b DNA from BCG mycobacteria was methylated with SssI methylase. In vitro differentiated macrophages were either left untreated (Co) or treated with methylated or mock-treated DNA. Expression levels for TNF-α were normalized on housekeeping genes and data are expressed as x-fold TNF-α induction. Data show means ± SEM of 3 experiments, each performed in triplicates. * p < 0.05 significantly different from Co; + p < 0.05 significantly different from BCG SssI methylated (t test). c Mouse alveolar macrophages from wild-type (WT) or TLR9CpG1/CpG1 (CpG1) mice were either left untreated (Co) or treated with LPS (1 µg/ml) or BCG DNA (5 µg/ml) before staining of p65/NF-κB (green) and nuclear staining with Toto-3 (blue) as described in Materials and Methods. Data show channels for each of the single fluorophores (left and middle panel) and a merged overlay (right panel).
Expression of TLR9 in Murine and Human Alveolar Macrophages
TLR9CpG1/CpG1 mice are on a pure C57BL/6 background, but have a point mutation in the TLR9 gene that renders it non-responsive to TLR9 agonists [34]. We therefore utilized homozygous TLR9CpG1/CpG1 mice to investigate the requirement for functional TLR9 on mycobacterial DNA-mediated activation of murine macrophages. Alveolar macrophages from TLR9CpG1/CpG1 and wild-type C57BL/6 mice were stimulated with BCG DNA and activation of NF-κB was assessed. Due to the relatively limited number of murine alveolar macrophages that were obtainable, we assessed NF-κB activation by visualizing nuclear translocation of the NF-κB subunit p65 by confocal microscopy. Stimulation of alveolar macrophages with BCG DNA induced NF-κB translocation in wild-type C57BL/6 but not in TLR9CpG1/CpG1 mice (fig. 3c). In contrast, stimulation of the alveolar macrophages with the TLR4 agonist LPS induced NF-κB translocation in both wild-type C57BL/6 and TLR9CpG1/CpG1 mice (fig. 3c). These experiments further confirmed the purity of our BCG DNA preparation, and at the same time demonstrated that murine alveolar macrophages both express TLR9 and require TLR9 for an optimal innate response to BCG DNA.
We also examined TLR9 mRNA and protein expression in human alveolar macrophages. TLR9 mRNA as well as TRL2 mRNA were detected by conventional RT-PCR (fig. 4a). Similar to the pattern seen with the in vitro differentiated macrophages, quantitative RT-PCR analysis of human alveolar macrophages showed that TLR9 mRNA was expressed at a much lower level than TLR2 or TLR4 mRNA (fig. 4b). The presence of TLR9 protein in human alveolar macrophages was then confirmed by confocal microscopy, Western blotting and quantitative laser scanning microscopy (fig. 4c).
Fig. 4.
Human alveolar macrophages express TLR9 mRNA and protein. TLR mRNA expression was analysed in human alveolar macrophages by qualitative (a) and quantitative RT-PCR (b) and TLR9 protein was detected by immunocytochemistry and Western blot (c). a, b RNA isolation, reverse transcription and qualitative RT-PCR for TLR2 and TLR9 as well as quantitative PCR for TLR2, TLR4 and TLR9 were performed as described in Materials and Methods. Qualitative RT-PCR was performed in the presence (+) or absence (–) of cDNA. Reaction products of qualitative RT-PCR reactions and a size marker (M) were electrophoresed and stained with ethidium bromide. Expression levels of TLR2, TLR4 and TLR9 are expressed as mRNA copies of the respective gene per 1,000 copies of GAPDH. Quantification was performed by comparing starting amounts of cDNAs with a standard curve of amplicon cloned into pGEM-T Easy. c Alveolar macrophages were stained for TLR9 (blue) and with propidium iodide (red) as described in Materials and Methods employing a custom-made antibody and a secondary antibody detected via biotin/streptavidin-Cy5. Stained cells were analysed by confocal microscopy and laser scanning cytometry. A laser scanning cytometer equipped with an argon-ion air-cooled laser and an HeNe laser equipped with a digital camera were used to measure Cy5 fluorescence. Nuclear propidium iodide fluorescence was used as the contouring parameter. Alveolar macrophages for the different analyses were each obtained from different donors. For Western blots, PMA-differentiated THP-1 macrophages or alveolar macrophages were lysed. TLR9 was detected by Western blot by a custom antibody and visualized by an IRDye 800 conjugated anti-mouse IgG. Actin served as a loading control.
