Abstract
Gestational diabetes mellitus (GDM), one of the major pregnancy-related complications, characterized as a transitory form of diabetes induced by insulin resistance accompanied by a low/absent pancreatic beta-cell compensatory adaptation to the increased insulin demand, causes the acute, long-term, and transgenerational health complications. The aim of the study was to assess if alterations in gene expression of microRNAs associated with diabetes/cardiovascular/cerebrovascular diseases are present in whole peripheral blood of children aged 3–11 years descending from GDM complicated pregnancies. A substantially altered microRNA expression profile was found in children descending from GDM complicated pregnancies. Almost all microRNAs with the exception of miR-92a-3p, miR-155-5p, and miR-210-3p were upregulated. The microRNA expression profile also differed between children after normal and GDM complicated pregnancies in relation to the presence of overweight/obesity, prehypertension/hypertension, and/or valve problems and heart defects. Always, screening based on the combination of microRNAs was superior over using individual microRNAs, since at 10.0% false positive rate it was able to identify a large proportion of children with an aberrant microRNA expression profile (88.14% regardless of clinical findings, 75.41% with normal clinical findings, and 96.49% with abnormal clinical findings). In addition, the higher incidence of valve problems and heart defects was found in children with a prior exposure to GDM. The extensive file of predicted targets of all microRNAs aberrantly expressed in children descending from GDM complicated pregnancies indicates that a large group of these genes is involved in ontologies of diabetes/cardiovascular/cerebrovascular diseases. In general, children with a prior exposure to GDM are at higher risk of later development of diabetes mellitus and cardiovascular/cerebrovascular diseases, and would benefit from dispensarisation as well as implementation of primary prevention strategies.
Keywords: BMI, bioinformatics, cardiovascular risk, children, echocardiography, gestational diabetes mellitus, microRNA expression, miRWalk2.0 database, prehypertension/hypertension, screening
1. Introduction
Gestational diabetes mellitus (GDM) represents a pregnancy-related complication with the onset during the second or third trimester of gestation with worldwide increasing prevalence ranging from 7 to 14% [1,2]. GDM is defined as a transitory form of diabetes induced by insulin resistance accompanied by a low/absent pancreatic beta-cell compensatory adaptation to the increased insulin demand. Pancreatic beta-cell dysfunction is just a central component of the pathogenesis of GDM [1,2,3,4,5,6].
MicroRNAs are important metabolic and developmental regulators during gestation that play a role in the onset of GDM [7]. Recent studies have evaluated the expression of microRNAs playing a role in glucose homeostasis, insulin sensitivity, and beta-cell function in different sample types (placenta, umbilical vein endothelial cells, whole blood, plasma, and serum) with the aim to assess their potential as diagnostic or prognostic biomarkers of GDM [7,8,9,10,11,12,13,14,15,16,17,18,19,20,21].
Accumulating data suggest that exposure to hyperglycemia in utero, as occurs in gestational diabetes mellitus, may expose offspring to short-term and long-term adverse effects [22,23,24,25].
Young offspring aged 3–5 years of GDM complicated pregnancies were reported to have increased BMI, skinfold thickness, body fat, blood pressure, altered lipid profiles, and glucose metabolism [26]. Several consecutive studies confirmed higher mean values of systolic blood pressure and higher prevalence of hypertension in young offspring (at the age of 3–6 years) of mothers with GDM compared with their counterparts born to mothers with a normal course of gestation [27,28]. An independent association between the occurrence of maternal GDM and abnormal glucose tolerance, obesity, and higher BP was confirmed in children at seven years of age [29]. Another study indicated that also children at the age of 10–14 years had an impaired glucose tolerance as a consequence of in utero exposition to maternal GDM [30].
Parallel, in 17-year-old offspring of mothers with GDM and higher mean BMI, systolic and diastolic BP values were observed when compared to no-recorded-GDM offspring [31]. Nevertheless, another recent study performed on youth (at the age of 10.6–16.9 years) demonstrated that maternal GDM was related to altered lipid profile and higher systolic blood pressure only in a sex-specific manner [32]. A higher total and LDL cholesterol were detected only in girls and a higher systolic blood pressure only in boys previously exposed in utero to GDM [32]. Even a significant association between maternal GDM and the rate of hospitalization for subsequent cardiovascular related morbidities (pre-defined set of ICD-9 codes) was noted in an 18-year follow-up study [33].
Another population-based cohort study with a 40-year follow-up revealed the GDM association with an increased occurrence of cardiovascular diseases in early adulthood. Varied increased rates of specific early onset cardiovascular diseases, particularly heart failure, hypertensive disease, deep vein thrombosis, and pulmonary embolism, were observed in the offspring previously affected with GDM [34]. Similarly, young adults descending from pregnancies with an impaired glucose tolerance were at an increased risk of development of type 2 diabetes [23,35,36]. The cumulative risk of type 2 diabetes reached 15% in the offspring at the age of 20 years, and about 30% at the age of 24 years [23,35].
Furthermore, children with in utero exposition to GDM were demonstrated to have an impaired neurophysiology (reduced cortical excitability, neuroplasticity, and salivary cortisol) [37] and to be at an increased risk of neurodevelopmental difficulties, including attention deficit hyperactivity disorder [38,39], autism spectrum disorders [40,41], impaired motor development [42], and neuropsychiatric disorders [43].
In addition, children descending from GDM complicated pregnancies were also reported to have an increased risk of paediatric ophthalmic morbidity [44]. Moreover, a small risk of childhood asthma was observed after exposure to GDM requiring medication only [45].
In general, the aim of this follow-up study was to assess if alterations in the gene expression of microRNAs are present in children descending from GDM complicated pregnancies.
In detail, the expression profile of microRNAs associated with diabetes/cardiovascular/cerebrovascular diseases was assessed in whole peripheral venous blood (white blood cells) of children aged 3 to 11 years prenatally exposed to GDM with the goal to assess to what extent fetal and environmental programming predisposed the affected children to the development of diabetes mellitus, cardiovascular, and cerebrovascular diseases.
The hypothesis of the assessment of potential diabetes/cardiovascular risk in children prenatally exposed to GDM was founded on the fact that particular microRNAs play a key role in the inducement and progress of diabetes mellitus and cardiovascular/cerebrovascular diseases (Table 1) [46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117,118,119,120,121,122,123,124,125,126,127,128,129,130,131,132,133,134,135,136,137,138,139,140,141,142,143,144,145,146,147,148,149,150,151,152,153,154,155,156,157,158,159,160,161,162,163,164,165,166,167,168,169,170,171,172,173,174,175,176,177,178,179,180,181,182,183,184,185,186,187,188,189,190,191,192,193,194,195,196,197,198,199,200,201,202,203,204,205,206,207,208,209,210,211,212,213,214,215,216,217,218,219,220,221].
Table 1.
The role of studied microRNAs in the pathogenesis of diabetes mellitus and cardiovascular/cerebrovascular diseases.
| miRBase ID | Gene Location on Chromosome | Role in the Pathogenesis of Diabetes Mellitus and Cardiovascular/Cerebrovascular Diseases |
|---|---|---|
| hsa-miR-1-3p | 20q13.3 [46] 18q11.2 | Acute myocardial infarction, heart ischemia, post-myocardial infarction complications [47], thoracic aortic aneurysm [48], diabetes mellitus [49,50], vascular endothelial dysfunction [51]. |
| hsa-miR-16-5p | 13q14.2 | Myocardial infarction [52,53], heart failure [54], acute coronary syndrome, cerebral ischaemic events [55], gestational diabetes mellitus [9,13,20], diabetes mellitus [56,57,58]. |
| hsa-miR-17-5p | 13q31.3 [59,60] | Cardiac development [61], ischemia/reperfusion-induced cardiac injury [62], kidney ischemia-reperfusion injury [63], diffuse myocardial fibrosis in hypertrophic cardiomyopathy [64], acute ischemic stroke [65], coronary artery disease [66], adipogenic differentiation [67], gestational diabetes mellitus [9,13], diabetes mellitus [58,68]. |
| hsa-miR-20a-5p | 13q31.3 [69] | Pulmonary hypertension [70], gestational diabetes mellitus [9,13,17], diabetic retinopathy [71], diabetes with abdominal aortic aneurysm [72]. |
| hsa-miR-20b-5p | Xq26.2 [69] | Hypertension-induced heart failure [73], insulin resistance [74], T2DM [75,76], diabetic retinopathy [77]. |
| hsa-miR-21-5p | 17q23.2 [78] | Homeostasis of the cardiovascular system [79], cardiac fibrosis and heart failure [80,81], thoracic aortic aneurysm [48], ascending aortic aneurysm [82], regulation of hypertension-related genes [83], myocardial infarction [84], insulin resistance [74], T2DM [85], T2DM with major cardiovascular events [86], T1DM [87,88,89], diabetic nephropathy [90]. |
| hsa-miR-23a-3p | 19p13.12 | Heart failure [91], coronary artery disease [92], cerebral ischemia-reperfusion [93], vascular endothelial dysfunction [51], small and large abdominal aortic aneurysm [94], obesity and insulin resistance [95]. |
| hsa-miR-24-3p | 19p13.12 | Asymptomatic carotid stenosis [96], familial hypercholesterolemia and coronary artery disease [97], angina pectoris [98], ischemic dilated cardiomyopathy [99], small and large abdominal aortic aneurysm [94], myocardial ischemia/reperfusion [100,101], diabetes mellitus [50,58,62,64]. |
| hsa-miR-26a-5p | 3p22.2 [102] 12q14.1 | Heart failure, cardiac hypertrophy [103], myocardial infarction [84,104,105], ischemia/reperfusion injury [106], pulmonary arterial hypertension [107], T1DM [108], diabetic nephropathy [90]. |
| hsa-miR-29a-3p | 7q32.3 | Ischemia/reperfusion-induced cardiac injury [109], cardiac cachexia, heart failure [110], atrial fibrillation [111], diffuse myocardial fibrosis in hypertrophic cardiomyopathy [64], coronary artery disease [112], pulmonary arterial hypertension [107], gestational diabetes mellitus [12], diabetes mellitus [49,57,113,114]. |
| hsa-miR-92a-3p | 13q31.3 Xq26.2 | Mitral chordae tendineae rupture [115], children with rheumatic carditis [116], myocardial infarction [117], heart failure [118], coronary artery disease [119], renal injury-associated atherosclerosis [120]. |
| hsa-miR-100-5p | 11q24.1 | Failing human heart, idiopathic dilated cardiomyopathy, ischemic cardiomyopathy [99], regulation of hypertension-related genes [83], T1DM [87]. |
| hsa-miR-103a-3p | 5q34 [121] 20p13 | Hypertension [122], hypoxia-induced pulmonary hypertension [123], myocardial ischemia/reperfusion injury, acute myocardial infarction [124], ischemic dilated cardiomyopathy [99], obesity, regulation of insulin sensitivity [125], T1DM [126]. |
| hsa-miR-125b-5p | 11q24.1 [126] 21q21.1 | Acute ischemic stroke [127], acute myocardial infarction [128,129], ischemic dilated cardiomyopathy [99], ascending aortic aneurysm [82], gestational diabetes mellitus [19], T1DM [130,131], T2DM [132]. |
| hsa-miR-126-3p | 9q34.3 [133] | Acute myocardial infarction [105], thoracic aortic aneurysm [48], T2DM [86,134], T2DM with major cardiovascular events [86], gestational diabetes mellitus [10]. |
| hsa-miR-130b-3p | 22q11.21 | Hypertriglyceridemia [135,136], intracranial aneurysms [137], hyperacute cerebral infarction [138], T2DM [85,139,140], gestational diabetes mellitus [10]. |
| hsa-miR-133a-3p | 18q11.2 [141] 20q13.33 | Heart failure [142], myocardial fibrosis in hypertrophic cardiomyopathy [64,143], arrhythmogenesis in the hypertrophic and failing hearts [144,145], coronary artery calcification [146], thoracic aortic aneurysm [48], ascending aortic aneurysm [82], diabetes mellitus [49,50]. |
| hsa-miR-143-3p | 5q33 | Intracranial aneurysms [147], coronary heart disease [148], myocardial infarction [149], myocardial hypertrophy [150], dilated cardiomyopathy [151], pulmonary arterial hypertension [152], acute ischemic stroke [127], ascending aortic aneurysm [82]. |
| hsa-miR-145-5p | 5q33 | Hypertension [153,154], dilated cardiomyopathy [155], myocardial infarction [156,157], stroke [157], acute cerebral ischemic/reperfusion [158], T2DM [58,159], T1DM [85], diabetic retinopathy [160], gestational diabetes mellitus [161]. |
| hsa-miR-146a-5p | 5q33.3 [162,163] | Angiogenesis [164], hypoxia, ischemia/reperfusion-induced cardiac injury [165], myocardial infarction [53], coronary atherosclerosis, coronary heart disease in patients with subclinical hypothyroidism [166], thoracic aortic aneurysm [48], acute ischemic stroke, acute cerebral ischemia [167], T2DM [58,85], T1DM [108], diabetic nephropathy [90]. |
| hsa-miR-155-5p | 21q21.3 | Thoracic aortic aneurysm [48], type 1 diabetes [125], gestational diabetes mellitus [20], adolescent obesity [168], diet-induced obesity and obesity resistance [168], atherosclerosis [170], hyperlipidemia—associated endotoxemia [171], coronary plaque rupture [172], children with cyanotic heart disease [173], chronic kidney disease and nocturnal hypertension [174], atrial fibrillation [175]. |
| hsa-miR-181a-5p | 1q32.1 [176] 9q33.3 | Regulation of hypertension-related genes [65], atherosclerosis [176], metabolic syndrome, coronary artery disease [177], non-alcoholic fatty liver disease [178], ischaemic stroke, transient ischaemic attack, acute myocardial infarction [179,180], obesity and insulin resistance [95,176,177], T1DM [85,181], T2DM [176,180]. |
| hsa-miR-195-5p | 17p13.1 [182] | Cardiac hypertrophy, heart failure [183,184], abdominal aortic aneurysms [185], aortic stenosis [186], T2DM [159], gestational diabetes mellitus [18]. |
| hsa-miR-199a-5p | 1q24.3 19p13.2 | T1DM, T2DM, gestational diabetes mellitus [187], diabetic retinopathy [188], cerebral ischemic injury [189], heart failure [190], hypertension [191,192], congenital heart disease [193], pulmonary artery hypertension [194], unstable angina [195], hypoxia in myocardium [193], acute kidney injury [196]. |
| hsa-miR-210-3p | 11p15.5 | Cardiac hypertrophy [197], acute kidney injury [198], myocardial infarction [199], atherosclerosis [200]. |
| hsa-miR-221-3p | Xp11.3 | Asymptomatic carotid stenosis [96], cardiac amyloidosis [201], heart failure [202], atherosclerosis [203,204], aortic stenosis [205], acute myocardial infarction [206], acute ischemic stroke [207], focal cerebral ischemia [208], pulmonary artery hypertension [209], obesity [210]. |
| hsa-miR-342-3p | 14q32.2 | Cardiac amyloidosis [201], obesity [211], T1DM [85,187,212], T2DM [187,213,214], gestational diabetes mellitus [187], endothelial dysfunction [215]. |
| hsa-miR-499a-5p | 20q11.22 | Myocardial infarction [53,216], hypoxia [217], cardiac regeneration [218], vascular endothelial dysfunction [51]. |
| hsa-miR-574-3p | 4p14 | Myocardial infarction [219], coronary artery disease [136], cardiac amyloidosis [201], stroke [220], T2DM [140,221]. |
T1DM: Diabetes mellitus type 1; T2DM: Diabetes mellitus type 2.
Previously, by searching the Medline database we identified a large scale of microRNAs playing a role in pathogenesis of diabetes mellitus and cardiovascular/cerebrovascular diseases. Finally, we selected for the study a shortlist of 29 microRNAs demonstrated repetitively by numerous scientific teams to be associated with normal stages (development and homeostasis of the cardiovascular system, angiogenesis, and adipogenesis) and pathological conditions and diseases (vascular endothelial dysfunction and inflammation, hypoxia, hypertension and regulation of hypertension-related genes, obesity, dyslipidaemia, atherosclerosis and atherosclerotic plaque formation, insulin resistance, diabetes mellitus and diabetes-related complications, metabolic syndrome, cardiovascular diseases involving the blood vessels (coronary and peripheral artery diseases, carotid artery disease, pulmonary arterial hypertension, cerebrovascular disease, aortic and intracranial aneurysms), cardiovascular diseases involving the heart (congenital heart disease, cardiomyopathies, cardiac dysrhythmias, hypertensive heart disease, myocardial disease, valvular heart disease, inflammatory heart disease, rheumatic heart disease, pulmonary heart disease, and heart failure), chronic kidney disease, ischemia/reperfusion injury, cardiac regeneration and cachexia) (Table 1) [46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117,118,119,120,121,122,123,124,125,126,127,128,129,130,131,132,133,134,135,136,137,138,139,140,141,142,143,144,145,146,147,148,149,150,151,152,153,154,155,156,157,158,159,160,161,162,163,164,165,166,167,168,169,170,171,172,173,174,175,176,177,178,179,180,181,182,183,184,185,186,187,188,189,190,191,192,193,194,195,196,197,198,199,200,201,202,203,204,205,206,207,208,209,210,211,212,213,214,215,216,217,218,219,220,221].
Furthermore, the MiRWalk database (http://www.umm.uni-heidelberg.de/apps/zmf/mirwalk/) was utilized to acquire data on the dysregulation of microRNAs, in particular, the abovementioned human disease and phenotype ontologies and OMIM disorders.
To our knowledge, no data on expression profiles of microRNAs associated with diabetes mellitus, cardiovascular, and cerebrovascular diseases in whole peripheral venous blood (white blood cells) of children descending from GDM affected pregnancies are currently available. The only study which was reported in this field was dedicated to profiling of mir-15 family members, affecting insulin signalling pathway, in skeletal muscle biopsies of adult offspring of women with a history of GDM [222].
2. Materials and Methods
2.1. Participants
The study had a prospective design, that ran from August 2016 to October 2019, and included Caucasian children aged 3 to 11 years descending from normal (n = 85) and GDM complicated pregnancies (n = 118). Children of equal age descending from pregnancies with a normal course of gestation were chosen as a control group.
Ninety-eight children were exposed to a GDM pregnancy on diet only, and 20 children descended from a GDM complicated pregnancy on the combination of diet and therapy (in nineteen pregnancies insulin was administrated and in one pregnancy metformin was prescribed). According to the Institutional Review Board (IRB) Guidelines for gestational diabetes mellitus mothers were divided into groups based on preconception BMI (below 18.5, 18.5–24.9, 25.0–29.9, above 30.0), the achievement of optimal total weight gain during pregnancy (12.5–18.0, 11.5–16.0, 7.0–11.5, 5.0–9.0 kg), and the achievement of optimal week weight gain during the second and the third trimesters of gestation (0.5–0.6, 0.4–0.5, 0.2–0.3, 0.2–0.3 kg) were monitored.
The clinical data of children descending from normal and GDM complicated pregnancies are summarized in Table 2. The preconceptional and gestational clinical data of mothers with normal and GDM complicated pregnancies are involved as well.
Table 2.
