Abstract
Anthracnose (ANT) and angular leaf spot (ALS) caused by Colletotrichum lindemuthianum and Pseudocercospora griseola, respectively, are devastating diseases of common bean around the world. Therefore, breeders are constantly searching for new genes with broad-spectrum resistance against ANT and ALS. This study aimed to characterize the genetic resistance of California Dark Red Kidney (CDRK) to C. lindemuthianum races 73, 2047, and 3481 and P. griseola race 63–39 through inheritance, allelism testing, and molecular analyses. Genetic analysis of response to ANT and ALS in recombinant inbred lines (RILs) from a CDRK × Yolano cross (CY) showed that the resistance of CDRK cultivar is conferred by a single dominant loci, which we named CoPv01CDRK/PhgPv01CDRK. Allelism tests performed with race 3481showed that the resistance gene in CDRK is independent of the Co-1 and Co-AC. We conducted co-segregation analysis in genotypes of 110 CY RILs and phenotypes of the RILs in response to different races of the ANT and ALS pathogens. The results revealed that CoPv01CDRK and PhgPv01CDRK are coinherited, conferring resistance to all races. Genetic mapping of the CY population placed the CoPv01CDRK/PhgPv01CDRK loci in a 245 Kb genomic region at the end of Pv01. By genotyping 19 RILs from the CY population using three additional markers, we fine-mapped the CoPv01CDRK/PhgPv01CDRK loci to a smaller genomic region of 33 Kb. This 33 Kb region harbors five predicted genes based on the common bean reference genome. These results can be applied in breeding programs to develop bean cultivars with ANT and ALS resistance using marker-assisted selection.
Introduction
Phaseolus vulgaris L. (common bean) is the most commonly consumed Phaseolus species worldwide [1, 2, 3], and it is an important primary source of protein in several countries. In particular, common beans are consumed in large quantities in many areas of Africa and Latin America and are part of traditional diets in the Middle East and Europe [4, 5]. However, this legume is susceptible to several diseases threatening its production worldwide [6]. For instance, anthracnose (ANT), caused by Colletotrichum lindemuthianum (Sacc. and Magnus) Briosi and Cavara, and angular leaf spot (ALS), caused by Pseudocercospora griseola (Sacc.) Crous and Braun, are the most widespread, recurrent, and devastating diseases of the common bean in Latin America and Africa [7, 8, 9, 10, 11]. Under favorable climatic conditions for the pathogen and with the use of susceptible cultivars or infected seeds, ANT and ALS can cause field losses of up to 100% and 80%, respectively [7, 12, 13, 14]. Importantly, ANT is not limited to the tropics and is one of the major diseases of beans throughout temperate regions as well [15, 16].
The most desirable strategy to control ANT and ALS is the use of resistant cultivars, which can reduce yield losses without the negative environmental impact of fungicide application [17, 18, 19]. However, the implementation of resistance is challenged by the recurrent emergence of virulence phenotypes in the pathogen population, usually referred to races of C. lindemuthianum and P. griseola. New virulence races of these pathogens have resulted in reduced or complete loss of yield in previously resistant commercial cultivars [17, 20, 21, 22]. Thus, new sources of durable resistance are highly desirable for effective breeding efforts. Gene pyramiding for durable resistance to diseases caused by highly variable pathogens is greatly facilitated by marker-assisted selection, and the development of highly accurate molecular markers that are tightly linked to important disease-resistance genes enables the pyramiding of these genes into single cultivars with broad-spectrum resistance.
Resistance to ANT is conferred by independently segregating individual loci in a series named Co. Currently, the known Co genes are Co-1 and its alleles, Co-14, Co-Pa, Co-x, and Co-won chromosome Pv01 [22, 23, 25, 27, 28, 29, 30, 31]; Co-u and CoPv02 on chromosome Pv02 [18, 22]; Co-13 and Co-17 on chromosome Pv03 [32]; Co-3, Co-32, Co-33, Co-34/Phg-3, Co-y, Co-z, and Co-RVI on chromosome Pv04 [24, 26]; Co-5, Co-6, and Co-v on chromosome Pv07; and Co-2 on chromosome Pv11 [32].
Although resistance to the ALS pathogen is largely conferred by single dominant resistance loci identified by classical genetic approaches, quantitative trait loci (QTLs) have recently been found as well [33, 34, 35, 36]. To date, five resistance loci (the Phg series) have been mapped to the integrated bean linkage map; the three independent loci Phg-1, Phg-2, and Phg-3 are located on chromosomes Pv01, Pv08, and Pv04, respectively [25, 26, 37], while the two major QTLs Phg-4 and Phg-5 are located on Pv04 and Pv10 [33, 34, 36, 38, 39, 40].
Importantly, these Co and Phg loci may be part of disease-resistance clusters on various chromosomes [23, 24, 25, 28, 32, 41]. For instance, linkage group Pv01 contains a gene cluster having ANT (Co-1, Co-AC, Co-14, Co-x, and Co-w), rust (Ur-9), and ALS (Phg-1) resistance genes [18, 25, 26, 28, 32, 29]. The major resistance genes for bean rust, ANT, common bacterial blight, and white mold were mapped in clusters on chromosomes Pv01, Pv04, Pv07, and Pv11 [42, 43]. In addition, a set of resistance gene analogs (RGAs) that are linked to resistance loci for different common bean pathogens were identified [44]. These observations highlight the opportunity for the simultaneous selection of resistance to multiple diseases, especially diseases that occur in similar geographic regions, such as ANT and ALS. The Andean common bean cultivar California Dark Red Kidney (CDRK) is resistant to Mesoamerican races of ANT reported in Brazil, Argentina, and Colombia; all of the Central American races; and races present in Africa and Europe [45, 46, 47, 48, 49, 50, 51].
The availability of reference genomes for the Andean G19833 [41] and Mesoamerican BAT93 [52] has enabled the fine-mapping of many resistance loci, including Co-AC, Co-1HY, Co-x, and Co-173-X on Pv01 [28, 29, 31, 53] and Co-42 on Pv08 [54]. In this study, we verified the resistance inheritance in CDRK. We also found that CDRK is an excellent source to test the hypothesis that ANT and ALS resistance are colocalized in the bean genome and to fine-map the resistance loci using the recombinant inbred line (RIL) mapping population California Dark Red Kidney × Yolano (CY), molecular markers and reference genome.
Material and methods
Plant material and growth conditions
To determine the genetic basis of disease resistance in the genotype CDRK, we used 110 RILs derived from the CY population described by [55]. Seeds were sown in plastic trays (50 × 30 × 9 cm) containing a mixture of the commercial substrate MecPlant (Register EP PB 09549-4/Mapa Brazil, MEC PREC—Ind. Com Ltda, Telemaco Borba, PR), which had been previously fertilized and sterilized. Seedlings were grown under natural light in greenhouses supplemented with 400 W high-pressure sodium lamps, providing a total light intensity of 115 μmoles m-2 s-1 for 15 days until the seedlings reached the first fully expanded trifoliate leaf stage V3 [56]. Plants were inoculated at this stage.
Additionally, cultivars including Michigan Dark Red Kidney (MDRK), Kaboon, Perry Marrow, AND 277, Widusa, Jalo Vermelho, Jalo Listras Pretas, Pitanga, Corinthiano, Paloma, Amendoim Cavalo, Jalo Pintado 2 and Jalo EEP 558 were evaluated for ANT resistance as well as ALS resistance to race 63–39. The experiments comply with the current laws of the country in which they were performed.
Pathogenesis assays
To determine the spectrum of resistance in the above bean genotypes, seedlings were inoculated with the Mesoamerican races 9, 64, 65, 73, 89, 445, 453, 1545, 2047, and 3481, as well as the Andean races 2, 7, 19, 23, 39, and 55 of C. lindemuthianum. Furthermore, seedlings of the CY RIL population were inoculated with races 73, 2047, and 3481 of C. lindemuthianum and race 63–39 of P. griseola to determine segregation patterns of the disease reaction in the CY RIL population.
Monosporic cultures of C. lindemuthianum and of P. griseola were prepared according to the methodologies proposed by Mathur et al. [57] and Sanglard et al. [58]. Inoculum of the ANT races were produced on green common bean pod medium [60] incubated at 22°C for 14 days. The inoculum of race 63–39 of ALS was multiplied in Petri dishes containing 2 mL tomato medium [58] and maintained in a BOD incubator at 24°C for 18 days.
Soon after the expansion of the first trifoliolate leaf, the leaflet of 10 seedlings of each line were inoculated with each race of C. lindemuthianum and the leaflet of 10 seedlings with P. griseola. Each pathogen was inoculated separately. A spore suspension containing 2.0 × 106 spores mL-1 of races 73, 2047 and 3481 of C. lindemuthianum were inoculated using a DeVilbiss number 15 atomizer powered by an electric air compressor (Schulz, SA, Joinville, Santa Catarina, Brazil). A similar procedure was employed for the inoculation with race 63–39 of P. griseola using a spore suspension of 1.2 × 104 conidia mL-1. Ten plants for resistant and susceptible control for each race were inoculated [59, 60].
After inoculation, the plants were maintained at >95% relative humidity, 20 ± 2°C, and 12 h of daily light (680 lux) in a mist chamber for three days. The plants were then transferred to open benches under the same conditions, except for the high humidity, for 7 days (ANT) and 17 days (ALS). ANT and ALS symptoms were evaluated using the disease severity scales (1 to 9) proposed by Pastor-Corrales et al. [61] and Inglis et al. [62]. Plants with disease reaction scores from 1 and 3 were considered resistant, whereas plants with scores from 4 to 9 were considered susceptible [61].
Inheritance of resistance
Studies of the inheritance of resistance in the CDRK genotype were conducted using 110 RILs derived from the CY population described by [55]. The parental line CDRK is resistant to races 73, 2047, and 3481 of C. lindemuthianum and race 63–39 of P. griseola, while the parental line Yolano is susceptible to all races.
