Abstract
COVID-19, a disease induced by SARS-CoV-2 (Severe Acute Respiratory Syndrome Coronavirus-2), has been the cause of a worldwide pandemic. Though extensive research works have been reported in recent days on the development of effective therapeutics against this global health crisis, there is still no approved therapy against SARS-CoV-2. In the present study, plant-synthesized secondary metabolites (PSMs) have been prioritized to make a review focusing on the efficacy of plant-originated therapeutics for the treatment of COVID-19. Plant metabolites are a source of countless medicinal compounds, while the diversity of multidimensional chemical structures has made them superior to treat serious diseases. Some have already been reported as promising alternative medicines and lead compounds for drug repurposing and discovery. The versatility of secondary metabolites may provide novel antibiotics to tackle MDR (Multi-Drug Resistant) microbes too. This review attempted to find out plant metabolites that have the therapeutic potential to treat a wide range of viral pathogens. The study includes the search of remedies belonging to plant families, susceptible viral candidates, antiviral assays, and the mode of therapeutic action; this attempt resulted in the collection of an enormous number of natural therapeutics that might be suggested for the treatment of COVID-19. About 219 plants from 83 families were found to have antiviral activity. Among them, 149 plants from 71 families were screened for the identification of the major plant secondary metabolites (PSMs) that might be effective for this pandemic. Our investigation revealed that the proposed plant metabolites can serve as potential anti- SARS-CoV-2 lead molecules for further optimization and drug development processes to combat COVID-19 and future pandemics caused by viruses. This review will stimulate further analysis by the scientific community and boost antiviral plant-based research followed by novel drug designing.
Keywords: medicinal plants, secondary metabolites, antiviral activities, natural therapeutics/alternative medicine, drug discovery, COVID-19
Introduction
Coronaviruses comprise a group of large, enveloped, positive-sensed, single-stranded RNA viruses that damage the respiratory tract of mammals including humans, bats, and other animals, leading to infections in the respiratory tract (1–5). The Coronavirus disease 2019 (COVID-19), initially called 2019 novel coronavirus (2019-nCoV), is an agile respiratory disease caused by a novel coronavirus primarily detected in Wuhan, China (6, 7). Now, it has spread to 216 countries and caused the death of more than 0.5 million people worldwide and was declared as a pandemic by the World Health Organization (WHO) (8, 9). Seven types of human coronaviruses have been reported so far, including HCoV-OC43, HCoV-229E, HCoV-HKU1, HCoV-NL63, severe acute respiratory syndrome (SARS)-CoV, Middle East respiratory syndrome (MERS-CoV), and 2019-novel coronavirus nCoV (10). Among them, MERS-CoV, SARS-CoV, and nCoV have taken the concern of scientists worldwide. In 2003, the severe acute respiratory syndrome (SARS) outbreak occurred in Guangdong (southern China) (6, 11) which infected 8,000 people and resulted in 800 deaths in 26 countries. Only a decade later, another coronavirus has attacked the world and caused another devastating outbreak, MERS, which infected 2,494 people and caused the deaths of 858 worldwide (12, 13). However, the COVID-19 pandemic caused by SARS CoV-2 resulted in remarkable levels of morbidity and mortality all over the world. Initially China, followed by the USA, Italy, France, Iran, Spain, Russia, Turkey, and the UK became hotspots for SARS CoV-2. The virus hotspot has now moved to Latin America and, at this time, Brazil, Mexico, and Peru are the new hotspots of SARS CoV-2. The important aspects of the pathobiology, a viral response phase, and a hyperbolic host response phase are linked with the morbidity and mortality in COVID-19 patients (14). However, the increased cytokine levels (IL-6, IL-10, and TNF-α), lymphopenia (in CD4+ and CD8+ T cells), and decreased IFN-γ expression in CD4+ T cells are the more risky and possibly life-threatening events related to severe COVID-19 (15–17). The infection rate of COVID-19 is increasing gradually but scientists have not been able to suggest any specific drug, vaccine, or any other certified therapeutic agents against SARS-CoV-2, which consequently leads to the significant morbidity and mortality.
On the other hand, plants have been essential to human welfare for their uses as therapeutics since ancient times (18, 19). According to the WHO, about 80% of the world's population depends on medicinal plants or herbs to fulfill their medicinal needs (20–22). A significant amount of antiviral compounds produced from numerous kinds of plants have been used in many studies (23–25). Researchers all around the world are screening therapeutic drugs from existing antiviral plant secondary metabolites (PSMs) and are also trying to find novel compounds from medicinal plants [(26–159); Supplementary Table 1] to avert this global crisis. Plant metabolites can halt the activity of enzymes involved in the replication cycle of CoVs including papain-like protease and 3CL protease, halt the fusion of the S protein of coronaviruses and ACE2 of the host, and also inhibit cellular signaling pathways (123, 144, 160). Screening from existing PSMs, researchers have been trying to find novel compounds from medicinal plants to prevent numerous diseases, including COVID-19 (Supplementary Table 1). Therefore, the current manuscript aims to describe potential metabolites from plant sources that have antiviral properties that might be aligned for the alternative approach against COVID-19. Hence, understanding the structure, life cycle, pathogenicity, cell signaling, epidemiology of the recently emerging virus, drug targets, and drug discovery process have become very important issues to find specific/effective therapeutics.
