Abstract
Heat Shock Protein 90 (Hsp90) is an essential molecular chaperone in eukaryotic cells, and it maintains the functional conformation of a subset of proteins that are typically key components of multiple regulatory and signaling networks mediating cancer cell proliferation, survival and metastasis. It is possible to selectively inhibit Hsp90 using natural products such as geldanamycin (GA) or radicicol (RD), which have served as prototypes for development of synthetic Hsp90 inhibitors. These compounds bind within the ADP/ATP-binding site of the Hsp90 N-terminal domain to inhibit its ATPase activity. As numerous N-terminal domain inhibitors are currently undergoing extensive clinical evaluation, it is important to understand the factors that may modulate in vivo susceptibility to these drugs. We recently reported that Wee1Swe1-mediated, cell cycle-dependent, tyrosine phosphorylation of Hsp90 affects GA binding and impacts cancer cell sensitivity to Hsp90 inhibition. This phosphoryfiglation also affects Hsp90 ATPase activity and its ability to chaperone a selected group of clients, comprised primarily of protein kinases. Wee1 regulates the G2/M transition. Here we present additional data demonstrating that tyrosine phosphorylation of Hsp90 by Wee1Swe1 is important for Wee1Swe1 association with Hsp90 and for Wee1Swe1 stability. Yeast expressing non-phosphorylatable yHsp90-Y24F, like swe1Δ yeast, undergo premature nuclear division that is insensitive to G2/M checkpoint arrest. These findings demonstrate the importance of Hsp90 phosphorylation for proper cell cycle regulation.
Keywords: heat shock protein 90, phosphorylation, wee1 kinase, molecular chaperones, post-translational modification, cell cycle
Introduction
Heat Shock Protein 90 (Hsp90) is one of the most abundant proteins in cells (1–2% of total cellular protein). The cellular functions of this essential molecular chaperone have been most clearly identified in mammalian cells, Drosophila and baker’s yeast.1–4 Hsp90 and a discrete set of co-chaperones create and maintain the functional conformation of a subset of proteins referred to as “clients” (www.picard.ch/downloads/Hsp90interactors.pdf).5,6 These targets are key mediators of signal transduction and cell cycle control.
Hsp90 function has been the subject of intense scrutiny because (1) drugs that inhibit its function are anti-tumor agents currently being evaluated in numerous clinical trials,7 and (2) Hsp90 acts as a ‘genetic buffering system’ to limit the effects of genetic variation in populations. Disturbance of Hsp90 function diminishes this buffering and allows genetic variation to manifest itself, principally as morphogenic alteration.4,8,9
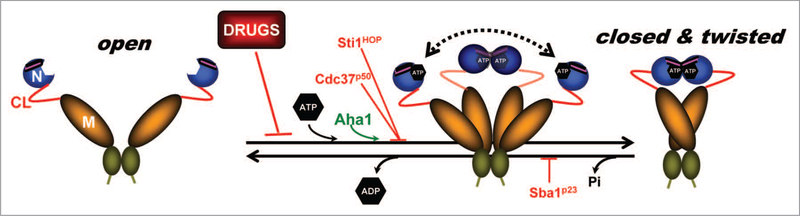
Hsp90 structure is highly conserved across species and it consists of: (1) an N-terminal domain, containing nucleotide and drug binding sites; (2) a middle (M) domain, which provides binding sites for client proteins and various co-chaperones; (3) a C-terminal domain, containing a dimerization region that provides for constitutive association of two Hsp90 protomers.10–12 Eukaryotic Hsp90 also possess an unstructured charged-linker region of significant but variable length connecting N and M domains13,14 (Fig. 1). Hsp90 function relies on ATP binding and hydrolysis, which in turn impact its conformational dynamics. Hsp90 inhibitors currently undergoing clinical trial halt the chaperone cycle by replacing ATP in Hsp90’s nucleotide binding pocket.12,15–18
Figure 1.
Model depicting the Hsp90 chaperone cycle. ATP binding to the N-terminal domains of Hsp90 (open) promotes repositioning of a “lid” segment followed by transient dimerization of the N-domains. Subsequent structural rearrangements result in the (closed and twisted) conformation of Hsp90 that is competent for ATP hydrolysis. Binding of the co-chaperone Aha1 enhances Hsp90 ATPase activity. The co-chaperones Sti1/HOP and Cdc37/p50, or pharmacologic inhibitors such as geldanamycin or radicicol, exert an opposite effect by blocking the initial structural changes necessary for N-domain dimerization. Sba1/p23 strengthens the late Hsp90 conformation and inhibits ATP hydrolysis. Domain labeling is as follows: N, N-domain (blue); CL, charged linker (red); M, M-domain (yellow); C, C-domain (green); ATP lid, (purple).
The regulation of Hsp90 function is complex and multifactorial.19 In eukaryotes, co-chaperones modulate its intrinsic ATPase activity.20,21 Post-translational modification of Hsp90 (e.g., phosphorylation, acetylation and S-nitrosylation) also impacts ATP and co-chaperone binding,22–24 thus providing a further layer of regulation to the Hsp90 cycle not found in bacteria, a requirement no doubt made necessary by the increasingly complex utilization of Hsp90 in maintaining cellular homeostasis in the face of diverse environmental fluctuation.
