Abstract
Physical resiliency declines with age and comorbid conditions. In humans, angiotensin-converting enzyme (ACE) has been associated with attenuation of the decline in physical performance with age. ACE-inhibitor compounds, commonly prescribed for hypertension, often have beneficial effects on physical performance however the generality of these effects are unclear. Here, we tested the effects of the ACE-inhibitor Lisinopril on life span, and age-specific speed, endurance, and strength using three genotypes of the Drosophila melanogaster Genetic Reference Panel. We show that age-related decline in physical performance and survivorship varies with genetic background. Lisinopril treatment increased mean life span in all Drosophila Genetic Reference Panel lines, but its effects on life span, speed, endurance, and strength depended on genotype. We show that genotypes with increased physical performance on Lisinopril treatment experienced reduced age-related protein aggregation in muscle. Knockdown of skeletal muscle-specific Ance, the Drosophila ortholog of ACE, abolished the effects of Lisinopril on life span, implying a role for skeletal muscle Ance in survivorship. Using transcriptome profiling, we identified genes involved in stress response that showed expression changes associated with genotype and age-dependent responsiveness to Lisinopril. Our results demonstrate that Ance is involved in physical decline and demonstrate genetic variation in phenotypic responses to an ACE inhibitor.
Keywords: Angiotensin, Aging, Locomotion, Muscle, Genetic variation
Advanced age confers high risk for disability and mortality. Approximately 20% of older adults living independently require the aid of another person or a walking device and experience higher incidence of falls, hospitalizations, and need for long-term care. In contrast, the ability of older adults to maintain physical activity and function in later life is a hallmark of those with longer health spans (1). Resilient individuals maintain physical function in the face of age-related losses or disease (2). What sets apart resilient from frail older adults is currently unclear (3).
Previous studies report significant variation among individuals in the age at which they begin to exhibit decline in physical ability and this variation has a genetic basis (4). Notably, the gene encoding angiotensin-converting enzyme (ACE), a regulatory enzyme of the renin-angiotensin system (RAS), has been associated with physical performance (5) and longevity (6). ACE inhibitors (ACEi), such as Lisinopril, are commonly prescribed for hypertension and their protective benefits are believed to arise from systemic effects on blood pressure (7). However, there is a muscle-specific RAS including a muscle-specific ACE which is crucial for the regulation of muscle bioenergetics and glucose homeostasis (7). ACEi have been reported to improve physical performance in the elderly persons (8,9). However, some studies report no beneficial effects of ACEi on physical performance (10–12) whereas others report detrimental effects (13,14). Although reasons for these inconsistencies are not known, an intriguing possibility is that genetic differences among individuals produce varied responses to drug treatment. Variable responses could make it difficult for studies to converge on a general conclusion about the effectiveness of treatment.
In this study, we used the fruit fly, Drosophila melanogaster, to test the hypothesis that genetic variation influences age-specific physical performance and that inconsistent responses to ACEi treatment can be due, in part, to genetic differences among individuals. The closest Drosophila ortholog to mammalian ACE is angiotensin-converting enzyme (Ance) (15). Ance is a single-domain protein with 76% amino acid sequence similarity to the active C-domain of human somatic ACE (16). Biochemical analyses support similar catalytic functions of Ance and ACE (15,17). Structural analysis indicates that the binding site of antihypertensive drugs is conserved, suggesting that the mechanism by which Lisinopril binds to fly Ance is similar to that of human ACE (16).
We tested whether Lisinopril treatment affects age-specific physical performance and longevity, using three genotypes of the Drosophila Genetic Reference Panel (DGRP) (18) that differ in life span (19). We evaluated genome-wide changes in gene expression in response to Lisinopril using RNA-sequencing (RNA-seq) of two DGRP lines at young and old ages to identify evolutionarily conserved loci that modulate the response to ACEi treatment. Finally, to investigate a potential physiological mechanism to explain the effects of ACEi on physical performance, we tested whether Lisinopril treatment affects protein aggregation in skeletal muscle. Protein aggregation is associated with many age-related diseases (20), and skeletal muscle aging with deterioration in muscle integrity (21). Prior research has shown that skeletal-muscle aggregates contribute to declines in muscle mass and function in humans and that there is twice as much aggregate in the older individuals (22). Although in vivo studies have shown that ACEi can reduce amyloid plaque deposition in Alzheimer’s models (23,24), the impact of ACEi on age-related buildup of skeletal muscle protein aggregates is unknown (22).
Methods
Drosophila Stocks and Maintenance
We used virgin males of DGRP_73, DGRP_229, and DGRP_304 from the Drosophila Genetic Reference Panel for all survivorship and physical performance assays. The dj667-Gal4 driver(w1118; P(w[+mW.hs]=GawB)DJ667, stock # 8171), UAS-RNAi-Ance (Harvard TRiP.GLC01369)attP2, mhc-Gal4 driver(w[*]; P(w[+mC]=Mhc-RFP.F3-580)2, P(w[+mC]=Mhc-GAL4.F3-580)2/SM6b, stock # 38464), UAS-GFP (y1, w*; P(w[+mC]=UAS-mCD8::GFP.L)LL5, P(UAS-mCD8::GFP.L)2, stock # 5137, or y*, w*; P(w[+mC]=UAS-2EGFP]AH3, stock # 6658), attP2 control stock (y1v1; P(CaryP)attP2, stock # 36303), and mCherry control stock (y1sc* v1; P(VALIUM20-mCherry)attP2, stock # 35785) were obtained from the Bloomington Drosophila Stock Center. All stocks and control groups were fed standard food medium (solid ingredients: 79% cornmeal, 16% yeast, and 5% agar). Flies were maintained in population cages at 25°C and 55% relative humidity under a 12-hour light/dark cycle. All physical performance assays were completed between 8 am and 2 pm.
Lisinopril Treatment
Treatment groups were administered 1-mM Lisinopril (Sandoz Pharmaceuticals, Princeton, NJ), which was homogenously mixed into the fly food. This dosage was based on the human dosing equation, dependent on body mass, and estimated for mass of fly (mean = 0.5 mg). Pilot studies tested serial doses of Lisinopril in the following concentrations: 0.2, 0.4, 1, and 10 mM. The 1-mM dose was used in this study because it had a large positive effect on both the mean and maximum life span (Supplementary Figure 1).
Feeding Rate Assay
We used the Capillary Feeder Assay (25) to account for potential differences among genotypes in feeding rates on the physical performance measures. Groups of four 5-day old virgin males from each genotype were placed in a vial and allowed to feed on either 1-mM sucrose or 1-mM Lisinopril with sucrose from capillary tubes (#53432–706, VWR) for 24 hours. To account for the effect of evaporation, we calculated mean evaporation from control 1-mM sucrose (n = 10) and sucrose plus Lisinopril (n = 10) capillaries using vials without flies. Food loss by evaporation or consumption by flies was measured using a digital caliper. We used the following formula to determine total consumption: food consumption of flies (μL) = (food loss [μL] − evaporative loss [μL])/total mg of flies in the vial.
Validation of Muscle-Specificity of Gal4 Lines
To confirm the muscle-specificity of the dj667-Gal4 (26) driver we crossed virgin females with male UAS-GFP. We carried out a similar cross with another commonly used muscle driver, mhc-Gal4 (27) for comparison. Offspring were collected and aged to 1 week. Flies were dissected along the dorsal midline and fixed. High resolution images were taken using a Leica SP5 confocal microscope using a ×10 objective. Images of live, whole flies were taken with ×20 magnification using a Leica M205 fluorescent stereoscope (Buffalo Grove, IL). Images were visually examined for presence and location of GFP fluorescence.