Epidemiology, Genomic Organization, and Life Cycle of SARS CoV-2
In December 2019, SARS CoV-2, one of the most devastating viral outbreaks since SARS CoV and MERS, originated from Wuhan city seafood market in China (161–163). The virus was found to be transmitted through close contact with infected people or through exposure to coughing, sneezing, and respiratory droplets (164, 165). It has already been reported to have spread to 216 countries and caused more than 0.5 million deaths. Brazil is now the new hotspot for SARS CoV-2 after the USA, Russia, France, Italy, Germany, Spain, and the UK, where more than 11 million people are infected (166, 167).
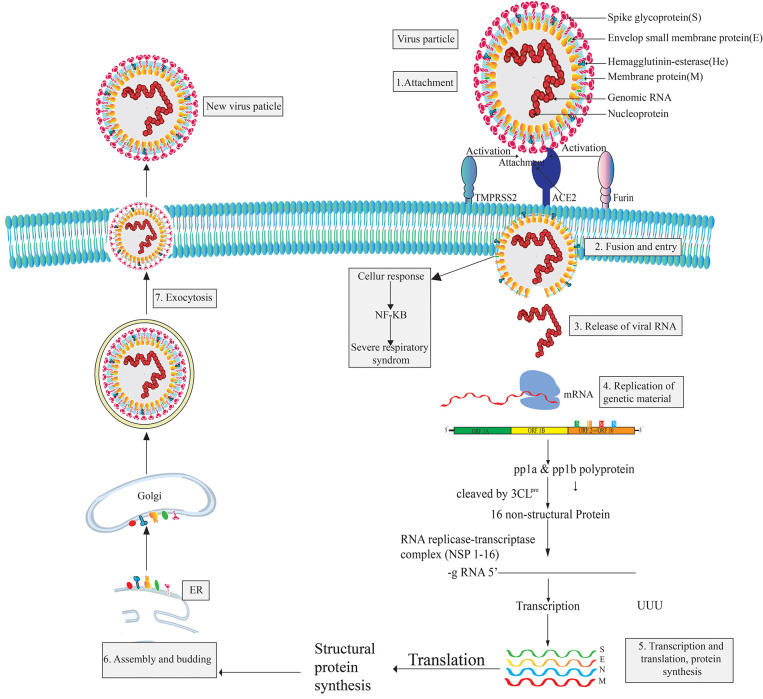
The pleomorphic or spherical shaped SARS COV-2 has a single-stranded RNA genome of 26.4–31.7 kb in length and a crown-like glycoproteins on its surface (168–173). It is more similar to SARS CoV (over 80%) than MERS (174, 175). However, the RNA genome of CoV-2 is considered as one of the largest genomes compared to those of other RNA viruses (176, 177). The largest open reading frame, ORF1ab, encodes non-structural proteins while the remaining ORFs encode four structural proteins, namely the envelope glycoprotein or spike protein (S), envelope (E) protein, membrane (M) protein, and nucleocapsid (N) protein. The S protein mediates attachment to the host cell while the E protein is involved in virus assembly, membrane permeability of the host cell, and virus-host cell interaction. The M protein is known as a central organizer for the coronavirus assembly and the nucleocapsid (N) protein is usually involved in the processing of helical ribonucleocapsid complex, including some accessory proteins (172, 178). Six types of mutations are found in the genome of SARS CoV-2 while three mutations have been reported in orf 1ab gene, two mutation in S gene, and the final one in the orf 7b and orf 8 (174, 175). Proteomic analysis revealed that SARS CoV-2 is vastly homologous to SARS CoV but two proteins, orf 8 and orf 10, are not homologous to SARS CoV (175). To complete its life cycle, SARS CoV-2 passes into the human body through the nose, mouth, or eyes and then attaches itself to the receptor-binding domain (RBD) using the surface glycoprotein (Spike-protein) of the virion which tries to attach with the hACE2 receptor (179, 180). The entry mechanism of SARS CoV-2 depends on cellular transmembrane serine protease 2 (TMPRSS2) and furin, along with viral receptor ACE2 (180–182). However, after the fusion of the SARS CoV-2 virion particle with the host cell membrane, the envelope and capsid part of the virus are removed. The virus releases its genetic material (RNA) into the host cell cytoplasm and acts as mRNA for the translation from ORF1a and ORF1ab to produce pp1a and pp1ab polypeptides (169, 183). Subsequently, chymotrypsin-like protease (3CLpro) slices these polypeptides into 16 non-structural proteins (NSPs) that are responsible for replication and transcription (184). Then, infected cells produce proteins when they become hijacked by SARS CoV-2. In this situation, the immune system supports the assembly of SARS CoV-2 into new copies of virion particles (185, 186). Freshly synthesized viral nucleic acids and proteins then assemble into the lumen of the ERGIC (Endoplasmic Reticulum Golgi Intermediate Compartment) and leave the cells through exocytosis [(187, 188); Figure 1]. Infected cells release virions and infect other human cells.
Figure 1.
