Abstract
Purpose
Schlemm’s canal (SC) endothelial cells derived from donors with or without glaucoma showed different mechanical properties and gene expression. As an important contributor to the regulation of intraocular pressure (IOP) and pathogenesis of primary open-angle glaucoma (POAG), the heritable key epigenetic changes, methylation may play an important role in the physiologic function of SC cells. This study aims to identify differentially methylated CpG sites (DMSs) in primary cultures of human SC cells with or without glaucoma.
Methods
We examined the methylation pattern of seven strains of primary human cells (two glaucoma and five normal SC cell samples), which were isolated and characterized using established protocols. DNA methylation was profiled using Illumina Human Methylation 450 BeadChip. Raw data were extracted and exported using Illumina GenomeStudio software. After quantile normalization, DNA methylation data were analyzed using R package RnBeads in Bioconductor. DMSs were filtered with p ≤ 1E-5, methylation change ≥ 0.1, and false discovery rate ≤ 0.05. The closest genes and the location of each CpG site were annotated using R package FDb.InfiniumMethylation.hg19. Gene Ontology and pathway analysis was performed using WebGestalt. Selected DMSs were validated using the Zymo qMethyl kit.
Results
We used five non-glaucoma and two glaucomatous SC cell samples to profile genome-wide DNA methylation using Illumina Infinium Methylation BeadChips. Principle component analysis showed the separation between the glaucoma and control samples. After quality control and differential analysis, we identified 298 highly significant DMSs (p ≤ 1E-5). Among them, 221 DMSs were within 1 kb of a nearby gene. Gene Ontology analysis demonstrated significant enrichment in positive regulation of cell migration, negative regulation of endothelial cell proliferation, and stress fiber and actin filament bundles. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis showed enrichment in cell adhesion and gap junctions. Several glaucoma-related genes were identified, including TGFBR3, THBS1, PITX2, DAXX, TBX3, TNXB, ANGPT1, and PLEKHA7. We also examined differentially methylated regions (DMRs) near these CpG sites and identified significant DMRs in TBX3, TNXB1, DAXX, and PITX2.
Conclusions
This study represents the first genome-wide DNA methylation profiling in cultured human primary SC cells. The DMSs were enriched in the pathways related to outflow resistance. Several DMRs were validated in glaucoma-associated genes, further suggesting the role of DNA methylation in glaucoma development. This study could provide comprehensive understanding of DNA methylation in glaucoma and its effect on aqueous humor outflow.
Introduction
Glaucoma is a progressive neurodegenerative disorder and the leading cause of irreversible blindness worldwide [1]. Primary open-angle glaucoma (POAG) is the most common subtype with an unspecified cause. POAG is characterized by the progressive loss of retinal ganglion cells, optic nerve cupping, and visual field loss [2]. Risk factors include advanced aging, elevated intraocular pressure (IOP), positive family history, and African ancestry [3]. Among them, elevated IOP is the main risk factor and the only effective target for clinical treatment [4]. The balance between secretion and drainage of aqueous humor in the anterior chamber determines IOP [5]. In most cases, elevated IOP is caused by increased outflow resistance, which is mainly generated by cells in the juxtacanalicular (JCT) region, where trabecular meshwork (TM) and Schlemm’s canal (SC) endothelial cells interact [6,7]. In glaucomatous eyes, the mechanical properties of JCT tissue are altered, becoming more fibrotic [8]. Significantly, cultured glaucomatous SC cells show greater stiffness and similar mechanical properties to glaucomatous JCT tissue [9]. Moreover, expression profiling of cultured SC cells with or without glaucoma revealed significant differential expression of genes involved in extracellular matrix (ECM) remodeling [10]. These corresponding changes suggested a heritable phenotype with cell division, which may be ascribed to epigenetic changes in cellular genome, such as histone modification and DNA methylation.
The contribution of genetic defects to POAG has been recognized for decades [11,12]. Several causal genes, such as myocilin (Gene ID 4653, OMIM 601652), optineurin (Gene ID 10133, OMIM 602432), and TBK1 (Gene ID 29110, OMIM 604834), have been identified [13-15]; however, mutations in these Mendelian genes account for ≤ 10% POAG cases worldwide [16,17]. Additionally, large-cohort genome-wide association studies (GWASs) have identified many associated single nucleotide polymorphisms (SNPs) associated with POAG and IOP [18-22]. Currently, POAG is considered to be a complex disease affected by genetic and environmental factors [17,23]. DNA methylation, the most well-studied epigenetic regulators in response to environmental factors, may inhibit gene transcription by covalent addition of methyl groups to DNA using DNA methyltransferase (DNMT), and may reverse the inhibition by cell division or removal of methyl groups through tet methylcytosine dioxygenase (TET) enzymes [24]. The methyl group is susceptibly added to CpG sites, where cytosine is followed by a guanine. The effect of DNA methylation is dependent on the density of CpG sites, especially high CG content regions named CpG islands, and their locations within a gene [25]. Although the mechanism of environmental factors in DNA methylation remains unclear, especially the gene-specific DNA methylation patterns, numerous studies support the susceptibility of DNA methylation to environmental risk factors in relation to eye diseases [26,27]. However, little is known about the effect of DNA methylation in the outflow facility glaucoma pathogenesis. In this study, we aimed to identify differentially methylated DNA loci in glaucomatous SC endothelial cells using genome-wide methylation profiling. The identification of such DNA methylation patterns holds promise to provide more information about how epigenetic factors play a role in glaucoma pathogenesis.
Methods
Isolation and culture of SC cells
Human SC endothelial cells were isolated from postmortem eyes provided by Miracles in Sight (Winston-Salem, NC), Midwest Eye Bank (Ann Arbor, MI), National Disease Research Interchange (Philadelphia, PA), or Life Legacy (Tucson, AZ) within 36 h of death with enucleation occurring ≤ 6 h after death. Isolation and culture of primary SC cells were processed according to the established techniques [28]. All strains of primary SC cells were characterized using five inclusion criteria: (1) the expression of vascular endothelial cadherin, (2) a net trans-endothelial electrical resistance ≥10 ohms·cm2, (3) lack of myocilin induction by dexamethasone, (4) monolayers that formed non-overlapping, linear-arranged morphology, and (5) contact-inhibited growth. SC cells were isolated from two donors with glaucoma and five donors without glaucoma. The determination of glaucoma was based on a combination of information provided by the eye banks: a history of ocular hypertension or glaucoma or the presence of glaucoma eye drops on the patient’s medication list. All the cultured SC cells were harvested between passages 3 and 5, as cell stiffness remains consistent through passage 6 [9].