Here we will briefly review serine, threonine and tyrosine phosphorylation of Hsp90, including our recent observation that Hsp90 is tyrosine phosphorylated by its client protein Wee1Swe.1 We will also present additional data showing that Hsp90 tyrosine phosphorylation is important for Wee1Swe1 stability, and that yeast expressing Wee1Swe1-non-phosphorylatable Hsp90 share a similar cell cycle defect as SWE1 delete yeast.25 These findings support an important role for Hsp90 in regulating the cell cycle.25,26
Serine/Threonine Phosphorylation of Hsp90
Hsp90 is a phosphoprotein.27–39 However our understanding of the role played by phosphorylation of distinct residues in regulating the chaperone function of Hsp90 remains incomplete. A number of serine and threonine phosphorylation sites on Hsp90 have been identified and studied for their impact on chaperone function (Table 1).22 Early work showed that treating cancer cells with the serine/threonine phosphatase inhibitor okadaic acid promoted Hsp90 hyperphosphorylation, which was accompanied by decreased association with its client kinase pp60v-src, suggesting a link between Hsp90 phosphorylation and chaperoning of its client proteins.27,35 Hsp90 is also a substrate for DNA-dependent protein kinase, Akt, B-Raf and casein kinase II (CKII).36,37,40,41 Further, PKA phosphorylation of Thr90 induced by 3-hydroxy-3-methylglutarylcoenzyme A reductase inhibitors has been reported to increase association of human Hsp90α with the client protein eNOS.42 Lastly, a study reported that protein phosphatase 5 (Pp5/Ppt1) can dephosphorylate Hsp90 in vitro.43 This study also showed that Ppt1 deletion in yeast compromised Hsp90 activity.
Table 1.
Yeast (Hsc82, Hsp82) and human (Hsp90α, Hsp90β) phosphorylation sites and identified kinases
| Residues | |||||
|---|---|---|---|---|---|
| yHsp90 | yHsc90 | hHsp90α | hHsp90β | Protein kinase | Reference |
| N/A | N/A | T5* | N/A | Ds-DNA activated protein kinase | 36 |
| N/A | N/A | T7* | N/A | Ds-DNA activated protein kinase | 36 |
| Y24* | Y24 | Y38* | Y33 | Swe1/Wee1 | 25 |
| Y47 | Y47 | Y61 | Y56* | Unknown | 57, 58 |
| S49 | N/A | S63* | S58 | Casein kinase II | 38 |
| S51 | S51 | T65* | T60 | Casein kinase II | 38 |
| N/A | N/A | S68* | S63 | Casein kinase II | 38 |
| T58 | T58 | S72* | S67 | Casein kinase II | 38 |
| N/A | N/A | T88* | T83 | Unknown | 42 |
| N/A | N/A | T90* | T85 | PKA | 59 |
| Y184 | Y184 | Y197 | Y192* | Unknown | 58 |
| N/A | N/A | S231* | S226* | Casein kinase II | 35, 37, 39 |
| N/A | N/A | S252* | N/A | Unknown | 60 |
| N/A | N/A | S263* | S255* | Casein kinase II, B-Raf | 35, 37, 39, 40 |
| S282* | S278 | N/A | N/A | Unknown | SGD |
| T285 | T281 | T305 | T297* | Unknown | 61 |
| S297* | S293 | N/A | N/A | Unknown | SGD |
| Y289 | Y285 | Y309 | Y300* | c-Src (does not exist in yeast) | 33 |
| S295 | S291 | S315 | S307* | Unknown | 62 |
| S334* | S330 | N/A | N/A | Unknown | SGD |
| S379* | S375 | S399 | S391 | Unknown | SGD |
| T443* | T429 | N/A | N/A | Unknown | SGD |
| N/A | N/A | S460 | S452* | PKA | 63 |
| Y472 | Y468 | Y492* | Y484* | Unknown | PhosphoSitePlus® |
| Y473 | Y469 | Y493* | Y485 | Unknown | PhosphoSitePlus® |
| N/A | N/A | N/A | S532* | Unknown | 62 |
| N/A | N/A | Y604* | Y596* | Unknown | PhosphoSitePlus® |
| Y606 | Y602 | Y627* | Y619* | Unknown | PhosphoSitePlus® |
| S616* | S612 | N/A | N/A | Unknown | SGD |
| S619* | S615 | N/A | N/A | Unknown | SGD |
| S657* | S653 | S677 | S668 | Unknown | 64 |
| S663* | S659 | N/A | N/A | Unknown | 64 |
| N/A | N/A | T725* | N/A | Unknown | 62 |
| N/A | N/A | S726* | S718* | Unknown | 62 |
Identified phosphorylation residues are marked by asterisk (*); orthologue residue is also shown when present. Data were taken from published literature and the websites PhosphoSitePlus® (www.phosphosite.org) and Saccharomyces Genome Database (SGD) (www.yeastgenome.org).