TLR9 in Macrophage Infection with Virulent versus Attenuated Mycobacterial Strains
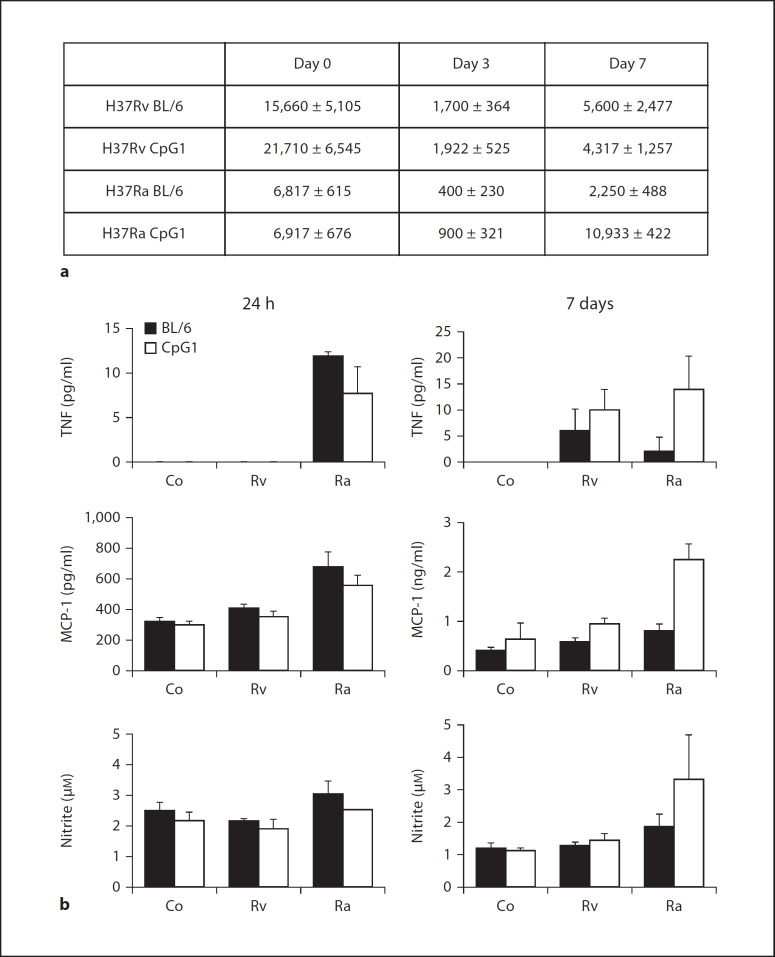
To assess the impact of macrophage TLR9 activation during infection with live mycobacteria, we measured bacterial replication in bone marrow-derived macrophages from wild-type or TLR9CpG1/CpG1 mice infected with either virulent (H37Rv) or attenuated (H37Ra) mycobacteria. As previously described [36], there was an initial decline in bacterial CFU during the first 3 days after infection, followed by rising CFU counts after 7 days (fig. 5a). For the virulent H37Rv bacteria, no difference in replication was seen irrespective of whether macrophages from wild-type and TLR9CpG1/CpG1 mice were used. In contrast, however, attenuated H37Ra bacteria replicated significantly faster in TLR9CpG1/CpG1 macrophages than in wild-type macrophages (fig. 5a). The difference in CFU counts was just over 2-fold after 3 days and more than 4-fold after 7 days (p < 0.0001). These results suggest that TLR9-mediated signalling may contribute to the ability of macrophages to control the intracellular survival of attenuated mycobacteria.
Fig. 5.
Mycobacterial infection of murine macrophages. Mouse bone marrow macrophages were derived from wild-type (BL/6) or TLR9CpG1/CpG1 mice (CpG1) and were either mock infected (Co) or infected with virulent H37Rv or attenuated H37Ra. a Cells were lysed 3 and 7 days after infection and plated in serial dilutions on 7H11 agar to enumerate CFU. Data show values from 2 independent experiments with 3–6 infections (means ± SEM, cell preparations from 6–9 mice per strain). b Cell supernatants from mock-infected cells (Co) or from cells infected with virulent H37Rv or attenuated H37Ra were taken off at days 1 and 7 after infection and TNF-α, MCP-1 and nitrite levels were determined as described in Materials and Methods. Data are expressed as means ± SD of cell preparations from 6–9 mice per strain, performed in triplicates.
We also measured the cytokine, chemokine and nitrite production responses of infected macrophages. Infection with H37Rv, despite higher infection rates at day 0, induced only a marginal increase in TNF-α at 7 days compared to non-infected macrophages (fig. 5b). In contrast, macrophages infected with H37Ra induced robust increases in TNF-α and MCP-1 as well as nitrite 1 day after infection (fig. 5b). TLR9CpG1/CpG1 macrophages were activated slightly less than wild-type macrophages 1 day after infection, most likely due to impaired TLR9 activation. However, after 7 days of H37Ra infection, cytokine, chemokine and nitrite levels were markedly higher in TLR9CpG1/CpG1 macrophages compared to wild-type macrophages, whereby only values for MCP-1 reached statistical significance (p < 0.01). The differences in activation of the macrophages after 7 days likely reflects the higher mycobacterial load found in the TLR9CpG1/CpG1 macrophages (fig. 5a).
In vivo Infection Experiments
Host defence in vivo is only partially mediated by macrophages and we therefore aimed to decipher the impact of our findings for tuberculosis infection in vivo. Thus, we infected wild-type and TLR9CpG1/CpG1 mice with either H37Ra or H37Rv. As expected, infection with H37Ra neither resulted in high bacterial load nor did it kill the animals (data not shown). Interestingly, however, cytokine production in TLR9-deficient animals was lower than in wild-type animals when infected with the attenuated strain (fig. 6).
