Abstract
Viral myocarditis (VMC) is a disease characterized as myocardial parenchyma or interstitium inflammation caused by virus infection, especially Coxsackievirus B3 (CVB3) infection, which has no accurate non-invasive examination for diagnosis and specific drugs for treatment. The mechanism of CVB3-induced VMC may be related to direct myocardial damage of virus infection and extensive damage of abnormal immune response after infection. Non-coding RNA (ncRNA) refers to RNA that is not translated into protein and plays a vital role in many biological processes. There is expanding evidence to reveal that ncRNAs regulate the occurrence and development of VMC, which may provide new treatment or diagnosis targets. In this review, we mainly demonstrate an overview of the potential role of ncRNAs in the pathogenesis, diagnosis and treatment of CVB3-induced VMC.
Keywords: viral myocarditis, non-coding RNA, microRNA, long non-coding RNA, circular RNA
Introduction
Viral myocarditis (VMC) is a non-ischemic inflammatory disease caused by viral infection and related autoimmune disorders, occupying the vast majority of myocarditis. Epidemiological studies have shown that the incidence of VMC is estimated at 1.0–2.2/million (Vos et al., 2015; Olejniczak et al., 2020). VMC usually presents with chest pain, palpitation and dyspnea, and some patients may progress to heart failure, which has bring huge economic burden on patients and societies (Grun et al., 2012). The gold diagnosis standard of VMC is endomyocardial biopsy (EMB), which has a limited clinical application, so the incidence rate of VMC is likely underestimated. So far, there is no specific blood test has been established to diagnose VMC reliably (Olejniczak et al., 2020). Besides, immunosuppressant, antiviral, and circulatory support are the major treatment methods of VMC, but there still are many patients who have progressed to heart failure after these treatments and thus need heart transplantation (Schultz et al., 2009; Pollack et al., 2015; Tschope et al., 2019). Therefore, the exploration of potentially less traumatic diagnosis methods and new therapeutic targets of VMC is important.
As we all know, lots of viruses can cause VMC, including enterovirus, adenovirus, cytomegalovirus, influenza virus, hepatitis C virus, parvovirus, and other viruses (Fairweather et al., 2012; Lobo et al., 2014; Verdonschot et al., 2016; Minhas et al., 2017; Ntusi, 2017; Spartalis et al., 2017). Coxsackievirus B3 (CVB3) is a single positive strand enterovirus, which is the most common pathogen in VMC etiology (Fairweather et al., 2012). Since the first time using CVB3 to induce myocarditis in mice in 1974 by Woodruff, most of the subsequent experimental models of VMC were induced by CVB3 (Gauntt and Huber, 2003). As the same pattern with many other viruses, CVB3 causes VMC through direct damage to host and indirect injury induced by the abnormal immune response of host immune system (Kindermann et al., 2012; Fung et al., 2016). Therefore, the molecular mechanism research of CVB3 induced VMC model can provide a theoretical basis for the clinical diagnosis and treatment.
Non-coding RNAs (ncRNAs) refer to RNAs that are not translated into protein, including transfer RNA (tRNA), ribosomal RNA (rRNA), small nuclear RNA (snRNA), piwi interacting RNA (piRNA), micro RNA (miRNA), long non-coding RNA (lncRNA), and circular RNA (circRNA) (Panni et al., 2019; Smolle et al., 2019; Zhang W. et al., 2019). ncRNAs were once regarded as transcriptional “garbage.” But recent studies suggested that ncRNAs play an important role in viral infection and host antiviral immunity (Chen et al., 2018). It is generally known that a series of pathophysiological changes after the viral infection is determined by the mutual battle between the virus and the host (Barbu et al., 2020). When the virus is in a dominant position, the host is damaged. On the contrary, the host clears the virus through immune response. Therefore, after viral infection, some ncRNAs may change to help the virus escape from the host's immune system, while the other ncRNAs may change to help the host clear the virus (Nicolas, 2017). Future therapies may focus on helping the host clear viruses or decreasing the viral replication by changing the expression of these important ncRNAs. In addition, considering that some ncRNAs of blood will change significantly in the state of disease and have good stability in peripheral circulation, so they have great potential in the field of VMC diagnosis (Regouc et al., 2020; Santos et al., 2020). In this review, we will mainly describe the potential roles of miRNAs, lncRNAs, and circRNAs in the pathogenesis, diagnosis, and treatment of CVB3-induced viral myocarditis.
VMC Model Induced by CVB3
Although there is no detailed epidemiological incidence data of CVB3-induced VMC, CVB3 is a leading causative pathogen of VMC (Gauntt and Huber, 2003). Therefore, many teams have studied the formation structure, life cycle, cell infection preference of CVB3 and the CVB3-induced VMC model (Fairweather et al., 2012; Garmaroudi et al., 2015).
Structure of CVB3
CVB3 virus particle is an icosahedral particle without coating, and its diameter is about 30 nm (Gauntt and Huber, 2003). Capsid protein comprises VP1, VP2, VP3, and VP4. Among these, the first three form the external capsid, while the VP4 located at the inner layer of capsid. The viral genome is a linear single-stranded RNA molecule of about 7.4 kb in size. The genome comprises an open reading frame (ORF) in the middle and an untranslated region on both sides. The ORF encodes 11 proteins (four capsid proteins and 2A, 2B, 2C, 3A, 3B, 3C, 3D) (Garmaroudi et al., 2015). Among them, 2A and 3C are viral proteases, which can induce cell death by cutting the host translation process related proteins such as elF4G and DAP5. Besides, 2B and 2C are involved in RNA synthesis, in which 2B can lead to cardiomyocyte death through largely increasing intracellular calcium level caused by inserting itself into endoplasmic reticulum and Golgi membrane. Moreover, 3A and 3B are vital for viral replication (Peischard et al., 2019). 3D acts as an RNA dependent RNA polymerase, which plays an important role in the synthesis of new viral RNA genome (Garmaroudi et al., 2015).
Life Cycle of CVB3
CVB3 life cycle mainly includes three critical steps: virus entry, viral translation and replication, virus assembly and release (Freimuth et al., 2008; Knowlton, 2008; Pinkert et al., 2009). DAF and CAR are membrane proteins required for CVB3 to enter the cell, and the entry mode is internalization (Bergelson et al., 1995; Shafren et al., 1995; Bewley et al., 1999; Coyne and Bergelson, 2006). After virus entry, viral genome transcribes, and translates various functional and structural proteins. The viral RNAs are copied by RNA polymerase 3D protein, and then the copied genome is transferred to the structural protein to assemble complete viral particles. Finally, the virus promotes host cell death to facilitate the release of virus particles.
Tissue Specificity and Gender Preference of CVB3 Infection
CVB3 susceptible cells are mainly distributed in the heart and pancreas, while the lung and kidney are relatively tolerant (Cheung et al., 2005; Harvala et al., 2005). In addition, CVB3 infection has a difference between males and females. Clinically, two-thirds of cases of myocarditis occur in men. It was also found that male mice were more likely to develop CVB3-induced myocarditis than females (Gauntt and Huber, 2003). The sex difference may be related to the different immune responses induced by sex hormones.
Mechanism of CVB3-Induced Myocarditis
As mentioned before, the pathogenesis of CVB3-induced VMC model includes direct myocardial injury by virus and indirect myocardial injury mediated by the immune system. In the process of direct myocardial injury, the functional proteins 2A and 3C of CVB3 block the translation process of host proteins and hinder the metabolism of host cells by cutting the important proteins elF4G and DAP5 (Chau et al., 2007; Lewis et al., 2008). Besides, the breakdown of the endoplasmic reticulum and Golgi apparatus leads to cell death during the process of virus release (van Kuppeveld et al., 1997). In the process of indirect myocardial injury, the virus promotes the release of inflammatory factors by activating innate immune cells (macrophages, NK cells) and acquired immune cells (Th cells). Moreover, the continuous synthesis of virus products in infected host cells can induce the production of cytokines, inflammatory factors and chemokines, thus stimulating the cardiac infiltration of inflammatory cells. In this process, IL-1α/−5 /−6 /−7 can be massively synthesized by infected cardiomyocytes, while IL-1β/−4 /−6 /−8, TNF α, and TNF β can be massively synthesized and released by inflammatory infiltrated leukocytes (Gauntt and Huber, 2003). Subsequently, a large number of inflammatory factors will further promote myocardial injury.
The Role of miRNAs in CVB3-Induced VMC
MiRNAs are a type of endogenous single-stranded RNAs constructed by about 20–25 nucleotides, which can regulate gene expression by binding to the target mRNAs (Dong et al., 2013; Ha and Kim, 2014; Jonas and Izaurralde, 2015; Bracken et al., 2016; Gebert and MacRae, 2019; Treiber et al., 2019). With the deeper and deeper research in recent years, the synthesis mechanism and the function of miRNAs are clarified. The synthesis process of miRNAs contains several steps: transcription, cleavage, export, further cleavage, strand selection, and interaction with mRNA. First, the gene encoding miRNA is transcribed into primary miRNA transcripts which named pri-miRNA with cap structure and polyadenylated tail by RNA polymerase II or RNA polymerase III (Lee et al., 2004). Then, Pri-miRNA is cleaved into about 70–80 nucleotide precursor miRNAs (pre-miRNAs) with hairpin structure by a protein complex consisted of Drosha and its cofactor Pasha (Lee et al., 2003; Denli et al., 2004; Gregory et al., 2004; Han et al., 2004; Landthaler et al., 2004). Subsequently, pre-miRNAs are transported from the nucleus to the cytoplasm by the transporter exportin-5 (Yi et al., 2003; Bohnsack et al., 2004; Kim, 2004; Lund et al., 2004). The pre-miRNAs are cut by Dicer enzyme to form 21 to 25 nt double-stranded miRNA (miRNA duplex) (Grishok et al., 2001; Ketting et al., 2001). After the unwinding of the double-strand, one strand of miRNA duplex was degraded and the other one became of the mature miRNA which then directed into RNA induced silencing complex (RISC) (Gregory et al., 2005; Kobayashi and Tomari, 2016). At last, mature miRNAs regulate target mRNA expression by binding to the target mRNA. When the miRNAs and target mRNAs are completely complementary, miRNAs can lead to degradation of target mRNAs, and the binding site is usually in the coding region of mRNAs (Lai, 2002). By contrast, when they are not completely complementary, miRNAs can inhibit the translation process by binding to the 3′ untranslated region (3′ UTR) of target mRNAs, and finally affect the expression of protein (Bartel, 2004). Recently, it has been found that some miRNAs can also promote gene expression in some cases (Xiao et al., 2017). In the course of VMC, miRNAs have been proved to be differentially expressed, which may regulate the disease development by affecting the life cycle of virus and host immune (Figure 1).
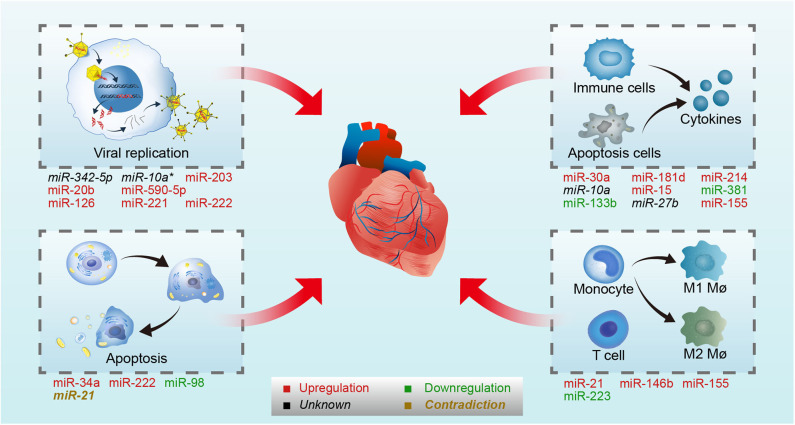
Figure 1.