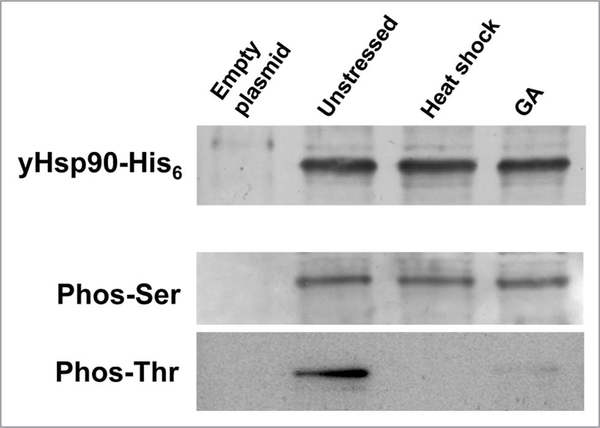
Simple baker’s yeast, Saccharomyces cerevisiae, is a well-established and valuable tool for studying various aspects of conserved protein chaperone machinery. Yeast have provided us with a powerful tool to study Hsp90 phosphorylation, since it readily allows plasmid exchange whereby any introduced Hsp90 gene—provided it is partially functional—can provide 100% of the Hsp90 of the cell. Such genetic manipulations are simply not achievable in cultured mammalian cells. Using the yeast system, it is possible to show that Hsp90 is constitutively phosphorylated on serine and theronine residues. However, Hsp90 threonine phosphorylation is lost upon either heat shock stress or treatment with the Hsp90 inhibitor geldanamycin (GA) (Fig. 2). These results agree with a previous study showing rapid dephosphorylation of Hsp90 in heat-shocked HeLa cells.44 Loss of threonine phosphorylation may impact Hsp90 function in response to heat shock stress or to inhibitory drugs.
Figure 2.
Yeast Hsp90 phosphorylation on serine (phos-Ser) and threonine (phos-Thr) residues. yHsp90-His6 was purified from yeast cells that were heat shocked at 39°C for 40 min or treated with 100 μM geldanamycin (GA) for 60 min. wild-type cells containing empty plasmid were used as negative control.
Tyrosine Phosphorylation of Hsp90
There are only few reports of Hsp90 tyrosine phosphorylation (Table 1).27 A recent study reported that c-Src directly phosphorylates Tyr300 of human Hsp90β.32,33 This is essential for VEGF-stimulated endothelial nitric oxide synthase (eNOS) association with Hsp90 and thus is necessary for nitric oxide release from endothelial cells.33
Another study reported that tyrosine-phosphorylated Hsp90 repressed the function of ionotropic P2X7 receptors. These receptors serve as ligand-gated ion channels and are responsible for ATP-dependent lysis of macrophages.31
Our recent work has shown that Wee1Swe1 phosphorylates Hsp90. Swe1 is the only “true” tyrosine kinase in budding yeast.45 It phosphorylates and inhibits the kinase activity of the main cell cycle cyclin-dependent kinase Cdc28 (human Cdc2) thereby regulating the G2/M transition.46–50 Initial studies showed that in Scizosaccharomyces pombe, formation of an active Wee1 tyrosine kinase depends on its interaction with Hsp90.26 Subsequent work showed that inhibiting Hsp90 chaperone function with GA led to the proteasome-mediated degradation of Wee1Swe1.51,52
Wee1Swe1 Dependent Tyrosine Phosphorylation of Hsp90 Regulates its Chaperone Function
Wee1Swe1 phosphorylates a conserved tyrosine residue (Y24 in yHsp90 and Y38 in hHsp90α) in the N-domain of a subpopulation of nuclear-localized Hsp90 in a cell cycle-dependent manner.25 Cytosolic relocation of phosphotyrosyl yHsp90 precedes its ubiquitination and degradation by proteasomes. This appears to be the “switching off” mechanism for this form of the chaperone, since we were unable to identify a tyrosine phosphatase capable of dephosphorylating yHsp90 (Fig. 3).
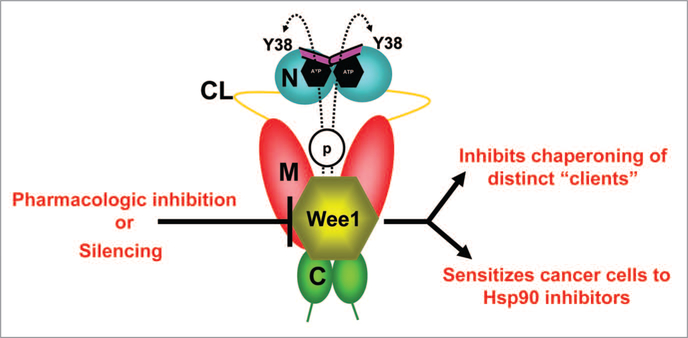
Figure 3.
wee1, an Hsp90 client protein, phosphorylates a conserved tyrosine residue (Y38) in the N-domain of a subpopulation of nuclear-localized yHsp90. Phosphorylation also leads to ubiquitination and degradation of Hsp90 by cytoplasmic proteasomes. Pharmacologic inhibition/molecular silencing of wee1 inhibits Hsp90 chaperoning of distinct clients and sensitizes cells to Hsp90 inhibitor-induced apoptosis. Domain labeling is as follows: N, N-domain (blue); CL, charged linker (yellow); M, M-domain (red); C, C-domain (green); ATP lid, (purple).
