Abstract
The outbreak of Coronavirus Disease-2019 (COVID-19) caused by Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) has threatened health worldwide. As of the end of 2020, there were nearly 10 million confirmed cases and nearly 5 million deaths associated with COVID-19. Rapid and early laboratory diagnosis of COVID-19 is the main focus of treatment and control. Molecular tests are the basis for confirmation of COVID-19, but serological tests for SARS-CoV-2 are widely available and play an increasingly important role in understanding the epidemiology of the virus and in identifying populations at higher risk for infection. Point-of-care tests have the advantage of rapid, accurate, portable, low cost and non-specific device requirements, which provide great help for disease diagnosis and detection. This review will discuss the performance of different laboratory diagnostic tests and platforms, as well as suitable clinical samples for testing, and related biosafety protection. This review shall guide for the diagnosis of COVID-19 caused by SARS-CoV-2.
Keywords: CoVs, SARS-CoV-2, COVID-19, nCoV-19, Laboratory diagnosis
1. Introduction
A novel virus caused an outbreak of pneumonia began from Wuhan, Hubei Province, China. The virus involved in the event was identified as the seventh coronavirus, becoming the third zoonotic human coronaviruses (HCoV) of the century, and posing serious threats to international health. The International Committee on Taxonomy of Viruses (ICTV) has announced that the novel coronavirus is officially classified as severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). The World Health Organization (WHO) announced that the official name of the disease caused by the virus is Coronavirus Disease 2019 (COVID-19).
Coronaviruses (CoVs) are enveloped viruses with a single-stranded, positive-sense RNA genome, which is the largest discovered genome of RNA virus [1]. Before the appearance of SARS-CoV-2, 6 HCoVs have been discovered by researchers, including HCoV-229E, HCoV-HKU1, HCoV-NL63, HCoV-OC43, Severe Acute Respiratory Syndrome (SARS) CoV and Middle East Respiratory Syndrome (MERS) CoV [2]. There are currently 7 CoVs (including SARS-CoV-2) that can cause human respiratory diseases, but to date, only SARS-CoV, MERS-CoV, and SARS-CoV-2 have caused a large outbreak with high mortality.
The ongoing continuation and spread of the COVID-19 pose challenges for public health control. On 30 January 2020, WHO has declared the prevalence of COVID-19 is a Public Health Emergency of International Concern (PHEIC). More than 3.5 million cases of laboratory-confirmed COVID-19 and 250,000 deaths worldwide have been reported globally in early May 2020 [3]. By the end of June 2020, there were nearly 10 million confirmed cases and nearly 5 million deaths of COVID-19, and the number of positive cases and deaths is still rising, posing a serious threat to international health [4].
During the COVID-19 pandemic, virus detection based on culture is an important tool for virus pathogenesis research, but the culture cycle is too long, so it is not suitable for the rapid detection of viruses related to acute infectious diseases. Nucleic acid tests can rapidly and sensitively identify pathogens in suspected or confirmed COVID-19 patients, but large amounts of genetic variation in RNA viruses, and mismatches of primers, probes, and target sequences may result in reduced detection performance and false-negative results [5], [6]. Suspects with positive chest computed tomography scans may present negative results for SARS-CoV-2 by reverse-transcriptase polymerase chain reaction (RT-PCR) [6]. In addition, nsopharyngeal samples that contain weak viral RNA were identified as negative by a point-of-care test, Abbott ID NOW [7]. Therefore, the interpretation of nucleic acid test results must be careful and negative results cannot be excluded. Serological surveys can aid the investigation of an ongoing outbreak and retrospective assessment of the attack rate or extent of an outbreak. Detection of SARS-CoV-2 IgM and IgG antibody in vivo as a supplement to molecular diagnostic methods [8]. Genomic sequencing has been playing an irreplaceable role in the identification of emerging viruses [9]. In this review, we describe the current laboratory diagnosis of SARS-CoV-2 infection. It is hoped that these information will contribute to the rapid and accurate laboratory diagnosis and treatment of COVID-19 in clinical laboratory.
2. Origin and transmission of SARS-CoV-2
SARS-CoV-2 is a beta-CoV belong to the subgenus sarbecovirus, Orthocoronavirinae subfamily based on sequence analysis [10]. SARS-CoV-2 binds to the same cell entry receptor, Angiotensin Converting Enzyme 2 (ACE2), to infect humans, as SARS-CoV [8]. SARS-CoV-2 ranges between 50 and 200 nm in diameter and contains a 29,881 bp genome [11]. SARS-COV-2 genome encodes at least four structural proteins called nucleocapsid (N), spikes (S), envelope (E), and membrane (M), respectively. The N protein holds the viral genome, while S, E and M construct the viral envelope. Among them, S protein mediates virus entry into the host cell and determines to a certain degree the host range during virus infection [12].
Upon virus genome sequencing, SARS-CoV-2 shares 88% identity to two bat SARS-like CoVs (bat-SL-CoVZC45 and bat-SL-CoVZXC21) and 79% identity to SARS-CoV, but only 50% identity to MERS-CoV [13]. The genome of SARS-CoV-2 was closely related to that of Bat CoV RaTG13, showing 96.2% overall genomic sequence identity [8], indicating that human SARS-CoV-2 and bat CoV may share the same ancestor. It was reported CoVs identified in pangolins with 90% sequence identity to SARS-CoV-2 by protein sequence alignments and phylogenetic analysis [14], [15], suggesting pangolins are the most likely intermediate hosts for SARS-COV-2. However, the outcome of phylogeny analyses does not necessarily support the view that pangolin is the exact intermediate host of SARS-CoV-2, and other animals may also serve as intermediate hosts [16]. A recent study reported that similar ACE2 receptor residues were found in some species other than pangolin, such as turtles and snakes, which supplies more possibilities for alternative intermediate hosts [17]. In sum, host ranges and animal reservoirs of SARS-CoV-2 still need to be explored.
SARS-CoV-2 is mainly transmitted through contact, respiratory droplets and the potential route of fecal-oral. The estimated reproductive number (R0) of SARS-CoV-2 ranges from 2.2 to 5.7 [18], [19], [20], while the reported R0 of SARS-CoV is around 3 [21]. It is speculated that the primary virus replication occurs in the mucosal epithelium of the upper respiratory tract (pharynx and nasal cavity), and further multiplies in the mucosa of the lower respiratory tract and gastrointestinal tract, causes mild viremia [22]. A study based- hospital survey found that the maximum propagation distance of aerosols containing SARS-CoV-2 virions might be 4 m from the patients with COVID-19 [23]. Neeltje et al. revealed that SARS-CoV-2 aerosols remained infectious in the tissue culture experiments, and the infectivity only decreased slightly during a 3-hour observation period [24]. Several recent studies have reported that SARS-COV-2 was detected in stool samples [25], [26], [27]. Although these evidence indicate that SARS-CoV-2 may also be an enterovirus that can be transmitted through the fecal-oral route, these discoveries are based on the situation of very few patients and more researches are warranted.
3. Clinical specimens for SARS-CoV-2 testing
Similar to other infectious diseases, appropriate specimen collection is the key step in the laboratory diagnosis of COVID-19. Acceptable specimens include upper respiratory tract specimens, lower respiratory tract specimens, stool specimens, whole blood specimens, and serum specimens, and the respiratory secretions is the most frequently sample for diagnosis [28]. Currently, SARS-CoV-2 has been detected in nasopharyngeal swabs [29], [30], oropharyngeal swabs [25], [30], throat swabs [29], [31], sputum [29], [31], [32], bronchoalveolar lavage fluid (BALF) [10], [11], [33], whole blood [25], serum [25], stool [25], [26], [27], urine [34], [35], saliva [36], [37], [38], rectal swabs [34], [39] and conjunctival swabs [40], [41].
With limited understanding of COVID-19, it is difficult to exclude SARS-CoV-2 infection based on a single negative PCR result, especially when testing was used for upper respiratory tract specimens. Collection and detection of lower respiratory tract specimens are strongly recommended even if the upper respiratory tract specimens are negative, especially in patients with severe or progressive conditions [42]. ACE2 is mainly distributed in alveolar type II epithelial cells [17], suggesting lower respiratory tract specimens (including sputum, tracheal aspirates, BALF) may contain high viral RNA loads. Yu et al. compared the average viral load in sputum, throat swabs and nasal swabs from 127 confirmed or suspected COVID-19 patients. The study found that the average viral load in sputum (17429 ± 6920 copies/test) was obviously higher than in nasal swabs (651 ± 501 copies/test) and throat swabs (2552 ± 1965 copies/test) [31]. Besides, a high viral load of 108 copies per milliliter was detected in the sputum of an asymptomatic patient five days after symptoms onset in Germany [32]. These findings indicate a higher viral load in lower respiratory tract samples. Zhang et al. analyzed the virus dynamics in oral and anal swabs of 16 COVID-19 patients who had been treated for about 10 days. Most of the cases were detected positive to SARS-CoV-2 from oral swabs (8/10, 80%) on the day of the first sampling. However, 5 days after the first sampling, the positive rate of the anal swab (6/8, 75%) was higher than that of the oral swab (4/8, 50%) by nucleic acid test, indicating a change from more oral swabs positive during the early period to more anal swabs positive during the later period [25].
In addition to the most common respiratory specimens, it should be noted that SARS-CoV-2 has been frequently detected in non-respiratory specimens. It was reported that SARS-CoV-2 was detected in the urine sample from one confirmed COVID-19 patient, although no urinary irritation was found [43]. Recently, SARS-CoV-2 was frequently detected in saliva samples of COVID-19 patients during infection [36], [37], [38]. SARS-CoV-2 was identified in the saliva specimens from 11 out of 12 COVID-19 patients by RT-PCR. The viral load monitoring of serial saliva generally showed a downward trend after hospitalization [36]. In another study, SARS-CoV-2 RNA was detected in the posterior oropharyngeal saliva of 20 (87%) patients. The viral load in saliva was highest in the first week after symptom onset and decreased over time [37]. Jonathan et al. simultaneously assessed posterior oropharyngeal saliva and nasopharyngeal swab specimens for the detection of SARS-CoV-2 using an automated Xpert® Xpress SARS-CoV-2 assay. In this study, 84.5% (49/58) patients tested positive in both saliva and nasopharyngeal swab, 5.2% (3/58) patients tested positive in saliva only and 10.3% (6/58) patients tested positive in nasopharyngeal swab only [38]. These studies indicated that saliva can be used as a promising non-invasive sample for diagnosis, monitoring and control of COVID-19. Furthermore, tear and conjunctival swab samples from 2 out of 60 COVID-19 patients were detected positive to SARS-CoV-2 by RT-PCR [41]. However, the virus was not successfully isolated and cultured in the conjunctival secretion, and more studies are needed to be evaluated.
Currently, the virus has not been detected in cerebrospinal fluid, pericardial effusion, peritoneal effusion, posterior fornix, joint fluid, peritoneal exudate, semen, female reproductive tract secretions and other samples. Although no virus has been detected in these specimens until now, it is also recommended that these be treated as infectious specimens. Further investigations are needed to identify whether these specimens may spread the virus.
4. Type of nucleic acid tests for SARS-CoV-2 detection
According to SARS-CoV-2 genomes released on public databases, specific primers and probes designed for the target genes can realize the laboratory diagnosis of COVID-19. So far, the gene targets were used to detect SARS-CoV-2 include nucleocapsid (N), envelope (E), spike (S), RNA-dependent RNA polymerase (RdRP) and open reading frame1ab (ORF1ab) genes [44]. As the gold standard test for SARS-CoV-2 identification, real time quantification RT-PCR (qRT-PCR) is the routine confirmation test recommended by WHO. However, due to the time-consuming process, the requirements of expensive equipment and biosafety conditions, this test is not suitable for point-of-care diagnosis. Point-of-care tests are not only suitable for clinical laboratories, but also can be performed by trained non-laboratory personnel in patient care facilities, such as physicians’ offices or Emergency Departments, making the diagnostic test of SARS-CoV-2 closer to the patient. The number of commercially available point-of-care tests for SARS-CoV-2 is increasing. A comparison of different nucleic acid amplification of SARS-CoV-2, including Lab-based tests and point-of-care tests, is shown in Table 1 .
Table 1.