DNA extraction
Genomic DNA was extracted using the AllPrep DNA Mini spin column kit (Qiagen, Valencia, CA) according to the manufacturer’s instructions. Briefly, cultured cells were harvested and lysed using provided buffer RLT. The lysate was collected and loaded to the spin column. After purification, 50 µl of RNase-free water was added to elute DNA from the spin column. The DNA concentration was measured using the TECAN Infinite M200PRO microplate reader (Tecan, Mannedorf, Switzerland).
Identification of differentially methylated CpG sites
DNA samples were labeled and hybridized to the Illumina Infinium HumanMethylation450 BeadChips and scanned by the Illumina iScan System following the manufacturer’s protocol. Briefly, 1 µg of high-quality DNA samples were treated with bisulfite to convert unmethylated cytosines to uracils while methylated cytosines were protected and remained cytosine. Predesigned probes for each CpG site, flagged with special fluorescence, were used to determine whether the base at a given locus was converted or not converted, leading to a profile of the DNA methylation status at this locus. Methylation status was calculated as the β value: the ratio of the signal from a methylated probe relative to the sum of the methylated and unmethylated probes. This value ranges from 0 (unmethylated) to 1 (fully methylated). This platform enabled sensitive and reproducible methylation profiling with quantitative measurements. Quality controls from the Illumina BeadChips were checked for experimental procedures, including bisulfite conversion, staining, hybridization, target removal, specificity, negative, and non-polymorphism. The internal bisulfite conversion control Infinium I and II probes indicated the successful conversion upon initial bisulfite treatment of the DNA samples. The experiment was performed by the Genotyping Core Facility at University of Miami (Coral Gables, FL).
Probe and intensity data for each sample were imported into R using package RnBeads in Bioconductor [29], followed by preprocessing to filter probes outside the CpG context, SNP probes, sex chromosome probes, probes without intensity value, and probes with a low standard deviation. Then preprocessed data were normalized using the subset quantile normalization (SWAN) function. Data from duplicated samples were verified to have similar β results and averaged for further analysis. The Rnbeads.hg19 package was used to annotate the location of each probe in chromosomes and CpG islands. Principle component analysis (PCA) was performed to show genome-wide methylation variation between each biologic sample. Next, the glaucoma samples were grouped and compared to the set of control samples to identify differentially methylated CpG sites (DMSs) using the limma package. The most significant DMSs were filtered as p ≤ 1E-5, false discovery rate (FDR) ≤ 0.05, and absolute difference in β values (|Δβ|) greater than 0.1. Because there were only two glaucoma SC samples, we assumed that the control and glaucoma groups had similar variations. The FDb.InfiniumMethylation.hg19 package [30] and Infinium Human-Methylation450K v1.2 product files were imported to annotate all CpG sites with nearest genes, distance to the gene, and location within the gene.
Gene Ontology and pathway analysis
First, the list of annotated genes within 1 kb of significant CpG sites (p ≤ 1E-5, |Δβ| ≥ 0.1) was uploaded into WebGestalt (WEB-based Gene SeT AnaLysis Toolkit) for Gene Ontology analysis, categorized by biologic process, molecular function, and cellular component. WebGestalt exhibited a network view in each category to reveal the relationship of each class and the location of proteins produced by target genes. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis was also performed with WebGestalt to explore the underlying biology that may contribute in the development of glaucoma.
Differentially methylated regions
The OneStep qMethyl Kit (Zymoresearch, Irvine, CA) was used to examine differentially methylated CpG regions containing selected candidate CpG sites. To perform quantitative real time polymerase chain reaction (RT-PCR) with methylation sensitive restriction enzymes (MSREs), we designed primers that included two or more MSRE targeting sites in the amplicon flanking targeted CpG sites (Table 1). The methylated sites in the region were protected from digestion of restriction enzymes and failed RT–PCR amplification. Then the percentage of methylation was estimated using the Ct between the test group (with restriction enzymes) and the reference group (without restriction enzymes). Statistical analysis was performed with GraphPad 8 using an unpaired t test with Welch’s correction (p ≤ 0.05).
Table 1. Selected primers for DMR validation using Zymo OneStep qMethyl Kit.
