Abstract
N6-methyladenosine (m6A) is one of the most common RNA modifications in eukaryotes, mainly in messenger RNA (mRNA). Increasing evidence shows that m6A methylation modification acts an essential role in various physiological and pathological bioprocesses. Noncoding RNAs (ncRNAs), including miRNAs, lncRNAs and circRNAs, are known to participate in regulating cell differentiation, angiogenesis, immune response, inflammatory response and carcinogenesis. m6A regulators, such as METTL3, ALKBH5 and IGF2BP1 have been reported to execute a m6A-dependent modification of ncRNAs involved in carcinogenesis. Meanwhile, ncRNAs can target or modulate m6A regulators to influence cancer development. In this review, we provide an insight into the interplay between m6A modification and ncRNAs in cancer.
Keywords: Noncoding RNAs, Cancer, m6A RNA methylation
Introduction
Up to now, more than 100 kinds of RNA modifications have been confirmed [1]. Among them, m6A RNA methylation is one of the most thoroughly studied modifications. m6A RNA modification occurs by methylation of the sixth N atom of adenine (A) in mRNAs or ncRNAs [2]. m6A modification sites tend to be found in the stop codons and 3′-Untranslated region (3′-UTR) of mRNA with a typical consensus sequence RRACH (R = G or A and H = A, C, or U) [3, 4]. Accumulating data show that m6A RNA methylation acts by modulating circadian rhythm, gene expression, cell differentiation, stress response, inflammatory response, and carcinogenesis [5–10]. According to the global cancer statistics, there were estimated 18.1 million new cases and 9.6 million deaths in 2018 [11]. Recent studies have shown that m6A modification acts a vital role in the diagnosis, treatment and prognosis of cancer patients as well as in carcinogenesis. It also regulates fly sex, virus genome, meiosis of yeast, tissue differentiation, germination, and collateral generation of Arabidopsis [12–15].
Noncoding RNAs (ncRNAs) including microRNAs (miRNAs), long non-coding RNAs (lncRNAs) and circular RNAs (circRNAs) act pivotal roles in cancer [16–18]. m6A modification can affect ncRNA splicing and maturation involved in carcinogenesis (Table 1). In this review, we summarize the latest progress about the interplay between m6A modification and ncRNAs in cancer.
Table 1.
m6A methylation modifies ncRNAs in cancers
| m6A component | Related non-coding RNA | Cancer | Function | Role in cancer | Regulation | References |
|---|---|---|---|---|---|---|
| METTL3 | miR-25-3p | PDAC | Writers | Oncogene | Up-regulation | [45] |
| miR-221、miR-222 | Bladder cancer | Writers | Oncogene | Up-regulation | [46] | |
| miR-106b, miR-18a/b, miR-3607, miR-423, miR-30a, miR-320b/d/e | arsenite-induced carcinogenesis | Writers | Oncogene | Up-regulation | [47] | |
| miR-1246 | CRC | Writers | Oncogene | Up-regulation | [49] | |
| miR-143-3p | Lung cancer | Writers | Oncogene | Up-regulation | [50] | |
| METTL14 | miR-126 | HCC | Writers | Anti-oncogene | Down-regulation | [48] |
| METTL3 | lncRNA FAM225A | NPC | Writers | Oncogene | Up-regulation | [67] |
| lncRNA LINC00958 | HCC | Writers | Oncogene | Up-regulation | [65] | |
| lncRNA RP11 | CRC | Writers | Oncogene | Up-regulation | [68] | |
| MALAT1 | NSCLC | Writers | Oncogene | Up-regulation | [69] | |
| METTL14 | XIST | CRC | Writers | Anti-oncogene | Down-regulation | [70] |
| METTL3/METTL14 | LNCAROD | HNSCC | Writers | Oncogene | Up-regulation | [66] |
| ALKBH5 | lncRNA NEAT1 | GC | Erasers | Oncogene | Up-regulation | [71] |
| lncRNA FOXM1-AS | glioblastoma | Erasers | Oncogene | Up-regulation | [57] | |
| YTHDF1 | LINC00278 | ESCC | Readers | Anti-oncogene | Down-regulation | [72] |
| IGF2BP2 | lncRNA DANCR | Pancreatic cancer | Readers | Oncogene | Up-regulation | [60] |
PDAC pancreatic ductal adenocarcinoma, HCC hepatocellular cancer, NPC nasopharyngeal cancer, GC gastric cancer, CRC colorectal cancer, NSCLC non-small cell lung cancer, HNSCC head and neck squamous cell carcinoma, ESCC esophageal squamous cell carcinoma
Molecular compositions of m6A RNA methylation
Molecular compositions of m6A RNA methylation include m6A methyltransferase, m6A demethylase, and m6A recognition factors (Fig. 1). m6A methyltransferases, called “writers” contain methyltransferase-like 3 (METTL3) [19], METTL14 [20], Wilms tumor 1-associated protein (WTAP) [2], KIAA1429 [21], METTL16 [22] and RNA-binding motif protein 15/15B (RBM15/15B) [23]. METTL3 regulates the circadian clock of hepatic lipid metabolism and hematopoiesis [24, 25]. METTL3/14 depletion promotes myeloid differentiation and suppresses the progression of acute myeloid leukemia (AML) [26, 27]. METTL16 maintains the levels of methyl donor S-adenosylmethionine (SAM) [28]. WTAP connects METTL3/14 to form a complex, anchored to the nucleus to catalyze m6A methyltransferase [2].
Fig. 1.
m6A modification is a dynamic and reversible process. m6A modification can be executed by “Writers” (METTL3/14, WTAP, KIAA1429, RBM15/15B, METTL16), demethylated by “Erasers” (FTO and ALKBH5) and regulated by “readers” (YTHDF1–3, YTHDC1–2, IGFBPs, eIF3 and HNRNPA2B1)
m6A methylation is dynamic and can be reversed by m6A demethylase, also named as m6A “erasers”, containing fat mass and obesity-associated protein (FTO) and alkB homologue 5 (ALKBH5) [29, 30]. FTO shares the motifs with Fe (II)- and 2-oxoglutarate-dependent oxygenase and is related to increased fat mass [31]. FTO harbors an efficient oxidative demethylation activity and reduces the m6A levels of mRNAs [30]. ALKBH5 is responsible for RNA splicing and stability and causes the degradation of abnormal transcripts in spermatocytes and round spermatids [32].
m6A recognition factors, known as “readers,” consist of YT521-B homology (YTH) domain family (YTHDF1/2/3) [33], YTH domain-containing proteins (YTHDC1/2) [12], heterogeneous nuclear ribonucleoprotein (HNRNP) protein families [33], eukaryotic translation initiation factor 3 (eIF3) [23], and insulin-like growth factor-2 mRNA-binding proteins 1/2/3 (IGF2BP1/2/3) [34]. m6A recognition factors act in oligodendrocyte progenitor cells and oligodendrocyte fate [35]. YTHDF1 controls pre-crossing axon guidance in the spinal cord by regulating m6A-modified Robo3.1 [36]. HNRNPA2B1 can initiate the immune response to DNA viruses by regulating interferon-α/β and stimulator of interferon genes (STING)-dependent antiviral signaling [37].
m6A modification of miRNAs in cancer
As is known to us, the dysregulation of miRNAs is involved in various bio-behaviors, such as mouse prenatal development, immune response, inflammatory response and carcinogenesis [38–41]. METTL3 or HNRNPA2B1 facilitates pri-miRNA processing by recruiting RNA-binding protein DiGeorge syndrome critical region 8 (DGCR8) [42, 43]. METTL3 suppresses osteogenic processes by promoting the maturation of miR-7212-5p and downregulating its target fibroblast growth factor receptor 3 (FGFR3) [44].