Generation of a Skeletal Muscle-Specific Ance Knockdown
dj667-Gal4 male flies were crossed with virgin female UAS-RNAi-Ance flies to knock down Ance expression in skeletal muscle. We denote the F1 generation as RNAi-Ance from here on. Flies used to control for the effects of RNAi knockdown were derived from the following crosses (1): dj667-Gal4 × y1,v1; P(CaryP)attP2 control (2), dj667-Gal4 × y1,sc*,v1; P(VALIUM20-mCherry)attP2 control for activation of RNAi machinery. Note that the mCherry line also contains attP2. We denote the F1 generation as attP2 or mCherry.
Validation of RNA Knockdown
We used real-time quantitative reverse transcriptase polymerase chain reaction to confirm that RNAi reduced Ance expression. One-week-old male flies from RNAi-Ance and controls were flash frozen with liquid nitrogen and stored at −80°C. RNA was extracted from homogenized tissue of 10 males per genotype using the RNeasy Mini Kit from Qiagen. DNA was removed from samples using the TURBO DNA-free Kit (ThermoFisher Scientific). Complementary DNA (cDNA) was synthesized using a BioRad iScript cDNA Synthesis Kit and 0.25 µg of RNA. 1X iTaq Universal SYBR Green Supermix (BioRad) was then mixed with 0.5 µL of the newly synthesized cDNA and 0.5 µM of the appropriate forward and reverse primers. Real-time amplification was performed on a BioRad CFX384 Real-time Detection System. Three biological replicates were run for every reaction, each with three technical replicates. Relative expression values were normalized to Ribosomal Protein L32 (rp49) expression levels. Primers for Ance and rp49 were designed according to the fly primer bank (http://www.flyrnai.org/cgi-bin/DRSC_primerbank.pl). Primers for Ance expression were as follows: Forward, GTGATACCACCAAGTTCCAATGG, Reverse, GGCATAGTCGTCTTCAGGTAGAG. Primers for rp49 expression were as follows: Forward, GTGAAGAAGCGCACCAAGCAC, Reverse, ACGCACTCTGTTGTCGATACCC.
Physical Performance Assays
We measured climbing speed, endurance, and strength as indicators of physical performance. We used 30 flies per genotype, per treatment (control or Lisinopril) at each of three ages (weeks 1, 3, and 5), per performance measure. Independent sets of flies were used for measurements of physical performance at each age, so a given fly was only measured for physical performance at one age.
Climbing speed was tested by placing a single fly into the bottom of a Costar 25-mL in 2/10, non-pyrogenic serological pipet, marked at 9 and 27 cm. Flies were tapped down to the bottom of the inverted pipet and a timer started once the body of the fly passed the zero mark on the pipet. The timer was stopped when the fly reached 9 cm. Nine centimeters was chosen based on a pilot study which found that more than 90% 1-week-old flies climbed 9 cm. Endurance was measured using the same technique but was based on the distance traveled in 15 seconds or the time it took for an individual to climb to a height of 27 cm. This is similar to measures of endurance in human frailty studies (28). The design of the endurance assay was based on pilot studies in which more than 90% of 1-week-old flies were able to climb 27 cm within 15 seconds, but less than 10% could do so at 5 weeks of age.
Strength was estimated by measuring the time it took for a fly to escape from a clear, colorless, 1 × 3 cm, strip of double-sided Scotch tape. Although this is not a direct measurement of muscle contractile activity or force, this measure is a surrogate for strength in the same way that grip strength is used to indicate frailty in human studies (28). For each trial, individual flies were held at −20°C for 60 seconds and then placed dorsal side-down onto the tape. Each wing was then gently tapped into place and the time to escape measured. Pilot studies indicated that the maximum length of time taken for 1-week-old flies to escape was 3 minutes. If flies were unable to escape from the tape within that time, a maximum score of 3 minutes was given.
Development of a Physical Performance Index in Drosophila
In humans, a frailty index is often used to identify patients at risk of adverse outcomes and as a predictor of life span (28). We took a similar approach, using a composite of climbing speed, endurance, and strength to establish a fly performance phenotype (FPP). These measures are comparable to criteria used to indicate frailty in humans and mice (28,29). Each fly was ranked for its performance on each of the three tests, and the ranked data were then divided into quartiles as follows. Individuals were classified as “high performers” and placed in the highest quartile if their score in each physical test did not fall 1.5 SD below the cohort mean for speed and endurance and did not fall 1.5 SD above the cohort mean for strength (larger values indicate poorer performance in the strength assay). Individuals were classified as “medium performers” if their performance score was 1.5 SD below the mean for climbing speed and endurance or above the mean for strength for either one or two of three tests; if ranked 1.5 SD away from the mean for one of the tests they were placed in the second quartile while if they ranked 1.5 SD away the mean for two of the tests they are placed in the third quartile. Individuals were classified as “low performers” if performances were 1.5 SD away from the mean in the direction of poor performance for all three tests.
Drosophila Survivorship Studies
We used population cages to assess differences in survivorship among genotypes, the effect of Lisinopril on survivorship, and any differences among genotypes in the response to Lisinopril. We established populations of newly emerged flies using 250–270 male virgin flies for each of six Plexiglas population cages (20l × 21w × 21.5h cm), two cages for each genotype, one population on control food and the other on Lisinopril. Forty milliliters of control or drug food was placed in 100 × 15 mm BD Falcon plastic petri dishes and replaced in their respective cages every other day. We then examined the effect of chronic Lisinopril treatment on survivorship of all three DGRP lines compared to untreated flies of these genotypes. Flies were monitored every other day and dead flies were removed until all individuals had died (30–33).
Whole-Mount Immunostaining of Drosophila Skeletal Muscle and Protein Analysis
To visualize and quantify protein aggregation we used standard procedures for immunostaining Drosophila skeletal muscle (34). In brief, 32 one- and five-week-old control and treated flies from each genotype were dissected by separating the thorax from the head and the abdomen. Thoraces were cut longitudinally into halves and cuticles removed. Thoraces were transferred, fixed, and stained using anti-ubiquitinylated proteins antibody, clone FK2 (1:100; Millipore, cat. no. 04-263) to mark protein aggregates, Alexa Fluor 488 phalloidin to label actin (1:200; cat no. A22284; Molecular Probes, Eugene, Oregon), and Cyanine3 anti-Rat secondary antibody (1:200; ThermoFisher Scientific, cat no. A10522). Images were taken using Zeiss LSM 78, ×63 oil immersion. Protein aggregate areas (µm2) were measured for set size regions of tissue within each whole tissue. We analyzed 30 regions from 32 to 40 individuals/genotype/treatment, at each age using Volocity 6.3 PerkinElmer cellular imaging.