Characteristics of cases and controls.
| Normal Pregnancies Normal Clinical Findings (n = 48) | Normal Pregnancies Abnormal Clinical Findings (n = 37) | Gestational Diabetes Mellitus (GDM) Normal Clinical Findings (n = 61) | GDM Abnormal Clinical Findings (n= 57) | p-Value1 | p-Value2 | p-Value3 | |
|---|---|---|---|---|---|---|---|
| Children at follow-up | |||||||
| Age (years) | 5 (3–11) | 5 (3–11) | 5 (3–10) | 5 (3–9) | 0.228 | 0.980 | 0.358 |
| Height (cm) | 115.5 (98–144.5) | 118.0 (100–153) | 114.0 (99–143.5) | 113.5 (98–153) | 0.332 | 0.546 | 0.274 |
| Weight (kg) | 20.8 (14–37) | 22.3 (14.7–40.8) | 19.5 (14.4–37.4) | 19.6 (15–47.1) | 0.104 | 0.604 | 0.565 |
| BMI (kg/m2) | 15.41 (13.22–18.09) | 15.80 (13.3–20) | 15.20 (13.53–18.09) | 15.58 (12.97–20.08) | 0.067 | 0.976 | 0.166 |
| Systolic BP (mmHg) | 98 (84–115) | 104 (89–123) | 99 (82–113) | 101 (87–125) | <0.001 | 0.864 | 0.027 |
| Diastolic BP (mmHg) | 60 (38–68) | 64.0 (43–81) | 60 (47–67) | 61 (49–79) | 0.002 | 0.795 | 0.008 |
| Heart rate (n/min) | 90 (67–110) | 90.0 (51–120) | 96 (64–118) | 98 (78–122) | 0.954 | 0.022 | <0.001 |
| During Gestation | |||||||
| Maternal age at delivery (years) | 31.5 (21–40) | 32 (25–46) | 34 (27–42) | 33 (27–45) | 0.233 | 0.009 | 0.046 |
| GA at delivery (weeks) | 39.86 (37.71–41.57) | 39.86 (37.86–41.86) | 39.72 (37.43–41.28) | 39.43 (37.00–41.14) | 0.852 | 0.111 | 0.020 |
| Fetal birth weight (g) | 3425 (2730–4220) | 3280 (2530–4450) | 3500 (2700-4330) | 3420 (2770–4400) | 0.989 | 0.157 | 0.252 |
| Mode of delivery | 0.698 | <0.001 | 0.001 | ||||
| Vaginal | 44 (91.67%) | 33 (89.19%) | 38 (62.30%) | 37 (64.91%) | |||
| CS | 4 (8.33%) | 4 (10.81%) | 23 (37.70%) | 20 (35.09%) | |||
| Fetal sex | 0.346 | 0.771 | 0.273 | ||||
| Boy | 27 (56.25%) | 17 (45.95%) | 36 (59.02%) | 38 (66.67%) | |||
| Girl | 21 (43.75%) | 20 (54.05%) | 25 (40.98%) | 19 (33.33%) | |||
| Primiparity | 0.069 | 0.181 | 0.091 | ||||
| Yes | 29 (60.42%) | 15 (40.54%) | 29 (47.54%) | 25 (43.86%) | |||
| No | 19 (39.58%) | 22 (59.46%) | 32 (52.46%) | 32 (56.14%) | |||
| Birth order of index pregnancy | 0.058 | 0.080 | 0.294 | ||||
| 1st | 26 (54.17%) | 11 (29.73%) | 20 (32.79%) | 22 (38.60%) | |||
| 2nd | 16 (33.33%) | 14 (37.84%) | 23 (37.70%) | 22 (38.60%) | |||
| 3rd | 4 (8.33%) | 10 (27.03%) | 10 (16.39%) | 6 (10.52%) | |||
| 4th+ | 2 (4.17%) | 2 (5.40%) | 8 (13.11%) | 7 (12.28%) | |||
| Infertility treatment | 0.852 | 0.014 | 0.084 | ||||
| Yes | 1 (2.08%) | 1 (2.70%) | 10 (16.39%) | 6 (10.53%) | |||
| No | 47 (97.92%) | 36 (97.30%) | 51 (83.61%) | 51 (89.47%) | |||
| Maternal BMI (kg/m2) | |||||||
| Prepregnancy BMI | 21.88 (14.77–30.3) | 21.22 (17.37–28.04) | 22.02 (16.3–30.85) | 22.64 (17.53–30.49) | 1.000 | 1.000 | 0.567 |
| BMI < 18.5 | 4 (8.33%) | 5 (13.51%) | 5 (8.20%) | 1 (1.75%) | - | - | - |
| BMI 18.5–24.9 | 38 (79.17%) | 31 (83.78%) | 43 (70.49%) | 37 (64.91%) | |||
| BMI 25.0–29.9 | 5 (10.42%) | 1 (2.70%) | 11 (18.03%) | 17 (29.82%) | |||
| BMI > 30 | 1 (2.08%) | 0 (0%) | 2 (3.28%) | 2 (3.51%) | |||
| BMI at admission for delivery | 26.17 (20.88–34.82) | 26.45 (20.72–33.17) | 25.97 (19.84–36.85) | 27.53 (20.18–36.73) | 1.000 | 1.000 | 1.000 |
| Total gestational weight gain (GWG) (kg) | 14.5 (8–25.5) | 14.5 (8–21) | 10 (2–21) | 11 (3–26) | 1.000 | <0.001 | 0.008 |
| BMI at follow-up | 23.26 (17.7–39.08) | 22.11 (18.17–29.17) | 23.82 (17.39–32.14) | 23.42 (17.39–34.37) | 0.414 | 1.000 | 1.000 |
| Serum Metabolic Biochemical Parameters of Mothers During the Third Trimester of Gestation | |||||||
| HbA1c | - | - | 32 (26–42) | 32.5 (26–40) | - | - | - |
| Creatinine (μmol/L) | 56.0 (44.0–70.0) | 52.5 (43.0–83.0) | 56.0 (40.0–78.0) | 55.0 (36.0–83.0) | 1.000 | 1.000 | 1.000 |
| Uric acid (μmol/L) | 286.5 (200.0–377.0) | 294.5 (221.0–345.0) | 298 (180.0–419.0) | 289 (157.0–471.0) | 1.000 | 1.000 | 1.000 |
| Total bilirubin (μmol/L) | 4.0 (3.0–7.0) | 4.0 (0.18–12.0) | 6.0 (2.70–20.0) | 6.0 (3.0–20.0) | 1.000 | 0.497 | 0.108 |
| ALT (μkat/L) | 0.22 (0.11–2.4) | 0.17 (0.07–0.36) | 0.23 (0.11–1.09) | 0.22 (0.11–0.46) | 1.000 | 1.000 | 1.000 |
| AST (μkat/L) | 0.38 (0.25–2.36) | 0.47 (0.22–2.29) | 0.38 (0.22–1.46) | 0.40 (0.24–0.84) | 1.000 | 1.000 | 1.000 |
| ALP (μkat/L) | 2.12 (1.86–4.66) | 2.51 (1.71–3.52) | 2.67 (1.35–6.15) | 2.36 (1.46–5.32) | 1.000 | 1.000 | 1.000 |
| Cholesterol (mmol/L) | 7.50 (6.30–11.40) | 9.2 (6.90–10.5)) | 7.71 (4.47–9.5) | 6.93 (6.13–9.10) | 1.000 | 1.000 | 1.000 |
| Triglyceride (mmol/L) | 3.10 (3.00–3.20) | 2.9 (2.7–3.3) | 3.05 (1.80–4.80) | 3.35 (2.50–8.90) | 1.000 | 1.000 | 1.000 |
| Total protein (g/L) | 63.7 (48.7–70.8) | 64,35 (47.2–68.9) | 60.3 (37.7–70.8) | 61.6 (43.8–67.9%) | 1.000 | 1.000 | 1.000 |
| Albumin (g/L) | 37.35 (28.9–40.6) | 36.9 (27.9–41.6) | 36.5 (4.40–42.1) | 36.2(26.3–40.9) | 1.000 | 1.000 | 1.000 |
| Blood glucose (mmol/L) | 4.7 (4.2–5.1) | 4.5 (4.3–5.8) | 4.65 (4.1–8.0) | 4.4 (3.8–5.6) | 1.000 | 1.000 | 1.000 |
Data are presented as a median (range) for continuous variables and as a number (percent) for categorical variables. Continuous variables were compared using the non-parametric Kruskal-Wallis test. P-value1: The comparison among children from the control group with normal and abnormal postnatal clinical findings; p-value2: The comparison among children descending from normal and GDM complicated pregnancies with normal postnatal clinical findings; p-value3: The comparison among children descending from normal pregnancies with normal postnatal clinical findings and children descending from GDM complicated pregnancies with abnormal postnatal clinical findings. Categorical variables were compared using a chi-square test. GDM: Gestational diabetes mellitus; BP: Blood pressure; CS: Caesarean section; GA: Gestational age; GWG: Gestational weight gain; HbA1c: Haemoglobin A1c; ALT: Alanine aminotransferase; AST: Aspartate aminotransferase; ALP: Alkaline phosphatase.
Children descending from normal pregnancies were healthy infants born after 37 completed weeks of gestation with the weight >2500 g. Normal pregnancies were those ones with the absence of medical, obstetrical, or surgical complications.
Gestational diabetes mellitus was diagnosed if any degree of glucose intolerance appeared for the first-time during gestation [223,224,225]. Concerning the diagnosis and classification of hyperglycemia in pregnancy the guidelines of The International Association of Diabetes and Pregnancy Study Groups (IADPSG) were followed [223]. The first screening was performed during the first trimester of gestation with the aim to detect women with overt diabetes (fasting plasma glucose level is ≥7.0 mmol/L) and women with GDM (fasting plasma glucose level ≥5.1–<7.0 mmol/L). The second screening phase, 2 h 75-g OGTT, was performed during 24–28 weeks of gestation only in those women not previously diagnosed to have overt diabetes or GDM. The second screening phase identified GDM if the fasting plasma glucose level was ≥5.1 mmol/L, or the 1 h plasma glucose was ≥10.0 mmol/L, or the 2-h plasma glucose was ≥8.5 mmol/L [223].
Children with inborn defects or chromosomal disorders, as well as children descending from pregnancies demonstrating other complications were excluded from the inclusion into the study.
Informed consent was obtained from all participants included in the study. Two independent Ethics Committees (one at the Institute for the Care of the Mother and Child, and the second one at the Third Faculty of Medicine, Charles University) approved the study (grant no. AZV 16-27761A, long-term monitoring of complex cardiovascular profile in the mother, foetus, and offspring descending from pregnancy-related complications, dates of approval: 27 March, 2014 and 28 May, 2015). All procedures were also in compliance with the Helsinki Declaration of 1975, as revised in 2000.
2.2. BP and Echocardiography Measurements, BMI Assessment
Standardized BP and echocardiography measurements, and BMI assessment were performed as previously described [226]. In brief, the average of the last two systolic blood pressure (SBP) and diastolic blood pressure (DBP) values was taken under consideration for data analyses. A normal BP was defined as SBP and DBP that was below the 90th percentile for gender, age, and height. Hypertension was defined as an average SBP or DBP ≥95th percentile on at least three separate occasions. Prehypertension was defined as an average SBP or DBP within the range of ≥90th percentile and <95th percentile [227].
The age- and sex-specific BMI Percentile Calculator was used to calculate BMI in children (https://www.cdc.gov/healthyweight/assessing/bmi/childrens_bmi/about_childrens_bmi.html). BMI above the 5th percentile and below the 85th percentile was considered as normal BMI. Children with BMI above the 85th percentile and below the 95th percentile were in the overweight category. Children with BMI equal to or greater than the 95th percentile were in the obese category.
A complete two-dimensional echocardiography was performed using the Philips HD15 ultrasound machine (Philips Ultrasound, Bothell, WA, USA) with a sector array transducer (3–8 MHz) incorporating colour flow, pulse wave, and continuous wave Doppler measurements with an adaptive technology. A complete two-dimensional echocardiography was performed by an investigator experienced with paediatric echocardiography. Children with abnormal findings were referred to a paediatric cardiologist.
2.3. Processing of Samples
Samples of unclotted whole peripheral venous blood (200 µL) were processed as previously described [226]. Briefly, homogenized cell lysates were prepared as soon as possible after blood collection using a QIAamp RNA Blood Mini Kit (Qiagen, Hilden, Germany, no. 52304).
Afterwards, total RNA was extracted using a mirVana microRNA Isolation kit (Ambion, Austin, TX, USA, no. AM1560). The isolated RNA was treated with DNase I (Thermo Fisher Scientific, Carlsbad, CA, USA, no. EN0521).
2.4. Reverse Transcription
Particular microRNAs were transcribed into cDNA using microRNA-specific stem-loop RT primers, components of TaqMan MicroRNA Assays and TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems, Branchburg, NJ, USA, no. 4366597) as previously described [226]. A total reaction volume of the reaction was 10 µL. The reverse transcriptase reaction was performed following the guidelines for a 7500 Real-Time PCR system (Applied Biosystems, Branchburg, NJ, USA): 30 min at 16 °C, 30 min at 42 °C, 5 min at 85 °C, and then held at 4 °C.
2.5. Relative Quantification of microRNAs
Relative quantification of microRNAs by real-time PCR was performed as previously described [226]. cDNA (3 µL) was mixed with specific TaqMan MGB primers and probes (TaqMan MicroRNA Assay, Applied Biosystems, Branchburg, NJ, USA), and the constituents of the TaqMan Universal PCR Master Mix (Applied Biosystems, Branchburg, NJ, USA, no: 4318157) in a total reaction volume of 15 µL. The samples were regarded as positive if Ct (threshold cycle) was below 40 (Ct < 40).
The comparative Ct method was used to determine the expression of each microRNA [228]. The expression of studied microRNAs was normalized to a geometric mean of RNU58A and RNU38B, endogenous controls with the lowest expression variability between studied samples [229]. A reference sample was used throughout the study for relative quantification. As a reference sample we used small RNAs enriched RNA fraction extracted from the fetal part of one randomly selected placenta of a normally ongoing pregnancy.
2.6. Statistical Analysis
Logistic regression was used to compare the presence of abnormal clinical findings (BMI in the category overweight or obesity and/or systolic or diastolic BP values in the category prehypertension or hypertension and/or the presence of valve problems and heart defects) among various groups of children.
The Shapiro-Wilk test was used to assess the data normality [230]. Our experimental data did not follow a normal distribution. Therefore, microRNA levels were compared among the groups of children using the Kruskal-Wallis one-way analysis of variance with a post-hoc test (K-W) for the comparison between multiple groups.
Receivers operating characteristic (ROC) curves were used to assess the areas under the curves (AUC), the optimal cut-off points, and the respective sensitivities at 10.0% false positive rate (FPR) for particular microRNAs (MedCalc Software bvba, Ostend, Belgium). The significance level was established at a p-value of p < 0.05.
To identify the optimal combinations of microRNA biomarkers logistic regression combined with the ROC curve analysis was used (MedCalc Software bvba, Ostend, Belgium). In brief, in this setting, the power of the model’s predicted values to discriminate between positive and negative cases is quantified by the area under the ROC curve. To perform a full ROC curve analysis the predicted probabilities are first saved and next used as a new variable in the ROC curve analysis. The dependent variable used in logistic regression then acts as the classification variable in the ROC curve analysis dialog box.
Statistica software (version 9.0; StatSoft, Inc., Tulsa, OK, USA) was used to generate box plots of log-normalized gene expression values (RT-qPCR expression, log10 2−ΔΔCt) for particular microRNAs. The box plots display the medians, the 75th and 25th percentiles (the upper and lower limits of the boxes), the maximum and minimum values (the upper and lower whiskers), outliers (circles), and extremes (asterisks). Dot plots, all observations are also displayed in the charts.
2.7. Bioinformatics Analysis—microRNA-Target Interactions on Disease Ontologies, Human Phenotype Ontologies, and OMIM Disorders
MiRWalk database (http://www.umm.uni-heidelberg.de/apps/zmf/mirwalk/) and the Predicted Target module [231] were utilized to acquire data on predicted targets for microRNAs that have been identified to be dysregulated in whole peripheral venous blood of children descending from GDM complicated pregnancies.
MiRWalk is a comprehensive database that provides in addition to other things also information on human microRNA predicted and/or validated target genes, information on microRNA-target interactions on disease ontologies, human phenotype ontologies and OMIM disorders, and information on possible interactions between microRNAs and genes associated with KEGG, Panther and Wiki pathways.
Only those predicted targets involved in ontologies of human diseases (obesity, hypertension, glucose intolerance, lipid metabolism disease, type 2 diabetes mellitus, heart septal defects and heart valve disease, heart disease, heart failure, venous insufficiency, and pulmonary embolism) reported by previous population-based cohort studies in children and young adults descending from GDM complicated pregnancies [26,27,28,29,30,31,32,33,34,35,36] are presented below.
2.8. Workflow of the Study
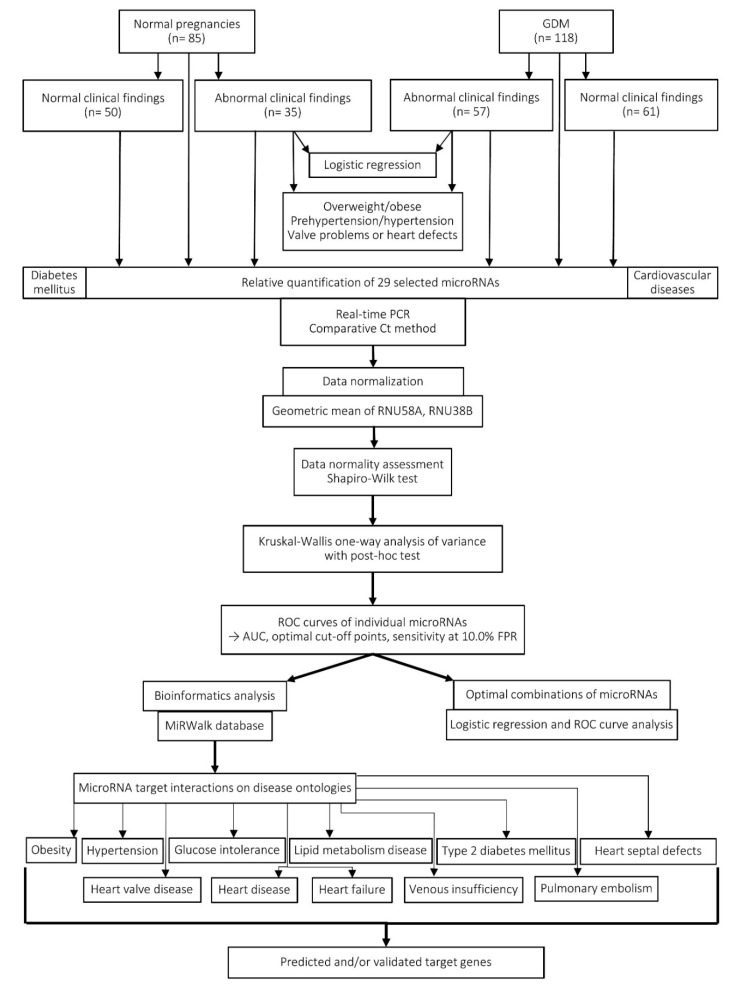
Figure 1 presents the workflow of the study including the particular groups of the participants, the performance of relative quantification of microRNAs and bioinformatics analyses, and the variety of statistical analyses applied.
Figure 1.
Workflow of the study.
3. Results
3.1. Higher Incidence of Valve Problems and Heart Defects in a group of Children Descending from GDM Complicated Pregnancies
We distributed children descending from normal and GDM complicated pregnancies into groups based on the results of anamnesis and the results of consecutive clinical examination. The group of children with abnormal clinical findings consisted of those ones who were already dispensarised in the department of paediatric cardiology, or those ones who were overweight/obese, had prehypertension/hypertension, and/or abnormal echocardiogram findings during the visit (in total: FG, n = 35; GDM, n = 57).
In detail, the studied groups of children with abnormal clinical findings consisted of those ones already dispensarised in the department of paediatric cardiology (FG, n = 8/85; GDM, n = 5/118), those ones indicated by the sonographer during the visit to have valve problems and heart defects (FG, n = 17/85; GDM, n = 42/118) (tricuspid valve regurgitation (FG, n = 8/85; GDM, n = 23/118), mitral valve regurgitation (FG, n = 1/85; GDM, n = 1/118), pulmonary valve regurgitation (FG, n = 2/85; GDM, n = 8/118), bicuspid aortic valve regurgitation (FG, n = 1/85; GDM, n = 0/118), ventricular septum defect (FG, n = 1/85; GDM, n = 0/118), atrial septum defect (FG, n = 1/85; GDM, n = 2/118), foramen ovale apertum (FG, n = 5/85; GDM, n = 11/118), arrhythmia (FG, n = 1/85; GDM, n = 1/118)], those ones confirmed over several visits to have a high BP (FG, n = 15/85; GDM, n = 19/118) (SBP and/or DBP ≥ 90th percentile evaluated by the Age-based Paediatric Blood Pressure Reference Charts calculator) and/or high BMI (FG, n = 8/85; GDM, n = 6/118) (BMI >85th percentile evaluated by the BMI Percentile Calculator for Child and Teens).
The group of children with normal clinical findings consisted of children with normal anamnesis, normal BP, normal BMI, and normal reference values of echocardiographic measurements (FG, n = 50/85; GDM, n = 61/118).
Logistic regression revealed no difference in the incidence of overweight/obesity and/or prehypertension/hypertension between the groups of children descending from normal and GDM complicated pregnancies. Nevertheless, a higher incidence of valve problems and heart defects was observed in a group of children descending from GDM complicated pregnancies (Table 3).
Table 3.
The results of clinical examination in children descending from normal and GDM complicated pregnancies.
| Normal Pregnancy | GDM Pregnancy | OR (95% CI) | p-Value | |
|---|---|---|---|---|
| Overweight/obese | 8/85 (9.41%) | 6/118 (5.08%) | 0.515 (0.172–1.545) | 0.237 |
| Prehypertension/ hypertension | 15/85 (17.65%) | 19/118 (16.10%) | 0.896 (0.426–1.883) | 0.771 |
| Valve problems or heart defects | 17/85 (20.0%) | 42/118 (35.59%) | 2.240 (1.167–4.300) | 0.015 |
Logistic regression was used to compare the presence of abnormal clinical findings between particular groups. The significance level was established at a p-value of p < 0.05. No difference in the incidence of overweight/obesity and/or prehypertension/hypertension was found between the groups of children descending from normal and GDM complicated pregnancies. A higher incidence of valve problems and heart defects was observed in a group of children descending from GDM complicated pregnancies.