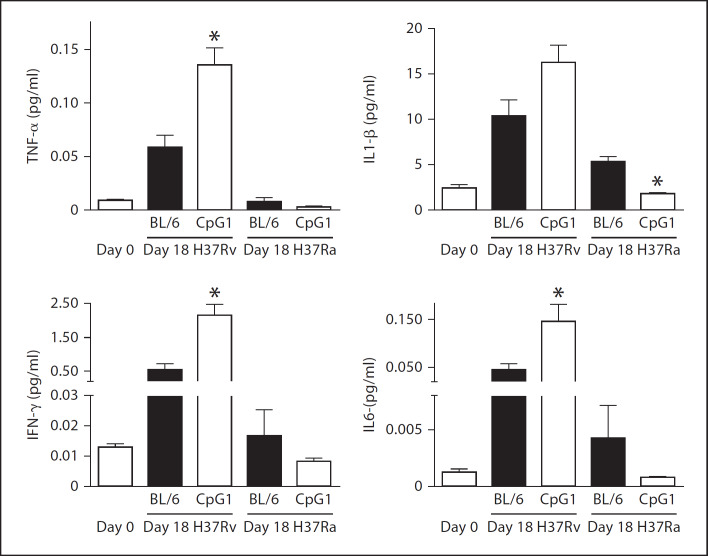
Fig. 6.
Altered cytokine production in TLR9-deficient animals infected with virulent or attenuated mycobacteria. Wild-type (BL/6) or TLR9CpG1/CpG1 mice (CpG1) were infected with M. tuberculosis H37Rv or H37Ra. Lungs were homogenized on day 0 or day 18 and TNF-α, IL-1β, IFN-γ and IL-6 levels were measured by a Luminex bead assay (Upstate Beadlyte). Data show means ± SEM of lungs from 4 animals. * p < 0.05 statistically different from values in wild-type animals.
Similar to what was demonstrated before by others, TLR9-deficient mice did not show differences in bacterial load of the H37Rv strain compared to wild-type mice (data not shown) [5]. Interestingly, in contrast to H37Ra, production of cytokines was significantly increased in mutant mice. Median survival of H37Rv-infected TLR9CpG1/CpG1 animals was slightly but not significantly shorter than that of wild-type animals (266 vs. 279 days).
Different Potency of Mycobacterial DNA to Activate Human Macrophages
We reasoned that there could be several mechanisms explaining the differences in macrophage activation in response to acute infection with H37Rv versus H37Ra mycobacteria as well as the diminished activation of macrophages in TLR9CpG1/CpG1 macrophages versus C57BL/6 macrophages following infection with H37Ra mycobacteria. One possibility is that virulent mycobacteria fail to activate TLR9 because less efficient intracellular bacterial killing of the virulent organisms results in less free genomic DNA able to interact with intracellular TLR9. Alternatively, DNA from virulent bacteria could have less potency to activate human macrophages than DNA from attenuated bacteria. To discriminate between these possible mechanisms, we isolated DNA from virulent and attenuated mycobacteria, and then stimulated human monocyte-derived macrophages with equal concentrations of the different DNA preparations. Remarkably, DNA from attenuated mycobacteria (H37Ra and M. bovis BCG) consistently stimulated a more vigorous macrophage TNF-α response than did DNA from virulent mycobacteria (H37Rv and M. bovis; fig. 7a). This observation was also confirmed in human alveolar macrophages where H37Ra DNA induced a much stronger induction of TNF-α than H37Rv (fig. 7b). In order to check whether the action of H37Ra DNA preparations results in fact from its DNA content and not from any contaminants, H37Ra DNA was digested with DNase. The effect of this preparation was compared to the stimulatory potency of non-digested DNA. We saw that digestion in fact abrogated the ability to stimulate TNF-α mRNA when added to human macrophages (fig. 7c). Taken together, our data suggest that virulent mycobacteria have evolved a mechanism of attenuated TLR9 activation by their genomic DNA.
Fig. 7.
Activation of human monocyte-derived and alveolar macrophages by DNA from virulent and attenuated mycobacteria. Monocyte-derived (a) or alveolar (b) macrophages were treated by DNA from virulent M. tuberculosis (H37Rv) and M. bovis or from attenuated H37Ra and M. bovis BCG strains for 3 h (a) or 2 h (b). RNA was isolated, followed by real-time PCR analysis for TNF-α. Data are expressed as TNF-α induction after normalization to housekeeping gene expression. Data show means ± SEM of 3–13 values employing cells from 3–5 different donors. * p < 0.05 significant difference between treatment with the same DNA concentration of a virulent and the respective attenuated strain (t test). c DNA was isolated from H37Ra mycobacteria and digested with DNase. Primary monocyte-derived macrophages were either left untreated (Co) or treated with digested or mock-treated DNA followed by RNA isolation and real-time PCR analysis for TNF-α. Data are expressed as x-fold TNF-α induction after normalization to GAPDH values. Data show means ± SEM of 3 independent experiments performed in triplicate. * p < 0.05 significantly different to Co; + p < 0.05 significantly different to DNase digested (t test).
Discussion
Control of infection by M. tuberculosis requires an effective combined innate plus adaptive immune response by the host. The innate response mounted by infected macrophages and the adaptive CD4+ T cell response have been highlighted as key contributors to an effective response [1]. Studies using MyD88 null mice have demonstrated that TLR signalling plays a critically important role in the success of the innate host response [35]. The specific role of individual TLRs, however, remains elusive. Our results suggest that TLR9, the sensor for bacterial CpG DNA, plays an important role in shaping the innate macrophage response to mycobacteria infection and thus helps determine the virulence of infection.