| Probe ID | Gene Symbol | Gene Description | Forward Primer | Reverse Primer | Amplicon Size, bp |
|---|---|---|---|---|---|
| cg13078798 |
TGFBR3 |
Transforming growth factor, β receptor III |
TGA GGA TGC ATT TGG ATG AG |
AAG GCC TGT AGG CTG CTG TA |
200 |
| cg16055869 |
CXCL5 |
Chemokine (C-X-C motif) ligand 5 |
CTC TAC CGA GAG GGA CGT TG |
AGG AGG AGC ATC TCC CAG AG |
171 |
| cg21951797 |
TNXB |
Tenascin XB |
GGA ACA CAA ATG ACC CGT AGA |
AGC CCA CGT TTT TCT AGT GA |
345 |
| cg07139946 |
GGC TCA GTC AGA CCA GGA GA |
AGG GCC AGT TTG ACT CCT TT |
255 |
||
| cg17982478 |
PLEKHA7 |
Pleckstrin homology domain contaning, family A member 7 |
CCC GAA CGC TCA GTA AAC C |
CCA GCT GAA AAC TTG GGA AA |
273 |
| cg03133735 |
PITX2 |
Paired-like homeodomain 2 |
TAC TAT GCG TTG CCG ATT CC |
AAA GAC CCC TGC TCC AAA AT |
268 |
| cg01733176 |
GTG TGG AAG GAG CTG GAC AT |
ATT GCT TGC TTT GCT TGG AC |
295 |
||
| cg26500914
cg24498636 |
DAXX |
Death-domain associated protein |
GTA CCC CAT CCA CAC CTC AC |
GGG CTG AGT GCT CTG ACT TT |
263 |
| cg05431670 |
GTC ACA GAG TTT CCG CCT TC |
GTA GAA GCA CCG GGT GAA AA |
268 |
||
| cg16277169 |
TBX3 |
T-Box 3 |
GAA ACT GGA CGA AAG GTG GA |
GGG CCA ACA GTT CTT CAA C |
300 |
| cg18161956 |
TCA GCA GCG AAA AGG TGA G |
CAG AGA GGC TAA GGG GCT TT |
300 |
||
| cg09413529 |
TGC CCG TTG AAG AAC TGT TG |
CAC CAT CTC GTC CAG CAC T |
227 |
||
| cg11246938 |
AGT GCT GGA CGA GAT GGT G |
GAG AGC AAA GAG GAG CAT GG |
201 |
||
| cg09053536 |
GAA TTC AGT TTC GGG GAA CA |
TTT GGC AAC TGA GGA GCA AT |
322 |
||
| cg13661397 | NET1 | Neuroepithelial cell transforming 1 | GAT GCG CTC AGG AGT TAA GG | CAG CGA TCA GCC AAT CAG T | 130 |
Results
Clinical phenotypes
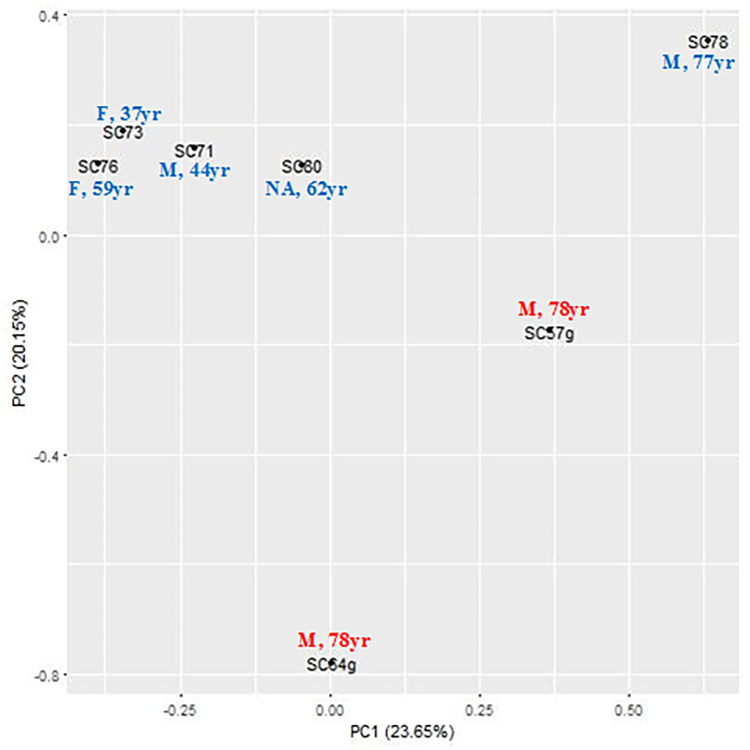
This study examined human primary SC endothelial cells from seven donors, two with glaucoma and five without glaucoma (Table 2). The average age was approximately 56 years for the controls, while the average age was 78 years for the patients with glaucoma. Although age was included as a cofactor using the R package limma in Bioconductor [31-35] to reduce the influence of varying age in the analysis, PCA showed variation of methylation pattern in samples with aging, as well as a clear separation between the control group and the glaucoma group (Figure 1).
Table 2. Clinical characteristics of donors to derive the primary endothelial cells.
| Group | Cell Strain No. | Gender | Age (years) |
|---|---|---|---|
| Glaucoma |
SC57 g |
Male |
78 |
| |
SC64 g |
Male |
78 |
| Control |
SC71 |
Male |
44 |
| |
SC73 |
Female |
37 |
| |
SC76 |
Female |
59 |
| |
SC78 |
Male |
77 |
| SC80 | NA | 62 |
Figure 1.
Principal component analysis of SC cells using genome-wide methylation data. F: female. M: male. NA: no gender information. Blue: non-glaucoma control SC samples. Red: glaucomatous SC samples.
Differential DNA methylation analysis
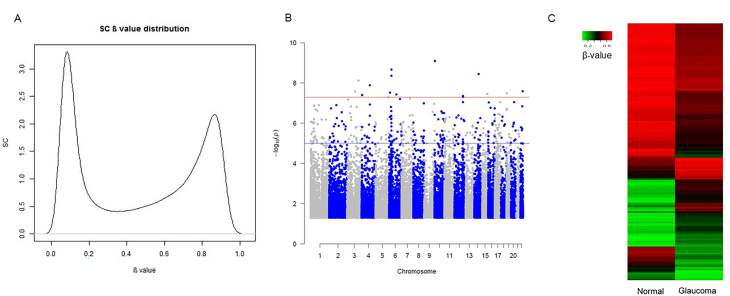
The Illumina internal control probes for bisulfite conversion indicated the successful bisulfite conversion of all DNA samples. After quality control, an average of 485,577 genomic loci were detected with methylation status in each sample. There were 383,976 identified methylation sites in the control and glaucomatous SC cells. The distribution of identified methylation status across these sites was binomial, meaning with two peaks nearing 0.1 or 0.9 of the β value, instead of one peak around 0.5 (Figure 2A).
Figure 2.
Differential methylation in human primary cultured SC cells. A: Binomial distribution of the Schlemm’s canal (SC) β values. Two peaks were found near 0.1 and 0.9. B: A Manhattan plot of differential methylation changes in single sites. The red line represents the threshold of a p value 5E-8 while the blue line represents the threshold of a p value 1E-5. C: The heat map shows different methylation patterns in SC cells with or without glaucoma.
The glaucoma samples were grouped (n = 2) and compared to the set of control samples (n = 5) to identify DMSs using the limma package. For greater accuracy of differential methylation analysis, the significance level (-logP value) of each DMS was plotted across all chromosomes in the Manhattan plots (Figure 2B). The highly statistically significant DMSs (p ≤ 1E-5, FDR < 0.05) were then filtered with significant changes ((|Δβ| ≥ 0.1) in the glaucomatous SC cells to identify 298 significant DMSs, including 106 increased DMSs and 192 decreased DMSs (Appendix 1). These sites were then used to generate a heat map to visualize the difference in methylation patterns between the two groups (Figure 2C).