3.2. Only Just miR-21-5p Indicates a Trend Towards Differentiation between Children with Normal and Abnormal Clinical Findings (BMI, Blood Pressure, and Echocardiogram Findings) Descending from Normal Pregnancies
Furthermore, we compared the microRNA expression profile between individual groups of children with respect to clinical findings. From the ROC curve analysis, it is apparent that only miR-21-5p (a sensitivity of 17.14% at a specificity of 90.0%) trended to differentiate between children descending from normal pregnancies with normal and abnormal clinical findings (BMI, blood pressure, and echocardiogram findings) (Figure S1).
3.3. Only Just miR-29a-3p and miR-92a-3p Indicate a Slight Differentiation Between Children with Normal and Abnormal Clinical Findings (BMI, Blood Pressure, and Echocardiogram Findings) Descending from GDM Complicated Pregnancies
The performance of the ROC curve analysis revealed that miR-29a-3p (12.28%) and miR-92a-3p (12.28%) were able to slightly differentiate at 10.0% FPR within children descending from GDM complicated pregnancies based on the presence of normal and abnormal clinical findings (BMI, blood pressure, and echocardiogram findings) (Figure S2).
3.4. Substantially Altered Expression Profile of Diabetes/Cardiovascular/Cerebrovascular Disease Associated microRNAs in Children Descending from Pregnancy Complicated by Gestational Diabetes Mellitus
With regard to the assessment of the effect of maternal pregnancy complication on postnatal microRNA expression profile, we further compared microRNA gene expression between children descending from normal and GDM complicated pregnancies irrespective of the clinical findings (overweight/obesity, prehypertension/hypertension, and/or valve problems and heart defects). The K-W and subsequent ROC curve analysis revealed a significant dysregulation of multiple microRNAs in children descending from GDM complicated pregnancies (Figure 2, Figure S3).
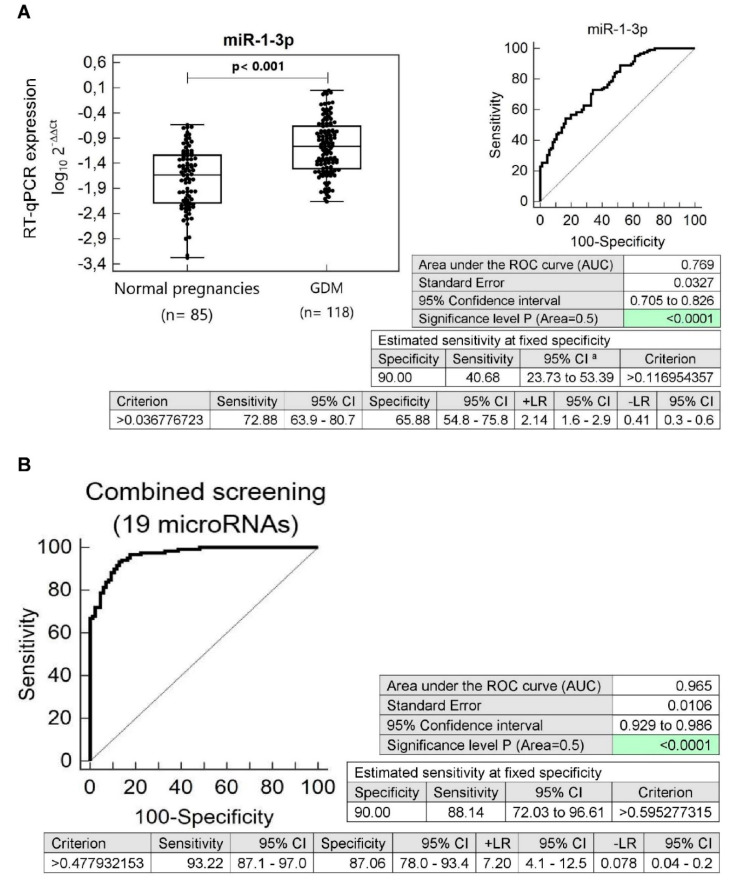
Figure 2.
Aberrant microRNA expression profile in children descending from GDM complicated pregnancies irrespective of the clinical findings (overweight/obesity, prehypertension/hypertension, and/or valve problems and heart defects). (A) Upregulation of miR-1-3p was observed in children descending from GDM complicated pregnancies when the comparison to the controls irrespective of the clinical findings was performed. Concerning individual microRNAs, miR-1-3p showed the highest accuracy for the identification of children at a higher risk of later development of diabetes mellitus and/or cardiovascular/cerebrovascular diseases. (B) Combined screening of microRNAs in the identification of children prenatally exposed to GDM at an increased risk of later development of diabetes mellitus and/or cardiovascular/cerebrovascular diseases. Screening based on the combination of microRNAs with a good sensitivity (miR-1-3p, miR-16-5p, miR-17-5p, miR-20a-5p, miR-20b-5p, miR-21-5p, miR-26a-5p, miR-29a-3p, miR-103a-3p, miR-125b-5p, miR-126-3p, miR-133a-3p, miR-143-3p, miR-181a-5p, miR-195-5p, miR-210-3p, miR-221-3p, miR-499a-5p, and miR-574-3p) showed the highest accuracy for the identification of children at a higher risk of later development of diabetes mellitus and/or cardiovascular/cerebrovascular diseases. GDM: Gestational diabetes mellitus.
The sensitivity at 10.0% FPR for miR-1-3p (40.68%), miR-16-5p (14.41%), miR-17-5p (22.88%), miR-20a-5p (27.97%), miR-20b-5p (29.66%), miR-21-5p (16.10%), miR-26a-5p (17.80%), miR-29a-3p (19.49%), miR-92a-3p (6.78%), miR-103a-3p (20.34%), miR-125b-5p (24.58%), miR-126-3p (25.42%), miR-133a-3p (22.03%), miR-143-3p (18.64%), miR-146a-5p (8.47%), miR-155-5p (5.08%), miR-181a-5p (16.10%), miR-195-5p (27.97%), miR-199a-5p (11.02%), miR-210-3p (15.25%), miR-221-3p (17.80%), miR-499a-5p (27.97%), and miR-574-3p (18.64%) was observed (Figure 2, Figure S3).
MicroRNAs with a poor sensitivity at 10.0% FPR (miR-92a-3p, miR-146a-5p, miR-155-5p, and miR-199a-5p) were not further used for diabetes mellitus and cardiovascular risk assessment in children descending from GDM complicated pregnancies.
Screening based on a combination of microRNAs with a good sensitivity (miR-1-3p, miR-16-5p, miR-17-5p, miR-20a-5p, miR-20b-5p, miR-21-5p, miR-26a-5p, miR-29a-3p, miR-103a-3p, miR-125b-5p, miR-126-3p, miR-133a-3p, miR-143-3p, miR-181a-5p, miR-195-5p, miR-210-3p, miR-221-3p, miR-499a-5p, and miR-574-3p) showed the highest accuracy for children prenatally exposed to GDM (AUC 0.965, p < 0.001, sensitivity 93.22%, specificity 87.06%, cut off >0,477932153). At 10.0% FPR, it was able to identify 88.14% children with an increased risk of later development of diabetes and/or cardiovascular/cerebrovascular diseases (Figure 2).
3.5. Dysregulation of Multiple microRNAs in Children Descending from GDM Complicated Pregnancies with Normal Clinical Findings (Normal BMI, Blood Pressure, and Echocardiogram Findings)
Furthermore, we compared the microRNA expression profile between individual groups of children with respect to clinical findings. Despite the presence of normal clinical findings in both groups, that were compared, the performance of the ROC curve analysis revealed that miR-1-3p (57.38%), miR-16-5p (9.84%), miR-17-5p (31.15%), miR-20a-5p (29.51%), miR-20b-5p (32.79%), miR-21-5p (21.31%), miR-23a-3p (26.23%), miR-26a-5p (8.20%), miR-29a-3p (22.95%), miR-100-5p (22.95%), miR-103a-3p (22.95%), miR-125b-5p (36.07%), miR-126-3p (24.59%), miR-133a-3p (27.87%), miR-143-3p (16.39%), miR-146a-5p (9.84%), miR-181a-5p (21.31%), miR-195-5p (19.67%), miR-210-3p (13.11%), miR-221-3p (18.03%), miR-499a-5p (29.51%), and miR-574-3p (16.39%) differentiated between children descending from GDM complicated and normal pregnancies with a various sensitivity at a specificity of 90.0% (Figure 3, Figure S4).
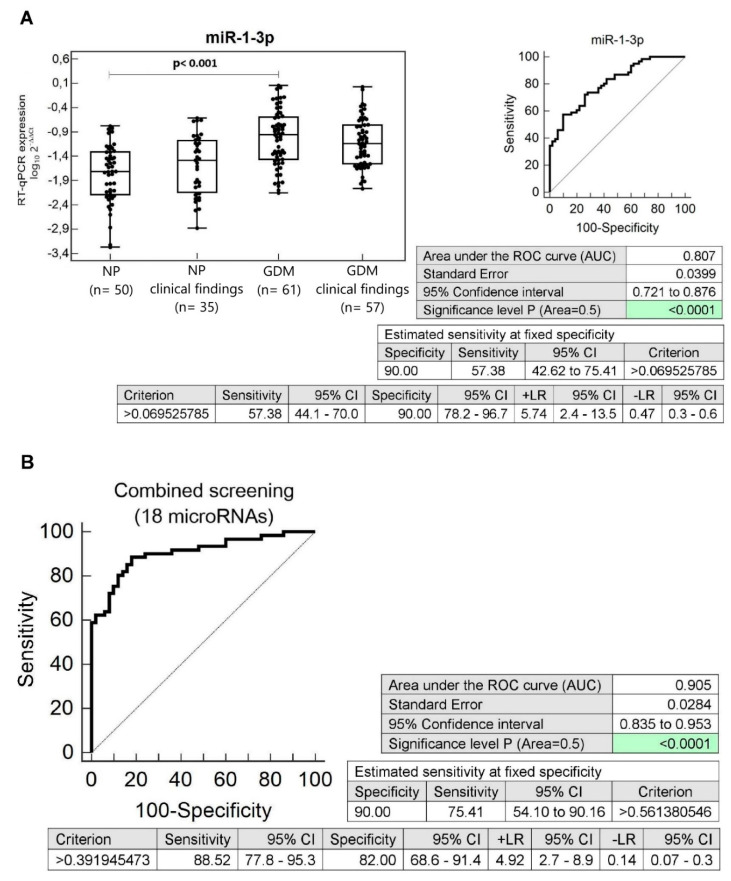
Figure 3.
Aberrant microRNA expression profile in children descending from GDM complicated pregnancies with normal clinical findings. (A) Upregulation of miR-1-3p was observed in children descending from GDM complicated pregnancies with normal clinical findings, when the comparison to the controls with normal clinical findings was performed. Concerning individual microRNAs, miR-1-3p showed the highest accuracy for the identification of children at a higher risk of later development of diabetes mellitus and/or cardiovascular/cerebrovascular diseases. (B) Combined screening of microRNAs in the identification of children prenatally exposed to GDM with normal clinical findings at an increased risk of later development of diabetes mellitus and/or cardiovascular/cerebrovascular diseases. Screening based on the combination of microRNAs with a good sensitivity (miR-1-3p, miR-17-5p, miR-20a-5p, miR-20b-5p, miR-21-5p, miR-23a-3p, miR-29a-3p, miR-100-5p, miR-103a-3p, miR-125b-5p, miR-126-3p, miR-133a-3p, miR-143-3p, miR-181a-5p, miR-195-5p, miR-221-3p, miR-499a-5p, and miR-574-3p) showed the highest accuracy for the identification of children prenatally exposed to GDM with normal clinical findings at a higher risk of later development of diabetes mellitus and/or cardiovascular/cerebrovascular diseases. The comparison to the controls with normal clinical findings was performed. NP: Normal pregnancies; GDM: Gestational diabetes mellitus.
Combined screening of miR-1-3p, miR-17-5p, miR-20a-5p, miR-20b-5p, miR-21-5p, miR-23a-3p, miR-29a-3p, miR-100-5p, miR-103a-3p, miR-125b-5p, miR-126-3p, miR-133a-3p, miR-143-3p, miR-181a-5p, miR-195-5p, miR-221-3p, miR-499a-5p, and miR-574-3p was superior over using individual microRNAs in the assessment of risk of later development of diabetes mellitus and cardiovascular/cerebrovascular diseases (AUC 0.905, p < 0.001, sensitivity 88.52%, specificity 82.0%, cut off >0.391945473) in a group of children descending from GDM complicated pregnancies with normal clinical findings. At 10.0% FPR, it was able to identify 75.41% children with an increased diabetic/cardiovascular risk (Figure 3).
3.6. Dysregulation of Multiple microRNAs in Children Descending from GDM Complicated Pregnancies with Abnormal Clinical Findings (Abnormal BMI, Blood Pressure, and Echocardiogram Findings)
In addition, it was observed that a set of microRNAs differentiated between the groups of children affected with GDM with abnormal clinical findings (abnormal BMI, blood pressure, and echocardiogram findings) and the controls, children descending from normal pregnancies with normal clinical findings. The sensitivity of individual microRNAs at 10.0% FPR was the following: miR-1-3p (50.88%), miR-17-5p (22.81%), miR-20a-5p (22.81%), miR-20b-5p (26.32%), miR-21-5p (22.81%), miR-29a-3p (15.79%), miR-92a-3p (19.30%), miR-103a-3p (17.54%), miR-126-3p (22.81%), miR-133a-3p (19.30%), miR-143-3p (15.79%), miR-155-5p (7.02%), miR-181a-5p (19.30%), miR-195-5p (29.82%), miR-210-3p (12.28%), miR-221-3p (15.79%), and miR-499a-5p (29.82%) (Figure 4, Figure S5).
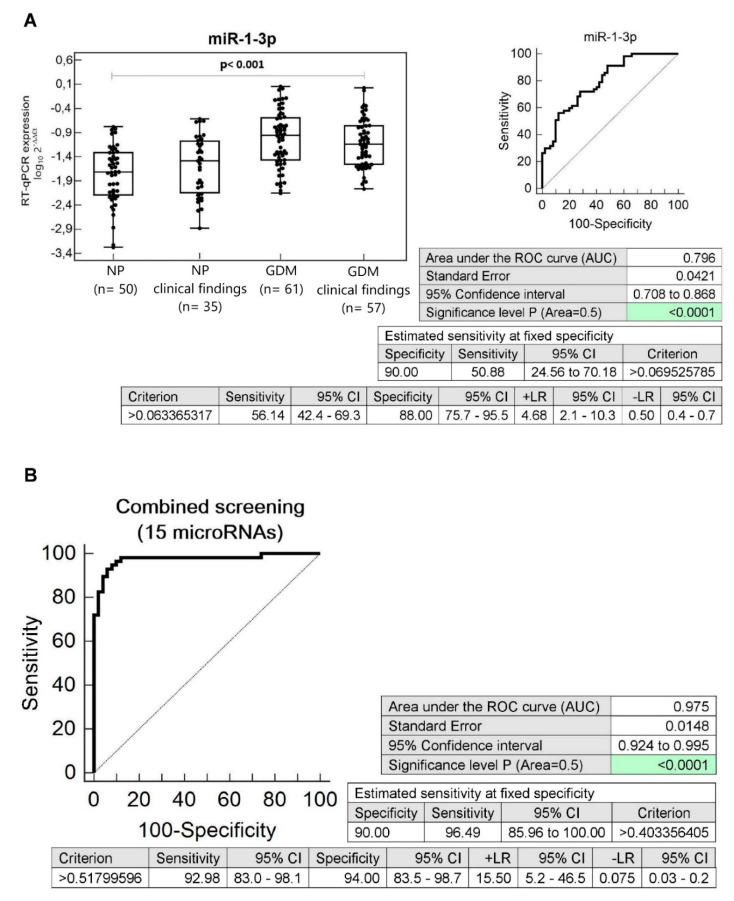
Figure 4.
Aberrant microRNA expression profile in children descending from GDM complicated pregnancies with abnormal clinical findings. (A) Upregulation of miR-1-3p was observed in children descending from GDM complicated pregnancies with abnormal clinical findings, when the comparison to the controls with normal clinical findings was performed. Concerning individual microRNAs, miR-1-3p showed the highest accuracy for the identification of children at a higher risk of later development of diabetes mellitus and/or cardiovascular/cerebrovascular diseases. (B) Combined screening of microRNAs in the identification of children prenatally exposed to GDM with abnormal clinical findings at an increased risk of later development of diabetes mellitus and/or cardiovascular/cerebrovascular diseases. Screening based on the combination of microRNAs with a good sensitivity (miR-1-3p, miR-17-5p, miR-20a-5p, miR-20b-5p, miR-21-5p, miR-29a-3p, miR-92a-3p, miR-103a-3p, miR-126-3p, miR-133a-3p, miR-143-3p, miR-181a-5p, miR-195-5p, miR-221-3p, and miR-499a-5p) showed the highest accuracy for the identification of children prenatally exposed to GDM with abnormal clinical findings at a higher risk of later development of diabetes mellitus and/or cardiovascular/cerebrovascular diseases. The comparison to the controls with normal clinical findings was performed. NP: Normal pregnancies; GDM: Gestational diabetes mellitus.
Screening based on the combination of microRNAs with a good sensitivity (miR-1-3p, miR-17-5p, miR-20a-5p, miR-20b-5p, miR-21-5p, miR-29a-3p, miR-92a-3p, miR-103a-3p, miR-126-3p, miR-133a-3p, miR-143-3p, miR-181a-5p, miR-195-5p, miR-221-3p, and miR-499a-5p) showed the highest accuracy for prediction of diabetes/cardiovascular risk (96.49% children at 10.0% FPR) in children with abnormal clinical findings with a prior exposure to GDM (AUC 0.975, p < 0.001, sensitivity 92.98%, specificity 94.0%, cut off >0.51799596) (Figure 4).
3.7. Information on microRNA-Target Interactions on Disease Ontologies, Human Phenotype Ontologies, and OMIM Disorders
The extensive file of predicted targets of all microRNAs with altered expression in whole peripheral blood of children descending from GDM complicated pregnancies indicates that a large group of these genes is involved in ontologies of human diseases (obesity, hypertension, glucose intolerance, lipid metabolism disease, type 2 diabetes mellitus, heart septal defects and heart valve disease, heart disease, heart failure, venous insufficiency, and pulmonary embolism) reported by several previous population-based cohort studies [5,6,7,8,9,10,11,12,13,14,15] (Tables S1–S11).
4. Discussion
A diabetes/cardiovascular/cerebrovascular disease associated microRNA expression profile was assessed in whole peripheral blood of children at the age of 3 to 11 years with a prior exposure to GDM with the aim to assess to what extent fetal and environmental programming predisposes the affected children to later development of diabetes mellitus, cardiovascular, and cerebrovascular diseases.
The hypothesis of the assessment of potential diabetes/cardiovascular risk in children prenatally exposed to GDM was based on the knowledge that a serious of microRNAs play a role in the pathogenesis of diabetes mellitus and cardiovascular/cerebrovascular diseases (Table 1) [46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117,118,119,120,121,122,123,124,125,126,127,128,129,130,131,132,133,134,135,136,137,138,139,140,141,142,143,144,145,146,147,148,149,150,151,152,153,154,155,156,157,158,159,160,161,162,163,164,165,166,167,168,169,170,171,172,173,174,175,176,177,178,179,180,181,182,183,184,185,186,187,188,189,190,191,192,193,194,195,196,197,198,199,200,201,202,203,204,205,206,207,208,209,210,211,212,213,214,215,216,217,218,219,220,221].
Surprisingly, a substantially altered expression profile of diabetes/cardiovascular/cerebrovascular disease associated microRNAs (23/29 studied microRNAs: miR-1-3p, miR-16-5p, miR-17-5p, miR-20a-5p, miR-20b-5p, miR-21-5p, miR-26a-5p, miR-29a-3p, miR-92a-3p, miR-103a-3p, miR-125b-5p, miR-126-3p, miR-133a-3p, miR-143-3p, miR-146a-5p, miR-155-5p, miR-181a-5p, miR-195-5p, miR-199a-5p, miR-210-3p, miR-221-3p, miR-499a-5p, and miR-574-3p) was found in children descending from GDM complicated pregnancies when compared with children descending from pregnancies with a normal course of gestation. Almost all microRNAs with the exception of miR-92a-3p, miR-155-5p, and miR-210-3p were upregulated in whole peripheral blood of children affected with GDM.