Allelism tests
To determine the independence of the ANT resistance allele present in California Dark Red Kidney from Co-1 and Co-AC previously characterized ANT resistance alleles, CDRK was crossed with Andean bean cultivars in the following list: MDRK (Co-1) and Amendoim Cavalo (Co-AC). In all cases, CDRK was used as the female parent. The F1 seeds were sown in polyethylene vases (48 × 30 × 11 cm) containing a mixture of the commercial substrate Plantmax®, which had been previously fertilized and sterilized. The plant vases were kept in a greenhouse until the F2 seeds were produced. The F2 individuals, obtained by selfing individual F1 plants, were used to characterize the plants for resistance to race 3481 of C. lindemuthianum. Seedlings were grown until pod maturation under natural light in greenhouses. The seedlings were maintained in a greenhouse until the first trifoliolate leaves [56] were fully expanded. Race 3481 of C. lindemuthianum was chosen to conduct allelism tests because all parental cultivars inoculated with these races yielded the R × R reaction type.
Statistical analysis
A goodness-of-fit test for the 1:1 segregation ratio was performed for races 73, 2047, 3481, and 63–39 in the CY RIL population. For allelism tests, segregation analysis of two F2 populations from the crosses of CDRK with Andean cultivars (MDRK and Amendoim Cavalo) was also performed using the χ2 test according to the Mendelian segregation null hypothesis of 15: 1 R/S ratio.
SNP genotyping
Total genomic DNA was isolated from the 110 RIL families (F10 generation) and parents (CDRK and Yolano) using the DNeasy Plant Mini Kit (Qiagen, CA, USA) following the manufacturer’s instructions. The DNA was quantified using 1.5% agarose gel (Agarose SFR, Amresco, IL, USA) with TBE buffer (tris-borate-ethylenediamine tetra acetic acid) and stained with 1 μg mL−1 ethidium bromide. The DNA samples were screened with 5,398 SNP DNA markers on the BARCBean6K_3 Illumina BeadChip [63] by following the Infinium HD Assay Ultra Protocol (Illumina, Inc., San Diego, CA, USA). The BeadChip was imaged using the Illumina BeadArray Reader to measure fluorescence intensity. Automatic allele calling for each locus was performed with the Genome Studio Genotyping Module v1.8.4 software (Illumina, San Diego, CA, USA), and all allele calls were visually inspected. Any errors in allele calling due to improper cluster identification were corrected, resulting in 4,633 high-quality SNPs.
Genome-wide linkage map analysis
SNP markers that were polymorphic between the parents CDRK and Yolano segregated at a 1:1 ratio in the RIL population, as measured by the χ2 test at p = 0.01, were used to create a linkage map using the default settings of the JoinMap 4.1 software [64]. Briefly, the regression-mapping algorithm based on the Kosambi map function was used to define the linkage order and genetic distances in centiMorgans (cM). A minimum likelihood of odds (LOD) ≥3.0 and a maximum distance of ≤50 cM were used to test linkages among markers. A genetic linkage map was created using the software MapChart [65]. SNP markers flanking the genomic locations associated with ANT and ALS disease reactions were used to define the physical region of these loci based on the bean reference genome v.1.0 [41] available in NCBI v.1.0 (http://phytozome.jgi.doe.gov).
Fine-mapping
A fine linkage map was developed with 17 SNPs, two additional SSRs (BARCPVSSR01358, BARCPVSSR01361) and the STS CV542014 markers (http://phaseolusgenes.bioinformatics.ucdavis.edu/markers/1009). The selected SSR and STS markers were amplified from the genomic DNA of the parents CDRK (resistant) and Yolano (susceptible) and 19 of the CY RILs to fine-map a genomic region of 245 Kb between the ss715645251 (50,301,592) and ss715645248 (50,546,985) markers in chromosome Pv01. The primer sequences used to genotype BARCPVSSR01358, BARCPVSSR01361 and STS CV542014 were 'TGGCTGGTTGGTGTTTATGA' (forward) and 'GGTCCCACCCTCTTCTCTTC' (reverse), ´GAATGGTTCATCGTTCATGG´ (forward), and ´TCGGCTGTTTAACGTGGTCT´ (reverse), and `CACTTTCCACTGACGGATTTGAACC`(forward) and `CAGAGGATGCTTCTCACGGT`(reverse), respectively. The PCR mixes contained 30 ng of genomic DNA, 0.25 μM of forward and reverse primers, 1 X PCR Buffer (200 mM Tris-HCl (pH 8.0), 500 mM KCl, 2 mM each dNTP, 10% glycerol, 15 mM MgCl2, and 20 ng/μL of single-strand binding protein (SSB)) and 0.1 unit of Taq DNA polymerase (Invitrogen). The PCR cycle consisted of 3 min at 92°C; followed by 38 cycles of 50 s at 90°C, 45 s at 58°C, and 45 s at 72°C; a 5 min extension at 72°C; and a hold at 10°C. A 2 μL aliquot of loading buffer (30% glycerol and 0.25% bromophenol blue) was added to the DNA products, which were then fractionated on 6% polyacrylamide gels at 3 W A−1 cm−1. The amplified fragments were stained using SYBR Safe (0.02%), and the DNA bands were visualized under ultraviolet light. Digital images were recorded using an L-PIX Image EX (Loccus Biotecnologia-Loccus do Brasil, Cotia, SP, Brazil).
Functional annotation of genes linked to ANT and ALS disease reactions
The P. vulgaris reference genome v.1.0 [41] was used to define the physical position of the markers flanking the CoPv01CDRK/PhgPv01CDRK resistance loci. The putative genes within this genomic region were annotated as candidate genes associated with resistance or susceptibility to ANT and ALS. The putative functional annotation of each candidate gene was based on the descriptions available in Phytozome v.1.0 (https://phytozome.jgi.doe.gov#). After fine-mapping, the new physical region was defined based on the genomic location of the markers flanking the new CDRK resistance loci. Genes predicted within the fine-mapped region were highlighted, and their putative homologs in Arabidopsis thaliana were identified using BLASTp in NCBI (National Center for Biotechnology Information; https://www.ncbi.nlm.nih.gov). The A. thaliana protein with the lowest E-value (<0.0) and highest identity (>40%) with each bean protein was considered a putative homolog and used to infer its molecular function.
Results
To identify sources of resistance against the ANT pathogen, a panel of 14 Andean cultivars were screened for their reactions to ten Mesoamerican (9, 64, 65, 73, 89, 445, 453, 1545, 2047, and 3481) and six Andean (2, 7, 19, 23, 39, and 55) races of C. lindemuthianum (Table 1). The genomic locations of the known Co genes in each cultivar, except for Jalo Vermelho, Jalo Pintado 2, and CDRK (Table 1), have been previously reported [25, 28, 29, 30, 31, 32, 53]. We observed that five genotypes known to carry alleles at the Co-1 locus (MDRK, Kaboon, Perry Marrow, AND 277, and Widusa) showed different spectra of ANT resistance (Table 1), supporting the hypothesis that they indeed carry different alleles of Co-1 or additional unidentified Co loci. All cultivars except Pitanga were resistant to race 1545 of C. lindemuthianum. By contrast, Pitanga and six other genotypes were resistant to race 2047. Kaboon (Co-12), AND 277 (Co-14), Jalo Vermelho (Co-12), Pitanga (Co-14), Corithiano (Co-15), and CDRK were the only cultivars that showed resistance and susceptibility to C. lindemuthianum races 2 and 7, respectively (Table 1). As CDRK showed resistance to highly virulent races of C. lindemuthianum (Table 1) and it is a parent of a well-described RIL population named CY [55], we further characterized the Co loci that it might carry. We observed that CDRK exhibited resistance to races 2, 9, 39, 55, 64, 65, 73, 89, 1545, 2047, and 3481 of C. lindemuthianum, as well as race 63–39 of P. griseola, while Yolano was susceptible to all races (Table 1). Thus, the CY RIL population was used for co-segregation analysis.
Table 1. Resistant (R) or Susceptible (S) reactions of 15 Phaseolus vulgaris cultivars to nine Mesoamerican (9, 65, 73, 89, 445, 453, 1545, 2047, and 3481) and six Andean (2, 7, 19, 23, 39, and 55) races of C. lindemuthianum and race 63–39 of P. griseola.
| Cultivar | Genes | P. griseola 63–39 | Races of Colletotrichum lindemuthianuma | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 2 | 7 | 9 | 19 | 23 | 39 | 55 | 64 | 65 | 73 | 89 | 449 | 453 | 1545 | 2047 | 3481 | |||
| MDRKb | Co-1 | dNe | S | S | R | S | S | S | S | R | R | R | R | R | R | R | S | R |
| Kaboon | Co-12 | dNe | R | R | R | R | R | S | S | R | R | R | R | R | R | R | S | R |
| Perry Marrow | Co-13 | dNe | R | S | R | R | S | S | S | R | R | R | R | R | S | R | S | R |
| AND 277 | Co-14 | S | R | S | R | S | R | S | R | R | R | R | R | R | R | R | R | R |
| Widusa | Co-15 | dNe | R | R | R | S | S | R | S | R | R | R | S | R | R | R | S | S |
| Jalo Vermelho | Co-12 | R | R | S | R | S | R | S | R | R | R | S | R | R | R | R | S | S |
| Jalo Listras Pretas | Co-13 | R | S | S | R | S | S | S | S | R | R | R | R | S | S | R | S | R |
| Pitanga | Co-14 | dNe | R | S | R | R | R | S | R | R | R | R | S | S | S | S | R | S |
| Corinthiano | Co-15 | R | R | S | S | S | R | dNe | S | R | R | R | R | R | S | R | R | S |
| Paloma | CoPv01PA | dNe | S | S | S | S | R | R | R | R | R | R | S | S | S | R | R | R |
| Amendoim Cavalo | CoPv01AC | R | R | R | R | R | R | R | R | R | R | R | R | S | S | R | R | R |
| Jalo Pintado 2 | CoPv04JP2 | R | R | R | R | S | S | S | S | R | R | R | S | R | R | R | R | R |
| Jalo EEP558 | Co-w, Co-x | R | S | S | dNe | S | dNe | dNe | S | R | R | R | S | R | R | R | S | R |
| Co-y, Co-z | ||||||||||||||||||
| CDRKc | CoPv01CDRK | R | R | S | R | S | S | R | R | R | R | R | R | S | dNe | R | R | R |
| Yolano e(MA) | S | S | S | S | R | S | S | S | S | S | S | S | S | dNe | S | S | S | |
aRaces = Ten Mesoamerican (9, 64, 65, 73, 89, 445, 453, 1545, 2047 and 3481) and six Andean (2, 7, 19, 23, 39, and 55) races of Colletotrichum lindemuthianum;
bMDRK = Michigan Dark Red Kidney;
cCDRK = California Dark Red Kidney;
dNe = Not evaluated;
eMA = Mesoamerican.