Structure, genomic organization, life cycle, and drug targets of SARS CoV-2.
SARS-CoV-2 viral infection can be divided into three stages: the asymptomatic period, non-severe symptomatic period, and the severe infection stage (17, 189). SARS CoV-2 patients are reported to have a significant amount of cytokines and chemokines; the levels of cytokines are especially highly increased in patients admitted to ICUs (Intensive Care Unit) (190, 191). These significantly high levels are what results in a patient reaching a critical stage. However, the main mediator of SARS CoV-2, the spike glycoprotein, is found in two conformations (192) and the enzyme 3CLpro of SARS-CoV-2 share a 99.02% sequence identity with 3CLpro of SARS-CoV, which is also highly similar to bat SARS CoV 3CLpro (193). SARS CoV-2 binds to the host cell receptor with a higher affinity than SARS CoV (194). SARS CoV-2 has shown some strategic alteration with the substrate-binding site of bat SARS CoV-2 and 12 point-mutations are found in SARS CoV-2 compared to SARS CoV. Mutations disrupt the significant hydrogen bonds and modify the receptor binding site (RBS) of SARS-CoV-2 3CLpro. However, the occurrence of recurrent mutations can lead to new strains with alterations in virulence, which one of the reasons discovering a suitable vaccine to combat SARS CoV-2 is challenging (175, 195).
Major Drug Targets of SARS CoV-2
A fundamental therapeutic approach to treat multi-viral infections is the interruption of human host-virus interactions (17). The major structural proteins of SARS CoV-2 can be obvious targets for drugs designed against COVID-19. In addition, 16 non-structural proteins (NSPs) can also be considered (169). However, the manifestation of recurrent recombination events is a major hindrance to develop SARS CoV-2 specific vaccines/drugs (176). Up-to-date studies revealed that, though SARS-CoV-2 and SARS-CoV identify a similar receptor (ACE2) in humans (194, 196), there is a noteworthy variation in the antigenicity between SARS-CoV and SARS-CoV-2 which has significance on the development of therapeutic options against this rapidly emerging virus (197). The SARS-CoV-2 spike protein exhibits a higher affinity to the ACE2 receptor in comparison to SARS-CoV, but hACE2 showed a lower binding affinity to RBD (Receptor Binding Domain) of SARS COV-2 when compared to SARS CoV (194, 198). The two most paramount enzymes of SARS CoV-2, proprotein convertase furin- potentiates cell fusion and serine protease TMPRSS2, are responsible for S-protein activation and are propitious drug targets for the treatment of COVID (180, 194, 199).
Sars-CoV-2 and Searching for Effective Therapeutics
Though extensive research works are being continued for the development of effective vaccines or drug compounds against SARS-CoV-2, efficacious therapeutics have not yet been attained (200). Moreover, interferon therapies, monoclonal antibodies, oligonucleotide-based therapies, peptides, small-molecule drugs, and vaccines, are regarded as some strategic approaches for controlling or preventing COVID-19 (201, 202). Existing drugs can be used as the first-line treatment for coronavirus outbreaks, but this is not the ultimate solution to eradicate the disease (203). Therefore, the development of therapeutic drugs for the treatment of the COVID-19 outbreak have gathered considerable attention. Scientists from different fields are trying to figure out the way to develop therapeutics. However, experimental implications of drug recombination might be both expensive and time-consuming, whereas computational evaluation may bring about testable hypotheses for systematic drug recombination (174).
PSMs Can be Effective Over Synthetic Drugs Against SARS CoV-2
Though there are approved, repurposed drugs currently in clinical use, there is still an urgent need for specific antiviral therapeutics and vaccines (199). Bioengineered and vectored antibodies and therapies based on cytokines and nucleic acid which target virus gene expression have been found as promising to treat coronavirus infections (204). For example, the repurposing drugs, including favipiravir, remdesivir, lopinavir, ritonavir, nebulized α-interferon, chloroquine, hydroxychloroquine, ribavirin, and interferon (IFN), have been shown to be effective for the treatment of COVID-19. Apart from this, some therapeutics are in clinical trials, such as peptide vaccine (mRNA-1273) (198) and antibody therapies (205). Recently, plasma therapy showed promising results for COVID-19 treatment (206, 207). But, application of these synthetic drugs are not efficient as they exhibit adverse direct or indirect side effects [(208–220); Table 1]. In addition, scientists all around the world are trying to find out some prominent drug and multi-epitope vaccine candidates against this deadly virus using various kinds of immuno-informatics approaches (221, 222). Therefore, the urgent need for safe, effective, and inexpensive therapies/drugs with negligible side effects against COVID-19 is imperative.
Table 1.
