Abstract
CD8 effector T cells with a CD127hi KLRG1- phenotype are considered precursors to the long-lived memory pool, while KLRG1+ CD127low cells are viewed as short-lived effectors. Nevertheless, we and others have shown that a KLRG1+ CD127low population persists into the memory phase and that these T cells (termed long-lived effector cells or LLEC) display robust protective function during acute re-challenge with bacteria or viruses. Whether these T cells represent a true memory population or are instead a remnant effector cell population that failed to undergo initial contraction has remained unclear. Here, we show that LLEC from mice express a distinct phenotypic and transcriptional signature that shares characteristics of both early effectors and long-lived memory cells. We also find that in contrast to KLRG1+ effector cells, LLEC undergo homeostatic proliferation and are not critically dependent on IL-15 for their maintenance. Furthermore, we find that LLEC are predominantly derived from KLRG1+ effector cells when isolated at day 12 of the response. Our work challenges the concept that the KLRG1+ CD127low population is dominated by short-lived cells and shows that KLRG1 downregulation is not a prerequisite to become a long-lived protective memory T cell.
Introduction
During acute infection, effector CD8 T cells help clear pathogens through robust cytokine production and cytotoxic clearance of infected cells. Following resolution of the primary infection and contraction of the effector pool, a small number of memory cells are maintained for long-term protection of the host. Effector T cells are heterogeneous, and a small population of cells expressing IL-7 receptor-α (CD127) but not Killer Cell Lectin-like Receptor G1 (KLRG1) was proposed to include effector phase cells with the potential to form the long-lived memory pool (and hence are called memory-precursor effector cells: MPEC) (1–3). In contrast, persistent expression of KLRG1, which is enforced by T-bet, Zeb-2, and Id2, has been associated with senescence and terminal differentiation (4–7). Therefore, effector cells with a KLRG1+ CD127low phenotype were termed ‘short lived effector cells’ (SLEC) and considered a transient cell population that succumbs to apoptotic death during contraction phase. A number of factors including inflammatory cues (e.g. IL-12 receptor signaling) and timing of arrival to sites of infection determine the relative generation of these subsets, and hence generation of durable CD8 T cell memory(5, 8).
However, KLRG1+ cells are not completely absent after contraction of the CD8 effector T cell population. We previously defined a memory CD8 T cell subset termed ‘long-lived effector cells’ (LLEC) which lack expression of CD62L, CD27, and CD43, and display reduced CD127 expression, but maintain expression of KLRG1 and granzyme B months after infection (9). LLEC have robust cytotoxic function, but weaker proliferative capacity upon acute challenges with bacterial or viral pathogens when compared with other memory CD8 T cells. We observed LLEC to be superior to other memory subsets in the clearance of systemic pathogens, and found that after intentional boosting or physiologic microbial exposure, LLEC comprise a dominant population among circulating memory CD8 T cells (9, 10). Furthermore, cells with the characteristics of LLEC are prominent in humans (11). In both humans and mice, CD8 T cells expressing the fractalkine receptor, CX3CR1, were shown to have a more effector-like phenotype. CX3CR1+ CD8 T cells express high levels of granzyme B and demonstrate enhanced killing of infected target cells (11). In mice, CX3CR1hi cells are localized primarily within the vasculature and often co-express KLRG1 (12). Furthermore, a subset of KLRG1+ CD8 T cells have a delayed contraction phase and express NK1.1+. These NK1.1+ have an increased capability to clear Listeria monocytogenes (LM) and produce interferon gamma (IFNγ) following re-stimulation (13). Using these various surface markers, these studies highlight the functional diversity of the CD8 memory T cell compartment as defined by cytokine production, proliferation, and cytotoxic killing. However, the derivation of LLEC and their relationship to early effector cell populations and other memory T cell subsets is unknown.
Although memory CD8 T cells are still classified into subsets based on location and expression of surface markers, the increasingly evident heterogeneous nature of memory T cells suggests that establishing distinct cell subsets with rigid definitions may be misguided (8, 14). Recent studies continue to add to the complexity by defining additional surface and transcriptional markers, trafficking patterns, metabolic, and functional features that refine our understanding of how a heterogeneous pool of CD8 T cells is formed and maintained. Diversification of the memory pool is likely necessary so that both the rapid effector responses and the proliferative burst necessary for sterilizing immunity can be achieved when faced with expanding microbial burdens. Recently, Flavell and colleagues used a fate-mapping model to demonstrate that KLRG1 can be transiently expressed during the effector phase by cells that subsequently differentiate into effector, central and tissue resident memory CD8 T cells (15). Still, these studies focused on derivation of memory CD8 T cells from a population that ceased KLRG1 expression (“exKLRG1” cells) hence provide little insight into the pool of cells that acquire or maintain expression of KLRG1 into the memory phase. Here, we investigate the derivation and characterization of KLRG1-expressing memory stage CD8 T cells, which will be referred to here as LLEC.
In this study we asked if KLRG1-expressing LLEC represent a distinct population of memory T cells or could simply be ‘leftover’ effector T cells that failed to die during the initial contraction phase. Through RNA-sequencing and in vivo adoptive transfer experiments, we find that LLEC are derived predominantly (and relatively efficiently) from KLRG1+ precursors present at day 12 of the effector phase and display a unique phenotype encompassing characteristics of both effector and memory T cells. We conclude that functional, long-lived memory T cell populations can originate from either SLECs or MPECs, KLRG1 downregulation is not required for memory generation, and that LLEC represent a distinct population encompassing properties of both the effector and memory phases of the immune response.
Materials and Methods
Mice and pathogens
C57Bl/6, and C57Bl/6.SJL (6–12 weeks) mice were purchased from Charles River/National Cancer Institute, and CX3CR1-GFP B6 mice were purchased from Jackson. All animal experiments were done with approved Institutional Animal Care and Use Committee protocols at the University of Minnesota. Lymphocytic choriomeningitis virus Armstrong strain (LCMV-Arm) and vesicular stomatitis virus expressing ovalbumin (VSV-OVA) were provided by Dr. David Masopust (University of Minnesota). Mice were infected with 2×105 plaque forming units of LCMV-Arm intraperitoneally (i.p.), and 1×106 PFU of VSV-OVA i.v. Listeria monocytogenes (LM) was thawed and grown to an optical density (OD600) of 0.1 in tryptic soy broth with 50 mg/ml. For challenges, 8*104 CFU of LM or LM-OVA were injected intravenously (i.v.).
Peptide/MHC class I tetramer enrichment
Enrichment of antigen-specific CD8 T cell precursors or LCMV-specific CD8 T cells at memory time points was performed as previously described(16). Spleens were homogenized and splenocytes were labeled with PE- or APC-conjugated tetramers, followed by magnetic enrichment over columns using anti-PE or anti-APC magnetic beads (Miltenyi). Enriched samples were stained with surface antibodies, and AccuCheck counting beads (Invitrogen) were used for calculating cell number.
Surface and intracellular flow cytometry
Single cell suspensions were generated from spleen and lymph nodes as designated. Cells were stained with the following extracellular antibodies: anti-CD4 (RM4–5), anti-CD8 (53–6.7), anti-CD44 (IM7), anti-CXCR3 (CXCR3–173), anti-CX3CR1 (SA011F11), anti-CD45.1 (A20), anti-CD45.2 (104), anti-KLRG1 (2F1), anti-CD27 (LG.7F9), anti-CD62L (MEL-14) and anti-CD127 (A7R34). Antibodies were purchased from eBioscience, BD bioscience, Biolegend, or Tonbo. LCMV Db GP33 tetramer was provided by the NIH Tetramer Facility. For intracellular staining, cells were fixed with the FoxP3 Fix/perm kit (eBioscience) and stained with the following antibodies: anti-Eomesodermin (Dan11mag), anti-T-bet (4B10). For IV labelling experiments, tissues were harvested 3 minutes post-antibody injection.