ANT and ALS co-segregation analyses in the CY RIL population
To determine the inheritance of resistance to races 73, 2047, and 3481 of C. lindemuthianum and 63–39 of P. griseola in CDRK, we inoculated 110 RILs (ten plants from each RIL) with each of these races and scored their disease symptoms (S1 Table). As expected for a single, dominant resistance locus, we observed a segregation ratio of 1RR:1rr in the RIL population, namely, 54 RILs were resistant (RR) and 56 RILs were susceptible (rr) (χ2 = 0.036, p-value = 0.849; Table 2). Interestingly, each RIL showed identical phenotypes in response to each race, indicating that resistances to these ANT and ALS races co-segregate in this population and that the sources of resistance in the CDRK genetic background are either tightly linked. Thus, we named this loci CoPv01CDRK/PhgPv01CDRK.
Table 2. Segregation for resistance to races 73, 2047, 3481 of Colletotrichum lindemuthianum and 63–39 of Pseudocercospora griseola in common bean F10 RIL population from the California Dark Red Kidney × Yolano cross.
| Parental cross | Generation | Observed Ratio (1R:1S)a | Expected Ratio (1R:1S) | χ2 | P value (1 df) |
|---|---|---|---|---|---|
| Race 73, 2047, and 3481 of Colletotrichum lindemuthianum | |||||
| CDRKb | RPc | 30:0 | |||
| Yolano | SPd | 0:30 | |||
| CDRK × Yolano | F10 | 54:56 | 55:55 | 0.036 | 0.849 |
| Race 63–39 of Pseudocercospora griseola | |||||
| CDRK | RPb | 30:0 | |||
| Yolano | SPc | 0:30 | |||
| CDRK × Yolano | F10 | 54:56 | 55:55 | 0.036 | 0.849 |
aR = Resistant; S = Susceptible;
bCDRK = California Dark Red Kidney;
cRP = Resistant Parent;
dSP = Susceptible Parent.
Allelism tests
The results of studies of the allelic relationship between the anthracnose resistance gene in the Andean common bean cultivar CDRK crossed with MDRK and Amendoim Cavalo revealed the absence of allelism (S4 Table). In the allelism test using the F2population from the cross CDRK × Amendoim Cavalo (χ2 = 0.026; p-value = 0.87) inoculated with race 3481, a segregation ratio of 15R:1S was obtained, indicating the presence of two independent dominant genes; one gene is Co-AC [66, 32], present in the cultivar Amendoim Cavalo, and the other gene originated in CDRK. The same ratio of 15R:1S was obtained using race 3481 in the cross CDRK × MDRK (χ2 = 0.022; p-value = 0.88), indicating the action of two dominant genes. In this case, the gene present in CDRK was shown to be independent of the gene Co-1 [67] present in MDRK.
Fine-mapping of the CoPv01CDRK/PhgPv01CDRK loci
Genetic linkage analysis between the CoPv01CDRK/PhgPv01CDRK loci and SNPs showing the expected segregation of 1:1 in the RIL population revealed that CoPv01CDRK/PhgPv01CDRK is flanked by the SNP markers ss715645251 and ss715645248 in a genomic region on chromosome Pv01 (Fig 1). The physical locations of the markers ss715645251 and ss715645248 are 50,301,532 bp and 50,546,985 bp, respectively, which correspond to a distance of 245.6 Kb based on the bean reference genome v1.0 (https://www.ncbi.nlm.nih.gov).
Fig 1. Genetic map of common bean linkage group Pv01 containing the anthracnose and angular leaf spot resistance loci and linked Single Nucleotide Polymorphism (SNPs) markers used to genotype the F10 population California Dark Red Kidney × Yolano.
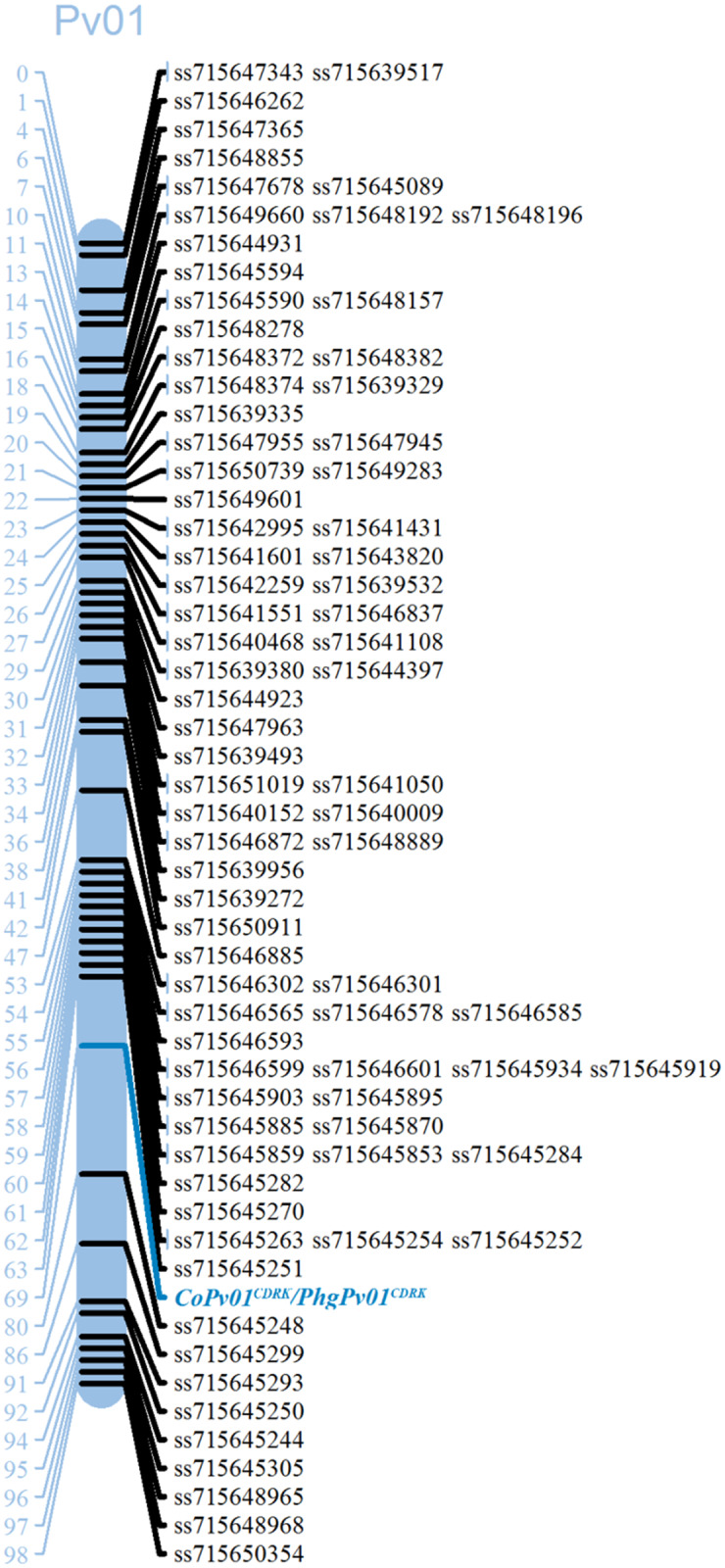

Recombination distances are indicated on the left side of the linkage group in centiMorgans (cM), and the marker names are shown on the right side. The CoPv01CDRK/PhgPv01CDRK resistance loci were flanked by SNP markers ss715645251 and ss715645248 in F10 mapping population. The map was drawn with MapChart (65).
To narrow the genomic region harboring the CoPv01CDRK/PhgPv01CDRK loci, we performed a fine-mapping analysis by genotyping 19 RILs that showed recombination events in the 245.6 Kb region identified. Recombination events were identified based on the genotypic data of all 110 RILs obtained with the BARCBEAN6K_3 BeadChip. Upon genotyping these 19 RILs with 12 SNPs, two SSRs, and one STS marker, we observed that the susceptible CY5 RIL and the resistant CY48 RIL contained recombination events (Table 3) that allowed us to delimit the CoPv01CDRK/PhgPv01CDRK region to the area between the CV542014 and ss715645248 markers. Based on the bean reference genome [41], these new CoPv01CDRK/PhgPv01CDRK flanking markers are located at positions 50,513,853 bp (CV542014) and 50,546,985 bp (ss715645248) of chromosome Pv01, spanning 33 Kb (Fig 2).
Table 3. Genotype and phenotype of 19 F10 recombinant events in the region of Pv01 used for fine mapping of the anthracnose and angular leaf spot resistance loci in CDRK.