Transcript Profiling of Control and Treated Drosophila
RNA extraction and sequencing
Virgin male flies from lines DGRP_229 and DGRP_73 were maintained on control or Lisinopril-treated food for either 1 or 5 weeks. We extracted RNA from at least 50 flies for each age and treatment combination. Prior to RNA extraction, flies were separated into two groups to yield two biological replicates for each age and treatment combination. Total RNA was extracted with QIAzol Lysis Reagent (Qiagen) and the Quick-RNA MiniPrep Zymo Research Kit (Zymo Research). Ribosomal RNA (rRNA) was depleted from 5 μg of total RNA using the Ribo-Zero Gold Kit (Illumina, Inc.). Depleted mRNA was fragmented and converted to first-strand cDNA using Superscript III reverse transcriptase (Invitrogen). Second-strand cDNA was synthesized using dUTP instead of dTTP to label the second-strand cDNA. cDNA from each sample was used to produce barcoded cDNA libraries using NEXTflex DNA Barcodes (Bioo Scientific) with an Illumina TruSeq compatible protocol. Each sample was subjected to end-repair (Enzymatics), adenylation of 3′-ends (Enzymatics), and ligation of indexed adapters (Enzymatics and Bioo Scientific). Each enzymatic reaction was purified using 1.8× Agencourt AMPure XP beads (Beckman-Coulter). Size selection of each library was performed using Agencourt AMPure XP beads (Beckman Coulter) to an approximate insert size of 130 bp and a total library size of approximately 250 bp. Second-strand cDNA was digested with Uracil-DNA Glycosylase prior to PCR-enrichment to produce directional cDNA libraries. PCR-enrichment of the purified barcoded DNA was carried out with KAPA HiFi Hot Start Mix (Kapa Biosystems) and NEXTflex Primer Mix (Bioo Scientific). Libraries were quantified using the Qubit dsDNA HS kit (Life Technologies) and their sizes determined with the 2100 Bioanalyzer (Agilent Technologies). Each sample was diluted to equal molarity, quantified, multiplexed, denatured, and diluted to 14 pM. Clonal clusters for each pooled library sample were generated on the Illumina cBot and then sequenced on the Illumina HiSeq2500 using 125 bp single-read v4 chemistry (Illumina, Inc.). We generated two multiplexed libraries containing eight samples each (1- or 5-week-old flies). Each multiplexed library was run on one lane of the HiSeq2500.
Barcoded sequence reads were demultiplexed using the Illumina pipeline v1.9. Adapter sequences were trimmed using cutadapt v1.6 (35) and trimmed sequences shorter than 50bp were discarded from further analysis. Trimmed sequences were then filtered for ribosomal RNA sequences by aligning against a database containing the complete 5S, 18S-5p8S-2S-28S, mt:lrRNA, and mt:srRNA sequences using BWA v0.7.10 (MEM algorithm with parameters “-v 2 –t 4”) (36). The remaining sequences were aligned to the D melanogaster genome (BDGP5) and known transcriptome (FlyBase v5.57) using STAR v2.4.0e (37). Read counts were computed for known gene models using HTSeq-count (38) with the “intersection-nonempty” assignment method. Tabulated read counts were then analyzed for all known genes across all samples using EdgeR (39) as follows. First, genes with low expression overall (<20 aligned reads in at least one replicate of every sample condition) were excluded from the analysis. Library sizes were recomputed as the sum of reads assigned to the remaining genes, and normalized using the Trimmed Mean of M-values method (40). We then used the generalized linear model (GLM)-based methods for estimating tag-wise dispersion and fit model parameters to the following model design: x = l + d + l × d + b + ε, where x = observed log2(read count), l = line effect (RAL_73 vs RAL_229), d = drug effect (Lisinopril vs Control), l × d = line by drug interaction effects, b = overall batch effects (each line and drug combination was analyzed using two biological replicates, with the first replicates processed in a separate batch from the second replicates), and ε = model error following a negative binomial distribution with estimated gene-wise dispersion. We then selected gene expression levels with significant line effects, drug effects and line by drug interactions passing a 10% false discovery rate threshold (based on Benjamini–Hochberg corrected p values) from the edgeR likelihood ratio test on the interaction term coefficient (41). This analysis was run separately and independently on 1- and 5-week-old flies.
Statistical Analyses
Climbing, and endurance data were analyzed with analysis of covariance (PROC GLM, SAS V9.3) using the model: y = c + m + g + t + a + all interactions + E, where c is a constant, m was mass, g tested for differences among DGRP lines, t tested the effects of Lisinopril treatment, and a is the effect of age. None of the interactions between mass and the main effects were significant so interaction terms involving mass were dropped from the model. Strength was analyzed in the same manner, however, mass was not measured so analysis of variance was used to analyze the effects of genotype, treatment, and age. Feeding rate data were analyzed in analysis of variance using the model: y = c + g + t + g × t + E, where c is a constant, g tested for differences among DGRP genotypes, t tested the effects of Lisinopril treatment, and E is the error. The Tukey test was used for post hoc comparisons when appropriate. Climbing speed and endurance data were transformed to natural logs to satisfy assumptions of analysis of variance. Survivorship data were analyzed by Cox regression proportional hazards models (PROC PHREG, SAS V9.3). We used reduced models to compare survivorship of control and Lisinopril-treated flies for each genotype separately. We also used a full model in which we compared the effects of genotype and treatment in a single model. We used the sensitivity index of Falconer (42) to compare the sensitivity of the performance traits to age. The Falconer sensitivity measure indicates the steepness of change in the intervals. To calculate the sensitivity of each trait to age for each genotype, we took the average phenotypic value of the trait for a given genotype at week 1 minus the average value at week 5 divided by the average change of this trait with age across all lines. We tested for the effects of Lisinopril treatment on protein aggregate intensity using a t test.
Results
Genetic Variation Insurvivorship and Health Span
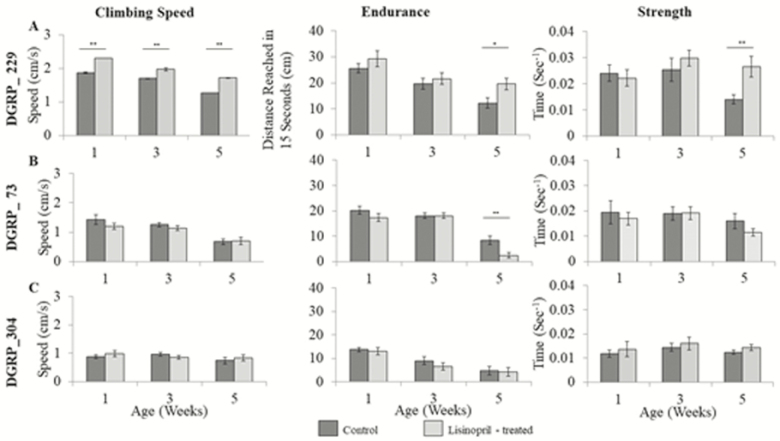
Genotypes differed significantly in survivorship (p < .0001). We found a significant decline in climbing speed, endurance, and strength (Figure 1, Supplementary Table 1) with age. However, the effect of age on performance measures varied across the genotypes (Supplementary Table 1). Using Falconer’s estimate of sensitivity (42), the climbing speed and endurance of DGRP_73 was much more sensitive to age than DGRP_229 and DGRP_304 (Table 1).
Figure 1.
Ace inhibition improves speed, endurance, and strength in an age- and genotype-specific manner. (A) DGRP_229 (n = 540). (B) DGRP_73 (n = 540). (C) DGRP_304 (n = 540). Data are means ± SEM. *p < .05; **p < .01. The measure of strength is the inverse of the amount of time in seconds it took for flies to escape.
Table 1.