MiRNAs involved in CVB3-induced VMC. CVB3 induced VMC through direct myocardial injury by virus and indirect myocardial injury mediated by the immune system. On one hand, miRNAs regulate viral life cycle (viral replication and host cell apoptosis) by targeting virus genome or host genes in CVB3-induced VMC. On the other hand, miRNAs control immune response by targeting host genes in CVB3-induced VMC. On the upper-left panel, miR-342-5p, /-10a*, /-203, /-20b, /-590-5p, /-126, /-221, and /- 222 involved in viral replication by targeting viral genome or host genes. On the lower-left panel, miR-34a, /-222, /-98, and /-21 involved in host apoptosis. On the upper-right panel, miR-30a, /-181d, /-214, /-10a, /-15, /-381, /-133b, /-27b, and /-155 involved in cytokines production. On the lower-right panel, miR-21, /-146b, /-155, and /-223 involved in immune cells differentiate and activation. Upregulated microRNAs in CVB3 infection are in red, while the downregulated are in green. The microRNA whose expression level were not stated in the original article is in black, and the microRNA whose expression level were contradictory in different studies is in brown. Mø, macrophages.
The Influence of miRNAs on the Life Cycle of CVB3
Viral infection initiates a battle between virus and host. As a type of micro-organism that completely depends on cells to survive, the integrity of life cycle is of great significance for viral infection and infection expansion. Viruses complete their life cycle depends on controlling the survival of host cells. Before the viral load reaches a certain threshold in the cell, the virus inhibits the apoptosis of the host cell, to facilitate viral replication, translation, and assembly. Once viral load reaches the threshold, the virus promotes cell apoptosis to release the amplified virus particles. At the same time, the body is also making efforts to fight against the virus. In the acute phase of virus infection, the body can reduce the infection by inhibiting virus replication. Recent studies have shown that CVB3 can regulate viral replication and host cell apoptosis by affecting miRNAs expression (Chen et al., 2018).
MiRNAs on the Replication of CVB3
It has been proven that the replication process is indispensable in the viral life cycle, so inhibiting this process may help to block VMC in the early stages. Undoubtedly, many miRNAs are involved in this process (Table 1; Sodroski et al., 2019; Su et al., 2020). MiRNAs have two patterns to regulate viral replication: one is the direct pattern, and the other is the indirect pattern.
Table 1.
MiRNAs involved in viral replication.
| MiRNA | Target | MiRNAs expression | Description | Process | Disease model | References |
|---|---|---|---|---|---|---|
| miR-342-5p | CVB3 2C region | Unknown | miR-342-5p inhibits CVB3 replication by targeting 2C-coding region | Viral replication | In vitro: CVB3 (Woodruff strain) infected Hela cells | Wang et al., 2012 |
| miR-10a* | CVB3 3D region | Unknown | miR-10a* promotes CVB3 replication by targeting 3D-coding region | Viral replication | In vitro: CVB3 (pKMS1) infected Hela cells | Tong et al., 2013 |
| miR-203 | ZFP-148 | Upregulation in CVB3 infected Hela cells and A/J mice | PKC/AP-1 activation promotes miR-203 expression miR-203 enhances CVB3 replication by targeting ZFP-148 |
Viral replication | In vitro: CVB3 (Gauntt strain) infected Hela cells; In vivo: CVB3 infected A/J mice | Hemida et al., 2013 |
| miR-20b | ZFP-148 | Upregulation in CVB3 infected BALB/c mice | miR-20b inhibits ZFP-148 expression and thus enhance CVB3 replication miR-20b promotes cardiomyocytes survival |
Viral replication/cell survival | In vivo: CVB3 (Nancy strain) infected BALB/c mice | Xu et al., 2017 |
| miR-590-5p | SPRY1 | Upregulation in extracellular vesicles released by CVB3 infected HL-1 cells | miR-590-5p enhances CVB3 replication by targeting SPRY1 | Viral replication | In vitro: CVB3 (pMKS1) infected HL-1 cells | Germano et al., 2019 |
| miR-126 | SPRED1, LPR6, WRCH1 | Upregulation in CVB3 infected Hela cells | miR-126 enhances CVB3 replication by targeting SPRED1 (ERK1/2 pathway) miR-126 promotes cell death and viral release by targeting LPR6 and WRCH1 (Wnt/β0catenin pathway) |
Viral replication/cell death | In vitro: CVB3 infected Hela cells | Ye et al., 2013 |
| miR-221/-222 | ETS1/2, IRF2, BCL2L11, TOX, BMF, and CXCL12 | Upregulation in CVB3 infected C3H and C57Bl6N mice | miR-221/-222 inhibition increases viral replication and immune cell infiltration miR-221/-222 targets ETS1/2, IRF2, BCL2L11, TOX, BMF, and CXCL12 TOX, CXCL12, and IRF2 inhibition suppressed inflammatory response |
Viral replication/ inflammation |
In vitro: CVB3 (Nancy strain) infected neonatal rat cardiomyocytes; In vivo: CVB3 (Nancy strain) infected C3H and C57Bl6N mice | Corsten et al., 2015 |
ZFP-148, zinc finger protein-148; SPRY1, sprout 1; SPRED1, sprouty-related EVH1 domain-containing protein 1; LRP6, lipoprotein receptor-related protein 6; WRCH1, Wnt responsive Cdc42 homolog 1; IRF2, interferon regulatory factor 2; BCL2L11, Bcl-2-like protein 11; TOX, Thymocyte selection-associated high-mobility group box; BMF, BCL-2-modifying factor; CXCL12, CXC chemokine ligand 12.
The direct pattern means that miRNAs directly target the viral genome sequence to inhibit or promote viral replication. Wang et al. have found that miR-342-5p may target the 2C-coding region of the viral genome by bioinformatic analysis. Further experiments confirmed that miR-342-5p can directly inhibit viral replication by targeting the 2C-coding region of CVB3 (Wang et al., 2012). Unlike with most negatively regulated traditional miRNAs, some miRNAs can promote mRNA expression. Wang's group found that miR-10a* directly promotes the viral biosynthesis by targeting 3D-coding region (nt6818-nt6941) of CVB3. Additionally, miR-10a* is abundant in heart of Balb/c mice which indicated that miR-10a* may affect VMC via inducing CVB3 replication (Tong et al., 2013). These findings implied that miRNAs might be a useful treatment by directly limiting viral replication.
The indirect pattern means that miRNAs indirectly control viral replication by targeting the host genome. Many genes, such as zinc finger protein (ZFP)-148, sprout 1 (SPRY1), sprouty-related EVH1 domain-containing protein 1 (SPRED1), and E-twenty six 1/2 (ETS1/2), regulate replication in viral infection. Studies have shown that the upregulation miRNAs, such as miR-203 and miR-20b, targeted ZFP-148 to promote viral replication and cell survival in CVB3 induced VMC (Hemida et al., 2013; Xu et al., 2017). The miRNAs secreted by cells may also act as complex regulators on viral infection. Germano et al. showed that the levels of miR-590-5p increased in vesicles shed from infected cells, and this miRNA prolong viral replication and enhance cell survival via targeting SPRY1 (Germano et al., 2019). Besides, extracellular signal-regulated kinase 1/2 (ERK1/2) and Wnt/β-catenin pathways were involved in the VMC. CVB3 infection promotes ERK1/2 phosphorylation, which activated ETS transcriptional protein activity, causing miR-126 expression upregulation. Then miR-126 blocked the target gene SPRED1, lipoprotein receptor-related protein 6 (LRP6) and Wnt responsive Cdc42 homolog 1 (WRCH1) to promote the crosstalk with ERK1/2 and Wnt/β-catenin pathways. Finally, the ERK1/2 pathway accelerated the viral replication, and the Wnt/β-catenin pathway promoted virus-mediated cell death and viral release (Ye et al., 2013). Therefore, CVB3 infection can induce the upregulation of miR-126 and promote virus replication and release by regulating ERK1/2 and Wnt/β-catenin pathways.
The body did not await its doom in CVB3 infection. Corsten et al. found that CVB3 infection promoted the miR-221/-222 expression in cardiac tissue. Further study demonstrated that miR-221/-222 could inhibit ETS1/2, interferon regulatory factor 2 (IRF2), Bcl-2-like protein 11 (BCL2L11), Thymocyte selection-associated high-mobility group box (TOX), BCL-2-modifying factor (BMF), and CXC chemokine ligand 12 (CXCL12) As we all know, those target genes are important in viral replication and inflammation (Welsh, 1986; Bosselut et al., 1990; Sieweke et al., 1998; Posada et al., 2000; Kühl et al., 2003; Zhan et al., 2005; Russell and Garrett-Sinha, 2010; Cheng et al., 2011; Johansson et al., 2011; Tanaka-Nakanishi et al., 2014; Okuyama et al., 2018). The in-vitro study confirmed that miR-221/-222 inhibited inflammation through those IRF2, CXCL12 and TOX in CVB3 infection (Corsten et al., 2015). This study indicated that the body tried to eliminate the virus and protect heart by increasing the expression of miR-221/-222 reactively during CVB3 infection.
All in all, these studies confirmed that the expression of miR-342-5p, miR-10a *, miR-203, miR-20b, miR-590-5p, and miR-126 could promote viral replication, and the body could also reactively promote the expression of miR-221/-222 to reduce viral replication. In the future, we can reduce CVB3 replication by targeting these miRNAs in VMC treatment.
The Effect of miRNAs on Host Cell Apoptosis During CVB3 Infection
The virus completes its life cycle by controlling apoptosis (Kvansakul, 2017). In the early stage of viral infection, the virus promotes cell survival by inhibiting cell apoptosis to ensure sufficient amplification. In the later stage of infection, the virus promotes cell apoptosis to facilitate release and spread. MiR-34a, miR-222, miR-98, and miR-21 play an important role in cell apoptosis (Zhou et al., 2017; Tong et al., 2019; Zheng et al., 2019; Shen et al., 2020). Therefore, many researchers focus on the role of miRNAs in virus-induced cell apoptosis. Jiang et al. found that inhibition of miR-34a expression largely decreased the level change of apoptosis-related protein, including Bax and Bcl-2 (Jiang et al., 2019). Sirtuin 1 (SIRT1) was a validated target gene of miR-34a (Castro et al., 2013; Yang et al., 2015; Carloni et al., 2016). Then the researchers confirmed miR-34a act as a pro-apoptotic molecule in VMC via targeting the SIRT1-p53 signaling pathway (Jiang et al., 2019). In contrast, miR-222, miR-98, and miR-21 may act as anti-apoptotic factors in VMC (He et al., 2013; Zhang B. Y. et al., 2016; Zhang X. et al., 2019). Zhang et al. found that the expression of adenosine deaminase, RNA-specific (ADAR1) and miR-222 was increased in CVB3 induced VMC model, and the study in vitro showed that ADAR1 combined Dicer increased cell viability by inducing miR-222 synthesis which decreased the confirmed target phosphatase and tensin homolog (PTEN) expression (Zhang X. et al., 2019). Previous studies have proven that PTEN is an apoptosis protein (Dupont et al., 2002; Lin et al., 2007; Cheng et al., 2009). Hence, ADAR1-Dicer Complex may inhibit cell apoptosis by miR-222 in CVB3 induced VMC. Additionally, MiR-98 decreased cell apoptosis by targeting the FAS/FASL gene (Zhang B. Y. et al., 2016), and miR-21 reduced cell apoptosis by targeting programmed cell death 4 (PDCD4) (He et al., 2013), a well-known apoptosis gene (Gaur et al., 2011; Stagakis et al., 2011; Junker et al., 2015). Furthermore, another study confirmed that miR-21 reduced cell apoptosis by targeting mitogen-activated protein kinase kinase 3 (MAP2K3) (He et al., 2019). Interestingly, although both studies showed that miR-21 has an anti-apoptosis effect, the expression trend of miR-21 was not consistent after CVB3 infection in different studies (Table 2). The conflicting results may be related to the different viral strains used and different sample acquisition time in two studies. All in all, these studies show that the virus can complete its life cycle by influencing the level of miRNAs to regulate host cell apoptosis. In the future, we can target these miRNAs to destroy the life cycle of the virus to reduce body injury.