Tumor proliferation and tumorigenesis
m6A methylation can modify the maturation of miRNAs involved in cell proliferation and tumorigenesis (Fig. 2). miR-25-3p acts as a pivotal role in pancreatic ductal adenocarcinoma (PDAC). Cigarette smoke condensate (CSC) mediates METTL3 to promote miR-25-3p maturation in PDAC tumorigenesis [45]. METTL3 also enhances the binding of pri-miR-221/222 with DGCR8 involved in the proliferation of bladder cancer [46]. m6A modification affects arsenite-induced carcinogenesis via modifying multiple miRNAs (miR-106b, miR-18a/b, miR-3607, miR-423, miR-30a, miR-320b/d/e) [47].
Fig. 2.
m6A methylation modifies miRNAs to regulate tumorigenesis and metastasis in multiple cancers including PDAC, lung cancer, HCC, bladder cancer, and CRC
Tumor invasion and metastasis
METTL14 promotes the maturation of pri-miR-126 and suppresses the invasion and metastasis of hepatocellular carcinoma (HCC) [48]. METTL3 facilitates the maturation of pri-miR-1246 to enhance the metastasis of colorectal cancer (CRC) [49]. METTL3 also accelerates the maturation of miR-143-3p, leading to the formation of METTL3/miR-143-3p/vasohibin-1 axis to favor the metastasis of lung cancers [50].
m6A modification of lncRNAs in cancer
LncRNAs, a subgroup of non-coding RNAs over 200 nucleotides in length can be modified by m6A methylation in cancer (Fig. 3). m6A methylation facilitates lncRNA X-inactive specific transcript (XIST)-mediated transcriptional repression [51–53]. YTHDC1 preferentially recognizes the m6A residues of XIST and RBM15/15B and participates in XIST-mediated gene silencing [53]. However, RBM15/m6A-MTase complex is reported to act a minor role in XIST-mediated gene silencing [54]. YTHDF2 recognizes m6A methylation site of lnc-Dpf3 to promote its degradation and enhances the binding of lnc-Dpf3 with hypoxia-inducible factor 1-alpha (HIF-1α), leading to the suppression of the glycolysis and migration of dendritic cells [55]. METTL3 can modify metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) to form the METTL3/MALAT1/miR-145/focal adhesion kinase (FAK) axis, contributing to the aggravation of renal fibrogenesis in obstructive nephropathy [56].
Fig. 3.
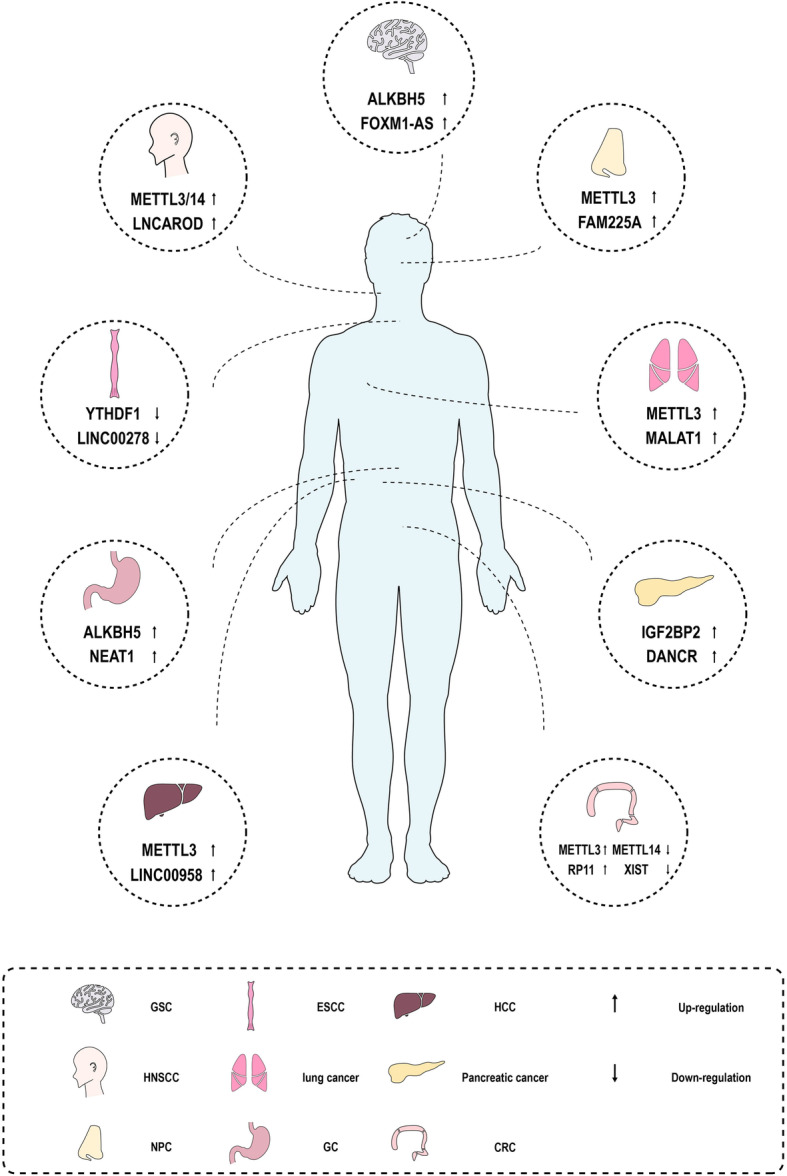
m6A methylation modifies lncRNAs to participate in tumorigenesis and metastasis in multiple cancers including GSC, HNSCC, NPC, ESCC, lung cancer, GC, HCC, pancreatic cancer and CRC
Tumor proliferation and tumorigenesis
ALKBH5 has been found upregulated in glioblastoma and prompts the proliferation of glioblastoma stem-like cells (GSCs). A lncRNA antisense to forkhead box M1 (FOXM1-AS) promotes the interaction of ALKBH5 with forkhead box M1 (FOXM1) nascent transcripts to increase FOXM1 expression and GSCs tumorigenesis [57]. LncRNA Differentiation antagonizing non-protein coding RNA (DANCR) contributes to the tumorigenesis of multiple cancers [58, 59]. IGF2BP2 serves as an m6A reader to modify DANCR and favors the oncogenicity of pancreatic cancer [60]. MALAT1, the first lncRNA to be found associated with lung cancer, possesses a triple helix structure at its 3’end [61–63]. METTL16 interacts directly with MALAT1 triple helix and promotes cancer cell proliferation [64].
Tumor invasion and metastasis
Long non-coding RNA 00958 (LINC00958) is upregulated by METTL3 and facilitates HCC cell migration and invasion by sponging miR-3619-5p [65]. METTL3/14 enhance the migration of head and neck squamous cell carcinoma (HNSCC) by upregulating lncRNA activating regulator of DKK1 (LNCAROD) [66]. METTL3-family with sequence similarity 225 member A (FAM225A)-integrin β3 (ITGB3)-FAK/PI3K/Akt axis facilitates the metastasis of nasopharyngeal cancer [67]. METTL3 mediates lncRNA RP11–138 J23.1 (RP11) or MALAT1-miR-1914-3p-Yes associated protein (YAP) axis to enhance the migration and invasion of CRC and non-small cell lung cancer (NSCLC) [68, 69]. METTL14 increases the m6A levels of XIST and suppresses the invasion of CRC [70]. ALKBH5 favors the invasion and metastasis of gastric cancer (GC) by demethylating lncRNA nuclear paraspeckle assembly transcript 1 (NEAT1) [71]. YTHDF1 restrains esophageal squamous cell carcinoma (ESCC) by interacting with long intergenic non-protein coding RNA 278 (LINC00278), but ALKBH5 harbors an opposite function [72].
m6A modification of circRNAs in cancer
CircRNAs, a novel subset of ncRNAs generated by back-splicing, play a crucial role in protein translation [73]. METTL3 and YTHDC1 are associated with the metabolism of circular RNA zinc finger protein 609 (circ-ZNF609) and promote its production [74]. Minigenes of ribosomes-circRNAs (Ribo-circRNAs) can facilitate protein translation in drosophila heads and circ-ZNF609 boosts protein translation and myoblasts cell proliferation [75, 76]. m6A methylation has been reported to affect protein translation of cricRNAs [77, 78]. m6A motifs are enriched in circRNAs, and a single m6A site is regarded as a trigger to initiate the translation of circRNAs. m6A regulators METTL3/14, FTO, YTHDF3, and initiation factor eIF4G2 are involved in m6A-driven protein translation [78]. Mammalian cells can recognize the m6A modification on circRNAs to inhibit innate immunity by abrogating immune gene activation and adjuvant activity [79].