Sensitivity to Effects of Aging in DGRP Genotypes
| Genotype | Assay | Week 1–3 Sensitivity | Week 3–5 Sensitivity | Week 1–5 Sensitivity |
|---|---|---|---|---|
| DGRP_229 | CS | 1.196 | 0.900 | 1.110 |
| EN | 0.511 | 1.201 | 1.447 | |
| ST | 1.051 | 2.518 | 1.539 | |
| DGRP_304 | CS | 0.857 | 0.587 | 0.300 |
| EN | 0.589 | 0.549 | 0.586 | |
| ST | 0.642 | 0.039 | 0.443 | |
| DGRP_73 | CS | 0.946 | 1.512 | 1.590 |
| EN | 1.900 | 1.250 | 0.967 | |
| ST | 1.307 | 0.443 | 1.018 |
Note: Higher Sensitivity Index score indicates greater effect of aging on climbing speed (CS), endurance (EN), and/or strength (ST). Overall decline, measured from week 1 to week 5 of age, is highest in EN and ST of DGRP_229 and in CS of DGRP_73. Overall decline is minimal in CS, EN, and ST of DGRP_304.
We noted that some genotypes performed well in one or more of the measures, but not in all three. Therefore, we used the FPP, to classify flies from each genotypeas exhibiting high capacity (HC), medium capacity (MC), or low capacity (LC) to perform physical assays. DGRP_73 flies displayed not only the shortest average life span but also had the highest prevalence of LC performers across all ages (49.8%). In contrast, DGRP_229 had the highest prevalence of HC performers (48.9%) and a mean life span that fell between the other two lines. DGRP_304 had the highest prevalence of MC performers (43.9%) and the longest average life span.
Lisinopril affects Life- and Health-Span Traits
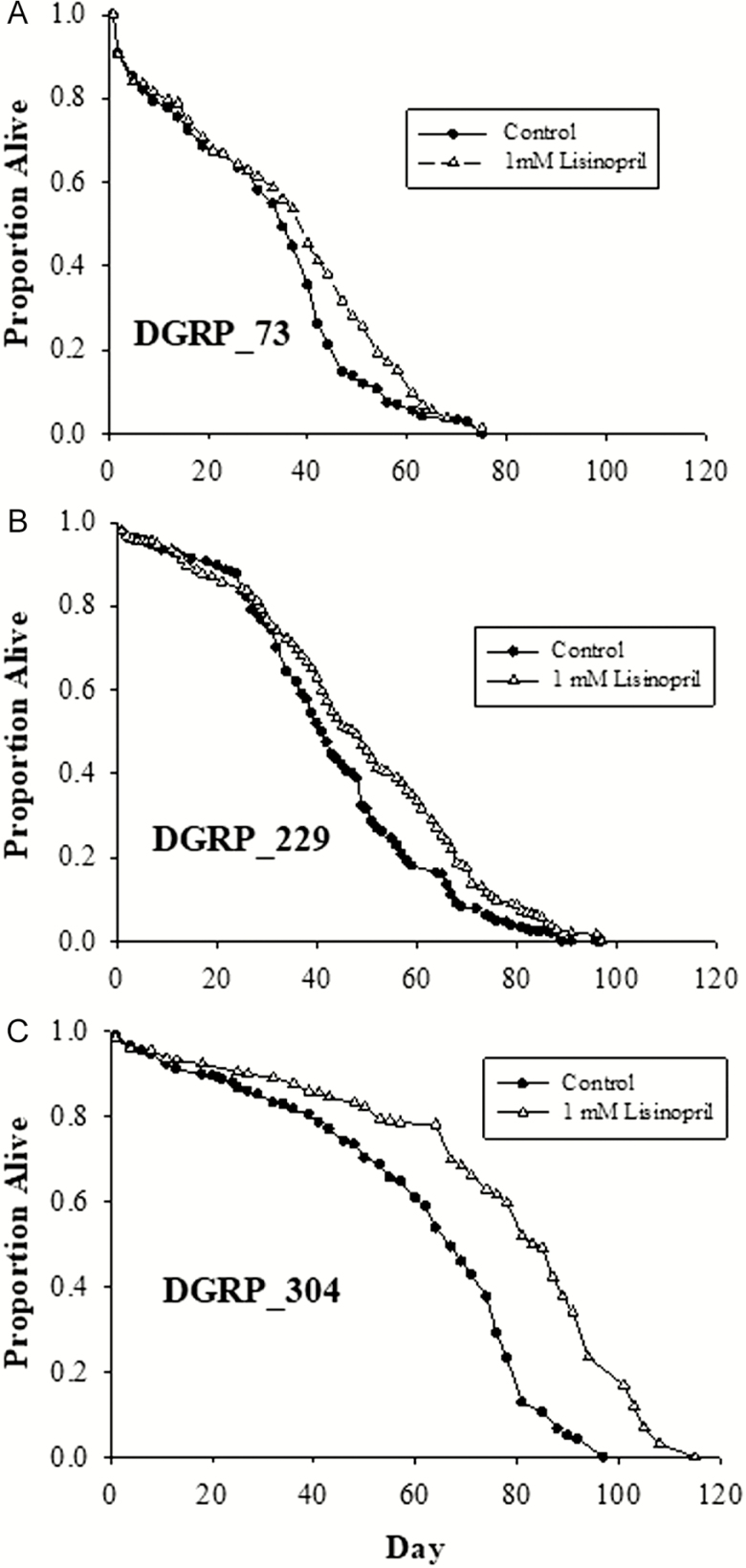
On the basis of the full model, Lisinopril treatment increased mean life span across all DGRP genotypes(p < .0001) but did so to different degrees among lines (Figure 2). DGRP_304 had the highest gain in life span in response to 1-mM Lisinopril followed by DGRP_229 and DGRP_73.
Figure 2.
Effects of Lisinopril on life span. (A) DGRP_73 (p < .01; n = 520). (B) DGRP_229 (p < .01; n = 520). (C) DGRP_304 (p < .001; n = 520). Solid black lines depict control and dashed lines depict Lisinopril-treated flies.
To ensure that the phenotypic differences in response to drug treatment was not due to differences in drug dosage due to different feeding rates, we performed the Capillary Feeder Assay (25). Interestingly, DGRP_304 flies that had the highest gains in survivorship consumed the least amount of Lisinopril(Supplementary Figure 2). In contrast, DGRP_73 flies had the highest consumption rate of Lisinopril, followed by DGRP_229. Adding Lisinopril to the food did not reduce food intake in any of the genotypes (Supplementary Figure 2).
The effect of Lisinopril on the age-specific climbing speed, endurance, and strength differed among genotypes (Figure 1, Supplementary Table 1). Lisinopril treatment improved performance across all ages and attenuated the decline of all three physical performance measures for DGRP_73 flies (Figure 1A). In contrast, DGRP_229 flies showed no significant effect of treatment on climbing speed or strength and a significant decrease in endurance (Figure 1B). Lisinopril treatment did not affect climbing speed, endurance, or strength in DGRP_304 flies (Figure 3C).
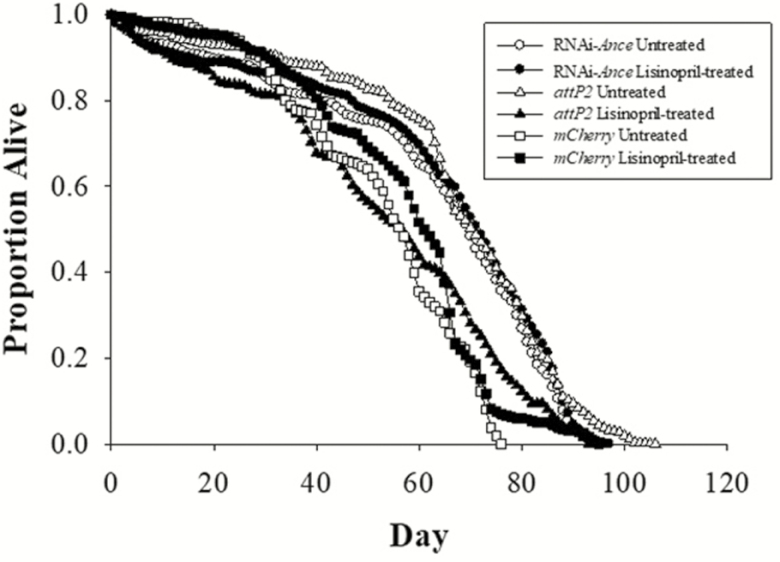
Figure 3.