Adoptive transfers
CD8 T cells were magnetically isolated as described above from congenic donor(s) and then subjected to 4-way FACS sorting on a FACSAria II sorter with a 70μm nozzle to isolate indicated populations based on CD44, CD62L, KLRG1 and CD27. Equal numbers of the indicated CD8 T cell populations were transferred into separate congenic recipients.
BrdU labelling
Mice were treated with BrdU through drinking water for 14 days. BrdU was diluted in drinking water at a concentration of 0.8mg/mL. Drinking water also contained 2% sucrose and was protected from light for the duration of study.
RNA-sequencing
Bulk RNA sequencing of splenic memory CD8 T cell subsets was performed 90 days post infection with LCMV-Arm. CD8 T cells were isolated from the spleen and FACS sorted. Samples were mini-pooled with 3 mice per sample, resulting in 3 samples total for each subset (LLEC, TEM, TCM, naïve). RNA was isolated using Qiagen RNeasy plus mini kit and submitted for sequencing by the University of Minnesota Genomics Center (UMGC) using the HiSeq 2500 instrument (high output sequencing, paired end read, 125 bp). For single-cell RNA sequencing P14 cells were harvested and FACS sorted from the spleen at 12 or 60 days post-LCMV-Arm infection. Samples were processed using the 10X Chromium platform to generate single-cell gel beads in emulsion (GEMs). Libraries were prepared using Chromium Single cell 3’ Reagent Kits v2 Library. (10X Genomics). Samples were sequenced using the UMGC HiSeq 2500 instrument (high output sequencing, paired end read, 125 bp), and demultiplexed with cellranger (ver 2.0.0) “mkfastq” function. Using the cellranger “count” function, reads were aligned with STAR (ver 2.5.1b) against GRCm38 genome with Ensembl release 84 GTF. Cells were filtered based on quality control metrics, clustered, and further analyzed using functions from the Seurat package (ver 2.3.4) within R software (ver 3.4.3). Differential gene expression testing was performed with Seurat using the Wilcoxon rank sum test. Unsupervised hierarchical clustering of differentially expressed genes was performed using Pearson correlation and Ward1 methods via the “aheatmap” function in the NMF R package (ver 0.22).
Data analysis and statistics
Flow cytometry data was analyzed using Flow Jo (Treestar). Statistics were calculated using Graphpad Prism software. P values are represented as follows: ****p<0.0001, ***p<0.001, **p<0.01, *p<0.05.
Online supplemental material
Fig. S1 shows that LLEC are maintained at 5 months post-infection at similar or greater frequency to TEM after LM, VSV-OVA, or LCMV-Arm infection. Fig. S2 shows the gating strategy and post-sort purity of sorted CD8 T cell subsets. Fig. S3 shows differential gene expression levels defining clusters of cells 60 days post LCMV-Arm infection and the transcriptional heterogeneity within the TEM population. Fig. S4 shows that unique gene expression patterns identify shared gene expression patterns between SLEC and LLEC at the single cell level.
Results
LLEC are a unique subset within the memory T cell compartment
Classically, CD62L expression is used to differentiate central memory T cells (TCM) from the conventionally described effector memory population(17). However, it is becoming increasingly apparent that the CD62L- memory compartment contains a heterogeneous population of cells(17). Here, we show that CD62L- memory cells are comprised of at least two distinct populations based on KLRG1 and CD27 surface expression (Fig 1A). Our previous data defined LLEC as a subset of antigen-specific CD8 T cells which were present at memory timepoints and had low expression of CD27 and the activation associated glycoform of CD43, and which we noted had a uniquely high expression of KLRG1 (9). Using a gating strategy based on expression of KLRG1, we further characterized the KLRG1+ fraction of the polyclonal repertoire after acute infection with LCMV. The KLRG1+ CD27- fraction had low CD127 and CXCR3 expression and high CX3CR1 and T-bet expression, relative to KLRG1- CD27+ cells (Fig 1B). Therefore, the traditionally described CD62L- memory population is heterogeneous and contains at least two phenotypically unique subsets: KLRG1+ CX3CR1+, T-bet+, CD127low LLEC, and KLRG1-, CXCR3+, T-betlow, CD127hi cells TEM. These two subsets were also observed among antigen specific (gp33/Db tetramer positive) cells at least 90 days post LCMV-Arm infection (Fig 1C). Furthermore, LLEC are not exclusive to endogenous T cells 3 months after LCMV-Arm infection, but are also observed at equal or greater frequency to TEM 5 months after adoptive transfer of transgenic T cells and infection with LCMV-Arm, LM, or Vesicular stomatitis virus (VSV) (Suppl. Fig 1).
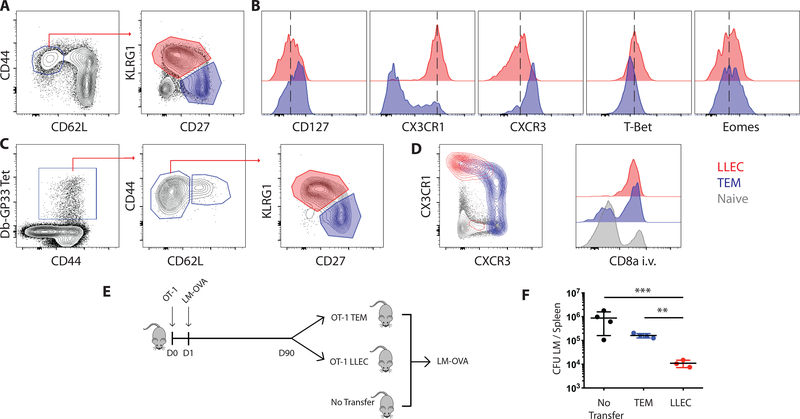
Figure 1: Long lived effector cells make up a distinct subset within the effector memory T cell compartment.
(A) Ninety days post infection with LCMV-Arm, spleens were harvested from B6 mice and long-lived effector cells (LLEC) and effector memory cells (TEM) were differentiated based on their expression of CD27 and KLRG1. (B) Polyclonal LLEC have differing expression of several surface markers and transcription factors, including high expression of CX3CR1 and T-bet, and low expression of CD127, CXCR3, and Eomes when compared to TEM. C) LLEC were also identified within GP33-H2-Db tetramer positive cells 30 days post-infection with LCMV-Arm. D) CX3CR1 expression correlated with iv antibody labeling of LLEC in the spleen which is indicative of localization to the red pulp. E) LLEC and TEM OT-I memory cells were generated in mice challenged with virulent recombinant Listeria monocytogenes expressing ova (LM-OVA). At 90 days post-infection TEM and LLEC OT-I were FACS sorted and 2*104 cells were transferred into congenially distinct naïve recipients. One day later, the recipient mice were challenged with LM-OVA and bacterial burden was determined in the spleen five days post challenge (F).
Using i.v. injection of antibodies to label CD8 T cells in the vasculature and red pulp of the spleen we observed striking and uniform localization of LLEC to the red pulp (indicated by antibody staining) as compared to TEM and naïve T cells (Fig 1D)(18, 19). Recent reports describe a CX3CR1+ CD8 T cell memory population of cells that is also restricted to the vasculature (12). These data define shared characteristics between LLEC and these cells by surface markers (CX3CR1+, KLRG1+) and vascular localization. As stated before, CX3CR1+ memory cells are also identified as a unique subset in humans capable of enhanced cytotoxic potential (11).
To confirm the functional features of LLEC with this gating strategy, we assessed the capacity of the memory subsets to protect against LM infection. LLEC (CD44+ CD62L-KLRG1+ CD27- ) and TEM (CD44+ CD62L- KLRG1- CD27+) OT-I cells were sorted from donor mice 90 days post LM-OVA and transferred to naïve mice to measure bacterial clearance after high dose challenge (Fig 1E). Mice receiving LLEC had the largest reduction in CFU, which was a significant improvement over TEM cells (Fig 1F). Thus, we conclude that LLEC are distinct from TEM, and can be defined by surface expression of KLRG1 and CX3CR1, strong localization to the splenic red pulp, and robust protective immunity against LM challenge.