Phosphorylation of Hsp90 by Wee1Swe1 is not essential for yeast survival but the non-phosphorylatable Hsp90 mutant (yHsp90-Y24F) fails to interact with the co-chaperone Aha1 and shows significantly reduced interaction with p23/Sba1 (Fig. 1). Identical results were also observed with the human Hsp90 mutant (Y38F) in mammalian cells. These data suggest that phosphorylation of Y24/Y38 affects the equilibrium between open and closed states of Hsp90. Importantly, some clients, including the protein kinases pp60v-Src, Raf-1 (Ste11 in yeast), ErbB2, Mpk1/Slt2 (yeast MAP kinase), and the transcription factor heat shock factor 1 (yeast Hsf1), seem to require Wee1Swe1mediated tyrosine phosphorylation of Hsp90. However, Hsp90 chaperoning of other clients, including the glucocorticoid receptor (GR) and androgen receptor (AR), appears not to be affected by the phosphorylation status of this residue. This is the first demonstration that phosphorylation permits Hsp90 to switch from an inactive to an active state for chaperoning of a subset of clients while not impacting the chaperoning of other client proteins.25
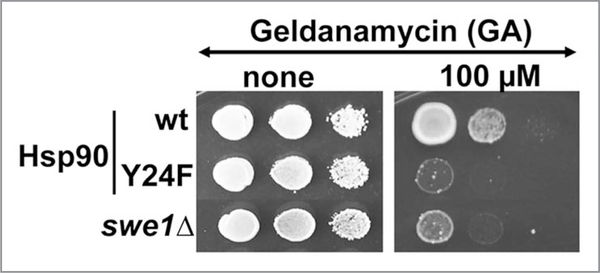
As part of this study, we found that yHsp90-Y24F, although having a similar in vitro affinity for GA (Kd 3.6 ± 0.3 μM) as wild-type yHsp90 (Kd 2.6 ± 0.3 μM), demonstrated more drug binding in vivo, as did wild-type Hsp90 expressed in swe1Δ yeast cells. These data are complemented by enhanced growth sensitivity of the mutants to GA compared to wild-type cells (Fig. 4). These observations in yeast were corroborated in cancer cells, where silencing of WEE1 or pharmacologic inhibition of Wee1 kinase sensitized cells to Hsp90 inhibitor (Fig. 3).25
Figure 4.
Yeast cells expressing yHsp90-Y24F and swe1Δ cells are hypersensitive to GA. Yeast cells were grown to mid-log and then a 1:10 dilution series were spotted on YPD agar containing 100 μM GA. Plates were incubated at 25°C for 4 days.
Cell Cycle Consequences of Hsp90 Tyrosine Phosphorylation
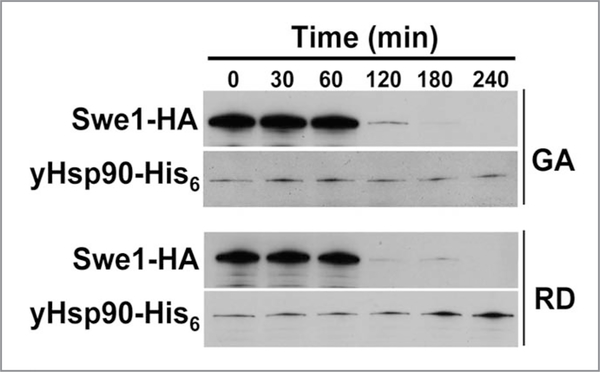
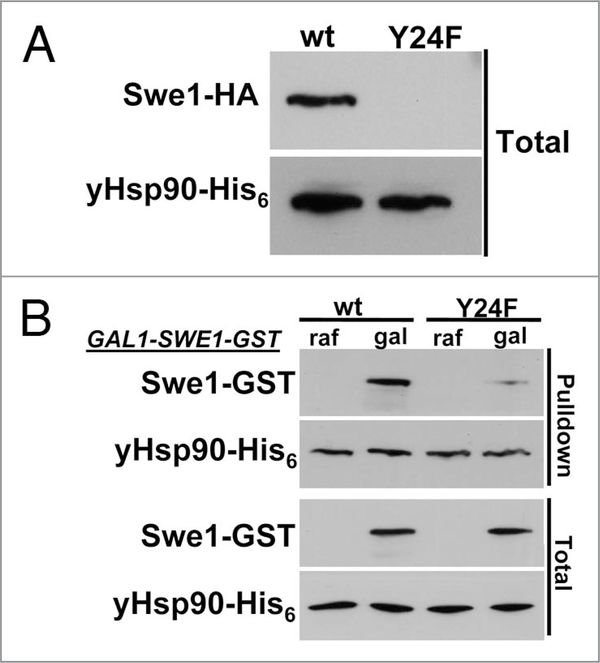
Treating yeast cells with Hsp90 inhibitors destabilizes Swe1 (Fig. 5). Interestingly, GA treatment of yeast cells for 1 hr, significantly reduces yHsp90 tyrosine phosphorylation and also increases the percentage of cells in G1, suggesting a possible direct impact on Swe1. Indeed, we found Swe1 to be poorly expressed in yHsp90-Y24F yeast (Fig. 6A). We next overexpressed Swe1-GST, under a galactose inducible promoter (GAL1), in wild-type and yHsp90-Y24F yeast. Association of Swe1-GST with yHsp90-Y24F was markedly reduced compared to wild-type yeast cells (Fig. 6B). Taken together, these data suggests that Wee1Swe1-dependent tyrosine phosphorylation of Hsp90 is important to strengthen the Wee1Swe1-Hsp90 chaperone complex and to permit Hsp90 to effectively chaperone this tyrosine kinase.
Figure 5.
Effect of Hsp90 inhibitors GA or radicicol (RD) on the stability of Swe1 tyrosine kinase. Yeast cells were grown to mid-log and then treated with 50 μM GA or RD. Swe1-HA in yeast lysate was detected by western blot using an anti-HA-monoclonal antibody. yHsp90-His6 was used as loading control.