The phenotype was obtained from the reaction of the 110 F10 RILs to races 73, 2047, and 3481 of Colletotrichum lindemuthianum and race 63–39 of P. griseola. Genotyping was achieved using the flanking markers 12 SNP, two SSR and one STS markers that enabled the positioning of the CoPv01CDRK/PhgPv01CDRK loci in a 33 kb genomic region flanked by markers CV542014 and ss715645248.
| Marker | SNP position | Recombinant lines from CDRK × Yolano | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 5 | 12 | 19 | 20 | 33 | 38 | 43 | 47 | 48 | 62 | 70 | 73 | 79 | 87 | 88 | 91 | 115 | 96 | 146 | ||
| ss715645260 | 50115685 | AA | AA | BB | BB | AA | BB | AA | AA | AA | BB | AA | AA | AA | BB | BB | AA | BB | AB | AB |
| ss715645259 | 50130201 | AA | AA | BB | BB | AA | BB | AA | AA | AA | BB | AA | AA | AA | BB | BB | AA | BB | AB | AB |
| ss715645258 | 50155987 | AA | AA | BB | BB | AA | BB | AA | AA | AA | BB | AA | AA | AA | BB | BB | AA | BB | AB | AB |
| ss715645257 | 50161526 | AA | AA | BB | BB | AA | BB | AA | AA | AA | BB | AA | AA | AA | BB | BB | AA | BB | AB | AB |
| ss715645256 | 50182775 | AA | AA | BB | BB | AA | BB | AA | AA | AA | BB | AA | AA | AA | BB | BB | AA | BB | AB | AB |
| ss715645254 | 50203547 | AA | AA | BB | BB | AA | BB | AA | AA | AA | BB | AA | AA | AA | BB | BB | AA | BB | AB | AB |
| ss715645252 | 50222584 | AA | AA | BB | BB | AA | BB | AA | AA | AA | BB | AA | AA | AA | BB | BB | AA | BB | AB | AB |
| ss715645251 | 50301592 | AA | AA | BB | BB | AA | BB | AA | AA | AA | BB | AA | AA | BB | BB | BB | AA | BB | AB | AB |
| BARCPVSSR01358 | 50350345 | AA | AA | BB | BB | AA | BB | - | AA | AA | BB | AA | AA | BB | BB | BB | AA | BB | BB | AA |
| BARCPVSSR01361 | 50388017 | AA | AA | BB | BB | AA | BB | - | AA | AA | BB | AA | AA | BB | BB | BB | AA | BB | BB | AA |
| CV542014 | 50513853 | AA | AA | BB | BB | AA | BB | AA | AA | AA | BB | AA | AA | BB | BB | BB | AA | BB | AB | AB |
| CoPv01CDRK/PhgPv01CDRK | BB | AA | BB | BB | AA | BB | AA | AA | AA | BB | AA | AA | BB | BB | BB | AA | BB | BB | AA | |
| ss715645248 | 50546985 | BB | AA | BB | BB | AA | BB | AA | AA | BB | BB | AA | AA | BB | BB | BB | AA | BB | AB | AB |
| ss715645299 | 51353193 | BB | BB | AA | BB | AA | AA | AA | BB | BB | BB | BB | BB | BB | BB | BB | BB | AA | AB | AA |
| ss715645293 | 51617802 | BB | BB | AA | AA | BB | AA | BB | BB | BB | AA | AA | BB | BB | AA | AA | BB | AA | AA | AA |
| ss715645250 | 51726047 | BB | BB | AA | AA | BB | AA | BB | BB | BB | AA | AA | BB | BB | AA | AA | BB | AA | BB | AA |
| ss715645244 | 51764167 | AA | BB | AA | AA | BB | AA | BB | BB | BB | AA | AA | BB | BB | AA | AA | BB | AA | BB | AA |
| ss715645305 | 51786948 | AA | BB | AA | AA | BB | AA | BB | BB | BB | AA | AA | BB | BB | AA | AA | BB | AA | BB | AA |
| ss715645301 | 51819821 | AA | BB | AA | AA | BB | AA | BB | BB | BB | AA | AA | BB | BB | AA | AA | BB | AA | BB | AA |
| ss715648967 | 51883712 | AA | BB | AA | AA | BB | AA | BB | BB | BB | AA | AA | BB | BB | AA | AA | BB | AA | BB | AA |
| ss715648965 | 51896315 | AA | BB | AA | AA | BB | AA | BB | BB | BB | AA | AA | BB | BB | AA | AA | BB | AA | BB | AA |
AA = Resistant; BB = Susceptible; AB = Heterozygous;— = not available.
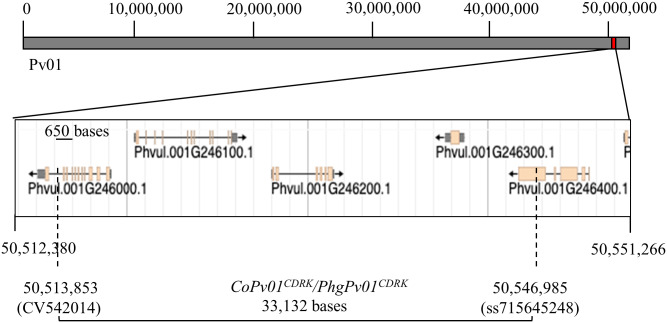
Fig 2. Fine mapped region for the CDRK resistance lociCoPv01CDRK/PhgPv01CDRK.
Upper bar represents the entire chromosome Pv01, in which the CoPv01CDRK/PhgPv01CDRK region is highlighted as red square at the end of the chromosome. The five predicted genes within this region are shown, where the location of the CoPv01CDRK/PhgPv01CDRK flanking markers CV542014 and ss715645248 are indicated by dashed lines, within the predicted genes Phvul.001G246000 and Phvul.001G246400, respectively. The genomic region between these markers is indicated by the lower bar and cover around 33 Kbp of the genome.
Predicted genes and functions associated with the CoPv01CDRK/PhgPv01CDRK loci
Based on the bean reference genome, the 33 Kb region contains five predicted genes (Table 4). The predicted genes encode the proteins Phvul_001G246000 (ATP-dependent RNA Helicase), Phvul.001G246100 (Cation-dependent mannose-6-phosphate receptor), Phvul.001G246200 (Protein Trichome Birefringence-like 33), Phvul.001G246300 (Abscisic Acid (ABA) receptor PYL5), and Phvul.001G246400 (SNF2 domain-containing protein classy 1-related). The putative orthologs for these genes were identified in the Arabidopsis genome (TAIR) for further functional reference (Table 4).
Table 4. Gene models found in delimitated region CDRK resistance loci against anthracnose and angular leaf spot were fine-mapped, gene position in reference genome v1.0 and annotation.
| Gene Model in P. vulgaris | Homolog in A. thaliana | E-valuea | Identitya | Functional annotation on TAIRb | Functional annotation on Phytozomec |
|---|---|---|---|---|---|
| Phvul.001G245300 | AT4G18640 | 8 × 10−169 | 43.40% | Male Discoverer 2 | Protein tyrosine kinase (pkinase_tyr) // leucine rich repeat n-terminal domain (lrrnt_2) |
| Phvul.001G246000 | AT5G05450 | 0 | 70.50% | RNA Helicase 18 | ATP-dependent RNA helicase ddx55/spb4 [ec:3.6.4.13] (ddx55, spb4) |
| Phvul.001G246100 | AT2G37390 | 1 × 10−105 | 52.60% | Sodium Potassium Root Defective 2 | Cation-dependent mannose-6-phosphate receptor |
| Phvul.001G246200 | AT2G40320 | 0 | 69.90% | Trichome Birefringence-Like 33 | Protein trichome birefringence-like 33 |
| Phvul.001G246300 | AT2G40330 | 1 × 10−76 | 56.90% | ABA Receptor PYL6 | Abscisic acid receptor pyl5 |
| Phvul.001G246400 | AT5G20400 | 0 | 48.30% | Flavanone 3 Hydroxylase-like | SNF2 domain-containing protein classy 1-related |
| Phvul.001G246800 | AT3G50740 | 0 | 60.60% | UDP-Glucosyl Transferase 72E1 | Leucine-rich repeat receptor-like protein kinase imk3-related |
aE-values and Identity for BLASTp analysis performed on NCBI (National Center for Biotechnology Information; https://www.ncbi.nlm.nih.gov)
b Functional gene annotation resource: TAIR—The Arabidopsis Information Resource (https://www.arabidopsis.org)
cFunctional gene annotation resource: Phytozome—Common bean reference genome v1.0 (https://phytozome.jgi.doe.gov#
Interestingly, the marker CV542014 is physically mapped at 946 bases upstream the stop codon of the Phvul.001G246000, at the last predicted intron (Fig 2). Furthermore, the marker ss715645248 is located at 1,283 bases from the stop codon of the predicted gene Phvul.001G246400 (Fig 2), resulting in a mutation in the last exon of this gene, which encodes for a putative DEXH-box Helicase Domain (DEXHc_ATRX-like; Conserved Domain cd18007, e-value = 9.8 × 10−73).
Additionally, the predicted gene Phvul.001G245300, which encodes a putative Leucine-Rich Repeat Protein Kinase (LRR-Kinase), was detected approximately 66 Kb upstream of the marker locus CV542014 (Table 4). An additional putative LRR-kinase (Phvul.001G246800) was found 26 Kb downstream of the marker ss715645248 (Table 4). These five genes are interesting candidates for CoPv01CDRK/PhgPv01CDRK and may confer resistance to ANT and ALS pathogens. The Fig 3 shows the ANT resistance cluster present at the end of chromosome Pv01.
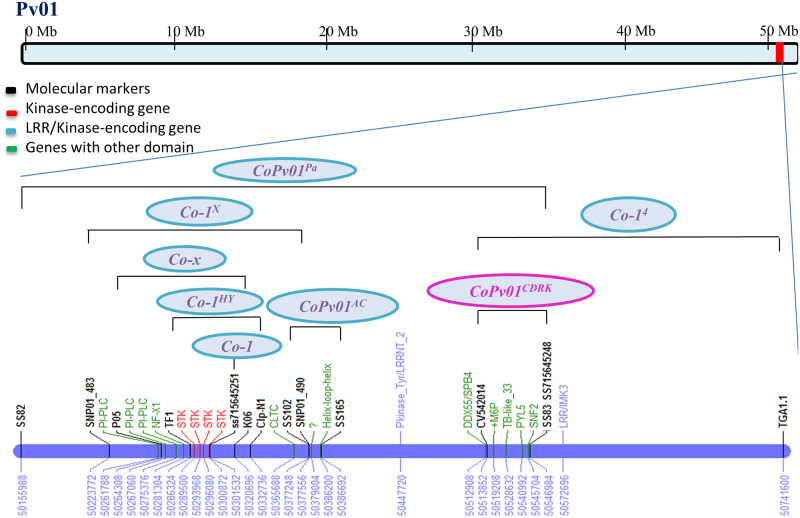
Fig 3. Anthracnose cluster 1.1 on Pv01 with specific candidate genes within and close to the genomic region where these resistance genes were mapped.