Screening based on the combination of microRNAs with a good sensitivity only in the ROC curve analysis was superior over using individual microRNAs for the prediction of potential risk of later development of diabetes mellitus and/or cardiovascular/cerebrovascular diseases, since it was able to identify 88.14% children with an aberrant microRNA expression profile at 10.0% FPR.
Subsequently, we performed analyses to see how the microRNA expression profile differed in relation to the current absence or presence of cardiovascular risk factors and cardiovascular complications (overweight/obesity, prehypertension/hypertension, and/or valve problems and heart defects) and simultaneously to the previous occurrence of maternal pregnancy complication (GDM).
Again, a set of microRNAs associated with diabetes/cardiovascular/cerebrovascular diseases (miR-1-3p, miR-17-5p, miR-20a-5p, miR-20b-5p, miR-21-5p, miR-29a-3p, miR-103a-3p, miR-126-3p, miR-133a-3p, miR-143-3p, miR-181a-5p, miR-195-5p, miR-210-3p, miR-221-3p, and miR-499a-5p) was dysregulated in both groups of children with a prior exposure to GDM regardless of the occurrence of postnatal clinical findings. In addition, seven microRNAs (miR-16-5p, miR-23a-3p, miR-26a-5p, miR-100-5p, miR-125b-5p, miR-146a-5p, and miR-574-3p) were dysregulated in children affected with GDM with normal postnatal clinical findings. An aberrant expression of two additional microRNAs, miR-92a-3p and miR-155-5p, was observed in children with a prior exposure to GDM, who were found to have abnormal clinical findings.
As expected, screening based on the combination of microRNAs with a good sensitivity in the ROC curve analysis only was superior over using individual microRNAs in the assessment of potential risk of later development of diabetes mellitus and cardiovascular/cerebrovascular diseases in groups of children descending from GDM pregnancies with either normal or abnormal clinical findings. At 10.0% FPR, it was able to identify 75.41% and 96.49% children with an aberrant microRNA expression profile.
Subsequently, to assess the impact of cardiovascular risk factors and cardiovascular complications (overweight/obesity, prehypertension/hypertension, and/or valve problems and heart defects) on the microRNA expression profile we further compared the microRNA gene expression profile between equal groups of children descending from pregnancies that had a normal or abnormal course of gestation.
Similarly, as in our previous study [226] dedicated to cardiovascular risk assessment in children descending from pregnancies affected with gestational hypertension, preeclampsia and/or fetal growth restriction, the expression profile of microRNAs was equal between the groups of children descending from uncomplicated pregnancies with normal and abnormal clinical findings, with an exception of miR-21-5p, which showed a trend towards upregulation in a proportion of children with abnormal clinical findings (17.14%).
Parallel, the presence of cardiovascular risk factors (overweight/obesity and/or prehypertension/hypertension) and cardiovascular complications (valve problems and heart defects) had little impact on the microRNA expression profile of children with a prior exposure to GDM, since only miR-29a-3p (12.28%) and miR-92a-3p (12.28%) differentiated between children with normal and abnormal clinical findings (BMI, blood pressure, and echocardiogram findings) with a poor sensitivity.
Previously, we demonstrated that a proportion of children affected with pregnancy-related complications such as gestational hypertension (GH), preeclampsia (PE), and/or fetal growth restriction (FGR) had alterations in the microRNA expression profile that may predispose these children to later development of cardiovascular/cerebrovascular diseases [226,232,233].
Likewise, as a previous occurrence of GH, PE and/or FGR as well as a previous occurrence of GDM was associated with the dysregulation of miR-1-3p, miR-17-5p, miR-20a-5p, miR-20b-5p, miR-21-5p, miR-23a-3p, miR-26a-5p, miR-29a-3p, miR-103a-3p, miR-125b-5p, miR-126-3p, miR-133a-3p, miR-146a-5p, miR-181a-5p, miR-195-5p, miR-210-3p, and miR-342-3p [226,232,233].
On the other hand, our study revealed that dysregulation of miR-16-5p, miR-92a-3p, miR-100-5p, miR-143-3p, miR-155-5p, miR-221-3p, miR-499a-5p, and miR-574-3p represented a unique feature of an aberrant expression profile of children with a prior exposure to GDM. From these findings, it is obvious, that a large proportion of children prenatally exposed to GDM has a substantially altered microRNA expression profile associated with diabetes mellitus and cardiovascular/cerebrovascular diseases.
Generally, epigenetics refers to DNA sequence independent alterations that are responsible for control of transcription. The classic epigenetic modifications include DNA methylation, post-translational modifications of histone proteins, silencing of the extra copy of the X chromosome in women, and genomic imprinting. In addition, proteins and protein complexes with epigenetic modifying capabilities have been involved under the definition of epigenetics. Furthermore, with the discovery of the RNA interference machinery, several classes of noncoding RNAs (microRNAs, small-interfering RNAs, and long noncoding RNAs) have been added to the definition of epigenetics [234]. Nevertheless, controversy still exists as to whether or not microRNAs should be considered as part of the epigenetic program [234,235,236]. While classical epigenetic mechanisms, such as histone modification and DNA methylation, regulate expression at the transcriptional level, microRNAs putatively function mainly at the posttranscriptional level [234,235,236].
Epigenetics mechanisms appear to be interconnected on multiple levels. DNA methylation may direct histone methylation or vice versa chromatin remodelling drives DNA methylation [235,237]. RNAi also appears to be interconnected with DNA methylation and histone modifications. In some species, the link between microRNAs and epigenetics is strong as shown in vitro by transfection of synthetic siRNAs. However, it remains to be seen whether endogenous microRNAs or other types of noncoding RNAs can be also linked to epigenetic mechanisms in vivo [235].
It is also well known that epigenetic modifications are heritable, can be stably transmitted through cell divisions, but can also be reset, since they are very sensitive to the environment and health status [238,239,240,241,242,243,244,245,246,247,248,249]. Recently, the microRNA expression patterns in placental [241,242,244] and germ cells [243,245] have been implicated in fetal programming, and increasing evidence is considering the function of microRNAs in mediating transgenerational epigenetic inheritance [244,246,247,248]. Notably, microRNAs control de novo DNA methylation by regulating transcriptional repressors during germ cell reprogramming [238]. Conversely, global suppression of microRNAs has been observed in mature oocytes and during early embryonic development [239,240]. Consistent with these data, oocytes lack DGCR8 (Pasha), which is necessary for microRNA pathways [239]. It was also demonstrated that many environmental factors contribute to the variations in the epigenome, but diet and early life experiences are key modulators of epigenome, which may initiate the development of the disease [234,249,250,251,252,253,254,255,256,257].
From our findings, it is obvious, that a large proportion of children prenatally exposed to GDM has substantially altered the microRNA expression profile associated with diabetes mellitus and cardiovascular/cerebrovascular diseases, which may be one of several possible reasons for an increased cardiovascular risk. In general, children with a prior exposure to GDM are at higher risk of later development of diabetes mellitus and cardiovascular/cerebrovascular diseases and would benefit from dispensarisation and implementation of primary prevention strategies.
The identification of valve problems and heart defects within the group of children descending from GDM complicated pregnancies more often than usual may support the suggestion for their dispensarisation and implementation of primary prevention strategies.
Similarly, as other studies [258,259,260] we observed that infertility and an infertility treatment may be associated with an increased risk of GDM onset. The incidence of infertility and infertility treatment was significantly higher in groups of mothers with a GDM complicated course of gestation when compared to the group of mothers with normal pregnancies (Table 2). In addition, GDM occurring after assisted reproductive technology conception had been reported to increase the risk of adverse obstetric and perinatal outcomes [259]. This finding may strengthen the idea of dispensarisation of children descending from GDM complicated pregnancies and implementation of primary prevention strategies in this risky group. With regard to a low number of children descending from GDM complicated pregnancies after an infertility treatment (n = 16), it is misleading to interpret the association between microRNA gene expression in whole peripheral venous blood of children and infertility treatment in mothers. Most of the significant results was achieved when the comparison between children descending from normal pregnancies and children descending from GDM complicated pregnancies regardless of infertility treatment was performed or when the comparison between the groups of children descending from normal and GDM complicated pregnancies without infertility treatment was made (83 NP and 102 GDM).
5. Conclusions
In conclusion, any of the tissue-specific and circulation-specific (plasma and/or serum) changes in the microRNA expression profile characteristic for patients with diabetes mellitus and cardiovascular/cerebrovascular diseases are also present in whole peripheral blood (white blood cells) of children previously exposed to GDM. This finding indicates that a previous occurrence of GDM may predispose affected children to later development of diabetes mellitus and cardiovascular/cerebrovascular diseases. Consecutive large-scale studies are needed to verify the findings resulting from this pilot study.
6. Patents
National patent granted—Industrial Property Office, Czech Republic (Patent no. 308102).
International patent filed—Industrial Property Office, Czech Republic (PCT/CZ2019/050050).
Acknowledgments
We thank the staff of the Institute for the Care of Mother and child (Jana Kumprichtova and Sarka Stranska) for assistance with the collection of patients’ biological samples.
Abbreviations
| GDM | Gestational Diabetes Mellitus |
| PE | Preeclampsia |
| FGR | Fetal growth restriction |
| GH | Gestational hypertension |
| AUC | Area under the Curve |
| ROC | Receive Operating Characteristic |
| FPR | False Positive Rate |
| CI | Confidence Interval |
| LR+ | Positive Likelihood Ratio |
| LR- | Negative Likelihood Ratio |
| NP | Normal Pregnancies |
| BP | Blood Pressure |
| OGTT | Oral glucose tolerance test |
| IADPSG | International Association of Diabetes and Pregnancy Study Groups |
| BMI | Body Mass Index |
| SBP | Systolic Blood Pressure |
| DBP | Diastolic Blood Pressure |
| LDL | Low-density Lipoprotein |
| GA | Gestational age |
| CS | Caesarean Section |
| EDTA | Ethylenediaminetetraacetic Acid |
| NTC | No template control |
| NAC | No amplification control |
Supplementary Materials
The following are available online at https://www.mdpi.com/2073-4409/9/6/1557/s1, Figure S1: Aberrant microRNA expression profile in children descending from normal pregnancies, Figure S2: Aberrant microRNA expression profile in children descending from GDM complicated pregnancies, Figure S3: Aberrant microRNA expression profile in children descending from GDM complicated pregnancies irrespective of the clinical findings (overweight/obesity, prehypertension/hypertension, and/or valve problems and heart defects), Figure S4: Aberrant microRNA expression profile in children descending from GDM complicated pregnancies with normal clinical findings., Figure S5: Aberrant microRNA expression profile in children descending from GDM complicated pregnancies with abnormal clinical findings.; Table S1: A list of predicted targets of appropriate microRNAs dysregulated in whole peripheral blood of children descending from GDM complicated pregnancies in relation to obesity using miRWalk2.0 database (putative microRNA binding sites predicted by miRWalk algorithm within mRNA selected regions), Table S2: A list of predicted targets of appropriate microRNAs dysregulated in whole peripheral blood of children descending from GDM complicated pregnancies in relation to hypertension using miRWalk2.0 database (putative microRNA binding sites predicted by miRWalk algorithm within mRNA selected regions); Table S3: A list of predicted targets of appropriate microRNAs dysregulated in whole peripheral blood of children descending from GDM complicated pregnancies in relation to glucose intolerance using miRWalk2.0 database (putative microRNA binding sites predicted by miRWalk algorithm within mRNA selected regions); Table S4: A list of predicted targets of appropriate microRNAs dysregulated in whole peripheral blood of children descending from GDM complicated pregnancies in relation to lipid metabolism disease using miRWalk2.0 database (putative microRNA binding sites predicted by miRWalk algorithm within mRNA selected regions); Table S5: A list of predicted targets of appropriate microRNAs dysregulated in whole peripheral blood of children descending from GDM complicated pregnancies in relation to type 2 diabetes using miRWalk2.0 database (putative microRNA binding sites predicted by miRWalk algorithm within mRNA selected regions); Table S6: A list of predicted targets of appropriate microRNAs dysregulated in whole peripheral blood of children descending from GDM complicated pregnancies in relation to heart septal defect using miRWalk2.0 database (putative microRNA binding sites predicted by miRWalk algorithm within mRNA selected regions); Table S7: A list of predicted targets of appropriate microRNAs dysregulated in whole peripheral blood of children descending from GDM complicated pregnancies in relation to heart valve disease using miRWalk2.0 database (putative microRNA binding sites predicted by miRWalk algorithm within mRNA selected regions); Table S8: A list of predicted targets of appropriate microRNAs dysregulated in whole peripheral blood of children descending from GDM complicated pregnancies in relation to heart disease using miRWalk2.0 database (putative microRNA binding sites predicted by miRWalk algorithm within mRNA selected regions); Table S9: A list of predicted targets of appropriate microRNAs dysregulated in whole peripheral blood of children descending from GDM complicated pregnancies in relation to heart failure using miRWalk2.0 database (putative microRNA binding sites predicted by miRWalk algorithm within mRNA selected regions); Table S10: A list of predicted targets of appropriate microRNAs dysregulated in whole peripheral blood of children descending from GDM complicated pregnancies in relation to venous insufficiency using miRWalk2.0 database (putative microRNA binding sites predicted by miRWalk algorithm within mRNA selected regions); Table S11: A list of predicted targets of appropriate microRNAs dysregulated in whole peripheral blood of children descending from GDM complicated pregnancies in relation to pulmonary embolism using miRWalk2.0 database (putative microRNA binding sites predicted by miRWalk algorithm within mRNA selected regions).
Author Contributions
Conceptualization, I.H. and L.K.; methodology, I.H., K.K., and L.D.; software, I.H., K.K., and J.S.; validation, I.H., L.K., and J.S.; formal analysis, I.H. and K.K.; investigation, K.K., L.D., and J.S.; resources, L.D.; data curation, I.H., K.K., and L.D.; writing—original draft preparation, I.H., K.K., and J.S.; writing—review and editing, I.H. and K.K.; visualization, K.K.; supervision, I.H. and L.K.; project administration, I.H. and L.K.; funding acquisition, I.H. and L.K. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Agency of Medical Research, Ministry of Health, Prague, Czech Republic, grant number AZV 16-27761A and by the Charles University, Prague, Czech Republic, grant number 260529/SVV/2020 and PROGRES Q34. All rights reserved.
Conflicts of Interest
The authors declare no conflict of interest.
References
- 1.American Diabetes Association Gestational diabetes mellitus. Diabetes Care. 2004;27:S88–S90. doi: 10.2337/diacare.27.2007.S88. [DOI] [PubMed] [Google Scholar]
- 2.International Diabetes Federation IDF Diabetes Atlas–Across the Globe. [(accessed on 12 April 2020)];2017 Available online: http://diabetesatlas.org/across-the-globe.html.