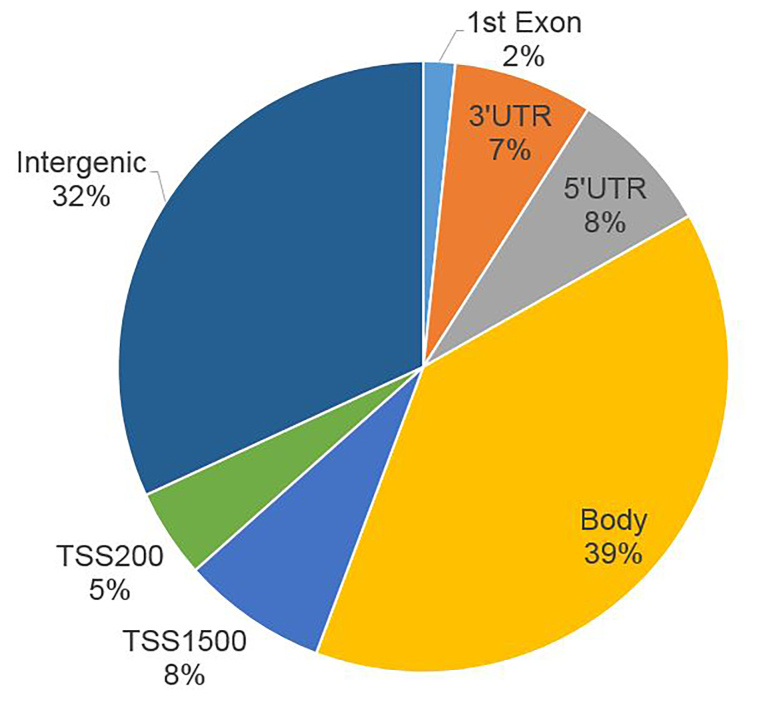
To identify glaucoma-related genes related with epigenetic alterations, we annotated all significant DMSs with their relative distance to the nearest genes. The majority of DMSs are located in the body of genes; 15% are in the untranslated region (UTR), and 13% within 1,500 bases of the transcription start site (TSS; Figure 3). According to previous studies, gene expression has a greater chance of being inhibited through methylation presented at or near the TSS, including the promoter region [25]. To identify methylation that potentially contributes to the altered expression of related genes, we filtered the distance to the nearest genes at 1,000 base pairs for greater chance of regulation. In this circumstance, we identified 221 significant DMSs located near or within 208 genes. Many of these genes are related to glaucoma, such as TGFBR3 (Gene ID 7049, OMIM 600742), LAMA3 (Gene ID 3909, OMIM 600805), PITX2 (Gene ID 5308, OMIM 601542), TNXB (Gene ID 7148, OMIM 600985), ANGPT1 (Gene ID 284, OMIM 601667), and PLEKHA7 (Gene ID 144100, OMIM 612686)[36-41]. Gene Ontology analysis indicated that these 208 genes were significantly enriched in functions related with positive regulation of cell migration, negative regulation of endothelial cell proliferation, and stress fiber and actin filament bundles (Appendix 2). KEGG pathway analysis showed significant enrichment in cell adhesion and gap junctions (Appendix 3). These results suggest that glaucoma SC cells may provide greater resistance to aqueous humor outflow with inheritable gene expression alteration.
Figure 3.
The percentage of DMS location related to the nearest genes. UTR: untranslated region. TSS: transcript start sites. Intergenic: no annotation information.
Differentially methylated region in glaucoma-related genes
The differential methylation of one CpG site has less chance to turn over gene expression, yet differential methylation of a CpG region (DMR) has a larger possibility [25]. Based on the distance of the genes, the relationship with the CpG islands, and the function of the nearest gene, we selected ten glaucoma-related genes to examine potential DMRs, including previously reported glaucoma associated genes CXCL5 (Gene ID 6374, OMIM 600324), TNXB (Gene ID 7148, OMIM 600985), ANGPT1 (Gene ID 284, OMIM 601667), PLEKHA7 (Gene ID 144100, OMIM 612686), THBS1 (Gene ID 7057, OMIM 188060), PITX2 (Gene ID 5308, OMIM 601542), DAXX (Gene ID 1616, OMIM 603186), and TBX3 (Gene ID 6926, OMIM 601621), and two genes that were differentially expressed in glaucoma SC cells, TGFBR3 (Gene ID 7049, OMIM 600742) and NET1 (Gene ID 10276, OMIM 606450)[10]. Distinct from bilsulfite sequencing, the Zymo OneStep qMethyl kit can measure the methylation level of a region with a small amount of DNA sample (Zymoresearch Cat No.D5310). Using MSRE-based RT–PCR, we examined DMRs containing one or multiple significant DMSs in these candidate genes (Table 3). Corresponding to the microarray data, significant DMRs showed the same direction of methylation status alteration and mainly located at the gene with multiple significant DMSs, such as TBX3, which has five significant DMSs and four significant DMRs flanking these CpG sites, suggesting a greater chance of gene expression inhibition. This result indicated different methylation levels between the DMSs and the DMRs.
Table 3. Selected genes with differential methylation and relative location of these CpG sites in the genes.