Advantages and disadvantages of the laboratory diagnostic methods for SARS-CoV-2.
| Diagnostic approach | Method | Testing scenarios | Advantages | Disadvantages | Reference |
|---|---|---|---|---|---|
| Neutralization tests | VNT and PVNT | BSL-2 or BSL-3 laboratories, pathogen laboratories | Authoritative, simple, low cost, reliable, high sensitivity | Time-consuming, long-period, laborious, perform in BSL-3 or BSL-2 lab | [78], [80], [94], [95] |
| PCR | qRT-PCR Nested RT-PCR |
BSL-2 laboratories, public health institutes, quarantine depots BSL-2 laboratories, prefectural and municipal public health institutes, quarantine depots |
High specificity, not require expensive equipment, time-saving High sensitivity, specificity was higher than that of RT-PCR, suitable for detect low-copy-number viruses, time-saving |
Complex pretreatment steps, requires skillful, false negative Complex pretreatment steps, requires skillful, manpower, the second PCR amplification may caused cross-contamination |
[104], [105], [51], [52] |
| ddPCR | BSL-2 laboratories, public health institutes, quarantine depots | Quantitative, sensitive, suitable for detect samples with low viral load, independent of a traditional standard curve | Susceptible to exogenous contamination, expensive than qRT-PCR, calibrant materials need to be defined | [54], [55] | |
| Nanoparticles -based amplification | BSL-2 laboratories, environmental testing institutions | High sensitivity, adopted in fully-automated RNA extraction systems, excellent RNA binding performances | Complex pretreatment steps, requires skillful, expensive than qRT-PCR, with the risk of photobleaching | [57], [59], [60] | |
| RT-LAMP | Basic laboratories, community nursing sites | Time-saving, thermostatic, sensitive, user-friendly, sophisticated equipment-free | Easy to be contaminated and cause false-positive, nonspecific amplification cannot be easily identified, requires skillful | [56], [57], [58] | |
| Portable benchtop-sized analyzers | Clinical laboratories, physicians' office, emergency departments | Automatic, portable, rapid, not requires trained staff | Inconsistent performance, may lack sensitivity in weakly positive samples | [62], [63], [64], [65], [66], [67], [68] | |
| Immunological diagnostic | ELISA | Clinical laboratories, public health institutes | Quantitative detection, simple, a low risk of infection, convenient, stable reagent |
Time-consuming, low sensitivity, cross-reactivity, expensive monoclonal antibody, low-throughput | [78], [79] |
| IFA | Clinical laboratories, pathogen laboratories, public health institutes | Avoid the interference of endogenous biotin and contamination of antigens in the blood | Non-specific fluorescence, subjective, low-throughput, time-consuming | [78], [80]. | |
| CLIA | Clinical laboratories, public health institutes | Automatic, rapid, quantitative, high sensitivity, broad linear range, stable results |
Sophisticated instruments, high requirements for equipment and environment, not suitable for detect whole blood samples, | [81], [82], [83], [84] | |
| LFA | Clinical laboratories, physicians' offices, emergency departments, community service stations | Rapid, convenient, on-site screening, inexpensive, small sample volume |
Low sensitivity, cross-reactivity, inconsistent performance, not suitable for early diagnosis, low-throughput | [85], [86], [87], [88] | |
| Microarray and microfluidic chip | Clinical laboratories, emergency departments, community service stations | Small size, high sensitivity, automatic, high-throughput, portable | Core technologies lack norms and standards, high cost, nonspecific binding of proteins | [89], [90], [91], [92] | |
| Genome sequencing | Metatranscriptomic sequencing | BSL-2 laboratories, genetic testing centres, research laboratories | Simple, reduce the cost, does not claim a reference sequence | Increase cost, sophisticated instruments, insufficient coverage and depth |
[11], [13] |
| Nanopore targeted sequencing | BSL-2 laboratories, genetic testing centres, research laboratories | Broad detection range, rapid turnaround time, long-read, high-accuracy, monitor the variation | Increase cost, sophisticated instruments, requires skillful |
[97] |
|
| Amplicon se quencing | BSL-2 laboratories, genetic testing centres, research laboratories | Convenient, high sensitivity, suitable for detect samples with low viral load, economical | Sophisticated instruments, not be used to sequence highly diverse or recombinant viruses |
[96] |
|
| Hybrid capture -based sequencing | BSL-2 laboratories, genetic testing centres, research laboratories | High sensitivity, suitable for detect intra-individual variations | Sophisticated instruments, not be used to sequence highly diverse or recombinant viruses |
[96] |
4.1. Manual laboratory-based nucleic acid tests
4.1.1. Real-time quantification RT-PCR
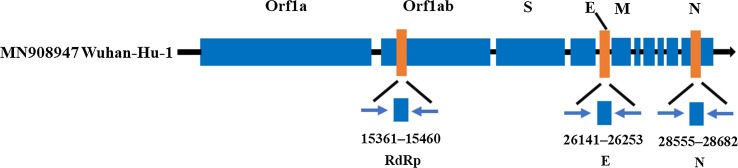
The WHO website provides several qRT-PCR protocols for detecting SARS-CoV-2 in different countries [44]. The gene targets for detection SARS-CoV-2 are different in China (ORF1ab and N genes), Germany (RdRP, E and N genes), United States (three targets in N gene), France (two targets in RdRP), Thailand (N gene), and Japan (pancorona and multiple targets, spike protein). RT-PCR tests/primers for SARS-CoV-2 in different institutions are shown in Table 2 . Relative positions of amplified targets on SARS-CoV-2 genome are shown in Fig. 1 . The centers for disease control and prevention (CDC) established a RT-PCR panel for specific detection SARS-CoV-2 and universal detection SARS-like beta-CoVs. Three sets of different primers were designed for the N gene, one set of primers/probes was used universally detect all beta-CoVs, while the other two were specific to identify SARS-CoV-2. The confirmation of COVID-19 must be positive for all three targets [45]. Two nucleic acid tests for detection RdRp and E genes of SARS-CoV-2, SARS-CoV and bat-like beta-CoVs were developed by Charite, Germany. Both tests are positive could enter the next step of the test, namely the SARS-CoV-2 specific RT-PCR test for RdRp gene [46].
Table 2.
RT-PCR tests/primers for SARS-CoV-2 in different institutions.
| Institute | Gene target | Probe (5′-3′) | Former primer (5′-3′) | Reverse primer (5′-3′) | Reference |
|---|---|---|---|---|---|
| China CDC | ORF1ab gene | FAM-CCGTCTGCGGTATGT GGA AAGGTTATGG-BHQ1 | CCCTGTGGGTTTTACACTTAA | ACGATTGTGCATCAGCTGA | [106] |
| N gene | FAM-TTGCTGCTGCTTGACAGA TT-TAMRA | GGGGAACTTCTCCTGCTAGAAT | CAGACATTTTGCTCTCAAGCTG | ||
| US CDC | N1 target | FAM-ACCCCGCATTAC GTT TGGTGGACC-BHQ1 | GAC CCC AAA ATC AGC GAA AT | TCT GGT TAC TGC CAG TTG AAT CTG | [45] |
| N2 target | FAM-ACAATTTGCCCCCAGCGC TTCAG-BHQ1 | TTA CAA ACA TTG GCC GCA AA | GCG CGA CAT TCC GAA GAA’ | ||
| N3 target | FAM-AYCACATTGGCACCCGCA ATCCTG-BHQ1 | GGG AGC CTT GAA TAC ACC AAA A | TGT AGC ACG ATT GCA GCA TTG | ||
| France Pasteur Institute | RdRP1 target | HEX-AGATGTCTTGTGCTGCCG GTA-BHQ1 | ATGAGCTTAGTCCTGTTG | CTCCCTTTGTTGTGTTGT | [107] |
| RdRP2 target | FAM-TCATACAAACCACGCCAG G-BHQ1 | GGTAACTGGTATGATTTCG | CTGGTCAAGGTTAATATAGG | ||
| Japan National Institute of Infectious Disease | N gene | FAM-ATGTCGCGCATTGGCATG GA-BHQ | AAATTTTGGGGACCAGGAAC | TGGCAGCTGTGTAGGTCAAC | [108] |
| Germany Charité | RdRP gene | FAM-CAGGTGGAACCTCATCAG GAGATGC-BBQ | GTGARATGGTCATGTGTGGCGG | CARATGTTAAASACACTATTAGCATA | [46] |
| E gene | FAM-ACACTAGCCATCCTTACTGCGCTTCG-BBQ | ACAGGTACGTTAATAGTTAATAGCGT | ATATTGCAGCAGTACGCACACA | ||
| Thailand National Institute of Health | N gene | FAM-CAACTGGCAGTAACCA-BQH1 | CGTTTGGTGGACCCTCAGAT- | CCCCACTGCGTTCTCCATT | [109] |
| Hong Kong University | ORF1b-nsp14 gene | FAM-TAGTTGTGATGCWATCATGACTAG-TAMRA | TGGGGYTTTACRGGTAACCT | AACRCGCTTAACAAAGCACTC | [110] |
| N gene | FAM-GCAAATTGTGCAATTTGCGG-TAMRA | TAATCAGACAAGGAACTGATTA | CGAAGGTGTGACTTCCATG |
Fig. 1.
The diffferent positions of amplicon targets on SARS-CoV-2 genome. Wuhan-Hu-1 genome (GenBank MN908947) as a reference. ORF: open reading frame; RdRp: RNA-dependent RNA polymerase gene; S: spike protein gene; E: envelope protein gene; M: membrane protein gene; N: nucleocapsid protein gene.
Although various institutions have developed different protocols for SARS-CoV-2 testing, it remains unclear whether the results from nucleic acid tests based on different targets are comparable. Chantal et al. [47] used RNA transcripts isolated from a COVID-19 patient to compare the analytical sensitivities of four qRT-PCR assays established in the United States, Germany, Hong Kong and China, respectively. The study found that SARS-CoV-2 could be detected in all primer-probe sets applied in the qRT-PCR tests, but significant discrepancy was observed in the detection limit and the ability to identify negatives and positives with a lower viral load. The primer-probe sets with the highest sensitivity were found in E-Sarbeco (Germany), 2019-nCoV_N1 (United States), and HKU-ORF1 (Hong Kong), but the lowest sensitivity was observed in RdRp-SARSr (Germany), which may be related to the mismatch in the reverse primer [47]. Besides, Konrad et al. compared qRT-PCR tests in different PCR systems and a commercial reagent using nasopharyngeal swab or sputum samples from COVID-19 patients in Germany [48]. When the same primers and probes were used, the distinctions in analytical sensitivities between different PCR systems were also observed. They found that the E gene target was more sensitive than the RdRp target when using a one-step qRT-PCR system. However, the high background of E gene target hindered the clear evaluation of the test, and further optimization of E gene assay may improve the sensitivity.
4.1.2. Nested RT-PCR
Real-time nested RT-PCR assay that connects the time-saving real-time instruments with high sensitivity of nested PCR has proven to be suitable for detecting low-copy-number SARS coronaviruses that present in the early stage of disease [49], [50]. In the early days of the outbreak, the detection of SARS-CoV-2 by nested RT-PCR method has been verified in Japan. As of 8 February 2020, the method developed has successfully identified 25 positive patients in Japan [51]. Recently, Ji et al. designed a one-step nested real-time RT-PCR (OSN-qRT-PCR) assay for targeting SARS-CoV-2 ORF1ab and N genes. The sensitivity of the assay was 1 copy/test and 10-fold higher than that of a commercial qRT-PCR assay (10 copies/test). Among 181 clinical samples, 14 samples with qRT-PCR-negative were confirmed by OSN-qRT-PCR. In addition, 7 samples with qRT-PCR- positive in the gray zone were confirmed to positive by OSN-qRT-PCR [52]. Compared with the qRT-PCR kit, nested RT-PCR analysis showed higher sensitivity and specificity, indicating that it is more suitable for clinical application to detect SARS-CoV-2 in cases with low viral load. However, nested RT-PCR can cause laboratory cross-contamination, which may lead to false-positive results [53].