- 3.Bowes S.B., Hennessy T.R., Umpleby A.M., Benn J.J., Jackson N.C., Boroujerdi M.A., Sönksen P.H., Lowy C. Measurement of glucose metabolism and insulin secretion during normal pregnancy and pregnancy complicated by gestational diabetes. Diabetologia. 1996;39:976–983. doi: 10.1007/BF00403918. [DOI] [PubMed] [Google Scholar]
- 4.Di Cianni G., Miccoli R., Volpe L., Lencioni C., Del Prato S. Intermediate metabolism in normal pregnancy and in gestational diabetes. Diabetes Metab. Res. Rev. 2003;19:259–270. doi: 10.1002/dmrr.390. [DOI] [PubMed] [Google Scholar]
- 5.Barbour L.A., McCurdy C.E., Hernandez T.L., Kirwan J.P., Catalano P.M., Friedman J.E. Cellular mechanisms for insulin resistance in normal pregnancy and gestational diabetes. Diabetes Care. 2007;30:S112–S119. doi: 10.2337/dc07-s202. Corrected in Diabetes Care2007, 30, 3154. [DOI] [PubMed] [Google Scholar]
- 6.Chen X.Y., Li G.M., Dong Q., Peng H. MiR-577 inhibits pancreatic β-cell function and survival by targeting fibroblast growth factor 21 (FGF-21) in pediatric diabetes. Genet. Mol. Res. 2015;14:15462–15470. doi: 10.4238/2015.November.30.24. [DOI] [PubMed] [Google Scholar]
- 7.Dias S., Pheiffer C., Abrahams Y., Rheeder P., Adam S. Molecular Biomarkers for Gestational Diabetes Mellitus. Int. J. Mol. Sci. 2018;19:2926. doi: 10.3390/ijms19102926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Zhao C., Dong J., Jiang T., Shi Z., Yu B., Zhu Y., Chen D., Xu J., Huo R., Dai J., et al. Early Second-Trimester Serum miRNA Profiling Predicts Gestational Diabetes Mellitus. PLoS ONE. 2011;6:e23925. doi: 10.1371/journal.pone.0023925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zhu Y., Tian F., Li H., Zhou Y., Lu J., Ge Q. Profiling maternal plasma microRNA expression in early pregnancy to predict gestational diabetes mellitus. Int. J. Gynaecol. Obstet. 2015;130:49–53. doi: 10.1016/j.ijgo.2015.01.010. [DOI] [PubMed] [Google Scholar]
- 10.Tryggestad J.B., Vishwanath A., Jiang S., Mallappa A., Teague A.M., Takahashi Y., Thompson D.M., Chernausek S.D. Influence of gestational diabetes mellitus on human umbilical vein endothelial cell miRNA. Clin. Sci. 2016;130:1955–1967. doi: 10.1042/CS20160305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Poirier C., Desgagné V., Guérin R., Bouchard L. MicroRNAs in Pregnancy and Gestational Diabetes Mellitus: Emerging Role in Maternal Metabolic Regulation. Curr. Diabetes Rep. 2017;17:35. doi: 10.1007/s11892-017-0856-5. [DOI] [PubMed] [Google Scholar]
- 12.Wander P.L., Boyko E.J., Hevner K., Parikh V.J., Tadesse M.G., Sorensen T.K., Williams M.A., Enquobahrie D.A. Circulating early- and mid-pregnancy microRNAs and risk of gestational diabetes. Diabetes Res. Clin. Pract. 2017;132:1–9. doi: 10.1016/j.diabres.2017.07.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Cao Y.L., Jia Y.J., Xing B.H., Shi D.D., Dong X.J. Plasma microRNA-16-5p, -17-5p and -20a-5p: Novel diagnostic biomarkers for gestational diabetes mellitus. J. Obstet. Gynaecol. Res. 2017;43:974–981. doi: 10.1111/jog.13317. [DOI] [PubMed] [Google Scholar]
- 14.Sebastiani G., Guarino E., Grieco G.E., Formichi C., Poggi C.D., Ceccarelli E., Dotta F. Circulating microRNA (miRNA) Expression Profiling in Plasma of Patients with Gestational Diabetes Mellitus Reveals Upregulation of miRNA miR-330-3p. Front. Endocrinol. 2017;8:345. doi: 10.3389/fendo.2017.00345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Stirm L., Huypens P., Sass S., Batra R., Fritsche L., Brucker S., Abele H., Hennige A.M., Theis F., Beckers J., et al. Maternal Whole Blood Cell miRNA-340 Is Elevated in Gestational Diabetes and Inversely Regulated by Glucose and Insulin. Sci. Rep. 2018;8:1366. doi: 10.1038/s41598-018-19200-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.He Y., Bai J., Liu P., Dong J., Tang Y., Zhou J., Han P., Xing J., Chen Y., Yu X. miR-494 Protects Pancreatic β-cell Function by Targeting PTEN in Gestational Diabetes Mellitus. EXCLI J. 2017;16:1297–1307. doi: 10.17179/excli2017-491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Pheiffer C., Dias S., Rheeder P., Adam S. Decreased Expression of Circulating miR-20a-5p in South African Women with Gestational Diabetes Mellitus. Mol. Diagn. Ther. 2018;22:345–352. doi: 10.1007/s40291-018-0325-0. [DOI] [PubMed] [Google Scholar]
- 18.Tagoma A., Alnek K., Kirss A., Uibo R., Haller-Kikkatalo K. MicroRNA profiling of second trimester maternal plasma shows upregulation of miR-195-5p in patients with gestational diabetes. Gene. 2018;672:137–142. doi: 10.1016/j.gene.2018.06.004. [DOI] [PubMed] [Google Scholar]
- 19.Lamadrid-Romero M., Solís K.H., Cruz-Reséndiz M.S., Pérez J.E., Díaz N.F., Flores-Herrera H., García-López G., Perichart O., Reyes-Muñoz E., Arenas-Huertero F., et al. Central nervous system development-related microRNAs levels increase in the serum of gestational diabetic women during the first trimester of pregnancy. Neurosci. Res. 2018;130:8–22. doi: 10.1016/j.neures.2017.08.003. [DOI] [PubMed] [Google Scholar]
- 20.Hocaoglu M., Demirer S., Senturk H., Turgut A., Komurcu-Bayrak E. Differential expression of candidate circulating microRNAs in maternal blood leukocytes of the patients with preeclampsia and gestational diabetes mellitus. Pregnancy Hypertens. 2019;17:5–11. doi: 10.1016/j.preghy.2019.04.004. [DOI] [PubMed] [Google Scholar]
- 21.Yoffe L., Polsky A., Gilam A., Raff C., Mecacci F., Ognibene A., Crispi F., Gratacós E., Kanety H., Mazaki-Tovi S., et al. Early Diagnosis of Gestational Diabetes Mellitus Using Circulating microRNAs. Eur. J. Endocrinol. 2019;181:565–577. doi: 10.1530/EJE-19-0206. [DOI] [PubMed] [Google Scholar]
- 22.Kawasaki M., Arata N., Miyazaki C., Mori R., Kikuchi T., Ogawa Y., Ota E. Obesity and abnormal glucose tolerance in offspring of diabetic mothers: A systematic review and meta-analysis. PLoS ONE. 2018;13:e0190676. doi: 10.1371/journal.pone.0190676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Garcia-Vargas L., Addison S.S., Nistala R., Kurukulasuriya D., Sowers J.R. Gestational Diabetes and the Offspring: Implications in the Development of the Cardiorenal Metabolic Syndrome in Offspring. Cardiorenal Med. 2012;2:134–142. doi: 10.1159/000337734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hammoud N.M., Visser G.H.A., van Rossem L., Biesma D.H., Wit J.M., de Valk H.W. Long-term BMI and growth profiles in offspring of women with gestational diabetes. Diabetologia. 2018;61:1037–1045. doi: 10.1007/s00125-018-4584-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Bianco M.E., Josefson J.L. Hyperglycemia During Pregnancy and Long-Term Offspring Outcomes. Curr. Diabetes Rep. 2020;19:143. doi: 10.1007/s11892-019-1267-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Lee H., Jang H.C., Park H.K., Cho N.H. Early manifestation of cardiovascular disease risk factors in offspring of mothers with previous history of gestational diabetes mellitus. Diabetes Res. Clin. Pract. 2007;78:238–245. doi: 10.1016/j.diabres.2007.03.023. [DOI] [PubMed] [Google Scholar]
- 27.Lu J., Zhang S., Li W., Leng J., Wang L., Liu H., Li W., Zhang C., Qi L., Tuomilehto J., et al. Maternal Gestational Diabetes Is Associated with Offspring’s Hypertension. Am. J. Hypertens. 2019;32:335–342. doi: 10.1093/ajh/hpz005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wright C.S., Rifas-Shiman S.L., Rich-Edwards J.W., Taveras E.M., Gillman M.W., Oken E. Intrauterine exposure to gestational diabetes, child adiposity, and blood pressure. Am. J. Hypertens. 2009;22:215–220. doi: 10.1038/ajh.2008.326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Tam W.H., Ma R.C.W., Ozaki R., Li A.M., Chan M.H.M., Yuen L.Y., Lao T.T.H., Yang X., Ho C.S., Tutino G.E., et al. In Utero Exposure to Maternal Hyperglycemia Increases Childhood Cardiometabolic Risk in Offspring. Diabetes Care. 2017;40:679–686. doi: 10.2337/dc16-2397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Lowe W.L., Jr., Scholtens D.M., Kuang A., Linder B., Lawrence J.M., Lebenthal Y., McCance D., Hamilton J., Nodzenski M., Talbot O., et al. HAPO Follow-up Study Cooperative Research Group. Hyperglycemia and Adverse Pregnancy Outcome Follow-up Study (HAPO FUS): Maternal Gestational Diabetes Mellitus and Childhood Glucose Metabolism. Diabetes Care. 2019;42:372–380. doi: 10.2337/dc18-1646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Tsadok M.A., Friedlander Y., Paltiel O., Manor O., Meiner V., Hochner H., Sagy Y., Sharon N., Yazdgerdi S., Siscovick D., et al. Obesity and blood pressure in 17-year-old offspring of mothers with gestational diabetes: Insights from the Jerusalem Perinatal Study. Exp. Diabetes Res. 2011;2011:906154. doi: 10.1155/2011/906154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Perng W., Hockett C.W., Sauder K.A., Dabelea D. In utero exposure to gestational diabetes mellitus and cardiovascular risk factors in youth: A longitudinal analysis in the EPOCH cohort. Pediatr. Obes. 2020:e12611. doi: 10.1111/ijpo.12611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Leybovitz-Haleluya N., Wainstock T., Landau D., Sheiner E. Maternal gestational diabetes mellitus and the risk of subsequent pediatric cardiovascular diseases of the offspring: A population-based cohort study with up to 18 years of follow up. Acta Diabetol. 2018;55:1037–1042. doi: 10.1007/s00592-018-1176-1. [DOI] [PubMed] [Google Scholar]
- 34.Yu Y., Arah O.A., Liew Z., Cnattingius S., Olsen J., Sørensen H.T., Qin G., Li J. Maternal diabetes during pregnancy and early onset of cardiovascular disease in offspring: Population based cohort study with 40 years of follow-up. BMJ. 2019;367:l6398. doi: 10.1136/bmj.l6398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Franks P.W., Looker H.C., Kobes S., Touger L., Tataranni P.A., Hanson R.L., Knowler W.C. Gestational glucose tolerance and risk of type 2 diabetes in young Pima Indian offspring. Diabetes. 2006;55:460–465. doi: 10.2337/diabetes.55.02.06.db05-0823. [DOI] [PubMed] [Google Scholar]
- 36.Clausen T.D., Mathiesen E.R., Hansen T., Pedersen O., Jensen D.M., Lauenborg J., Damm P. High prevalence of type 2 diabetes and pre-diabetes in adult offspring of women with gestational diabetes mellitus or type 1 diabetes: The role of intrauterine hyperglycemia. Diabetes Care. 2008;31:340–346. doi: 10.2337/dc07-1596. [DOI] [PubMed] [Google Scholar]
- 37.Van Dam J.M., Garrett A.J., Schneider L.A., Hodyl N.A., Goldsworthy M.R., Coat S., Rowan J.A., Hague W.M., Pitcher J.B. Reduced Cortical Excitability, Neuroplasticity, and Salivary Cortisol in 11-13-Year-Old Children Born to Women with Gestational Diabetes Mellitus. EBioMedicine. 2018;31:143–149. doi: 10.1016/j.ebiom.2018.04.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Xiang A.H., Wang X., Martinez M.P., Getahun D., Page K.A., Buchanan T.A., Feldman K. Maternal Gestational Diabetes Mellitus, Type 1 Diabetes, and Type 2 Diabetes During Pregnancy and Risk of ADHD in Offspring. Diabetes Care. 2018;41:2502–2508. doi: 10.2337/dc18-0733. [DOI] [PubMed] [Google Scholar]
- 39.Nomura Y., Marks D.J., Grossman B., Yoon M., Loudon H., Stone J., Halperin J.M. Exposure to gestational diabetes mellitus and low socioeconomic status: Effects on neurocognitive development and risk of attention-deficit/hyperactivity disorder in offspring. Arch. Pediatr. Adolesc. Med. 2012;166:337–343. doi: 10.1001/archpediatrics.2011.784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Xiang A.H., Wang X., Martinez M.P., Walthall J.C., Curry E.S., Page K., Buchanan T.A., Coleman K.J., Getahun D. Association of maternal diabetes with autism in offspring. JAMA. 2015;313:1425–1434. doi: 10.1001/jama.2015.2707. [DOI] [PubMed] [Google Scholar]
- 41.Xu G., Jing J., Bowers K., Liu B., Bao W. Maternal diabetes and the risk of autism spectrum disorders in the offspring: A systematic review and meta-analysis. J. Autism Dev. Disord. 2014;44:766–775. doi: 10.1007/s10803-013-1928-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ornoy A., Wolf A., Ratzon N., Greenbaum C., Dulitzky M. Neurodevelopmental outcome at early school age of children born to mothers with gestational diabetes. Arch. Dis. Child. Fetal Neonatal Ed. 1999;81:F10–F14. doi: 10.1136/fn.81.1.F10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Nahum Sacks K., Friger M., Shoham-Vardi I., Abokaf H., Spiegel E., Sergienko R., Landau D., Sheiner E. Prenatal exposure to gestational diabetes mellitus as an independent risk factor for long-term neuropsychiatric morbidity of the offspring. Am. J. Obstet. Gynecol. 2016;215:380e1–380e7. doi: 10.1016/j.ajog.2016.03.030. [DOI] [PubMed] [Google Scholar]
- 44.Walter E., Tsumi E., Wainstock T., Spiegel E., Sheiner E. Maternal gestational diabetes mellitus: Is it associated with long-term pediatric ophthalmic morbidity of the offspring? J. Maternal Fetal Neonatal Med. 2019;32:2529–2538. doi: 10.1080/14767058.2018.1439918. [DOI] [PubMed] [Google Scholar]
- 45.Martinez M.P., Lin J., Chow T., Chung J., Wang X., Xiang A.H. Maternal Gestational Diabetes and Type 2 Diabetes During Pregnancy and Risk of Childhood Asthma in Offspring. J. Pediatr. 2020;219:173–179. doi: 10.1016/j.jpeds.2019.12.053. [DOI] [PubMed] [Google Scholar]
- 46.Li J., Dong X., Wang Z., Wu J. MicroRNA-1 in Cardiac Diseases and Cancers. Korean J. Physiol. Pharmacol. 2014;18:359–363. doi: 10.4196/kjpp.2014.18.5.359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Li Y.Q., Zhang M.F., Wen H.Y., Hu C.L., Liu R., Wei H.Y., Ai C.M., Wang G., Liao X.X., Li X. Comparing the diagnostic values of circulating microRNAs and cardiac troponin T in patients with acute myocardial infarction. Clinics. 2013;68:75–80. doi: 10.6061/clinics/2013(01)OA12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Gasiulė S., Stankevičius V., Patamsytė V., Ražanskas R., Zukovas G., Kapustina Z., Zaliaduonytė D., Benetis R., Lesauskaitė V., Vilkaitis G. Tissue-Specific miRNAs Regulate the Development of Thoracic Aortic Aneurysm: The Emerging Role of KLF4 Network. J. Clin. Med. 2019;8:1609. doi: 10.3390/jcm8101609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Gerlinger-Romero F., Yonamine C.Y., Junior D.C., Esteves J.V., Machado U.F. Dysregulation between TRIM63/FBXO32 expression and soleus muscle wasting in diabetic rats: Potential role of miR-1-3p, -29a/b-3p, and -133a/b-3p. Mol. Cell Biochem. 2017;427:187–199. doi: 10.1007/s11010-016-2910-z. [DOI] [PubMed] [Google Scholar]
- 50.Kokkinopoulou I., Maratou E., Mitrou P., Boutati E., Sideris D.C., Fragoulis E.G., Christodoulou M.I. Decreased expression of microRNAs targeting type-2 diabetes susceptibility genes in peripheral blood of patients and predisposed individuals. Endocrine. 2019;66:226–239. doi: 10.1007/s12020-019-02062-0. [DOI] [PubMed] [Google Scholar]
- 51.Hromadnikova I., Kotlabova K., Dvorakova L., Krofta L. Evaluation of Vascular Endothelial Function in Young and Middle-Aged Women with Respect to a History of Pregnancy, Pregnancy-Related Complications, Classical Cardiovascular Risk Factors, and Epigenetics. Int. J. Mol. Sci. 2020;21:430. doi: 10.3390/ijms21020430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Wang X., Shang Y., Dai S., Wu W., Yi F., Cheng L. MicroRNA-16-5p aggravates myocardial infarction injury by targeting expression of insulin receptor substrates 1 and mediating myocardial apoptosis and angiogenesis. Curr. Neurovascular Res. 2020;17:11–17. doi: 10.2174/1567202617666191223142743. [DOI] [PubMed] [Google Scholar]
- 53.O’Sullivan J.F., Neylon A., McGorrian C., Blake G.J. miRNA-93-5p and other miRNAs as predictors of coronary artery disease and STEMI. Int. J. Cardiol. 2016;224:310–316. doi: 10.1016/j.ijcard.2016.09.016. [DOI] [PubMed] [Google Scholar]
- 54.Vegter E.L., Schmitter D., Hagemeijer Y., Ovchinnikova E.S., van der Harst P., Teerlink J.R., O’Connor C.M., Metra M., Davison B.A., Bloomfield D., et al. Use of biomarkers to establish potential role and function of circulating microRNAs in acute heart failure. Int. J. Cardiol. 2016;224:231–239. doi: 10.1016/j.ijcard.2016.09.010. [DOI] [PubMed] [Google Scholar]
- 55.Gacoń J., Badacz R., Stępień E., Karch I., Enguita F.J., Żmudka K., Przewłocki T., Kabłak-Ziembicka A. Diagnostic and prognostic micro-RNAs in ischaemic stroke due to carotid artery stenosis and in acute coronary syndrome: A four-year prospective study. Kardiol. Polska. 2018;76:362–369. doi: 10.5603/KP.a2017.0243. [DOI] [PubMed] [Google Scholar]
- 56.Duan Y.R., Chen B.P., Chen F., Yang S.X., Zhu C.Y., Ma Y.L., Li Y., Shi J. Exosomal microRNA-16-5p from human urine-derived stem cells ameliorates diabetic nephropathy through protection of podocyte. J. Cell Mol. Med. 2019 doi: 10.1111/jcmm.14558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Assmann T.S., Recamonde-Mendoza M., Costa A.R., Puñales M., Tschiedel B., Canani L.H., Bauer A.C., Crispim D. Circulating miRNAs in diabetic kidney disease: Case-control study and in silico analyses. Acta Diabetol. 2019;56:55–65. doi: 10.1007/s00592-018-1216-x. [DOI] [PubMed] [Google Scholar]
- 58.Alicka M., Major P., Wysocki M., Marycz K. Adipose-Derived Mesenchymal Stem Cells Isolated from Patients with Type 2 Diabetes Show Reduced “Stemness” through an Altered Secretome Profile, Impaired Anti-Oxidative Protection, and Mitochondrial Dynamics Deterioration. J. Clin. Med. 2019;8:765. doi: 10.3390/jcm8060765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Mogilyansky E., Rigoutsos I. The miR-17/92 cluster: A comprehensive update on its genomics, genetics, functions and increasingly important and numerous roles in health and disease. Cell Death Differ. 2013;20:1603–1614. doi: 10.1038/cdd.2013.125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Zhou L., Qi R.Q., Liu M., Xu Y.P., Li G., Weiland M., Kaplan D.H., Mi Q.S. microRNA miR-17-92 cluster is highly expressed in epidermal Langerhans cells but not required for its development. Genes Immun. 2014;15:57–61. doi: 10.1038/gene.2013.61. [DOI] [PubMed] [Google Scholar]
- 61.Danielson L.S., Park D.S., Rotllan N., Chamorro-Jorganes A., Guijarro M.V., Fernandez-Hernando C., Fishman G.I., Phoon C.K., Hernando E. Cardiovascular dysregulation of miR-17-92 causes a lethal hypertrophic cardiomyopathy and arrhythmogenesis. FASEB J. 2013;27:1460–1467. doi: 10.1096/fj.12-221994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Du W., Pan Z., Chen X., Wang L., Zhang Y., Li S., Liang H., Xu C., Zhang Y., Wu Y., et al. By targeting Stat3 microRNA-17-5p promotes cardiomyocyte apoptosis in response to ischemia followed by reperfusion. Cell Physiol. Biochem. 2014;34:955–965. doi: 10.1159/000366312. [DOI] [PubMed] [Google Scholar]