LLEC express a transcriptional profile enriched for effector genes
To further define the characteristics of LLEC, we performed RNA-sequencing of sorted LLEC, TEM, and TCM memory populations (Suppl. Fig 2). Our analysis revealed a unique transcriptional profile exclusively defining LLEC. When plotting the variance using principal component analysis (PCA), LLEC group distinctly from naïve, TCM, and TEM (Fig 2A). An examination of genes differentially expressed between memory subsets reveals LLEC have increased expression of genes associated with cytotoxicity (GzmA and GzmB) and terminal effector differentiation (Zeb2)(6). Additionally, expression of the effector associated fractalkine receptor, Cx3cr1, and sphingosine-1-phosphate receptor 5, S1pr5, are elevated in LLEC (Fig 2B). In contrast, LLEC have reduced expression of genes encoding the lymphoid homing chemokines L-selectin and CCR7. Interestingly, TEM had few uniquely expressed genes, but tended to exhibit intermediate expression of many genes with opposing expression in LLEC and TCM subsets. Cumulatively, these results indicate that LLEC exhibit a transcriptional profile that is distinct from other memory T cell subsets, that is characterized by high expression of effector genes and transcription factors such as Zeb2 that enforce terminal differentiation (4, 6, 20).
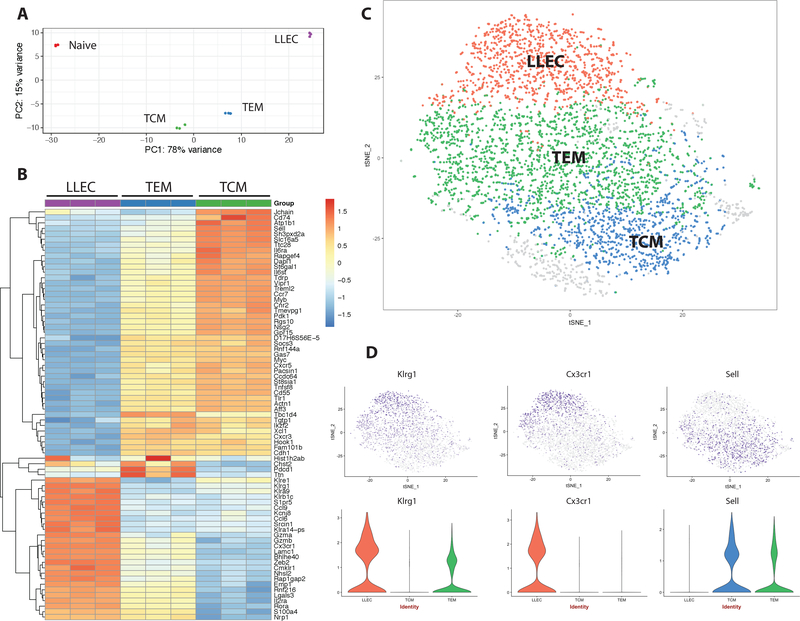
Figure 2: LLEC maintain a unique transcriptional profile enriched for an effector affiliated gene signature.
Splenic LLEC (CD44+ CD62L-KLRG1+ CD27-), TEM (CD44+ CD62L- KLRG1- CD27+), and TCM (CD44+CD62L+KLRG1-CD27+), and naïve CD8 T cells were FACS sorted from mice at least 90 days post infection with LCMV-Arm. Samples were mini-pooled with 3 mice per sample, resulting in 3 samples total. (A) Principal component analysis of all populations. (B) Hierarchical clustering depicting gene expression of the 75 most variable genes between memory subsets (LLEC, TEM, and TCM). C) t-distributed stochastic neighbor embedding (t-SNE) dimensional reduction was used to display unique clusters classified by a shared nearest neighbor algorithm (Seurat) via single cell sequencing (scRNA-seq) of the splenic P14 T cell population 60 days post LCMV-Arm infection. P14 cells were FACS sorted using Thy1.1 prior to sequencing. D) Gene expression levels of Klrg1, Cx3cr1, and Sell visualized by color intensity (t-SNE plot) and violin plots.
We next utilized single cell RNA sequencing (scRNA-seq) to gain a perspective on the transcriptional profile of antigen specific memory T cells without bias by surface expression of phenotypic markers, and to determine whether the defined memory subsets could be further resolved. At 60 days post LCMV-Arm infection, antigen specific P14 cells were sorted and 4,471 cells were sequenced. PCA of the normalized expression data was used to reduce the gene dimensionality prior to cell clustering by the Seurat method. The cells in PCA space were then projected into a t-distributed stochastic neighbor embedding (t-SNE) dimensional reduction space, plotted in 2D, and labeled by cluster assignment (Fig. 2C). This revealed a cluster of cells uniquely expressing LLEC associated genes (Fig 2C, Suppl. Fig 2A). The effector associated genes Klrg1, Cx3cr1, GzmB and Zeb2 define this population (Fig 2D, Suppl. Fig 3B), supporting the findings from our other analysis methods (Figures 1 and 2). LLEC most distinctly clustered from TCM, which were defined by elevated translational machinery as well as L-selectin (Sell) (Fig 2D, Suppl. Fig 3A). In agreement with surface protein expression, LLEC also had reduced expression of Il7r relative to both TCM and TEM memory subsets (Suppl. Fig 3B).
Our scRNA-seq identified TEM cells as exhibiting an intermediate transcriptional phenotype compared to the more distinct differences identified between LLEC and TCM groups. Further dissection of the TEM population revealed additional heterogeneity within this subset of cells (Suppl. Fig 3C). Intermediate expression of Cx3cr1 identified one cluster (orange) within the TEM compartment that displays characteristics of the recently described peripheral memory cells (TPM) responsible for surveilling tissues (Suppl. Fig3D) (12). Differential gene expression analysis of the TEM population revealed heterogeneity based on gene expression of Cx3cr1, S1pr5, Zeb2, Gzmb, Il7r, and Sell among others (Suppl. Fig 3D). However, unlike TPM, we were unable to identify a unique set of markers to clearly define the remaining subsets (red, purple) (Suppl. Fig 3C,D). Transcriptional analysis from both our unbiased scRNA-seq clustering as well as sorted bulk RNA sequencing indicate that TEM cells lie on a spectrum between LLEC and TCM. Overall, our sequencing analysis reveals a high level of heterogeneity among memory T cells, particularly within the TEM compartment. However, unbiased clustering identifies LLEC as a distinct population with a transcriptional signature unique from other memory populations.
LLEC share aspects with conventional memory CD8 T cells
Our data show that LLEC represent a unique population of memory cells by both surface protein expression and transcriptional analysis. However, many of the genes highly expressed by LLEC at memory stage (i.e. Zeb2, GzmA, GzmB, Klrg1, Cx3cr1) are also expressed by the SLEC population during the effector response (2, 3, 21). This led us to examine whether LLEC are truly a memory T cell subset or are instead simply the residual population of SLEC at the end of a protracted contraction phase. One of the key properties that differentiates memory cells from early effector cells is their longevity. Memory CD8 T cells rely on IL-2 and IL-15 cytokine signals for maintenance in order to survive for months to years after antigen exposure (22, 23). We find that LLEC express increased levels of proteins associated with enhanced CD8 T cell survival as compared to day 12 SLECs. At 100 days post-infection LLEC express elevated levels of cytokine receptors CD25, CD122, and CD127 relative to day 12 post-infection SLEC (Fig 3A,B). TEM also express increased levels of these cytokine receptors as compared to day 12 MPEC or SLEC. Additionally, TEM have elevated expression of the transcription factor Eomes and reduced T-bet relative to effector cells (Fig 3A,B). LLEC mirror these differences relative to SLEC (although LLEC show slight but significantly reduced Eomes expression relative to TEM) indicating a subset of cells better equipped for long-term survival.