Molecular markers linked to the resistance genes are displayed in black color. In red candidate genes that encode kinases, in blue candidate genes encoding NB-LRR and in green candidate genes with other domains. The resistance gene in CDRK cultivar was fine mapped in a region having five candidate genes and close to LRR. Co-1HY, Co-x and Co-1X harbor the same kinases. CoPv01AC, Co-14 and CoPv01PA are the resistance genes of the Amendoim Cavalo, AND 277 and Paloma cultivars, respectively.
Discussion
The spectra of resistance observed for CDRK were different from those of the four Andean cultivars that have ANT resistance alleles at the Co-1 locus. CDRK is resistant to race 2 while MDRK is susceptible. In relation to races 39, 55 and 2047 CDRK was resistant and MDRK, Kaboon and Perry Marrow were susceptible. Additionally, AND 277 showed compatible reaction to race 39. The resistance of CDRK to races 2, 9, 64, 65, and 73 of C. lindemuthianum and race 63–39 of P. griseola is important for common bean breeding programs in Brazil, where these races have high rates of occurrence [45, 46, 47, 49].
The results of inoculating the 110 CY RILs with races 73, 2047, and 3481 of C. lindemuthianum and 63–39 of P. griseola showed that 54 RILs were resistant, while 56 were susceptible. These results fit a segregation ratio of 1R:1S, revealing a monogenic inheritance. It is important to note that all RILs resistant to race 73 were also resistant to races 2047, 3481, and 63–39, while all RILs susceptible to race 73 were also susceptible to races 2047, 3481, and 63–39.
The present study established that the resistance of CDRK to races 73, 2047, and 3481 of C. lindemuthianum and race 63–39 of P. griseola is conferred by a dominant gene. The dominant nature of resistance in the CDRK cultivar suggests that resistance is transferable to commercial cultivars.
In addition, allelism tests conducted in two F2 populations from crosses R × R between the CDRK × MDRK, and CDRK × Amendoim Cavalo inoculated with race 3481 revealed the presence of two dominant genes in each population that conferred resistance to anthracnose. One of these genes (Co-1) is present in MDRK [67], and the other gene is present in CDRK. Notably, the segregation results for the F2 population from the cross CDRK × Amendoim Cavalo highlighted the presence of a dominant gene in CDRK that is independent of the Co-AC genes present in Amendoim Cavalo. The monogenic nature of Amendoim Cavalo resistance to C. lindemuthianum race 3481 was reported by [31]. These results support the hypothesis that the gene conferring resistance to race 3481 of this fungal pathogen, present in CDRK, is independent from other genes (Co-1 and Co-AC genes), harbored in Michigan Dark Red Kidney and Amendoim Cavalo, respectively.
In this study, we elucidated the inheritance of anthracnose resistance in the Andean bean cultivar CDRK and established the genetic relationship between this resistance locus and two known ANT resistance genes mapped to Cluster 1.1 at the end of Pv01. Additional segregation analysis in populations derived from crossing CDRK with two Andean bean cultivars, each carrying different ANT resistance genes, revealed that the ANT resistance of CDRK is conferred by a new gene that is distinct from the previously reported resistance genes Co-1 and Co-AC in common bean. Based on this evidence, the authors propose that the symbol for anthracnose resistance loci in the CDRK cultivar be named CoPv01CDRK/PhgPv01CDRK.
The co-segregation analysis of the ANT and ALS resistance genes was observed in 110 CY RILs originating from a CDRK × Yolano cross, which showed identical phenotypes in response to both diseases. A total of 54 RILs that were resistant to the ANT pathogen were also resistant to all races of the ALS pathogen; 56 RILs that were susceptible (S) to the ANT pathogen were also susceptible to the ALS pathogen. These results suggest that the CoPv01CDRK and PhgPv01CDRK loci are closely linked.
ANT and ALS are two of the most widespread and severe diseases of common bean in the Americas and Africa, which are considered the two largest producers and consumers of this crop [8, 11]. Moreover, these genes protect common bean against two pathogens possessing high and possibly rapidly changing virulence diversity. Thus, the availability of accurate molecular markers to transfer these genes into commercial common bean cultivars would probably increase resistance durability in these cultivars against highly variable pathogens. In the present study, we conducted a co-segregation analysis of the CoPv01CDRK and PhgPv01CDRK loci using two sets of CY RILs, which were inoculated independently with specific races of the ANT and ALS pathogens, to ensure accurate phenotypic evaluations. We combined separate co-segregation tests for ALS and ANT using a large set of phenotyped plants (4,400 F10 plants) with the power of an Illumina BeadChip (containing over 5,398 SNPs). These large sets of information enabled the creation of a genetic linkage map and revealed the genetic distance between the genes CoPv01CDRK and PhgPv01CDRK at the end of chromosome Pv01. Most of the disease-resistance genes of common bean that have already been characterized are located in distal chromosome regions with high levels of recombination, thus favoring the identification of molecular markers closely linked to disease-resistance genes. However, the presence of repeated sequences, such as khipu satellites, at several large resistance clusters of Pv04, Pv10, and Pv11 might complicate the process of designing locus-specific primers [32].
The candidate region containing the CoPv01CDRK and PhgPv01CDRK loci in linkage group Pv01 is located close to one of the most important clusters of ANT resistance genes in the common bean genome [32]. The Pv01 cluster includes the following ANT disease-resistance genes: Co-1, Co-1HY, Co-14; Co-x, Co-Pa, Co-AC, and Co-1X [23, 25, 28, 29, 30, 31, 53, 68]. The CoPv01CDRK/PhgPv01CDRK loci was found in a genomic region flanked by the markers ss715645251 (50,301,532 bp) and ss715645248 (50,546,925 bp) on Pv01 (Fig 1).
Furthermore, the kinase Phvul.001G243800 was significantly associated with responses to races 65, 73, and 3481 in a genome-wide association study of Andean bean lines [68]. This study used an Andean diversity panel (ADP) and identified the SNP marker ss715645251, which was associated with ANT, in the gene Phvul.001G243800 at position 50,301,532; the authors attributed this gene to the Co-1 locus [68]. Moreover, the Co1HY resistance gene in the Hongyundou cultivar was fine-mapped between the markers TF1 and Clp-N1, positioned at 50,286,325 and 50,332,737, respectively. This 46 Kb region harbors four candidate genes: Phvul.001G243500, Phvul.001G243600, Phvul.001G243700, and the previously mentioned candidate gene for the Co-1 locus, Phvul.001G243800 [29].
The resistance gene Co-x was identified in the Jalo EEP558 cultivar and fine-mapped to a genomic region (56 Kb) flanked by the markers P05 and K06 at positions 50,264,307 and 50,320,695. This 56 Kb region overlaps the 34 Kb of Co-1HY, which contains the four candidate genes mentioned above, and the larger region harbors three additional genes, Phvul.001G243200, Phvul.001G243300, and Phvul.001G243400 [28]. The resistance gene in the Xana cultivar, named Co-1X, was mapped between the markers SNP01_483 and SNP01_490, located in a 153 Kb region from 50,223,771 and 50,377,556. A total of 17 candidate genes were identified in this region, which also covers part of the Co-1HY and Co-x region [53].
Furthermore, the resistance gene of Amendoim Cavalo was recently fine-mapped between the markers SS102 and SS165, located at 50,377,247 and 50,386,692. This 9 Kb interval harbors three candidate genes: Phvul.001G244300, Phvul.001G244400, and Phvul.001G244500. The first of these genes is also present in the Co-1X interval, sharing 0.309 kb.
The AND 277 resistance allele (Co-14) was mapped between the markers CV542014450 (50,513,853) at 0.7 cM and TGA1.1570 (50,741,598) at 1.3 cM. Fine-mapping studies are being conducted to reduce this 227 Kb interval. Mapping analysis located the CoPv01PA resistance gene in a 390 Kb region flanked by the SNP markers SS82 (50,155,987) and SS83 (50,546,985) at distances of 1.3 and 2.1 cM, respectively. This region contains 46 annotated genes, nine of which contain domains with functions related to pathogen resistance: Phvul.001G243200, Phvul.001G243300, Phvul.001G243500, Phvul.001G243600, Phvul.001G243700, the abovementioned Phvul.001G243800, Phvul.001G243100, Phvul.001G245100 and Phvul.001G245300.
CoPv01CDRK was fine-mapped to a 33 Kb interval between CV542014 (50,513,853) and ss715645248 (50,546,985) harboring Phvul.001G246000, Phvul.001G246100, Phvul.001G246200, Phvul.001G246300 and Phvul.001G246400 and close to the genes Phvul.001G245300 and Phvul.001G246800 that encode NB-LRR domains. In this context and based on the physical positions of the aforementioned markers, CoPv01CDRK is clearly positioned downstream of the Co-1, Co-1HY, Co-x, CoPv01AC, and Co-1X loci (Fig 3). These results suggest that the CoPv01CDRK locus is different from the aforementioned loci.