Figure 6.
Swe1 destabilization in yHsp90-Y24F-expressing yeast. (A) western blotting was used to detect Swe1-HA in yeast cell lysate expressing either wild-type yHsp90 or yHsp90-Y24F. yHsp90-His6 was used as loading control. (B) Association of GST-tagged Swe1 with wild-type yHsp90 and yHsp90-Y24F. GST-tagged Swe1 under galactose (gal) inducible promoter (GAL1) was expressed in yeast cells containing either wild-type yHsp90 or yHsp90-Y24F. Swe1-GST co-precipitating with yHsp90-His6 was detected by western blotting.
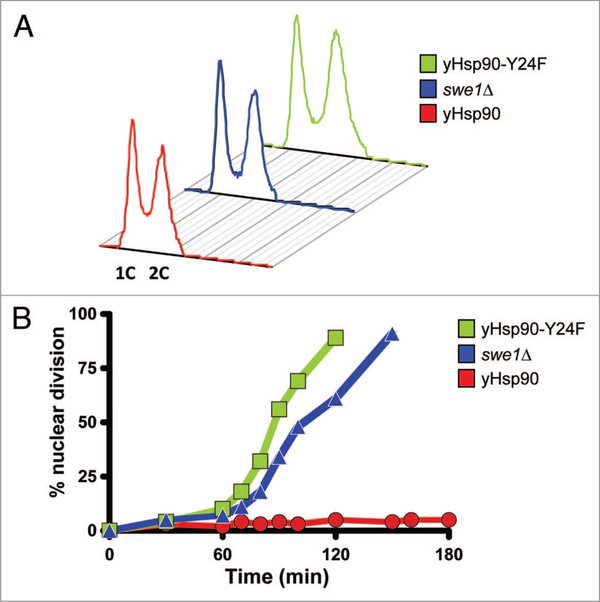
Deletion of SWE1 causes a short delay in entry into mitosis but the length of G2 is unaltered. Flow cytometric analysis (FACS) showed that asynchronously growing yHsp90-Y24F mutants and swe1Δ cells both had a similar proportion of cells with 1C and 2C DNA content compared to wild-type cells (Fig. 7A). We then arrested these cells in G1-phase with α-factor and then released them by incubation in fresh media containing 50 μM Latrunculin-A (Lat-A) in order to trigger checkpoint-mediated G2 arrest. Unlike wild-type cells, the yHsp90-Y24F mutants underwent premature nuclear division, as did swe1Δ cells (Fig. 7B). These data suggest that yHsp90-Y24F mutants, like swe1Δ cells, have a defective G2/M cell cycle checkpoint. This is fully consistent with the observed destabilization of Swe1 in yHsp90-Y24F cells. Previous reports have suggested that proteolytic destruction of Swe1 is the key step in its deactivation and allows entry into mitosis.53,54 Our data implicate Hsp90 phosphorylation status (because it regulates Hsp90-Swe1 association) in this process.
Figure 7.
Lack of G2/M checkpoint-induced delay of nuclear division in yHsp90-Y24F and swe1Δ cells. (A) Flow cytometric analysis of the DNA content of asynchronously growing wild-type, swe1Δ, and yHsp90-Y24F yeast cells. Occupancy of G2 is unaltered in the two mutants when compared to wild-type cells (wild-type, 48.7%; swe1Δ, 49.0%; yHsp90-Y24F, 51.8%). (B) Cells were released from α-factor-induced cell cycle arrest into fresh medium containing 50 μM Lat-A. inclusion of Lat-A causes arrest at the G2/M checkpoint. At the indicated times, cell aliquots were removed, fixed and stained with DAPi to visualize DNA, and 100 cells were scored. Premature nuclear division is apparent in both yHsp90-Y24F mutant and swe1Δ cells.
Concluding Remarks
In eukaryotes, the regulation of Hsp90 function is complex. Phosphorylation events have been shown to fine tune Hsp90 chaperone activity.2,27,33,55,56 Our recent work uncovered a unique role for Wee1Swe1 in regulating Hsp90. We identified a single conserved tyrosine residue in the N-domain of Hsp90, whose phosphorylation status likely permits prolonged association of Hsp90 with some of its client proteins. We also demonstrated that lack of phosphorylation at this tyrosine residue enhanced Hsp90 binding to inhibitory drugs. Here, we show that, as is the case in cancer cells, prevention of this tyrosine phosphorylation makes yeast cells hypersensitive to Hsp90 inhibition. We also provide additional data suggesting that the stability of Wee1Swe1 not only depends on its interaction with Hsp90, but also on its ability to phosphorylate this molecular chaperone. These observations demonstrate an unexpected role for Wee1Swe1 in regulating Hsp90 function and, consequently, in determining its own ability to regulate the G2/M checkpoint.
Acknowledgements
We thank our colleagues and collaborators, Professors Laurence H. Pearl and Peter W. Piper, Drs. Chris Prodromou, Jane Trepel, Brian Blagg, William G. Stetler-Stevenson, Giorgio Colombo, Barry Panaretou, Dimitra Bourboulia, Min-Jung Lee, Giulia Morra, and Bradley T. Scroggins, and Kristin Beebe, Sunmin Lee, and Alison C. Donnelly for their scientific contributions and stimulating discussions. We are grateful to Dr. Dimitra Bourboulia for the FACS analyses. The Intramural Research Program of the National Cancer Institute supported this work.