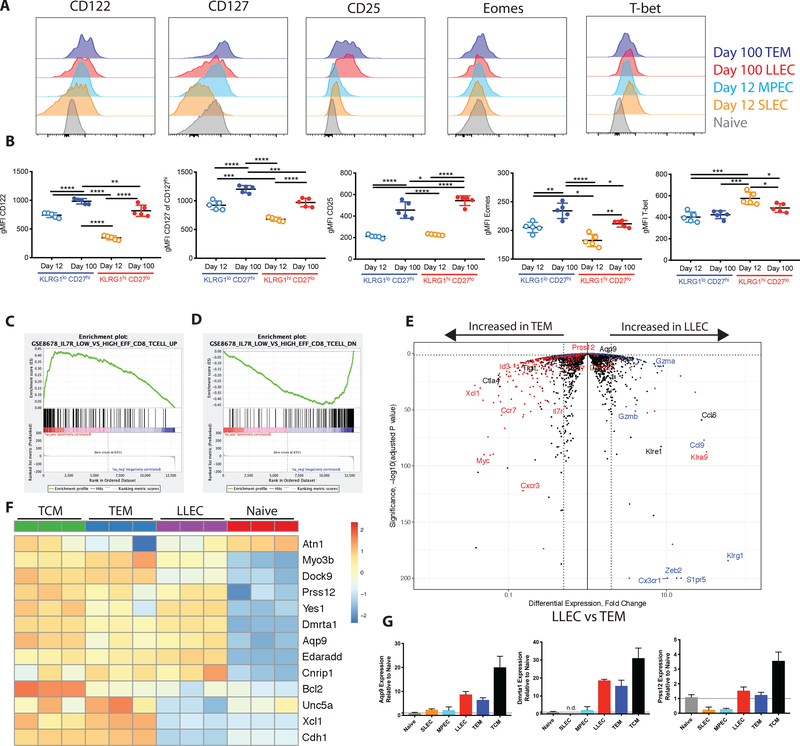
Figure 3: Upregulation of memory associated genes differentiates LLEC from short lived precursors.
(A) CD122, CD127, CD25, Eomes, and T-bet protein expression was compared in KLRG1+ CD27- and KLRG1- CD27+ P14 CD8 T cells were compared 12 days and 100 days post LCMV-Arm infection, with representative staining shown in (A) and quantification shown in (B). (C) Gene set enrichment analysis (GSEA) comparing genes differentially expressed in LLEC vs TEM to genes upregulated in short-lived effector CD8 T cells compared to memory precursor cells (GSE8678) reveals enrichment for short-lived effector transcriptional phenotype. (D) Gene set enrichment analysis (GSEA) comparing genes differentially expressed in LLEC vs TEM to genes downregulated in short-lived effector CD8 T cells compared to memory precursor cells (GSE8678) reveals enrichment for short-lived effector transcriptional phenotype. (E) Volcano plot showing gene differential expression by fold change and p-value. LLEC and TEM gene list generated in Fig 2. (F) Heatmap comparing differential gene expression of hallmark memory genes between TCM, TEM, LLEC, and naïve t cells. (G) qPCR confirmation of Dmrta1, Prss12, and Aqp9 expression in T cell subsets.
Given this phenotypic dichotomy, we considered whether the transcriptional profile of LLEC might align more closely with SLECs or MPECs. Gene set enrichment analysis (GSEA) reveals that the differential gene expression profile distinguishing LLEC from TEM is enriched for many of the same genes which distinguish SLECs from MPEC (Fig 3C). Additionally, genes downregulated by SLEC relative to MPEC are enriched within the TEM transcriptional profile (Fig 3D). Thus, LLEC share a transcriptional profile with SLEC while also displaying characteristics of long-lived memory cells. Directly comparing the differential gene expression profiles between LLEC and TEM shows that LLEC express higher levels of “effector genes” (Gzma, Gzmb, Cx3cr1, Zeb2, S1pr5) (Fig 3E) than TEM.
We also investigated expression of a group of ‘hallmark memory genes’ that are characteristic of CD8 T cells at very late memory timepoints (24). These include Bcl2 and Aqp9, both of which are important for long-term memory CD8 T cell survival (25, 26). Notably, TCM and TEM but also LLEC upregulate a majority of these genes relative to naïve cells (Fig 3F). A subset of ‘hallmark memory genes’ (Dmrta1, Prss12, and Aqp9) was confirmed by qPCR to be elevated in LLEC, TEM, and TCM relative to SLEC and MPEC (Fig 3G). The finding that LLEC express genes that regulate maintenance of the memory pool supports the concept that these cells form a stable, relatively long-lived memory population. It is interesting to note that expression of three “hallmark” genes (Xcl1, Unc5a and Cdh1) are expressed in TEM and TCM but not LLEC, suggesting upregulation of those genes are specific for some, but not all memory populations. Overall, our data illustrate that LLEC make up a discrete component of the memory T cell pool, rather than merely cells with a SLEC gene expression profile that had escaped initial contraction.
LLEC have a distinct gene expression profile compared to SLEC
Our studies identified characteristics shared between LLEC and both early effector cells and memory cells. We also compared the transcriptional profile of P14 T cells at 12 days and 60 days post infection by scRNA-seq (Fig 4A) in order to observe changes in gene expression as the LLEC population emerges. Analysis of the day 12 cells identified SLEC and MPEC populations (Suppl. Fig 4A). Transcription of Klrg1, Cx3cr1, Zeb2, Il7r, and Bcl2 readily differentiated SLEC and MPEC populations (Suppl. Fig 4B). Following alignment and merging of the two timepoints (27), we performed t-SNE dimensional reduction on the PCA data to visualize the clusters (Fig 4B). An effector-like profile defined a group of cells which contained both day 12 and day 60 P14 cells (Suppl. Fig4D). Elevated expression of Klrg1, Cx3cr1, Gzma, S1pr5, and Zeb2 as well as low Il7r expression confirmed this cluster encompasses SLEC and LLEC cells from the combined dataset (Fig 4C, Suppl. Fig4E). This effector-like cluster contains 30.7% of all day 12 cells and 20.9% of day 60 cells (Fig 4D).
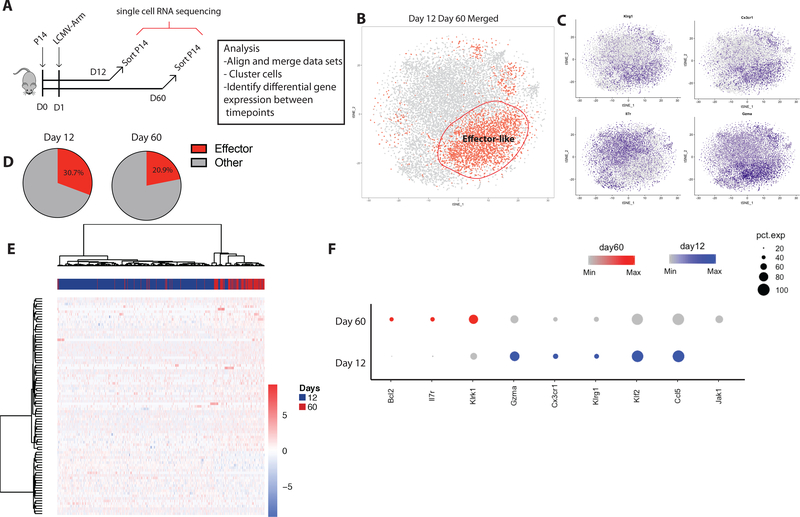
Figure 4: Effector cell gene expression is reduced in LLEC compared to SLEC.