Table 2.
MiRNAs involved in cell apoptosis.
| MiRNA | Target | MiRNAs expression | Description | Process | Disease model | References |
|---|---|---|---|---|---|---|
| miR-34a | SIRT1 | Upregulation in CVB3 infected neonatal rat cardiomyocytes | miR-34a promotes cell apoptosis via SIRT/p53pathway | Apoptosis | In vitro: CVB3 (Nancy strain) infected neonatal rat cardiomyocytes | Jiang et al., 2019 |
| miR-222 | PTEN | Upregulation in CVB3 infected H9C2 cells, primary cardiac cells, and BALB/c mice | ADAR1 combined Dicer induced miR-222 synthesis miR-222 decreases PTEN expression ADAR1 increases cell viability via regulating PTEN expression |
Apoptosis | In vitro: CVB3 infected H9C2 and primary cardiac cells; In vivo: CVB3 infected BALB/c mice | Zhang X. et al., 2019 |
| miR-98 | FAS/FASL | Downregulation in blood of VMC patients | miR-98 decreases and FAS/FASL increased in VMC patients miR-98 inhibits apoptosis via targeting FAS/FASL |
Apoptosis | VMC patients | Zhang B. Y. et al., 2016 |
| miR-21 | PDCD4 | Downregulation in CVB3 infected BALB/c mice | miR-21 inhibits cell apoptosis via PDCD4 | Apoptosis | In vivo: CVB3 (Nancy strain) infected BALB/c mice | He et al., 2013 |
| miR-21 | MAP2K3 | Upregulation in CVB3 infected BALB/c mice and Hela cells | miR-21 inhibits cell apoptosis via MAP2K3/p38 MAPK pathway miR-21 does not affect CVB3 replication |
Apoptosis | In vitro: pCVB3M strain infected Hela cells; In vivo: pCVB3M strain infected BALB/c mice | He et al., 2019 |
SIRT1, Sirtuin 1; ADAR1, adenosine deaminase, RNA-specific; PTEN, phosphatase and tensin homolog; PDCD4, programmed cell death 4; MAP2K3, mitogen-activated protein kinase kinase 3.
MiRNAs on Host Immune and Inflammatory Response
The CVB3-induced immune response is very complex, including the activation of innate immunity and acquired immunity. In the early stage of CVB3 infection, innate immune-related cells (macrophages and NK cells) infiltrate the heart, producing chemokines and Interferon (IFNs). CVB3 infection can promote the polarization of M2 macrophages toward M1 macrophages, which increases the inflammation (Zhang Y. et al., 2016; Gou et al., 2018). Besides, the 2A protein of CVB3 inhibits the production of IFNs by targeting TLR3 and MDA5/RIF-I pathways, which are vital pathways in host antiviral immunity (Lind et al., 2016). In the subsequent stage, acquired immune-related cells (Th1, Th2, and Th17 cells) are activated to produce cytokines and inflammatory factors (Gauntt and Huber, 2003). Th1 cells mainly produce IFN-γ, which can promote inflammation. Moreover, Th2 cells mainly produce IL-10, which has an anti-inflammatory effect. Furthermore, Th17 cells mainly produce IL-17, which plays a key role in promoting CVB3 induced infection (Garmaroudi et al., 2015). In recent years, researchers also focus on the field that miRNAs participate in the inflammation process by activating innate and acquired immune responses during CVB3 infection (Table 3).
Table 3.
MiRNAs involved in immune and inflammation.
| MiRNA | Target | MiRNAs expression | Description | Process | Disease model | References |
|---|---|---|---|---|---|---|
| miR-21, miR-146b | ROR-γt | Upregulation in CVB3 infected BALB/c mice | Inhibition of miR-21 or miR-146b decreased the proportion of Th17 cells via targeting ROR-γt miR-21 or miR-146b decreased IL-17, IL-6, and TGF-β levels |
Th17 cells differentiation | In vivo: CVB3 (Nancy strain) infected BALB/c mice | Liu et al., 2013 |
| miR-155 | PU.1 | Upregulation in CVB3 infected C3H mice, C57Bl6 mice and VMC patients | Inhibition of miR-155 relieved cardiac injury by inhibiting macrophage infiltration and T cell activation PU.1 is the direct target gene of miR-155 Inhibition of miR-155 does not affect viral replication |
Macrophage infiltration and T cell activation | In vivo: CVB3 infected C3H mice and C57Bl6 mice; VMC patients | Corsten M. et al., 2012 |
| miR-155 | – | Upregulation in CVB3 infected miR-155 knock out C57Bl6 mice | Silencing miR-155 suppresses M1 macrophages polarization, and promotes macrophages toward to M2 phenotype | Macrophages polarization | In vivo: CVB3 (Nancy strain) infected C57Bl6 mice | Zhang Y. et al., 2016 |
| miR-155 | RelA | Upregulation in myocardial tissue of VMC patients | miR-155 overexpression decreases inflammatory factor to reduce cardiac injury by targeting RelA (NF-κB pathway) | NF-κB inflammatory pathway | In vivo: CVB3 (Nancy strain) infected BALB/c mice; VMC patients | Bao and Lin, 2014 |
| miR-223 | PKNOX1 | Downregulation in CVB3 infected BALB/c mice | miR-223 overexpression suppresses M1 macrophages polarization, and promotes macrophages toward to M2 phenotype via targeting PKNOX1 | Macrophages polarization | In vivo: CVB3 (Nancy strain) infected BALB/c mice | Gou et al., 2018 |
| miR-30a, miR-181d | SOCS3 | Upregulation in the blood of VMC patients and CVB3 infected Hela cells | miR-30a and miR-181d enhanced the level of IL-6 by targeting SOCS-3 | Proinflammatory factor | In vitro: CVB3 (Nancy strain) infected Hela cells; VMC patients | Fan et al., 2019 |
| miR-214 | ITCH | Upregulation in the right ventricular septal specimens of VMC | miR-214 increases TNF-α, IL-1β, MCP-1, and IL-6 by targeting ITCH (NF-κB pathway) | Proinflammatory factor | In vitro: CVB3 infected Hela cells; VMC patients | Chen et al., 2015 |
| miR-10a | ITCH | Unknown | Allele A of rs3809783 in pri-miR-10a coding region in the VMC population was related to VMC occurrence miR-10a promoted IL-6 expression by targeting ITCH (NF-κB pathway) |
Proinflammatory factor | In vitro: CVB3 infected Hela cells; VMC patients | Liao et al., 2015 |
| miR-15 | NLRX-1 | Upregulation in CVB3 infected H9C2 cells | miR-15 promotes the expression of IL-1β, IL-6, and IL-18 by targeting NLRX-1 to activate NLRP3 inflammasomes miR-15 inhibition suppresses CVB3-induced cell apoptosis |
Proinflammatory factor | In vitro: CVB3 (Nancy strain) infected H9C2 cells; | Tong et al., 2020 |
| miR-381 | COX-2 | Downregulation in the blood of children with VMC and CVB3 infected BALB/c mice | miR-381 decreases myocardial injury via targeting COX-2 | Anti-inflammatory factor | In vivo: CVB3 infected BALB/c mice; VMC children | Zhang Y. et al., 2018 |
| miR-133b | Rab27B | Downregulation in the blood of VMC patients | miR-133b reduced IL-6 and TNF-α by directly targeting Rab27B | Anti-inflammatory factor | In vitro: CVB3 infected cardiomyocytes; VMC patients | Zhang et al., 2017 |
| miR-27b | MCP1 | Unknown | miR-27b inhibited the level of MCP1 in IL-17 treated H9C2 cells | Anti-inflammatory factor | In vitro: IL-17 treated H9C2 cells | Huang et al., 2016 |
ROR-γt, retinoid-related orphan receptor gamma-t; PU.1, purine-rich box 1; RelA, v-rel avian reticuloendotheliosis viral oncogene homolog A; PKNOX1, PBX/knotted 1 homeobox 1; SOCS3, suppressor of cytokine signaling 3; ITCH, itchy E3 ubiquitin-protein ligase; NLRX-1, NLR family member X1; COX-2, cytochrome c oxidase subunit II; Rab27B, RAB27B, member RAS oncogene family; MCP-1, macrophage chemo-attractant protein-1; TNF-α, tumor necrosis factor-alpha; IL, interleukin.
MiRNAs on T Cells and Macrophages During CVB3 Infection
Monocytes-macrophages and T cells are representative cells in innate and adaptive immune in VMC, respectively (Gauntt and Huber, 2003). Monocytes differentiate into macrophages after they reach the tissues, among which M1 macrophages have the effect of promoting inflammation, while M2 macrophages have the effect of anti-inflammation (Huang et al., 2018a). Th cells (mainly including Th1, Th2, and Th17 cells) also play a role in CVB3 infection (Garmaroudi et al., 2015). Previous studies have shown that IFN-γ knockout mice are free from VMC after CVB3 treatment, while IFN-γ overexpression is susceptible (Huber and Sartini, 2005). Therefore, Th1 cells can secrete IFN-γ to promote the development of VMC. Besides, IL-10 is a vital anti-inflammatory factor, and Th2 cells mainly secrete IL-10 to act as an anti-inflammatory role. As we all know, Th17 cells can secrete IL-17. Previous studies have found that inhibition of IL-17 by neutralizing antibodies can reduce viral replication and myocardial damage in myocarditis, suggesting that Th17 cells are important pro-inflammatory cells (Fan et al., 2011). Proinflammatory factors, such as miR-21 and miR-146b, have been identified that were increased in VMC. Inhibition of miR-21 or miR-146b decreased Th17 cells and relieved myocardial injury by targeting retinoid-related orphan receptor gamma-t (ROR-γt) (Liu et al., 2013). Moreover, miR-155 also acts as a pro-inflammatory factor in some cases and it was upregulated in CVB3 induced VMC model and human myocarditis specimen (Corsten M.F. et al., 2012; Zhang Y. et al., 2016). A research conducted by Corsten et al. illustrated that inhibition of miR-155 relieved cardiac injury by inhibiting macrophage infiltration and T cell activation, and the direct target of miR-155, purine-rich box 1 (PU.1), may function in this process (Corsten M.F. et al., 2012). A further study directed by Zhang et al. showed that inhibition of miR-155 increased M2 macrophages and decreased M1 macrophages to attenuate cardiac inflammatory (Zhang Y. et al., 2016). However, Bao et al. found that miR-155 acts as an anti-inflammatory factor to reduce cardiac injury by targeting v-rel avian reticuloendotheliosis viral oncogene homolog A (RelA) (Bao and Lin, 2014), a vital component of nuclear factor kappa-B (NF-κB) inflammatory pathways (Gasparini et al., 2014; Mukherjee et al., 2015; Kabacaoglu et al., 2019). These contradictory results may be due to the use of different models. Moreover, miR-223, which also acts as an anti-inflammatory factor, has been verified that was decreased in VMC. Forced expression of miR-223 activated M1 macrophages toward M2 anti-inflammatory phenotype through targeting PBX/knotted 1 homeobox 1 (PKNOX1), and thus relieving myocardial inflammation (Gou et al., 2018). Therefore, CVB3 infection can promote immune cells-mediated inflammation by regulating the expression of miRNAs. In the future, we can reduce the degree of myocardial inflammation by targeting these miRNAs.