References
- 1.Pearl LH, Prodromou C, Workman P. The Hsp90 molecular chaperone: an open and shut case for treatment. Biochem J 2008; 410:439–53. [DOI] [PubMed] [Google Scholar]
- 2.Wandinger SK, Richter K, Buchner J. The Hsp90 chaperone machinery. J Biol Chem 2008; 283:18473–7. [DOI] [PubMed] [Google Scholar]
- 3.Neckers L Heat shock protein 90: the cancer chaperone. J Biosci 2007; 32:517–30. [DOI] [PubMed] [Google Scholar]
- 4.Rutherford SL, Lindquist S. Hsp90 as a capacitor for morphological evolution. Nature 1998; 396:336–42. [DOI] [PubMed] [Google Scholar]
- 5.Zhao R, Davey M, Hsu YC, Kaplanek P, Tong A, Parsons AB, et al. Navigating the chaperone network: an integrative map of physical and genetic interactions mediated by the hsp90 chaperone. Cell 2005; 120:715–27. [DOI] [PubMed] [Google Scholar]
- 6.Millson SH, Truman AW, Wolfram F, King V, Panaretou B, Prodromou C, et al. Investigating the protein-protein interactions of the yeast Hsp90 chaperone system by two-hybrid analysis: potential uses and limitations of this approach. Cell Stress Chaperones 2004; 9:359–68. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kim YS, Alarcon SV, Lee S, Lee MJ, Giaccone G, Neckers L, et al. Update on Hsp90 inhibitors in clinical trial. Curr Top Med Chem 2009; 9:1479–92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Sollars V, Lu X, Xiao L, Wang X, Garfinkel MD, Ruden DM. Evidence for an epigenetic mechanism by which Hsp90 acts as a capacitor for morphological evolution. Nat Genet 2003; 33:70–4. [DOI] [PubMed] [Google Scholar]
- 9.Yeyati PL, van Heyningen V. Incapacitating the evolutionary capacitor: Hsp90 modulation of disease. Curr Opin Genet Dev 2008; 18:264–72. [DOI] [PubMed] [Google Scholar]
- 10.Ali MM, Roe SM, Vaughan CK, Meyer P, Panaretou B, Piper PW, et al. Crystal structure of an Hsp90-nucleotide-p23/Sba1 closed chaperone complex. Nature 2006; 440:1013–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Prodromou C, Roe SM, O’Brien R, Ladbury JE, Piper PW, Pearl LH. Identification and structural characterization of the ATP/ADP-binding site in the Hsp90 molecular chaperone. Cell 1997; 90:65–75. [DOI] [PubMed] [Google Scholar]
- 12.Graf C, Stankiewicz M, Kramer G, Mayer MP. Spatially and kinetically resolved changes in the conformational dynamics of the Hsp90 chaperone machine. EMBO J 2009; 28:602–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hainzl O, Lapina MC, Buchner J, Richter K. The charged linker region is an important regulator of Hsp90 function. J Biol Chem 2009; 284:22559–67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Tsutsumi S, Mollapour M, Graf C, Lee CT, Scroggins BT, Xu W, et al. Hsp90 charged-linker truncation reverses the functional consequences of weakened hydrophobic contacts in the N domain. Nat Struct Mol Biol 2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Panaretou B, Prodromou C, Roe SM, O’Brien R, Ladbury JE, Piper PW, et al. ATP binding and hydrolysis are essential to the function of the Hsp90 molecular chaperone in vivo. EMBO J 1998; 17:4829–36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Shiau AK, Harris SF, Southworth DR, Agard DA. Structural Analysis of E. coli hsp90 reveals dramatic nucleotide-dependent conformational rearrangements. Cell 2006; 127:329–40. [DOI] [PubMed] [Google Scholar]
- 17.Hessling M, Richter K, Buchner J. Dissection of the ATP-induced conformational cycle of the molecular chaperone Hsp90. Nat Struct Mol Biol 2009; 16:287–93. [DOI] [PubMed] [Google Scholar]
- 18.Mickler M, Hessling M, Ratzke C, Buchner J, Hugel T. The large conformational changes of Hsp90 are only weakly coupled to ATP hydrolysis. Nat Struct Mol Biol 2009; 16:281–6. [DOI] [PubMed] [Google Scholar]
- 19.Southworth DR, Agard DA. Species-dependent ensembles of conserved conformational states define the Hsp90 chaperone ATPase cycle. Mol Cell 2008; 32:631–40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Zuehlke A, Johnson JL. Hsp90 and co-chaperones twist the functions of diverse client proteins. Biopolymers 2009; 93:211–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Phillips JJ, Yao ZP, Zhang W, McLaughlin S, Laue ED, Robinson CV, et al. Conformational dynamics of the molecular chaperone Hsp90 in complexes with a co-chaperone and anticancer drugs. J Mol Biol 2007; 372:1189–203. [DOI] [PubMed] [Google Scholar]
- 22.Scroggins BT, Neckers L. Post-translational modification of heat shock protein 90: impact on chaperone function. Expert Opin Drug Discov 2007; 2:1403–14. [DOI] [PubMed] [Google Scholar]