4.1.3. Droplet digital PCR
Droplet digital PCR (ddPCR) has been shown to improve the lower LOD, sensitivity and accuracy in detecting SARS-CoV-2 [54], [55]. Suo et al. analyzed the feasibility of ddPCR for SARS-CoV-2 RNA detection compared with qRT-PCR using the same primer/probe sets issued by China CDC targeting ORF1ab or N gene. 26 COVID-19 patients with negative RT-PCR results were confirmed positive by ddPCR. The sensitivity and accuracy were improved from 40% and 47% for RT-PCR to 94% and 95% for ddPCR, respectively. Within 5–12 days after discharge, 6/14 patients (42.9%) were demonstrated to positive by ddPCR [55]. The study demonstrated that ddPCR could largely reduce the false negatives results caused by qRT-PCR. The team further stringently analyzed the performance of ddPCR and qRT-PCR using 8 primer/probe sets with the same samples and conditions. The results showed that all the 8 primers/probes used in qRT-PCR were unable to significantly distinguish positive and negative at low viral load (10−4 dilution). False-positive results of qRT-PCR tests of US CDC-N1, N2 and China CDC-N primers/probes sets were discovered. In contrast, the overall performance of ddPCR was significantly better than qRT-PCR, especially for samples with low viral load [54]. However, ddPCR also has some shortcomings. To ensure commutability between molecular diagnostic laboratories, gold standards or calibrant materials still need to be precisely defined. Besides, ddPCR is currently more costly than qRT-PCR for each test performed using dedicated instruments and consumables.
4.2. Rapid and point-of-care nucleic acid tests
4.2.1. Loop-mediated isothermal amplification
Loop-mediated isothermal amplification (LAMP) has the advantages of rapid amplification at a single temperature, which is efficient in the rapid and reliable diagnosis of CoVs. Zhang et al. designed full LAMP primers targeting the 5′ region of the ORF1a and N genes of SARS-CoV-2 and detected by a visual, colorimetric RT-LAMP alongside a commercial RT-PCR assay. Six of seven samples exhibited visible color change suggesting positive amplification, while the single sample remained pink color and was confirmed negative. The colorimetric RT-LAMP analysis was 100% consistent with RT-PCR results across a range of Cq values, and matched with RT-PCR in the field and point-of-care settings without sophisticated instrumentation [56]. Yu et al. [57] developed an isothermal LAMP-based detection method for ORF1ab gene –(isothermal LAMP-based method for COVID-19, ILACO). ILACO showed species-specificity by sequence comparison of 11 respiratory viruses (including 7 similar CoVs, 2 influenza viruses and 2 normal CoVs). In addition, iLACO's sensitivity was comparable to that of Taqman-based RT-PCR, which was able to detect as low as 10 copies of SARS-CoV-2. Besides. Mohamed et al. described a highly sensitive, point-of-care test based on LAMP and nested-like amplification assay, rapid isothermal amplification assay (Penn-RAMP) [58]. The sensitivity of RAMP was 10 times better than that of LAMP and RT-PCR for testing purified targets, and 100 times better than that of LAMP and RT-PCR for samples with minimally processed. This method is suitable for use at home, point-of-care and in the clinic with the least trained using and the least instrumentation. It has the potential to reduce false negatives from routine nucleic acid tests.
4.2.2. Nanoparticles-based amplification
Nanoparticles have been introduced to the nucleic acid amplification system to enhance the sensitivity and specificity of SARS-CoV-2 detection [57], [59], [60]. Parikshit et al. designed a naked-eye colorimetric test based on gold nanoparticles (AuNPs) with thiol-modified antisense oligonucleotides (ASOs) targeting the SARS-CoV-2 N-gene. The limit of detection (LOD) was found to be 0.18 ng/μL of SARS-CoV-2 viral load [60]. In addition, a LOD of 12 copies/test was observed in a one-step nanoparticles-based biosensor (NBS) coupled with RT-LAMP [59]. Among laboratory-confirmed COVID-19 patients, the analytical sensitivity of SARS-CoV-2 was 100% (33/33) in the oropharynx swab specimens, and the specificity of the assay was also 100% (96/96) when analyzed the RNA templates from non-COVID-19 patients. The unique properties of nanoparticles give them an advantage over traditional methods that are usually expensive and laborious. Nanoparticles-based amplification is a promising method for diagnosing SARS-CoV-2 infection in first-line clinical laboratories, especially in areas with challenged medical resources. However, Nanoparticles-based amplification is more expensive than qRT-PCR and the pretreatment steps are complex. Beside, as with any system using traditional organic carriers, the risk of photobleaching may lead to reduced sensitivity and false-negative results [61].
4.2.3. Portable benchtop-sized analyzers
The automated molecular diagnostic platform demonstrates a highly sensitive, powerful and accurate method for the rapid identification of SARS-CoV-2. Even in a laboratory without PCR training or point-of-care testing, the assay also can achieve rapid decisions and technological innovation. The inconsistent performance of the different portable benchtop-sized analyzers for SARS-CoV-2 detection was reported.
Benoit et al. evaluated the QIAstat-Dx Respiratory SARS-CoV-2 Panel for SARS-CoV-2 detection. This platform showed a comparable sensitivity to RT-PCR with a LOD at 1000 copies/mL. In 69 clinical samples analysis, the overall consistent percentage of QIAstat-DX SARS and RT-PCR recommended by WHO was 97%, with a specificity of 93% (27/29) and a sensitivity of 100% (40/40). No cross-reactions of other respiratory viruses or bacteria were observed in this assay [62]. The results indicated that QIAstat-Dx Respiratory SARS-CoV-2 Panel has a comparable sensitivity to RT-PCR assay.
Rhoads et al. found that there is a 94% positive percent agreement (PPA) between the ID NOW tests and an improved test developed by the CDC laboratory [63], while other assessments indicated that ID NOW has a lower PPA, ranging from 75% to 87%, compared with the laboratory-developed reference methods [64], [65], [66]. Compared to Xpert Xpress assay (~46 min) and ePlex assay (~1.5 h), ID NOW generates the fastest detection time for each sample (~17 min), but it comes at the expense of analysis and clinical performance, with the highest LOD and the lowest PPA [67]. In addition, Stephanie et al. showed the specificity of this assay was 100%, but 13 of the 46 SARS-CoV-2 positive samples were false-negative resulting in a lower sensitivity of 71.7%. All of the false-negatives equal to those weak positive samples [7]. This indicates that ID NOW has acceptable performance for samples contain high or moderate viral RNA, but may lack sensitivity when performing weakly positive samples.
Femke et al. evaluated Xpert Xpress point-of-care assay (Cepheid GeneXpert systems) for targeting SARS-CoV-2 E-gene and N2-gene in three medical laboratories in the Netherlands. The assay can detect SARS-CoV-2 with a LOD of 8.26 copies/mL in all three laboratories [68]. While Wei et al. reported Xpert Xpress test for targeting SARS-CoV-2 E-gene and N2-gene with a LOD of 100 copies/mL [67]. This difference may be caused by the different methods used to determine the input concentration and requires further verification. In Wei’s study, Xpert Xpress had the lowest LOD (100 copies/mL) compared to ID now (20,000 copies/mL) and ePlex (1000 copies/mL), and highest PPA (98.3%) compared to ID now (87.7%) and ePlex (91.4%). The information about inconsistent performance in these assays and workflow is vital to make timely and informed decisions on laboratory testing platform.
5. Antigen tests for diagnosis of COVID-19
SARS-CoV-2 consists of multiple virus-encoded proteins, including S, N, E, and M proteins. Among those proteins, S and N were the two main antigenic targets of SARS-CoV-2 antibodies [69]. Under the action of a host cell furin-like protease, the S protein is sliced into two separate polypeptides in most CoVs, S1 and S2 [70]. Although S protein is critical for virus entry and exists on the surface of the virus, N protein is the most richly expressed immune dominant protein that interacts with RNA and is more conserved than S protein [70]. Within S protein, the S1 submit is less conserved and more highly specific to SARS-CoV-2, suggesting S1 submit is more specific than full-length S or S2 submit as an antigen for COVID-19 serologic detection [70]. In addition, the RBD domain of S1 protein, is more conserved than S1 or full-length S, and has much less cross-reactivity with other CoVs [71]. Multiple forms of N protein or S protein (full-length S, S1 domain, S2 domain or receptor-binding domain [RBD]) — are used as targets [70], [71], [72], [73], [74].
Immunochromatographic assay is the most commonly used method for the detection of SARS-CoV-2 antigens [74], [75]. Thomas et al. compared four lateral flow antigen-detection kits (RapiGEN, Liming bio, Savant, and Bioeasy) for the detection of SARS-CoV-2 and showed test performances with significant differences. Among these tests, the Bioeasy test showed the highest accuracy of 89.2% and Kappa coefficient of 0.8, while Liming bio test was discontinued during testing because of poor performance. Sensitivities of other kits ranged from 16.7% for the Savant assay to 85% for the Bioeasy test [75].
Compared with the lower sensitivity of immunochromatography, highly sensitive biosensors-based tests have been demonstrated in the detection of SARS-CoV-2. Sophie et al. developed a portable, rapid cell-based biosensor with a human chimeric spike S1 antibody for detection of the SARS-CoV-2 S1 protein [76]. The biosensor allows tests completed within 3 min with a detection limit of 1 fg/mL and a semi-linear response range of 10 fg to 1 µg/mL. Similarly, Subhasis et al. constructed a biosensor device (eCovSens) to target SARS-CoV-2 S1 protein and compared it with a commercial potentiostat sensor. The LOD were found 90 fM for eCovSens and 120 fM for commercial potentiostat sensor in saliva samples [73]. These platforms can be used to monitor SARS-CoV-2 antigen on a large scale, providing a promising scheme for timely monitoring and eventual control of the global pandemic.
6. Antibody tests for diagnosis of COVID-19
With specific reagents, individual antibody types, like IgG, IgM, and IgA, can be determined. After SARS-CoV-2 infects the human body, IgM antibody can be produced within 5–7 days and is most useful for determining recent infection, while IgG antibody can be produced within 10–15 days and may remain detectable for months or years [8]. IgA is important for mucosal immunity and can be detected in mucous secretions within 6–8 days [77], though its significance in this disease is still to be determined.
In cases where RT-PCR assays are negative and there is a strong epidemiological link to SARS-CoV-2 infection, paired serum samples (in the acute and convalescent-phase) could support diagnosis once validated serology tests are available with the initial samples collected in the first week of COVID-19 and the second collected after 2–4 weeks [28]. Tests that detect binding antibodies fall into two broad categories, laboratory tests and point-of-care tests. A comparison of different serology tests for the detection of SARS-CoV-2 is shown in Table 1. The performance of currently available detection tests is compared in Table 3 .
Table 3.
The performance comparison analysis of serological detection tests.