Among the candidate genes identified in the CoPv01CDRK/PhgPv01CDRK loci, the genomic sequences of Phvul.001G246000 and Phvul.001G246400 contain the locus-flanking markers CV542014 and ss715645248, respectively (Fig 2, Table 4). Phvul.001G246000 is a homolog of A. thaliana RNA Helicase 18 (RH18), which was linked to spontaneous chlorosis in hybrids [69]. Several members of the RNA helicase family were found to be involved in the chlorotic phenotype in young leaves by affecting chloroplast biogenesis and reducing photosynthesis [70, 71]. Interestingly, C. lindemuthianum was previously shown to modulate the expression of several genes predicted to be located in the chloroplast [33], and a decrease in plant photosynthetic rates was observed after ANT infection [72]. Indeed, chlorosis can arise due to a delay in chloroplast biogenesis, functionality, or metabolism [73]. Therefore, Phvul.001G246000 could affect photosynthesis to control the chlorosis induced by ANT and ALS, causing resistant plants not to present disease symptoms. Similarly, Phvul.001G246400 is a homolog of Arabidopsis Flavanone 3 Hydroxylase-like (F3H-like), a flavonol synthase involved in the accumulation of flavonoids under light [74]. Hence, it is possible that Phvul.001G246400 could be involved in the biosynthesis of flavonoids to avoid light stress, preventing damage in the chloroplast and contributing to the absence of necrosis after fungal infection. Moreover, flavonoids are secondary metabolites known to be induced upon pathogenic attack, mainly by the induction of the phytohormone jasmonic acid (JA) [75]. JA is well known as an antifungal plant hormone and has previously been linked to defense against ANT in common bean [33] and ALS [36]. The CoPv01CDRK/PhgPv01CDRK loci also contains Phvul.001G246300, a homolog of Arabidopsis ABA Receptor PYL6 (Table 4), which plays a central role in the crosstalk between the ABA and JA responses [76]. This suggests that Phvul.001G246300 may have a role in JA/ABA responses during common bean interaction with C. lindemuthianum and P. griseola.
Moreover, Phvul.001G246100 in the CoPv01CDRK/PhgPv01CDRK loci is a homolog of Arabidopsis Sodium Potassium Root Defective 2 (NAKR2), which belongs to a family of proteins known to have a heavy-metal-associated domain that is linked to the cell division rate in the root meristem [77]. Another gene in the CoPv01CDRK/PhgPv01CDRK loci is Phvul.001G246200, a homolog of Arabidopsis Trichome Birefringence-Like 33 (TBL33), a protein known to be involved in the synthesis and deposition of secondary wall cellulose [78]. Finally, receptor-like kinases are known to function in plant defense against pathogens [79], including common bean response to ANT and ALS [33, 36]. Therefore, we also investigated the closest LRR-Kinases to the fine-mapped CoPv01CDRK/PhgPv01CDRK loci (Table 4). Phvul.001G245300 was shown to be a homolog of Arabidopsis Male Discoverer 2 (MDIS2), a receptor located in the pollen tube that perceives a female attractant signal to promote fertilization [80]. Phvul.001G246800 is homologous to Arabidopsis UDP-Glucosyl Transferase 72E1 (UGT72E1), an enzyme thought to be involved in lignin metabolism [81]. Although none of these proteins has any directly established function in plant defense against pathogens, it is possible that NAKR2, TBL33, and UGT72E1 have roles in plant cell wall strengthening to avoid fungal penetration. MDIS2 is involved in plant fertilization [80]. Another protein putatively involved in plant fertility and linked to ANT resistance in beans is Feronia-like, which is closely related to the ANT resistance gene COK-4 [54] and is thought to function in the regulation of both plant growth and defense [82]. Therefore, it seems that proteins involved in pollen–gametophyte recognition have roles in common bean defense against ANT and possibly also against ALS.
Conclusions
The results presented here showed that CDRK cultivar has the CoPv01CDRK and PhgPv01CDRK genes conferring resistance to races 73, 2047, and 3481 of C. lindemuthianum and race 63–39 of P. griseola. The CoPv01CDRK and PhgPv01CDRK loci co-segregated and were completely linked on chromosome Pv01. New resistance loci, CoPv01CDRK and PhgPv01CDRK, against C. lindemuthianum and P. griseola were fine-mapped in a genomic region of 33 Kb on chromosome Pv01 that harbors five predicted genes. Allelism tests showed that CoPv01CDRKresistance gene is different from the Co-1 and Co-AC loci mapped on Pv01; the physical distances of these genes from CoPv01CDRK are 211,376 bp and 126,216 bp, respectively. In addition, CoPv01CDRK and PhgPv01CDRK resistance alleles are inherited together and can be monitored with high efficiency using SNP markers. These results can be applied to breeding programs aimed at developing bean cultivars with ANT and ALS resistance using marker-assisted selection.
Supporting information
(TIF)
Ten plants per each RIL were evaluated. Lines written in italics underlined carry recombinant events in the predicted location of the CoCDRK/PhgCDRK loci.
(DOC)
Gene positions and their functional annotations are based on the bean reference genome available at Phytozome.
(DOC)
*Genetic position in centiMorgans (cM), based on the map developed by Song et al. (2015).
(DOC)
(DOC)
Acknowledgments
M.C. Gonçalves-Vidigal is grateful for grant from Capes. The authors acknowledge Dr Paul Gepts of the University of California Davis, USA, for kindly provided the seeds of CY RILS, and Dr Pedrina Gonçalves Vidigal for critical reading of the manuscript.
Abbreviations
- ARS
Agricultural Research Service
- NB-LRR
nucleotide-binding leucine-rich repeat
- Nupagri
Núcleo de Pesquisa Aplicada à Agricultura
- PTK
protein tyrosine kinase
- SNP
single nucleotide polymorphism
- STK
serine-threonine protein kinase
- TNL
N-terminal Toll-interleukin-1 receptor (TIR)-like domain
- UEM
Universidade Estadual de Maringá
Data Availability
All relevant data are within the paper and its Supporting Information files.
Funding Statement
This research was supported by Brazilian Federal Funding Institutions National Council for Scientific and Technological Development (CNPq) for financial support and scholarship grants and the Coordination for the Improvement of Higher Education Personnel (Capes). M.C. Gonçalves-Vidigal is grateful for grant from Capes number BEX 88881.170662//2018-01. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Beebe S. Common bean breeding in the tropics. Beebe S. Common bean breeding in the tropics In: Janick J, Editor, Plant Breeding Reviews. Hoboken: Wiley‐Blackwell; 2012. pp. 357–426. 10.1002/9781118358566.ch5 [DOI] [Google Scholar]
- 2.Blair MW, Galeano CH, Tovar E, Muñoz-Torres MC, Castrillón AV, Beebe SE, et al. Development of a Mesoamerican intra-gene pool genetic map for quantitative trait loci detection in a drought tolerant × susceptible common bean (Phaseolus vulgaris L.) cross. Mol Breed. 2012;29: 71–88. 10.1007/s11032-010-9527-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Gepts P. Beans: Origins and development In: Smith C. (Editor) Encyclopedia of Global Archaeology. New York: Springer; 2014. pp. 822–827. 10.1007/978-1-4419-0465-2-2169 [DOI] [Google Scholar]
- 4.Broughton WJ, Hernández G, Blair M, Beebe S, Gepts P, Vanderleyden J. Beans (Phaseolus spp.)—model food legumes. Plant Soil. 2003;252: 55–128. 10.1023/A:1024146710611 [DOI] [Google Scholar]
- 5.Casquero PA, Lema M, Santalla M, De Ron AM. Performance of common bean landraces from Spain in the Atlantic and Mediterranean environments. Genet Resour Crop Evol. 2006;53: 1021–1032. 10.1007/s10722-004-7794-1 [DOI] [Google Scholar]
- 6.Schwartz HF, Pastor-Corrales MA. Bean production problems in the tropics. Cali: CIAT; 1989. p 654. [Google Scholar]
- 7.Correa-Victoria FJ, Pastor-Corrales MA, Saettler AW. Angular leaf spot In: Schwartz H, Pastor-Corrales MA, editors. Bean production problems in the tropics. Cali: CIAT; 1989. pp. 59–75. [Google Scholar]
- 8.Pastor-Corrales MA, Erazo OA, Estrada EI, Singh SP. Inheritance of anthracnose resistance in common bean accession G 2333. Plant Dis. 1994;78: 959–962. 10.1094/PD-78-0959 [DOI] [Google Scholar]
- 9.Guzmán P, Gilbertson RL, Nodari R, Johnson WC, Temple SR, Mandala D, et al. Characterization of variability in the fungus Phaeoisariopsis griseola suggest coevolution with the common bean (Phaseolus vulgaris). Phytopathology. 1995;85: 600–607. 10.1094/Phyto-85-600 [DOI] [Google Scholar]
- 10.Liebenberg MM, Pretorius ZA. A review of angular leaf spot of common bean (Phaseolus vulgaris L). African Plant Prot. 1997;3: 81–106. [Google Scholar]
- 11.Aggarwal VD, Pastor-Corrales MA, Chirwa RM, Buruchara RA. Andean beans (Phaseolus vulgaris L.) with resistance to the angular leaf spot pathogen (Phaeoisariopsis griseola) in southern and eastern Africa. Euphytica. 2004;136: 201–210. 10.1023/B:EUPH.0000030671.03694.bb [DOI] [Google Scholar]