| Probe ID | Gene | Description | ∆β in array | p in array | ∆β in validation | p in validation | Location in gene |
|---|---|---|---|---|---|---|---|
| cg13078798 |
TGFBR3 |
Transforming growth factor, β receptor III |
0.54 |
2.17E-07 |
−0.08 |
0.376 |
Body |
| cg16055869 |
CXCL5 |
Chemokine (C-X-C motif) ligand 5 |
0.24 |
4.58E-06 |
0.08 |
0.138 |
TSS200 |
| cg21951797 |
TNXB |
Tenascin XB |
0.47 |
7.13E-06 |
0.06 |
0.002 |
Body |
| cg07139946 |
−0.18 |
1.42E-06 |
0.17 |
0.16 |
|||
| cg15724328 |
ANGPT1 |
Angiopoietin 1 |
−0.31 |
5.76E-06 |
N/A* |
|
Body |
| cg17982478 |
PLEKHA7 |
Pleckstrin homology domain contaning, family A member 7 |
0.36 |
5.35E-07 |
0.005 |
0.42 |
TSS1500 |
| cg04827020 |
THBS1 |
Thrombospondin 1 |
0.27 |
1.74E-06 |
N/A* |
|
Body |
| cg16933229 |
PITX2 |
Paired–like homeodomain 2 |
0.6 |
3.72E-07 |
N/A* |
|
|
| cg03133735 |
0.6 |
1.17E-07 |
0.02 |
0.001 |
|
||
| cg01733176 |
0.38 |
1.34E-08 |
0.1 |
0.06 |
|
||
| cg26500914 |
DAXX |
Death-domain associated protein |
0.5 |
4.11E-06 |
0.32 |
0.08 |
Body |
| cg24498636 |
0.48 |
2.26E-06 |
0.32 |
0.08 |
|||
| cg05431670 |
0.28 |
2.37E-09 |
0.07 |
0.001 |
TSS200 |
||
| cg16277169 |
TBX3 |
T-Box 3 |
−0.32 |
1.10E-07 |
−0.56 |
0.04 |
Body |
| cg18161956 |
−0.44 |
4.25E-08 |
−0.14 |
0.734 |
|||
| cg09413529 |
−0.49 |
7.55E-06 |
−0.82 |
0.001 |
|||
| cg11246938 |
−0.39 |
5.36E-08 |
−0.19 |
0.051 |
|||
| cg09053536 |
−0.51 |
7.33E-06 |
−0.89 |
0.001 |
|||
| cg13661397 | NET1 | Neuroepithelial cell transforming 1 | 0.47 | 8.37E-10 | −0.00043 | 0.861 | TSS200 |
* Not available due to shortage of sites for methylation sensitive restriction enzyme. TSS200: 200 base pairs upstream of the transcription start site; TSS1500: 1500 base pairs upstream of the transcription start site.
Discussion
To our knowledge, this is the first genome-wide DNA methylation profiling of primary human SC endothelial cells derived from patients with or without glaucoma. The glaucomatous SC cells displayed a significantly different methylation pattern than control SC cells. The majority of DMSs are located at the body of genes. Significant DMSs were enriched in genes related with stress fiber and actin filament bundles, cell adhesion and gap junctions, which may contribute to the outflow resistance. Using a microarray and the Zymo OneStep qMethyl kit, we identified several DMSs and DMRs in glaucoma-related genes, showing the potential contribution of epigenetic regulation in glaucoma development.
Despite being the most studied epigenetic regulators related to environmental factors, the role of DNA methylation is understudied in human ocular diseases, especially those related with the anterior chamber. Due to the relatively large number of TM cells and easy access in the eye, it is easier to isolate and culture primary TM cells than SC cells [42]. A detection of DNMT1 (Gene ID 1786, OMIM 126375)transcripts in TM tissues and primary TM cells showed consistent DNMT expression in vitro [43]. The global DNA methylation was increased in age-matched glaucoma TM cells, accompanied with increased expression of TGFβ1 (Gene ID 7040, OMIM 190180) [44]. The treatment of normal TM cells with TGFβ1 leads to increased expression of DNMT1 (Gene ID 1786, OMIM 126375) and decreased RASAL1 (Gene ID 8437, OMIM 604118) expression, similar to those in glaucoma TM cells [44]. Reduced TGFβ1 promoter methylation and enhanced global DNA methylation were also examined in glaucomatous lamina cribrosa cells [45], further suggesting the association of alteration in DNA methylation and development of glaucoma. However, there was no significant difference between global DNA methylation level of glaucoma and normal SC cells in this study, which may be ascribed to the loss of global methylation in aging samples of both groups [46].
The deep location and limited number of cells in the SC lead to an extremely low successful rate of primary human SC cell culture using postmortem human donor tissues [28]. Moreover, the late onset nature of POAG raises greater difficulty in populating cells derived from older donors with glaucoma. The small sample size and age variation limited the statistical analysis of profiling, so we focused on the methylation of the previously reported glaucoma-related genes. Integrated with these SC expression data [10], TGFBR3 and NET1 showed potential correlation of DMSs with the opposite direction of expression change. The regions around the significant DMSs of TGFBR3 and NET1 were not significantly different. The variations between the DMS and DMR validation were probably a result of probe-based microarray that CpG sites were differentially methylated, but the level of the region was not significantly different. In the whole view, there were 40 DMSs located in the body of TGFBR3, but only four DMSs (10%) with a p value of less than 0.05. The comparison of multiple probes in a consecutive genetic region showed the potential failure in validation of DMRs. However, genes with multiple significant DMSs have several validated DMRs, including PLEKHA7, PITX2, DAXX, and TBX3 (Table 3). These results indicated the complexity of methylation regulation by altering at several locations, but not the whole gene. There was no significant expression change in correspondence with these DMRs, which may be due to different set of samples used for methylation and expression profiling or other expression-regulating mechanisms such as histone modifications. Among these identified genes with DMRs, PLEKHA7 and PITX2 were associated with primary glaucoma [41,47]; while TBX3 was crucial in retina development and DAXX was a key component mediating retinal cell death [48,49]. The different methylation level on these genes may propose the existence of inheritable gene malfunction instead of mutation in patients with glaucoma.
This study was limited by several factors, especially the small sample size. Age variation may have contributed to the differences in methylation. Additionally, due to the limited amount of DNA for methylation profiling, we used Illumina BeadChips instead of DNA methylation sequencing, so we may have missed some DMSs. Thus, we increased the significance of DMSs, targeted on the glaucoma-related genes, and validated DMRs containing DMSs. It will be critical to derive more human primary cells, which was limited by the decreasing availability and high cost of human donor eyes. We believed that this study was able to provide comprehensive understanding of glaucoma methylation in outflow facility. By updating our knowledge of the mechanism of methylation regulation, we can integrate these human data with genetics, expression, and functions.
Acknowledgments
We thank the financial support from The Glaucoma Foundation, The Glaucoma Research Foundation, The BrightFocus Foundation, NIH R01EY023242, R21EY028671, R01EY022359, P30EY031631, and P30EY005722. The funding sources have no influences on the experimental design, data generation and analysis. We sincerely thank all the donors for their ocular samples. This study would not be feasible without these precious samples.
Appendix 1. The list of significant DMSs (|Δβ|≥0.1, p<1X10-5).