Significant controversy exists regarding whether human macrophages express functional TLR9. While mouse macrophage TLR9 activation and signalling is well established, a number of studies failed to find evidence for functional TLR9 in human monocytes or macrophages [10, 11, 12]. Miettinen et al. [9] failed to detect any TLR9 mRNA by Northern blot in human macrophages; however, their results also showed very weak mRNA levels of TLR2 (a TLR known to be highly expressed and functional in human macrophages), suggesting that the sensitivity of Northern blot may not have been sufficient to detect the TLR9 mRNA levels present in human macrophages. Even employing qualitative RT-PCR, Takeshita et al. [12] failed to detect TLR9 mRNA in THP-1 cells. In contrast, other studies have provided convincing data that human cells of the monocytic lineage do express TLR9. Real-time PCR analysis revealed detectable levels of TLR9 in THP-1 cells as well as in peripheral monocytes in 36 donors [36], supporting our findings of low but readily detectable levels of TLR9 in THP-1 macrophages, monocyte-derived macrophages and alveolar macrophages. With TLR9 mRNA levels being lower than those of TLR2 or TLR4 in both in vitro differentiated as well as alveolar macrophages, we confirm the quantitative RT-PCR analyses previously reported using primary human monocytes [10] and murine [37] as well as human [38] alveolar macrophages.
We also clearly detected TLR protein in THP-1 macrophages, human monocyte-derived macrophages and human alveolar macrophages. These findings corroborate other reports of TLR9 protein expressed in monocytic CD14/TLR9 double-positive cells [39] as well as alveolar macrophages from human tuberculous lung granulomas [17].
Several reports suggested that human monocytic cells contained functional TLR9 receptors. Sawamura et al. [16] showed that monocytes were primarily responsible for the inflammatory reaction induced by injection of plasmid DNA into human skin. Mao et al. [19] reported that human monocytes showed marked responsiveness to CpG stimulation and two groups reported THP-1 monocytes as target cells for ISS [32, 40]. Furthermore, addition of CpG sequences was recently demonstrated to attenuate growth of virulent M. tuberculosis (Erdman) in human monocyte-derived macrophages [41]. We now demonstrate that human monocyte-derived macrophages express TLR9 and are activated by treatment with bacterial DNA. Interestingly, we found that human macrophages respond far more vigorously to bacterial DNA than they do to CpG ODN at the same concentration. Many of the published effects ascribed to specific CpG sequences were recently proven to be due to CpG-independent effects of phosphodiesters [42]. As shown by the employment of GpC ODN controls, such non-specific effects seem to be of little importance in human macrophages, leading to a weak activation by the use of phosphorothioate-modified ISS. In addition, multimerization of DNA has been reported crucial for optimal TLR9 activation, suggesting that genomic DNA has a higher potential of aggregation [43]. This was confirmed by the observation that bacterial genomic DNA is much more efficient than oligonucleotides in activating mouse macrophages [20]. We show here for the first time that such a difference can also be observed in human macrophages.
We also report similar TLR9-mediated signalling after stimulation with bacterial DNA in human and mouse alveolar macrophages. This contrasts to the report by Suzuki et al. [37] who described impaired TLR9 signalling in mouse alveolar macrophages despite detectable TLR9 expression. Like other publications that failed to find evidence of TLR9 signalling in human monocytes or macrophages [10, 11, 13], Suzuki et al. [37] only tested the effect of CpG ODN, not that of larger, bacterial DNA. Since mouse macrophages have been shown to require larger DNA molecules in order to be sufficiently activated [20], the use of the suboptimal stimulus might explain the lack of responsiveness. The use of bacterial DNA in fact seems to more adequately reflect the in vivo situation. This possibility is largely supported by our demonstration of different responses of TLR9CpG1/CpG1-derived macrophages towards mycobacterial infection compared to cells from wild-type animals. In addition, theoretical calculations indicate that the small volume of endosomes (the organelle known TLR9 signalling takes place in) promotes signalling even from a limited number of TLR agonists [44].
In order to determine whether macrophage TLR9 activation in fact plays a role in mycobacterial infection, macrophages from wild-type (C57BL/6) or TLR9-deficient (TLR9CpG1/CpG1) mice [34] were infected with mycobacteria. There was no difference in intracellular replication of the virulent H37Rv strain between wild-type and TLR9-deficient macrophages, a result in keeping with a recent in vivo study demonstrating that TLR9-deficient animals did not show differences in survival compared to wild-type animals when infected with a low dose of mycobacteria [5]. In contrast, when macrophages were infected with the attenuated H37Ra strain, bacteria replicated much faster in TLR9CpG1/CpG1 macrophages than in wild-type cells. The TLR9CpG1/CpG1 macrophages also showed a decreased activation pattern (assessed by TNF-α, MCP-1 and nitrite concentrations in the supernatant) compared to wild-type macrophages 1 day after infection. We suggest that these findings are explained by an impaired TLR9 activation. Seven days after infection, however, TLR9CpG1/CpG1 macrophage cytokine, chemokine and nitrite production was markedly higher than that found from wild-type macrophages. This late increase in macrophage activation is likely due to the increased bacterial load leading to strong activation of macrophages via other receptors of the innate immune response, such as TLR2 and TLR4.
Previous studies have suggested that the innate immune response against tuberculosis involves multiple TLRs and that abrogation of individual TLRs (such as TLR2, TLR, TLR6 or TLR9) does not substantially abrogate the response to M. tuberculosis infection [5, 45, 46, 47]. It is therefore not surprising that our in vivo data employing animals deficient in only TLR9 do not show significant differences in bacterial replication compared to wild-type animals and only slightly reduced survival. Still, it seems of special importance that knockdown of TLR9 leads to completely different cytokine responses upon infection with H37Ra or H37Rv. Whereas TLR9CpG1/CpG1 mice show reduced cytokine production after H37Ra infection, it is increased after H37Rv infection. While it is not possible to determine the cellular source of cytokine production in vivo, this observation definitely shows differences in TLR9 activation between virulent and attenuated mycobacterial strains.