A) P14 cells were transferred to C57Bl/6 mice and challenged with LCMV-Arm. At 12 days post-infection(dpi) and 60 dpi P14 cells were isolated from the spleen and processed for scRNA-seq using the 10X Chromium platform (10X Genomics). Datasets from 12dpi and 60dpi were aligned and merged using the Seurat R package. Following PCA, t-distributed stochastic neighbor embedding (t-SNE) dimensional reduction was completed and cells were labeled based on clustering using Seurat. B) t-SNE plot of aligned merged P14 data sets reveals family of cells defined by effector associated expression profile. C) Expression distribution of Klrg1, Cx3cr1, Il7r, and Gzma by cells. D) Proportion of total cells at day 12 and day 60 post infection timepoints that cluster within the ‘effector-like’ merged cluster. (E). Differential gene expression analysis comparing day 12 and day 60 cells (from only the effector cell cluster) resulted in 85 DE genes. Unsupervised hierarchical clustering of these genes with the scaled dataset highlights the gene expression differences between the two timepoints. F) Split dot plots comparing differentially expressed genes between day 12 and day 60 cells within effector-like cell cluster. Size of dots represents frequency of cells within the cluster expressing and the intensity of the color indicates the expression level of the indicated gene.
We next investigated this effector-like cell cluster for genes differentially expressed between the day 12 SLEC and day 60 LLEC. This query revealed differential expression in 85 genes. Hierarchical clustering using these 85 genes distinctly separated day 12 SLEC from day 60 LLEC (Fig 4E). Therefore, while overall similarities in transcriptional profiles drive effector-like cells to cluster together, there are distinctions between SLEC and LLEC. Differences were revealed in pathways necessary for cell survival and effector function. For example, IL7r and Bcl2 were elevated in day 60 LLECs relative to the day 12 effector cells indicating upregulation of a cell survival transcriptional pathway in effector cells at memory timepoints (Fig 4F). Interestingly, although LLECs express Gzma, Cx3cr1, and Klrg1 like SLECs, they have a reduced magnitude of expression for these genes (Fig 4F). This indicates that although LLEC maintain transcription of canonical effector differentiation genes, they do so at a lower expression level than SLEC. This raises the interesting possibility that the most terminally differentiated effector cells are programmed to die, and that LLEC arise from SLEC that were either less differentiated or able to downregulate these pathways and diverted away from death during contraction.
IL-15 dependency and homeostatic turnover of LLEC
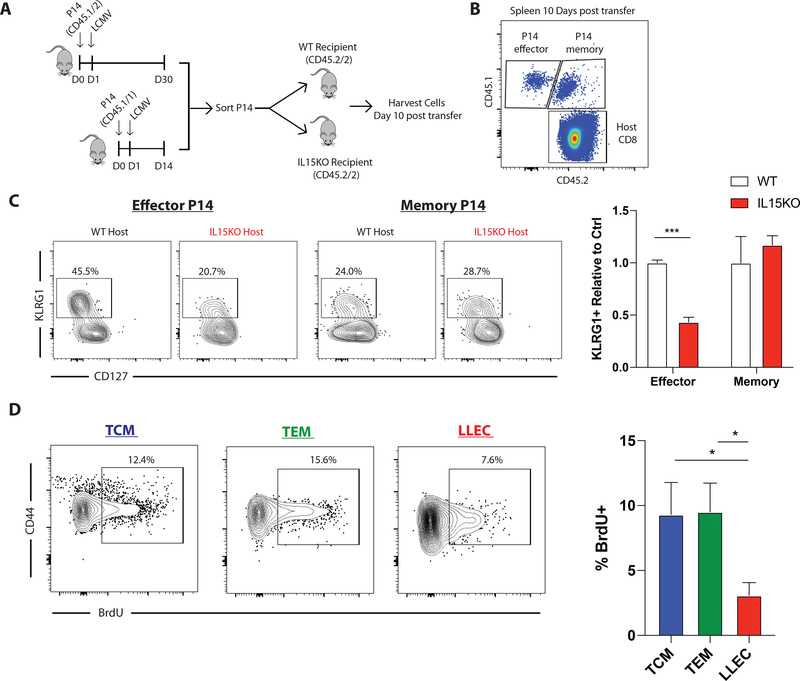
We next tested whether LLECs respond to cytokine cues like IL-15 for their maintenance. Previous reports have demonstrated that KLRG1+ effector cells require IL-15 for survival and rapidly deteriorate within IL-15 deficient hosts(1). To identify whether LLEC similarly depend upon IL-15 we co-transferred effector and memory P14 cells into IL-15 knockout or sufficient hosts (Fig 5A). 10 days post transfer spleens were harvested from recipients and donor cells were characterized (Fig 5B). While memory cells maintained similar frequencies of KLRG1+ cells from both WT and IL-15 knockout hosts, KLRG1+ effector cells were lost upon transfer to IL-15 knockout hosts as compared to WT hosts. KLRG1+ effector cells exhibited a > 2-fold reduction after 10 days in IL-15 deficient hosts relative to WT mice (Fig 5C). This demonstrates a differential survival requirement for IL-15 between KLRG1+ effector cells and LLEC.
Figure 5: The IL15 dependency of LLEC is similar to other memory cells, but homeostatic turnover is low.
A) Congenically labelled P14 cells were isolated from LCMV-Armstrong challenged mice. D14 effector cells (CD45.1/1) and D30 memory cells (CD45.½) were transferred into C57BL/6 or IL15-KO recipients (CD45.2/2). B) 10 days post transfer spleens were harvested for flow cytometry. C) Donor P14 cells were analyzed for phenotype of surviving cells. P14 memory cells had similar frequencies of KLRG1+ cells in both C57BL/6 and IL15 knockout recipients. Effector P14 cells transferred to IL15 knockout recipients had reduced KLRG1+ cells relative to those transferred to C57BL/6 recipients. n=3 mice, representative from two independent experiments D) Memory P14 cells were established by transferring naïve P14 cells and challenging mice with LCMV-Armstrong infection. 30 days post infection mice were treated with BrdU in drinking water for two weeks. 14 days post transfer BrdU incorporation into donor cells was quantified by flow cytometry. n=4 mice, representative of two independent experiments.
We have shown that LLEC make up approximately half of the CD62L- memory population and are relatively stable over an extended period of time (Fig 1 and Suppl. Fig 1). We next assessed whether these cells undergo homeostatic proliferation similarly to other memory cells. P14 memory cells were generated by acute LCMV infection and rested for 30 days. Mice were then treated daily with BrdU in the drinking water for two weeks. TCM, TEM, and LLEC all proliferated as measured by BrdU incorporation (Fig 5D). However, LLEC divide more slowly than TEM and TCM subsets. These data identify the ability of LLEC to homeostatically divide >30 days post-infection and show that their dependency on IL-15 for maintenance is reduced as compared to effector cells.
LLEC are derived from Day 12 KLRG1hi effector cells
LLEC possess traits characteristic of both effector and memory cells, representing a functionally and transcriptionally “hybrid” phenotype. These cells are functionally similar to terminally differentiated effector cells yet remain alive long after the resolution of infection as a protective memory population. This could arise either by KLRG1- MPEC (or their progeny) acquiring expression of KLRG1 and other LLEC markers, or by some “short-lived” effector cells acquiring the memory characteristics found in LLEC. This issue is not resolved by the recent fate-mapping studies demonstrating “ex-KLRG1” can populate the long-lived memory pool(15), since LLEC are (by phenotypic definition) not included in the ex-KLRG1 pool.