In addition, the dysregulation of circRNAs is associated with the progression of multiple cancers, such as breast cancer, gastric cancer (GC), gallbladder cancer and cervical cancer [80–83]. YTHDC1 interacts with circRNA NOP2/Sun RNA methyltransferase 2 (circNSUN2) to facilitate its cytoplasmic export, which leads to colorectal liver metastasis by forming a circNUSN2/IGF2BP2/high mobility group AT-hook 2 (HMGA2) RNA-protein ternary complex in the cytoplasm [84]. m6A modification can be involved in the progression of GC by regulating circRNA poliovirus receptor-related 3 (circPVRL3) [85].
m6A regulators are regulated by ncRNAs in cancer
NcRNAs have the capabilities to affect m6A levels involved in multiple biological processes (Table 2). miRNAs can modulate the binding between METTL3 and its target mRNAs to participate in the reprogramming efficiency of mouse embryonic fibroblasts (MEFs) [86]. miR-149-3p inhibits adipogenesis lineage differentiation and potentiates osteogenic lineage differentiation by targeting FTO [87]. miR-1266 inhibits CRC progression by targeting FTO [88]. miR-145 suppresses the proliferation of HCC by targeting YTHDF2 [89]. Similarly, miR-33a and miR-448 suppress the proliferation of NSCLC by targeting METTL3 and eIF3a [90, 91]. METTL3 is also downregulated by miR-600, which induces the apoptosis of lung cancer [92]. miR-141 suppresses the proliferation of pancreatic cancer by forming the miR-141/IGF2BP2/P13K/Akt axis [93]. Hepatitis B X-interacting protein (HBXIP) inhibits let-7 g expression to upregulate IGF2BP2, thus leading to the formation of a positive feedback loop of HBXIP/let-7 g/IGF2BP2/HBXIP to accelerate cell proliferation in breast cancer [94]. miR-497 partially reverses transforming growth factor beta 1 (TGFβ1)-induced epithelial-mesenchymal transition (EMT) and pulmonary fibroblast proliferation through inhibiting eIF3a in alveolar epithelial cells [95].
Table 2.
NcRNAs modulate m6A regulators in cancers
| Related non-coding RNA | m6A component | Cancer | Function | Role in cancer | Regulation | References |
|---|---|---|---|---|---|---|
| miR-33a | METTL3 | NSCLC | Writers | Oncogene | Up-regulation | [90] |
| miR-600 | METTL3 | Lung cancer | Writers | Oncogene | Up-regulation | [92] |
| miRNA let-7g | METTL3 | Breast cancer | Writers | Oncogene | Up-regulation | [94] |
| miR-1266 | FTO | CRC | Erasers | Oncogene | Up-regulation | [88] |
| miR-145 | YTHDF2 | HCC | Readers | Oncogene | Up-regulation | [89] |
| miR-488 | eIF3a | NSCLC | Readers | Oncogene | Up-regulation | [91] |
| miR-141 | IGF2BP2 | Pancreatic cancer | Readers | Oncogene | Up-regulation | [93] |
| lncRNA LINC00470 | METTL3 | GC | Writers | Oncogene | Up-regulation | [97] |
| lncRNA GATA3-AS | KIAA1429 | HCC | Writers | Oncogene | Up-regulation | [104] |
| lncRNA GAS5-AS1 | ALKBH5 | Cervical cancer | Erasers | Anti-oncogene | Down-regulation | [102] |
| lncRNA GAS5 | YTHDF3 | CRC | Readers | Oncogene | Up-regulation | [103] |
| lncRNA LIN28B-AS1 | IGF2BP1 | LUAD | Readers | Oncogene | Up-regulation | [100] |
| lncRNA LINRIS | IGF2BP2 | CRC | Readers | Oncogene | Up-regulation | [101] |
| lncRNA miR503HG | HNRNPA2B1 | HCC | Readers | Oncogene | Up-regulation | [98] |
| lncRNA LINC01234 | HNRNPA2B1 | NSCLC | Readers | Oncogene | Up-regulation | [99] |
HCC hepatocellular cancer, GC gastric cancer, CRC colorectal cancer, LUAD lung adenocarcinoma, NSCLC non-small cell lung cancer
lncRNAs also regulate m6A methylation in cancer. LncRNA derived from hepatocytes (lnc-HC) interacts with HNRNPA2B1 to inhibit cholesterol metabolism in hepatocytes [96]. Long intergenic non-protein coding RNA 470 (LINC00470) interacts with METTL3 to suppress the stability of phosphatase and tensin homolog (PTEN) to facilitate GC progression [97]. LncRNA miR503 host gene (miR503HG) also interacts with HNRNPA2B1 to promote its degradation through an ubiquitin-proteasome pathway in HCC [98]. Similarly, long intergenic non-protein coding RNA 1234 (LINC01234) interacts with HNRNPA2B1 to facilitate cell proliferation and inhibit cell apoptosis in NSCLC [99]. Lin-28 homolog B antisense RNA 1 (LIN28B-AS1) interacts with IGF2BP1 to promote the proliferation and metastasis of lung adenocarcinoma (LUAD) [100]. Long intergenic Noncoding RNA for IGF2BP2 Stability (LINRIS) promotes CRC proliferation by stabilizing IGF2BP2 [101]. The antisense RNA of growth arrest special 5 (GAS5-AS1) depends on ALKBH5 to suppresses the growth and metastasis of cervical cancer [102]. Growth arrest special 5 (GAS5) can suppress YAP-mediated YTHDF3 to restrain the proliferative behavior of CRC [103]. Antisense strand of the GATA binding protein 3 gene (GATA3-AS) enhances the interaction between KIAA1429 and GATA binding protein 3 (GATA3) pre-mRNA, leading to the formation of the GATA3-AS/KIAA1429/GATA3 axis in HCC [104].
Clinical application of m6A methylation in cancer
m6A methylation serves as new biomarkers for diagnosis and prognosis in cancer. m6A regulators METTL3, YTHDC2 and HNRNPC are used to predict the prognosis in patients with HNSCC [105]. Upregulated METTL3/FTO or downregulated YTHDF2 and METTL14 can indicate a poor survival in GC, CRC, and HCC [48, 70, 106]. Low expression of METTL14 is associated with tumor differentiation, clinical stage, and microvascular invasion [48]. Low expression of ALKBH5 or FTO predicts an unfavorable marker in lung cancer and HCC [107, 108]. IGF2BP2 is considered as a prognostic marker in pancreatic cancer, esophagogastric junction adenocarcinoma and CRC [60, 109, 110].
m6A methylation also participates in drug resistance and cancer treatment. METTL3 stabilizes YAP and Rho GTPase activating protein 5 (ARHGAP5) to induce cisplatin resistance in NSCLC and in GC [69, 111]. HNRNPA2B1 is overexpressed in tamoxifen-resistant breast cancer and reduces 4-hydroxytamoxifen sensitivity [112]. In addition to METTL3 and METTL14, FTO and YTHDF2 are overexpressed in AML [26, 27, 113, 114]. A recent study shows that FTO inhibitor (FB23) and its derivative (FB23–2) promote myeloid differentiation and apoptosis in AML by targeting FTO [115]. m6A methylation is also involved in estimating tumor microenvironment and TME infiltration characterization so as to provide insights into an effective immunotherapy for cancer [116]. YTHDF2 is correlated with inflammation infiltration, vascular reconstruction and distant metastasis and predicts a poor prognosis in HCC [117].
In summary, the role of m6A modification in clinical application has been widely validated. As for the core members of m6A methylation, METTL3/14 exert their functions in many biological processes. METTL3/14 can be regarded as the most important and promising m6A regulator and arouse our attention about their modifications on ncRNAs and the clinical application in cancer diagnosis.