- 23.Martinez-Ruiz A, Villanueva L, Gonzalez de Orduna C, Lopez-Ferrer D, Higueras MA, Tarin C, et al. S-nitrosylation of Hsp90 promotes the inhibition of its ATPase and endothelial nitric oxide synthase regulatory activities. Proc Natl Acad Sci USA 2005; 102:8525–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Fiskus W, Ren Y, Mohapatra A, Bali P, Mandawat A, Rao R, et al. Hydroxamic acid analogue histone deacetylase inhibitors attenuate estrogen receptor-alpha levels and transcriptional activity: a result of hyperacetylation and inhibition of chaperone function of heat shock protein 90. Clin Cancer Res 2007; 13:4882–90. [DOI] [PubMed] [Google Scholar]
- 25.Mollapour M, Tsutsumi S, Donnelly AC, Beebe K, Tokita MJ, Lee MJ, et al. Swe1Wee1-dependent tyrosine phosphorylation of Hsp90 regulates distinct facets of chaperone function. Mol Cell 2010; 37:333–43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Aligue R, Akhavan-Niak H, Russell P. A role for Hsp90 in cell cycle control: Wee1 tyrosine kinase activity requires interaction with Hsp90. EMBO J 1994; 13:6099–106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Mimnaugh EG, Worland PJ, Whitesell L, Neckers LM. Possible role for serine/threonine phosphorylation in the regulation of the heteroprotein complex between the hsp90 stress protein and the pp60v-src tyrosine kinase. J Biol Chem 1995; 270:28654–9. [DOI] [PubMed] [Google Scholar]
- 28.Garnier C, Lafitte D, Jorgensen TJ, Jensen ON, Briand C, Peyrot V. Phosphorylation and oligomerization states of native pig brain HSP90 studied by mass spectrometry. Eur J Biochem 2001; 268:2402–7. [DOI] [PubMed] [Google Scholar]
- 29.Zhao YG, Gilmore R, Leone G, Coffey MC, Weber B, Lee PW. Hsp90 phosphorylation is linked to its chaperoning function. Assembly of the reovirus cell attachment protein. J Biol Chem 2001; 276:32822–7. [DOI] [PubMed] [Google Scholar]
- 30.Fontana J, Fulton D, Chen Y, Fairchild TA, McCabe TJ, Fujita N, et al. Domain mapping studies reveal that the M domain of hsp90 serves as a molecular scaffold to regulate Akt-dependent phosphorylation of endothelial nitric oxide synthase and NO release. Circ Res 2002; 90:866–73. [DOI] [PubMed] [Google Scholar]
- 31.Adinolfi E, Kim M, Young MT, Di Virgilio F, Surprenant A. Tyrosine phosphorylation of HSP90 within the P2X7 receptor complex negatively regulates P2X7 receptors. J Biol Chem 2003; 278:37344–51. [DOI] [PubMed] [Google Scholar]
- 32.Wang C, Chen J. Phosphorylation and hsp90 binding mediate heat shock stabilization of p53. J Biol Chem 2003; 278:2066–71. [DOI] [PubMed] [Google Scholar]
- 33.Duval M, Le Boeuf F, Huot J, Gratton JP. Src-mediated phosphorylation of Hsp90 in response to vascular endothelial growth factor (VEGF) is required for VEGF receptor-2 signaling to endothelial NO synthase. Mol Biol Cell 2007; 18:4659–68. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Miyata Y, Yahara I. The 90-kDa heat shock protein, HSP90, binds and protects casein kinase II from self-aggregation and enhances its kinase activity. J Biol Chem 1992; 267:7042–7. [PubMed] [Google Scholar]
- 35.Ogiso H, Kagi N, Matsumoto E, Nishimoto M, Arai R, Shirouzu M, et al. Phosphorylation analysis of 90 kDa heat shock protein within the cytosolic arylhydrocarbon receptor complex. Biochemistry 2004; 43:15510–9. [DOI] [PubMed] [Google Scholar]
- 36.Lees-Miller SP, Anderson CW. The human double-stranded DNA-activated protein kinase phosphorylates the 90-kDa heat-shock protein, hsp90 alpha at two NH2-terminal threonine residues. J Biol Chem 1989; 264:17275–80. [PubMed] [Google Scholar]
- 37.Lees-Miller SP, Anderson CW. Two human 90-kDa heat shock proteins are phosphorylated in vivo at conserved serines that are phosphorylated in vitro by casein kinase II. J Biol Chem 1989; 264:2431–7. [PubMed] [Google Scholar]
- 38.Rose DW, Wettenhall RE, Kudlicki W, Kramer G, Hardesty B. The 90-kilodalton peptide of the heme-regulated eIF-2alpha kinase has sequence similarity with the 90-kilodalton heat shock protein. Biochemistry 1987; 26:6583–7. [DOI] [PubMed] [Google Scholar]
- 39.Kurokawa M, Zhao C, Reya T, Kornbluth S. Inhibition of apoptosome formation by suppression of Hsp90beta phosphorylation in tyrosine kinase-induced leukemias. Mol Cell Biol 2008; 28:5494–506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Old WM, Shabb JB, Houel S, Wang H, Couts KL, Yen CY, et al. Functional proteomics identifies targets of phosphorylation by B-Raf signaling in melanoma. Mol Cell 2009; 34:115–31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Barati MT, Rane MJ, Klein JB, McLeish KR. A proteomic screen identified stress-induced chaperone proteins as targets of Akt phosphorylation in mesangial cells. J Proteome Res 2006; 5:1636–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Lei H, Venkatakrishnan A, Yu S, Kazlauskas A. Protein kinase A-dependent translocation of Hsp90 alpha impairs endothelial nitric-oxide synthase activity in high glucose and diabetes. J Biol Chem 2007; 282:9364–71. [DOI] [PubMed] [Google Scholar]