Taken together, the infection data suggest that virulent H37Rv mycobacteria elicit an altered TLR9 response compared to attenuated H37Ra mycobacteria. To address the mechanism underlying this difference, we stimulated macrophages with DNA preparations made from virulent (H37Rv and M. bovis) or attenuated (H37Ra and M. bovis BCG) mycobacteria and demonstrated that DNA from attenuated mycobacteria is more efficient in activating human monocyte-derived and alveolar macrophages than DNA from virulent mycobacteria. These results might explain data from the literature reporting that the ability of mycobacteria to induce the secretion of TNF-α is inversely related to their virulence in murine macrophages [48].
A recent genome-wide screening analysed ISS in genomic mycobacterial DNA [49]. The data showed that not only the CpG motifs but also the context of the sequence surrounding the CpG motif are important for the immunostimulatory activities of genomic DNA of M. bovis. The generally higher CG content in mycobacteria compared to other bacteria such as Escherichia coli (65 vs. 51%) together with an even higher frequency of CpG sequences in mycobacteria compared to E. coli suggested a specifically high potential of mycobacteria to activate TLR9. Since the different mycobacterial strains have a >99% DNA homology and possess a comparable CG content of about 65% [50, 51], one might wonder how differences in the extent of TLR9 activation can occur between genomic DNA from different strains. There are at least 3 possible explanations for the observed differences in the efficiency of TLR9 activation by different mycobacterial DNA and at this point we can only speculate about the mechanisms involved: (1) attenuated strains might have a reduced frequency of activatory CpG sequences based on subtle differences in the surrounding DNA sequences; (2) virulent strains might have an increased number of immunosuppressive DNA sequences in their genomes; (3) virulent strains may have developed strategies to mask their CpGs, such as by expressing DNA methylases. Since the genomes of attenuated strains have not been sequenced yet, whole-genome sequence comparisons cannot be performed in order to check hypotheses (1) and (2). Most interestingly, however, data from the literature suggest that the virulent H37Rv strain is able to methylate cytosines, whereas the H37Ra strain is not [52, 53]. This observation might explain why DNA from virulent strains has a lower potential to activate TLR9 in human macrophages than DNA from attenuated strains. Further studies need to be undertaken to prove whether structural alterations of mycobacterial DNA of different strains might in fact be responsible for the decreased ability of DNA from virulent mycobacteria to stimulate human macrophages.
Taken together, these studies establish that human macrophages express functional TLR9 and suggest that signalling through TLR9 contributes to the innate macrophage response to infection by mycobacteria. Furthermore, the differences between the stimulatory activity of DNA from virulent versus attenuated mycobacteria suggest that TLR9-induced macrophage activation may influence the host's ability to limit and then kill intracellular mycobacteria. The altered TLR9 activation by virulent mycobacteria might represent a mechanism how they evade the host immune response.
Acknowledgements
We thank Dr. Riaz Zuberi for help in mouse alveolar macrophage isolation, Dr. Jane Eddleston for support in real-time PCR setup, Christoph Meyer for help in macrophage isolation, Dr. Hanno Huwer for supply with human lung tissue as well as Michael Bur and Dr. Claus-Michael Lehr for helpful advice and support in the isolation of human alveolar macrophages. This work was supported, in part, by the Public Health Service grant PO1 CI000095. A.K.K. was supported by the Alexander von Humboldt foundation, J.H. by the Landesgraduiertenkolleg des Saarlandes.
References
- 1.Doffinger R, Patel SY, Kumararatne DS. Host genetic factors and mycobacterial infections: lessons from single gene disorders affecting innate and adaptive immunity. Microbes Infect. 2006;8:1141–1150. doi: 10.1016/j.micinf.2005.10.028. [DOI] [PubMed] [Google Scholar]
- 2.Beutler B. Inferences, questions and possibilities in Toll-like receptor signalling. Nature. 2004;430:257–263. doi: 10.1038/nature02761. [DOI] [PubMed] [Google Scholar]
- 3.Shimada S, Yano O, Inoue H, Kuramoto E, Fukuda T, Yamamoto H, Kataoka T, Tokunaga T. Antitumor activity of the DNA fraction from Mycobacterium bovis BCG. II. Effects on various syngeneic mouse tumors. J Natl Cancer Inst. 1985;74:681–688. [PubMed] [Google Scholar]