- 12.Stenglein S, Ploper LD, Vizgarra O, Balatti P. Angular leaf spot: A disease caused by the fungus Phaeoisariopsis griseola (Sacc.) Ferraris on Phaseolus vulgaris L. Adv. Appl. Microbiol. 2003;52: 209–243. 10.1016/s0065-2164(03)01009-8 [DOI] [PubMed] [Google Scholar]
- 13.Singh SP, Schwartz HF. Breeding common bean for resistance to diseases: A review. Crop Sci. 2010;50: 2199–2223. 10.2135/cropsci2009.03.0163 [DOI] [Google Scholar]
- 14.Porch TG, Beaver JS, Debouck DG, Jackson SA, Kelly JD, Dempewolf H. Use of wild relatives and closely related species to adapt common bean to climate change. Agronomy. 2013;3: 433–461. 10.3390/agronomy3020433 [DOI] [Google Scholar]
- 15.Kelly JD, Afanador L, Cameron LS. New races of Colletotrichum lindemuthianum in Michigan and implications in dry bean resistance breeding. Plant Dis. 1994;78: 892–894. 10.1094/PD-78-0892 [DOI] [Google Scholar]
- 16.Dean R, VabKan JAL, Pretorius ZA, Hammond-Kosack KE, Pietro AD, Spanu PD, et al. The Top 10 fungal pathogens in molecular plant pathology. Mol Plant Pathology. 2012;13: 414–430. 10.1111/j.1364-3703.2011.00783.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Pastor-Corrales MA, Jara C, Singh SP. Pathogenic variation in, sources of, and breeding for resistance to Phaeoisariopsis griseola causing angular leaf spot in common bean. Trop. Agric. 1998;103: 161–171. 10.1023/A:1018350826591 [DOI] [Google Scholar]
- 18.Kelly JD, Vallejo VA. A comprehensive review of the major genes conditioning resistance to anthracnose in common bean. Hort Science. 2004;39: 1196–1207. 10.21273/HORTSCI.39.6.1196 [DOI] [Google Scholar]
- 19.Correa FJ, Saettler AW. Angular leaf spot of red kidney beans in Michigan. Plant. Dis. 1987;71: 915–918. 10.1094/PD-71-0915 [DOI] [Google Scholar]
- 20.Busogoro JP, Jijakli MH, Lepoivre P. Identification of a novel source of resistance to angular leaf spot disease of common bean within the secondary gene pool. Plant Breed. 1999;118: 417–423. 10.1046/j.1439-0523.1999.00413.x [DOI] [Google Scholar]
- 21.Mahuku GS, Jara C, Cuasquer JB, Castellanos G. Genetic variability within Phaeoisariopsis griseola from Central America and its implications for resistance breeding of common bean. Plant Pathol. 2002;51: 594–604. 10.1046/j.1365-3059.2002.00742.x [DOI] [Google Scholar]
- 22.Geffroy V, Sevignac M, Billant P, Dron M, Langin T. Resistance to Colletotrichum lindemuthianum in Phaseolus vulgaris: a case study for mapping two independent genes. Theor Appl Genet. 2008;116: 407–415. 10.1007/s00122-007-0678-y [DOI] [PubMed] [Google Scholar]
- 23.Mahiya-Farooq, Padder BA, Bhat NN, Shah MD, Shikari AB, Awale HE, et al. Temporal expression of candidate genes at the Co-1 locus and their interaction with other defense related genes in common bean. Physiol Mol Plant Pathol. 2019; 108: 101424 10.1016/j.pmpp.2019.101424 [DOI] [Google Scholar]
- 24.David P, Sévignac M, Thareau V, Catillon Y, Kami J, Gepts P, et al. BAC end sequences corresponding to the B4 resistance gene cluster in common bean: a resource for markers and synteny analyses. Mol Gen Genomics. 2008;280: 521–533. 10.1007/s00438-008-0384-8 [DOI] [PubMed] [Google Scholar]
- 25.Gonçalves-Vidigal MC, Cruz AS, Garcia A, Kami J, Vidigal Filho PS, Sousa LL, et al. Linkage mapping of the Phg-1 and Co-14 genes for resistance to angular leaf spot and anthracnose in the common bean cultivar AND 277. Theor Appl Genet. 2011;122: 893–903. 10.1007/s00122-010-1496-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gonçalves-Vidigal MC, Cruz AS, Lacanallo GF, Vidigal Filho PS, Sousa LL, Pacheco CMNA et al. Co-segregation analysis and mapping of the anthracnose Co-10 and angular leaf spot Phg-ON disease-resistance genes in the common bean cultivar Ouro Negro. Theor Appl Genet. 2013;126: 2245–2255. 10.1007/s00122-013-2131-8 [DOI] [PubMed] [Google Scholar]
- 27.Gonçalves-Vidigal MC, Pacheco CMNA, Vidigal Filho PS, Lacanallo GF, Sousa LL, Martins VSR. Genetic mapping of the anthracnose resistance gene Co-14 in the common bean cultivar Pitanga. Ann Rep Bean Improv Coop. 2016;59: 85–86. [Google Scholar]
- 28.Richard MM, Pflieger S, Sévignac M, Thareau V, Blanchet S, Li Y, et al. Fine mapping of Co-x, an anthracnose resistance gene to a highly virulent strain of Colletotrichum lindemuthianum in common bean. Theor Appl Genet, 2014;127: 1653–1666. 10.1007/s00122-014-2328-5 [DOI] [PubMed] [Google Scholar]
- 29.Chen M, Wu J, Wang L, Mantri N, Zhang X, Zhu Z, et al. Mapping and genetic structure analysis of the anthracnose resistance locusCo-1HY in the common bean (Phaseolus vulgaris L.). PLoS One. 2017; 12(1): e0169954 10.1371/journal.pone.0169954 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Lima-Castro SA, Gonçalves-Vidigal MC, Gilio TAS, Lacanallo GF, Valentini G, Martins VSR et al. Genetics and mapping of a new anthracnose resistance locus in Andean common bean Paloma. BMC Genomics. 2017; 18: 306 10.1186/s12864-017-3685-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Gilio TAS, Hurtado-Gonzales OP, Valentini G, Castro SAL, Elias HT, Song Q, et al. Fine mapping the broad spectrum anthracnose resistance gene in Amendoim Cavalo. Ann Rep Bean Improv Coop. 2017;60: 3–4. [Google Scholar]
- 32.Meziadi C, Richard MMS, Derquennes A, Thareau V, Blanchet S, Gratias A et al. Development of molecular markers linked to disease resistance genes in common bean based on whole genome sequence. Plant Sci. 2016;242: 351–357. 10.1016/j.plantsci.2015.09.006 [DOI] [PubMed] [Google Scholar]
- 33.Oblessuc PR, Baroni RM, Garcia AAF, Chioratto AF, Carbonell SAM, Camargo LEA, et al. Mapping of angular leaf spot resistance QTL in common bean (Phaseolus vulgaris L.) under different environments. BMC Genet. 2012;13: 50 10.1186/1471-2156-13-50 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Keller B, Manzanares C, Jara C, Lobaton JD, Studer B, Raatz B. Fine mapping of a major QTL controlling angular leaf spot resistance in common bean (Phaseolus vulgaris L.). Theor. Appl. Genet. 2015;128: 813–826. 10.1007/s00122-015-2472-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Mahuku GS, Iglesias AM, Jara C. Genetics of angular leaf spot resistance in the Andean common bean accession G5686 and identification of markers linked to the resistance genes. Euphytica. 2009;167: 381–396. 10.1007/s10681-009-9897-4 [DOI] [Google Scholar]
- 36.Oblessuc PR, Perseguini JMKC, Baroni RM, Chiorato AF, Carbonell SAM, Mondego JMC, et al. Increasing the density of markers around a major QTL controlling resistance to angular leaf spot in common bean. Theor Appl. Genet. 2013; 126:1–15. [DOI] [PubMed] [Google Scholar]
- 37.Sartorato A, Nietsche S, Barros EG, Moreira MA. Inheritance of angular leaf spot resistance and RAPD markers linked to disease resistance gene in common beans. Ann Rep Bean Improv Coop. 1999;42: 21–22. [Google Scholar]
- 38.Corrêa RX, Good-God PIV, Oliveira MLP, Nietsche S, Moreira MA, Barros EG. Inheritance of resistance to the common bean angular leaf spot and identification of molecular markers flanking the resistance locus. Fitopatol. Bras. 2001;26: 27–32. 10.1590/S0100-41582001000100005 [DOI] [Google Scholar]
- 39.Nay MM, Souza TLPO, Raatz B, Mukankusi CM, Gonçalves-Vidigal MC, Abreu AFB, et al. A review of angular leaf spot resistance in common bean. Crop Sci. 2019;59: 1376–1391. 10.2135/cropsci2018.09.0596 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Gil J, Solarte D, Lobaton JD, Mayor V, Barrera S, Jara C, et al. Fine-mapping of angular leaf spot resistance gene Phg-2 in common bean and development of molecular breeding tools. Theor Appl Genet. 2019;132: 2003–2016. 10.1007/s00122-019-03334-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Schmutz J, McClean PE, Mamidi S, Wu GA, Cannon SB, Grimwood J, et al. A reference genome for common bean and genome-wide analysis of dual domestications. Nat Genet 2014;46: 707–713. 10.1038/ng.3008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Gepts P. Development of an integrated genetic linkage map in common bean (Phaseolus vulgaris L.) and its use In: Singh S (ed). Bean breeding for the 21st century. Dordrecht: Springer; 1999. pp 53–91. 10.1007/978-94-015-9211-6_3 [DOI] [Google Scholar]
- 43.Kelly JD, Gepts P, Miklas PN, Coyne DP. Tagging and mapping of genes and QTL and molecular marker-assisted selection for traits of economic importance in bean and cowpea. Field Crops Res. 2003;82: 135–154. 10.1016/S0378-4290(03)00034-0 [DOI] [Google Scholar]
- 44.Lopez CE, Acosta IF, Jara C, Pedraza F, Gaitan-Solis E, Gallego G, et al. Identifying resistance gene analogs associated with resistances to different pathogens in common bean. Phytopathology. 2003;93: 88–95. 10.1094/PHYTO.2003.93.1.88 [DOI] [PubMed] [Google Scholar]
- 45.Balardin RS, Jarosz AM, Kelly JD. Virulence and molecular diversity in Colletotrichum lindemuthianum from South, Central, and North America. Phytopathology. 1997;87: 1184–1191. 10.1094/PHYTO.1997.87.12.1184 [DOI] [PubMed] [Google Scholar]