- 63.Kaucsár T., Révész C., Godó M., Krenács T., Albert M., Szalay C.I., Rosivall L., Benyó Z., Bátkai S., Thum T., et al. Activation of the miR-17 family and miR-21 during murine kidney ischemia-reperfusion injury. Nucleic Acid Ther. 2013;23:344–354. doi: 10.1089/nat.2013.0438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Fang L., Ellims A.H., Moore X.L., White D.A., Taylor A.J., Chin-Dusting J., Dart A.M. Circulating microRNAs as biomarkers for diffuse myocardial fibrosis in patients with hypertrophic cardiomyopathy. J. Transl. Med. 2015;13:314. doi: 10.1186/s12967-015-0672-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Wu J., Du K., Lu X. Elevated expressions of serum miR-15a, miR-16, and miR-17-5p are associated with acute ischemic stroke. Int. J. Clin. Exp. Med. 2015;8:21071–21079. [PMC free article] [PubMed] [Google Scholar]
- 66.Chen J., Xu L., Hu Q., Yang S., Zhang B., Jiang H. MiR-17-5p as circulating biomarkers for the severity of coronary atherosclerosis in coronary artery disease. Int. J. Cardiol. 2015;197:123–124. doi: 10.1016/j.ijcard.2015.06.037. [DOI] [PubMed] [Google Scholar]
- 67.Tian L., Song Z., Shao W., Du W.W., Zhao L.R., Zeng K., Yang B.B., Jin T. Curcumin represses mouse 3T3-L1 cell adipogenic differentiation via inhibiting miR-17-5p and stimulating the Wnt signalling pathway effector Tcf7l2. Cell Death Dis. 2017;8:e2559. doi: 10.1038/cddis.2016.455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Chen T.C., Sung M.L., Kuo H.C., Chien S.J., Yen C.K., Chen C.N. Differential regulation of human aortic smooth muscle cell proliferation by monocyte-derived macrophages from diabetic patients. PLoS ONE. 2014;9:e113752. doi: 10.1371/journal.pone.0113752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Mendell J.T. miRiad roles for the miR-17-92 cluster in development and disease. Cell. 2008;133:217–222. doi: 10.1016/j.cell.2008.04.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Brock M., Samillan V.J., Trenkmann M., Schwarzwald C., Ulrich S., Gay R.E., Gassmann M., Ostergaard L., Gay S., Speich R., et al. AntagomiR directed against miR-20a restores functional BMPR2 signalling and prevents vascular remodelling in hypoxia-induced pulmonary hypertension. Eur. Heart J. 2014;35:3203–3211. doi: 10.1093/eurheartj/ehs060. [DOI] [PubMed] [Google Scholar]
- 71.Platania C.B.M., Maisto R., Trotta M.C., D’Amico M., Rossi S., Gesualdo C., D’Amico G., Balta C., Herman H., Hermenean A., et al. Retinal and circulating miRNA expression patterns in diabetic retinopathy: An in silico and in vivo approach. Br. J. Pharmacol. 2019;176:2179–2194. doi: 10.1111/bph.14665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Lareyre F., Clément M., Moratal C., Loyer X., Jean-Baptiste E., Hassen-Khodja R., Chinetti G., Mallat Z., Raffort J. Differential micro-RNA expression in diabetic patients with abdominal aortic aneurysm. Biochimie. 2019;162:1–7. doi: 10.1016/j.biochi.2019.03.012. [DOI] [PubMed] [Google Scholar]
- 73.Dickinson B.A., Semus H.M., Montgomery R.L., Stack C., Latimer P.A., Lewton S.M., Lynch J.M., Hullinger T.G., Seto A.G., van Rooij E. Plasma microRNAs serve as biomarkers of therapeutic efficacy and disease progression in hypertension-induced heart failure. Eur. J. Heart Fail. 2013;15:650–659. doi: 10.1093/eurjhf/hft018. [DOI] [PubMed] [Google Scholar]
- 74.Flowers E., Aouizerat B.E., Abbasi F., Lamendola C., Grove K.M., Fukuoka Y., Reaven G.M. Circulating microRNA-320a and microRNA-486 predict thiazolidinedione response: Moving towards precision health for diabetes prevention. Metabolism. 2015;64:1051–1059. doi: 10.1016/j.metabol.2015.05.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Katayama M., Wiklander O.P.B., Fritz T., Caidahl K., El-Andaloussi S., Zierath J.R., Krook A. Circulating Exosomal miR-20b-5p Is Elevated in Type 2 Diabetes and Could Impair Insulin Action in Human Skeletal Muscle. Diabetes. 2019;68:515–526. doi: 10.2337/db18-0470. [DOI] [PubMed] [Google Scholar]
- 76.Xiong Y., Chen L., Yan C., Zhou W., Endo Y., Liu J., Hu L., Hu Y., Mi B., Liu G. Circulating Exosomal miR-20b-5p Inhibition Restores Wnt9b Signaling and Reverses Diabetes-Associated Impaired Wound Healing. Small. 2020;16:e1904044. doi: 10.1002/smll.201904044. [DOI] [PubMed] [Google Scholar]
- 77.Zhu K., Hu X., Chen H., Li F., Yin N., Liu A.L., Shan K., Qin Y.W., Huang X., Chang Q., et al. Downregulation of circRNA DMNT3B contributes to diabetic retinal vascular dysfunction through targeting miR-20b-5p and BAMBI. EBioMedicine. 2019;49:341–353. doi: 10.1016/j.ebiom.2019.10.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Sekar D., Venugopal B., Sekar P., Ramalingam K. Role of microRNA 21 in diabetes and associated/related diseases. Gene. 2016;582:14–18. doi: 10.1016/j.gene.2016.01.039. [DOI] [PubMed] [Google Scholar]
- 79.Suárez Y., Fernández-Hernando C., Pober J.S., Sessa W.C. Dicer dependent microRNAs regulate gene expression and functions in human endothelial cells. Circ. Res. 2007;100:1164–1173. doi: 10.1161/01.RES.0000265065.26744.17. [DOI] [PubMed] [Google Scholar]
- 80.Dong S., Ma W., Hao B., Hu F., Yan L., Yan X., Wang Y., Chen Z., Wang Z. microRNA-21 promotes cardiac fibrosis and development of heart failure with preserved left ventricular ejection fraction by up-regulating Bcl-2. Int. J. Clin. Exp. Pathol. 2014;7:565–574. [PMC free article] [PubMed] [Google Scholar]
- 81.Zhang J., Xing Q., Zhou X., Li J., Li Y., Zhang L., Zhou Q., Tang B. Circulating miRNA 21 is a promising biomarker for heart failure. Mol. Med. Rep. 2017;16:7766–7774. doi: 10.3892/mmr.2017.7575. [DOI] [PubMed] [Google Scholar]
- 82.Licholai S., Blaż M., Kapelak B., Sanak M. Unbiased Profile of MicroRNA Expression in Ascending Aortic Aneurysm Tissue Appoints Molecular Pathways Contributing to the Pathology. Ann. Thorac. Surg. 2016;102:1245–1252. doi: 10.1016/j.athoracsur.2016.03.061. [DOI] [PubMed] [Google Scholar]
- 83.Kriegel A.J., Baker M.A., Liu Y., Liu P., Cowley A.W., Jr., Liang M. Endogenous microRNAs in human microvascular endothelial cells regulate mRNAs encoded by hypertension-related genes. Hypertension. 2015;66:793–799. doi: 10.1161/HYPERTENSIONAHA.115.05645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Velle-Forbord T., Eidlaug M., Debik J., Sæther J.C., Follestad T., Nauman J., Gigante B., Røsjø H., Omland T., Langaas M., et al. Circulating microRNAs as predictive biomarkers of myocardial infarction: Evidence from the HUNT study. Atherosclerosis. 2019;289:1–7. doi: 10.1016/j.atherosclerosis.2019.07.024. [DOI] [PubMed] [Google Scholar]
- 85.Demirsoy İ.H., Ertural D.Y., Balci Ş., Çınkır Ü., Sezer K., Tamer L., Aras N. Profiles of Circulating MiRNAs Following Metformin Treatment in Patients with Type 2 Diabetes. J. Med. Biochem. 2018;37:499–506. doi: 10.2478/jomb-2018-0009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Olivieri F., Spazzafumo L., Bonafè M., Recchioni R., Prattichizzo F., Marcheselli F., Micolucci L., Mensà E., Giuliani A., Santini G., et al. MiR-21-5p and miR-126a-3p levels in plasma and circulating angiogenic cells: Relationship with type 2 diabetes complications. Oncotarget. 2015;6:35372–35382. doi: 10.18632/oncotarget.6164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Assmann T.S., Recamonde-Mendoza M., De Souza B.M., Crispim D. MicroRNA expression profiles and type 1 diabetes mellitus: Systematic review and bioinformatic analysis. Endocr. Connect. 2017;6:773–790. doi: 10.1530/EC-17-0248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Lakhter A.J., Pratt R.E., Moore R.E., Doucette K.K., Maier B.F., DiMeglio L.A., Sims E.K. Beta cell extracellular vesicle miR-21-5p cargo is increased in response to inflammatory cytokines and serves as a biomarker of type 1 diabetes. Diabetologia. 2018;61:1124–1134. doi: 10.1007/s00125-018-4559-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Grieco G.E., Cataldo D., Ceccarelli E., Nigi L., Catalano G., Brusco N., Mancarella F., Ventriglia G., Fondelli C., Guarino E., et al. Serum Levels of miR-148a and miR-21-5p Are Increased in Type 1 Diabetic Patients and Correlated with Markers of Bone Strength and Metabolism. Noncoding RNA. 2018;4:37. doi: 10.3390/ncrna4040037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Gholaminejad A., Abdul Tehrani H., Gholami Fesharaki M. Identification of candidate microRNA biomarkers in diabetic nephropathy: A meta-analysis of profiling studies. J. Nephrol. 2018;31:813–831. doi: 10.1007/s40620-018-0511-5. [DOI] [PubMed] [Google Scholar]
- 91.Long B., Gan T.Y., Zhang R.C., Zhang Y.H. miR-23a Regulates Cardiomyocyte Apoptosis by Targeting Manganese Superoxide Dismutase. Mol. Cells. 2017;40:542–549. doi: 10.14348/molcells.2017.0012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Wang S., He W., Wang C. MiR-23a Regulates the Vasculogenesis of Coronary Artery Disease by Targeting Epidermal Growth Factor Receptor. Cardiovasc. Ther. 2016;34:199–208. doi: 10.1111/1755-5922.12187. [DOI] [PubMed] [Google Scholar]
- 93.Cong X., Li Y., Lu N., Dai Y., Zhang H., Zhao X., Liu Y. Resveratrol attenuates the inflammatory reaction induced by ischemia/reperfusion in the rat heart. Mol. Med. Rep. 2014;9:2528–2532. doi: 10.3892/mmr.2014.2090. [DOI] [PubMed] [Google Scholar]
- 94.Černá V., Ostašov P., Pitule P., Moláček J., Třeška V., Pešta M. The Expression Profile of MicroRNAs in Small and Large Abdominal Aortic Aneurysms. Cardiol. Res. Pract. 2019;2019:8645840. doi: 10.1155/2019/8645840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Lozano-Bartolomé J., Llauradó G., Portero-Otin M., Altuna-Coy A., Rojo-Martínez G., Vendrell J., Jorba R., Rodríguez-Gallego E., Chacón M.R. Altered Expression of miR-181a-5p and miR-23a-3p Is Associated with Obesity and TNFα-Induced Insulin Resistance. J. Clin. Endocrinol. Metab. 2018;103:1447–1458. doi: 10.1210/jc.2017-01909. [DOI] [PubMed] [Google Scholar]
- 96.Dolz S., Górriz D., Tembl J.I., Sánchez D., Fortea G., Parkhutik V., Lago A. Circulating MicroRNAs as Novel Biomarkers of Stenosis Progression in Asymptomatic Carotid Stenosis. Stroke. 2017;48:10–16. doi: 10.1161/STROKEAHA.116.013650. [DOI] [PubMed] [Google Scholar]
- 97.De Gonzalo-Calvo D., Cenarro A., Garlaschelli K., Pellegatta F., Vilades D., Nasarre L., Camino-Lopez S., Crespo J., Carreras F., Leta R., et al. Translating the microRNA signature of microvesicles derived from human coronary artery smooth muscle cells in patients with familial hypercholesterolemia and coronary artery disease. J. Mol. Cell Cardiol. 2017;106:55–67. doi: 10.1016/j.yjmcc.2017.03.005. [DOI] [PubMed] [Google Scholar]
- 98.Gecys D., Tatarunas V., Veikutiene A., Lesauskaite V. New potential modulators of CYP4F2 enzyme activity in angina pectoris: Hsa-miR-24-3p and hsa-miR-34a-5p. Biomarkers. 2020;25:40–47. doi: 10.1080/1354750X.2019.1690580. [DOI] [PubMed] [Google Scholar]
- 99.Onrat S.T., Onrat E., Ercan Onay E., Yalım Z., Avşar A. The Genetic Determination of the Differentiation between Ischemic Dilated Cardiomyopathy and Idiopathic Dilated Cardiomyopathy. Genet. Test. Mol. Biomark. 2018;22:644–651. doi: 10.1089/gtmb.2018.0188. [DOI] [PubMed] [Google Scholar]
- 100.Tan H., Qi J., Fan B.Y., Zhang J., Su F.F., Wang H.T. MicroRNA-24-3p Attenuates Myocardial Ischemia/Reperfusion Injury by Suppressing RIPK1 Expression in Mice. Cell Physiol. Biochem. 2018;51:46–62. doi: 10.1159/000495161. [DOI] [PubMed] [Google Scholar]
- 101.Xiao X., Lu Z., Lin V., May A., Shaw D.H., Wang Z., Che B., Tran K., Du H., Shaw P.X. MicroRNA miR-24-3p Reduces Apoptosis and Regulates Keap1-Nrf2 Pathway in Mouse Cardiomyocytes Responding to Ischemia/Reperfusion Injury. Oxid. Med. Cell Longev. 2018;2018:7042105. doi: 10.1155/2018/7042105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Gao J., Liu Q.G. The role of miR-26 in tumors and normal tissues. Oncol. Lett. 2011;2:1019–1023. doi: 10.3892/ol.2011.413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Zheng L., Lin S., Lv C. MiR-26a-5p regulates cardiac fibroblasts collagen expression by targeting ULK1. Sci. Rep. 2018;8:2104. doi: 10.1038/s41598-018-20561-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Bye A., Røsjø H., Nauman J., Silva G.J., Follestad T., Omland T., Wisløff U. Circulating microRNAs predict future fatal myocardial infarction in healthy individuals - The HUNT study. J. Mol. Cell Cardiol. 2016;97:162–168. doi: 10.1016/j.yjmcc.2016.05.009. [DOI] [PubMed] [Google Scholar]
- 105.Hsu A., Chen S.J., Chang Y.S., Chen H.C., Chu P.H. Systemic approach to identify serum microRNAs as potential biomarkers for acute myocardial infarction. Biomed. Res. Int. 2014;2014:418628. doi: 10.1155/2014/418628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Xing X., Guo S., Zhang G., Liu Y., Bi S., Wang X., Lu Q. miR-26a-5p protects against myocardial ischemia/reperfusion injury by regulating the PTEN/PI3K/AKT signaling pathway. Braz. J. Med. Biol. Res. 2020;53:e9106. doi: 10.1590/1414-431x20199106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Chouvarine P., Geldner J., Giagnorio R., Legchenko E., Bertram H., Hansmann G. Trans-Right-Ventricle and Transpulmonary MicroRNA Gradients in Human Pulmonary Arterial Hypertension. Pediatr. Crit. Care Med. 2020;21:340–349. doi: 10.1097/PCC.0000000000002207. [DOI] [PubMed] [Google Scholar]
- 108.Garavelli S., Bruzzaniti S., Tagliabue E., Prattichizzo F., Di Silvestre D., Perna F., La Sala L., Ceriello A., Mozzillo E., Fattorusso V., et al. Blood Co-Circulating Extracellular microRNAs and Immune Cell Subsets Associate with Type 1 Diabetes Severity. Int. J. Mol. Sci. 2020;21:477. doi: 10.3390/ijms21020477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Ye Y., Hu Z., Lin Y., Zhang C., Perez-Polo J.R. Downregulation of microRNA-29 by antisense inhibitors and a PPAR-gamma agonist protects against myocardial ischaemia-reperfusion injury. Cardiovasc. Res. 2010;87:535–544. doi: 10.1093/cvr/cvq053. [DOI] [PubMed] [Google Scholar]
- 110.Moraes L.N., Fernandez G.J., Vechetti-Júnior I.J., Freire P.P., Souza R.W.A., Villacis R.A.R., Rogatto S.R., Reis P.P., Dal-Pai-Silva M., Carvalho R.F. Integration of miRNA and mRNA expression profiles reveals microRNA-regulated networks during muscle wasting in cardiac cachexia. Sci. Rep. 2017;7:6998. doi: 10.1038/s41598-017-07236-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Zhao Y., Yuan Y., Qiu C. Underexpression of CACNA1C Caused by Overexpression of microRNA-29a Underlies the Pathogenesis of Atrial Fibrillation. Med. Sci. Monit. 2016;22:2175–2181. doi: 10.12659/MSM.896191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Zhang L., Zhang Y., Xue S., Ding H., Wang Y., Qi H., Wang Y., Zhu W., Li P. Clinical significance of circulating microRNAs as diagnostic biomarkers for coronary artery disease. J. Cell Mol. Med. 2020;24:1146–1150. doi: 10.1111/jcmm.14802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Kong L., Zhu J., Han W., Jiang X., Xu M., Zhao Y., Dong Q., Pang Z., Guan Q., Gao L., et al. Significance of serum microRNAs in pre-diabetes and newly diagnosed type 2 diabetes: A clinical study. Acta Diabetol. 2011;48:61–69. doi: 10.1007/s00592-010-0226-0. [DOI] [PubMed] [Google Scholar]
- 114.Widlansky M.E., Jensen D.M., Wang J., Liu Y., Geurts A.M., Kriegel A.J., Liu P., Ying R., Zhang G., Casati M., et al. miR-29 contributes to normal endothelial function and can restore it in cardiometabolic disorders. EMBO Mol. Med. 2018;10:e8046. doi: 10.15252/emmm.201708046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Bulent Vatan M., Kalaycı Yigin A., Akdemir R., Tarik Agac M., Akif Cakar M., Aksoy M., Tatli E., Kilic H., Gunduz H., Guzel D., et al. Altered Plasma MicroRNA Expression in Patients with Mitral Chordae Tendineae Rupture. J. Heart Valve Dis. 2016;25:580–588. [PubMed] [Google Scholar]
- 116.Gumus G., Giray D., Bobusoglu O., Tamer L., Karpuz D., Hallioglu O. MicroRNA values in children with rheumatic carditis: A preliminary study. Rheumatol. Int. 2018;38:1199–1205. doi: 10.1007/s00296-018-4069-2. [DOI] [PubMed] [Google Scholar]
- 117.Rogg E.M., Abplanalp W.T., Bischof C., John D., Schulz M.H., Krishnan J., Fischer A., Poluzzi C., Schaefer L., Bonauer A., et al. Analysis of Cell Type-Specific Effects of MicroRNA-92a Provides Novel Insights into Target Regulation and Mechanism of Action. Circulation. 2018;138:2545–2558. doi: 10.1161/CIRCULATIONAHA.118.034598. [DOI] [PubMed] [Google Scholar]
- 118.Marques F.Z., Vizi D., Khammy O., Mariani J.A., Kaye D.M. The transcardiac gradient of cardio-microRNAs in the failing heart. Eur. J. Heart Fail. 2016;18:1000–1008. doi: 10.1002/ejhf.517. [DOI] [PubMed] [Google Scholar]
- 119.Liu Y., Li Q., Hosen M.R., Zietzer A., Flender A., Levermann P., Schmitz T., Frühwald D., Goody P., Nickenig G., et al. Atherosclerotic Conditions Promote the Packaging of Functional MicroRNA-92a-3p Into Endothelial Microvesicles. Circ. Res. 2019;124:575–587. doi: 10.1161/CIRCRESAHA.118.314010. [DOI] [PubMed] [Google Scholar]
- 120.Wiese C.B., Zhong J., Xu Z.Q., Zhang Y., Ramirez Solano M.A., Zhu W., Linton M.F., Sheng Q., Kon V., Vickers K.C. Dual inhibition of endothelial miR-92a-3p and miR-489-3p reduces renal injury-associated atherosclerosis. Atherosclerosis. 2019;282:121–131. doi: 10.1016/j.atherosclerosis.2019.01.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Moncini S., Salvi A., Zuccotti P., Viero G., Quattrone A., Barlati S., De Petro G., Venturin M., Riva P. The role of miR-103 and miR-107 in regulation of CDK5R1 expression and in cellular migration. PLoS ONE. 2011;6:e20038. doi: 10.1371/journal.pone.0020038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Huang L., Li L., Chen X., Zhang H., Shi Z. MiR-103a targeting Piezo1 is involved in acute myocardial infarction through regulating endothelium function. Cardiol. J. 2016;23:556–562. doi: 10.5603/CJ.a2016.0056. [DOI] [PubMed] [Google Scholar]
- 123.Deng B., Du J., Hu R., Wang A.P., Wu W.H., Hu C.P., Li Y.J., Li X.H. MicroRNA-103/107 is involved in hypoxia-induced proliferation of pulmonary arterial smooth muscle cells by targeting HIF-1β. Life Sci. 2016;147:117–124. doi: 10.1016/j.lfs.2016.01.043. [DOI] [PubMed] [Google Scholar]
- 124.Trajkovski M., Hausser J., Soutschek J., Bhat B., Akin A., Zavolan M., Heim M.H., Stoffel M. MicroRNAs 103 and 107 regulate insulin sensitivity. Nature. 2011;474:649–653. doi: 10.1038/nature10112. [DOI] [PubMed] [Google Scholar]
- 125.Assmann T.S., Recamonde-Mendoza M., Puñales M., Tschiedel B., Canani L.H., Crispim D. MicroRNA expression profile in plasma from type 1 diabetic patients: Case-control study and bioinformatic analysis. Diabetes Res. Clin. Pract. 2018;141:35–46. doi: 10.1016/j.diabres.2018.03.044. [DOI] [PubMed] [Google Scholar]
- 126.Shaham L., Binder V., Gefen N., Borkhardt A., Izraeli S. MiR-125 in normal and malignant hematopoiesis. Leukemia. 2012;26:2011–2018. doi: 10.1038/leu.2012.90. [DOI] [PubMed] [Google Scholar]
- 127.Tiedt S., Prestel M., Malik R., Schieferdecker N., Duering M., Kautzky V., Stoycheva I., Böck J., Northoff B.H., Klein M., et al. RNA-Seq Identifies Circulating miR-125a-5p, miR-125b-5p, and miR-143-3p as Potential Biomarkers for Acute Ischemic Stroke. Circ. Res. 2017;121:970–980. doi: 10.1161/CIRCRESAHA.117.311572. [DOI] [PubMed] [Google Scholar]