| Tests | Detected antibodies | Specificity | Sensitivity (0-7d) | Sensitivity (8-14d) | Sensitivity (>=15d) | Total Sensitivity | Reference |
|---|---|---|---|---|---|---|---|
| Euroimmun (ELISA) | IgA | 86.1% | 65.5% | 87.09% | 93.93% | 83.65 | [84] |
| IgG | 98.6% | 17.2% | 66.12% | 90.9% | 61.7% | ||
| IgA or IgG | 87.5% | 65.5% | 88.7% | 93.935 | 84.4% | ||
| Maglumi™ (CLIA) | IgM | 100.0% | 14.3% | 64.51% | 87.5% | 58.7% | [84] |
| IgG | 100.0% | 7.1% | 54.8% | 87.5% | 53.2% | ||
| IgM or IgG | 100.0% | 17.9% | 69.4% | 93.8% | 64.3% | ||
| Alltest (LFA) | IgM | 100.0% | 21.1% | 19.0% | 60.0% | 28.9% | [87] |
| IgG | 100.0% | 21.1% | 33.3% | 86.7% | 60.0% | ||
| IgM or IgG | 100.0% | 26.3% | 42.9% | 93.3% | 64.4% | ||
| Clungene (LFA) | IgM | 91.3% | 16.2% | 42.3% | 55.3% | 39.2% | [88] |
| IgG | 98.1% | 29.7% | 60.3% | 97.4% | 62.1% | ||
| IgM or IgG | 90.3% | 35.1% | 64.1% | 97.4% | 65.4% | ||
| IgM and IgG | 99.0% | 10.8% | 38.5% | 55.3% | 35.9% | ||
| OrientGene (LFA) | IgM | 95.1% | 40.5% | 75.6% | 97.4% | 72.5% | [88] |
| IgG | 93.2% | 40.5% | 69.2% | 92.1% | 68.0% | ||
| IgM or IgG | 91.3% | 46.0% | 80.8% | 97.4% | 76.5% | ||
| IgM and IgG | 97.1% | 35.1% | 64.1% | 92.1% | 64.1% | ||
| VivaDiag (LFA) | IgM | 100.0% | 35.1% | 64.1% | 97.4% | 65.4% | [88] |
| IgG | 99.0% | 35.1% | 60.3% | 94.7% | 62.8% | ||
| IgM or IgG | 99.0% | 35.1% | 64.1% | 97.4% | 65.4% | ||
| IgM and IgG | 100.0% | 35.1% | 60.3% | 94.7% | 62.8% | ||
| StrongStep (LFA) | IgM | 99.0% | 10.8% | 33.3% | 50.0% | 32.0% | [88] |
| IgG | 99.0% | 32.4% | 64.1% | 97.4% | 64.7% | ||
| IgM or IgG | 98.1% | 35.1% | 66.7% | 97.4% | 66.7% | ||
| IgM and IgG | 100.0% | 8.1% | 30.8% | 50.0% | 30.1% | ||
| Dynamiker (LFA) | IgM | 95.1% | 46.0% | 66.7% | 97.4% | 69.3% | [88] |
| IgG | 99.0% | 27.0% | 61.5% | 94.7% | 61.4% | ||
| IgM or IgG | 95.2% | 46.0% | 66.7% | 97.4% | 69.3% | ||
| IgM and IgG | 99.0% | 27.0% | 61.5% | 94.7% | 61.4% | ||
| Multi-G (LFA) | IgM | 91.3% | 27.0% | 44.9% | 57.9% | 43.8% | [88] |
| IgG | 97.1% | 29.7% | 65.4% | 97.4% | 64.7% | ||
| IgM or IgG | 88.3% | 43.2% | 71.8% | 97.4% | 71.2% | ||
| IgM and IgG | 100.0% | 13.5% | 38.5% | 57.9% | 37.3% | ||
| Prima (LFA) | IgM | 93.2% | 43.2% | 56.4% | 68.4% | 56.2% | [88] |
| IgG | 90.3% | 40.5% | 71.8% | 100.0% | 71.2% | ||
| IgM or IgG | 85.4% | 56.8% | 79.5% | 100.0% | 79.1% | ||
| IgM and IgG | 98.1% | 27.0% | 48.7% | 68.4% | 48.4% | ||
| pGOLD assay (Nano-Plasmonic Platform) | IgM | 99.8% | 43.8% | 66.7% | 100.0% | 77.1% | [111] |
| IgG | 99.8% | 12.5% | 47.6% | 100.0% | 64.3% | ||
| IgM and IgG | 99.8% | 43.8% | 66.7% | 100.0% | 77.1% |
6.1. Manual laboratory-based antibody tests
6.1.1. Enzyme-linked immunosorbent assay
Niko et al. compared the performance of Vircell COVID-19 IgG (recombinant N protein) and Euroimmun SARS-CoV-2 IgG (recombinant S protein) assays at different stages of SARS-CoV-2 infection. The sensitivity was 70.6% for Vircell ELISA for and 58.8% for Euroimmun ELISA on days 5–9 after PCR-confirmed COVID-19. During 10–18 days after confirmation, the sensitivity was 100% for Vircell ELISA and 93.8% for Euroimmun ELISA [78]. Similarly, Liu et al. evaluated two ELISA kits based on SARS-CoV-2 S protein and N protein for detecting IgG and IgM antibodies. The team detected SARS-CoV-2 IgG antibodies in less than 60% of samples using S protein-based ELISA test on 6–10 days after the onset of the disease, and the sensitivity of the sample increases to >90% on 16 to 20 days after onset [79]. In addition, the sensitivity of 74.3% and 77.1% for S protein-based IgG and IgM ELISA, of 70.1% and 68.2% for N protein-based IgG and IgM ELISA, respectively. The overall sensitivity of the S protein-based and N protein-based ELISA tests were 82.2% and 80.4%. A higher sensitivity was observed in S protein-based ELISA compared to N protein-based ELISA [79].
6.1.2. Immunofluorescence assay
SARS-CoV-2 IgM and IgG antibodies in serum from the first COVID-19 case in Finland were identified by Immunofluorescence assay (IFA). The patient's serum was continuously diluted and incubated for 30 min for IgG and 2 h for IgM in Vero E6 cells. Although the antibodies were not detected on day 4 since the first symptoms, IgM and IgG titer both increased to 1:80 on day 9, 1:32 and 1:128 on day 20, respectively. The serum samples from the control group were not observed specific binding at dilutions greater than 1:20 [80]. Niko et al. compared the performance of IFA and neutralization assays at different stages of SARS-CoV-2 infection. The sensitivity of IFA and neutralization assays increased from 76.5% on days 5–9 after confirmation by PCR to 100% on days 10–18 [78]. Although IFA is promising in early diagnosis, non-specific fluorescence may lead to a wrong judgment and the sensitivity of this approach needs to be further evaluated.
6.1.3. Chemiluminescence immunoassay
Chemiluminescence immunoassay (CLIA) is one of the most popular immunology assays in identifying infectious diseases with the advantage of quantitative detection. The advantage of CLIA has recently been demonstrated in the detection of SARS-CoV-2 [81], [82], [83], [84]. Cai et al. developed a peptide-based Magnetic Chemiluminescence Enzyme Immunoassay (MCLIA) to detect SARS-CoV-2 antibodies against ORF1a/b, N and S proteins [83]. The sensitivity of IgM and IgG was 57.2% and 71.4%, respectively. A combination of IgM and IgG increased the detection rate to 81.5%. Isabel et al. evaluated Maglumi™ IgG/IgM CLIA assay and other four rapid IgG/IgM tests (three LFA tests and one ELISA test) for the detection of SARS-CoV-2 IgG, IgM and IgA antibodies in serum samples from 128 COVID-19 patients confirmed by RT-PCR. Compared with the Euroimmun IgG/IgA ELISA assay, the Maglumi™ IgG/IgM CLIA assay shows a lower overall sensitivity (84.4% vs. 64.3%). Both assays exhibited very similar specificities of IgG, which were 99% and 100% for ELISA and CLIA assays, respectively [84].
6.2. Rapid and point-of-care antibody tests
6.2.1. Lateral flow assay
The qualitative or semi-quantitative detection for SARS-CoV-2 IgM and IgG antibodies in serum, plasma and venous blood samples in vitro were proved by Lateral flow assay (LFA) [85], [86], [87], [88]. Isabel et al. evaluated three IgG/IgM LFA tests (Avioq, QuickZen and LaboOn Time) and two quantitative automated immunoassays (Euroimmun IgG/IgA ELISA test and Maglumi™ IgG/IgM CLIA assay) for the detection of SARS-CoV-2 IgG, IgM and IgA antibodies in serum samples from 128 COVID-19 patients confirmed by RT-PCR. The test presented a specificity of 100% in the analysis of negative control samples. The overall sensitivity of the three LFA tests was approximately 70% and no significant difference was observed. The sensitivity of all tests increased during the second week after the onset of symptoms, and all tests achieved a similar level of sensitivity (91% to 94%) after 14 days [84].
Jan et al. evaluate the diagnostic performance of 7 IgG/IgM LFA kits and the Euroimmun IgG/IgA ELISA test for detecting SARS-CoV-2 antibodies in COVID-19 patients. The specificity, sensitivity and dynamic trend to seropositivity were analyzed from 103 negative controls and 94 RT-PCR-confirmed COVID-19 patients. The specificity of LFA test was ≥90.3% for IgG, ≥ 91.3% for IgM, ≥ 85.4% IgM or IgG, and ≥97.1% for the combination IgM and IgG. The specificity of the ELISA test was 96.1% for IgG and only 73.8% for IgA. The sensitivity (14–25 days after onset of symptoms) of IgG LFA was ≥92.1%, while that of IgG ELISA was 89.5%. The positivity of IgM antibody for LFA yielded a decrease in specificity compared to IgG antibody alone and does not improve diagnostic performance [88]. The study indicated that measuring only LFA IgG can avoid false-positive results of IgM.
6.2.2. Microarray and microfluidic chip
Microarray and microfluidic platforms can be easily converted to point-of-care settings by portability, miniaturization, automation and integration of multiple functions onto chips. With small size, high sensitivity, and high throughput analysis, microarray and microfluidic chips are powerful tools for pathogen identification. Rafael et al. describe an antigen microarray containing 65 antigens that are causes of acute respiratory infections from SARS-CoV-2, SARS-CoV, MERS-CoV, and other common respiratory viruses. The SARS-CoV-2 antigens coated on this array include N protein, S protein, RBD, S1 and S2 domains. The samples of confirmed COVID-19 cases were highly reactive to SARS-CoV-2 antigens, and the reactivity in IgG is more obvious than IgA. Similar to IgG, IgA showed higher reactivity to SARS-CoV-2 N protein, S2 domain, and full-length S, cross-reactivity with SARS-CoV N protein, but not with MERS-CoV antigens [89]. The team further developed a modular microarray imaging system for the detection of SARS-CoV-2 antibody. The imaging platform can produce similar results to the commercial imager (ArrayCAM 400-S, Grace Biolabs), which are 100 times more expensive. Linear regression analysis of microarray fluorescence intensities revealed R-squared values > 0.85 between different imaging platforms. This platform has the advantages of low-cost, high-throughput, and can handle more than 100,000 samples, potentially valuable for serosurveillance in COVID-19 patients [90]. In addition, Tan et al. designed a microfluidic ELISA test for quantitative detection of SARS-CoV-2 S1 protein and anti-SARS-CoV-2 S1 IgG using humanized SARS-CoV-2 IgG and recombinant S1 protein, respectively. The lower LOD of 2 ng/mL for anti-SARS-CoV-2 S1 IgG and of 0.4 ng/mL for S1 antigen was achieved in serum [91]. These studies demonstrated high performance of analyzing SARS-CoV-2 specific antibodies and antigens in COVID-19 patients by microarray and microfluidic chip. However, some proteins on the microarray were observed not expressed in mammalian cell systems and need to be further optimized [92].
7. Virological tests
As the gold standard for virological diagnosis, a culture-based virus detection is an important tool for virus discovery, pathogenesis research and strategy evaluation. Neutralization tests evaluated the capacity of serum from COVID-19 patients to reduce cytopathic effects (CPE) caused by SARS-CoV-2 susceptible cells in vitro. It involves incubating the serum or plasma with live virus followed by infection and incubation of cells. It mainly includes virus neutralization tests (VNT) and pseudovirus neutralization tests (PVNT) [93].
The first COVID-19 case in Finland was confirmed by cytopathic effect (CPE)-based VNT to analyze SARS-CoV-2 antibody levels. At day 9–10 after the first symptom onset, an increase of at least 4-fold in the neutralizing antibodies was observed, and the titer of antibodies was still increasing on day 20 [80]. Wang et al. found that the titers of antibodies bind to N and S proteins was observably higher in severe patients. Meanwhile, the average titers of neutralizing antibody against SARS-CoV-2 pseudovirus and live virus was higher in the sicker cases, by ~5-fold and ~7-fold, respectively [94]. Renata et al. detected the ability of serum from COVID-19 patients to reduce the CPE caused by SARS-CoV-2 infection of susceptible cells by a micro-neutralization assay. Neutralizing antibodies were detected in 16 out of 20 patients, with titers ranging from 10 to 1920. There is a strong positive correlation between neutralizing antibody titers and total SARS-CoV-2 IgG antibody levels in COVID-19 patients were observed. In addition, anti-S1-IgA, anti-S1-IgG and anti-N-IgM levels were also related to the neutralizing antibody titers of SARS-CoV-2. These studies indicating that SARS-CoV-2 antibody titers in COVID- 19 patients may reflect the capacity of serum to neutralize SARS-CoV-2 [95].
Focusing on IgG antibodies, the performance of in-house plaque reduction neutralization test (PRNT), in-house IFA, Euroimmun ELISA, Vircell ELISA and FaStep LFA were analyzed. In the early stages of infection, the sensitivity to PRNT and IFA tests was 76.5%, higher than two ELISA tests and LFA test. At the later stage of 10–18 days, IFA, PRNT and Vircell ELISA showed a sensitivity of 100%, while the Euroimmun ELISA and Assure LFA of 93.8% [78]. The study demonstrated the superior sensitivity of the neutralization test in the diagnosis of COVID-19. Neutralization assay is specific for SARS-CoV-2 antibodies and could be used to monitor patient immunity to the virus. However, compared with serological tests, neutralization tests are long-period, laborious, expensive, time-consuming, and limited to perform in biosafety level 3 (BSL-3) or BSL-2 laboratories.