- 4.Tokunaga T, Yamamoto H, Shimada S, Abe H, Fukuda T, Fujisawa Y, Furutani Y, Yano O, Kataoka T, Sudo T. Antitumor activity of deoxyribonucleic acid fraction from Mycobacterium bovis BCG. I. Isolation, physicochemical characterization, and antitumor activity. J Natl Cancer Inst. 1984;72:955–962. [PubMed] [Google Scholar]
- 5.Bafica A, Scanga CA, Feng CG, Sher A, Leifer C, Cheever A. TLR9 regulates Th1 responses and cooperates with TLR2 in mediating optimal resistance to Mycobacterium tuberculosis. J Exp Med. 2005;202:1715–1724. doi: 10.1084/jem.20051782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hemmi H, Takeuchi O, Kawai T, Kaisho T, Sato S, Sanjo H, Matsumoto M, Hoshino K, Wagner H, Takeda K, Akira S. A Toll-like receptor recognizes bacterial DNA. Nature. 2000;408:740–745. doi: 10.1038/35047123. [DOI] [PubMed] [Google Scholar]
- 7.Leifer CA, Kennedy MN, Mazzoni A, Lee C, Kruhlak MJ, Segal DM. TLR9 is localized in the endoplasmic reticulum prior to stimulation. J Immunol. 2004;173:1179–1183. doi: 10.4049/jimmunol.173.2.1179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Latz E, Schoenemeyer A, Visintin A, Fitzgerald KA, Monks BG, Knetter CF, Lien E, Nilsen NJ, Espevik T, Golenbock DT. TLR9 signals after translocating from the ER to CpG DNA in the lysosome. Nat Immunol. 2004;5:190–198. doi: 10.1038/ni1028. [DOI] [PubMed] [Google Scholar]
- 9.Miettinen M, Sareneva T, Julkunen I, Matikainen S. IFNs activate Toll-like receptor gene expression in viral infections. Genes Immun. 2001;2:349–355. doi: 10.1038/sj.gene.6363791. [DOI] [PubMed] [Google Scholar]
- 10.Hornung V, Rothenfusser S, Britsch S, Krug A, Jahrsdorfer B, Giese T, Endres S, Hartmann G. Quantitative expression of toll-like receptor 1-10 mRNA in cellular subsets of human peripheral blood mononuclear cells and sensitivity to CpG oligodeoxynucleotides. J Immunol. 2002;168:4531–4537. doi: 10.4049/jimmunol.168.9.4531. [DOI] [PubMed] [Google Scholar]
- 11.Doyle SE, O'Connell RM, Vaidya SA, Liu PT, Cheng G, Miranda GA, Lane TF, Chow EK, Modlin RL, Suzuki S, Suzuki N, Yeh WC. Toll-like receptors induce a phagocytic gene program through p38. J Exp Med. 2004;199:81–90. doi: 10.1084/jem.20031237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Takeshita F, Suzuki K, Sasaki S, Ishii N, Klinman DM, Ishii KJ. Transcriptional regulation of the human TLR9 gene. J Immunol. 2004;173:2552–2561. doi: 10.4049/jimmunol.173.4.2552. [DOI] [PubMed] [Google Scholar]
- 13.Remer KA, Brcic M, Sauter KS, Jungi TW. Human monocytoid cells as a model to study Toll-like receptor-mediated activation. J Immunol Methods. 2006;313:1–10. doi: 10.1016/j.jim.2005.07.026. [DOI] [PubMed] [Google Scholar]
- 14.Hartmann G, Krieg AM. CpG DNA and LPS induce distinct patterns of activation in human monocytes. Gene Ther. 1999;6:893–903. doi: 10.1038/sj.gt.3300880. [DOI] [PubMed] [Google Scholar]
- 15.Hornung V, Guenthner-Biller M, Bourquin C, Ablasser A, Schlee M, Uematsu S, Noronha A, Manoharan M, Akira S, de Fougerolles A, Endres S, Hartmann G. Sequence-specific potent induction of IFN-α by short interfering RNA in plasmacytoid dendritic cells through TLR7. Nat Med. 2005;11:263–270. doi: 10.1038/nm1191. [DOI] [PubMed] [Google Scholar]
- 16.Sawamura D, Abe R, Goto M, Akiyama M, Hemmi H, Akira S, Shimizu H. Direct injection of plasmid DNA into the skin induces dermatitis by activation of monocytes through toll-like receptor 9. J Gene Med. 2005;7:664–671. doi: 10.1002/jgm.709. [DOI] [PubMed] [Google Scholar]
- 17.Fenhalls G, Squires GR, Stevens-Muller L, Bezuidenhout J, Amphlett G, Duncan K, Lukey PT. Associations between Toll-like receptors and interleukin-4 in the lungs of patients with tuberculosis. Am J Respir Cell Mol Biol. 2003;29:28–38. doi: 10.1165/rcmb.2002-0163OC. [DOI] [PubMed] [Google Scholar]
- 18.Hoene V, Peiser M, Wanner R. Human monocyte-derived dendritic cells express TLR9 and react directly to the CpG-A oligonucleotide D19. J Leukoc Biol. 2006;80:1328–1336. doi: 10.1189/jlb.0106011. [DOI] [PubMed] [Google Scholar]
- 19.Mao TK, Lian ZX, Selmi C, Ichiki Y, Ashwood P, Ansari AA, Coppel RL, Shimoda S, Ishibashi H, Gershwin ME. Altered monocyte responses to defined TLR ligands in patients with primary biliary cirrhosis. Hepatology. 2005;42:802–808. doi: 10.1002/hep.20859. [DOI] [PubMed] [Google Scholar]
- 20.Roberts TL, Dunn JA, Terry TD, Jennings MP, Hume DA, Sweet MJ, Stacey KJ. Differences in macrophage activation by bacterial DNA and CpG-containing oligonucleotides. J Immunol. 2005;175:3569–3576. doi: 10.4049/jimmunol.175.6.3569. [DOI] [PubMed] [Google Scholar]