- 43.Wandinger SK, Suhre MH, Wegele H, Buchner J. The phosphatase Ppt1 is a dedicated regulator of the molecular chaperone Hsp90. EMBO J 2006; 25:367–76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Legagneux V, Morange M, Bensaude O. Heat shock increases turnover of 90 kDa heat shock protein phosphate groups in HeLa cells. FEBS Lett 1991; 291:359–62. [DOI] [PubMed] [Google Scholar]
- 45.Kellogg DR. Wee1-dependent mechanisms required for coordination of cell growth and cell division. J Cell Sci 2003; 116:4883–90. [DOI] [PubMed] [Google Scholar]
- 46.Booher RN, Deshaies RJ, Kirschner MW. Properties of Saccharomyces cerevisiae wee1 and its differential regulation of p34CDC28 in response to G1 and G2 cyclins. EMBO J 1993; 12:3417–26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Harvey SL, Kellogg DR. Conservation of mechanisms controlling entry into mitosis: budding yeast wee1 delays entry into mitosis and is required for cell size control. Curr Biol 2003; 13:264–75. [DOI] [PubMed] [Google Scholar]
- 48.Jorgensen P, Nishikawa JL, Breitkreutz BJ, Tyers M. Systematic identification of pathways that couple cell growth and division in yeast. Science 2002; 297:395–400. [DOI] [PubMed] [Google Scholar]
- 49.Lew DJ. The morphogenesis checkpoint: how yeast cells watch their figures. Curr Opin Cell Biol 2003; 15:648–53. [DOI] [PubMed] [Google Scholar]
- 50.McGowan CH, Russell P. Human Wee1 kinase inhibits cell division by phosphorylating p34cdc2 exclusively on Tyr15. EMBO J 1993; 12:75–85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Goes FS, Martin J. Hsp90 chaperone complexes are required for the activity and stability of yeast protein kinases Mik1, Wee1 and Swe1. Eur J Biochem 2001; 268:2281–9. [DOI] [PubMed] [Google Scholar]
- 52.Munoz MJ, Bejarano ER, Daga RR, Jimenez J. The identification of Wos2, a p23 homologue that interacts with Wee1 and Cdc2 in the mitotic control of fission yeasts. Genetics 1999; 153:1561–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.McMillan JN, Sia RA, Bardes ES, Lew DJ. Phosphorylation-independent inhibition of Cdc28p by the tyrosine kinase Swe1p in the morphogenesis checkpoint. Mol Cell Biol 1999; 19:5981–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Sia RA, Bardes ES, Lew DJ. Control of Swe1p degradation by the morphogenesis checkpoint. EMBO J 1998; 17:6678–88. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Aoyagi S, Archer TK. Modulating molecular chaperone Hsp90 functions through reversible acetylation. Trends Cell Biol 2005; 15:565–7. [DOI] [PubMed] [Google Scholar]
- 56.Scroggins BT, Prince T, Shao J, Uma S, Huang W, Guo Y, et al. High affinity binding of Hsp90 is triggered by multiple discrete segments of its kinase clients. Biochemistry 2003; 42:12550–61. [DOI] [PubMed] [Google Scholar]
- 57.Guo A, Villen J, Kornhauser J, Lee KA, Stokes MP, Rikova K, et al. Signaling networks assembled by oncogenic EGFR and c-Met. Proc Natl Acad Sci USA 2008; 105:692–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Rikova K, Guo A, Zeng Q, Possemato A, Yu J, Haack H, et al. Global survey of phosphotyrosine signaling identifies oncogenic kinases in lung cancer. Cell 2007; 131:1190–203. [DOI] [PubMed] [Google Scholar]
- 59.Molina H, Horn DM, Tang N, Mathivanan S, Pandey A. Global proteomic profiling of phosphopeptides using electron transfer dissociation tandem mass spectrometry. Proc Natl Acad Sci USA 2007; 104:2199–204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Olsen JV, Blagoev B, Gnad F, Macek B, Kumar C, Mortensen P, et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 2006; 127:635–48. [DOI] [PubMed] [Google Scholar]
- 61.Matsuoka S, Ballif BA, Smogorzewska A, McDonald ER, 3rd,, Hurov KE, Luo J, et al. ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science 2007; 316:1160–6. [DOI] [PubMed] [Google Scholar]
- 62.Dephoure N, Zhou C, Villen J, Beausoleil SA, Bakalarski CE, Elledge SJ, et al. A quantitative atlas of mitotic phosphorylation. Proc Natl Acad Sci USA 2008; 105:10762–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Huang SY, Tsai ML, Chen GY, Wu CJ, Chen SH. A systematic MS-based approach for identifying in vitro substrates of PKA and PKG in rat uteri. J Proteome Res 2007; 6:2674–84. [DOI] [PubMed] [Google Scholar]
- 64.Albuquerque CP, Smolka MB, Payne SH, Bafna V, Eng J, Zhou H. A multidimensional chromatography technology for in-depth phosphoproteome analysis. Mol Cell Proteomics 2008; 7:1389–96. [DOI] [PMC free article] [PubMed] [Google Scholar]