Conclusions and perspectives
Accumulating studies have been focused on how m6A methylation modifies the stability, splicing and translation of ncRNAs or ncRNAs regulate m6A regulators in cancer. The interaction between m6A methylation and ncRNAs can impact the different life activities including cancer cell proliferation, invasion and metastasis. As for the clinical application of m6A methylation, they can be regarded as the potential targets for cancer diagnosis, prognosis and treatment. The latest findings show that lncRNA long intergenic non-protein coding RNA 266–1 (LINC00266–1) interacts with IGF2BP1 by encoding a 71-amino acid peptide, named RNA-binding regulatory peptide, thereby promoting tumorigenesis [118]. However, the specific binding sites between m6A methylation and ncRNAs need be further investigated.
Acknowledgements
None.
Abbreviations
- 3′-UTR
3′ -untranslated region
- ALKBH5
alkB homologue 5
- AML
Acute myeloid leukemia
- ARHGAP5
Rho GTPase activating protein 5
- circRNAs
circular RNAs
- circNSUN2
circular RNA NOP2/Sun RNA methyltransferase 2
- circPVRL3
circular RNA poliovirus receptor-related 3
- circ-ZNF609
circular RNA zinc finger protein 609
- CRC
Colorectal cancer
- CSC
Cigarette smoke condensate
- DANCR
Differentiation antagonizing non-protein coding RNA
- DGCR8
DiGeorge syndrome critical region 8
- eIF3
eukaryotic translation initiation factor 3
- EMT
Epithelial-mesenchymal transition
- ESCC
Esophageal squamous cell carcinoma
- FAK
Focal adhesion kinase
- FGFR3
Fibroblast growth factor receptor 3
- FAM225A
Family with sequence similarity 225 member A
- FTO
Fat mass and obesity-associated protein
- FOXM1
Forkhead box M1
- FOXM1-AS
antisense to forkhead box M1
- GAS5
Growth arrest special 5
- GAS5-AS1
the antisense RNA of GAS5
- GATA3
GATA binding protein 3
- GATA3-AS
Antisense strand of the GATA binding protein 3 gene
- GC
Gastric cancer
- GSCs
Glioblastoma stem-like cells
- HBXIP
Hepatitis B X-interacting protein
- HCC
Hepatocellular carcinoma
- HIF-1α
Hypoxia-inducible factor 1-alpha
- HMGA2
High mobility group AT-hook 2
- HNRNP
Heterogeneous nuclear ribonucleo protein
- HNSCC
Head and neck squamous cell carcinoma
- IGF2BP1/2/3
Insulin-like growth factor-2 mRNA-binding proteins 1/2/3
- ITGB3
Integrin β3
- LINC00266–1
Long intergenic non-protein coding RNA 266–1
- LINC00278
Long intergenic non-protein coding RNA 278
- LINC00470
Long intergenic non-protein coding RNA 470
- LINC00958
Long non-coding RNA 00958
- LINC01234
Long intergenic non-protein coding RNA 1234
- LIN28B-AS1
Lin-28 homolog B antisense RNA 1
- LINRIS
Long intergenic Noncoding RNA for IGF2BP2 Stability
- LNCAROD
lncRNA activating regulator of DKK1
- lnc-HC
LncRNA derived from hepatocytes
- lncRNAs
Long non-coding RNAs
- LUAD
Lung adenocarcinoma
- m6A
N6-methyladenosine
- m6A-seq
N6-methyladenosine-sequensing
- MALAT1
Metastasis-associated lung adenocarcinoma transcript-1
- MEFs
Mouse embryonic fibroblasts
- METTL3/14/16
Methyltransferase-like 3/14/16
- miR503HG
miR503 host gene
- miRNAs
Micro RNAs
- mRNA
Messenger RNA
- ncRNAs
Noncoding RNAs
- NEAT1
Nuclear paraspeckle assembly transcript 1
- NSCLC
Non-small cell lung cancer
- PDAC
Pancreatic ductal adenocarcinoma
- PTEN
Phosphatase and tensin homolog
- RBM15/15B
RNA-binding motif protein 15/15B
- Ribo-circRNAs
Ribosomes-circRNAs
- RP11
RP11–138 J23.1
- SAM
S-adenosylmethionine
- STING
Stimulator of interferon genes
- TGFβ1
Transforming growth factor beta 1
- WTAP
Wilms tumor 1-associated protein
- XIST
X-inactive specific transcript
- YAP
Yes associated protein
- YTH
YT521-B homology
- YTHDC1/2
YTH domain-containing proteins 1/2
- YTHDF1/2/3
YTH domain family 1/2/3
Authors’ contributions
JZ and JSZ designed this study and YCY drafted the manuscript. YCY and XYC collected the data and conducted the picture processing. JZ revised the paper and all authors read and approved the final manuscript.
Funding
Our work was supported by the grants from National Natural Science Foundation of China (No. 81873143) and Double-Hundred Talent Plan of Shanghai Jiao Tong University School of Medicine (No. 20191831).
Availability of data and materials
All data generated or analyzed during this study are included in this published article and its additional files.
Ethics approval and consent to participate
None.
Consent for publication
Consent for publication has been obtained from the authors.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
You-Cai Yi, Email: 15521120969@163.com.
Xiao-Yu Chen, Email: xiaoyu643@163.com.
Jing Zhang, Email: jing5522724@vip.163.com.
Jin-Shui Zhu, Email: zhujs1803@163.com.
References
- 1.Boccaletto P, Machnicka MA, Purta E, Piatkowski P, Baginski B, Wirecki TK, et al. MODOMICS: a database of RNA modification pathways. 2017 update. Nucleic Acids Res. 2018;46:D303–D307. doi: 10.1093/nar/gkx1030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ping X-L, Sun B-F, Wang L, Xiao W, Yang X, Wang W-J, et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 2014;24:177–189. doi: 10.1038/cr.2014.3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature. 2012;485:201–206. doi: 10.1038/nature11112. [DOI] [PubMed] [Google Scholar]
- 4.Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR. Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell. 2012;149:1635–1646. doi: 10.1016/j.cell.2012.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Fustin J-M, Doi M, Yamaguchi Y, Hida H, Nishimura S, Yoshida M, et al. RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell. 2013;155:793–806. doi: 10.1016/j.cell.2013.10.026. [DOI] [PubMed] [Google Scholar]
- 6.Zhao BS, Roundtree IA, He C. Post-transcriptional gene regulation by mRNA modifications. Nat Rev Mol Cell Biol. 2017;18:31–42. doi: 10.1038/nrm.2016.132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Edens BM, Vissers C, Su J, Arumugam S, Xu Z, Shi H, et al. FMRP Modulates Neural Differentiation through m6A-Dependent mRNA Nuclear Export. Cell Rep. 2019;28:845–854.e5. doi: 10.1016/j.celrep.2019.06.072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Engel M, Eggert C, Kaplick PM, Eder M, Röh S, Tietze L, et al. The Role of m6A/m-RNA Methylation in Stress Response Regulation. Neuron. 2018;99:389. doi: 10.1016/j.neuron.2018.07.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Yu R, Li Q, Feng Z, Cai L, Xu Q. m6A Reader YTHDF2 Regulates LPS-Induced Inflammatory Response. Int J Mol Sci. 2019;20:1323. doi: 10.3390/ijms20061323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chen X-Y, Zhang J, Zhu J-S. The role of m6A RNA methylation in human cancer. Mol Cancer. 2019;18:103. doi: 10.1186/s12943-019-1033-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424. doi: 10.3322/caac.21492. [DOI] [PubMed] [Google Scholar]
- 12.Haussmann IU, Bodi Z, Sanchez-Moran E, Mongan NP, Archer N, Fray RG, et al. m6A potentiates Sxl alternative pre-mRNA splicing for robust Drosophila sex determination. Nature. 2016;540:301–304. doi: 10.1038/nature20577. [DOI] [PubMed] [Google Scholar]