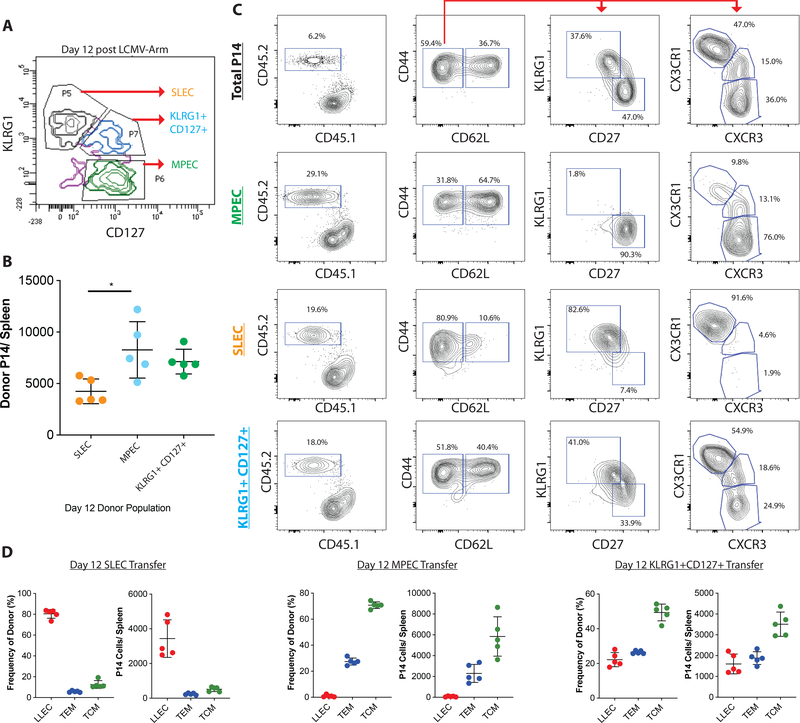
To address this, KLRG1+ CD127- SLEC, KLRG1- CD127+ MPEC, and KLRG1+ CD127+ P14 cells were isolated 12 days post LCMV-Arm infection and transferred to congenically distinct naïve hosts to define their capacity to generate memory CD8 T cells (Fig 6A). Cells were rested in the uninfected recipient hosts for thirty days before analysis of donor cell phenotype. Consistent with previous reports, transferred MPEC produced significantly more total memory cells than transferred SLEC or KLRG1+ CD127+ populations(3, 28). However, this difference was only around 2-fold, suggesting many of the donor SLEC and KLRG1+ CD127+ cells also persist into memory phase (Fig 6B).
Figure 6: LLEC are derived from KLRG1+ short-lived effector cells.
P14 CD8 T cells were transferred into congenically distinct recipients. One day later, the recipients were infected with LCMV-Arm. (A) On day 12 post infection, P14 cells were FACS sorted based on KLRG1 and CD127 expression (A). These populations were transferred into naïve recipients. Thirty days post transfer, mice were harvested and donor cells enriched by tetramer pulldown. (B) Thirty days post transfer the Day 12 KLRG1- CD127+ (MPEC) CD8 T cells produced more long-lived cells than KLRG1+CD127- (SLEC) and KLRG1+CD127+ precursors. (C) Representative flow cytometry plots depicting donor cell phenotype at day 30 post transfer. (D) Frequency and number of TCM (CD44+ CD62L+), TEM (CD44+ CD62L- KLRG1- CD27+), and LLEC (CD44+ CD62L- KLRG1+ CD27-) phenotype transferred cells.
As expected, transfer of unsorted P14 day 12 effector cells generated a diverse population comprised of TCM, TEM, and LLEC (Fig 6C, 6D). However, transferred MPEC strictly produced TCM and TEM populations with no significant generation of LLEC. In contrast, SLEC-phenotype cells almost exclusively produced LLEC (Fig 6C, 5D). Transfer of the KLRG1+CD127+ double positive effector cell population generated TCM, TEM, and LLEC much like total P14 transfer (Fig 6B). These findings identify the KLRG1+CD127+ population at day 12 post infection as the precursor with the potential to produce the recently described ex-KLRG1 memory population(15). Our results show that the cell fate decision to differentiate into LLEC is programmed by 12 dpi; we next asked how early this program was imparted onto the precursor cells.
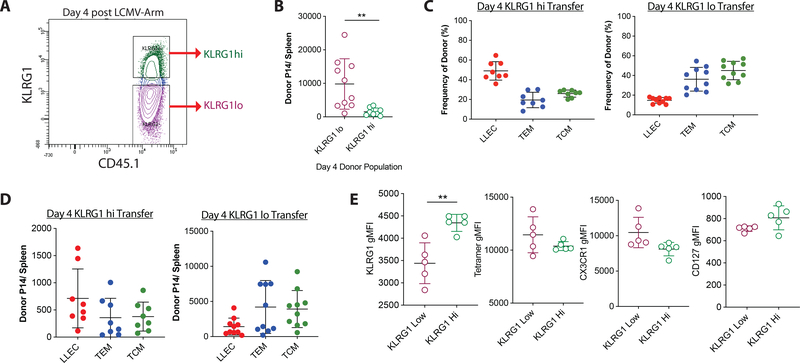
We also isolated KLRG1hi and KLRG1low P14 cells from mice at 4 days post LCMV-Arm infection, the earliest day SLEC vs MPEC distinctions can be made based on KLRG1 surface expression. Cells were transferred into infection matched recipients to assess potential to generate memory populations (Fig 7A). As previously described, KLRG1low effector cells produced more overall memory (Fig 7B, 7D) (2). However, we find that approximately 50% of cells generated from KLRG1hi precursors formed KLRG1+ memory cells, whereas less than 20% of KLRG1low cells generated KLRG1+ cells (Fig 7C), showing that day 4 KLRG1hi effectors still favor the formation of LLEC. We next asked whether LLEC derived from KLRG1hi or KLRG1low effector cells are different. LLEC derived from day 4 KLRG1hi precursors displayed elevated KLRG1 surface expression compared to LLEC from KLRG1low precursors (Fig 7E). No differences were observed in tetramer staining, CX3CR1, or CD127 expression levels (Fig 7E). Transfer of donor effector cells at this early day 4 timepoint demonstrates that memory-fate decisions are starting to be made, but are not yet strictly defined. We chose these two separate time points (day 4 and day 12) to capture the resulting memory populations that form from effector cells during these different phases of the response to acute infection. We conclude that at day 4, LLEC can still form from either KLRG1hi or KLRG1low effector cells. However, by day 12 post-infection, LLEC are now derived overwhelmingly from the KLRG1+ effector cell pool.
Figure 7. KLRG1+ effectors preferentially generate LLEC.
A) Naïve P14 CD8 T cells were transferred to congenically distinct recipients and one day later infected with LCMV-Arm. 4 days post infection, P14 cells were sorted based on KLRG1 expression and transferred into LCMV-Arm infection matched recipients. Thirty days post transfer donor cells were enriched by tetramer pulldown and analyzed by flow cytometry. B) The total number of P14 memory cells generated from KLRG1hi and KLRG1low donor cells. Frequency (C) and number (D) of TCM (CD44+ CD62L+), TEM (CD44+ CD62L- KLRG1- CD27+), and LLEC (CD44+ CD62L- KLRG1+ CD27-) phenotype cells. E) gMFI of KLRG1, GP33-Tetramer, CX3CR1, and CD127 of LLEC that resulted from either KLRG1hi and KLRG1low donor cells.
Discussion
Our data demonstrate that a population of memory T cells, LLEC, balance the dual roles of rapid functionality and longevity. Moreover, these cells are predominantly derived from KLRG1+ progenitor cells, illustrating that the early fate decisions that generate ‘SLECs’ do not preclude their development into stable and long-lived memory cells. Thus, ‘short-lived’ appears to be a misnomer when applied indiscriminately to all KLRG1+ effector cells. The relative efficiency with which day 12 SLEC can become LLEC (only around 2-fold reduced numbers compared to production of conventional memory cells from transferred MPEC – Fig. 6B) is consistent with previous works (2)(29) and suggests many SLEC-phenotype cells present after the peak of the effector response may persist to become LLEC in the memory phase. It remains unclear, however, whether there is a KLRG1+ effector cell with specialized capacity to seed the LLEC population or whether all SLEC at this stage have equivalent potential of acquiring the memory characteristics associated with LLEC.
Our results also illustrate that LLEC share phenotypic and transcriptional characteristics with memory cells that contrast from KLRG1+ effector cells, supporting that LLEC are an independent population distinct from SLEC. LLEC dependency on IL-15 is consistent with other circulating memory subsets and contrasts with the rapid decline of KLRG1+ effector cells in an IL-15 deficient host (1). Importantly, these experiments do not imply that LLEC (and other memory populations) do not depend upon IL-15, but rather that the requirement for IL-15 by LLEC aligns more closely with TCM and TEM than it does with effector cells. Further studies will be helpful to characterize the cytokine signals required for LLEC survival and their slow turnover.