Recently used synthetic drugs and their side effects.
| Drug | Side effects | References |
|---|---|---|
| Arbidol | Side effects in children include sensitization to the drug | (208) |
| Darunavir | Liver problems and severe skin reactions or rash | (209) |
| Flavipir | – | (210) |
| Hydroxychloroquine | One of the most serious side effects of hydroxychloroquine is a risk of heart rhythm problems, which can result in heart failure and in some cases death. Hydroxychloroquine can upset the stomach. Severe, permanent damage to the retina has been reported with the use of hydroxychloroquine | (211) |
| Ivermectin | Eye or eyelid irritation, pain, redness, or swelling | (212) |
| Lopinavir | Drowsiness, dizziness, a bad taste in the mouth, and trouble sleeping | (213) |
| Loprazolam | Paradoxical increase in aggression, lightheadedness, blood disorders, and jaundice | (214) |
| Lurasidone | Drowsiness, lightheadedness, weight gain, mask-like facial expression, and agitation | (214) |
| Oseltamivir | Phlegm-producing cough, wheezing, abdominal or stomach cramps or tenderness, bloating | (215) |
| Remdesivir | Increased liver enzyme levels that may indicate possible liver damage | (209, 211) |
| Ribavirin | Allergic reaction, anemia, stabbing chest pain, wheezing | (220) |
| Ritonavir | Diarrhea, nausea, vomiting, heartburn, stomach pain, dizziness, tiredness | (215) |
| Salmeterol | Hoarseness, throat irritation, rapid heartbeat, cough, dry mouth/throat, or upset stomach | (217) |
| Saquinavir | Hyperglycemia, increased bleeding in people with hemophilia, increases in the levels of certain fats | (209) |
| Talampicillin | – | (214) |
| Teicoplanin | Maculopapular or erythematous rash and drug-related fever | (218) |
| Andrographolide (PSM) | – | (219) |
| Rubitecan | – | (214) |
PSMs are a source of natural antiviral compounds that could be an effective option, as most of them are safer and more cost-effective compared to orthodox drugs (223), though some PSMs are toxic too. The dependency on and popularity of plant-based drugs are increasing day by day (224). Due to the presence of multiple compounds in crude plant extracts, it can be either beneficial or not, depending on the amounts used each time; if properly regulated, better activity might be shown. It was also found that crude extracts can target multiple sites at a time in a virion particle (225). However, this is yet to be tested against SARS-CoV-2. PSMs can affect the disruption of cell membrane functions and structures (226), interference with intermediary metabolisms (227), interruption of DNA/RNA synthesis and function (228), interruption of normal cell communication (quorum sensing) (229), and the induction of coagulation of cytoplasmic constituents (230). Different kinds of plant metabolites act against SARS CoV (Supplementary Table 1). Plant-based products affect several key events in the pathogenic process. For example, curcumin is effective for its antineoplastic, anti-proliferative, anti-aging, anti-inflammatory, anti-angiogenic, antiviral and anti-oxidant effects, and can regulate redox status, protein kinases, transcription factors, adhesion molecules, and cytokines in the human body (231). In silico analysis revealed that anti-SARS CoV PSMs could be one of the most valuable drug targets against SARS CoV-2 [(232); Table 2]. A huge amount of plant metabolites have remained unexplored due to the extensive process of isolation of the target compound. Now, various types of modern techniques have been developed for the isolation of lead compounds from crude extracts including maceration, percolation, decoction, reflux extraction, soxhlet extraction, pressurized liquid extraction, supercritical fluid extraction, ultrasound assisted extraction, microwave-assisted extraction, pulsed electric field extraction, enzyme assisted extraction, hydro distillation, and steam distillation (179). These techniques can lead us to find out novel anti-SARS CoV-2 compounds earlier than traditional techniques. In addition, plant metabolomics are used as a tool for the discovery of novel drugs from plant resources (262, 263).
Table 2.
Probable drug candidates against SARS CoV-2 obtained through virtual screening.
| Drug targets | Major metabolites | References |
|---|---|---|
| ANTIVIRAL PSMs THAT CAN INHIBIT SARS CoV-2 AT DIFFERENT TARGET | ||
| Spike protein | Magnoflorine, tinosponone, cirsimaritin, chrysoeriol, vasicinone, quercetin, luteolin | (233) |
| Spike protein | Epigallocatechingallate (EGCG), curcumin, apigenin, chrysophanol | (234) |
| Spike protein, main protease | Spike protein, main protease | (235) |
| Spike protein and ACE-2 | Hesperidin, emodin, and chrysin | (236) |
| Spike protein and ACE-2 | Curcumin, nimbin, withaferin A, piperine, mangiferin, thebaine, berberine, and andrographolide | (222) |
| Spike protein and ACE-2 | Chebulagic acid | (237) |
| Spike protein, MPro, and RdRp | Silybin, withaferin A, cordioside, catechin, and quercetin | (238) |
| RdRp | Protopine, allocryptopine, and (±) 6-acetonyldihydrochelerythrine | (239) |
| Main Protease (MPro) | Crocin, digitoxigenin, and b–eudesmol | (240) |