- 13.Fleming AM, Nguyen NLB, Burrows CJ. Colocalization of m6A and G-Quadruplex-Forming Sequences in Viral RNA (HIV, Zika, Hepatitis B, and SV40) Suggests Topological Control of Adenosine N6-Methylation. ACS Cent Sci. 2019;5:218–228. doi: 10.1021/acscentsci.8b00963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Shah JC, Clancy MJ. IME4, a gene that mediates MAT and nutritional control of meiosis in Saccharomyces cerevisiae. Mol Cell Biol. 1992;12:1078–1086. doi: 10.1128/mcb.12.3.1078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Zhong S, Li H, Bodi Z, Button J, Vespa L, Herzog M, et al. MTA is an Arabidopsis messenger RNA adenosine methylase and interacts with a homolog of a sex-specific splicing factor. Plant Cell. 2008;20:1278–1288. doi: 10.1105/tpc.108.058883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Cesarini V, Silvestris DA, Tassinari V, Tomaselli S, Alon S, Eisenberg E, et al. ADAR2/miR-589-3p axis controls glioblastoma cell migration/invasion. Nucleic Acids Res. 2018;46:2045–2059. doi: 10.1093/nar/gkx1257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Luo H, Zhu G, Xu J, Lai Q, Yan B, Guo Y, et al. HOTTIP lncRNA promotes hematopoietic stem cell self-renewal leading to AML-like disease in mice. Cancer Cell. 2019;36:645–659. doi: 10.1016/j.ccell.2019.10.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Li J, Huang C, Zou Y, Ye J, Yu J, Gui Y. CircTLK1 promotes the proliferation and metastasis of renal cell carcinoma by sponging miR-136-5p. Mol Cancer. 2020;19:103. doi: 10.1186/s12943-020-01225-2. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 19.Schumann U, Shafik A, Preiss T. METTL3 Gains R/W Access to the Epitranscriptome. Mol Cell. 2016;62:323–324. doi: 10.1016/j.molcel.2016.04.024. [DOI] [PubMed] [Google Scholar]
- 20.Liu J, Yue Y, Han D, Wang X, Fu Y, Zhang L, et al. A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat Chem Biol. 2014;10:93–95. doi: 10.1038/nchembio.1432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Schwartz S, Mumbach MR, Jovanovic M, Wang T, Maciag K, Bushkin GG, et al. Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5’ sites. Cell Rep. 2014;8:284–296. doi: 10.1016/j.celrep.2014.05.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Koh CWQ, Goh YT, Goh WSS. Atlas of quantitative single-base-resolution N6-methyl-adenine methylomes. Nat Commun. 2019;10:5636. doi: 10.1038/s41467-019-13561-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Meyer KD, Jaffrey SR. Rethinking m6A Readers, Writers, and Erasers. Annu Rev Cell Dev Biol. 2017;33:319–342. doi: 10.1146/annurev-cellbio-100616-060758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Zhong X, Yu J, Frazier K, Weng X, Li Y, Cham CM, et al. Circadian Clock Regulation of Hepatic Lipid Metabolism by Modulation of m6A mRNA Methylation. Cell Rep. 2018;25:1816–1828.e4. doi: 10.1016/j.celrep.2018.10.068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Cheng Y, Luo H, Izzo F, Pickering BF, Nguyen D, Myers R, et al. m6A RNA Methylation Maintains Hematopoietic Stem Cell Identity and Symmetric Commitment. Cell Rep. 2019;28:1703–1716.e6. doi: 10.1016/j.celrep.2019.07.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Weng H, Huang H, Wu H, Qin X, Zhao BS, Dong L, et al. METTL14 Inhibits Hematopoietic Stem/Progenitor Differentiation and Promotes Leukemogenesis via mRNA m6A Modification. Cell Stem Cell. 2018;22:191–205.e9. doi: 10.1016/j.stem.2017.11.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Vu LP, Pickering BF, Cheng Y, Zaccara S, Nguyen D, Minuesa G, et al. The N6-methyladenosine (m6A)-forming enzyme METTL3 controls myeloid differentiation of normal hematopoietic and leukemia cells. Nat Med. 2017;23:1369–1376. doi: 10.1038/nm.4416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Pendleton KE, Chen B, Liu K, Hunter OV, Xie Y, Tu BP, et al. The U6 snRNA m6A Methyltransferase METTL16 Regulates SAM Synthetase Intron Retention. Cell. 2017;169:824–835.e14. doi: 10.1016/j.cell.2017.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Zheng G, Dahl JA, Niu Y, Fedorcsak P, Huang C-M, Li CJ, et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell. 2013;49:18–29. doi: 10.1016/j.molcel.2012.10.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Jia G, Fu Y, Zhao X, Dai Q, Zheng G, Yang Y, et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol. 2011;7:885–887. doi: 10.1038/nchembio.687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Gerken T, Girard CA, Tung Y-CL, Webby CJ, Saudek V, Hewitson KS, et al. The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science. 2007;318:1469–1472. doi: 10.1126/science.1151710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Tang C, Klukovich R, Peng H, Wang Z, Yu T, Zhang Y, et al. ALKBH5-dependent m6A demethylation controls splicing and stability of long 3’-UTR mRNAs in male germ cells. Proc Natl Acad Sci U S A. 2018;115:E325–E333. doi: 10.1073/pnas.1717794115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Shi H, Wei J, He C. Where, When, and How: Context-Dependent Functions of RNA Methylation Writers, Readers, and Erasers. Mol Cell. 2019;74:640–650. doi: 10.1016/j.molcel.2019.04.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, et al. Recognition of RNA N6-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018;20:285–295. doi: 10.1038/s41556-018-0045-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Wu R, Li A, Sun B, Sun J-G, Zhang J, Zhang T, et al. A novel m6A reader Prrc2a controls oligodendroglial specification and myelination. Cell Res. 2019;29:23–41. doi: 10.1038/s41422-018-0113-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zhuang M, Li X, Zhu J, Zhang J, Niu F, Liang F, et al. The m6A reader YTHDF1 regulates axon guidance through translational control of Robo3.1 expression. Nucleic Acids Res. 2019;47:4765–4777. doi: 10.1093/nar/gkz157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wang L, Wen M, Cao X. Nuclear hnRNPA2B1 initiates and amplifies the innate immune response to DNA viruses. Science. 2019;365. 10.1126/science.aav0758. [DOI] [PubMed]
- 38.Rahmanian S, Murad R, Breschi A, Zeng W, Mackiewicz M, Williams B, et al. Dynamics of microRNA expression during mouse prenatal development. Genome Res. 2019;29:1900–1909. doi: 10.1101/gr.248997.119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wu C-J, Cho S, Huang H-Y, Lu C-H, Russ J, Cruz LO, et al. MiR-23~27~24-mediated control of humoral immunity reveals a TOX-driven regulatory circuit in follicular helper T cell differentiation. Sci Adv. 2019;5:eaaw1715. doi: 10.1126/sciadv.aaw1715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Cho YK, Son Y, Kim S-N, Song H-D, Kim M, Park J-H, et al. MicroRNA-10a-5p regulates macrophage polarization and promotes therapeutic adipose tissue remodeling. Mol Metab. 2019;29:86–98. doi: 10.1016/j.molmet.2019.08.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Cen B, Lang JD, Du Y, Wei J, Xiong Y, Bradley N, et al. Prostaglandin E2 induces MIR675-5p to promote colorectal tumor metastasis via modulation of p53 expression. Gastroenterology. 2019;158:971–984.e10. doi: 10.1053/j.gastro.2019.11.