Recent work demonstrated that KLRG1+ effector cells can downregulate expression of KLRG1 early after infection and contribute to memory T cell generation (15). The convincing characterization of these ‘ex-KLRG1’ cells showed that they contribute to the diversity of the memory T cell pool with regard to proliferation and cytotoxicity (15). Our study illustrates another mechanism by which KLRG1+ effector cells add to the memory pool separate from the ex-KLRG1 population. Herndler-Branstetter et al. found that the ex-KLRG1 pool was characterized by reduced expression of Zeb2, a transcription factor that has been associated with the terminal differentiation of short-lived effector cells (4, 6, 15, 20). Our studies indicate that downregulation of Klrg1, Zeb2 and several other genes associated with SLEC is not a requirement for long-term survival and the ability of LLEC to confer robust protection upon pathogen exposure. Thus, LLEC are a unique subset of cells predominantly derived from KLRG1+ effector cells that maintain a high level of functionality (9) yet persist due to expression of key genes associated with durable memory in CD8 T cells.
The use of CX3CR1 expression to interrogate memory function has led to the definition of CX3CR1hi cells forming the ‘effector memory’ compartment (12). These cells are proposed to surveille the blood and are derived from CX3CR1+ effector cells. There are obvious similarities between this population and the LLEC described here. Bottcher et al. showed that human CX3CR1+ memory cells have increased Granzyme B expression and cytotoxic activity against antigen specific target cells in vitro and in vivo (30). However, Gerlach et al. demonstrated that while all KLRG1+ memory cells are CX3CR1hi, the reciprocal is not true. Anywhere from 40% to 70% of CX3CR1hi peripheral memory cells were KLRG1+ depending on the infectious challenge (12). Given the similarities between CX3CR1+ cells and LLEC, a direct comparison between these populations would be required to define differences between these two groups of cells. In future experiments, it will be interesting to functionally compare CX3CR1+ KLRG1- cells with LLEC. The identification of analogous cells to LLEC in humans provides hope that the potent protective ability of LLEC that we have identified in mice (9) may also be a prominent feature in human LLEC. Functional studies must be performed to determine whether human LLEC share the enhanced pathogen clearing functionality with their murine counter parts in addition to their similar transcriptional signatures. Elevated expression of effector genes such as granzyme B and perforin provide clues that they may, but these alone do not provide definitive evidence of function. The translational potential for intentionally generating a memory T cell response optimized to target infected cells makes LLEC an important subset for additional study.
The finding that LLEC are derived from day 12 SLEC provides intriguing possibilities for modulating the composition of the immune response. Our data suggest that enriching for KLRG1+ effector cells may preferentially increase the LLEC population. Given that LLEC have enhanced capability to effectively clear systemic infections, a better understanding of the signals required for their generation could be exploited to optimize CD8 T cell vaccination strategies. These experiments show that by day 12 post- infection, KLRG1+ CD127lo cells have already differentiated from memory precursors that can seed the TEM and TCM populations. In the early effector phase (day 4) both KLRG1hi and KLRG1low effector cells are still capable of forming LLEC. However, at this time point effector cells are still likely impacted by the ongoing response to acute infection and KLRG1low cells may still upregulate this marker after transfer into infection-matched recipients. Recent work has established that sustained enforcement of the effector-like transcriptional program via Id2 is required to maintain a terminal effector phenotype(7). During effector differentiation, T-bet- induced Zeb2 is required for generating KLRG1+ effector cells, whereas Eomes, Id3, and FOXO1 are responsible for generating CD127+ memory precursor cells(4, 6, 31, 32). The plasticity of CD8 T cell subsets has been highlighted in recent years. Competing networks of transcription factor gradients (ex: Id2/Id3, Zeb1/Zeb2) intricately regulate the differentiation state of the memory population(6, 20). However, the existence of LLEC suggest that the fate decision for terminal effector differentiation is not mutually exclusive from the acquisition of memory properties. LLEC maintain high Zeb2 expression and acquire increased memory associated gene expression (Bcl2, Dmrta, Prss12, Aqp9, Fig 3G). Future work identifying transcription factors regulating expression of these genes while maintaining transcription of effector genes are needed. Important questions remain as to whether there is a specialized subset of KLRG1+ effector cells that are LLEC precursors, and which signals promote their longevity into the memory pool.
Supplementary Material
Key points.
LLEC exhibit transcriptional characteristics common to effector and memory T cells.
LLEC undergo homeostatic proliferation, but do not critically depend on IL-15.
LLEC are predominantly derived from KLRG1hi effector cells at 12 days post-infection.
Acknowledgements
We thank Dr. Stephen Jameson for numerous helpful discussions during the course of this project and critical review of the manuscript. We also thank the members of the Jamequist lab, and the UMN Center for Immunology for key input. We thank the University Flow Cytometry Resource and the University Genomics Center for their expertise and contribution to this work.
This work was supported by R01-AI116678 (to S.E.H.), an AAI careers in Immunology award (to M.A.H. and S.E.H.), a postdoctoral fellowship- UMN Cancer Biology Training Grant (T32 CA009138 to KR), and a Paul C. Shiverick/CRI Irvington fellowship (to H.BdS.).
Abbreviations
- LCMV-arm
lymphocytic choriomeningitis virus Armstrong strain
- LLEC
long-lived effector cell
- LM
Listeria monocytogenes
- MPEC
memory precursor effector cell
- OVA
ovalbumin
- PCA
principal component analysis
- scRNA-seq
single cell RNA sequencing
- SLEC
short-lived effector cell
- TCM
central memory cell
- TEM
effector memory cell
- t-SNE
t-distributed stochastic neighbor embedding
- VSV
vesicular stomatitis virus
References
- 1.Joshi NS, Cui W, Chandele A, Lee HK, Urso DR, Hagman J, Gapin L, and Kaech SM. 2007. Inflammation Directs Memory Precursor and Short-Lived Effector CD8+ T Cell Fates via the Graded Expression of T-bet Transcription Factor. Immunity 27: 281–295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Sarkar S, Kalia V, Haining WN, Konieczny BT, Subramaniam S, and Ahmed R. 2008. Functional and genomic profiling of effector CD8 T cell subsets with distinct memory fates. J. Exp. Med. 205: 625–640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kaech SM, Tan JT, Wherry EJ, Konieczny BT, Surh CD, and Ahmed R. 2003. Selective expression of the interleukin 7 receptor identifies effector CD8 T cells that give rise to long-lived memory cells. Nat. Immunol. 4: 1191–1198. [DOI] [PubMed] [Google Scholar]