| Main Protease (MPro) | Oolonghomobisflavan-A, theasinensin D, theaflavin-30-O-gallate | (241) |
| Main Protease (MPro) | Andrographolide | (219) |
| Main Protease (MPro) | Hispidin, lepidine E, and folic acid | (242) |
| Main Protease (MPro) | Ursolic acid, carvacrol, and oleanolic acid | (243) |
| Main Protease (MPro) | Hypericin, cyanidin 3-glucoside, baicalin, glabridin | (244) |
| Main Protease (MPro) | Cetylglucopetunidin, isoxanthohumol, and ellagic acid | (245) |
| Main Protease (MPro) | Benzylidenechromanones | (246) |
| Main Protease (MPro) | Carnosol, arjunglucoside-I, and rosmanol | (247) |
| Main Protease (MPro) | Leucoefdin | (248) |
| Main Protease (MPro) | (1E,6E)-1,2,6,7-tetrahydroxy-1,7-bis(4-hydroxy-3-methoxyphenyl)hepta-1,6-diene-3,5-dione) and (4Z,6E)-1,5-dihydroxy-1,7-bis(4-hydroxyphenyl)hepta-4,6-dien-3-one | (249) |
| Mpro and ACE2 | Quercetin 3-glucuronide-7-glucoside, and Quercetin 3-vicianoside | (250) |
| Mpro, hACE-2 and RdRp | d-Viniferin, myricitrin, chrysanthemin, myritilin, taiwanhomoflavone A, lactucopicrin 15-oxalate, nympholide A, afzelin, biorobin, hesperidin, and phyllaemblicin B | (251) |
| Mpro, spike protein, and non-structural proteins (NSP-9, 15) | Arzanol, ferulic acid, genistein, resveratrol, rosmanol | (252) |
| ACE-2 receptor | Resveratrol, pterostilbene, pinosylvin, piceatannol | (253) |
| ACE-2 receptor | Isothymol, chloroquine, captopril | (254) |
| ACE-2 receptor | Resveratrol, quercetin, luteolin, naringenin, zingiberene, and gallic acid | (222) |
| Envelope protein | Belachinal, macaflavanone E, vibsanol B | (249) |
| PLpro, 3CLpro | Cryptotanshinone, quercetin, tanshinone IIa, coumaroyltyramine, N-cis-feruloyltyramine | (178) |
| PLpro, 3CLpro, RdRp, and spike protein | Andrographolide (AGP1), 14-deoxy 11,12-didehydro andrographolide (AGP2), neoandrographolide (AGP3), and 14-deoxy andrographolide (AGP4) | (255) |
| 3CLpro | 10-hydroxyusambarensine, cryptoquindoline, 6-oxoisoiguesterin, 22-hydroxyhopan-3-one, cryptospirolepine, isoiguesterin, and 20-epibryonolic acid | (256) |
| 3CLpro | Flavone and coumarine | (209) |
| 3CLpro | Myricitrin, methyl rosmarinat, calceolarioside B, licoleafol, amaranthin, colistin | (191) |
| 6LU7 and 6Y2E proteases | Apigenin, glabridin, glycoumarin, oleanolic acid, glucobrassicin | (257) |
| Transmembrane protease serine 2 (TMPRSS2) | Withanone and withaferin-A | (258) |
| Membrane (M) and Envelope (E) proteins | Nimbolin A, nimocin, and cycloartanols | (259) |
| ANTIVIRAL PSMs THAT CAN INHIBIT SARS CoV-2 AT DIFFERENT LIFE CYCLE | ||
| Viral attachment | Phytoestrogens (diadiazin, genistein, formontein, and biochanin A), chlorogenic acid, linolenic acid, palmitic acid, caffeic acid, caffeic acid phenethyl ester, hydroxytyrosol, cis-p-Coumaric acid, cinnamaldehyde, thymoquinone, and some physiological hormones such as estrogens, progesterone, testosterone, and cholesterol | (260) |
| Entry | Dihydrotanshinone – 1, desmethoxyreserpine | (241) |
| Multiplication | Betulinic acid, desmethoxyreserpine, lignan, sugiol | (241) |
| Viraus–host interaction | Dithymoquinone (DTQ) | (261) |
PSMs Having Antiviral Properties as Alternatives to Synthetic Drugs and Hope For CoVID-19
Plants produce diversified low molecular weight PSMs to protect them from different herbivores and microbes (264). Before the discovery of allopathic drugs, these leading natural sources were extensively used for treating several kinds of human diseases (265, 266). Due to the increased resistance of microbial pathogens against allopathic drugs, researchers have now returned to natural resources, focusing especially on plant metabolites, to find out lead compounds to fight against human pathogens (175). Moreover, about 35% of the global medicine market (which accounts for 1.1 trillion US dollars) have been shared by medicinal products prepared using natural plants or herbs (265). Investigations are undergoing for the finding of novel and modern drugs from numerous herbal preparations to fight against this microbial resistance war. Many similarities have been found between SARS CoV and SARS CoV-2 (both of them belong to beta family, containing the same genetic material-RNA, and using the same receptor for viral attachment-ACE2, with an 86% identity and 96% similarity of genome, with almost the same pathogenesis). Thus, previously reported antiviral plant metabolites for SARS CoV can be considered as emerging drug candidates for COVID-19. Right now, the setbacks arising from viral infection around the world have placed budget constraints on researchers trying to discover effective antiviral drugs. However, some PSMs have already shown anti-SARS CoV activity where other antiviral activities are also reported (Supplementary Table 1). These results suggest that there is a scope to find alternative medicines and specific compounds. So, plants could be a vital resource in the fight against COVID-19. Our study suggests that around 76 natural metabolites from different plant species can be efficiently active against COVID-19 (Table 3 and Supplementary Figure 1).