MiRNAs on Cytokines Expression After CVB3 Infection
CVB3-induced cardiomyocytes apoptosis and activated immune cells release many cytokines, such as Interferons (IFNs), tumor necrosis factor-alpha (TNF-α), Interleukin-1 beta (IL-1β), Interleukin-6 (IL-6), Interleukin-10 (IL-10), Interleukin-12 (IL-12) (Kanda et al., 1996; Miyamoto et al., 2001; Matsumori et al., 2004; Lindner et al., 2014; Wei et al., 2019; Yang et al., 2019). Some of these cytokines (Type I IFNs, Type II IFNs, and IL-10, etc.) are anti-inflammatory factors while others (TNF-a, IL-6, and IL-12, etc.) are proinflammatory factors (Corsten M. et al., 2012). MiRNAs also regulated the expression of cytokines during CVB3 infection. An investigation conducted by Fan et al. found that miR-30a and miR-181d increased in CVB3 induced VMC mice model. Further research confirmed that those miRNAs enhanced the level of IL-6 by targeting SOCS-3 (Fan et al., 2019). Analogously, miR-214, increased in VMC, enhanced the level of IL-1β, IL-6, macrophage chemo-attractant protein (MCP)-1, and TNF-α by targeting itchy E3 ubiquitin-protein ligase (ITCH) (Chen et al., 2015), an NF-κB pathway suppressor (Chen et al., 2001; Shembade et al., 2008; Perez et al., 2015). ITCH is also the direct target of miR-10b. Liao et al. found that allele A of rs3809783 in pri-miR-10a coding region in the VMC patients was related to VMC occurrence, and this mutation site reduced mature miR-10a expression. Further research identified that miR-10a promoted IL-6 expression by targeting ITCH (Liao et al., 2015). MiRNAs may promote the development of inflammation through the activation of inflammasomes. Tong et al. found that miR-15 was up-regulated after CVB3 infection. This up-regulated miRNA promotes the expression of IL-1β, IL-6, and IL-18 by targeting NLR family member X1 (NLRX-1) to activate NLRP3 inflammasomes (Tong et al., 2020). On the contrary, miR-381, miR-133b, and miR-27b act as anti-inflammatory factors. MiR-381 relieved myocardial injury by targeting cytochrome c oxidase subunit II (COX-2) (Zhang Y. et al., 2018), miR-133b reduced IL-6 and TNF-α by directly targeting RAB27B, member RAS oncogene family (Rab27B) (Zhang et al., 2017), and miR-27b inhibited the level of MCP1 in IL-17 treated H9C2 cells (Huang et al., 2016). These results indicate that CVB3-induced inflammation was regulated by miRNAs targeted inflammatory signaling pathways.
Potentiality of miRNAs in the Treatment of CVB3-Induced VMC
As mentioned above, after CVB3 infection, miRNAs function by influencing the viral life cycle and immune response. Therefore, miRNAs may be potential therapeutic targets for VMC. For example, miRNAs such as miR-342-5p, miR-10a *, miR-203, miR-20b, miR-590-5p, and miR-126 can directly or indirectly affect virus replication. We can design inhibitors or analogs of these miRNAs to inhibit CVB3 replication, thereby reducing the viral load in infected host cells. In addition, miRNAs such as miR-21, miR-146b, miR-233, miR-155, miR-27b, miR-30a, miR-181d, miR-214, miR-10a, miR-381, and miR-133b can regulate the CVB3-induced inflammatory response by affecting the differentiation of immune cells or the release of cytokines, so we can design inhibitors or analogs of miRNAs to reduce myocardial inflammation and damage. Clinical trials of miRNAs based therapeutics have been used in other diseases, so it may become a reliable treatment for VMC in the future (Beg et al., 2017).
Although miRNAs-based therapeutics might be a promising treatment in VMC, there are still some problems. First, finding the method to help the miRNAs targeted transport to the heart is critical. MiRNAs can reach all organs of the body through blood circulation, which may cause other organs injury under miRNAs untargeted transporting. Second, one miRNA can target multiple genes at the same time, and one gene may be controlled by multiple miRNAs. MiRNAs may hence have other unpredictable side effects. Thirdly, a large number of miRNAs are only studied in animals. As we all know, mice and the human body have great differences. MiRNAs with functions in mice may not work in human body. Finally, miRNAs are also expressed in physiological state, so the determination of its therapeutic dose is also a difficulty. Therefore, further research may be needed to solve these problems.
Potentiality of miRNAs in VMC Diagnosis
In VMC, miRNAs are differentially expressed not only in cells but also in blood. The stability of miRNAs in the blood can be maintained by binding RNA binding proteins and enveloping them within extracellular vesicles (Santos et al., 2020). There has been other disease using miRNAs as candidate diagnostic tool (Chen et al., 2019), but VMC has not yet been. Therefore, miRNAs in the blood may be used as a disease marker for VMC. Expression change of miRNAs in the circulation may help us assess the severity of VMC. Wang et al. evaluated the predictive value of miR-1 and miR-146b in VMC. MiR-1 decreased significantly in VMC, which was negatively correlated with the left ventricular shortening fraction and left ventricular ejection fraction. MiR-146b was significantly up-regulated in VMC, which was positively correlated with those two indexes (Wang D. et al., 2016). Therefore, miR-1 and miR-146b may be potential disease markers of VMC. Moreover, Corsten et al. Found that miR-208b and miR-499 were significantly increased in VMC, and correlated with blood troponin levels (Corsten et al., 2010). Furthermore, in a study on the correlation between miRNAs and child VMC pathology, the level of miRNA-208a increased significantly in the acute phase of the disease. Compared with the chronic phase, the level of miRNA-21 in the acute phase was also significantly higher (Goldberg et al., 2018). These studies indicated that miR-208a, miR-21, miR-1, miR-146b, miR-208b, and miR-499 will be promising biomarkers for VMC diagnosis. Compared with EMB, circulating miRNAs have the characteristics of less trauma, low cost, good sensitivity, and high specificity, so they may be used as potential disease markers for VMC differential diagnosis in the future. However, as a disease marker, miRNAs also meet some challenges. First of all, the quantitative methods, units of measurement and thresholds of miRNAs are different in different laboratories, which may limit their further use. Secondly, different methods of extracting miRNAs in different laboratories may produce contradictory results. Thirdly, the level of miRNAs has been fluctuating in different stage of VMC, so the point in time of obtaining miRNAs still needs further study.
The Role of lncRNAs in VMC
LncRNAs belongs to non-coding RNAs with a length longer than 200 nucleotides (Li J. et al., 2018; Barangi et al., 2019; Chi et al., 2019). In many ways, lncRNAs are similar to mRNAs. However, compared with mRNAs, the transcription number and species conservation of lncRNA is low. The secondary structure of lncRNA, such as hairpin structure and stem ring structure formed by post-transcriptional modification, interacts with chromatin and protein to function. Based on the relative position to the protein-coding gene, lncRNAs can be divided into the following five categories: bidirectional lncRNA, the intron lncRNA, intergenic lncRNA, sense lncRNA, and antisense lncRNA. LncRNAs function mainly through chromatin remodeling, transcription and post-transcriptional regulation to influence gene expression (Atianand and Fitzgerald, 2014; Quinn and Chang, 2016; Dykes and Emanueli, 2017; Huang et al., 2018b). Previous studies have found that lncRNAs are involved in many pathophysiological processes, such as growth and development, immune, inflammation and tumor (Li Y. et al., 2018; Alessio et al., 2019; Chi et al., 2019; Wang et al., 2019; Zhang and Wang, 2019; Zhang K. et al., 2019).
Liu et al. confirmed that lncRNAs were differentially expressed in child acute fulminant myocarditis. Among them, 1645 lncRNAs were up-regulated and 1456 lncRNAs were down-regulated. Further bioinformatic analysis showed that T cell activation, T cell receptor complex, negative regulation of complement activation, immune response and T-helper 17 cell differentiation process may participate in the progress of VMC (Liu Q. et al., 2019). In addition, Zhang et al. recently found that lncRNA Ak085865 can promote the expression of anti-inflammatory M2 macrophages, reduce the number of proinflammatory M1 macrophages. LcRNA Ak085865 knockout mice were more susceptible to CVB3 induced VMC, suggesting that lncRNA Ak085865 may play a vital role in the pathogenesis of VMC by affecting macrophage polarization (Zhang et al., 2020). Furthermore, in isoproterenol-induced myocardial fibrosis model of VMC, lncRNA ROR promotes myocardial fibrosis by regulating the expression of C-Myc, suggesting that lncRNA act a vital factor in the chronicity of VMC (Zhang and Sun, 2019). From these existing studies, we can conclude that lncRNAs may be a potential therapeutic target by influencing macrophage polarization and myocardial fibrosis. However, there are few studies on lncRNAs in VMC, and more studies are needed to elucidate the role of lncRNAs.
The Role of circRNAs in VMC
CircRNAs are non-coding RNAs widely expressed in eukaryotic cells, and different from the traditional linear RNAs because of they are circular molecules (Kristensen et al., 2019; Liu C. et al., 2019; Vo et al., 2019; Wilusz, 2019; Zlotorynski, 2019). The characteristics of circRNA are high abundance, structure stable and highly tissue-specific expression. They are mainly produced by the splicing of exon or intron sequences. Besides, reverse complementary sequences or RNA binding proteins (RBPs) are necessary for circRNAs production (Kristensen et al., 2019). Recent studies have shown that circRNAs act as a miRNA sponge to adsorb miRNA, protein sponge, protein scaffold and coding small peptide to complete their biological function (Xie et al., 2018; Zhang M. et al., 2018; Dong et al., 2019; Jiang and Ning, 2019; Liu J. et al., 2019; Lu et al., 2019). Many researchers have found that circRNAs play an important role in cardiovascular diseases such as myocardial infarction, heart failure, and atherosclerosis (Wang K. et al., 2016; Wang et al., 2017; Zhang F. et al., 2018). Zhang et al. confirmed that circRNAs were differentially expressed in fulminant myocarditis, and further bioinformatic analysis showed that many inflammatory or immune pathways, including TNF signaling pathway, Th1 and Th2 cell differentiation and T cell receptor signaling pathway, were involved in VMC (Zhang L. et al., 2019). Therefore, circRNAs may be a new therapeutic target for VMC by regulating the immune-inflammatory pathway. In addition, considering the stability of circRNAs, circulating circRNAs are also potential biomarkers for VMC. However, there is no detailed study on the mechanism of circRNAs affecting the development of VMC.
Conclusions and Perspective
VMC is characterized as a localized or diffuse disease of myocardial parenchyma or interstitium caused by virus infection. The mechanism of VMC may be related to direct myocardial damage of virus infection and extensive damage of abnormal immune response after infection. Although large progress on VMC has been obtained in recent decades, the diagnosis, and treatment of viral myocarditis are still facing great challenges. With the discovery of more and more non-coding RNAs, non-coding RNAs have become the vital molecules of gene regulation in our cognitive range now. In the progress of VMC, non-coding RNAs regulate virus life cycle, immune and inflammatory response by targeting virus or host genes. By reviewing the mechanism of ncRNAs in CVB3-induced myocarditis, we summarized the non-coding RNA related potential disease markers and therapeutic targets of VMC. MiRNAs of VMC have been largely studied by researchers. However, the lncRNA and circRNA field of VMC is still virgin land to be developed. In the future, our researchers should pay more attention to the lncRNA and circRNA area.
Author Contributions
ZY, CZ, YX, ZP, and LZ conceptualized the study. CZ and YX prepared the draft. CZ, YX, and ZY reviewed and edited the manuscript. CZ, YX, and ZL contributed to the visualization. ZY and HZ supervised the study. All authors contributed to the article and approved the submitted version.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors would like to thank the laboratory members for their contributions and the funding support from the sources indicated.