8. Genomic sequencing in SARS-CoV-2
Genomic sequencing of viruses is a powerful tool for analyzing virus evolution, and genetic association to diseases, tracking outbreaks and developing new therapies and vaccines. As of March 2020, hundreds of SARS-CoV-2 genomes were released on public databases including Global initiative on sharing all influenza data (GISAID), NCBI GenBank and China National GeneBank DataBase (CNGBdb). At present, sequencing methods of SARS-CoV-2 include metatranscriptomics sequencing [27], hybrid capture-based sequencing [96], amplicon sequencing [96], and nanopore targeted sequencing [97]. The advantages and disadvantages of SARS-CoV-2 genomic sequencing are shown in Table 1.
The first research team got the SARS-CoV-2 genomic sequences by metatranscriptomic sequencing, supplemented by PCR and Sanger sequencing of a connection between BALF and virus culture [8], [13] or from BALF directly [11], [33]. Lu et al. obtained ten genome sequences of SARS-CoV-2 from nine patients’ BALF and culture samples by metatranscriptomic sequencing. The ten genome sequences were nearly identical, displaying more than 99.98% sequence identity [13]. This finding indicates that SARS-CoV-2 may originate from the same source within a short period. In addition, it was reported that whole-genome sequences of SARS-CoV-2 from nasopharyngeal and oropharyngeal samples from a COVID-19 case by Sanger and metatranscriptomic sequencing with both MinIon and Illumina in the United States [27]. The entire genome sequences of the nasopharyngeal and oropharyngeal samples were identical to each other and were highly similar to other available SARS-CoV-2 sequences.
Although the amplicon sequencing and hybrid capture sequencing with high sensitivity but low accuracy, and neither of them can be applied to sequence viruses with highly diverse or recombinant due to primers and probes are designed to the known viral genomes [96]. Compared with metatranscriptomic sequencing, the results of amplicons and capture sequencing showed that significant increases in the ratio of SARS-CoV-2 reads out of the total reads, which indicates the enrichment was efficient 5710-fold in amplicon sequencing and 5596-fold in hybrid capture sequencing for each specimen on average. The alleles identified by hybrid capture sequencing showed lower frequencies than amplicon sequencing and metatranscriptomic sequencing, especially for viruses with lower load. Capture sequencing neglected a single nucleotide variation (SNV) when the cutoff of SNV calling was set as 80% allele frequency [96].
Combining the strengths of target amplification and long-read, real-time nanopore sequencing, nanopore targeted sequencing was established to simultaneously detect SARS-CoV-2 and 10 types of other respiratory viruses at LoD of 10 copies/ml. NTS identified positive to 22 of 61 suspected COVID-19 specimens that were either negative or inconclusive by RT-PCR. Screening mutations showed that 4 of 19 throat swab samples from COVID-19 patients were found single-nucleotide mutations at seven sites [97]. These findings suggested that NTS is suitable for the identification and mutation monitoring of SARS-CoV-2 from clinical samples.
9. Biosafety precaution and protection
Laboratory diagnosis of clinical specimens from suspected or confirmed COVID-19 patients should be conducted adopting practices and procedures described by WHO [98]. Specimen collection, storage, packaging, and transportation under standard operating procedures. SARS-CoV-2 was detected in unusual types of specimens such as saliva and conjunctival swabs [36], [37], [38], [40], so it is recommended that all specimens collected for laboratory investigation should be considered potentially infectious. RNA extraction, nucleic acid amplification assays and sequencing should be conducted in BSL-2 laboratory, while neutralization assays and virus culture should be carried out in BSL-3 or BSL-2 laboratory [28].
According to the current evidence, COVID-19 virus is primarily transmitted between people via respiratory droplets and contact routes. The droplets can also be deposited directly on individual next to the infected case. Hence, washing hands frequently and keeping a distance of at least one meter are considered the main measure to prevent infection [99]. Personal protective equipment (PPE) can protect people against infection, especially in susceptible people. PPE recommended by WHO include medical masks, gloves, gowns, goggles or a face shield, respirators and aprons [100]. Among the general public, persons with respiratory symptoms or those caring for COVID-19 patients at home should receive medical masks, but N95 respirators is not recommended [101]. Using N95 masks compared with the use of medical masks was not related to any statistically significant lower risk of laboratory-confirmed viral infections [102], [103]. With the development of globalization, emerging infectious diseases continually occur and may cause an outbreak globally. Understanding biosecurity and personal protection can prepare for the prevention and control of emerging infectious diseases.
10. Conclusion
COVID-19 has caused a large-scale epidemic in countries around the world and posed a serious threat to international health. The rapid and early laboratory diagnosis is critical to diagnose infection and control transmission. As the gold standard test for SARS-CoV-2 identification, RT-PCR is the routine confirmation test performed by WHO. Although RT-PCR is designed for conserved regions of the viral genome, RNA viruses display a large number of genetic variations, mismatches between primers, probes, and target sequences may lead to reduced detection performance and false-negative results. Serological surveys can aid the investigation of an ongoing outbreak and retrospective assessment of the attack rate or extent of an outbreak. Detection of SARS-CoV-2 IgM and IgG antibody in vivo as a supplement to molecular diagnostic methods. However, serological tests require seroconversion and cannot be applied to early infection.
Laboratory-based tests both for molecular and immunological tests are not suitable for point-of-care diagnosis due to time-consuming, expensive equipment and biosafety requirements. Point-of-care tests are not only suitable for clinical laboratories, but also performed by trained non-laboratory personnel in patient care facilities, such as physicians' offices or Emergency Departments, making the diagnostic test of SARS-CoV-2 closer to the patient. Portable benchtop-sized analyzers for molecular tests such as Xpert Xpress assay and ID NOW with the advantage of automatic, portable, rapid and not requires trained staff, should be recommended for real-time patient management and infection control decisions. Lateral flow immunoassay is the widely used serological test for point-of-care detection, but the performance of commercial kits still need to be improved. In conclusion, serological tests can serve as indicators of the spread of the virus, while RT-PCR tests can show who is currently infected with the disease. A combination of molecular and serological tests is needed to improve the diagnostic accuracy of COVID-19.
Author contributions
Chenxi Li and Chengxue Zhao conceived the essay; Chenxi Li researched the literature and wrote the manuscript; Jingfeng Bao and Bo Tang made the tables and figures; Yunfeng Wang and Bing Gu revised the manuscript.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgements
This research was supported by the Jiangsu Provincial Medical Talent (ZDRCA2016053), Six talent peaks project of Jiangsu Province (WSN-135), Advanced health talent of six-one project of Jiangsu Province (LGY2016042).
References
- 1.Su S., Wong G., Shi W., Liu J., Lai A.C.K., Zhou J., Liu W., Bi Y., Gao G.F. Epidemiology, genetic recombination, and pathogenesis of coronaviruses. Trends Microbiol. 2016;24(6):490–502. doi: 10.1016/j.tim.2016.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Huang P., Wang H., Cao Z., Jin H., Chi H., Zhao J., Yu B., Yan F., Hu X., Wu F., Jiao C., Hou P., Xu S., Zhao Y., Feng N., Wang J., Sun W., Wang T., Gao Y., Yang S., Xia X. A rapid and specific assay for the detection of MERS-CoV. Front. Microbiol. 2018;9:1101. doi: 10.3389/fmicb.2018.01101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.World Health Organization, Coronavirus disease (COVID-19) situation reports. https://www.who.int/docs/default-source/coronaviruse/situation-reports/20200507covid-19-sitrep-108.pdf?sfvrsn=44cc8ed8_2 (accessed 7 May 2020).
- 4.World Health Organization, Coronavirus disease (COVID-2019) situation reports. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports.
- 5.Tahamtan A., Ardebili A. Real-time RT-PCR in COVID-19 detection: issues affecting the results. Expert Rev. Mol. Diagn. 2020;20(5):453–454. doi: 10.1080/14737159.2020.1757437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Xie X., Zhong Z., Zhao W., Zheng C., Wang F., Liu J. Chest CT for typical 2019-nCoV pneumonia: relationship to negative RT-PCR testing. Radiology. 2020 doi: 10.1148/radiol.2020200343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Mitchell S.L., George K.S. Evaluation of the COVID19 ID NOW EUA assay. J. Clin. Virol. 2020;128 doi: 10.1016/j.jcv.2020.104429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Zhou P., Yang X.L., Wang X.G., Hu B., Zhang L., Zhang W., Si H.R., Zhu Y., Li B., Huang C.L., Chen H.D., Chen J., Luo Y., Guo H., Jiang R.D., Liu M.Q., Chen Y., Shen X.R., Wang X., Zheng X.S., Zhao K., Chen Q.J., Deng F., Liu L.L., Yan B., Zhan F.X., Wang Y.Y., Xiao G.F., Shi Z.L. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579(7798):270–273. doi: 10.1038/s41586-020-2012-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Maurier F., Beury D., Fléchon L., Varré J.S., Touzet H., Goffard A., Hot D., Caboche S. A complete protocol for whole-genome sequencing of virus from clinical samples: Application to coronavirus OC43. Virology. 2019;531:141–148. doi: 10.1016/j.virol.2019.03.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Zhu N., Zhang D., Wang W., Li X., Yang B., Song J., Zhao X., Huang B., Shi W., Lu R., Niu P., Zhan F., Ma X., Wang D., Xu W., Wu G., Gao G.F., Tan W. A novel coronavirus from patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020;382(8):727–733. doi: 10.1056/NEJMoa2001017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chen L., Liu W., Zhang Q., Xu K., Ye G., Wu W., Sun Z., Liu F., Wu K., Zhong B., Mei Y., Zhang W., Chen Y., Li Y., Shi M., Lan K., Liu Y. RNA based mNGS approach identifies a novel human coronavirus from two individual pneumonia cases in 2019 Wuhan outbreak. Emerg. Microbes Infect. 2020;9(1):313–319. doi: 10.1080/22221751.2020.1725399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wu C., Liu Y., Yang Y., Zhang P., Zhong W., Wang Y., Wang Q., Xu Y., Li M., Li X., Zheng M., Chen L., Li H. Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharm. Sin. B. 2020;10(5):766–788. doi: 10.1016/j.apsb.2020.02.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lu R., Zhao X., Li J., Niu P., Yang B., Wu H., Wang W., Song H., Huang B., Zhu N., Bi Y., Ma X., Zhan F., Wang L., Hu T., Zhou H., Hu Z., Zhou W., Zhao L., Chen J., Meng Y., Wang J., Lin Y., Yuan J., Xie Z., Ma J., Liu W.J., Wang D., Xu W., Holmes E.C., Gao G.F., Wu G., Chen W., Shi W., Tan W. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395(10224):565–574. doi: 10.1016/S0140-6736(20)30251-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lam T.T., Jia N., Zhang Y.W., Shum M.H., Jiang J.F., Zhu H.C., Tong Y.G., Shi Y.X., Ni X.B., Liao Y.S., Li W.J., Jiang B.G., Wei W., Yuan T.T., Zheng K., Cui X.M., Li J., Pei G.Q., Qiang X., Cheung W.Y., Li L.F., Sun F.F., Qin S., Huang J.C., Leung G.M., Holmes E.C., Hu Y.L., Guan Y., Cao W.C. Identifying SARS-CoV-2-related coronaviruses in Malayan pangolins. Nature. 2020 doi: 10.1038/s41586-020-2169-0. [DOI] [PubMed] [Google Scholar]
- 15.Zhang T., Wu Q., Zhang Z. Probable pangolin origin of SARS-CoV-2 associated with the COVID-19 outbreak. Curr. Biol. 2020;30(8):1578. doi: 10.1016/j.cub.2020.03.063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Y. Liu, G. Hu, Y. Wang, X. Zhao, F. Ji, W. Ren, M. Gong, X. Ju, C. Li, J. Hong, Y. Zhu, X. Cai, J. Wu, X. Lan, Y. Xie, X. Wang, Z. Yuan, R. Zhang, Q. Ding, Functional and genetic analysis of viral receptor ACE2 orthologs reveals a broad potential host range of SARS-CoV-2, bioRxiv (2020) 2020.04.22.046565. [DOI] [PMC free article] [PubMed]
- 17.Liu Z., Xiao X., Wei X., Li J., Yang J., Tan H., Zhu J., Zhang Q., Wu J., Liu L. Composition and divergence of coronavirus spike proteins and host ACE2 receptors predict potential intermediate hosts of SARS-CoV-2. J. Med. Virol. 2020 doi: 10.1002/jmv.25726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Riou J., Althaus C.L. Pattern of early human-to-human transmission of Wuhan 2019 novel coronavirus (2019-nCoV), December 2019 to January 2020. Euro Surveill. 2020;25(4) doi: 10.2807/1560-7917.ES.2020.25.4.2000058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhao S., Lin Q., Ran J., Musa S.S., Yang G., Wang W., Lou Y., Gao D., Yang L., He D., Wang M.H. Preliminary estimation of the basic reproduction number of novel coronavirus (2019-nCoV) in China, from 2019 to 2020: a data-driven analysis in the early phase of the outbreak. Int. J. Infect. Dis. 2020;92:214–217. doi: 10.1016/j.ijid.2020.01.050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sanche S., Lin Y.T., Xu C., Romero-Severson E., Hengartner N., Ke R. High contagiousness and rapid spread of severe acute respiratory syndrome coronavirus 2. Emerg. Infect. Dis. 2020;26(7) doi: 10.3201/eid2607.200282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.World Health Organization, Consensus document on theepidemiology of severe acuterespiratory syndrome (SARS). Available at: https://www.who.int/csr/sars/en/WHOconsensus.pdf.