Effect of Lisinopril on life span depends on Ance expression in skeletal muscle. Lisinopril treatment extended the life span of mCherry control flies, and reduced life span of attP2 flies. There is no significant difference in life span between untreated and treated dj667-Gal4 × RNAi-Ance males.
Because the three genotypes displayed differential responses to treatment with respect to physical performance traits, we compared the effects of Lisinopril on the composite measure, FPP, at 3 and 5 weeks of age. Consistent with the results from the individual traits, DGRP_229 exhibited increased incidence of HC flies and a decreased incidence of MC and LC flies at both ages, as well as an increased overall prevalence of HC flies, when treated with Lisinopril (Supplementary Figure 3A–C). However, Lisinopril treatment had little effect on the prevalence of HC flies for DGRP_73 at either age. Lisinopril treatment decreased the incidence of MC flies in this genotype and increased the incidence of LC flies (Supplementary Figure 3D–I). In DGRP_304, we observed a slight increase in HC and LC flies and a reduction in MC flies at 3 weeks of age. At 5 weeks of age, Lisinopril treatment also caused a slight increase in HC flies but, in contrast to week 3 of age, we saw a slight decrease in LC flies.
Effect of Lisinopril on Survival is Influenced by Expression of Ance in Skeletal Muscle
We used RNA interference (RNAi) to examine the effects of Lisinopril on life span when Ance expression was reduced in skeletal muscle. The effect of Lisinopril varied between the control lines, significantly extending life span for the mCherry genotype and causing a significant reduction in life span in the attP2 genotype. We attribute these different responses to Lisinopril to the different genetic backgrounds of the attP2 and mCherry stocks. However, Lisinopril treatment had no effect on life span relative to the control when expression of Ance in skeletal muscle was reduced (Figure 3).
To ensure muscle specificity of the RNAi in this experiment, we characterized the expression of GFP under the control of the dj667 driver, (dj667-Gal4 × UAS-gfp, n = 10) and compared GFP expression with another commonly used driver for muscle, mhc(mhc-Gal4 × UAS-gfp, n =10). In agreement with previous studies (26,43), we found that both lines induce high expression of GFP in muscle, and no detectable expression elsewhere (Supplementary Figure 4). We used real-time quantitative reverse transcriptase polymerase chain reaction to test the efficacy of the Ance RNAi and observed a 2- to 5-fold decrease compared to controls (Supplementary Figure 5).
Changes in Protein Aggregation in Skeletal Muscle With Age and Treatment
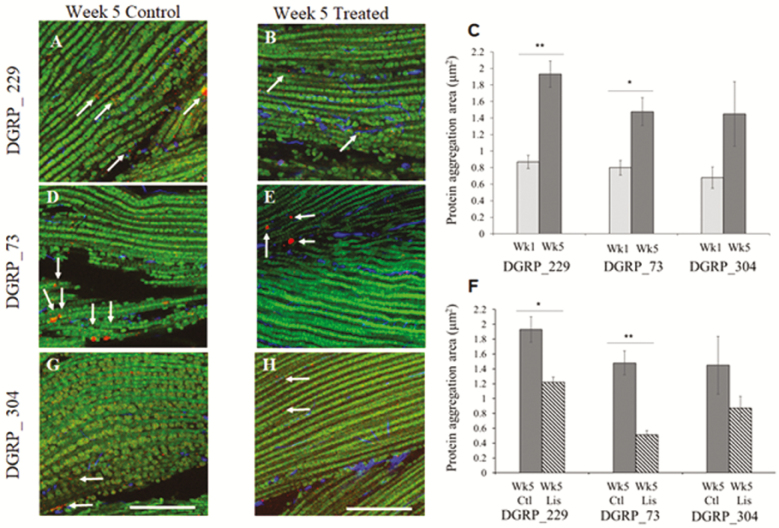
Previous studies suggest that protein aggregation contributes to the decline in muscle function (34,44) and protein aggregation is affected by ACEi (45–48). We hypothesized that treatment with Lisinopril augments repair and increases the turnover of dysfunctional proteins in skeletal muscle. If so, then the degree of improvement in physical performance in response to Lisinopril would dependent on the degree of protein accumulation with age in each genotype. We used standard procedures for immunostaining Drosophila skeletal muscle (34) and measured protein aggregation measured from set size regions of tissue within each whole tissue. We found a significant increase in protein aggregation area with age in the fibrillar muscles of DGRP_229 flies (p < .0001) (Figure 4). At old age, treatment with Lisinopril significantly reduced protein aggregation area in DGRP_229 (p = .0002) (Figure 4 A, B, and F). Similarly, protein aggregation area also significantly increased with age in line DGRP_73 (p = .0004) and treatment with Lisinopril significantly reduced protein aggregation area (p < .0001) (Figure 4 D, E, and F). In contrast, in DGRP_304 there was a slight but not significant change in protein aggregation area with age (p = .0680) and treatment with Lisinopril had no effect on aggregation area (Figure 4 G, H, and F). These patterns parallel the physical performance results, demonstrating that treatment with Lisinopril improved speed, endurance, and strength in DGRP_229 flies, had little to no influence on speed, endurance, and strength in DGRP_73 flies, and no effect on physical performance in DGRP_304 flies.
Figure 4.
Lisinopril reduces protein aggregation with age and line. Immunostaining of indirect flight muscles from control and treated (A and B) DGRP_229, (D and E) DGRP_73, and (G and H) DGRP_304 flies at 5 weeks of age. Poly-Ubiquitin (Cy3, red) immunoreactivity reveals deposition of aggregates (arrows), phalloidin staining (green) labels F-actin, and DAPI (blue) marks nuclei. (C) Mean intensity of protein aggregates increases with age in DGRP_229 and DGRP_73. Light gray bar is 1 week of age, dark gray is 5 weeks of age (F) At old age, treatment reduces mean intensity of protein aggregates in DGRP_229 and nearly reduces mean intensity in DGRP_73. Dark gray bar is control, striped gray is Lisinopril-treated; data are means and SEM bars (n = 8 to 12 flies). Scale bar, 100 µm. *p < .05; **p < .01 unpaired t test. Note that the 5-week control data are used for the comparisons shown in panels C and D, so the p values shown are not adjusted for use of the data in two comparisons.
Transcript Profiling of Control and Treated Drosophila
We used RNA-seq to evaluate the effects of Lisinopril treatment on gene expression, comparing DGRP_229 and DGRP_73, two of the lines whose physical performance responded differently to Lisinopril. We first assessed the average effect of drug treatment on transcript levels, when we pooled data from both lines at each age. For all results reported below we used 5% false discovery rate corrected p values. We used gene ontology (GO) cluster analyses (41) to identify genes in resultant gene lists with similar molecular or biological function. For 1-week-old flies, 25 genes were differentially expressed between drug-treated and control flies (Supplementary Table 2A). With this small number of genes, no GO terms were significantly overrepresented in this gene list. However, four genes have been implicated in stress response (Cytochrome P450-4e3, Invadolysin, Turandot A and Turandot C), and Troponin C isoform 4 is involved in muscle activation. In contrast to the results from 1-week-old flies, 192 genes were differentially expressed between Lisinopril-treated and control flies when they were 5 weeks old (Supplementary Table 2B). GO analysis identified six distinct clusters of functionally related genes that were overrepresented (Supplementary Table 3). These clusters include genes involved in detoxifying xenobiotics (CYP genes), immunity, and metabolism.