- 46.Thomazella C, Gonçalves-Vidigal MC, Vidigal Filho PS, Nunes WMC, Vida JB. Characterization of Colletotrichum lindemuthianum races in Paraná state, Brazil. Crop Breed App Biotechnology. 2002;2: 55–60. 10.12702/1984-7033.v02n01a08 [DOI] [Google Scholar]
- 47.del Rio LE, Lamppa RS, Gross PL, Brolley B, Prischmann J. Identification of Colletotrichum lindemuthianum race 73 in Manitoba, Canada. Can J Plant Pathol. 2003;25: 104–107. 10.1080/07060660309507055 [DOI] [Google Scholar]
- 48.Gonçalves-Vidigal MC, Thomazella C, Vidigal Filho PS, Kvitschal MV, Elias HT. Characterization of Colletotrichum lindemuthianum isolates using differential cultivars of common bean in Santa Catarina State, Brazil. Braz. Arch. Biol. Technol. 2008a;51: 883–888. 10.1590/S1516-89132008000500002 [DOI] [Google Scholar]
- 49.Nunes MP, Gonçalves-Vidigal MC, Lacanallo GF, Coimbra GK. Comprehension of genetic variability and virulence of Colletotrichum lindemuthianum in common bean In: Biennial Meeting of the Bean Improvement Cooperative, Portland, Oregon, USA: 2013. [Google Scholar]
- 50.Felipin-Azevedo R, Gonçalves-Vidigal MC, Lacanallo GF, Souza MCM, Castro SAL, Caixeta MP, et al. Analysis of diverse Colletotrichum lindemuthianum isolates of common bean (Phaseolus vulgaris L.) from Mato Grosso State, Brazil. Ann Rep Bean Improv Coop. 2014;57: 143–144. [Google Scholar]
- 51.Padder BA, Sharma PN, Awale HE, Kelly JD. Colletotrichum lindemuthianum, the causal agent of bean anthracnose. J Plant Pathol. 2017;99: 317–330. 10.4454/jpp.v99i2.3867 [DOI] [Google Scholar]
- 52.Vlasova A, Capella-Gutiérrez S, Rendón-Anaya M, Hernández-Oñate M, Minoche AE, Erb I, et al. Genome and transcriptome analysis of the mesoamerican common bean and the role of gene duplications in establishing tissue and temporal specialization of genes. Genome Biology 2016;17: 1–18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Murube E, Campa A, Ferreira JJ. Integrating genetic and physical positions of the anthracnose resistance genes described in bean chromosomes Pv01 and Pv04. PLoS One. 2019; 14(2): e0212298 10.1371/journal.pone.0212298 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Oblessuc PR, Matiolli CC, Chiorato AF, Camargo LEA, Benchimol-Reis LL, Melotto M. Common bean reaction to angular leaf spot comprises transcriptional modulation of genes in the ALS10.1 QTL. Front Plant Sci. 2015;6: 1–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Johnson WC, Gepts P. Segregation for performance in recombinant inbred populations resulting from inter-gene pool crosses of common bean (Phaseolus vulgaris L.). Euphytica. 1999;106: 45–56. 10.1023/A:1003541201923 [DOI] [Google Scholar]
- 56.Gepts P. Phenology and development In: Wisiol K, Hesketh JD. Plant growth modeling for resource management. Boca Raton: CRC Press; 1987;2 Pp.1–24. [Google Scholar]
- 57.Mathur RS, Barnett HL, Lilly VG. Sporulation of Colletotrichum lindemuthianum in culture. Phytopathol 1950;40: 104–114. [Google Scholar]
- 58.Sanglard DA, Balbi BP, Barros EG, Moreira MA. An efficient protocol for isolation, sporulation and maintenance of Pseudocercospora griseola. Ann Rep Bean Improv Coop. 2009;52: 62–63. [Google Scholar]
- 59.Cárdenas F, Adams MW, Andersen A. The genetic system for reaction of field beans (Phaseolus vulgaris L.) to infection by three physiologic races of Colletotrichum lindemuthianum. Euphytica. 1964;13: 178–186. 10.1007/BF00033307 [DOI] [Google Scholar]
- 60.Castellanos G, Jara C, Mosquera G. Guía práctica de laboratorio para el manejo de patógenos del fríjol. N° 6. Phaeoisariopsis griseola. Cali: CIAT; 2011.
- 61.Pastor-Corrales MA, Otoya MM, Molina A, Singh SP. Resistance to Colletotrichum lindemuthianum isolates from Middle America and Andean South America in different common bean races. Plant Dis. 1995;79: 63–67. 10.1094/PD-79-0063 [DOI] [Google Scholar]
- 62.Inglis DA, Hagedorn J, Rand RE. Use of dry inoculum to evaluate beans for resistance to anthracnose and angular leaf spot. Plant Dis. 1988;72: 771–774. 10.1094/PD-72-0771 [DOI] [Google Scholar]
- 63.Song Q, Jia G, Hyten DL, Jenkins J, Hwang EY, Schroeder SG et al. SNP assay development for linkage map construction, anchoring whole genome sequence and other genetic and genomic applications in common bean. G3: Genes, Genomes, Genetics. 2015;5: 2285–2290. 10.1534/g3.115.020594 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Van Ooijen JW. JoinMap® 4, Software for the calculation of genetic linkage maps in experimental populations. Kyazma BV, Wageningen: 2006. [Google Scholar]
- 65.Voorrips RE. MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered. 2002;93: 77–78. 10.1093/jhered/93.1.77 [DOI] [PubMed] [Google Scholar]
- 66.Nanami DSY, Gonçalves-Vidigal MC, Castro SAL, Frias AAT, Vidigal Filho PS, Elias HT. Characterization of genetic resistance in Andean common bean cultivar Amendoim Cavalo to Colletotrichum lindemuthianum. Agronomy Science and Biotechnology. 2017;3: 43–52. 10.33158/ASB.2017v3i1p43 [DOI] [Google Scholar]
- 67.Vallejo VA, Kelly JD. Molecular tagging and characterization of alleles at the Co-1 anthracnose resistance locus in common. ICFAI Univ. J. Genetics Evol. 2008;1:7–20. [Google Scholar]
- 68.Zuiderveen GH, Padder BA, Kamfwa K, Song Q, Kelly JD. Genome-wide association study of anthracnose resistance in Andean beans (Phaseolus vulgaris). PLoS One. 2016; 11(6): e0156391 10.1371/journal.pone.0156391 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Plötner B, Nurmi M, Fischer A, Watanabe M, Schneeberger K, Holm S, et al. Chlorosis caused by two recessively interacting genes reveals a role of RNA helicase in hybrid breakdown in Arabidopsis thaliana. The Plant Journal. 2017;91: 251–262. 10.1111/tpj.13560 [DOI] [PubMed] [Google Scholar]
- 70.Chi W, He B, Mao J, Li Q, Ma J, Ji D, et al. The function of RH22, a DEAD RNA helicase, in the biogenesis of the 50S ribosomal subunits of Arabidopsis chloroplasts. Plant Physiol. 2012;158: 693–707. 10.1104/pp.111.186775 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Fristedt R, Scharff LB, Clarke CA, Wang Q, Lin C, Merchant SS, et al. RBF1, a plant homolog of the bacterial ribosome-binding factor RbfA, acts in processing of the chloroplast 16S ribosomal RNA. Plant Physiol. 2014;164: 201–215. 10.1104/pp.113.228338 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Lopes DB, Berger RD. The effects of rust and anthracnose on the photosynthetic competence of diseased bean leaves. Phytopathology. 2001;91: 212–220. 10.1094/PHYTO.2001.91.2.212 [DOI] [PubMed] [Google Scholar]
- 73.Stern DB, Goldschmidt-Clermont M, Hanson MR. Chloroplast RNA metabolism. Ann Rev Plant Bio. 2010;61: 125–155. 10.1146/annurev-arplant-042809-11224 [DOI] [PubMed] [Google Scholar]
- 74.Nakamura Y, Kimura A, Saga H, Oikawa A, Shinbo Y, Kai K, et al. Differential metabolomics unraveling light/dark regulation of metabolic activities in Arabidopsis cell culture. Planta. 2007;227: 57–66. 10.1007/s00425-007-0594-z [DOI] [PubMed] [Google Scholar]
- 75.Wasternack C, Strnad M. Jasmonates are signals in the biosynthesis of secondary metabolites-Pathways, transcription factors and applied aspects- A brief review. New Biotechnology. 2019;48: 1–11. 10.1016/j.nbt.2017.09.007 [DOI] [PubMed] [Google Scholar]
- 76.Aleman F, Yazaki J, Lee M, Takahashi Y, Kim AY, Li Z, et al. ABA-increased interaction of the PYL6 ABA receptor with MYC2 transcription factor: a putative link of ABA and JA signaling. Scientific Reports. 2016;6: 28941 10.1038/srep28941 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Tian H, Baxter IR, Lahner B, Reinders A, Salt DE, Ward JM. Arabidopsis NPCC6/NaKR1 is a phloem mobile metal binding protein necessary for phloem function and root meristem maintenance. The Plant Cell. 2010;22: 3963–3979. 10.1105/tpc.110.080010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Yuan Y, Teng Q, Zhong R, Haghighat M, Richardson EA, Ye ZH. Mutations of Arabidopsis TBL32 and TBL33 affect xylan acetylation and secondary wall deposition. PLoS One. 2016;11(1): e0146460 10.1371/journal.pone.0146460 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Tang D, Wang G, Zhou JM. Receptor kinases in plant-pathogen interactions: more than pattern recognition. The Plant Cell. 2017;29:618–637. 10.1105/tpc.16.00891 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Wang T, Liang L, Xue Y, Jia PF, Chen W, Zhang MX, et al. A receptor heteromer mediates the male perception of female attractants in plants. Nature. 2016; 531: 241–244. 10.1038/nature16975 [DOI] [PubMed] [Google Scholar]
- 81.Lim EK, Jackson RG, Bowles DJ. Identification and characterisation of Arabidopsis glycosyl transferases capable of glucosylating coniferyl aldehyde and sinapyl aldehyde. FEBS Lett. 2005;579: 2802–2806. 10.1016/j.febslet.2005.04.016 [DOI] [PubMed] [Google Scholar]
- 82.Azevedo RF, Gonçalves-Vidigal MC, Oblessuc PR, Melotto M. The common bean COK-4 and the Arabidopsis FER kinase domain share similar functions in plant growth and defense. Mol Plant Pathol. 2018;19: 1765–1778. 10.1111/mpp.12659 [DOI] [PMC free article] [PubMed] [Google Scholar]