To access the data, click or select the words “Appendix 1.”
Appendix 2. The gene ontology of significant DMSs nearest genes.
To access the data, click or select the words “Appendix 2.”
Appendix 3. Significant KEGG pathway of nearest genes.
To access the data, click or select the words “Appendix 3.”
References
- 1.Resnikoff S, Pascolini D, Etya’ale D, Kocur I, Pararajasegaram R, Pokharel GP, Mariotti SP. Global data on visual impairment in the year 2002. Bull World Health Organ. 2004;82:844–51. [PMC free article] [PubMed] [Google Scholar]
- 2.Tham YC, Li X, Wong TY, Quigley HA, Aung T, Cheng CY. Global prevalence of glaucoma and projections of glaucoma burden through 2040: a systematic review and meta-analysis. Ophthalmology. 2014;121:2081–90. doi: 10.1016/j.ophtha.2014.05.013. [DOI] [PubMed] [Google Scholar]
- 3.Liu Y, Allingham RR. Molecular genetics in glaucoma. Exp Eye Res. 2011;93:331–9. doi: 10.1016/j.exer.2011.08.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Jonas JB, Aung T, Bourne RR, Bron AM, Ritch R, Panda-Jonas S. Glaucoma. Lancet. 2017;390:2183–93. doi: 10.1016/S0140-6736(17)31469-1. [DOI] [PubMed] [Google Scholar]
- 5.Roy Chowdhury U, Hann CR, Stamer WD, Fautsch MP. Aqueous humor outflow: dynamics and disease. Invest Ophthalmol Vis Sci. 2015;56:2993–3003. doi: 10.1167/iovs.15-16744. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Stamer WD, Braakman ST, Zhou EH, Ethier CR, Fredberg JJ, Overby DR, Johnson M. Biomechanics of Schlemm’s canal endothelium and intraocular pressure reduction. Prog Retin Eye Res. 2015;44:86–98. doi: 10.1016/j.preteyeres.2014.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Stamer WD, Acott TS. Current understanding of conventional outflow dysfunction in glaucoma. Curr Opin Ophthalmol. 2012;23:135–43. doi: 10.1097/ICU.0b013e32834ff23e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Last JA, Pan T, Ding Y, Reilly CM, Keller K, Acott TS, Fautsch MP, Murphy CJ, Russell P. Elastic modulus determination of normal and glaucomatous human trabecular meshwork. Invest Ophthalmol Vis Sci. 2011;52:2147–52. doi: 10.1167/iovs.10-6342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Overby DR, Zhou EH, Vargas-Pinto R, Pedrigi RM, Fuchshofer R, Braakman ST, Gupta R, Perkumas KM, Sherwood JM, Vahabikashi A, Dang Q, Kim JH, Ethier CR, Stamer WD, Fredberg JJ, Johnson M. Altered mechanobiology of Schlemm’s canal endothelial cells in glaucoma. Proc Natl Acad Sci USA. 2014;111:13876–81. doi: 10.1073/pnas.1410602111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cai J, Perkumas KM, Qin X, Hauser MA, Stamer WD, Liu Y. Expression Profiling of Human Schlemm’s Canal Endothelial Cells From Eyes With and Without Glaucoma. Invest Ophthalmol Vis Sci. 2015;56:6747–53. doi: 10.1167/iovs.15-17720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Tielsch JM, Katz J, Sommer A, Quigley HA, Javitt JC. Family history and risk of primary open angle glaucoma. The Baltimore Eye Survey. Arch Ophthalmol. 1994;112:69–73. doi: 10.1001/archopht.1994.01090130079022. [DOI] [PubMed] [Google Scholar]
- 12.Wolfs RC, Klaver CC, Ramrattan RS, van Duijn CM, Hofman A, de Jong PT. Genetic risk of primary open-angle glaucoma. Population-based familial aggregation study. Arch Ophthalmol. 1998;116:1640–5. doi: 10.1001/archopht.116.12.1640. [DOI] [PubMed] [Google Scholar]
- 13.Stone EM, Fingert JH, Alward WL, Nguyen TD, Polansky JR, Sunden SL, Nishimura D, Clark AF, Nystuen A, Nichols BE, Mackey DA, Ritch R, Kalenak JW, Craven ER, Sheffield VC. Identification of a gene that causes primary open angle glaucoma. Science. 1997;275:668–70. doi: 10.1126/science.275.5300.668. [DOI] [PubMed] [Google Scholar]
- 14.Rezaie T, Child A, Hitchings R, Brice G, Miller L, Coca-Prados M, Heon E, Krupin T, Ritch R, Kreutzer D, Crick RP, Sarfarazi M. Adult-onset primary open-angle glaucoma caused by mutations in optineurin. Science. 2002;295:1077–9. doi: 10.1126/science.1066901. [DOI] [PubMed] [Google Scholar]
- 15.Fingert JH, Miller K, Hedberg-Buenz A, Roos BR, Lewis CJ, Mullins RF, Anderson MG. Transgenic TBK1 mice have features of normal tension glaucoma. Hum Mol Genet. 2017;26:124–32. doi: 10.1093/hmg/ddw372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Fan BJ, Wang DY, Lam DS, Pang CP. Gene mapping for primary open angle glaucoma. Clin Biochem. 2006;39:249–58. doi: 10.1016/j.clinbiochem.2005.11.001. [DOI] [PubMed] [Google Scholar]