Footnotes
Funding. This study was supported by the grants from the National Natural Science Foundation of China (81670220), the project of Guangdong Provincial Science and Technology Plan (2015A020212013), the Science and Technology Project of Guangzhou City (201803010008), and the International Scientific and Technological Cooperation Project of Guangzhou Economic and Technological Development Zone (2017GH13).
References
- Alessio E., Buson L., Chemello F., Peggion C., Grespi F., Martini P., et al. (2019). Single cell analysis reveals the involvement of the long non-coding RNA Pvt1 in the modulation of muscle atrophy and mitochondrial network. Nucleic Acids Res. 47, 1653–1670. 10.1093/nar/gkz007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Atianand M., Fitzgerald K. (2014). Long non-coding RNAs and control of gene expression in the immune system. Trends Mol. Med. 20, 623–631. 10.1016/j.molmed.2014.09.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bao J. L., Lin L. (2014). MiR-155 and miR-148a reduce cardiac injury by inhibiting NF-κB pathway during acute viral myocarditis. Eur. Rev. Med. Pharmacol. Sci. 18, 2349–2356. Available online at: https://www.europeanreview.org/article/7707 [PubMed] [Google Scholar]
- Barangi S., Hayes A., Reiter R., Karimi G. (2019). The therapeutic role of long non-coding RNAs in human diseases: a focus on the recent insights into autophagy. Pharmacol. Res. 142, 22–29. 10.1016/j.phrs.2019.02.010 [DOI] [PubMed] [Google Scholar]
- Barbu M. G., Condrat C. E., Thompson D. C., Bugnar O. L., Cretoiu D., Toader O. D., et al. (2020). MicroRNA involvement in signaling pathways during viral infection. Front. Cell Dev. Biol. 8:143. 10.3389/fcell.2020.00143 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartel D. (2004). MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297. 10.1016/S0092-8674(04)00045-5 [DOI] [PubMed] [Google Scholar]
- Beg M. S., Brenner A. J., Sachdev J., Borad M., Kang Y. K., Stoudemire J., et al. (2017). Phase I study of MRX34, a liposomal miR-34a mimic, administered twice weekly in patients with advanced solid tumors. Invest. New. Drugs 35, 180–188. 10.1007/s10637-016-0407-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bergelson J. M., Mohanty J. G., Crowell R. L., St John N. F., Lublin D. M., Finberg R. W. (1995). Coxsackievirus B3 adapted to growth in RD cells binds to decay-accelerating factor (CD55). J. Virol. 69, 1903–1906. 10.1128/JVI.69.3.1903-1906.1995 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bewley M. C., Springer K., Zhang Y. B., Freimuth P., Flanagan J. M. (1999). Structural analysis of the mechanism of adenovirus binding to its human cellular receptor, CAR. Science 286, 1579–1583. 10.1126/science.286.5444.1579 [DOI] [PubMed] [Google Scholar]
- Bohnsack M., Czaplinski K., Gorlich D. (2004). Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA 10, 185–191. 10.1261/rna.5167604 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bosselut R., Duvall J., Gégonne A., Bailly M., Hémar A., Brady J., et al. (1990). The product of the c-ets-1 proto-oncogene and the related Ets2 protein act as transcriptional activators of the long terminal repeat of human T cell leukemia virus HTLV-1. EMBO J. 9, 3137–3144. 10.1002/j.1460-2075.1990.tb07511.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bracken C., Scott H., Goodall G. (2016). A network-biology perspective of microRNA function and dysfunction in cancer. Nat Rev. Genet. 17, 719–732. 10.1038/nrg.2016.134 [DOI] [PubMed] [Google Scholar]
- Carloni S., Favrais G., Saliba E., Albertini M., Chalon S., Longini M., et al. (2016). Melatonin modulates neonatal brain inflammation through endoplasmic reticulum stress, autophagy, and miR-34a/silent information regulator 1 pathway. J. Pineal. Res. 61, 370–380. 10.1111/jpi.12354 [DOI] [PubMed] [Google Scholar]
- Castro R., Ferreira D., Afonso M., Borralho P., Machado M., Cortez-Pinto H., et al. (2013). miR-34a/SIRT1/p53 is suppressed by ursodeoxycholic acid in the rat liver and activated by disease severity in human non-alcoholic fatty liver disease. J. Hepatol. 58, 119–125. 10.1016/j.jhep.2012.08.008 [DOI] [PubMed] [Google Scholar]
- Chau D. H., Yuan J., Zhang H., Cheung P., Lim T., Liu Z., et al. (2007). Coxsackievirus B3 proteases 2A and 3C induce apoptotic cell death through mitochondrial injury and cleavage of eIF4GI but not DAP5/p97/NAT1. Apoptosis 12, 513–524. 10.1007/s10495-006-0013-0 [DOI] [PubMed] [Google Scholar]
- Chen B., Xia Z., Deng Y. N., Yang Y., Zhang P., Zhu H., et al. (2019). Emerging microRNA biomarkers for colorectal cancer diagnosis and prognosis. Open Biol. 9:180212. 10.1098/rsob.180212 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen L., Zhou Y., Li H. (2018). LncRNA, miRNA and lncRNA-miRNA interaction in viral infection. Virus Res. 257, 25–32. 10.1016/j.virusres.2018.08.018 [DOI] [PubMed] [Google Scholar]
- Chen X., Wen S., Fukuda M., Gavva N., Hsu D., Akama T., et al. (2001). Human ITCH is a coregulator of the hematopoietic transcription factor NF-E2. Genomics 73, 238–241. 10.1006/geno.2001.6512 [DOI] [PubMed] [Google Scholar]
- Chen Z. G., Liu H., Zhang J. B., Zhang S. L., Zhao L. H., Liang W. Q. (2015). Upregulated microRNA-214 enhances cardiac injury by targeting ITCH during coxsackievirus infection. Mol. Med. Rep. 12, 1258–1264. 10.3892/mmr.2015.3539 [DOI] [PubMed] [Google Scholar]
- Cheng C., Tempel D., Den Dekker W., Haasdijk R., Chrifi I., Bos F., et al. (2011). Ets2 determines the inflammatory state of endothelial cells in advanced atherosclerotic lesions. Circ. Res. 109, 382–395. 10.1161/CIRCRESAHA.111.243444 [DOI] [PubMed] [Google Scholar]
- Cheng S., Hsia C., Feng B., Liou M., Fang X., Pandolfi P., et al. (2009). BCR-mediated apoptosis associated with negative selection of immature B cells is selectively dependent on Pten. Cell Res. 19, 196–207. 10.1038/cr.2008.284 [DOI] [PubMed] [Google Scholar]
- Cheung P. K., Yuan J., Zhang H. M., Chau D., Yanagawa B., Suarez A., et al. (2005). Specific interactions of mouse organ proteins with the 5'untranslated region of coxsackievirus B3: potential determinants of viral tissue tropism. J. Med. Virol. 77, 414–424. 10.1002/jmv.20470 [DOI] [PubMed] [Google Scholar]
- Chi Y., Wang D., Wang J., Yu W., Yang J. (2019). Long non-coding RNA in the pathogenesis of Cancers. Cells 8:1015 10.3390/cells8091015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corsten M., Schroen B., Heymans S. (2012). Inflammation in viral myocarditis: friend or foe? Trends Mol. Med. 18, 426–437. 10.1016/j.molmed.2012.05.005 [DOI] [PubMed] [Google Scholar]
- Corsten M. F., Dennert R., Jochems S., Kuznetsova T., Devaux Y., Hofstra L., et al. (2010). Circulating MicroRNA-208b and MicroRNA-499 reflect myocardial damage in cardiovascular disease. Circ. Cardiovasc. Genet. 3, 499–506. 10.1161/CIRCGENETICS.110.957415 [DOI] [PubMed] [Google Scholar]
- Corsten M. F., Heggermont W., Papageorgiou A. P., Deckx S., Tijsma A., Verhesen W., et al. (2015). The microRNA-221/-222 cluster balances the antiviral and inflammatory response in viral myocarditis. Eur. Heart J. 36, 2909–2919. 10.1093/eurheartj/ehv321 [DOI] [PubMed] [Google Scholar]
- Corsten M. F., Papageorgiou A., Verhesen W., Carai P., Lindow M., Obad S., et al. (2012). MicroRNA profiling identifies microRNA-155 as an adverse mediator of cardiac injury and dysfunction during acute viral myocarditis. Circ. Res. 111, 415–425. 10.1161/CIRCRESAHA.112.267443 [DOI] [PubMed] [Google Scholar]
- Coyne C. B., Bergelson J. M. (2006). Virus-induced Abl and Fyn kinase signals permit coxsackievirus entry through epithelial tight junctions. Cell 124, 119–131. 10.1016/j.cell.2005.10.035 [DOI] [PubMed] [Google Scholar]
- Denli A., Tops B., Plasterk R., Ketting R., Hannon G. (2004). Processing of primary microRNAs by the Microprocessor complex. Nature 432, 231–235. 10.1038/nature03049 [DOI] [PubMed] [Google Scholar]
- Dong H., Lei J., Ding L., Wen Y., Ju H., Zhang X. (2013). MicroRNA: function, detection, and bioanalysis. Chem. Rev. 113, 6207–6233. 10.1021/cr300362f [DOI] [PubMed] [Google Scholar]
- Dong W., Bi J., Liu H., Yan D., He Q., Zhou Q., et al. (2019). Circular RNA ACVR2A suppresses bladder cancer cells proliferation and metastasis through miR-626/EYA4 axis. Mol. Cancer 18:95. 10.1186/s12943-019-1025-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dupont J., Renou J., Shani M., Hennighausen L., Leroith D. (2002). PTEN overexpression suppresses proliferation and differentiation and enhances apoptosis of the mouse mammary epithelium. J. Clin. Invest. 110, 815–825. 10.1172/JCI0213829 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dykes I., Emanueli C. (2017). Transcriptional and post-transcriptional gene regulation by long non-coding RNA. Genomics Proteomics Bioinformatics 15, 177–186. 10.1016/j.gpb.2016.12.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fairweather D., Stafford K. A., Sung Y. K. (2012). Update on coxsackievirus B3 myocarditis. Curr. Opin. Rheumatol. 24, 401–407. 10.1097/BOR.0b013e328353372d [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fan K. L., Li M. F., Cui F., Feng F., Kong L., Zhang F. H., et al. (2019). Altered exosomal miR-181d and miR-30a related to the pathogenesis of CVB3 induced myocarditis by targeting SOCS3. Eur. Rev. Med. Pharmacol. Sci. 23, 2208–2215. 10.26355/eurrev_201903_17268 [DOI] [PubMed] [Google Scholar]
- Fan Y., Weifeng W., Yuluan Y., Qing K., Yu P., Yanlan H. (2011). Treatment with a neutralizing anti-murine interleukin-17 antibody after the onset of coxsackievirus b3-induced viral myocarditis reduces myocardium inflammation. Virol. J. 8:17. 10.1186/1743-422X-8-17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Freimuth P., Philipson L., Carson S. D. (2008). The coxsackievirus and adenovirus receptor. Curr. Top. Microbiol. Immunol. 323, 67–87. 10.1007/978-3-540-75546-3_4 [DOI] [PubMed] [Google Scholar]
- Fung G., Luo H., Qiu Y., Yang D., Mcmanus B. (2016). Myocarditis. Circ. Res. 118, 496–514. 10.1161/CIRCRESAHA.115.306573 [DOI] [PubMed] [Google Scholar]
- Garmaroudi F. S., Marchant D., Hendry R., Luo H., Yang D., Ye X., et al. (2015). Coxsackievirus B3 replication and pathogenesis. Future Microbiol. 10, 629–653. 10.2217/fmb.15.5 [DOI] [PubMed] [Google Scholar]