Table 3.
Probable promising secondary metabolites of medicinal plants against COVID-19.
| Compounds | Plant source | Family | References | |
|---|---|---|---|---|
| 1. | Diterpneoid | Andrographis paniculata | Acanthaceae | (26) |
| 2. | Alkaloids, flavonoids, and coumarins | Sambucus nigra | Adoxaceae | (29) |
| 3. | Alkaloids, anthraquinones, glycosides, flavonoids, saponins, phenols, terpenoids, sugar bearing compound, protein, thiols, and inferences | Iresine herbstii | Amaranthaceae | (31) |
| 4. | Tannins, Flavonoids, Terpenes, and Saponins | Anacardium occidentale | Anacardiaceae | (33) |
| 5. | Tannins, gallic acid, flavonoids like quercetin and quercitrin, phenolics, triterpenes | Rhus aromatica | Anacardiaceae | (34) |
| 6. | Gallic acid, quercetin, kaempferol, glycosides | Rhus parviflora | Anacardiaceae | (35) |
| 7. | Tannins and flavonoids | Spondias lutea | Anacardiaceae | (33) |
| 8. | Flavonoids | Spondias lutea L. | Anacardiaceae | (33) |
| 9. | Apigenin and luteolin | Arisaema tortuosum | Araceae | (40) |
| 10. | Phenolic acids, flavonoids (apigenin, apigeninglucoside, luteolin, cirsiliol, diosmetin), lignans, terpenic lactones, and alkamides | Achillea fragrantissima | Asteraceae | (47, 48) |
| 11. | Flavonoids, clerodane diterpenoids, phenolics, hydroxycinnamic acids | Baccharis gaudichaudiana DC | Asteraceae | (49) |
| 12. | Diterpenoids | Baccharis spicata (Lam.) Baill | Asteraceae | (49) |
| 13. | Triterpenoids, Steroids | Bidens subalternans DC | Asteraceae | (49) |
| 14. | Flavonoid glycosides and caffeoyl quinic acids | Eupatorium perfoliatum | Asteraceae | (50) |
| 15. | Flavonoids and terpenes | Jasonia montana | Asteraceae | (47) |
| 16. | Phenylpropanoids, flavonoids, essential oils, polyphenols, tannins, triterpenes | Pluchea sagittalis (Lam.) Cabrera | Asteraceae | (49) |
| 17. | Silymarin, quercetin, and kaempferol | Silybum marianum | Asteraceae | (51) |
| 18. | terpenoids, flavonoids, essential oils | Tagetes minuta L. | Asteraceae | (49) |
| 19. | phenolic acids (chlorogenic acids), and sesquiterpene lactones (parthenolide) | Tanacetum parthenium | Asteraceae | (52) |
| 20. | Flavonoids, D-glucopyranoside, quercetin, luteolin | Taraxacum officinale | Asteraceae | (53) |
| 21. | Flavonoids (apigenin, quercetin, kaempferol, falcarinol, selinene, limonene, and zerumbone) | Tridax procumbens | Asteraceae | (55) |
| 22. | Carbohydrates, lipids, proteins, alkaloids, flavonoids, saponins, and organic acids | Balanites aegyptiaca | Balanitaceae | (56, 57) |
| 23. | Icariin and quercetin | Epimedium koreanum Nakai | Berberidaceae | (58) |
| 24. | Flavonoids (quercetin, isoquercetin, and rutin) | Capparis sinaica | Capparaceae | (47, 64) |
| 25. | Tannins, flavonoids, carbohydrates and/or glycosides, resins, sterol, saponins, and alkaloids | Capparis sinaica | Capparaceae | (47, 65) |
| 26. | Natural lupane triterpenoids | Cassine xylocarpa | Celastraceae | (67) |
| 27. | Pentacyclic lupane-type triterpenoids | Maytenus cuzcoina | Celastraceae | (67) |
| 28. | Flavonoids, terpenoids, alkaloids, tannins, glycosides, and saponins | Combretum adenogonium | Combretaceae | (72) |
| 29. | Triterpenes, flavonoids, ellagitannins | Terminalia mollis | Combretaceae | (56, 73) |
| 30. | Lignans, diterpenes, flavonoids, proanthocyanidins, and sterols | Taxodium distichum | Cupressaceae | (75) |
| 31. | Monoterpenoids, sesquiterpenoids, triterpenoids, sterols, alkaloids, flavonoids, and phenolic compounds | Cyperus rotundus | Cyperaceae | (76) |
| 32. | Protocatecuic acid, caffeic acid, epicatechin, rutin, resveratrol, quercitin, kaempferol | Ephedra alata | Ephedraceae | (47, 77) |
| 33. | Isoflavonoid, indoles, phytosterols, polysaccharides, sesquiterpenes, alkaloids, glucans, and tannins | Equisetum giganteum | Equisetaceae | (78) |
| 34. | Triterpenes and steroids | Euphorbia denticulata | Euphorbiaceae | (79) |
| 35. | Tannins, diterpenes | Euphorbia hirta | Euphorbiaceae | (80) |
| 36. | Diterpenoids, jatrophane-type diterpenoids, and coumarino-type lignoids, lathyrane-type diterpenoids, multifidone, multifidanol, and multifidenol | Jatropha multifida | Euphorbiaceae | (82) |