As the two lines responded differently to Lisinopril, we also tested for genes that responded differently to Lisinopril treatment (those genes that exhibited significant genotype by drug treatment interaction) at each age. At 1 week of age, 117 genes exhibited a significant genotype by drug treatment interaction (Supplementary Table 4A). GO analysis revealed three clusters of genes that were functionally overrepresented in this list (Supplementary Table 5A). The first cluster was enriched for CHK (Check Point Kinase) genes which have been implicated in stress responses (49). The second cluster contained genes in the Turandot family, a family of genes also associated with stress response (50), including the immune response (51). The last cluster included many genes involved in membrane transport. At 5 weeks of age we found far fewer genes that exhibited genotype-specific responses to the drug treatment (Supplementary Table 4B). This may explain the relatively smaller phenotypic differences between the control and drug-treated flies among genotypes at older compared to younger ages. GO analysis identified one cluster of genes significantly overrepresented in this list and these were primarily involved in proteolysis (Supplementary Table 5B).
Discussion
Medications commonly used in older individuals, such as ACEi, have been reported to attenuate age-related decline in physical performance (8,23,24,52) and increase life span (53–55). However, treatments with ACEi are not always effective (13) and determinants of inter-individual variation in response to ACEi are largely unknown. Conflicting results between studies can potentially be explained by genetic differences among individuals. The complexity of the RAS and the significant cross talk and interaction (both positive and negative feedback loops) between different parts of the system as well as interaction with other hormonal systems added to the difficulty in tracking direct from indirect effects of targeting the system. Although the RAS system has only been identified in vertebrates, many genes regulating RAS are also found in Drosophila (15,16,19,34). This indicates that these genes serve other physiological functions and that amelioration of age-related declines in vertebrates by treatment with Lisinopril may be due to additional physiological effects that are not solely due to blockade of the RAS pathway.
As physical performance and life span are closely linked in humans (56,57), we compared changes in physical performance, FPP, and life span in our three fly lines to test the impact of genetic background on the effects of ACE inhibition. Lisinopril treatment influenced age-related decline of climbing speed, endurance, and strength that was dependent on genotype. Treatment of DGRP_229 flies significantly attenuated the decline of all three measures of physical performance: climbing speed, endurance, and strength. In contrast, treated flies of DGRP_73 and DGRP_304 showed no effect on climbing speed nor endurance, but rather only on strength. To further segregate responders from non-responders, we tested the effects of Lisinopril on the composite measure, FPP. We noted a decline in prevalence of LC performance in DGRP_229 and an increase in percentage of HC flies with treatment. As decline in physical function is associated with rate of mortality, we tested the relationship between change in physical function and mortality with treatment. While treatment with Lisinopril significantly extended the average life span of all DGRP genotypes, this reduction in mortality was associated with improvement of physical function only in DGRP_229 flies. We also note that Lisinopril provided the biggest extension of longevity to DGRP_304 flies, even though there was little change in performance in response to drug treatment. These results seem to indicate that the genetic factors, and by extension physiological factors, regulating longevity are largely independent of those influencing age-specific physical performance measures. Interestingly, DGRP_304 had no detectable increase in protein aggregation in muscle with age. Unfortunately, we do not know the causes of death in the flies. Such information would be useful for understanding why these traits appear to be disconnected.
We tested whether survivorship is affected by the expression of Ance in muscles. Although reduced Ance expression in muscles did not influence life span, we did show that alterations in life span resulting from Lisinopril treatment depended on Ance expression. Recent work on the same three DGRP genotypes in our study found that Lisinopril alters mitochondrial respiration and the production of ROS levels in an age and genotype-specific manner (58). Whether the Lisinopril induced changes in mitochondrial characteristics and related changes in metabolism explain the genotype-specific effects of Lisinopril on life span in this study is not known, but offers a promising direction to explore in future studies. One limitation to our study is the possibility that there is a functional Lisinopril level for each DGRP line that would yield the maximum impact on life span and that this dose may or may not have the same impact on physical function. Furthermore, the onset of administration and the duration of administration adds another layer of complexity. Further studies are needed to dissect the intersect between onset and dose of Lisinopril in each line.
At a molecular level, aging is associated with changes in muscle fiber type and accumulation of protein aggregates (21), potentially leading to defects in physical performance. Our data suggest that the differential effect of Lisinopril on climbing speed, endurance, and strength in the three DGRP lines is driven by differences in the accumulation of protein aggregates in muscles. Morphologically, there are two major muscle types in adult Drosophila: fibrillar muscles, which are exclusively present as indirect flight muscles and provide power for oscillatory flight, and tubular muscles, such as the jump muscles and leg muscles, which are neurogenic and used for activities including climbing and the initiation of flight (15). Although we specifically concentrated on the flight muscles, protein aggregation appears to be a general contributor to the decline of adult muscle function. As such, future studies should assess the effects of Lisinopril treatment on protein aggregation in other muscle types. We also suggest assessment of protein aggregation in other locations, such as nervous or cardiac tissue, as this might provide additional insight into the variable effects of Lisinopril on traits such as life span.
Results from the RNA-seq experiment identified several genes that responded to Lisinopril treatment. Many of these have been implicated in some aspect of stress and immune responses. These include genes in the Turandot family, CHK kinases and genes involved in the humoral response to infection. Additional experiments will be required to determine the functional effects of these genes on the phenotypes examined. This experiment also identified genes whose expression in response to Lisinopril depended on genotype in an age-specific manner. Many of these genes are also involved in stress responses, suggesting that genetically based variation in the phenotypic response to drug treatment may depend on the extent to which stress response pathways are activated in different genotypes. Given the fact that stress responses have also been associated with protein aggregation (20,46,48), additional experiments directed at elucidating the interrelationships between Lisinopril, stress response, protein aggregation and life span offer a promising line of future research that could have direct application to personalizing medical treatment for patients taking this and related medications. Although these RNA-seq data are descriptive in nature, they indicate that the response to Lisinopril treatment depends on the age and genotype of the individual. In this case, genetically based differences in the transcriptional response to drug treatment may contribute to the phenotypic differences we observed.
With the number of humans older than 60 years expected to double over the next 40 years, lack of physical ability is a major public health issue. A major gap in our knowledge is the role that genetic variation plays in contributing to individual differences in age-related decline of physical ability and the response to treatment. The biological functions of many of the genes that responded to Lisinopril treatment are unknown. This reflects the broader fact, that for most organisms, the biological roles of many genes in the genome are unknown. In this study, we demonstrate that Drosophila are a strategic model to elucidate the functions of particular genes relevant to human health.
Funding
This study was supported by the Johns Hopkins Older Americans Independence Center National Institute on Aging (grants P30 AG021334, R21AG043284, R01AG046441 and K23 AG035005), and the Nathan Shock in Aging Scholarship Award (P.M.A.), and Maryland Technology Development Grant phase 1&2 (J.W. and P.M.A.). Fly stocks were made available from the Bloomington Stock Center and the Harvard RNAi project. FlyBase provided genetic information for Drosophila genes.
Conflict of Interest
None reported.