- 22.Jin Y., Yang H., Ji W., Wu W., Chen S., Zhang W., Duan G. Virology, epidemiology, pathogenesis, and control of COVID-19. Viruses. 2020;12(4) doi: 10.3390/v12040372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Guo Z.D., Wang Z.Y., Zhang S.F., Li X., Li L., Li C., Cui Y., Fu R.B., Dong Y.Z., Chi X.Y., Zhang M.Y., Liu K., Cao C., Liu B., Zhang K., Gao Y.W., Lu B., Chen W. Aerosol and surface distribution of severe acute respiratory syndrome coronavirus 2 in hospital wards, Wuhan, China, 2020. Emerg. Infect. Dis. 2020;26(7) doi: 10.3201/eid2607.200885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.van Doremalen N., Bushmaker T., Morris D.H., Holbrook M.G., Gamble A., Williamson B.N., Tamin A., Harcourt J.L., Thornburg N.J., Gerber S.I., Lloyd-Smith J.O., de Wit E., Munster V.J. Aerosol and surface stability of SARS-CoV-2 as compared with SARS-CoV-1. N. Engl. J. Med. 2020;382(16):1564–1567. doi: 10.1056/NEJMc2004973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Zhang W., Du R.H., Li B., Zheng X.S., Yang X.L., Hu B., Wang Y.Y., Xiao G.F., Yan B., Shi Z.L., Zhou P. Molecular and serological investigation of 2019-nCoV infected patients: implication of multiple shedding routes. Emerg. Microbes Infect. 2020;9(1):386–389. doi: 10.1080/22221751.2020.1729071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Chen W., Lan Y., Yuan X., Deng X., Li Y., Cai X., Li L., He R., Tan Y., Deng X., Gao M., Tang G., Zhao L., Wang J., Fan Q., Wen C., Tong Y., Tang Y., Hu F., Li F., Tang X. Detectable 2019-nCoV viral RNA in blood is a strong indicator for the further clinical severity. Emerg. Microbes Infect. 2020;9(1):469–473. doi: 10.1080/22221751.2020.1732837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Holshue M.L., DeBolt C., Lindquist S., Lofy K.H., Wiesman J., Bruce H., Spitters C., Ericson K., Wilkerson S., Tural A., Diaz G., Cohn A., Fox L., Patel A., Gerber S.I., Kim L., Tong S., Lu X., Lindstrom S., Pallansch M.A., Weldon W.C., Biggs H.M., Uyeki T.M., Pillai S.K. First case of 2019 novel coronavirus in the United States. N Engl J Med. 2020;382(10):929–936. doi: 10.1056/NEJMoa2001191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.World Health Organization, Laboratory testing for 2019 novel coronavirus (2019-nCoV) in suspected human cases. https://www.who.int/publications-detail/laboratory-testing-for-2019-novel-coronavirus-in-suspected-human-cases-20200117 (accessed 19 March 2020).
- 29.Chan J.F., Yuan S., Kok K.H., To K.K., Chu H., Yang J., Xing F., Liu J., Yip C.C., Poon R.W., Tsoi H.W., Lo S.K., Chan K.H., Poon V.K., Chan W.M., Ip J.D., Cai J.P., Cheng V.C., Chen H., Hui C.K., Yuen K.Y. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet. 2020;395(10223):514–523. doi: 10.1016/S0140-6736(20)30154-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kim J.M., Chung Y.S., Jo H.J., Lee N.J., Kim M.S., Woo S.H., Park S., Kim J.W., Kim H.M., Han M.G. Identification of Coronavirus Isolated from a Patient in Korea with COVID-19. Osong Public Health Res. Perspect. 2020;11(1):3–7. doi: 10.24171/j.phrp.2020.11.1.02. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Yu F., Yan L., Wang N., Yang S., Wang L., Tang Y., Gao G., Wang S., Ma C., Xie R., Wang F., Tan C., Zhu L., Guo Y., Zhang F. Quantitative detection and viral load analysis of SARS-CoV-2 in infected patients. Clin. Infect. Dis. 2020 doi: 10.1093/cid/ciaa345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Rothe C., Schunk M., Sothmann P., Bretzel G., Froeschl G., Wallrauch C., Zimmer T., Thiel V., Janke C., Guggemos W., Seilmaier M., Drosten C., Vollmar P., Zwirglmaier K., Zange S., Wölfel R., Hoelscher M. Transmission of 2019-nCoV infection from an asymptomatic contact in Germany. N. Engl. J. Med. 2020;382(10):970–971. doi: 10.1056/NEJMc2001468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wu F., Zhao S., Yu B., Chen Y.M., Wang W., Song Z.G., Hu Y., Tao Z.W., Tian J.H., Pei Y.Y., Yuan M.L., Zhang Y.L., Dai F.H., Liu Y., Wang Q.M., Zheng J.J., Xu L., Holmes E.C., Zhang Y.Z. A new coronavirus associated with human respiratory disease in China. Nature. 2020;579(7798):265–269. doi: 10.1038/s41586-020-2008-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Guan W.J., Ni Z.Y., Hu Y., Liang W.H., Ou C.Q., He J.X., Liu L., Shan H., Lei C.L., Hui D.S.C., Du B., Li L.J., Zeng G., Yuen K.Y., Chen R.C., Tang C.L., Wang T., Chen P.Y., Xiang J., Li S.Y., Wang J.L., Liang Z.J., Peng Y.X., Wei L., Liu Y., Hu Y.H., Peng P., Wang J.M., Liu J.Y., Chen Z., Li G., Zheng Z.J., Qiu S.Q., Luo J., Ye C.J., Zhu S.Y., Zhong N.S. Clinical characteristics of coronavirus disease 2019 in China. N. Engl. J. Med. 2020;382(18):1708–1720. doi: 10.1056/NEJMoa2002032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Peng L., Liu J., Xu W., Luo Q., Chen D., Lei Z., Huang Z., Li X., Deng K., Lin B., Gao Z. SARS-CoV-2 can be detected in urine, blood, anal swabs, and oropharyngeal swabs specimens. J. Med. Virol. 2020 doi: 10.1002/jmv.25936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.K.K.-W. To, O.T.-Y. Tsang, C. Chik-Yan Yip, K.-H. Chan, T.-C. Wu, J.M.C. Chan, W.-S. Leung, T.S.-H. Chik, C.Y.-C. Choi, D.H. Kandamby, D.C. Lung, A.R. Tam, R.W.-S. Poon, A.Y.-F. Fung, I.F.-N. Hung, V.C.-C. Cheng, J.F.-W. Chan, K.-Y. Yuen, Consistent detection of 2019 novel coronavirus in saliva, Clin. Infect. Dis.: Off. Publ. Infect. Dis. Soc. Am. (2020) ciaa149. [DOI] [PMC free article] [PubMed]
- 37.To K.K., Tsang O.T., Leung W.S., Tam A.R., Wu T.C., Lung D.C., Yip C.C., Cai J.P., Chan J.M., Chik T.S., Lau D.P., Choi C.Y., Chen L.L., Chan W.M., Chan K.H., Ip J.D., Ng A.C., Poon R.W., Luo C.T., Cheng V.C., Chan J.F., Hung I.F., Chen Z., Chen H., Yuen K.Y. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. Lancet Infect. Dis. 2020;20(5):565–574. doi: 10.1016/S1473-3099(20)30196-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Chen J.H., Yip C.C., Poon R.W., Chan K.H., Cheng V.C., Hung I.F., Chan J.F., Yuen K.Y., To K.K. Evaluating the use of posterior oropharyngeal saliva in a point-of-care assay for the detection of SARS-CoV-2. Emerg. Microbes Infect. 2020:1–14. doi: 10.1080/22221751.2020.1775133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.L.V. Tan, N.M. Ngoc, B.T.T. That, L.T.T. Uyen, N.T.T. Hong, N.T.P. Dung, L.N.T. Nhu, T.T. Thanh, D.N.H. Man, N.T. Phong, T.T. Hien, N.T. Truong, G. Thwaites, N.V.V. Chau, Duration of viral detection in throat and rectum of a patient with COVID-19, medRxiv (2020) 2020.03.07.20032052.
- 40.Zhou Y., Zeng Y., Tong Y., Chen C. Ophthalmologic evidence against the interpersonal transmission of 2019 novel coronavirus through conjunctiva. medRxiv. 2020 2020.02.11.20021956. [Google Scholar]
- 41.Xia J., Tong J., Liu M., Shen Y., Guo D. Evaluation of coronavirus in tears and conjunctival secretions of patients with SARS-CoV-2 infection. J. Med. Virol. 2020 doi: 10.1002/jmv.25725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Hong K.H., Lee S.W., Kim T.S., Huh H.J., Lee J., Kim S.Y., Park J.S., Kim G.J., Sung H., Roh K.H., Kim J.S., Kim H.S., Lee S.T., Seong M.W., Ryoo N., Lee H., Kwon K.C., Yoo C.K. Guidelines for laboratory diagnosis of coronavirus disease 2019 (COVID-19) in Korea. Ann. Lab. Med. 2020;40(5):351–360. doi: 10.3343/alm.2020.40.5.351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.L. Peng, J. Liu, W. Xu, Q. Luo, K. Deng, B. Lin, Z. Gao, 2019 Novel Coronavirus can be detected in urine, blood, anal swabs and oropharyngeal swabs samples, medRxiv (2020) 2020.02.21.20026179. [DOI] [PMC free article] [PubMed]
- 44.World Health Organization, Coronavirus disease (COVID-19) technical guidance: Laboratory testing for 2019-nCoV in humans. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/technical-guidance/laboratory-guidance (2020).
- 45.Centers for Disease Control and Prevention, Real-time RT-PCR panel for detection 2019-Novel Coronavirus. https://www.cdc.gov/coronavirus/2019-ncov/downloads/rt-pcr-panel-for-detection-instructions.pdf (accessed 4 February 2020).