- Gasparini C., Celeghini C., Monasta L., Zauli G. (2014). NF-κB pathways in hematological malignancies. Cell. Mol. Life Sci. 71, 2083–2102. 10.1007/s00018-013-1545-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gauntt C., Huber S. (2003). Coxsackievirus experimental heart diseases. Front. Biosci. 8, e23–e35. 10.2741/928 [DOI] [PubMed] [Google Scholar]
- Gaur A., Holbeck S., Colburn N., Israel M. (2011). Downregulation of Pdcd4 by mir-21 facilitates glioblastoma proliferation in vivo. Neuro oncol. 13, 580–590. 10.1093/neuonc/nor033 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gebert L., MacRae I. (2019). Regulation of microRNA function in animals. Nat. Rev. Mol. Cell Biol. 20, 21–37. 10.1038/s41580-018-0045-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Germano J. F., Sawaged S., Saadaeijahromi H., Andres A. M., Feuer R., Gottlieb R. A., et al. (2019). Coxsackievirus B infection induces the extracellular release of miR-590-5p, a proviral microRNA. Virology 529, 169–176. 10.1016/j.virol.2019.01.025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldberg L., Tirosh-Wagner T., Vardi A., Abbas H., Pillar N., Shomron N., et al. (2018). Circulating MicroRNAs: a potential biomarker for cardiac damage, inflammatory response, and left ventricular function recovery in pediatric viral myocarditis. J. Cardiovasc. Transl. Res. 11, 319–328. 10.1007/s12265-018-9814-0 [DOI] [PubMed] [Google Scholar]
- Gou W., Zhang Z., Yang C., Li Y. (2018). MiR-223/Pknox1 axis protects mice from CVB3-induced viral myocarditis by modulating macrophage polarization. Exp. Cell. Res. 366, 41–48. 10.1016/j.yexcr.2018.03.004 [DOI] [PubMed] [Google Scholar]
- Gregory R., Chendrimada T., Cooch N., Shiekhattar R. (2005). Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell 123, 631–640. 10.1016/j.cell.2005.10.022 [DOI] [PubMed] [Google Scholar]
- Gregory R., Yan K., Amuthan G., Chendrimada T., Doratotaj B., Cooch N., et al. (2004). The Microprocessor complex mediates the genesis of microRNAs. Nature 432, 235–240. 10.1038/nature03120 [DOI] [PubMed] [Google Scholar]
- Grishok A., Pasquinelli A., Conte D., Li N., Parrish S., Ha I., et al. (2001). Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 106, 23–34. 10.1016/S0092-8674(01)00431-7 [DOI] [PubMed] [Google Scholar]
- Grun S., Schumm J., Greulich S., Wagner A., Schneider S., Bruder O., et al. (2012). Long-term follow-up of biopsy-proven viral myocarditis: predictors of mortality and incomplete recovery. J. Am. Coll. Cardiol. 59, 1604–1615. 10.1016/j.jacc.2012.01.007 [DOI] [PubMed] [Google Scholar]
- Ha M., Kim V. (2014). Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 15, 509–524. 10.1038/nrm3838 [DOI] [PubMed] [Google Scholar]
- Han J., Lee Y., Yeom K., Kim Y., Jin H., Kim V. (2004). The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev. 18, 3016–3027. 10.1101/gad.1262504 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harvala H., Kalimo H., Bergelson J., Stanway G., Hyypia T. (2005). Tissue tropism of recombinant coxsackieviruses in an adult mouse model. J. Gen. Virol. 86, 1897–1907. 10.1099/vir.0.80603-0 [DOI] [PubMed] [Google Scholar]
- He F., Xiao Z., Yao H., Li S., Feng M., Wang W., et al. (2019). The protective role of microRNA-21 against coxsackievirus B3 infection through targeting the MAP2K3/P38 MAPK signaling pathway. J. Transl. Med. 17:335. 10.1186/s12967-019-2077-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- He J., Yue Y., Dong C., Xiong S. (2013). MiR-21 confers resistance against CVB3-induced myocarditis by inhibiting PDCD4-mediated apoptosis. Clin. Invest. Med. 36, E103–E111. 10.25011/cim.v36i2.19573 [DOI] [PubMed] [Google Scholar]
- Hemida M. G., Ye X., Zhang H. M., Hanson P. J., Liu Z., Mcmanus B. M., et al. (2013). MicroRNA-203 enhances coxsackievirus B3 replication through targeting zinc finger protein-148. Cell. Mol. Life Sci. 70, 277–291. 10.1007/s00018-012-1104-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang K. D., Shen Y., Wei X., Zhang F. Q., Liu Y. Y., Ma L. (2016). Inhibitory effect of microRNA-27b on interleukin 17 (IL-17)-induced monocyte chemoattractant protein-1 (MCP1) expression. Genet. Mol. Res. 15:gmr.15027784. 10.4238/gmr.15027784 [DOI] [PubMed] [Google Scholar]
- Huang X., Xiu H., Zhang S., Zhang G. (2018a). The role of macrophages in the pathogenesis of ALI/ARDS. Mediat. Inflamm. 2018:1264913. 10.1155/2018/1264913 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang X., Zhou X., Hu Q., Sun B., Deng M., Qi X., et al. (2018b). Advances in esophageal cancer: a new perspective on pathogenesis associated with long non-coding RNAs. Cancer Lett. 413, 94–101. 10.1016/j.canlet.2017.10.046 [DOI] [PubMed] [Google Scholar]
- Huber S., Sartini D. (2005). T cells expressing the Vgamma1 T-cell receptor enhance virus-neutralizing antibody response during coxsackievirus B3 infection of BALB/c mice: differences in male and female mice. Viral Immunol. 18, 730–739. 10.1089/vim.2005.18.730 [DOI] [PubMed] [Google Scholar]
- Jiang D., Li M., Yu Y., Shi H., Chen R. (2019). microRNA-34a aggravates coxsackievirus B3-induced apoptosis of cardiomyocytes through the SIRT1-p53 pathway. J. Med. Virol. 91, 1643–1651. 10.1002/jmv.25482 [DOI] [PubMed] [Google Scholar]
- Jiang X., Ning Q. (2019). Circular RNAs as novel regulators, biomarkers and potential therapies in fibrosis. Epigenomics 11, 1107–1116. 10.2217/epi-2019-0001 [DOI] [PubMed] [Google Scholar]
- Johansson S., Brauner H., Hinkula J., Wahren B., Berg L., Johansson M. (2011). Accumulation and activation of natural killer cells in local intraperitoneal HIV-1/MuLV infection results in early control of virus infected cells. Cell. Immunol. 272, 71–78. 10.1016/j.cellimm.2011.09.005 [DOI] [PubMed] [Google Scholar]
- Jonas S., Izaurralde E. (2015). Towards a molecular understanding of microRNA-mediated gene silencing. Nat. Rev. Genet. 16, 421–433. 10.1038/nrg3965 [DOI] [PubMed] [Google Scholar]
- Junker F., Chabloz A., Koch U., Radtke F. (2015). Dicer1 imparts essential survival cues in Notch-driven T-ALL via miR-21-mediated tumor suppressor Pdcd4 repression. Blood 126, 993–1004. 10.1182/blood-2014-12-618892 [DOI] [PubMed] [Google Scholar]
- Kabacaoglu D., Ruess D., Ai J., Algül H. (2019). NF-κB/Rel transcription factors in pancreatic cancer: focusing on RelA, c-Rel, and RelB. Cancers 11:937. 10.3390/cancers11070937 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kanda T., Mcmanus J., Nagai R., Imai S., Suzuki T., Yang D., et al. (1996). Modification of viral myocarditis in mice by interleukin-6. Circ. Res. 78, 848–856. 10.1161/01.RES.78.5.848 [DOI] [PubMed] [Google Scholar]
- Ketting R., Fischer S., Bernstein E., Sijen T., Hannon G., Plasterk R. (2001). Dicer functions in RNA interference and in synthesis of small RNA involved in developmental timing in C. elegans. Genes Dev. 15, 2654–2659. 10.1101/gad.927801 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim V. (2004). MicroRNA precursors in motion: exportin-5 mediates their nuclear export. Trends Cell Biol. 14, 156–159. 10.1016/j.tcb.2004.02.006 [DOI] [PubMed] [Google Scholar]
- Kindermann I., Barth C., Mahfoud F., Ukena C., Lenski M., Yilmaz A., et al. (2012). Update on myocarditis. J. Am. Coll. Cardiol. 59, 779–792. 10.1016/j.jacc.2011.09.074 [DOI] [PubMed] [Google Scholar]
- Knowlton K. U. (2008). CVB infection and mechanisms of viral cardiomyopathy. Curr. Top. Microbiol. Immunol. 323, 315–335. 10.1007/978-3-540-75546-3_15 [DOI] [PubMed] [Google Scholar]
- Kobayashi H., Tomari Y. (2016). RISC assembly: coordination between small RNAs and argonaute proteins. Biochim. Biophys. Acta 1859, 71–81. 10.1016/j.bbagrm.2015.08.007 [DOI] [PubMed] [Google Scholar]
- Kristensen L., Andersen M., Stagsted L., Ebbesen K., Hansen T., Kjems J. (2019). The biogenesis, biology and characterization of circular RNAs. Nat. Rev. Genet. 20, 675–691. 10.1038/s41576-019-0158-7 [DOI] [PubMed] [Google Scholar]
- Kühl U., Pauschinger M., Schwimmbeck P., Seeberg B., Lober C., Noutsias M., et al. (2003). Interferon-beta treatment eliminates cardiotropic viruses and improves left ventricular function in patients with myocardial persistence of viral genomes and left ventricular dysfunction. Circulation 107, 2793–2798. 10.1161/01.CIR.0000072766.67150.51 [DOI] [PubMed] [Google Scholar]
- Kvansakul M. (2017). Viral infection and apoptosis. Viruses 9:356 10.3390/v9120356 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lai E. (2002). Micro RNAs are complementary to 3' UTR sequence motifs that mediate negative post-transcriptional regulation. Nat. Genet. 30, 363–364. 10.1038/ng865 [DOI] [PubMed] [Google Scholar]
- Landthaler M., Yalcin A., Tuschl T. (2004). The human DiGeorge syndrome critical region gene 8 and Its D. melanogaster homolog are required for miRNA biogenesis. Curr. Biol. 14, 2162–2167. 10.1016/j.cub.2004.11.001 [DOI] [PubMed] [Google Scholar]
- Lee Y., Ahn C., Han J., Choi H., Kim J., Yim J., et al. (2003). The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419. 10.1038/nature01957 [DOI] [PubMed] [Google Scholar]
- Lee Y., Kim M., Han J., Yeom K., Lee S., Baek S., et al. (2004). MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 23, 4051–4060. 10.1038/sj.emboj.7600385 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis S. M., Cerquozzi S., Graber T. E., Ungureanu N. H., Andrews M., Holcik M. (2008). The eIF4G homolog DAP5/p97 supports the translation of select mRNAs during endoplasmic reticulum stress. Nucleic Acids Res. 36, 168–178. 10.1093/nar/gkm1007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li J., Li Z., Leng K., Xu Y., Ji D., Huang L., et al. (2018). ZEB1-AS1: a crucial cancer-related long non-coding RNA. Cell Prolif. 51:e12423. 10.1111/cpr.12423 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y., Zhou L., Lu C., Shen Q., Su Y., Zhi Z., et al. (2018). Long non-coding RNA FAL1 functions as a ceRNA to antagonize the effect of miR-637 on the down-regulation of AKT1 in Hirschsprung's disease. Cell Prolif. 51:e12489. 10.1111/cpr.12489 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liao Y., Chen K. H., Dong X. M., Fang Y., Li W. G., Huang G. Y., et al. (2015). A role of pre-mir-10a coding region variant in host susceptibility to coxsackie virus-induced myocarditis. Eur. Rev. Med. Pharmacol. Sci. 19, 3500–3507. Available online at: https://www.europeanreview.org/article/9525 [PubMed] [Google Scholar]