- 21.Kiemer AK, Hartung T, Vollmar AM. cGMP-mediated inhibition of TNF-α production by the atrial natriuretic peptide in murine macrophages. J Immunol. 2000;165:175–181. doi: 10.4049/jimmunol.165.1.175. [DOI] [PubMed] [Google Scholar]
- 22.Plesner A. Increasing the yield of human mononuclear cells and low serum conditions for in vitro generation of macrophages with M-CSF. J Immunol Methods. 2003;279:287–295. doi: 10.1016/s0022-1759(03)00234-5. [DOI] [PubMed] [Google Scholar]
- 23.Kiemer AK, Vollmar AM. Autocrine regulation of inducible nitric-oxide synthase in macrophages by atrial natriuretic peptide. J Biol Chem. 1998;273:13444–13451. doi: 10.1074/jbc.273.22.13444. [DOI] [PubMed] [Google Scholar]
- 24.Santos AR, de Miranda AB, Lima LM, Suffys N, Degrave WM. Method for high yield preparation in large and small scale of nucleic acids from mycobacteria. J Microbiol Methods. 1992;15:83–94. [Google Scholar]
- 25.Cotten M, Baker A, Saltik M, Wagner E, Buschle M. Lipopolysaccharide is a frequent contaminant of plasmid DNA preparations and can be toxic to primary human cells in the presence of adenovirus. Gene Ther. 1994;1:239–246. [PubMed] [Google Scholar]
- 26.Buryanov Y, Shevchuk T. The use of prokaryotic DNA methyltransferases as experimental and analytical tools in modern biology. Anal Biochem. 2005;338:1–11. doi: 10.1016/j.ab.2004.02.048. [DOI] [PubMed] [Google Scholar]
- 27.Kiemer AK, Weber NC, Vollmar AM. Induction of IκB: atrial natriuretic peptide as a regulator of the NF-κB pathway. Biochem Biophys Res Commun. 2002;295:1068–1076. doi: 10.1016/s0006-291x(02)00807-0. [DOI] [PubMed] [Google Scholar]
- 28.Schwende H, Fitzke E, Ambs P, Dieter P. Differences in the state of differentiation of THP-1 cells induced by phorbol ester and 1,25-dihydroxyvitamin D3. J Leukoc Biol. 1996;59:555–561. [PubMed] [Google Scholar]
- 29.Baltathakis I, Alcantara O, Boldt DH. Expression of different NF-κB pathway genes in dendritic cells (DCs) or macrophages assessed by gene expression profiling. J Cell Biochem. 2001;83:281–290. doi: 10.1002/jcb.1231. [DOI] [PubMed] [Google Scholar]
- 30.Krug A, Towarowski A, Britsch S, Rothenfusser S, Hornung V, Bals R, Giese T, Engelmann H, Endres S, Krieg AM, Hartmann G. Toll-like receptor expression reveals CpG DNA as a unique microbial stimulus for plasmacytoid dendritic cells which synergizes with CD40 ligand to induce high amounts of IL-12. Eur J Immunol. 2001;31:3026–3037. doi: 10.1002/1521-4141(2001010)31:10<3026::aid-immu3026>3.0.co;2-h. [DOI] [PubMed] [Google Scholar]
- 31.Rutz M, Metzger J, Gellert T, Luppa P, Lipford GB, Wagner H, Bauer S. Toll-like receptor 9 binds single-stranded CpG-DNA in a sequence- and pH-dependent manner. Eur J Immunol. 2004;34:2541–2550. doi: 10.1002/eji.200425218. [DOI] [PubMed] [Google Scholar]
- 32.Sanjuan MA, Rao N, Lai KTA, Gu Y, Sun S, Fuchs A, Fung-Leung WP, Colonna M, Karlsson L. CpG-induced tyrosine phosphorylation occurs via a TLR9-independent mechanism and is required for cytokine secretion. J Cell Biol. 2006;172:1057–1068. doi: 10.1083/jcb.200508058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lee J, Chuang TH, Redecke V, She L, Pitha PM, Carson DA, Raz E, Cottam HB. Molecular basis for the immunostimulatory activity of guanine nucleoside analogs: activation of Toll-like receptor 7. Proc Natl Acad Sci USA. 2003;100:6646–6651. doi: 10.1073/pnas.0631696100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Tabeta K, Georgel P, Janssen E, Du X, Hoebe K, Crozat K, Mudd S, Shamel L, Sovath S, Goode J, Alexopoulou L, Flavell RA, Beutler B. Toll-like receptors 9 and 3 as essential components of innate immune defense against mouse cytomegalovirus infection. Proc Natl Acad Sci USA. 2004;101:3516–3521. doi: 10.1073/pnas.0400525101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Fremond CM, Yeremeev V, Nicolle DM, Jacobs M, Quesniaux VF, Ryffel B. Fatal Mycobacterium tuberculosis infection despite adaptive immune response in the absence of MyD88. J Clin Invest. 2004;114:1790–1799. doi: 10.1172/JCI21027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zarember KA, Godowski PJ. Tissue expression of human Toll-like receptors and differential regulation of Toll-like receptor mRNAs in leukocytes in response to microbes, their products, and cytokines. J Immunol. 2002;168:554–561. doi: 10.4049/jimmunol.168.2.554. [DOI] [PubMed] [Google Scholar]