| 37. | Flavonoid and polyphenol | Acacia arabica | Fabaceae | (83) |
| 38. | Luteolin and vitexin | Aspalathus linearis | Fabaceae | (85) |
| 39. | Saponins and flavonoids | Vachellia nilotica | Fabaceae | (87) |
| 40. | Catechin, kaempferol, quercetin, 3,4′,7-trihydroxyl-3′,5-dimethoxyflavone, rutin, isorhamnetin, epicatechin, afzelechin, epiafzelechin, mesquitol, ophioglonin, aromadendrin, and phenol | Acacia catechu | Fabaceae | (88) |
| 41. | Flavonoids, phenolics, and tannins | Quercus persica | Fagaceae | (90) |
| 42. | Phenolic, flavonoid, and flavonol compounds | Quercus persica | Fagaceae | (90) |
| 43. | Gallic acid, protocatechuic acid, corilagin, geraniin, ellagic acid, kaempferitrin, kaempferol 7-O-rhamnoside, quercetin, kaempferol | Geranium thunbergii | Geraniaceae | (91) |
| 44. | Flavonoids (orientin and vicenin) | Ocimum sanctum | Lamiaceae | (26, 99) |
| 45. | Terpenoid and polyphenol | Ocimum sanctum | Lamiaceae | (83) |
| 46. | Baicalin, flavonoids | Scutellaria baicalensis | Lamiaceae | (104) |
| 47. | Opuntin B, triterpene saponin, seroids, and phenylethanoids | Lindernia crustacea | Linderniaceae | (107) |
| 48. | Quercetin 3-O-methyl ether (3MQ) and strychnobiflavone (SBF) | Strychnos pseudoquina | Loganiaceae | (108) |
| 49. | Alkaloids, flavonoids, tannins, volatile oils, and glycosides | Cissampelos pareira Linn | Menispermaceae | (113) |
| 50. | Flavonoids, tannins, terpenes, saponins, and nitrogenous compounds | Artocarpus integrifolia | Moraceae | (33) |
| 51. | Flavonoids, rutin, kaempferol 3-O-rutinoside, and kaempferol 3-O-robinobioside | Ficus benjamina | Moraceae | (114) |
| 52. | N-arginine, luteolin, caffeic acid | Ficus carica | Moraceae | (115) |
| 53. | Flavonoids, tannins, saponins, alkaloids, and steroids/triterpenoids | Ficus religiosa | Moraceae | (116) |
| 54. | Tannins, flavonoid, saponin, glycoside | Ficus sycomorus | Moraceae | (56, 118) |
| 55. | Alkaloids, tannins, phenolics, and saponins | Moringa peregrina | Moringaceae | (47) |
| 56. | Flavonoids | Myristica fragrans | Myristicaceae | (33) |
| 57. | Tannins and flavonoids | Psidium guajava | Myrtaceae | (33) |
| 58. | Sesquiterpenes, monoterpenes, hydrocarbon, and phenolic compounds, eugenyl acetate, eugenol, and β-caryophyllene | Syzygium aromaticum L. | Myrtaceae | (119) |
| 59. | Paeoniflorin, monoterpene glycosides, albiflorin, benzoylpaeoniflorin, gallic acid, ethyl gallate | Paeonia delavayi | Paeoniaceae | (121) |
| 60. | Flavonoids, tomentin A, B, C, D, and E | Paulownia tomentosa | Paulowniaceae | (123) |
| 61. | Highly oxygenated norbisabolane sesquiterpenoids, phyllanthacidoid acid, methyl ester | Phyllanthus acidus | Phyllanthaceae | (124) |
| 62. | Alkaloids, flavonoids, lignans, phenols, and terpenes | Phyllanthus amarus | Phyllanthaceae | (125) |
| 63. | Geraniin, rutin, gallic acid, caffeolquinic acid, corilagen, galloylglucopyronoside, digalloylglucopyronoside, and quercetin glucoside | Phyllanthus amarus | Phyllanthaceae | (126) |
| 64. | Geraniin, rutin, gallic acid, caffeolquinic acid, corilagen, galloylglucopyronoside, digalloylglucopyronoside, and quercetin glucoside | Phyllanthus niruri | Phyllanthaceae | (126) |
| 65. | Trigalloylglucopyronoside, quercetin rhamnoside, geraniin, rutin, gallic acid, caffeolquinic acid, corilagen, galloylglucopyronoside, digalloylglucopyronoside, and quercetin glucoside | Phyllanthus urinaria | Phyllanthaceae | (126) |
| 66. | Quercetin rhamnoside, geraniin, rutin, gallic acid, caffeolquinic acid, corilagen, galloylglucopyronoside, digalloylglucopyronoside, and quercetin glucoside | Phyllanthus watsonii | Phyllanthaceae | (126) |
| 67. | Plumbagin, allicin, carbohydrates, flavonoids, proteins, saponins, fats and oils, alkaloids, steroids, phenols, and tannins | Plumbago indica | Plumbaginaceae | (129) |
| 68. | Flavonoids (catechin, hyperoside, quercitrin, quercetin, and rutin), tannins, and triterpenoids | Agrimonia pilosa | Rosaceae | (135) |
| 69. | Hydroxycinnamic acids, eriodictyol, isorhamnetin, quercetin, kaempferol, isorhamnetin, epicatechin, catechin | Prunus dulcis | Rosaceae | (136) |