Supplementary Material
Acknowledgments
We thank the following undergraduates who helped with fly assays: Mehnaz Ali Khan, Priyanka Ochaney, Parsa Khosravian, Shiv Parmar, Laura Powell, Jeanice Hwang, Saiah Yates, Jason Sumpter, and Sean Cordova. We thank Hoku West-Foyle and Fernando Vonhoff for help with the confocal imaging. We thank Meagan Jezek for help with the real-time quantitative reverse transcriptase polymerase chain reaction.
References
- 1. Hogan M. Physical and cognitive activity and exercise for older adults: a review. Int J Aging Hum Dev. 2005;60:95–126. doi: 10.2190/PTG9-XDVM-YETA-MKXA [DOI] [PubMed] [Google Scholar]
- 2. Resnick B, Galik E, Dorsey S, Scheve A, Gutkin S. Reliability and validity testing of the physical resilience measure. Gerontologist. 2011;51:643–652. doi: 10.1093/geront/gnr016 [DOI] [PubMed] [Google Scholar]
- 3. Whitson HE, Duan-Porter W, Schmader KE, Morey MC, Cohen HJ, Colon-Emeric CS. Physical resilience in older adults: systematic review and development of an emerging construct. J Gerontol A Biol Sci Med Sci. 2016;71(4):489–495. doi: 10.1093/gerona/glv202 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Cluett C, Melzer D. Human genetic variations: Beacons on the pathways to successful ageing. Mech Ageing Dev. 2009;130:553–563. doi: 10.1016/j.mad.2009.06.009 [DOI] [PubMed] [Google Scholar]
- 5. Montgomery HE, Marshall R, Hemingway H, et al. Human gene for physical performance. Nature. 1998;393:221–222. doi: 10.1038/30374 [DOI] [PubMed] [Google Scholar]
- 6. Zajc Petranović M, Skarić-Jurić T, Smolej Narančić N, et al. Angiotensin-converting enzyme deletion allele is beneficial for the longevity of Europeans. Age (Dordr). 2012;34:583–595. doi: 10.1007/s11357-011-9270-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Dietze GJ, Henriksen EJ. Angiotensin-converting enzyme in skeletal muscle: sentinel of blood pressure control and glucose homeostasis. J Renin Angiotensin Aldosterone Syst. 2008;9:75–88. doi: 10.3317/jraas.2008.011 [DOI] [PubMed] [Google Scholar]
- 8. Buford TW, Manini TM, Hsu FC, et al. Angiotensin-converting enzyme inhibitor use by older adults is associated with greater functional responses to exercise. J Am Geriatr Soc. 2012;60:1244–1252. doi: 10.1111/j.1532-5415.2012.04045.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Carter CS, Onder G, Kritchevsky SB, Pahor M. Angiotensin-converting enzyme inhibition intervention in elderly persons: effects on body composition and physical performance. J Gerontol A Biol Sci Med Sci. 2005;60:1437–1446. doi: 10.1093/gerona/60.11.1437 [DOI] [PubMed] [Google Scholar]
- 10. Gray SL, LaCroix AZ, Aragaki AK, et al. ; Women’s Health Initiative Observational Study Angiotensin-converting enzyme inhibitor use and incident frailty in women aged 65 and older: prospective findings from the Women’s Health Initiative Observational Study. J Am Geriatr Soc. 2009;57:297–303. doi: 10.1111/j.1532-5415.2008.02121.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Shrikrishna D, Tanner RJ, Lee JY, et al. A randomized controlled trial of angiotensin-converting enzyme inhibition for skeletal muscle dysfunction in COPD. Chest. 2014;146:932–940. doi: 10.1378/chest.13-2483 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Sumukadas D, Band M, Miller S, et al. Do ACE inhibitors improve the response to exercise training in functionally impaired older adults? A randomized controlled trial. J Gerontol A Biol Sci Med Sci. 2014;69:736–743. doi: 10.1093/gerona/glt142 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Tinetti ME, Han L, Lee DS, et al. Antihypertensive medications and serious fall injuries in a nationally representative sample of older adults. JAMA Intern Med. 2014;174(4):588–595. doi: 10.1001/jamainternmed.2013.14764 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. George C, Verghese J. Polypharmacy and gait performance in community-dwelling older adults. J Am Geriatr Soc. 2017;65:2082–2087. doi: 10.1111/jgs.14957 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Coates D, Isaac RE, Cotton J, et al. Functional conservation of the active sites of human and Drosophila angiotensin I-converting enzyme. Biochemistry. 2000;39:8963–8969. doi: 10.1021/bi000593q [DOI] [PubMed] [Google Scholar]
- 16. Akif M, Georgiadis D, Mahajan A, et al. High-resolution crystal structures of Drosophila melanogaster angiotensin-converting enzyme in complex with novel inhibitors and antihypertensive drugs. J Mol Biol. 2010;400:502–517. doi: 10.1016/j.jmb.2010.05.024 [DOI] [PubMed] [Google Scholar]
- 17. Williams TA, Michaud A, Houard X, Chauvet MT, Soubrier F, Corvol P. Drosophila melanogaster angiotensin I-converting enzyme expressed in Pichia pastoris resembles the C domain of the mammalian homologue and does not require glycosylation for secretion and enzymic activity. Biochem J. 1996;318 (Pt 1):125–131. doi: 10.1042/bj3180125 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Mackay TF, Richards S, Stone EA, et al. The Drosophila melanogaster genetic reference panel. Nature. 2012;482:173–178. doi: 10.1038/nature10811 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Durham MF, Magwire MM, Stone EA, Leips J. Genome-wide analysis in Drosophila reveals age-specific effects of SNPs on fitness traits. Nat Commun. 2014;5:4338. doi: 10.1038/ncomms5338 [DOI] [PubMed] [Google Scholar]
- 20. Squier TC. Oxidative stress and protein aggregation during biological aging. Exp Gerontol. 2001;36:1539–1550. doi: 10.1016/S0531-5565(01)00139-5 [DOI] [PubMed] [Google Scholar]
- 21. Stefani M, Dobson CM. Protein aggregation and aggregate toxicity: new insights into protein folding, misfolding diseases and biological evolution. J Mol Med (Berl). 2003;81:678–699. doi: 10.1007/s00109-003-0464-5 [DOI] [PubMed] [Google Scholar]
- 22. Ayyadevara S, Balasubramaniam M, Suri P, et al. Proteins that accumulate with age in human skeletal-muscle aggregates contribute to declines in muscle mass and function in Caenorhabditis elegans. Aging (Albany NY). 2016;8:3486–3497. doi: 10.18632/aging.101141 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. AbdAlla S, Langer A, Fu X, Quitterer U. ACE inhibition with captopril retards the development of signs of neurodegeneration in an animal model of Alzheimer’s disease. Int J Mol Sci. 2013;14:16917–16942. doi: 10.3390/ijms140816917 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Gebre AK, Altaye BM, Atey TM, Tuem KB, Berhe DF. Targeting renin-angiotensin system against Alzheimer’s disease. Front Pharmacol. 2018;9:440. doi: 10.3389/fphar.2018.00440 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Ja WW, Carvalho GB, Mak EM, et al. Prandiology of Drosophila and the CAFE assay. Proc Natl Acad Sci U S A. 2007;104:8253–8256. doi: 10.1073/pnas.0702726104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Seroude L, Brummel T, Kapahi P, Benzer S. Spatio-temporal analysis of gene expression during aging in Drosophila melanogaster. Aging Cell. 2002;1:47–56. doi: 10.1046/j.1474-9728.2002.00007.x [DOI] [PubMed] [Google Scholar]