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Alarcón CR, Lee H, Goodarzi H, Halberg N, Tavazoie SF. N6-methyladenosine marks primary microRNAs for processing. Nature. 2015;519:482–485. doi: 10.1038/nature14281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Alarcón CR, Goodarzi H, Lee H, Liu X, Tavazoie S, Tavazoie SF. HNRNPA2B1 Is a Mediator of m(6)A-Dependent Nuclear RNA Processing Events. Cell. 2015;162:1299–1308. doi: 10.1016/j.cell.2015.08.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Mi B, Xiong Y, Yan C, Chen L, Xue H, Panayi AC, et al. Methyltransferase-like 3-mediated N6-methyladenosine modification of miR-7212-5p drives osteoblast differentiation and fracture healing. J Cell Mol Med. 2020;24:6385–6396. doi: 10.1111/jcmm.15284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Zhang J, Bai R, Li M, Ye H, Wu C, Wang C, et al. Excessive miR-25-3p maturation via N6-methyladenosine stimulated by cigarette smoke promotes pancreatic cancer progression. Nat Commun. 2019;10:1858. doi: 10.1038/s41467-019-09712-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Han J, Wang J-Z, Yang X, Yu H, Zhou R, Lu H-C, et al. METTL3 promote tumor proliferation of bladder cancer by accelerating pri-miR221/222 maturation in m6A-dependent manner. Mol Cancer. 2019;18:110. doi: 10.1186/s12943-019-1036-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Gu S, Sun D, Dai H, Zhang Z. N6-methyladenosine mediates the cellular proliferation and apoptosis via microRNAs in arsenite-transformed cells. Toxicol Lett. 2018;292:1–11. doi: 10.1016/j.toxlet.2018.04.018. [DOI] [PubMed] [Google Scholar]
- 48.Ma J-Z, Yang F, Zhou C-C, Liu F, Yuan J-H, Wang F, et al. METTL14 suppresses the metastatic potential of hepatocellular carcinoma by modulating N6 -methyladenosine-dependent primary MicroRNA processing. Hepatol Baltim Md. 2017;65:529–543. doi: 10.1002/hep.28885. [DOI] [PubMed] [Google Scholar]
- 49.Peng W, Li J, Chen R, Gu Q, Yang P, Qian W, et al. Upregulated METTL3 promotes metastasis of colorectal Cancer via miR-1246/SPRED2/MAPK signaling pathway. J Exp Clin Cancer Res CR. 2019;38:393. doi: 10.1186/s13046-019-1408-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Wang H, Deng Q, Lv Z, Ling Y, Hou X, Chen Z, et al. N6-methyladenosine induced miR-143-3p promotes the brain metastasis of lung cancer via regulation of VASH1. Mol Cancer. 2019;18:181. doi: 10.1186/s12943-019-1108-x. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 51.Chu C, Zhang QC, da Rocha ST, Flynn RA, Bharadwaj M, Calabrese JM, et al. Systematic discovery of Xist RNA binding proteins. Cell. 2015;161:404–416. doi: 10.1016/j.cell.2015.03.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Monfort A, Di Minin G, Postlmayr A, Freimann R, Arieti F, Thore S, et al. Identification of Spen as a Crucial Factor for Xist Function through Forward Genetic Screening in Haploid Embryonic Stem Cells. Cell Rep. 2015;12:554–561. doi: 10.1016/j.celrep.2015.06.067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Patil DP, Chen C-K, Pickering BF, Chow A, Jackson C, Guttman M, et al. m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature. 2016;537:369–373. doi: 10.1038/nature19342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Nesterova TB, Wei G, Coker H, Pintacuda G, Bowness JS, Zhang T, et al. Systematic allelic analysis defines the interplay of key pathways in X chromosome inactivation. Nat Commun. 2019;10:3129. doi: 10.1038/s41467-019-11171-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Liu J, Zhang X, Chen K, Cheng Y, Liu S, Xia M, et al. CCR7 Chemokine Receptor-Inducible lnc-Dpf3 Restrains Dendritic Cell Migration by Inhibiting HIF-1α-Mediated Glycolysis. Immunity. 2019;50:600–615.e15. doi: 10.1016/j.immuni.2019.01.021. [DOI] [PubMed] [Google Scholar]
- 56.Liu P, Zhang B, Chen Z, He Y, Du Y, Liu Y, et al. m6A-induced lncRNA MALAT1 aggravates renal fibrogenesis in obstructive nephropathy through the miR-145/FAK pathway. Aging. 2020;12:5280–5299. doi: 10.18632/aging.102950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Zhang S, Zhao BS, Zhou A, Lin K, Zheng S, Lu Z, et al. m6A Demethylase ALKBH5 Maintains Tumorigenicity of Glioblastoma Stem-like Cells by Sustaining FOXM1 Expression and Cell Proliferation Program. Cancer Cell. 2017;31:591–606.e6. doi: 10.1016/j.ccell.2017.02.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Zhang K-J, Tan X-L, Guo L. The long non-coding RNA DANCR regulates the inflammatory phenotype of breast cancer cells and promotes breast cancer progression via EZH2-dependent suppression of SOCS3 transcription. Mol Oncol. 2020;14:309–328. doi: 10.1002/1878-0261.12622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Lin X, Yang F, Qi X, Li Q, Wang D, Yi T, et al. LncRNA DANCR promotes tumor growth and angiogenesis in ovarian cancer through direct targeting of miR-145. Mol Carcinog. 2019;58:2286–2296. doi: 10.1002/mc.23117. [DOI] [PubMed] [Google Scholar]
- 60.Hu X, Peng W-X, Zhou H, Jiang J, Zhou X, Huang D, et al. IGF2BP2 regulates DANCR by serving as an N6-methyladenosine reader. Cell Death Differ. 2019;27:1782–1794. doi: 10.1038/s41418-019-0461-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Ji P, Diederichs S, Wang W, Böing S, Metzger R, Schneider PM, et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene. 2003;22:8031–8041. doi: 10.1038/sj.onc.1206928. [DOI] [PubMed] [Google Scholar]
- 62.Amodio N, Raimondi L, Juli G, Stamato MA, Caracciolo D, Tagliaferri P, et al. MALAT1: a druggable long non-coding RNA for targeted anti-cancer approaches. J Hematol Oncol. 2018;11:63. doi: 10.1186/s13045-018-0606-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Brown JA, Bulkley D, Wang J, Valenstein ML, Yario TA, Steitz TA, et al. Structural insights into the stabilization of MALAT1 noncoding RNA by a bipartite triple helix. Nat Struct Mol Biol. 2014;21:633–640. doi: 10.1038/nsmb.2844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Brown JA, Kinzig CG, DeGregorio SJ, Steitz JA. Methyltransferase-like protein 16 binds the 3’-terminal triple helix of MALAT1 long noncoding RNA. Proc Natl Acad Sci U S A. 2016;113:14013–14018. doi: 10.1073/pnas.1614759113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Zuo X, Chen Z, Gao W, Zhang Y, Wang J, Wang J, et al. M6A-mediated upregulation of LINC00958 increases lipogenesis and acts as a nanotherapeutic target in hepatocellular carcinoma. J Hematol Oncol. 2020;13:5. doi: 10.1186/s13045-019-0839-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Ban Y, Tan P, Cai J, Li J, Hu M, Zhou Y, et al. LNCAROD is stabilized by m6A methylation and promotes cancer progression via forming a ternary complex with HSPA1A and YBX1 in head and neck squamous cell carcinoma. Mol Oncol. 2020;14:1282–1296. doi: 10.1002/1878-0261.12676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Zheng Z-Q, Li Z-X, Zhou G-Q, Lin L, Zhang L-L, Lv J-W, et al. Long Noncoding RNA FAM225A Promotes Nasopharyngeal Carcinoma Tumorigenesis and Metastasis by Acting as ceRNA to Sponge miR-590-3p/miR-1275 and Upregulate ITGB3. Cancer Res. 2019;79:4612–4626. doi: 10.1158/0008-5472.CAN-19-0799. [DOI] [PubMed] [Google Scholar]