- 17.Allingham RR, Liu Y, Rhee DJ. The genetics of primary open-angle glaucoma: a review. Exp Eye Res. 2009;88:837–44. doi: 10.1016/j.exer.2008.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Nakano M, Ikeda Y, Taniguchi T, Yagi T, Fuwa M, Omi N, Tokuda Y, Tanaka M, Yoshii K, Kageyama M, Naruse S, Matsuda A, Mori K, Kinoshita S, Tashiro K. Three susceptible loci associated with primary open-angle glaucoma identified by genome-wide association study in a Japanese population. Proc Natl Acad Sci USA. 2009;106:12838–42. doi: 10.1073/pnas.0906397106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Thorleifsson G, Walters GB, Hewitt AW, Masson G, Helgason A, DeWan A, Sigurdsson A, Jonasdottir A, Gudjonsson SA, Magnusson KP, Stefansson H, Lam DS, Tam PO, Gudmundsdottir GJ, Southgate L, Burdon KP, Gottfredsdottir MS, Aldred MA, Mitchell P, St Clair D, Collier DA, Tang N, Sveinsson O, Macgregor S, Martin NG, Cree AJ, Gibson J, Macleod A, Jacob A, Ennis S, Young TL, Chan JC, Karwatowski WS, Hammond CJ, Thordarson K, Zhang M, Wadelius C, Lotery AJ, Trembath RC, Pang CP, Hoh J, Craig JE, Kong A, Mackey DA, Jonasson F, Thorsteinsdottir U, Stefansson K. Common variants near CAV1 and CAV2 are associated with primary open-angle glaucoma. Nat Genet. 2010;42:906–9. doi: 10.1038/ng.661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wiggs JL, Yaspan BL, Hauser MA, Kang JH, Allingham RR, Olson LM, Abdrabou W, Fan BJ, Wang DY, Brodeur W, Budenz DL, Caprioli J, Crenshaw A, Crooks K, Delbono E, Doheny KF, Friedman DS, Gaasterland D, Gaasterland T, Laurie C, Lee RK, Lichter PR, Loomis S, Liu Y, Medeiros FA, McCarty C, Mirel D, Moroi SE, Musch DC, Realini A, Rozsa FW, Schuman JS, Scott K, Singh K, Stein JD, Trager EH, Vanveldhuisen P, Vollrath D, Wollstein G, Yoneyama S, Zhang K, Weinreb RN, Ernst J, Kellis M, Masuda T, Zack D, Richards JE, Pericak-Vance M, Pasquale LR, Haines JL. Common variants at 9p21 and 8q22 are associated with increased susceptibility to optic nerve degeneration in glaucoma. PLoS Genet. 2012;8:e1002654. doi: 10.1371/journal.pgen.1002654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Gharahkhani P, Burdon KP, Fogarty R, Sharma S, Hewitt AW, Martin S, Law MH, Cremin K, Bailey JNC, Loomis SJ, Pasquale LR, Haines JL, Hauser MA, Viswanathan AC, McGuffin P, Topouzis F, Foster PJ, Graham SL, Casson RJ, Chehade M, White AJ, Zhou T, Souzeau E, Landers J, Fitzgerald JT, Klebe S, Ruddle JB, Goldberg I, Healey PR. Wellcome Trust Case Control Consortium Nc, Mills RA, Wang JJ, Montgomery GW, Martin NG, RadfordSmith G, Whiteman DC, Brown MA, Wiggs JL, Mackey DA, Mitchell P, MacGregor S, Craig JE. Common variants near ABCA1, AFAP1 and GMDS confer risk of primary open-angle glaucoma. Nat Genet. 2014;46:1120–5. doi: 10.1038/ng.3079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Khawaja AP, Cooke Bailey JN, Wareham NJ, Scott RA, Simcoe M, Igo RP, Jr, Song YE, Wojciechowski R, Cheng CY, Khaw PT, Pasquale LR, Haines JL, Foster PJ, Wiggs JL, Hammond CJ, Hysi PG, Eye UKB, Vision C, Consortium N. Genome-wide analyses identify 68 new loci associated with intraocular pressure and improve risk prediction for primary open-angle glaucoma. Nat Genet. 2018;50:778–82. doi: 10.1038/s41588-018-0126-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Sieving PA, Collins FS. Genetic ophthalmology and the era of clinical care. JAMA. 2007;297:733–6. doi: 10.1001/jama.297.7.733. [DOI] [PubMed] [Google Scholar]
- 24.Smith ZD, Meissner A. DNA methylation: roles in mammalian development. Nat Rev Genet. 2013;14:204–20. doi: 10.1038/nrg3354. [DOI] [PubMed] [Google Scholar]
- 25.Jones PA. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet. 2012;13:484–92. doi: 10.1038/nrg3230. [DOI] [PubMed] [Google Scholar]
- 26.Sobrin L, Seddon JM. Nature and nurture- genes and environment- predict onset and progression of macular degeneration. Prog Retin Eye Res. 2014;40:1–15. doi: 10.1016/j.preteyeres.2013.12.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kowluru RA, Kowluru A, Mishra M, Kumar B. Oxidative stress and epigenetic modifications in the pathogenesis of diabetic retinopathy. Prog Retin Eye Res. 2015;48:40–61. doi: 10.1016/j.preteyeres.2015.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Stamer WD, Roberts BC, Howell DN, Epstein DL. Isolation, culture, and characterization of endothelial cells from Schlemm’s canal. Invest Ophthalmol Vis Sci. 1998;39:1804–12. [PubMed] [Google Scholar]
- 29.Assenov Y, Muller F, Lutsik P, Walter J, Lengauer T, Bock C. Comprehensive analysis of DNA methylation data with RnBeads. Nat Methods. 2014;11:1138–40. doi: 10.1038/nmeth.3115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Triche JT. FDb.InfiniumMethylation.hg19: Annotation package for Illumina Infinium DNA methylation probes. 2014.