- 128.Jia K., Shi P., Han X., Chen T., Tang H., Wang J. Diagnostic value of miR-30d-5p and miR-125b-5p in acute myocardial infarction. Mol. Med. Rep. 2016;14:184–194. doi: 10.3892/mmr.2016.5246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Bayoumi A.S., Park K.M., Wang Y., Teoh J.P., Aonuma T., Tang Y., Su H., Weintraub N.L., Kim I.M. A carvedilol-responsive microRNA, miR-125b-5p protects the heart from acute myocardial infarction by repressing pro-apoptotic bak1 and klf13 in cardiomyocytes. J. Mol. Cell Cardiol. 2018;114:72–82. doi: 10.1016/j.yjmcc.2017.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Satake E., Pezzolesi M.G., Md Dom Z.I., Smiles A.M., Niewczas M.A., Krolewski A.S. Circulating miRNA Profiles Associated with Hyperglycemia in Patients with Type 1 Diabetes. Diabetes. 2018;67:1013–1023. doi: 10.2337/db17-1207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Samandari N., Mirza A.H., Kaur S., Hougaard P., Nielsen L.B., Fredheim S., Mortensen H.B., Pociot F. Influence of Disease Duration on Circulating Levels of miRNAs in Children and Adolescents with New Onset Type 1 Diabetes. Noncoding RNA. 2018;4:35. doi: 10.3390/ncrna4040035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Yu C.Y., Yang C.Y., Rui Z.L. MicroRNA-125b-5p improves pancreatic β-cell function through inhibiting JNK signaling pathway by targeting DACT1 in mice with type 2 diabetes mellitus. Life Sci. 2019;224:67–75. doi: 10.1016/j.lfs.2019.01.031. [DOI] [PubMed] [Google Scholar]
- 133.Wu X.J., Zhao Z.F., Kang X.J., Wang H.J., Zhao J., Pu X.M. MicroRNA-126-3p suppresses cell proliferation by targeting PIK3R2 in Kaposi’s sarcoma cells. Oncotarget. 2016;7:36614–36621. doi: 10.18632/oncotarget.9311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Matsha T.E., Kengne A.P., Hector S., Mbu D.L., Yako Y.Y., Erasmus R.T. MicroRNA profiling and their pathways in South African individuals with prediabetes and newly diagnosed type 2 diabetes mellitus. Oncotarget. 2018;9:30485–30498. doi: 10.18632/oncotarget.25271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Lan X., Wu L., Wu N., Chen Q., Li Y., Du X., Wei C., Feng L., Li Y., Osoro E.K., et al. Long Noncoding RNA lnc-HC Regulates PPARγ-Mediated Hepatic Lipid Metabolism through miR-130b-3p. Mol. Ther. Nucleic Acids. 2019;18:954–965. doi: 10.1016/j.omtn.2019.10.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Zhang J., Jazii F.R., Haghighi M.M., Alvares D., Liu L., Khosraviani N., Adeli K. miR-130b is a potent stimulator of hepatic very-low-density lipoprotein assembly and secretion via marked induction of microsomal triglyceride transfer protein. Am. J. Physiol. Endocrinol. Metab. 2020;318:E262–E275. doi: 10.1152/ajpendo.00276.2019. [DOI] [PubMed] [Google Scholar]
- 137.Li P., Zhang Q., Wu X., Yang X., Zhang Y., Li Y., Jiang F. Circulating microRNAs serve as novel biological markers for intracranial aneurysms. J. Am. Heart Assoc. 2014;3:e000972. doi: 10.1161/JAHA.114.000972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Tian C., Li Z., Yang Z., Huang Q., Liu J., Hong B. Plasma MicroRNA-16 Is a Biomarker for Diagnosis, Stratification, and Prognosis of Hyperacute Cerebral Infarction. PLoS ONE. 2016;11:e0166688. doi: 10.1371/journal.pone.0166688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Prabu P., Rome S., Sathishkumar C., Aravind S., Mahalingam B., Shanthirani C.S., Gastebois C., Villard A., Mohan V., Balasubramanyam M. Circulating MiRNAs of ‘Asian Indian Phenotype’ Identified in Subjects with Impaired Glucose Tolerance and Patients with Type 2 Diabetes. PLoS ONE. 2015;10:e0128372. doi: 10.1371/journal.pone.0128372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Feng T., Li K., Zheng P., Wang Y., Lv Y., Shen L., Chen Y., Xue Z., Li B., Jin L., et al. Weighted Gene Coexpression Network Analysis Identified MicroRNA Coexpression Modules and Related Pathways in Type 2 Diabetes Mellitus. Oxid. Med. Cell Longev. 2019;2019:9567641. doi: 10.1155/2019/9567641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Liang H.W., Yang X., Wen D.Y., Gao L., Zhang X.Y., Ye Z.H., Luo J., Li Z.Y., He Y., Pang Y.Y., et al. Utility of miR 133a 3p as a diagnostic indicator for hepatocellular carcinoma: An investigation combined with GEO, TCGA, meta analysis and bioinformatics. Mol. Med. Rep. 2018;17:1469–1484. doi: 10.3892/mmr.2017.8040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Van Rooij E., Olson E.N. MicroRNAs: Powerful new regulators of heart disease and provocative therapeutic targets. J. Clin. Investig. 2007;117:2369–2376. doi: 10.1172/JCI33099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Wang J., Xu R., Lin F., Zhang S., Zhang G., Hu S., Zheng Z. MicroRNA: Novel regulators involved in the remodeling and reverse remodeling of the heart. Cardiology. 2009;113:81–88. doi: 10.1159/000172616. [DOI] [PubMed] [Google Scholar]
- 144.Kukreja R.C., Yin C., Salloum F.N. MicroRNAs: New players in cardiac injury and protection. Mol. Pharmacol. 2011;80:558–564. doi: 10.1124/mol.111.073528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Duisters R.F., Tijsen A.J., Schroen B., Leenders J.J., Lentink V., van der Made I., Herias V., van Leeuwen R.E., Schellings M.W., Barenbrug P., et al. miR-133 and miR-30 regulate connective tissue growth factor: Implications for a role of microRNAs in myocardial matrix remodeling. Circ. Res. 2009;104:170–178. doi: 10.1161/CIRCRESAHA.108.182535. [DOI] [PubMed] [Google Scholar]
- 146.Liu W., Ling S., Sun W., Liu T., Li Y., Zhong G., Zhao D., Zhang P., Song J., Jin X., et al. Circulating microRNAs correlated with the level of coronary artery calcification in symptomatic patients. Sci. Rep. 2015;5:16099. doi: 10.1038/srep16099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Jiang Y., Zhang M., He H., Chen J., Zeng H., Li J., Duan R. MicroRNA/mRNA profiling and regulatory network of intracranial aneurysm. BMC Med. Genom. 2013;6:36. doi: 10.1186/1755-8794-6-36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Liu H., Xiong W., Liu F., Lin F., He J., Liu C., Lin Y., Dong S. Significant role and mechanism of microRNA-143-3p/KLLN axis in the development of coronary heart disease. Am. J. Transl. Res. 2019;11:3610–3619. [PMC free article] [PubMed] [Google Scholar]
- 149.Li C., Li J., Xue K., Zhang J., Wang C., Zhang Q., Chen X., Gao C., Yu X., Sun L. MicroRNA-143-3p promotes human cardiac fibrosis via targeting sprouty3 after myocardial infarction. J. Mol. Cell Cardiol. 2019;129:281–292. doi: 10.1016/j.yjmcc.2019.03.005. [DOI] [PubMed] [Google Scholar]
- 150.Yu B., Zhao Y., Zhang H., Xie D., Nie W., Shi K. Inhibition of microRNA-143-3p attenuates myocardial hypertrophy by inhibiting inflammatory response. Cell Biol. Int. 2018;42:1584–1593. doi: 10.1002/cbin.11053. [DOI] [PubMed] [Google Scholar]
- 151.Jiao M., You H.Z., Yang X.Y., Yuan H., Li Y.L., Liu W.X., Jin M., Du J. Circulating microRNA signature for the diagnosis of childhood dilated cardiomyopathy. Sci. Rep. 2018;8:724. doi: 10.1038/s41598-017-19138-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Deng L., Blanco F.J., Stevens H., Lu R., Caudrillier A., McBride M., McClure J.D., Grant J., Thomas M., Frid M., et al. MicroRNA-143 Activation Regulates Smooth Muscle and Endothelial Cell Crosstalk in Pulmonary Arterial Hypertension. Circ. Res. 2015;117:870–883. doi: 10.1161/CIRCRESAHA.115.306806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Shi L., Tian C., Sun L., Cao F., Meng Z. The lncRNA TUG1/miR-145-5p/FGF10 regulates proliferation and migration in VSMCs of hypertension. Biochem. Biophys. Res. Commun. 2018;501:688–695. doi: 10.1016/j.bbrc.2018.05.049. [DOI] [PubMed] [Google Scholar]
- 154.Yang X., Niu X., Xiao Y., Lin K., Chen X. MiRNA expression profiles in healthy OSAHS and OSAHS with arterial hypertension: Potential diagnostic and early warning markers. Respir. Res. 2018;19:194. doi: 10.1186/s12931-018-0894-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Toro R., Blasco-Turrión S., Morales-Ponce F.J., Gonzalez P., Martínez-Camblor P., López-Granados A., Brugada R., Campuzano O., Pérez-Serra A., Rosa Longobardo F., et al. Plasma microRNAs as biomarkers for Lamin A/C-related dilated cardiomyopathy. J. Mol. Med. 2018;96:845–856. doi: 10.1007/s00109-018-1666-1. [DOI] [PubMed] [Google Scholar]
- 156.Yuan M., Zhang L., You F., Zhou J., Ma Y., Yang F., Tao L. MiR-145-5p regulates hypoxia-induced inflammatory response and apoptosis in cardiomyocytes by targeting CD40. Mol. Cell Biochem. 2017;431:123–131. doi: 10.1007/s11010-017-2982-4. [DOI] [PubMed] [Google Scholar]
- 157.Wu G., Tan J., Li J., Sun X., Du L., Tao S. miRNA-145-5p induces apoptosis after ischemia-reperfusion by targeting dual specificity phosphatase 6. J. Cell Physiol. 2019;234:16281–16289. doi: 10.1002/jcp.28291. [DOI] [PubMed] [Google Scholar]
- 158.Xie X., Peng L., Zhu J., Zhou Y., Li L., Chen Y., Yu S., Zhao Y. miR-145-5p/Nurr1/TNF-α Signaling-Induced Microglia Activation Regulates Neuron Injury of Acute Cerebral Ischemic/Reperfusion in Rats. Front. Mol. Neurosci. 2017;10:383. doi: 10.3389/fnmol.2017.00383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Nunez Lopez Y.O., Retnakaran R., Zinman B., Pratley R.E., Seyhan A.A. Predicting and understanding the response to short-term intensive insulin therapy in people with early type 2 diabetes. Mol. Metab. 2019;20:63–78. doi: 10.1016/j.molmet.2018.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Zhang J., Cui C., Xu H. Downregulation of miR-145-5p elevates retinal ganglion cell survival to delay diabetic retinopathy progress by targeting FGF5. Biosci. Biotechnol. Biochem. 2019;83:1655–1662. doi: 10.1080/09168451.2019.1630251. [DOI] [PubMed] [Google Scholar]
- 161.Mona Zamanian A., Rezaei-Tavirani M., Rezaei-Tavirani M., Robati R.M. Gestational Diabetes Mellitus Regulatory Network Identifies hsa-miR-145-5p and hsa-miR-875-5p as Potential Biomarkers. Int. J. Endocrinol. Metab. 2019;17:e86640. doi: 10.5812/ijem.86640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Taganov K.D., Boldin M.P., Chang K.J., Baltimore D. NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl. Acad. Sci. USA. 2006;103:12481–12486. doi: 10.1073/pnas.0605298103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Paterson M.R., Kriegel A.J. MiR-146a/b: A family with shared seeds and different roots. Physiol. Genom. 2017;49:243–252. doi: 10.1152/physiolgenomics.00133.2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Zhang X., Ye Z.H., Liang H.W., Ren F.H., Li P., Dang Y.W., Chen G. Down-regulation of miR-146a-5p and its potential targets in hepatocellular carcinoma validated by a TCGA- and GEO-based study. FEBS Open Bio. 2017;7:504–521. doi: 10.1002/2211-5463.12198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165.Wang X., Ha T., Liu L., Zou J., Zhang X., Kalbfleisch J., Gao X., Williams D., Li C. Increased expression of microRNA-146a decreases myocardial ischaemia/reperfusion injury. Cardiovasc. Res. 2013;97:432–442. doi: 10.1093/cvr/cvs356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Quan X., Ji Y., Zhang C., Guo X., Zhang Y., Jia S., Ma W., Fan Y., Wang C. Circulating MiR-146a May be a Potential Biomarker of Coronary Heart Disease in Patients with Subclinical Hypothyroidism. Cell Physiol. Biochem. 2018;45:226–236. doi: 10.1159/000486769. [DOI] [PubMed] [Google Scholar]
- 167.Li S.H., Chen L., Pang X.M., Su S.Y., Zhou X., Chen C.Y., Huang L.G., Li J.P., Liu J.L. Decreased miR-146a expression in acute ischemic stroke directly targets the Fbxl10 mRNA and is involved in modulating apoptosis. Neurochem. Int. 2017;107:156–167. doi: 10.1016/j.neuint.2017.01.011. [DOI] [PubMed] [Google Scholar]
- 168.Barberio M.D., Kasselman L.J., Playford M.P., Epstein S.B., Renna H.A., Goldberg M., DeLeon J., Voloshyna I., Barlev A., Salama M., et al. Cholesterol efflux alterations in adolescent obesity: Role of adipose-derived extracellular vesical microRNAs. J. Transl. Med. 2019;17:232. doi: 10.1186/s12967-019-1980-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Gaudet A.D., Fonken L.K., Gushchina L.V., Aubrecht T.G., Maurya S.K., Periasamy M., Nelson R.J., Popovich P.G. miR-155 Deletion in Female Mice Prevents Diet-Induced Obesity. Sci. Rep. 2016;6:22862. doi: 10.1038/srep22862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Chen L., Zheng S.Y., Yang C.Q., Ma B.M., Jiang D. MiR-155-5p inhibits the proliferation and migration of VSMCs and HUVECs in atherosclerosis by targeting AKT1. Eur. Rev. Med. Pharmacol. Sci. 2019;23:2223–2233. doi: 10.26355/eurrev_201903_17270. [DOI] [PubMed] [Google Scholar]
- 171.Zhu M., Wei Y., Geißler C., Abschlag K., Corbalán Campos J., Hristov M., Möllmann J., Lehrke M., Karshovska E., Schober A. Hyperlipidemia-Induced MicroRNA-155-5p Improves β-Cell Function by Targeting Mafb. Diabetes. 2017;66:3072–3084. doi: 10.2337/db17-0313. [DOI] [PubMed] [Google Scholar]
- 172.Li S., Lee C., Song J., Lu C., Liu J., Cui Y., Liang H., Cao C., Zhang F., Chen H. Circulating microRNAs as potential biomarkers for coronary plaque rupture. Oncotarget. 2017;8:48145–48156. doi: 10.18632/oncotarget.18308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 173.Mukai N., Nakayama Y., Murakami S., Tanahashi T., Sessler D.I., Ishii S., Ogawa S., Tokuhira N., Mizobe T., Sawa T., et al. Potential contribution of erythrocyte microRNA to secondary erythrocytosis and thrombocytopenia in congenital heart disease. Pediatr. Res. 2018;83:866–873. doi: 10.1038/pr.2017.327. [DOI] [PubMed] [Google Scholar]
- 174.Klimczak D., Kuch M., Pilecki T., Żochowska D., Wirkowska A., Pączek L. Plasma microRNA-155-5p is increased among patients with chronic kidney disease and nocturnal hypertension. J. Am. Soc. Hypertens. 2017;11:831–841.e4. doi: 10.1016/j.jash.2017.10.008. [DOI] [PubMed] [Google Scholar]
- 175.Wang M., Sun L., Ding W., Cai S., Zhao Q. Ablation alleviates atrial fibrillation by regulating the signaling pathways of endothelial nitric oxide synthase/nitric oxide via miR-155-5p and miR-24-3p. J. Cell Biochem. 2019;120:4451–4462. doi: 10.1002/jcb.27733. [DOI] [PubMed] [Google Scholar]
- 176.Sun X., Sit A., Feinberg M.W. Role of miR-181 family in regulating vascular inflammation and immunity. Trends Cardiovasc. Med. 2014;24:105–112. doi: 10.1016/j.tcm.2013.09.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.Hulsmans M., Sinnaeve P., Van der Schueren B., Mathieu C., Janssens S., Holvoet P. Decreased miR-181a expression in monocytes of obese patients is associated with the occurrence of metabolic syndrome and coronary artery disease. J. Clin. Endocrinol. Metab. 2012;97:E1213–E1218. doi: 10.1210/jc.2012-1008. [DOI] [PubMed] [Google Scholar]
- 178.Du X., Yang Y., Xu C., Peng Z., Zhang M., Lei L., Gao W., Dong Y., Shi Z., Sun X., et al. Upregulation of miR-181a impairs hepatic glucose and lipid homeostasis. Oncotarget. 2017;8:91362–91378. doi: 10.18632/oncotarget.20523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179.Wu J., Fan C.L., Ma L.J., Liu T., Wang C., Song J.X., Lv Q.S., Pan H., Zhang C.N., Wang J.J. Distinctive expression signatures of serum microRNAs in ischaemic stroke and transient ischaemic attack patients. Thromb. Haemost. 2017;117:992–1001. doi: 10.1160/TH16-08-0606. [DOI] [PubMed] [Google Scholar]
- 180.Zhu J., Yao K., Wang Q., Guo J., Shi H., Ma L., Liu H., Gao W., Zou Y., Ge J. Circulating miR-181a as a Potential Novel Biomarker for Diagnosis of Acute Myocardial Infarction. Cell Physiol. Biochem. 2016;40:1591–1602. doi: 10.1159/000453209. [DOI] [PubMed] [Google Scholar]
- 181.Nabih E.S., Andrawes N.G. The Association Between Circulating Levels of miRNA-181a and Pancreatic Beta Cells Dysfunction via SMAD7 in Type 1 Diabetic Children and Adolescents. J. Clin. Lab. Anal. 2016;30:727–731. doi: 10.1002/jcla.21928. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 182.He J.F., Luo Y.M., Wan X.H., Jiang D. Biogenesis of MiRNA-195 and its role in biogenesis, the cell cycle, and apoptosis. J. Biochem. Mol. Toxicol. 2011;25:404–408. doi: 10.1002/jbt.20396. [DOI] [PubMed] [Google Scholar]
- 183.Van Rooij E., Sutherland L.B., Liu N., Williams A.H., McAnally J., Gerard R.D., Richardson J.A., Olson E.N. A signature pattern of stress-responsive microRNAs that can evoke cardiac hypertrophy and heart failure. Proc. Natl. Acad. Sci. USA. 2006;103:18255–18260. doi: 10.1073/pnas.0608791103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 184.You X.Y., Huang J.H., Liu B., Liu S.J., Zhong Y., Liu S.M. HMGA1 is a new target of miR-195 involving isoprenaline-induced cardiomyocyte hypertrophy. Biochemistry. 2014;79:538–544. doi: 10.1134/S0006297914060078. [DOI] [PubMed] [Google Scholar]
- 185.Zampetaki A., Attia R., Mayr U., Gomes R.S., Phinikaridou A., Yin X., Langley S.R., Willeit P., Lu R., Fanshawe B., et al. Role of miR-195 in aortic aneurysmal disease. Circ. Res. 2014;115:857–866. doi: 10.1161/CIRCRESAHA.115.304361. [DOI] [PubMed] [Google Scholar]
- 186.Du J., Zheng R., Xiao F., Zhang S., He K., Zhang J., Shao Y. Downregulated MicroRNA-195 in the Bicuspid Aortic Valve Promotes Calcification of Valve Interstitial Cells via Targeting SMAD7. Cell Physiol. Biochem. 2017;44:884–896. doi: 10.1159/000485356. [DOI] [PubMed] [Google Scholar]
- 187.Collares C.V., Evangelista A.F., Xavier D.J., Rassi D.M., Arns T., Foss-Freitas M.C., Foss M.C., Puthier D., Sakamoto-Hojo E.T., Passos G.A., et al. Identifying common and specific microRNAs expressed in peripheral blood mononuclear cell of type 1, type 2, and gestational diabetes mellitus patients. BMC Res. Notes. 2013;6:491. doi: 10.1186/1756-0500-6-491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 188.Massaro J.D., Polli C.D., Costa E., Silva M., Alves C.C., Passos G.A., Sakamoto-Hojo E.T., Rodrigues de Holanda Miranda W., Bispo Cezar N.J., Rassi D.M., et al. Post-transcriptional markers associated with clinical complications in Type 1 and Type 2 diabetes mellitus. Mol. Cell Endocrinol. 2019;490:1–14. doi: 10.1016/j.mce.2019.03.008. [DOI] [PubMed] [Google Scholar]
- 189.Li M., Luan L., Liu Q., Liu Y., Lan X., Li Z., Liu W. MiRNA-199a-5p Protects Against Cerebral Ischemic Injury by Down-Regulating DDR1 in Rats. World Neurosurg. 2019;131:e486–e494. doi: 10.1016/j.wneu.2019.07.203. [DOI] [PubMed] [Google Scholar]
- 190.Yan M., Yang S., Meng F., Zhao Z., Tian Z., Yang P. MicroRNA 199a-5p induces apoptosis by targeting JunB. Sci. Rep. 2018;8:6699. doi: 10.1038/s41598-018-24932-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Lynch S.M., Ward M., McNulty H., Angel C.Z., Horigan G., Strain J.J., Purvis J., Tackett M., McKenna D.J. Serum levels of miR-199a-5p correlates with blood pressure in premature cardiovascular disease patients homozygous for the MTHFR 677C > T polymorphism. Genom. 2020;112:669–676. doi: 10.1016/j.ygeno.2019.04.019. [DOI] [PubMed] [Google Scholar]