- 68.Wu Y, Yang X, Chen Z, Tian L, Jiang G, Chen F, et al. m6A-induced lncRNA RP11 triggers the dissemination of colorectal cancer cells via upregulation of Zeb1. Mol Cancer. 2019;18:87. doi: 10.1186/s12943-019-1014-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Jin D, Guo J, Wu Y, Du J, Yang L, Wang X, et al. m6A mRNA methylation initiated by METTL3 directly promotes YAP translation and increases YAP activity by regulating the MALAT1-miR-1914-3p-YAP axis to induce NSCLC drug resistance and metastasis. J Hematol Oncol. 2019;12:135. doi: 10.1186/s13045-019-0830-6. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 70.Yang X, Zhang S, He C, Xue P, Zhang L, He Z, et al. METTL14 suppresses proliferation and metastasis of colorectal cancer by down-regulating oncogenic long non-coding RNA XIST. Mol Cancer. 2020;19:46. doi: 10.1186/s12943-020-1146-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Zhang J, Guo S, Piao H-Y, Wang Y, Wu Y, Meng X-Y, et al. ALKBH5 promotes invasion and metastasis of gastric cancer by decreasing methylation of the lncRNA NEAT1. J Physiol Biochem. 2019;75:379–389. doi: 10.1007/s13105-019-00690-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Wu S, Zhang L, Deng J, Guo B, Li F, Wang Y, et al. A novel micropeptide encoded by Y-Linked LINC00278 links cigarette smoking and AR signaling in male esophageal squamous cell carcinoma. Cancer Res. 2020;80:2790–2803. doi: 10.1158/0008-5472.CAN-19-3440. [DOI] [PubMed] [Google Scholar]
- 73.Li X, Yang L, Chen L-L. The Biogenesis, Functions, and Challenges of Circular RNAs. Mol Cell. 2018;71:428–442. doi: 10.1016/j.molcel.2018.06.034. [DOI] [PubMed] [Google Scholar]
- 74.Di Timoteo G, Dattilo D, Centrón-Broco A, Colantoni A, Guarnacci M, Rossi F, et al. Modulation of circRNA Metabolism by m6A Modification. Cell Rep. 2020;31:107641. doi: 10.1016/j.celrep.2020.107641. [DOI] [PubMed] [Google Scholar]
- 75.Pamudurti NR, Bartok O, Jens M, Ashwal-Fluss R, Stottmeister C, Ruhe L, et al. Translation of CircRNAs. Mol Cell. 2017;66:9–21.e7. doi: 10.1016/j.molcel.2017.02.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Legnini I, Di Timoteo G, Rossi F, Morlando M, Briganti F, Sthandier O, et al. Circ-ZNF609 Is a Circular RNA that Can Be Translated and Functions in Myogenesis. Mol Cell. 2017;66:22–37.e9. doi: 10.1016/j.molcel.2017.02.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Zhou C, Molinie B, Daneshvar K, Pondick JV, Wang J, Van Wittenberghe N, et al. Genome-Wide Maps of m6A circRNAs Identify Widespread and Cell-Type-Specific Methylation Patterns that Are Distinct from mRNAs. Cell Rep. 2017;20:2262–2276. doi: 10.1016/j.celrep.2017.08.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Yang Y, Fan X, Mao M, Song X, Wu P, Zhang Y, et al. Extensive translation of circular RNAs driven by N6-methyladenosine. Cell Res. 2017;27:626–641. doi: 10.1038/cr.2017.31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Chen YG, Chen R, Ahmad S, Verma R, Kasturi SP, Amaya L, et al. N6-Methyladenosine Modification Controls Circular RNA Immunity. Mol Cell. 2019;76:96–109.e9. doi: 10.1016/j.molcel.2019.07.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Du WW, Yang W, Li X, Awan FM, Yang Z, Fang L, et al. A circular RNA circ-DNMT1 enhances breast cancer progression by activating autophagy. Oncogene. 2018;37:5829–5842. doi: 10.1038/s41388-018-0369-y. [DOI] [PubMed] [Google Scholar]
- 81.Ding L, Zhao Y, Dang S, Wang Y, Li X, Yu X, et al. Circular RNA circ-DONSON facilitates gastric cancer growth and invasion via NURF complex dependent activation of transcription factor SOX4. Mol Cancer. 2019;18:45. doi: 10.1186/s12943-019-1006-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Wang S, Zhang Y, Cai Q, Ma M, Jin LY, Weng M, et al. Circular RNA FOXP1 promotes tumor progression and Warburg effect in gallbladder cancer by regulating PKLR expression. Mol Cancer. 2019;18:145. doi: 10.1186/s12943-019-1078-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Zhao J, Lee EE, Kim J, Yang R, Chamseddin B, Ni C, et al. Transforming activity of an oncoprotein-encoding circular RNA from human papillomavirus. Nat Commun. 2019;10:2300. doi: 10.1038/s41467-019-10246-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Chen R-X, Chen X, Xia L-P, Zhang J-X, Pan Z-Z, Ma X-D, et al. N6-methyladenosine modification of circNSUN2 facilitates cytoplasmic export and stabilizes HMGA2 to promote colorectal liver metastasis. Nat Commun. 2019;10:4695. doi: 10.1038/s41467-019-12651-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Sun H-D, Xu Z-P, Sun Z-Q, Zhu B, Wang Q, Zhou J, et al. Down-regulation of circPVRL3 promotes the proliferation and migration of gastric cancer cells. Sci Rep. 2018;8:10111. doi: 10.1038/s41598-018-27837-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Chen T, Hao Y-J, Zhang Y, Li M-M, Wang M, Han W, et al. m(6)A RNA methylation is regulated by microRNAs and promotes reprogramming to pluripotency. Cell Stem Cell. 2015;16:289–301. doi: 10.1016/j.stem.2015.01.016. [DOI] [PubMed] [Google Scholar]
- 87.Li Y, Yang F, Gao M, Gong R, Jin M, Liu T, et al. miR-149-3p Regulates the Switch between Adipogenic and Osteogenic Differentiation of BMSCs by Targeting FTO. Mol Ther Nucleic Acids. 2019;17:590–600. doi: 10.1016/j.omtn.2019.06.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Shen X-P, Ling X, Lu H, Zhou C-X, Zhang J-K, Yu Q. Low expression of microRNA-1266 promotes colorectal cancer progression via targeting FTO. Eur Rev Med Pharmacol Sci. 2018;22:8220–8226. doi: 10.26355/eurrev_201812_16516. [DOI] [PubMed] [Google Scholar]
- 89.Yang Z, Li J, Feng G, Gao S, Wang Y, Zhang S, et al. MicroRNA-145 Modulates N6-Methyladenosine Levels by Targeting the 3’-Untranslated mRNA Region of the N6-Methyladenosine Binding YTH Domain Family 2 Protein. J Biol Chem. 2017;292:3614–3623. doi: 10.1074/jbc.M116.749689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Du M, Zhang Y, Mao Y, Mou J, Zhao J, Xue Q, et al. MiR-33a suppresses proliferation of NSCLC cells via targeting METTL3 mRNA. Biochem Biophys Res Commun. 2017;482:582–589. doi: 10.1016/j.bbrc.2016.11.077. [DOI] [PubMed] [Google Scholar]
- 91.Fang C, Chen Y-X, Wu N-Y, Yin J-Y, Li X-P, Huang H-S, et al. MiR-488 inhibits proliferation and cisplatin sensibility in non-small-cell lung cancer (NSCLC) cells by activating the eIF3a-mediated NER signaling pathway. Sci Rep. 2017;7:40384. doi: 10.1038/srep40384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Wei W, Huo B, Shi X. miR-600 inhibits lung cancer via downregulating the expression of METTL3. Cancer Manag Res. 2019;11:1177–1187. doi: 10.2147/CMAR.S181058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Xu X, Yu Y, Zong K, Lv P, Gu Y. Up-regulation of IGF2BP2 by multiple mechanisms in pancreatic cancer promotes cancer proliferation by activating the PI3K/Akt signaling pathway. J Exp Clin Cancer Res CR. 2019;38:497. doi: 10.1186/s13046-019-1470-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Cai X, Wang X, Cao C, Gao Y, Zhang S, Yang Z, et al. HBXIP-elevated methyltransferase METTL3 promotes the progression of breast cancer via inhibiting tumor suppressor let-7g. Cancer Lett. 2018;415:11–19. doi: 10.1016/j.canlet.2017.11.018. [DOI] [PubMed] [Google Scholar]