- Lin P., Fosmire S., Park S., Park J., Baksh S., Modiano J., et al. (2007). Attenuation of PTEN increases p21 stability and cytosolic localization in kidney cancer cells: a potential mechanism of apoptosis resistance. Mol. Cancer 6:16. 10.1186/1476-4598-6-16 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lind K., Svedin E., Domsgen E., Kapell S., Laitinen O. H., Moll M., et al. (2016). Coxsackievirus counters the host innate immune response by blocking type III interferon expression. J. Gen. Virol. 97, 1368–1380. 10.1099/jgv.0.000443 [DOI] [PubMed] [Google Scholar]
- Lindner D., Li J., Savvatis K., Klingel K., Blankenberg S., Tschöpe C., et al. (2014). Cardiac fibroblasts aggravate viral myocarditis: cell specific coxsackievirus B3 replication. Mediat. Inflamm. 2014:519528. 10.1155/2014/519528 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu C., Li X., Nan F., Jiang S., Gao X., Guo S., et al. (2019). Structure and degradation of circular RNAs regulate PKR activation in innate immunity. Cell 177, 865–880.e21. 10.1016/j.cell.2019.03.046 [DOI] [PubMed] [Google Scholar]
- Liu J., Zhao K., Huang N., Zhang N. (2019). Circular RNAs and human glioma. Cancer Biol. Med. 16, 11–23. 10.20892/j.issn.2095-3941.2018.0425 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Q., Kong Y., Han B., Jiang D., Jia H., Zhang L. (2019). Long non-coding RNA expression profile and functional analysis in children with acute fulminant myocarditis. Front. Pediatr. 7:283. 10.3389/fped.2019.00283 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y. L., Wu W., Xue Y., Gao M., Yan Y., Kong Q., et al. (2013). MicroRNA-21 and−146b are involved in the pathogenesis of murine viral myocarditis by regulating TH-17 differentiation. Arch. Virol. 158, 1953–1963. 10.1007/s00705-013-1695-6 [DOI] [PubMed] [Google Scholar]
- Lobo M. L., Taguchi A., Gaspar H. A., Ferranti J. F., De Carvalho W. B., Delgado A. F. (2014). Fulminant myocarditis associated with the H1N1 influenza virus: case report and literature review. Rev. Bras. Ter. Intensiva 26, 321–326. 10.5935/0103-507X.20140046 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu Q., Liu T., Feng H., Yang R., Zhao X., Chen W., et al. (2019). Circular RNA circSLC8A1 acts as a sponge of miR-130b/miR-494 in suppressing bladder cancer progression via regulating PTEN. Mol. Cancer 18:111. 10.1186/s12943-019-1040-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lund E., Güttinger S., Calado A., Dahlberg J., Kutay U. (2004). Nuclear export of microRNA precursors. Science 303, 95–98. 10.1126/science.1090599 [DOI] [PubMed] [Google Scholar]
- Matsumori A., Nunokawa Y., Yamaki A., Yamamoto K., Hwang M., Miyamoto T., et al. (2004). Suppression of cytokines and nitric oxide production, and protection against lethal endotoxemia and viral myocarditis by a new NF-kappaB inhibitor. Eur. J. Heart Fail 6, 137–144. 10.1016/j.ejheart.2003.10.007 [DOI] [PubMed] [Google Scholar]
- Minhas A. M., Nayab A., Iyer S., Narmeen M., Fatima K., Khan M. S., et al. (2017). Association of zika virus with myocarditis, heart failure, and arrhythmias: a literature review. Cureus 9:e1399. 10.7759/cureus.1399 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyamoto T., Matsumori A., Hwang M., Nishio R., Ito H., Sasayama S. (2001). Therapeutic effects of FTY720, a new immunosuppressive agent, in a murine model of acute viral myocarditis. J. Am. Coll. Cardiol. 37, 1713–1718. 10.1016/S0735-1097(01)01204-9 [DOI] [PubMed] [Google Scholar]
- Mukherjee N., Houston T., Cardenas E., Ghosh R. (2015). To be an ally or an adversary in bladder cancer: the NF-κB story has not unfolded. Carcinogenesis 36, 299–306. 10.1093/carcin/bgu321 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicolas F. E. (2017). Role of ncRNAs in development, diagnosis and treatment of human cancer. Recent Pat. Anticancer. Drug Discov. 12, 128–135. 10.2174/1574892812666170105113415 [DOI] [PubMed] [Google Scholar]
- Ntusi N. A. B. (2017). HIV and myocarditis. Curr. Opin. HIV AIDS 12, 561–565. 10.1097/COH.0000000000000416 [DOI] [PubMed] [Google Scholar]
- Okuyama N., Cezar-Dos-Santos F., Pereira É., Trugilo K., Cebinelli G., Sena M., et al. (2018). Genetic variant in CXCL12 gene raises susceptibility to HPV infection and squamous intraepithelial lesions development: a case-control study. J. Biomed. Sci. 25:69. 10.1186/s12929-018-0472-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olejniczak M., Schwartz M., Webber E., Shaffer A., Perry T. E. (2020). Viral myocarditis-incidence, diagnosis and management. J. Cardiothorac. Vasc. Anesth. 34, 1591–1601. 10.1053/j.jvca.2019.12.052 [DOI] [PubMed] [Google Scholar]
- Panni S., Lovering R. C., Porras P., Orchard S. (2019). Non-coding RNA regulatory networks. Biochim Biophys Acta Gene Regul. Mech. 1863:194417. 10.1016/j.bbagrm.2019.194417 [DOI] [PubMed] [Google Scholar]
- Peischard S., Ho H. T., Theiss C., Strutz-Seebohm N., Seebohm G. (2019). A kidnapping story: how coxsackievirus B3 and its host cell interact. Cell. Physiol. Biochem. 53, 121–140. 10.33594/000000125 [DOI] [PubMed] [Google Scholar]
- Perez J., Chirieleison S., Abbott D. (2015). An IκB kinase-regulated feedforward circuit prolongs inflammation. Cell Rep. 12, 537–544. 10.1016/j.celrep.2015.06.050 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pinkert S., Westermann D., Wang X., Klingel K., Dorner A., Savvatis K., et al. (2009). Prevention of cardiac dysfunction in acute coxsackievirus B3 cardiomyopathy by inducible expression of a soluble coxsackievirus-adenovirus receptor. Circulation 120, 2358–2366. 10.1161/CIRCULATIONAHA.108.845339 [DOI] [PubMed] [Google Scholar]
- Pollack A., Kontorovich A. R., Fuster V., Dec G. W. (2015). Viral myocarditis–diagnosis, treatment options, and current controversies. Nat. Rev. Cardiol. 12, 670–680. 10.1038/nrcardio.2015.108 [DOI] [PubMed] [Google Scholar]
- Posada R., Pettoello-Mantovani M., Sieweke M., Graf T., Goldstein H. (2000). Suppression of HIV type 1 replication by a dominant-negative Ets-1 mutant. AIDS Res. Hum. Retroviruses 16, 1981–1989. 10.1089/088922200750054710 [DOI] [PubMed] [Google Scholar]
- Quinn J., Chang H. (2016). Unique features of long non-coding RNA biogenesis and function. Nat. Rev. Genet. 17, 47–62. 10.1038/nrg.2015.10 [DOI] [PubMed] [Google Scholar]
- Regouc M., Belge G., Lorch A., Dieckmann K. P., Pichler M. (2020). Non-coding microRNAs as novel potential tumor markers in testicular cancer. Cancers 12:749. 10.3390/cancers12030749 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Russell L., Garrett-Sinha L. (2010). Transcription factor Ets-1 in cytokine and chemokine gene regulation. Cytokine 51, 217–226. 10.1016/j.cyto.2010.03.006 [DOI] [PubMed] [Google Scholar]
- Santos J. M. O., Peixoto Da Silva S., Gil Da Costa R. M., Medeiros R. (2020). The emerging role of MicroRNAs and other non-coding RNAs in cancer cachexia. Cancers 12:1004. 10.3390/cancers12041004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schultz J. C., Hilliard A. A., Cooper L. T., Jr., Rihal C. S. (2009). Diagnosis and treatment of viral myocarditis. Mayo Clin. Proc. 84, 1001–1009. 10.1016/S0025-6196(11)60670-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shafren D. R., Bates R. C., Agrez M. V., Herd R. L., Burns G. F., Barry R. D. (1995). Coxsackieviruses B1, B3, and B5 use decay accelerating factor as a receptor for cell attachment. J. Virol. 69, 3873–3877. 10.1128/JVI.69.6.3873-3877.1995 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shembade N., Harhaj N., Parvatiyar K., Copeland N., Jenkins N., Matesic L., et al. (2008). The E3 ligase itch negatively regulates inflammatory signaling pathways by controlling the function of the ubiquitin-editing enzyme A20. Nat. Immunol. 9, 254–262. 10.1038/ni1563 [DOI] [PubMed] [Google Scholar]
- Shen X., Zhang X., Ru W., Huang Y., Lan X., Lei C., et al. (2020). circINSR promotes proliferation and reduces apoptosis of embryonic myoblasts by sponging miR-34a. Mol. Ther. Nucleic Acids 19, 986–999. 10.1016/j.omtn.2019.12.032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sieweke M., Tekotte H., Jarosch U., Graf T. (1998). Cooperative interaction of ets-1 with USF-1 required for HIV-1 enhancer activity in T cells. EMBO J. 17, 1728–1739. 10.1093/emboj/17.6.1728 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smolle M. A., Prinz F., Calin G. A., Pichler M. (2019). Current concepts of non-coding RNA regulation of immune checkpoints in cancer. Mol. Aspects Med. 70, 117–126. 10.1016/j.mam.2019.09.007 [DOI] [PubMed] [Google Scholar]
- Sodroski C., Lowey B., Hertz L., Jake Liang T., Li Q. (2019). MicroRNA-135a modulates hepatitis C virus genome replication through downregulation of host antiviral factors. Virol. Sin. 34, 197–210. 10.1007/s12250-018-0055-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spartalis M., Tzatzaki E., Spartalis E., Damaskos C., Mavrogeni S., Voudris V. (2017). Parvovirus B19 myocarditis of fulminant evolution. Cardiol. Res. 8, 172–175. 10.14740/cr580w [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stagakis E., Bertsias G., Verginis P., Nakou M., Hatziapostolou M., Kritikos H., et al. (2011). Identification of novel microRNA signatures linked to human lupus disease activity and pathogenesis: miR-21 regulates aberrant T cell responses through regulation of PDCD4 expression. Ann. Rheum. Dis. 70, 1496–1506. 10.1136/ard.2010.139857 [DOI] [PubMed] [Google Scholar]
- Su Y. C., Huang Y. F., Wu Y. W., Chen H. F., Wu Y. H., Hsu C. C., et al. (2020). MicroRNA-155 inhibits dengue virus replication by inducing heme oxygenase-1-mediated antiviral interferon responses. FASEB J. 34, 7283–7294. 10.1096/fj.201902878R [DOI] [PubMed] [Google Scholar]
- Tanaka-Nakanishi A., Yasunaga J., Takai K., Matsuoka M. (2014). HTLV-1 bZIP factor suppresses apoptosis by attenuating the function of FoxO3a and altering its localization. Cancer Res. 74, 188–200. 10.1158/0008-5472.CAN-13-0436 [DOI] [PubMed] [Google Scholar]