- 31.Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47. doi: 10.1093/nar/gkv007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Phipson B, Lee S, Majewski IJ, Alexander WS, Smyth GK. Robust Hyperparameter Estimation Protects against Hypervariable Genes and Improves Power to Detect Differential Expression. Ann Appl Stat. 2016;10:946–63. doi: 10.1214/16-AOAS920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Smyth GK, Michaud J, Scott HS. Use of within-array replicate spots for assessing differential expression in microarray experiments. Bioinformatics. 2005;21:2067–75. doi: 10.1093/bioinformatics/bti270. [DOI] [PubMed] [Google Scholar]
- 34.Smyth GK, Speed T. Normalization of cDNA microarray data. Methods. 2003;31:265–73. doi: 10.1016/s1046-2023(03)00155-5. [DOI] [PubMed] [Google Scholar]
- 35.Oshlack A, Emslie D, Corcoran LM, Smyth GK. Normalization of boutique two-color microarrays with a high proportion of differentially expressed probes. Genome Biol. 2007;8:R2. doi: 10.1186/gb-2007-8-1-r2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Li Z, Allingham RR, Nakano M, Jia L, Chen Y, Ikeda Y, Mani B, Chen LJ, Kee C, Garway-Heath DF, Sripriya S, Fuse N, Abu-Amero KK, Huang C, Namburi P, Burdon K, Perera SA, Gharahkhani P, Lin Y, Ueno M, Ozaki M, Mizoguchi T, Krishnadas SR, Osman EA, Lee MC, Chan AS, Tajudin LS, Do T, Goncalves A, Reynier P, Zhang H, Bourne R, Goh D, Broadway D, Husain R, Negi AK, Su DH, Ho CL, Blanco AA, Leung CK, Wong TT, Yakub A, Liu Y, Nongpiur ME, Han JC, Hon DN, Shantha B, Zhao B, Sang J, Zhang N, Sato R, Yoshii K, Panda-Jonas S, Ashley Koch AE, Herndon LW, Moroi SE, Challa P, Foo JN, Bei JX, Zeng YX, Simmons CP, Bich Chau TN, Sharmila PF, Chew M, Lim B, Tam PO, Chua E, Ng XY, Yong VH, Chong YF, Meah WY, Vijayan S, Seongsoo S, Xu W, Teo YY, Cooke Bailey JN, Kang JH, Haines JL, Cheng CY, Saw SM, Tai ES. Consortium IC-G, Consortium N, Richards JE, Ritch R, Gaasterland DE, Pasquale LR, Liu J, Jonas JB, Milea D, George R, Al-Obeidan SA, Mori K, Macgregor S, Hewitt AW, Girkin CA, Zhang M, Sundaresan P, Vijaya L, Mackey DA, Wong TY, Craig JE, Sun X, Kinoshita S, Wiggs JL, Khor CC, Yang Z, Pang CP, Wang N, Hauser MA, Tashiro K, Aung T, Vithana EN. A common variant near TGFBR3 is associated with primary open angle glaucoma. Hum Mol Genet. 2015;24:3880–92. doi: 10.1093/hmg/ddv128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Vranka JA, Kelley MJ, Acott TS, Keller KE. Extracellular matrix in the trabecular meshwork: intraocular pressure regulation and dysregulation in glaucoma. Exp Eye Res. 2015;133:112–25. doi: 10.1016/j.exer.2014.07.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Strungaru MH, Dinu I, Walter MA. Genotype-phenotype correlations in Axenfeld-Rieger malformation and glaucoma patients with FOXC1 and PITX2 mutations. Invest Ophthalmol Vis Sci. 2007;48:228–37. doi: 10.1167/iovs.06-0472. [DOI] [PubMed] [Google Scholar]
- 39.Pena JD, Varela HJ, Ricard CS, Hernandez MR. Enhanced tenascin expression associated with reactive astrocytes in human optic nerve heads with primary open angle glaucoma. Exp Eye Res. 1999;68:29–40. doi: 10.1006/exer.1998.0577. [DOI] [PubMed] [Google Scholar]
- 40.Thomson BR, Heinen S, Jeansson M, Ghosh AK, Fatima A, Sung HK, Onay T, Chen H, Yamaguchi S, Economides AN, Flenniken A, Gale NW, Hong YK, Fawzi A, Liu X, Kume T, Quaggin SE. A lymphatic defect causes ocular hypertension and glaucoma in mice. J Clin Invest. 2014;124:4320–4. doi: 10.1172/JCI77162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Lee MC, Chan AS, Goh SR, Hilmy MH, Nongpiur ME, Hong W, Aung T, Hunziker W, Vithana EN. Expression of the primary angle closure glaucoma (PACG) susceptibility gene PLEKHA7 in endothelial and epithelial cell junctions in the eye. Invest Ophthalmol Vis Sci. 2014;55:3833–41. doi: 10.1167/iovs.14-14145. [DOI] [PubMed] [Google Scholar]
- 42.Grierson I, Marshall J, Robins E. Human trabecular meshwork in primary culture: a morphological and autoradiographic study. Exp Eye Res. 1983;37:349–65. doi: 10.1016/0014-4835(83)90172-0. [DOI] [PubMed] [Google Scholar]
- 43.Bonnin N, Belville C, Chiambaretta F, Sapin V, Blanchon L. DNA methyl transferases are differentially expressed in the human anterior eye segment. Acta Ophthalmol. 2014;92:e366–71. doi: 10.1111/aos.12365. [DOI] [PubMed] [Google Scholar]
- 44.McDonnell F, Irnaten M, Clark AF, O’Brien CJ, Wallace DM. Hypoxia-Induced Changes in DNA Methylation Alter RASAL1 and TGFbeta1 Expression in Human Trabecular Meshwork Cells. PLoS One. 2016;11:e0153354. doi: 10.1371/journal.pone.0153354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.McDonnell FS, McNally SA, Clark AF, O’Brien CJ, Wallace DM. Increased Global DNA Methylation and Decreased TGFbeta1 Promoter Methylation in Glaucomatous Lamina Cribrosa Cells. J Glaucoma. 2016;25:e834–42. doi: 10.1097/IJG.0000000000000453. [DOI] [PubMed] [Google Scholar]
- 46.Fraga MF, Esteller M. Epigenetics and aging: the targets and the marks. Trends Genet. 2007;23:413–8. doi: 10.1016/j.tig.2007.05.008. [DOI] [PubMed] [Google Scholar]
- 47.Reis LM, Tyler RC, Volkmann Kloss BA, Schilter KF, Levin AV, Lowry RB, Zwijnenburg PJ, Stroh E, Broeckel U, Murray JC, Semina EV. PITX2 and FOXC1 spectrum of mutations in ocular syndromes. European journal of human genetics. Eur J Hum Genet. 2012;20:1224–33. doi: 10.1038/ejhg.2012.80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Motahari Z, Martinez-De Luna RI, Viczian AS, Zuber ME. Tbx3 represses bmp4 expression and, with Pax6, is required and sufficient for retina formation. Development. 2016;143:3560–72. doi: 10.1242/dev.130955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Lee YS, Dayma Y, Park MY, Kim KI, Yoo SE, Kim E. Daxx is a key downstream component of receptor interacting protein kinase 3 mediating retinal ischemic cell death. FEBS Lett. 2013;587:266–71. doi: 10.1016/j.febslet.2012.12.004. [DOI] [PubMed] [Google Scholar]