- 95.Guo R, Lv Y, Ouyang Y, Liu S, Li D. The Role of miR-497/EIF3A Axis in TGFβ1-Induced Epithelial-Mesenchymal Transition and Extracellular Matrix in Rat Alveolar Epithelial Cells and Pulmonary Fibroblasts. J Cell Biochem. 2017;118:3401–3408. doi: 10.1002/jcb.25997. [DOI] [PubMed] [Google Scholar]
- 96.Lan X, Yan J, Ren J, Zhong B, Li J, Li Y, et al. A novel long noncoding RNA Lnc-HC binds hnRNPA2B1 to regulate expressions of Cyp7a1 and Abca1 in hepatocytic cholesterol metabolism. Hepatol Baltim Md. 2016;64:58–72. doi: 10.1002/hep.28391. [DOI] [PubMed] [Google Scholar]
- 97.Yan J, Huang X, Zhang X, Chen Z, Ye C, Xiang W, et al. LncRNA LINC00470 promotes the degradation of PTEN mRNA to facilitate malignant behavior in gastric cancer cells. Biochem Biophys Res Commun. 2019;521:887–893. doi: 10.1016/j.bbrc.2019.11.016. [DOI] [PubMed] [Google Scholar]
- 98.Wang H, Liang L, Dong Q, Huan L, He J, Li B, et al. Long noncoding RNA miR503HG, a prognostic indicator, inhibits tumor metastasis by regulating the HNRNPA2B1/NF-κB pathway in hepatocellular carcinoma. Theranostics. 2018;8:2814–2829. doi: 10.7150/thno.23012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Chen Z, Chen X, Lei T, Gu Y, Gu J, Huang J, et al. Integrative Analysis of NSCLC Identifies LINC01234 as an Oncogenic lncRNA that Interacts with HNRNPA2B1 and Regulates miR-106b Biogenesis. Mol Ther J Am Soc Gene Ther. 2020;28:1479–1493. doi: 10.1016/j.ymthe.2020.03.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Wang C, Gu Y, Zhang E, Zhang K, Qin N, Dai J, et al. A cancer-testis non-coding RNA LIN28B-AS1 activates driver gene LIN28B by interacting with IGF2BP1 in lung adenocarcinoma. Oncogene. 2019;38:1611–1624. doi: 10.1038/s41388-018-0548-x. [DOI] [PubMed] [Google Scholar]
- 101.Wang Y, Lu J-H, Wu Q-N, Jin Y, Wang D-S, Chen Y-X, et al. LncRNA LINRIS stabilizes IGF2BP2 and promotes the aerobic glycolysis in colorectal cancer. Mol Cancer. 2019;18:174. doi: 10.1186/s12943-019-1105-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Wang X, Zhang J, Wang Y. Long noncoding RNA GAS5-AS1 suppresses growth and metastasis of cervical cancer by increasing GAS5 stability. Am J Transl Res. 2019;11:4909–4921. [PMC free article] [PubMed] [Google Scholar]
- 103.Ni W, Yao S, Zhou Y, Liu Y, Huang P, Zhou A, et al. Long noncoding RNA GAS5 inhibits progression of colorectal cancer by interacting with and triggering YAP phosphorylation and degradation and is negatively regulated by the m6A reader YTHDF3. Mol Cancer. 2019;18:143. doi: 10.1186/s12943-019-1079-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Lan T, Li H, Zhang D, Xu L, Liu H, Hao X, et al. KIAA1429 contributes to liver cancer progression through N6-methyladenosine-dependent post-transcriptional modification of GATA3. Mol Cancer. 2019;18:186. doi: 10.1186/s12943-019-1106-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Zhao X, Cui L. Development and validation of a m6A RNA methylation regulators-based signature for predicting the prognosis of head and neck squamous cell carcinoma. Am J Cancer Res. 2019;9:2156–2169. [PMC free article] [PubMed] [Google Scholar]
- 106.Liu T, Yang S, Sui J, Xu S-Y, Cheng Y-P, Shen B, et al. Dysregulated N6-methyladenosine methylation writer METTL3 contributes to the proliferation and migration of gastric cancer. J Cell Physiol. 2020;235:548–562. doi: 10.1002/jcp.28994. [DOI] [PubMed] [Google Scholar]
- 107.Jin D, Guo J, Wu Y, Yang L, Wang X, Du J, et al. m6A demethylase ALKBH5 inhibits tumor growth and metastasis by reducing YTHDFs-mediated YAP expression and inhibiting miR-107/LATS2-mediated YAP activity in NSCLC. Mol Cancer. 2020;19:40. doi: 10.1186/s12943-020-01161-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Liu X, Liu J, Xiao W, Zeng Q, Bo H, Zhu Y, et al. SIRT1 regulates N6 -methyladenosine RNA modification in hepatocarcinogenesis by inducing RANBP2-dependent FTO SUMOylation. Hepatol Baltim Md. 2020. 10.1002/hep.31222. [DOI] [PubMed]
- 109.Wu X-L, Lu R-Y, Wang L-K, Wang Y-Y, Dai Y-J, Wang C-Y, et al. Long noncoding RNA HOTAIR silencing inhibits invasion and proliferation of human colon cancer LoVo cells via regulating IGF2BP2. J Cell Biochem. 2018;120:1221–1231. doi: 10.1002/jcb.27079. [DOI] [PubMed] [Google Scholar]
- 110.Tang W, Chen S, Liu J, Liu C, Wang Y, Kang M. Investigation of IGF1, IGF2BP2, and IGFBP3 variants with lymph node status and esophagogastric junction adenocarcinoma risk. J Cell Biochem. 2019;120:5510–5518. doi: 10.1002/jcb.27834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Zhu L, Zhu Y, Han S, Chen M, Song P, Dai D, et al. Impaired autophagic degradation of lncRNA ARHGAP5-AS1 promotes chemoresistance in gastric cancer. Cell Death Dis. 2019;10:383. doi: 10.1038/s41419-019-1585-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Klinge CM, Piell KM, Tooley CS, Rouchka EC. HNRNPA2/B1 is upregulated in endocrine-resistant LCC9 breast cancer cells and alters the miRNA transcriptome when overexpressed in MCF-7 cells. Sci Rep. 2019;9:9430. doi: 10.1038/s41598-019-45636-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Li Z, Weng H, Su R, Weng X, Zuo Z, Li C, et al. FTO Plays an Oncogenic Role in Acute Myeloid Leukemia as a N 6 -Methyladenosine RNA Demethylase. Cancer Cell. 2017;31:127–141. doi: 10.1016/j.ccell.2016.11.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Paris J, Morgan M, Campos J, Spencer GJ, Shmakova A, Ivanova I, et al. Targeting the RNA m6A Reader YTHDF2 Selectively Compromises Cancer Stem Cells in Acute Myeloid Leukemia. Cell Stem Cell. 2019;25:137–148.e6. doi: 10.1016/j.stem.2019.03.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Huang Y, Su R, Sheng Y, Dong L, Dong Z, Xu H, et al. Small-Molecule Targeting of Oncogenic FTO Demethylase in Acute Myeloid Leukemia. Cancer Cell. 2019;35:677–691.e10. doi: 10.1016/j.ccell.2019.03.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Zhang B, Wu Q, Li B, Wang D, Wang L, Zhou YL. m6A regulator-mediated methylation modification patterns and tumor microenvironment infiltration characterization in gastric cancer. Mol Cancer. 2020;19:53. doi: 10.1186/s12943-020-01170-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Hou J, Zhang H, Liu J, Zhao Z, Wang J, Lu Z, et al. YTHDF2 reduction fuels inflammation and vascular abnormalization in hepatocellular carcinoma. Mol Cancer. 2019;18:163. doi: 10.1186/s12943-019-1082-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Zhu S, Wang J-Z, Chen D, He Y-T, Meng N, Chen M, et al. An oncopeptide regulates m6A recognition by the m6A reader IGF2BP1 and tumorigenesis. Nat Commun. 2020;11:1685. doi: 10.1038/s41467-020-15403-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All data generated or analyzed during this study are included in this published article and its additional files.