- Tong L., Lin L., Wu S., Guo Z., Wang T., Qin Y., et al. (2013). MiR-10a* up-regulates coxsackievirus B3 biosynthesis by targeting the 3D-coding sequence. Nucleic Acids Res. 41, 3760–3771. 10.1093/nar/gkt058 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tong R., Jia T., Shi R., Yan F. (2020). Inhibition of microRNA-15 protects H9c2 cells against CVB3-induced myocardial injury by targeting NLRX1 to regulate the NLRP3 inflammasome. Cell. Mol. Biol. Lett. 25:6. 10.1186/s11658-020-00203-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tong Z., Tang Y., Jiang B., Wu Y., Liu Y., Li Y., et al. (2019). Phosphorylation of nucleolin is indispensable to upregulate miR-21 and inhibit apoptosis in cardiomyocytes. J. Cell. Physiol. 234, 4044–4053. 10.1002/jcp.27191 [DOI] [PubMed] [Google Scholar]
- Treiber T., Treiber N., Meister G. (2019). Regulation of microRNA biogenesis and its crosstalk with other cellular pathways. Nat. Rev. Mol. Cell Biol. 20, 5–20. 10.1038/s41580-018-0059-1 [DOI] [PubMed] [Google Scholar]
- Tschope C., Cooper L. T., Torre-Amione G., Van Linthout S. (2019). Management of myocarditis-related cardiomyopathy in adults. Circ. Res. 124, 1568–1583. 10.1161/CIRCRESAHA.118.313578 [DOI] [PubMed] [Google Scholar]
- van Kuppeveld F. J., Hoenderop J. G., Smeets R. L., Willems P. H., Dijkman H. B., Galama J. M., et al. (1997). Coxsackievirus protein 2B modifies endoplasmic reticulum membrane and plasma membrane permeability and facilitates virus release. EMBO J. 16, 3519–3532. 10.1093/emboj/16.12.3519 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Verdonschot J., Hazebroek M., Merken J., Debing Y., Dennert R., Brunner-La Rocca H. P., et al. (2016). Relevance of cardiac parvovirus B19 in myocarditis and dilated cardiomyopathy: review of the literature. Eur. J. Heart Fail 18, 1430–1441. 10.1002/ejhf.665 [DOI] [PubMed] [Google Scholar]
- Vo J., Cieslik M., Zhang Y., Shukla S., Xiao L., Zhang Y., et al. (2019). The landscape of circular RNA in cancer. Cell 176, 869–881.e13. 10.1016/j.cell.2018.12.021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vos T., Barber R. M., Bell B., Bertozzi-Villa A., Biryukov S., Bolliger I., et al. (2015). Global, regional, and national incidence, prevalence, and years lived with disability for 301 acute and chronic diseases and injuries in 188 countries, 1990-2013: a systematic analysis for the Global Burden of Disease Study 2013. Lancet 386, 743–800. 10.1016/S0140-6736(15)60692-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang D., Li T., Cui H., Zhang Y. (2016). Analysis of the indicating value of cardiac troponin I, tumor necrosis factor-alpha, interleukin-18, mir-1 and mir-146b for viral myocarditis among children. Cell. Physiol. Biochem. 40, 1325–1333. 10.1159/000453185 [DOI] [PubMed] [Google Scholar]
- Wang K., Gan T. Y., Li N., Liu C. Y., Zhou L. Y., Gao J. N., et al. (2017). Circular RNA mediates cardiomyocyte death via miRNA-dependent upregulation of MTP18 expression. Cell Death Differ. 24, 1111–1120. 10.1038/cdd.2017.61 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang K., Long B., Liu F., Wang J. X., Liu C. Y., Zhao B., et al. (2016). A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. Eur. Heart J. 37, 2602–2611. 10.1093/eurheartj/ehv713 [DOI] [PubMed] [Google Scholar]
- Wang L., Qin Y., Tong L., Wu S., Wang Q., Jiao Q., et al. (2012). MiR-342-5p suppresses coxsackievirus B3 biosynthesis by targeting the 2C-coding region. Antiviral Res. 93, 270–279. 10.1016/j.antiviral.2011.12.004 [DOI] [PubMed] [Google Scholar]
- Wang Q., Roy B., Dwivedi Y. (2019). Co-expression network modeling identifies key long non-coding RNA and mRNA modules in altering molecular phenotype to develop stress-induced depression in rats. Transl. Psychiatry 9:125. 10.1038/s41398-019-0448-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei B., Deng Y., Huang Y., Gao X., Wu W. (2019). IL-10-producing B cells attenuate cardiac inflammation by regulating Th1 and Th17 cells in acute viral myocarditis induced by coxsackie virus B3. Life Sci. 235:116838. 10.1016/j.lfs.2019.116838 [DOI] [PubMed] [Google Scholar]
- Welsh R. (1986). Regulation of virus infections by natural killer cells. a review. Nat. Immun. Cell Growth Regul. 5, 169–199. [PubMed] [Google Scholar]
- Wilusz J. (2019). Circle the wagons: circular RNAs control innate immunity. Cell 177, 797–799. 10.1016/j.cell.2019.04.020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiao M., Li J., Li W., Wang Y., Wu F., Xi Y., et al. (2017). MicroRNAs activate gene transcription epigenetically as an enhancer trigger. RNA Biol. 14, 1326–1334. 10.1080/15476286.2015.1112487 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xie F., Li Y., Wang M., Huang C., Tao D., Zheng F., et al. (2018). Circular RNA BCRC-3 suppresses bladder cancer proliferation through miR-182-5p/p27 axis. Mol. Cancer 17:144. 10.1186/s12943-018-0892-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu H. F., Gao X. T., Lin J. Y., Xu X. H., Hu J., Ding Y. J., et al. (2017). MicroRNA-20b suppresses the expression of ZFP-148 in viral myocarditis. Mol. Cell. Biochem. 429, 199–210. 10.1007/s11010-017-2947-7 [DOI] [PubMed] [Google Scholar]
- Yang X., Yue Y., Xiong S. (2019). Dpep2 emerging as a modulator of macrophage inflammation confers protection against CVB3-induced viral myocarditis. Front. Cell. Infect. Microbiol. 9:57. 10.3389/fcimb.2019.00057 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang Y., Cheng H., Qiu Y., Dupee D., Noonan M., Lin Y., et al. (2015). MicroRNA-34a plays a key role in cardiac repair and regeneration following myocardial infarction. Circ. Res. 117, 450–459. 10.1161/CIRCRESAHA.117.305962 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ye X., Hemida M. G., Qiu Y., Hanson P. J., Zhang H. M., Yang D. (2013). MiR-126 promotes coxsackievirus replication by mediating cross-talk of ERK1/2 and Wnt/beta-catenin signal pathways. Cell. Mol. Life Sci. 70, 4631–4644. 10.1007/s00018-013-1411-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yi R., Qin Y., Macara I., Cullen B. (2003). Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 17, 3011–3016. 10.1101/gad.1158803 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhan Y., Brown C., Maynard E., Anshelevich A., Ni W., Ho I., et al. (2005). Ets-1 is a critical regulator of Ang II-mediated vascular inflammation and remodeling. J. Clin. Invest. 115, 2508–2516. 10.1172/JCI24403 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang B. Y., Zhao Z., Jin Z. (2016). Expression of miR-98 in myocarditis and its influence on transcription of the FAS/FASL gene pair. Genet. Mol. Res. 15:gmr.15027627. 10.4238/gmr.15027627 [DOI] [PubMed] [Google Scholar]
- Zhang F., Zhang R., Zhang X., Wu Y., Li X., Zhang S., et al. (2018). Comprehensive analysis of circRNA expression pattern and circRNA-miRNA-mRNA network in the pathogenesis of atherosclerosis in rabbits. Aging 10, 2266–2283. 10.18632/aging.101541 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang K., Han Y., Hu Z., Zhang Z., Shao S., Yao Q., et al. (2019). SCARNA10, a nuclear-retained long non-coding RNA, promotes liver fibrosis and serves as a potential biomarker. Theranostics 9, 3622–3638. 10.7150/thno.32935 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang L., Han B., Wang J., Liu Q., Kong Y., Jiang D., et al. (2019). Differential expression profiles and functional analysis of circular RNAs in children with fulminant myocarditis. Epigenomics 11, 1129–1141. 10.2217/epi-2019-0101 [DOI] [PubMed] [Google Scholar]
- Zhang L., Wang H. (2019). Long non-coding RNA in CNS injuries: a new target for therapeutic intervention. Mol Therapy Nucleic Acids 17, 754–766. 10.1016/j.omtn.2019.07.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang M., Zhao K., Xu X., Yang Y., Yan S., Wei P., et al. (2018). A peptide encoded by circular form of LINC-PINT suppresses oncogenic transcriptional elongation in glioblastoma. Nat. Commun. 9:4475. 10.1038/s41467-018-06862-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang N., Sun Y. (2019). LncRNA ROR facilitates myocardial fibrosis in rats with viral myocarditis through regulating C-Myc expression. Eur. Rev. Med. Pharmacol. Sci. 23, 10982–10988. 10.26355/eurrev_201912_19803 [DOI] [PubMed] [Google Scholar]
- Zhang W., Xu W., Feng Y., Zhou X. (2019). Non-coding RNA involvement in the pathogenesis of diabetic cardiomyopathy. J. Cell. Mol. Med. 23, 5859–5867. 10.1111/jcmm.14510 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X., Gao X., Hu J., Xie Y., Zuo Y., Xu H., et al. (2019). ADAR1p150 forms a complex with dicer to promote miRNA-222 activity and regulate PTEN expression in CVB3-induced viral myocarditis. Int. J. Mol. Sci. 20:407. 10.3390/ijms20020407 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y., Li X., Kong X., Zhang M., Wang D., Liu Y., et al. (2020). Long non-coding RNA AK085865 ablation confers susceptibility to viral myocarditis by regulating macrophage polarization. J. Cell. Mol. Med. 24, 5542–5554. 10.1111/jcmm.15210 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y., Sun L., Sun H., Liu X., Luo X., Li C., et al. (2017). Overexpression of microRNA-133b reduces myocardial injuries in children with viral myocarditis by targeting Rab27B gene. Cell. Mol. Biol. 63, 80–86. 10.14715/cmb/2017.63.10.13 [DOI] [PubMed] [Google Scholar]
- Zhang Y., Sun L., Sun H., Yu Z., Liu X., Luo X., et al. (2018). MicroRNA-381 protects myocardial cell function in children and mice with viral myocarditis via targeting cyclooxygenase-2 expression. Exp. Ther. Med. 15, 5510–5516. 10.3892/etm.2018.6082 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y., Zhang M., Li X., Tang Z., Wang X., Zhong M., et al. (2016). Silencing MicroRNA-155 attenuates cardiac injury and dysfunction in viral myocarditis via promotion of M2 phenotype polarization of macrophages. Sci. Rep. 6:22613. 10.1038/srep22613 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng Y. F., Luo J., Gan G. L., Li W. (2019). Overexpression of microRNA-98 inhibits cell proliferation and promotes cell apoptosis via claudin-1 in human colorectal carcinoma. J. Cell. Biochem. 120, 6090–6105. 10.1002/jcb.27895 [DOI] [PubMed] [Google Scholar]
- Zhou Z., Zhou L., Jiang F., Zeng B., Wei C., Zhao W., et al. (2017). Downregulation of miR-222 induces apoptosis and cellular migration in adenoid cystic carcinoma cells. Oncol. Res. 25, 207–214. 10.3727/096504016X14732772150460 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zlotorynski E. (2019). The innate function of circular RNAs. Nat. Rev. Mol. Cell Biol. 20:387. 10.1038/s41580-019-0146-y [DOI] [PubMed] [Google Scholar]