- 192.Tian X., Yu C., Shi L., Li D., Chen X., Xia D., Zhou J., Xu W., Ma C., Gu L., et al. MicroRNA-199a-5p aggravates primary hypertension by damaging vascular endothelial cells through inhibition of autophagy and promotion of apoptosis. Exp. Ther. Med. 2018;16:595–602. doi: 10.3892/etm.2018.6252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193.Zhou Y., Pang B., Xiao Y., Zhou S., He B., Zhang F., Liu W., Peng H., Li P. The protective microRNA-199a-5p-mediated unfolded protein response in hypoxic cardiomyocytes is regulated by STAT3 pathway. J. Physiol. Biochem. 2019;75:73–81. doi: 10.1007/s13105-018-0657-6. [DOI] [PubMed] [Google Scholar]
- 194.Liu Y., Liu G., Zhang H., Wang J. MiRNA-199a-5p influences pulmonary artery hypertension via downregulating Smad3. Biochem. Biophys. Res. Commun. 2016;473:859–866. doi: 10.1016/j.bbrc.2016.03.140. [DOI] [PubMed] [Google Scholar]
- 195.Wang J., Yu G. A Systems Biology Approach to Characterize Biomarkers for Blood Stasis Syndrome of Unstable Angina Patients by Integrating MicroRNA and Messenger RNA Expression Profiling. Evid. Based Complement. Altern. Med. 2013;2013:510208. doi: 10.1155/2013/510208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 196.Yu L., Gu T., Shi E., Wang Y., Fang Q., Wang C. Dysregulation of renal microRNA expression after deep hypothermic circulatory arrest in rats. Eur J. Cardiothorac. Surg. 2016;49:1725–1731. doi: 10.1093/ejcts/ezv460. [DOI] [PubMed] [Google Scholar]
- 197.Hirt M.N., Werner T., Indenbirken D., Alawi M., Demin P., Kunze A.C., Stenzig J., Starbatty J., Hansen A., Fiedler J., et al. Deciphering the microRNA signature of pathological cardiac hypertrophy by engineered heart tissue- and sequencing-technology. J. Mol. Cell Cardiol. 2015;81:1–9. doi: 10.1016/j.yjmcc.2015.01.008. [DOI] [PubMed] [Google Scholar]
- 198.Aguado-Fraile E., Ramos E., Conde E., Rodríguez M., Martín-Gómez L., Lietor A., Candela Á., Ponte B., Liaño F., García-Bermejo M.L. A Pilot Study Identifying a Set of microRNAs As Precise Diagnostic Biomarkers of Acute Kidney Injury. PLoS ONE. 2015;10:e0127175. doi: 10.1371/journal.pone.0127175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 199.Ma H., Chen P., Sang C., Huang D., Geng Q., Wang L. Modulation of apoptosis-related microRNAs following myocardial infarction in fat-1 transgenic mice vs wild-type mice. J. Cell Mol. Med. 2018;22:5698–5707. doi: 10.1111/jcmm.13846. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 200.Qiao X.R., Wang L., Liu M., Tian Y., Chen T. MiR-210-3p attenuates lipid accumulation and inflammation in atherosclerosis by repressing IGF2. Biosci. Biotechnol. Biochem. 2020;84:321–329. doi: 10.1080/09168451.2019.1685370. [DOI] [PubMed] [Google Scholar]
- 201.Derda A.A., Pfanne A., Bwangär C., Schimmel K., Kennel P.J., Xiao K., Schulze P.C., Bauersachs J., Thum T. Blood-based microRNA profiling in patients with cardiac amyloidosis. PLoS ONE. 2018;13:e0204235. doi: 10.1371/journal.pone.0204235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 202.Verjans R., Peters T., Beaumont F.J., van Leeuwen R., van Herwaarden T., Verhesen W., Munts C., Bijnen M., Henkens M., Diez J., et al. MicroRNA-221/222 Family Counteracts Myocardial Fibrosis in Pressure Overload-Induced Heart Failure. Hypertension. 2018;71:280–288. doi: 10.1161/HYPERTENSIONAHA.117.10094. [DOI] [PubMed] [Google Scholar]
- 203.Zhuang X., Li R., Maimaitijiang A., Liu R., Yan F., Hu H., Gao X., Shi H. miR-221-3p inhibits oxidized low-density lipoprotein induced oxidative stress and apoptosis via targeting a disintegrin and metalloprotease-22. J. Cell Biochem. 2019;120:6304–6314. doi: 10.1002/jcb.27917. [DOI] [PubMed] [Google Scholar]
- 204.Pereira-da-Silva T., Coutinho Cruz M., Carrusca C., Cruz Ferreira R., Napoleão P., Mota Carmo M. Circulating microRNA profiles in different arterial territories of stable atherosclerotic disease: A systematic review. Am. J. Cardiovasc. Dis. 2018;8:1–13. [PMC free article] [PubMed] [Google Scholar]
- 205.Coffey S., Williams M.J., Phillips L.V., Galvin I.F., Bunton R.W., Jones G.T. Integrated microRNA and messenger RNA analysis in aortic stenosis. Sci. Rep. 2016;6:36904. doi: 10.1038/srep36904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 206.Coskunpinar E., Cakmak H.A., Kalkan A.K., Tiryakioglu N.O., Erturk M., Ongen Z. Circulating miR-221-3p as a novel marker for early prediction of acute myocardial infarction. Gene. 2016;591:90–96. doi: 10.1016/j.gene.2016.06.059. [DOI] [PubMed] [Google Scholar]
- 207.Sørensen S.S., Nygaard A.B., Nielsen M.Y., Jensen K., Christensen T. miRNA expression profiles in cerebrospinal fluid and blood of patients with acute ischemic stroke. Transl. Stroke Res. 2014;5:711–718. doi: 10.1007/s12975-014-0364-8. [DOI] [PubMed] [Google Scholar]
- 208.Gusar V.A., Timofeeva A.V., Zhanin I.S., Shram S.I., Pinelis V.G. Estimation of Time-Dependent microRNA Expression Patterns in Brain Tissue, Leukocytes, and Blood Plasma of Rats under Photochemically Induced Focal Cerebral Ischemia. Mol. Biol. 2017;51:683–695. doi: 10.1134/S0026893317040100. [DOI] [PubMed] [Google Scholar]
- 209.Nie X., Chen Y., Tan J., Dai Y., Mao W., Qin G., Ye S., Sun J., Yang Z., Chen J. MicroRNA-221-3p promotes pulmonary artery smooth muscle cells proliferation by targeting AXIN2 during pulmonary arterial hypertension. Vascul. Pharmacol. 2019;116:24–35. doi: 10.1016/j.vph.2017.07.002. [DOI] [PubMed] [Google Scholar]
- 210.Villard A., Marchand L., Thivolet C., Rome S. Diagnostic Value of Cell-free Circulating MicroRNAs for Obesity and Type 2 Diabetes: A Meta-analysis. J. Mol. Biomark. Diagn. 2015;6:251. doi: 10.4172/2155-9929.1000251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211.Wang L., Xu L., Xu M., Liu G., Xing J., Sun C., Ding H. Obesity-Associated MiR-342-3p Promotes Adipogenesis of Mesenchymal Stem Cells by Suppressing CtBP2 and Releasing C/EBPα from CtBP2 Binding. Cell Physiol. Biochem. 2015;35:2285–2298. doi: 10.1159/000374032. [DOI] [PubMed] [Google Scholar]
- 212.Hezova R., Slaby O., Faltejskova P., Mikulkova Z., Buresova I., Raja K.R., Hodek J., Ovesna J., Michalek J. microRNA-342, microRNA-191 and microRNA-510 are differentially expressed in T regulatory cells of type 1 diabetic patients. Cell Immunol. 2010;260:70–74. doi: 10.1016/j.cellimm.2009.10.012. [DOI] [PubMed] [Google Scholar]
- 213.Eissa S., Matboli M., Bekhet M.M. Clinical verification of a novel urinary microRNA panal: 133b, -342 and -30 as biomarkers for diabetic nephropathy identified by bioinformatics analysis. Biomed. Pharmacother. 2016;83:92–99. doi: 10.1016/j.biopha.2016.06.018. [DOI] [PubMed] [Google Scholar]
- 214.Cheng S., Cui Y., Fan L., Mu X., Hua Y. T2DM inhibition of endothelial miR-342-3p facilitates angiogenic dysfunction via repression of FGF11 signaling. Biochem. Biophys. Res. Commun. 2018;503:71–78. doi: 10.1016/j.bbrc.2018.05.179. [DOI] [PubMed] [Google Scholar]
- 215.Khalyfa A., Kheirandish-Gozal L., Bhattacharjee R., Khalyfa A.A., Gozal D. Circulating microRNAs as Potential Biomarkers of Endothelial Dysfunction in Obese Children. Chest. 2016;149:786–800. doi: 10.1378/chest.15-0799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 216.Hoekstra M. MicroRNA-499-5p: A therapeutic target in the context of cardiovascular disease. Ann. Transl. Med. 2016;4:539. doi: 10.21037/atm.2016.11.61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 217.Zhao L., Wang B., Zhang W., Sun L. Effect of miR-499a-5p on damage of cardiomyocyte induced by hypoxia-reoxygenation via downregulating CD38 protein. J. Cell Biochem. 2020;121:996–1004. doi: 10.1002/jcb.29334. [DOI] [PubMed] [Google Scholar]
- 218.Neshati V., Mollazadeh S., Fazly Bazzaz B.S., de Vries A.A.F., Mojarrad M., Naderi-Meshkin H., Neshati Z., Mirahmadi M., Kerachian M.A. MicroRNA-499a-5p Promotes Differentiation of Human Bone Marrow-Derived Mesenchymal Stem Cells to Cardiomyocytes. Appl. Biochem. Biotechnol. 2018;186:245–255. doi: 10.1007/s12010-018-2734-2. [DOI] [PubMed] [Google Scholar]
- 219.Boštjančič E., Zidar N., Glavač D. MicroRNAs and cardiac sarcoplasmic reticulum calcium ATPase-2 in human myocardial infarction: Expression and bioinformatic analysis. BMC Genom. 2012;13:552. doi: 10.1186/1471-2164-13-552. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 220.Salinas J., Lin H., Aparico H.J., Huan T., Liu C., Rong J., Beiser A., Himali J.J., Freedman J.E., Larson M.G., et al. Whole blood microRNA expression associated with stroke: Results from the Framingham Heart Study. PLoS ONE. 2019;14:e0219261. doi: 10.1371/journal.pone.0219261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 221.Baldeón Rojas L., Weigelt K., de Wit H., Ozcan B., van Oudenaren A., Sempértegui F., Sijbrands E., Grosse L., Van Zonneveld A.J., Drexhage H.A., et al. Study on inflammation-related genes and microRNAs, with special emphasis on the vascular repair factor HGF and miR-574-3p, in monocytes and serum of patients with T2D. Diabetol. Metab. Syndr. 2016;8:6. doi: 10.1186/s13098-015-0113-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 222.Houshmand-Oeregaard A., Schrölkamp M., Kelstrup L., Hansen N.S., Hjort L., Thuesen A.C.B., Broholm C., Mathiesen E.R., Clausen T.D., Vaag A., et al. Increased Expression of microRNA-15a and microRNA-15b in Skeletal Muscle from Adult Offspring of Women with Diabetes in Pregnancy. Hum. Mol. Genet. 2018;27:1763–1771. doi: 10.1093/hmg/ddy085. [DOI] [PubMed] [Google Scholar]
- 223.International Association of Diabetes and Pregnancy Study Groups Consensus Panel. Metzger B.E., Gabbe S.G., Persson B., Buchanan T.A., Catalano P.A., Damm P., Dyer A.R., Leiva A.D., Hod M., et al. International association of diabetes and pregnancy study groups recommendations on the diagnosis and classification of hyperglycemia in pregnancy. Diabetes Care. 2010;33:676–682. doi: 10.2337/dc10-0719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 224.American Diabetes Association Diagnosis and classification of diabetes mellitus (Position Statement) Diabetes Care. 2009;32:S62–S67. doi: 10.2337/dc09-S062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 225.Metzger B.E., Coustan D.R. Summary and recommendations of the Fourth International Workshop-Conference on Gestational Diabetes Mellitus. The Organizing Committee. Diabetes Care. 1998;21:B161–B167. [PubMed] [Google Scholar]
- 226.Hromadnikova I., Kotlabova K., Dvorakova L., Krofta L., Sirc J. Postnatal Expression Profile of microRNAs Associated with Cardiovascular and Cerebrovascular Diseases in Children at the Age of 3 to 11 Years in Relation to Previous Occurrence of Pregnancy-Related Complications. Int. J. Mol. Sci. 2019;20:654. doi: 10.3390/ijms20030654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 227.National High Blood Pressure Education Program Working Group on High Blood Pressure in Children and Adolescents The fourth report on the diagnosis, evaluation, and treatment of high blood pressure in children and adolescents. Pediatrics. 2004;114:555–576. doi: 10.1542/peds.114.2.S2.555. [DOI] [PubMed] [Google Scholar]
- 228.Livak K.J., Schmittgen T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25:402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
- 229.Vandesompele J., de Preter K., Pattyn F., Poppe B., Van Roy N., de Paepe A., Speleman F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002;3:research0034. doi: 10.1186/gb-2002-3-7-research0034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 230.Shapiro S.S., Wilk M.B. An Analysis of Variance Test for Normality (Complete Samples) Biometrika. 1965;52:591–611. doi: 10.1093/biomet/52.3-4.591. [DOI] [Google Scholar]
- 231.Dweep H., Sticht C., Pandey P., Gretz N. miRWalk—database: Prediction of possible miRNA binding sites by “walking” the genes of three genomes. J. Biomed. Inform. 2011;44:839–847. doi: 10.1016/j.jbi.2011.05.002. [DOI] [PubMed] [Google Scholar]
- 232.Hromadnikova I. Postnatální Epigenetický Profil Kardiovaskulárních MikroRNA U Dětí Narozených Z Komplikovaných Gravidit-Nové Biomarkery Kardiovaskulárního Rizika. 308102. CZ Patent. 2018 Oct 31; Issued 20 November 2019. Industrial Property Office, Czech Republic.
- 233.Hromadnikova I. A Method of Predicting a Risk of Cardiovascular Disease for a Child Born from a Complicated Pregnancy. PCT International Application No. PCT/CZ2019/050050. 2019 Oct 30; Industrial Property Office, Czech Republic.
- 234.Rivera R.M., Bennett L.B. Epigenetics in Humans: An Overview. Curr. Opin. Endocrinol. Diabetes Obes. 2010;17:493–499. doi: 10.1097/MED.0b013e3283404f4b. [DOI] [PubMed] [Google Scholar]
- 235.Saetrom P., Snøve O., Jr., Rossi J.J. Epigenetics and microRNAs. Pediatr. Res. 2007;61:17R–23R. doi: 10.1203/pdr.0b013e318045760e. [DOI] [PubMed] [Google Scholar]
- 236.Tost J. Epigenetics. Caister Academic Press; Norfolk, UK: 2008. pp. 1–404. [Google Scholar]
- 237.Li E. Chromatin modification and epigenetic reprogramming in mammalian development. Nat. Rev. Genet. 2002;3:662–673. doi: 10.1038/nrg887. [DOI] [PubMed] [Google Scholar]
- 238.Sinkkonen L., Hugenschmidt T., Berninger P., Gaidatzis D., Mohn F., Artus-Revel C.G., Zavolan M., Svoboda P., Filipowicz W. MicroRNAs Control De Novo DNA Methylation Through Regulation of Transcriptional Repressors in Mouse Embryonic Stem Cells. Nat. Struct. Mol. Biol. 2008;15:259–267. doi: 10.1038/nsmb.1391. [DOI] [PubMed] [Google Scholar]
- 239.Ma J., Flemr M., Stein P., Berninger P., Malik R., Zavolan M., Svoboda P., Schultz R.M. MicroRNA Activity Is Suppressed in Mouse Oocytes. Curr. Biol. 2010;20:265–270. doi: 10.1016/j.cub.2009.12.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 240.Suh N., Baehner L., Moltzahn F., Melton C., Shenoy A., Chen J., Blelloch R. MicroRNA Function Is Globally Suppressed in Mouse Oocytes and Early Embryos. Curr. Biol. 2010;20:271–277. doi: 10.1016/j.cub.2009.12.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 241.Gu Y., Sun J., Groome L.J., Wang Y. Differential miRNA expression profiles between the first and third trimester human placentas. Am. J. Physiol. Endocrinol. Metab. 2013;304:E836–E843. doi: 10.1152/ajpendo.00660.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 242.Maccani M.A., Padbury J.F., Lester B.M., Knopik V.S., Marsit C.J. Placental miRNA expression profiles are associated with measures of infant neurobehavioral outcomes. Pediatr. Res. 2013;74:272–278. doi: 10.1038/pr.2013.102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 243.Soni K., Choudhary A., Patowary A., Singh A.R., Bhatia S., Sivasubbu S., Chandrasekaran S., Pillai B. miR-34 Is Maternally Inherited in Drosophila Melanogaster and Danio Rerio. Nucleic Acids Res. 2013;41:4470–4480. doi: 10.1093/nar/gkt139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 244.Babenko O., Kovalchuka I., Metz G.A. Epigenetic programming of neurodegenerative diseases by an adverse environment. Brain Res. 2015;1444:96–111. doi: 10.1016/j.brainres.2012.01.038. [DOI] [PubMed] [Google Scholar]
- 245.Rodgers A.B., Morgan C.P., Leu N.A., Bale T.L. Transgenerational epigenetic programming via sperm microRNA recapitulates effects of paternal stress. Proc. Natl. Acad. Sci. USA. 2015;112:13699–13704. doi: 10.1073/pnas.1508347112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 246.Fraser R., Lin C.J. Epigenetic reprogramming of the zygote in mice and men: On your marks, get set, go! Reproduction. 2016;152:R211–R222. doi: 10.1530/REP-16-0376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 247.Pang T.Y.C., Short A.K., Bredy T.W., Hannan A.J. Transgenerational paternal transmission of acquired traits: Stress-induced modification of the sperm regulatory transcriptome and offspring phenotypes. Curr. Opin. Behav. Sci. 2017;14:140–147. doi: 10.1016/j.cobeha.2017.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 248.Yeshurun S., Hannan A.J. Transgenerational epigenetic influences of paternal environmental exposures on brain function and predisposition to psychiatric disorders. Mol. Psychiatry. 2019;24:536–548. doi: 10.1038/s41380-018-0039-z. [DOI] [PubMed] [Google Scholar]
- 249.Lacal I., Ventura R. Epigenetic Inheritance: Concepts, Mechanisms and Perspectives. Front. Mol. Neurosci. 2018;11:292. doi: 10.3389/fnmol.2018.00292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 250.Mathers J.C., Strathdee G., Relton C.L. Induction of epigenetic alterations by dietary and other environmental factors. Adv. Genet. 2010;71:3–39. doi: 10.1016/B978-0-12-380864-6.00001-8. [DOI] [PubMed] [Google Scholar]
- 251.Christensen B.C., Marsit C.J. Epigenomics in environmental health. Front. Genet. 2011;2:84. doi: 10.3389/fgene.2011.00084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 252.Talikka M., Sierro N., Ivanov N.V., Chaudhary N., Peck M.J., Hoeng J., Coggins C.R.E., Peitsch C.M. Genomic Impact of Cigarette Smoke, With Application to Three Smoking-Related Diseases. Crit. Rev. Toxicol. 2012;42:877–889. doi: 10.3109/10408444.2012.725244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 253.Cabib S., Puglisi-Allegra S. The mesoaccumbens dopamine in coping with stress. Neurosci. Biobehav. Rev. 2012;36:79–89. doi: 10.1016/j.neubiorev.2011.04.012. [DOI] [PubMed] [Google Scholar]
- 254.Van Otterdijk S.D., Mathers J.C., Strathdee G. Do age-related changes in DNA methylation play a role in the development of age-related diseases? Biochem. Soc. Trans. 2013;41:803–807. doi: 10.1042/BST20120358. [DOI] [PubMed] [Google Scholar]
- 255.Jung M., Pfeifer G.P. Aging and DNA methylation. BMC Biol. 2015;13:7. doi: 10.1186/s12915-015-0118-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 256.Deans C., Maggert K.A. What Do You Mean, “Epigenetic”? Genetics. 2015;199:887–896. doi: 10.1534/genetics.114.173492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 257.Zoghbi H.Y., Beaudet A.L. Epigenetics and Human Disease. Cold Spring Harb. Perspect. Biol. 2016;8:a019497. doi: 10.1101/cshperspect.a019497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 258.Tobias D.K., Chavarro J.E., Williams M.A., Buck Louis G.M., Hu F.B., Rich-Edwards J., Missmer S.A., Zhang C. History of Infertility and Risk of Gestational Diabetes Mellitus: A Prospective Analysis of 40,773 Pregnancies. Am. J. Epidemiol. 2013;178:1219–1225. doi: 10.1093/aje/kwt110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 259.Kouhkan A., Khamseh M.E., Pirjani R., Moini A., Arabipoor A., Maroufizadeh S., Hosseini R., Baradaran H.R. Obstetric and perinatal outcomes of singleton pregnancies conceived via assisted reproductive technology complicated by gestational diabetes mellitus: A prospective cohort study. BMC Pregnancy Childbirth. 2018;18:495. doi: 10.1186/s12884-018-2115-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 260.Mohammadi M., Morasae E.K., Maroufizadeh S., Almasi-Hashiani A., Navid B., Amini P., Omani-Samani R., Alizadeh A. Assisted reproductive technology and the risk of gestational diabetes mellitus: A systemic review and meta-analysis. Middle East Fertil. Soc. J. 2020;25:6. doi: 10.1186/s43043-020-0018-6. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.