- 46.Corman V.M., Landt O., Kaiser M., Molenkamp R., Meijer A., Chu D.K., Bleicker T., Brünink S., Schneider J., Schmidt M.L., Mulders D.G., Haagmans B.L., van der Veer B., van den Brink S., Wijsman L., Goderski G., Romette J.L., Ellis J., Zambon M., Peiris M., Goossens H., Reusken C., Koopmans M.P., Drosten C. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 2020;25(3) doi: 10.2807/1560-7917.ES.2020.25.3.2000045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Vogels C.B.F., Brito A.F., Wyllie A.L., Fauver J.R., Ott I.M., Kalinich C.C., Petrone M.E., Landry M.-L., Foxman E.F., Grubaugh N.D. Analytical sensitivity and efficiency comparisons of SARS-COV-2 qRT-PCR assays. medRxiv. 2020 doi: 10.1038/s41564-020-0761-6. 2020.03.30.20048108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Konrad R., Eberle U., Dangel A., Treis B., Berger A., Bengs K., Fingerle V., Liebl B., Ackermann N., Sing A. Rapid establishment of laboratory diagnostics for the novel coronavirus SARS-CoV-2 in Bavaria, Germany, February 2020. Euro Surveill. 2020;25(9) doi: 10.2807/1560-7917.ES.2020.25.9.2000173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Lau L.T., Fung Y.-W.-W., Wong F.-P.-F., Lin S.-S.-W., Wang C.R., Li H.L., Dillon N., Collins R.A., Tam J.-S.-L., Chan P.K.S., Wang C.G., Yu A.-C.-H. A real-time PCR for SARS-coronavirus incorporating target gene pre-amplification. Biochem. Biophys. Res. Commun. 2003;312(4):1290–1296. doi: 10.1016/j.bbrc.2003.11.064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Jiang S.S., Chen T.-C., Yang J.-Y., Hsiung C.A., Su I.-J., Liu Y.-L., Chen P.-C., Juang J.-L. Sensitive and quantitative detection of severe acute respiratory syndrome coronavirus infection by real-time nested polymerase chain reaction. Clin. Infect. Dis.: Off. Publ. Infect. Dis. Soc. Am. 2004;38(2):293–296. doi: 10.1086/380841. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Shirato K., Nao N., Katano H., Takayama I., Saito S., Kato F., Katoh H., Sakata M., Nakatsu Y., Mori Y., Kageyama T., Matsuyama S., Takeda M. Development of genetic diagnostic methods for novel coronavirus 2019 (nCoV-2019) in Japan. Japanese J. Infect. Dis. 2020 doi: 10.7883/yoken.JJID.2020.061. [DOI] [PubMed] [Google Scholar]
- 52.Wang J., Cai K., Zhang R., He X., Shen X., Liu J., Xu J., Qiu F., Lei W., Wang J., Li X., Gao Y., Jiang Y., Xu W., Ma X. Novel one-step single-tube nested quantitative real-time PCR assay for highly sensitive detection of SARS-CoV-2. Anal. Chem. 2020 doi: 10.1021/acs.analchem.0c01884. [DOI] [PubMed] [Google Scholar]
- 53.Peiris J.S.M., Chu C.M., Cheng V.C.C., Chan K.S., Hung I.F.N., Poon L.L.M., Law K.I., Tang B.S.F., Hon T.Y.W., Chan C.S., Chan K.H., Ng J.S.C., Zheng B.J., Ng W.L., Lai R.W.M., Guan Y., Yuen K.Y., H.U.S.S. Group Clinical progression and viral load in a community outbreak of coronavirus-associated SARS pneumonia: a prospective study. Lancet (London, England) 2003;361(9371):1767–1772. doi: 10.1016/S0140-6736(03)13412-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Liu X., Feng J., Zhang Q., Guo D., Zhang L., Suo T., Hu W., Guo M., Wang X., Huang Z., Xiong Y., Chen G., Chen Y., Lan K. Analytical comparisons of SARS-COV-2 detection by qRT-PCR and ddPCR with multiple primer/probe sets. Emerg. Microbes Infect. 2020;9(1):1175–1179. doi: 10.1080/22221751.2020.1772679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Suo T., Liu X., Feng J., Guo M., Hu W., Guo D., Ullah H., Yang Y., Zhang Q., Wang X., Sajid M., Huang Z., Deng L., Chen T., Liu F., Xu K., Liu Y., Zhang Q., Liu Y., Xiong Y., Chen G., Lan K., Chen Y. ddPCR: a more accurate tool for SARS-CoV-2 detection in low viral load specimens. Emerg. Microbes Infect. 2020;9(1):1259–1268. doi: 10.1080/22221751.2020.1772678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Zhang Y., Odiwuor N., Xiong J., Sun L., Nyaruaba R.O., Wei H., Tanner N.A. Rapid molecular detection of SARS-CoV-2 (COVID-19) Virus RNA using colorimetric LAMP. medRxiv. 2020 2020.02.26.20028373. [Google Scholar]
- 57.Z. Zhao, H. Cui, W. Song, X. Ru, W. Zhou, X. Yu, A simple magnetic nanoparticles-based viral RNA extraction method for efficient detection of SARS-CoV-2, bioRxiv (2020) 2020.02.22.961268.
- 58.M. El-Tholoth, H.H. Bau, J. Song, A single and two-stage, closed-tube, molecular test for the 2019 novel coronavirus (COVID-19) at home, clinic, and points of entry, ChemRxiv (2020).
- 59.X. Zhu, X. Wang, L. Han, T. Chen, L. Wang, H. Li, S. Li, L. He, X. Fu, S. Chen, X. Mei, H. Chen, Y. Wang, Reverse transcription loop-mediated isothermal amplification combined with nanoparticles-based biosensor for diagnosis of COVID-19, medRxiv (2020) 2020.03.17.20037796. [DOI] [PMC free article] [PubMed]
- 60.Moitra P., Alafeef M., Dighe K., Frieman M.B., Pan D. Selective naked-eye detection of SARS-CoV-2 mediated by N gene targeted antisense oligonucleotide capped plasmonic nanoparticles. ACS Nano. 2020 doi: 10.1021/acsnano.0c03822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Halfpenny K.C., Wright D.W. Nanoparticle detection of respiratory infection. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2010;2(3):277–290. doi: 10.1002/wnan.83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Visseaux B., Le Hingrat Q., Collin G., Bouzid D., Lebourgeois S., Le Pluart D., Deconinck L., Lescure F.X., Lucet J.C., Bouadma L., Timsit J.F., Descamps D., Yazdanpanah Y., Casalino E., Houhou-Fidouh N. Evaluation of the QIAstat-Dx respiratory SARS-CoV-2 panel, the first rapid multiplex PCR commercial assay for SARS-CoV-2 detection. J. Clin. Microbiol. 2020 doi: 10.1128/JCM.00630-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Rhoads D.D., Cherian S.S., Roman K., Stempak L.M., Schmotzer C.L., Sadri N. Comparison of Abbott ID Now, Diasorin Simplexa, and CDC FDA EUA methods for the detection of SARS-CoV-2 from nasopharyngeal and nasal swabs from individuals diagnosed with COVID-19. J. Clin. Microbiol. 2020 doi: 10.1128/JCM.00760-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Zhen W., Smith E., Manji R., Schron D., Berry G.J. Clinical evaluation of three sample-to-answer platforms for the detection of SARS-CoV-2. J. Clin. Microbiol. 2020 doi: 10.1128/JCM.00783-20. JCM.00783-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Harrington A., Cox B., Snowdon J., Bakst J., Ley E., Grajales P., Maggiore J., Kahn S. Comparison of Abbott ID Now and Abbott m2000 methods for the detection of SARS-CoV-2 from nasopharyngeal and nasal swabs from symptomatic patients. J. Clin. Microbiol. 2020 doi: 10.1128/JCM.00798-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Hogan C.A., Sahoo M.K., Huang C., Garamani N., Stevens B., Zehnder J., Pinsky B.A. Five-minute point-of-care testing for SARS-CoV-2: Not there yet. J. Clin. Virol. 2020;128 doi: 10.1016/j.jcv.2020.104410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Zhen W., Smith E., Manji R., Schron D., Berry G.J. Clinical evaluation of three sample-to-answer platforms for the detection of SARS-CoV-2. J. Clin. Microbiol. 2020 doi: 10.1128/JCM.00783-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Wolters F., van de Bovenkamp J., van den Bosch B., van den Brink S., Broeders M., Chung N.H., Favié B., Goderski G., Kuijpers J., Overdevest I., Rahamat-Langedoen J., Wijsman L., Melchers W.J., Meijer A. Multi-center evaluation of cepheid xpert® xpress SARS-CoV-2 point-of-care test during the SARS-CoV-2 pandemic. J. Clin. Virol. 2020;128 doi: 10.1016/j.jcv.2020.104426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Meyer B., Drosten C., Müller M.A. Serological assays for emerging coronaviruses: challenges and pitfalls. Virus Res. 2014;194:175–183. doi: 10.1016/j.virusres.2014.03.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Ward S., Lindsley A., Courter J., Assa'ad A. Clinical testing for Covid-19. J. Allergy Clin. Immunol. 2020 doi: 10.1016/j.jaci.2020.05.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.M.A.O. Nisreen, A.M. Marcel, L. Wentao, W. Chunyan, H.G. Corine, M.C. Victor, M.L. Mart, S.S. Reina, B. Erwin de, D.C. Felicity, Y. Yazdan, H. Quentin Le, D. Diane, H.-F. Nadhira, B.E.M.R. Chantal, B. Berend-Jan, D. Christian, P.G.K. Marion, L.H. Bart, Severe Acute Respiratory Syndrome Coronavirus 2−Specific Antibody Responses in Coronavirus Disease 2019 Patients, Emerg. Infect. Dis. J. 26(7) (2020). [DOI] [PMC free article] [PubMed]
- 72.H. Lv, N.C. Wu, O.T. Tsang, M. Yuan, R. Perera, W.S. Leung, R.T.Y. So, J.M.C. Chan, G.K. Yip, T.S.H. Chik, Y. Wang, C.Y.C. Choi, Y. Lin, W.W. Ng, J. Zhao, L.L.M. Poon, J.S.M. Peiris, I.A. Wilson, C.K.P. Mok, Cross-reactive antibody response between SARS-CoV-2 and SARS-CoV infections, bioRxiv (2020).
- 73.S. Mahari, A. Roberts, D. Shahdeo, S. Gandhi, eCovSens-ultrasensitive novel in-house built printed circuit board based electrochemical device for rapid detection of nCovid-19 antigen, a spike protein domain 1 of SARS-CoV-2, bioRxiv (2020) 2020.04.24.059204.