- 37.Suzuki K, Suda T, Naito T, Ide K, Chida K, Nakamura H. Impaired Toll-like receptor 9 expression in alveolar macrophages with no sensitivity to CpG DNA. Am J Respir Crit Care Med. 2005;171:707–713. doi: 10.1164/rccm.200408-1078OC. [DOI] [PubMed] [Google Scholar]
- 38.Maris NA, Dessing MC, de Vos AF, Bresser P, van der Zee JS, Jansen HM, Spek CA, van der., Poll T Toll-like receptor mRNA levels in alveolar macrophages after inhalation of endotoxin. Eur Respir J. 2006;28:622–626. doi: 10.1183/09031936.06.00010806. [DOI] [PubMed] [Google Scholar]
- 39.Eaton-Bassiri A, Dillon SB, Cunningham M, Rycyzyn MA, Mills J, Sarisky RT, Mbow ML. Toll-like receptor 9 can be expressed at the cell surface of distinct populations of tonsils and human peripheral blood mononuclear cells. Infect Immun. 2004;72:7202–7211. doi: 10.1128/IAI.72.12.7202-7211.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kuo CC, Kuo CW, Liang CM, Liang SM. A transcriptomic and proteomic analysis of the effect of CpG-ODN on human THP-1 monocytic leukemia cells. Proteomics. 2005;5:894–906. doi: 10.1002/pmic.200401144. [DOI] [PubMed] [Google Scholar]
- 41.Wang JP, Hayashi T, Datta SK, Kornbluth RS, Raz E, Guiney DG. CpG oligonucleotides partially inhibit growth of Mycobacterium tuberculosis, but not Salmonella or Listeria, in human monocyte-derived macrophages. FEMS Immunol Med Microbiol. 2005;45:303–310. doi: 10.1016/j.femsim.2005.05.007. [DOI] [PubMed] [Google Scholar]
- 42.Roberts TL, Sweet MJ, Hume DA, Stacey KJ. Cutting edge: species-specific TLR9-mediated recognition of CpG and non-CpG phosphorothioate-modified oligonucleotides. J Immunol. 2005;174:605–608. doi: 10.4049/jimmunol.174.2.605. [DOI] [PubMed] [Google Scholar]
- 43.Wu CC, Lee J, Raz E, Corr M, Carson DA. Necessity of oligonucleotide aggregation for Toll-like receptor 9 activation. J Biol Chem. 2004;279:33071–33078. doi: 10.1074/jbc.M311662200. [DOI] [PubMed] [Google Scholar]
- 44.Crozat K, Beutler B. TLR7: a new sensor of viral infection. Proc Natl Acad Sci USA. 2004;101:6835–6836. doi: 10.1073/pnas.0401347101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Reiling N, Holscher C, Fehrenbach A, Kroger S, Kirschning CJ, Goyert S, Ehlers S. Cutting edge: Toll-like receptor (TLR)2- and TLR4-mediated pathogen recognition in resistance to airborne infection with Mycobacterium tuberculosis. J Immunol. 2002;169:3480–3484. doi: 10.4049/jimmunol.169.7.3480. [DOI] [PubMed] [Google Scholar]
- 46.Kamath AB, Alt J, Debbabi H, Behar SM. Toll-like receptor 4-defective C3h/HeJ mice are not more susceptible than other C3H substrains to infection with Mycobacterium tuberculosis. Infect Immun. 2003;71:4112–4118. doi: 10.1128/IAI.71.7.4112-4118.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Sugawara I, Yamada H, Li C, Mizuno S, Takeuchi O, Akira S. Mycobacterial infection in tlr2 and tlr6 knockout mice. Microbiol Immunol. 2003;47:327–336. doi: 10.1111/j.1348-0421.2003.tb03404.x. [DOI] [PubMed] [Google Scholar]
- 48.Falcone V, Bassey EB, Toniolo A, Conaldi PG, Collins FM. Differential release of tumor necrosis factor-α from murine peritoneal macrophages stimulated with virulent and avirulent species of mycobacteria. FEMS Immunol Med Microbiol. 1994;8:225–232. doi: 10.1111/j.1574-695X.1994.tb00447.x. [DOI] [PubMed] [Google Scholar]
- 49.Lee KW, Jung J, Lee Y, Kim TY, Choi SY, Park J, Kim DS, Kwon HJ. Immunostimulatory oligodeoxynucleotide isolated from genome wide screening of Mycobacterium bovis chromosomal DNA. Mol Immunol. 2006;43:2107–2118. doi: 10.1016/j.molimm.2005.12.004. [DOI] [PubMed] [Google Scholar]
- 50.Baess I, Mansa B. Determination of genome size and base ratio on deoxyribonucleic acid from mycobacteria. Acta Pathol Microbiol Scand. 1978;86B:309–312. doi: 10.1111/j.1699-0463.1978.tb00049.x. [DOI] [PubMed] [Google Scholar]
- 51.Garnier T, Eiglmeier K, Camus JC, Medina N, Mansoor H, Pryor M, Duthoy S, Grondin S, Lacroix C, Monsempe C, Simon S, Harris B, Atkin R, Doggett J, Mayes R, Keating L, Wheeler PR, Parkhill J, Barrell BG, Cole ST, Gordon SV, Hewinson RG. The complete genome sequence of Mycobacterium bovis. Proc Natl Acad Sci USA. 2003;100:7877–7882. doi: 10.1073/pnas.1130426100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Srivastava R, Gopinathan KP, Ramakrishnan T. Deoxyribonucleic acid methylation in mycobacteria. J Bacteriol. 1981;148:716–719. doi: 10.1128/jb.148.2.716-719.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Hemavathy KC, Nagaraja V. DNA methylation in mycobacteria: absence of methylation at GATC (Dam) and CCA/TGG (Dcm) sequences. FEMS Immunol Med Microbiol. 1995;11:291–296. doi: 10.1111/j.1574-695X.1995.tb00159.x. [DOI] [PubMed] [Google Scholar]