- 4.Dominguez CX, Amezquita RA, Guan T, Marshall HD, Joshi NS, Kleinstein SH, and Kaech SM. 2015. The transcription factors ZEB2 and T-bet cooperate to program cytotoxic T cell terminal differentiation in response to LCMV viral infection. J. Exp. Med. 212: 2041–2056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kaech SM, and Cui W. 2012. Transcriptional control of effector and memory CD8+ T cell differentiation. Nat. Rev. Immunol. 12: 749–761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Omilusik KD, Best JA, Yu B, Goossens S, Weidemann A, Nguyen JV, Seuntjens E, Stryjewska A, Zweier C, Roychoudhuri R, Gattinoni L, Bird LM, Higashi Y, Kondoh H, Huylebroeck D, Haigh J, and Goldrath AW. 2015. Transcriptional repressor ZEB2 promotes terminal differentiation of CD8 + effector and memory T cell populations during infection. J. Exp. Med. 212: 2027–2039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Omilusik KD, Nadjsombati MS, Shaw LA, Yu B, Milner JJ, and Goldrath AW. 2018. Sustained Id2 regulation of E proteins is required for terminal differentiation of effector CD8 + T cells. J. Exp. Med. 215: 773–783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Jameson SC, and Masopust D. 2018. Understanding Subset Diversity in T Cell Memory. Immunity 48: 214–226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Olson JA, McDonald-Hyman C, Jameson SC, and Hamilton SE. 2013. Effector-like CD8+ T Cells in the Memory Population Mediate Potent Protective Immunity. Immunity 38: 1250–1260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Beura LK, Hamilton SE, Bi K, Schenkel JM, Odumade OA, Casey KA, Thompson EA, Fraser KA, Rosato PC, Filali-Mouhim A, Sekaly RP, Jenkins MK, Vezys V, Nicholas Haining W, Jameson SC, and Masopust D. 2016. Normalizing the environment recapitulates adult human immune traits in laboratory mice. Nature 532: 512–516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Böttcher JP, Beyer M, Meissner F, Abdullah Z, Sander J, Höchst B, Eickhoff S, Rieckmann JC, Russo C, Bauer T, Flecken T, Giesen D, Engel D, Jung S, Busch DH, Protzer U, Thimme R, Mann M, Kurts C, Schultze JL, Kastenmüller W, and Knolle PA. 2015. Functional classification of memory CD8 + T cells by CX 3 CR1 expression. Nat. Commun. 6: 8306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gerlach C, Moseman EA, Loughhead SM, Garg R, De La Torre JC, Von UH, Correspondence A, Alvarez D, Zwijnenburg AJ, Waanders L, and Von Andrian UH. 2016. The Chemokine Receptor CX3CR1 Defines Three Antigen-Experienced CD8 T Cell Subsets with Distinct Roles in Immune Surveillance and Homeostasis Article The Chemokine Receptor CX3CR1 Defines Three Antigen-Experienced CD8 T Cell Subsets with Distinct Roles in Immune Surveillance and Homeostasis. Immunity 45: 1270–1284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ruiz AL, Soudja SM, Deceneux C, Lauvau G, and Marie JC. 2014. NK1.1+ CD8+ T cells escape TGF-β control and contribute to early microbial pathogen response. Nat. Commun. 5: 5150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Omilusik KD, and Goldrath AW. 2019. Remembering to remember: T cell memory maintenance and plasticity. Curr. Opin. Immunol. 58: 89–97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Herndler-Brandstetter D, Ishigame H, Shinnakasu R, Plajer V, Stecher C, Zhao J, Lietzenmayer M, Kroehling L, Takumi A, Kometani K, Inoue T, Kluger Y, Kaech SM, Kurosaki T, Okada T, and Flavell RA. 2018. KLRG1+ Effector CD8+ T Cells Lose KLRG1, Differentiate into All Memory T Cell Lineages, and Convey Enhanced Protective Immunity. Immunity 48: 716–729.e8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Haluszczak C, Akue AD, Hamilton SE, Johnson LDS, Pujanauski L, Teodorovic L, Jameson SC, and Kedl RM. 2009. The antigen-specific CD8 + T cell repertoire in unimmunized mice includes memory phenotype cells bearing markers of homeostatic expansion. J. Exp. Med. 206: 435–448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Jameson SC, and Masopust D. 2018. Understanding Subset Diversity in T Cell Memory. Immunity 48: 214–226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Arnon TI, Horton RM, Grigorova IL, and Cyster JG. 2013. Visualization of splenic marginal zone B-cell shuttling and follicular B-cell egress. Nature 493: 684–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Anderson KG, Mayer-Barber K, Sung H, Beura L, James BR, Taylor JJ, Qunaj L, Griffith TS, Vezys V, Barber DL, and Masopust D. 2014. Intravascular staining for discrimination of vascular and tissue leukocytes. Nat. Protoc. 9: 209–22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Guan T, Dominguez CX, Amezquita RA, Laidlaw BJ, Cheng J, Henao-Mejia J, Williams A, Flavell RA, Lu J, and Kaech SM. 2018. ZEB1, ZEB2, and the miR-200 family form a counterregulatory network to regulate CD8 + T cell fates. J. Exp. Med. 215: 1153–1168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Dominguez CX, Amezquita RA, Guan T, Marshall HD, Joshi NS, Kleinstein SH, and Kaech SM. 2015. The transcription factors ZEB2 and T-bet cooperate to program cytotoxic T cell terminal differentiation in response to LCMV viral infection. J. Exp. Med. 212: 2041–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lefrancois L 2006. Development, trafficking, and function of memory T-cell subsets. Immunol. Rev. 211: 93–103. [DOI] [PubMed] [Google Scholar]
- 23.Sallusto F, Geginat J, and Lanzavecchia A. 2004. Central Memory and Effector Memory T Cell Subsets : Function, Generation, and Maintenance. Annu. Rev. Immunol. 22: 745–763. [DOI] [PubMed] [Google Scholar]
- 24.Best JA, Blair DA, Knell J, Yang E, Mayya V, Doedens A, Dustin ML, Goldrath AW, Monach P, Shinton SA, Hardy RR, Jianu R, Koller D, Collins J, Gazit R, Garrison BS, Rossi DJ, Narayan K, Sylvia K, Kang J, Fletcher A, Elpek K, Bellemare-Pelletier A, Malhotra D, Turley S, Best JA, Jojic V, Koller D, Shay T, Regev A, Cohen N, Brennan P, Brenner M, Kreslavsky T, Bezman NA, Sun JC, Kim CC, Lanier LL, Miller J, Brown B, Merad M, Gautier EL, Jakubzick C, Randolph GJ, Kim F, Rao TN, Wagers A, Heng T, Painter M, Ericson J, Davis S, Ergun A, Mingueneau M, Mathis D, and Benoist C. 2013. Transcriptional insights into the CD8 + T cell response to infection and memory T cell formation. Nat. Immunol. 14: 404–412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Cui G, Staron MM, Gray SM, Ho P-C, Amezquita RA, Wu J, and Kaech SM. 2015. IL-7-Induced Glycerol Transport and TAG Synthesis Promotes Memory CD8+ T Cell Longevity. Cell 161: 750–61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Grayson JM, Zajac AJ, Altman JD, and Ahmed R. 2000. Cutting edge: increased expression of Bcl-2 in antigen-specific memory CD8+ T cells. J. Immunol. 164: 3950–4. [DOI] [PubMed] [Google Scholar]
- 27.Butler A, Hoffman P, Smibert P, Papalexi E, and Satija R. 2018. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36: 411–420. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Joshi NS, Cui W, Chandele A, Lee HK, Urso DR, Hagman J, Gapin L, and Kaech SM. 2007. Inflammation Directs Memory Precursor and Short-Lived Effector CD8+ T Cell Fates via the Graded Expression of T-bet Transcription Factor. Immunity 27: 281–295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Arsenio J, Kakaradov B, Metz PJ, Kim SH, Yeo GW, and Chang JT. 2014. Early specification of CD8 + T lymphocyte fates during adaptive immunity revealed by single-cell gene-expression analyses. Nat. Immunol. Vol. 15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Böttcher JP, Beyer M, Meissner F, Abdullah Z, Sander J, Höchst B, Eickhoff S, Rieckmann JC, Russo C, Bauer T, Flecken T, Giesen D, Engel D, Jung S, Busch DH, Protzer U, Thimme R, Mann M, Kurts C, Schultze JL, Kastenmüller W, and Knolle PA. 2015. Functional classification of memory CD8+ T cells by CX3CR1 expression. Nat. Commun. 6: 8306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Yang CY, Best JA, Knell J, Yang E, Sheridan AD, Jesionek AK, Li HS, Rivera RR, Lind KC, D’Cruz LM, Watowich SS, Murre C, and Goldrath AW. 2011. The transcriptional regulators Id2 and Id3 control the formation of distinct memory CD8+ T cell subsets. Nat. Immunol. 12: 1221–1229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ji Y, Pos Z, Rao M, Klebanoff CA, Yu Z, Sukumar M, Reger RN, Palmer DC, Borman ZA, Muranski P, Wang E, Schrump DS, Marincola FM, Restifo NP, and Gattinoni L. 2011. Repression of the DNA-binding inhibitor Id3 by Blimp-1 limits the formation of memory CD8+ T cells. Nat. Immunol. 12: 1230–1237. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.