- 74.Diao B., Wen K., Chen J., Liu Y., Yuan Z., Han C., Chen J., Pan Y., Chen L., Dan Y., Wang J., Chen Y., Deng G., Zhou H., Wu Y. Diagnosis of acute respiratory syndrome coronavirus 2 infection by detection of nucleocapsid protein. medRxiv. 2020 2020.03.07.20032524. [Google Scholar]
- 75.Weitzel T., Legarraga P., Iruretagoyena M., Pizarro G., Vollrath V., Araos R., Munita J.M., Porte L. Head-to-head comparison of four antigen-based rapid detection tests for the diagnosis of SARS-CoV-2 in respiratory samples. bioRxiv. 2020 doi: 10.1016/j.ijid.2020.05.098. 2020.05.27.119255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Mavrikou S., Moschopoulou G., Tsekouras V., Kintzios S. Development of a Portable, Ultra-Rapid and Ultra-Sensitive Cell-Based Biosensor for the Direct Detection of the SARS-CoV-2 S1 Spike Protein Antigen. Sensors. 2020;20(11) doi: 10.3390/s20113121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Padoan A., Sciacovelli L., Basso D., Negrini D., Zuin S., Cosma C., Faggian D., Matricardi P., Plebani M. IgA-Ab response to spike glycoprotein of SARS-CoV-2 in patients with COVID-19: a longitudinal study. Clin. Chim. Acta. 2020;507:164–166. doi: 10.1016/j.cca.2020.04.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Kohmer N., Westhaus S., Rühl C., Ciesek S., Rabenau H.F. Clinical performance of SARS-CoV-2 IgG antibody tests and potential protective immunity. bioRxiv. 2020 doi: 10.1002/jmv.26145. 2020.05.08.085506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Liu W., Liu L., Kou G., Zheng Y., Ding Y., Ni W., Wang Q., Tan L., Wu W., Tang S., Xiong Z., Zheng S. Evaluation of nucleocapsid and spike protein-based enzyme-linked immunosorbent assays for detecting antibodies against SARS-CoV-2. J. Clin. Microbiol. 2020;58(6) doi: 10.1128/JCM.00461-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Haveri A., Smura T., Kuivanen S., Österlund P., Hepojoki J., Ikonen N., Pitkäpaasi M., Blomqvist S., Rönkkö E., Kantele A., Strandin T., Kallio-Kokko H., Mannonen L., Lappalainen M., Broas M., Jiang M., Siira L., Salminen M., Puumalainen T., Sane J., Melin M., Vapalahti O., Savolainen-Kopra C. Serological and molecular findings during SARS-CoV-2 infection: the first case study in Finland, January to February 2020. Euro Surveill. 2020;25(11) doi: 10.2807/1560-7917.ES.2020.25.11.2000266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Long Q.-X., Deng H.-J., Chen J., Hu J., Liu B.-Z., Liao P., Lin Y., Yu L.-H., Mo Z., Xu Y.-Y., Gong F., Wu G.-C., Zhang X.-X., Chen Y.-K., Li Z.-J., Wang K., Zhang X.-L., Tian W.-G., Niu C.-C., Yang Q.-J., Xiang J.-L., Du H.-X., Liu H.-W., Lang C., Luo X.-H., Wu S.-B., Cui X.-P., Zhou Z., Wang J., Xue C.-J., Li X.-F., Wang L., Tang X.-J., Zhang Y., Qiu J.-F., Liu X.-M., Li J.-J., Zhang D.-C., Zhang F., Cai X.-F., Wang D., Hu Y., Ren J.-H., Tang N., Liu P., Li Q., Huang A.-L. Antibody responses to SARS-CoV-2 in COVID-19 patients: the perspective application of serological tests in clinical practice. medRxiv. 2020 2020.03.18.20038018. [Google Scholar]
- 82.Zhang J., Liu J., Li N., Liu Y., Ye R., Qin X., Zheng R. Serological detection of 2019-nCoV respond to the epidemic: A useful complement to nucleic acid testing. medRxiv. 2020 doi: 10.1016/j.intimp.2020.106861. 2020.03.04.20030916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Cai X.F., Chen J., Hu J.L., Long Q.X., Deng H.J., Fan K., Liao P., Liu B.Z., Wu G.C., Chen Y.K., Li Z.J., Wang K., Zhang X.L., Tian W.G., Xiang J.L., Du H.X., Wang J., Hu Y., Tang N., Lin Y., Ren J.H., Huang L.Y., Wei J., Gan C.Y., Chen Y.M., Gao Q.Z., Chen A.M., He C.L., Wang D.X., Hu P., Zhou F.C., Huang A.L., Liu P., Wang D.Q. A peptide-based magnetic chemiluminescence enzyme immunoassay for serological diagnosis of coronavirus disease 2019 (COVID-19) J. Infect. Dis. 2020 doi: 10.1093/infdis/jiaa243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Montesinos I., Gruson D., Kabamba B., Dahma H., Van den Wijngaert S., Reza S., Carbone V., Vandenberg O., Gulbis B., Wolff F., Rodriguez-Villalobos H. Evaluation of two automated and three rapid lateral flow immunoassays for the detection of anti-SARS-CoV-2 antibodies. J. Clin. Virol. 2020;128 doi: 10.1016/j.jcv.2020.104413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Jia X., Zhang P., Tian Y., Wang J., Zeng H., Wang J., Jiao L., Chen Z., Zhang L., He H., He K., Liu Y. Clinical significance of IgM and IgG test for diagnosis of highly suspected COVID-19 infection. medRxiv. 2020 doi: 10.3389/fmed.2021.569266. 2020.02.28.20029025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Li Z., Yi Y., Luo X., Xiong N., Liu Y., Li S., Sun R., Wang Y., Hu B., Chen W., Zhang Y., Wang J., Huang B., Lin Y., Yang J., Cai W., Wang X., Cheng J., Chen Z., Sun K., Pan W., Zhan Z., Chen L., Ye F. Development and clinical application of a rapid IgM-IgG combined antibody test for SARS-CoV-2 infection diagnosis. J. Med. Virol. 2020 doi: 10.1002/jmv.25727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Pérez-García F., Pérez-Tanoira R., Romanyk J., Arroyo T., Gómez-Herruz P., Cuadros-González J. Alltest rapid lateral flow immunoassays is reliable in diagnosing SARS-CoV-2 infection from 14 days after symptom onset: a prospective single-center study. J. Clin. Virol. 2020;129 doi: 10.1016/j.jcv.2020.104473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Van Elslande J., Houben E., Depypere M., Brackenier A., Desmet S., André E., Van Ranst M., Lagrou K., Vermeersch P. Diagnostic performance of seven rapid IgG/IgM antibody tests and the Euroimmun IgA/IgG ELISA in COVID-19 patients. Clin. Microbiol. Infect. 2020 doi: 10.1016/j.cmi.2020.05.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.de Assis R.R., Jain A., Nakajima R., Jasinskas A., Felgner J., Obiero J.M., Adenaiye O., Tai S., Hong F., Norris P.J., Stone M., Simmons G., Bagri A., Schreiber M., Buser A., Holbro A., Battegay M., Hosimer P., Noesen C., Milton D.K., Davies D.H., Contestable P., Corash L.M., Busch M.P., Felgner P.L., Khan S. Analysis of SARS-CoV-2 antibodies in COVID-19 convalescent blood using a coronavirus antigen microarray. bioRxiv. 2020 doi: 10.1038/s41467-020-20095-2. 2020.04.15.043364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Hedde P.N., Abram T.J., Jain A., Nakajima R., de Assis R.R., Pearce T., Jasinskas A., Toosky M.N., Khan S., Felgner P.L., Gratton E., Zhao W. A modular microarray imaging system for highly specific COVID-19 antibody testing. bioRxiv. 2020 doi: 10.1039/d0lc00547a. 2020.05.22.111518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Tan X., Lin C., Zhang J., Khaing Oo M.K., Fan X. Rapid and quantitative detection of COVID-19 markers in micro-liter sized samples. bioRxiv. 2020 2020.04.20.052233. [Google Scholar]
- 92.Jiang H.-W., Li Y., Zhang H.-N., Wang W., Men D., Yang X., Qi H., Zhou J., Tao S.-C. Global profiling of SARS-CoV-2 specific IgG/IgM responses of convalescents using a proteome microarray. medRxiv. 2020 doi: 10.1038/s41467-020-17488-8. 2020.03.20.20039495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Centers for Disease Control and Prevention, Interim Guidelines for COVID-19 Antibody Testing. https://www.cdc.gov/coronavirus/2019-ncov/lab/resources/antibody-tests-guidelines.html.
- 94.Wang P., Liu L., Nair M.S., Yin M.T., Luo Y., Wang Q., Yuan T., Mori K., Solis A.G., Yamashita M., Purpura L.J., Laracy J.C., Yu J., Sodroski J., Huang Y., Ho D.D. SARS-CoV-2 neutralizing antibody responses are more robust in patients with severe disease. bioRxiv. 2020 doi: 10.1080/22221751.2020.1823890. 2020.06.13.150250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Varnaitė R., García M., Glans H., Maleki K.T., Sandberg J.T., Tynell J., Christ W., Lagerqvist N., Asgeirsson H., Ljunggren H.-G., Ahlén G., Frelin L., Sällberg M., Blom K., Klingström J., Gredmark-Russ S. Expansion of SARS-CoV-2-specific antibody-secreting cells and generation of neutralizing antibodies in hospitalized COVID-19 patients. bioRxiv. 2020 doi: 10.4049/jimmunol.2000717. 2020.05.28.118729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Xiao M., Liu X., Ji J., Li M., Li J., Yang L., Sun W., Ren P., Yang G., Zhao J., Liang T., Ren H., Chen T., Zhong H., Song W., Wang Y., Deng Z., Zhao Y., Ou Z., Wang D., Cai J., Cheng X., Feng T., Wu H., Gong Y., Yang H., Wang J., Xu X., Zhu S., Chen F., Zhang Y., Chen W., Li Y., Li J. Multiple approaches for massively parallel sequencing of HCoV-19 (SARS-CoV-2) genomes directly from clinical samples. bioRxiv. 2020 doi: 10.1186/s13073-020-00751-4. 2020.03.16.993584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Wang M., Fu A., Hu B., Tong Y., Liu R., Gu J., Liu J., Jiang W., Shen G., Zhao W., Men D., Yu L., Deng Z., Li Y., Liu T. Nanopore target sequencing for accurate and comprehensive detection of SARS-CoV-2 and other respiratory viruses. medRxiv. 2020 doi: 10.1002/smll.202002169. 2020.03.04.20029538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.World Health Organization, Laboratory biosafety guidance related to coronavirus disease 2019 (COVID-19). https://www.who.int/publications-detail/laboratory-biosafety-guidance-related-to-coronavirus-disease-2019-(covid-19) (accessed 18 March 2020).
- 99.World Health Organization, Coronavirus disease (COVID-19) advice for the public. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/advice-for-public.
- 100.World Health Organization, Rational use of personal protective equipment for coronavirus disease 2019 (COVID-19). 2020. https://apps.who.int/iris/bitstream/handle/10665/331215/WHO-2019-nCov-IPCPPE_use-2020.1-eng.pdf (accessed 7 March 2020).
- 101.World Health Organization, (Advice on the use of masks in the context of COVID-19. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/advice-for-public/when-and-how-to-use-masks (accessed 5 June 2020)).
- 102.Long Y., Hu T., Liu L., Chen R., Guo Q., Yang L., Cheng Y., Huang J., Du L. Effectiveness of N95 respirators versus surgical masks against influenza: a systematic review and meta-analysis. J. Evid. Based Med. 2020;13(2):93–101. doi: 10.1111/jebm.12381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Jefferson T., Jones M., Al Ansari L.A., Bawazeer G., Beller E., Clark J., Conly J., Del Mar C., Dooley E., Ferroni E., Glasziou P., Hoffman T., Thorning S., Van Driel M. Physical interventions to interrupt or reduce the spread of respiratory viruses. Part 1 - Face masks, eye protection and person distancing: systematic review and meta-analysis. medRxiv. 2020 2020.03.30.20047217. [Google Scholar]
- 104.Corman V.M., Landt O., Kaiser M., Molenkamp R., Meijer A., Chu D.K., Bleicker T., Brünink S., Schneider J., Schmidt M.L. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance. 2020;25(3):2000045. doi: 10.2807/1560-7917.ES.2020.25.3.2000045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Li Q., Guan X., Wu P., Wang X., Zhou L., Tong Y., Ren R., Leung K.S.M., Lau E.H.Y., Wong J.Y., Xing X., Xiang N., Wu Y., Li C., Chen Q., Li D., Liu T., Zhao J., Li M., Tu W., Chen C., Jin L., Yang R., Wang Q., Zhou S., Wang R., Liu H., Luo Y., Liu Y., Shao G., Li H., Tao Z., Yang Y., Deng Z., Liu B., Ma Z., Zhang Y., Shi G., Lam T.T.Y., Wu J.T.K., Gao G.F., Cowling B.J., Yang B., Leung G.M., Feng Z. Early transmission dynamics in Wuhan, China, of novel coronavirus-infected pneumonia. New Engl. J. Med. 2020 doi: 10.1056/NEJMoa2001316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Specific Primers and Probes for Detection 2019 Novel Coronavirus, China National Institute For Viral Disease Control and Prevention: Beijing, 2020.
- 107.Institut Pasteu 2020 Institut Pasteur, France, Real-time RT-PCR assays for the detection of SARS-CoV-2. https://www.who.int/docs/default-source/coronaviruse/real-time-rt-pcr-assays-for-the-detection-of-sars-cov-2-institut-pasteurparis.pdf (accessed 2 March 2020).
- 108.N. Naganori, K. Shirato, Detection of Second Case of 2019-nCoV Infection in Japan; Department of Virology III, National Institute of Infectious Diseases, Japan, 2020.
- 109.Department of Medical Sciences, Ministry of Public Health, Thailand, Diagnostic Detection of Novel Coronavirus 2019 by Real Time RT-PCR, https://www.who.int/docs/default-source/coronaviruse/conventional-rt-pcr-followed-by-sequencing-for-detection-of-ncov-rirl-nat-inst-health-t.pdf (accessed 23 January 2020).
- 110.Detection of 2019 Novel Coronavirus (2019-nCoV) in Suspected Human Cases by RT-PCR, School of Public Health, Hong Kong University: Hong Kong, 2020.
- 111.Liu T., Hsiung J., Zhao S., Kost J., Sreedhar D., Olson K., Keare D., Roche J., Hanson C.V., Press C., Boggs J., Rodriguez-Soto J.P., Montoya J.G., Tang M., Dai H. High-accuracy multiplexed SARS-CoV-2 antibody assay with avidity and saliva capability on a nano-plasmonic platform. bioRxiv. 2020 2020.06.16.155580. [Google Scholar]