- 27. Lee JH, Bassel-Duby R, Olson EN. Heart- and muscle-derived signaling system dependent on MED13 and Wingless controls obesity in Drosophila. Proc Natl Acad Sci U S A. 2014;111:9491–9496. doi: 10.1073/pnas.1409427111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Fried LP, Tangen CM, Walston J, et al. ; Cardiovascular Health Study Collaborative Research Group Frailty in older adults: evidence for a phenotype. J Gerontol A Biol Sci Med Sci. 2001;56:M146–M156. doi: 10.1093/gerona/56.3.m146 [DOI] [PubMed] [Google Scholar]
- 29. Liu H, Graber TG, Ferguson-Stegall L, Thompson LV. Clinically relevant frailty index for mice. J Gerontol A Biol Sci Med Sci. 2014;69:1485–1491. doi: 10.1093/gerona/glt188 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Bjedov I, Toivonen JM, Kerr F, et al. Mechanisms of life span extension by rapamycin in the fruit fly Drosophila melanogaster. Cell Metab. 2010;11:35–46. doi: 10.1016/j.cmet.2009.11.010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Clancy DJ, Gems D, Harshman LG, et al. Extension of life-span by loss of CHICO, a Drosophila insulin receptor substrate protein. Science. 2001;292:104–106. doi: 10.1126/science.1057991 [DOI] [PubMed] [Google Scholar]
- 32. Giannakou ME, Goss M, Jünger MA, Hafen E, Leevers SJ, Partridge L. Long-lived Drosophila with overexpressed dFOXO in adult fat body. Science. 2004;305:361. doi: 10.1126/science.1098219 [DOI] [PubMed] [Google Scholar]
- 33. Promislow DEL, Tatar M, Pletcher S, Carey JR. Below-threshold mortality: implications for studies in evolution, ecology and demography. J Evol Biol. 1999;12(2):314–328. doi: 10.1046/j.1420-9101.1999.00037.x [DOI] [Google Scholar]
- 34. Demontis F, Piccirillo R, Goldberg AL, Perrimon N. Mechanisms of skeletal muscle aging: insights from Drosophila and mammalian models. Dis Model Mech. 2013;6:1339–1352. doi: 10.1242/dmm.012559 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet journal; Next Generation Sequencing Data Analysis. 2011;17(1):10–12. doi: 10.14806/ej.17.1.200 [DOI] [Google Scholar]
- 36. Li H, Durbin R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics. 2010;26(5):589–595. doi: 10.1093/bioinformatics/btp698 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Dobin A, Davis CA, Schlesinger F, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21. doi: 10.1093/bioinformatics/bts635 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Anders S, Pyl PT, Huber W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–169. doi: 10.1093/bioinformatics/btu638 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–140. doi: 10.1093/bioinformatics/btp616 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Robinson MD, Oshlack A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 2010;11:R25. doi: 10.1186/gb-2010-11-3-r25 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Huang dW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57. doi: 10.1038/nprot.2008.211 [DOI] [PubMed] [Google Scholar]
- 42. Falconer DS. Selection in different environments: effects on environmental sensitivity (reaction norm) and on mean performance. Genet Res. 1990;56(1):57–70. doi: 10.1017/S0016672300028883 [DOI] [Google Scholar]
- 43. Azad P, Zhou D, Russo E, Haddad GG. Distinct mechanisms underlying tolerance to intermittent and constant hypoxia in Drosophila melanogaster. PLoS One. 2009;4:e5371. doi: 10.1371/journal.pone.0005371 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Demontis F, Piccirillo R, Goldberg AL, Perrimon N. The influence of skeletal muscle on systemic aging and lifespan. Aging Cell. 2013;12:943–949. doi: 10.1111/acel.12126 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Vasconcellos LR, Dutra FF, Siqueira MS, et al. Protein aggregation as a cellular response to oxidative stress induced by heme and iron. Proc Natl Acad Sci U S A. 2016;113:E7474–E7482. doi: 10.1073/pnas.1608928113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Hu J, Igarashi A, Kamata M, Nakagawa H. Angiotensin-converting enzyme degrades Alzheimer amyloid beta-peptide (A beta); retards A beta aggregation, deposition, fibril formation; and inhibits cytotoxicity. J Biol Chem. 2001;276:47863–47868. doi: 10.1074/jbc.M104068200 [DOI] [PubMed] [Google Scholar]
- 47. Hemming ML, Selkoe DJ. Amyloid beta-protein is degraded by cellular angiotensin-converting enzyme (ACE) and elevated by an ACE inhibitor. J Biol Chem. 2005;280:37644–37650. doi: 10.1074/jbc.M508460200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Ogen-Shtern N, Ben DT, Lederkremer GZ. Protein aggregation and ER stress. Brain Res. 2016;1648(Pt B):658–666. doi: 10.1016/j.brainres.2016.03.044 [DOI] [PubMed] [Google Scholar]
- 49. Zhou BB, Elledge SJ. The DNA damage response: putting checkpoints in perspective. Nature. 2000;408:433–439. doi: 10.1038/35044005 [DOI] [PubMed] [Google Scholar]
- 50. Ekengren S, Hultmark D. A family of Turandot-related genes in the humoral stress response of Drosophila. Biochem Biophys Res Commun. 2001;284:998–1003. doi: 10.1006/bbrc.2001.5067 [DOI] [PubMed] [Google Scholar]
- 51. Brun S, Vidal S, Spellman P, Takahashi K, Tricoire H, Lemaitre B. The MAPKKK Mekk1 regulates the expression of Turandot stress genes in response to septic injury in Drosophila. Genes Cells. 2006;11:397–407. doi: 10.1111/j.1365-2443.2006.00953.x [DOI] [PubMed] [Google Scholar]
- 52. Coelho VA, Probst VS, Nogari BM, et al. Angiotensin-II blockage, muscle strength, and exercise capacity in physically independent older adults. J Phys Ther Sci. 2016;28:547–552. doi: 10.1589/jpts.28.547 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Linz W, Heitsch H, Schölkens BA, Wiemer G. Long-term angiotensin II type 1 receptor blockade with fonsartan doubles lifespan of hypertensive rats. Hypertension. 2000;35:908–913. doi: 10.1161/01.HYP.35.4.908 [DOI] [PubMed] [Google Scholar]
- 54. Kumar S, Dietrich N, Kornfeld K. Angiotensin converting enzyme (ACE) inhibitor extends caenorhabditis elegans life span. PLoS Genet. 2016;12:e1005866. doi: 10.1371/journal.pgen.1005866 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Santos EL, de Picoli SK, da Silva ED, et al. Long term treatment with ACE inhibitor enalapril decreases body weight gain and increases life span in rats. Biochem Pharmacol. 2009;78(8):951–958. doi: 10.1016/j.bcp.2009.06.018 [DOI] [PubMed] [Google Scholar]
- 56. van de Vijver PL, van BD, Westendorp RG. Early and extraordinary peaks in physical performance come with a longevity cost. Aging (Albany NY). 2016;8(8):1822–1829. doi: 10.18632/aging.101023 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Hall KS, Cohen HJ, Pieper CF, et al. Physical performance across the adult life span: correlates with age and physical activity. J Gerontol A Biol Sci Med Sci. 2017;72:572–578. doi: 10.1093/gerona/glw120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Ederer KA, Jin K, Bouslog S, et al. Age- and genotype-specific effects of the angiotensin-converting enzyme inhibitor lisinopril on mitochondrial and metabolic parameters in Drosophila melanogaster. Int J Mol Sci. 2018;19(11):3351– 3368. doi: 10.1038/35044005 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.