| 70. | Saponins, flavonoids, and alkaloids | Pavetta tomentosa | Rubiaceae | (138) |
| 71. | Saponins, flavonoids, and alkaloids | Tarenna asiatica | Rubiaceae | (138) |
| 72. | Triterpenes, tannins, flavonoids, and carbohydrates | Dimocarpus longan | Sapindaceae | (140) |
| 73. | Organic acids, terpenoids, and flavonoids | Illicium verum Hook. f. | Schisandraceae | (142) |
| 74. | Nilocitin, ellagic acid, gallic acid, flavonoids | Tamarix nilotica | Tamaricaceae | (47, 143) |
| 75. | Diterpenoids, biflavonoids (biflavone amentoflavone, apigenin, luteolin, and quercetin) | Torreya nucifera | Taxaceae | (144) |
| 76. | Friedelolactones, 2β-hydroxy-3, 4-seco-friedelolactone-27-oic acid flavonoids, coumarins, terpenoids, sterols, polypeptides | Viola diffusa | Violaceae | (147) |
Plant-Based Antiviral Compounds: Group Basis Mechanism of Action and PSMs Structure
A wide variety of antiviral compounds were found from 219 medicinal plants (26–159) belonging to 83 plant families (Supplementary Table 1). First and foremost are polyphenols, which contain multiple phenolic rings, and are classified as phenols, flavonoids, lignans, hydroxycinnamic acid, stilbenes, and hydroxybenzoic acid (267). We found polyphenols in numerous plants (Table 4) which exerted antiviral activity (269–271) against a wide range of viruses including HIV-1, HIV-2, HSV-1, HSV-2, Influenza virus, Dengue virus, HBV, HCV, Infectious bronchitis virus (IBV), Murbarg virus, Ebola virus, Newcastle disease virus (NDV), Poliomyelitis-1 virus, Lentivirus, and Coronavirus. Polyphenols work against coronaviruses using diverse mechanisms including actuating or inhibiting cellular signaling pathways or halting papain-like protease (PLpro) and 3-chymotripsin-like protease (3CLpro) enzyme (269, 272). Some polyphenol compounds (30-(3-methylbut-2-enyl)-30, 4-hydroxyisolonchocarpin, broussochalcone A, 4,7-trihydroxyflavane, broussochalcone B, papyriflavonol A, kazinol A, kazinol B, kazinol F, kazinol J, and broussoflavan A) isolated from Broussonetia papyrifera showed promising activity against SARS CoV. Higher efficiency against PLpro as observed by these compounds though activity against Mpro or 3CLpro is not up to the mark. Specially, papyriflavonol A possesses impressive activity against SARS CoV (IC50 3.7, l M) (272). In silico analysis revealed that polyphenols can inhibit SARS CoV-2 Mpro and RdRp effectively (273, 274). In our study, we have found another widely distributed, low molecular weight phenolic compound named as a flavonoid which showed strong antiviral activity against SARS CoV, Influenza virus, HBV, HSV, HCV, HIV, Dengue virus, Simian virus, Human rotavirus, Bovine viral diarrhea virus, Poliomyelitis-1 virus, Vesicular stomatitis virus (VSV), and Newcastle disease virus (NDV) (Table 4). Flavonoid type compounds, such as apigenin and quercetin, showed activity against SARS CoV virion particles through the inhibition of Mpro enzymes with an IC50 of 38.4 ± 2.4 μM and 23.8 μM, respectively (144, 150, 275). According to in silico analysis, flavonoid compounds can terminate the activity of Mpro of SARS CoV-2 (276, 277).
Table 4.
Major group basis antiviral PSMs obtained from medicinal plants.
| Major compounds | Plant source | Family | Target pathogen | References |
|---|---|---|---|---|
| Polyphenols | Avicennia marina | Acanthaceae | Human immunodeficiency virus (HIV) and herpes simplex virus (HSV) | (27) |
| Sambucus nigra | Adoxaceae | Dengue virus serotype-2 (DENV-2) | (29) | |
| Sambucus nigra | Adoxaceae | Infectious bronchitis virus (IBV)—chicken coronavirus | (30) | |
| Iresine Herbstii | Amaranthaceae | Newcastle disease virus (NDV) | (31) | |
| Anacardium occidentale | Anacardiaceae | Simian (SA-11) virus | (33) | |
| Artocarpus integrifolia | Moraceae | (SA-11) and human (HCR3) rotaviruses | (33) | |
| Myristica fragrans | Myristicaceae | Human (HCR3) rotaviruses | (33) | |
| Psidium guajava | Myrtaceae | Simian (SA-11) virus | (33) | |
| Spondias lutea | Anacardiaceae | Human (HCR3) rotaviruses | (33) | |
| Spondias lutea L. | Anacardiaceae | Simian (SA-11) and human (HCR3) rotaviruses | (33) | |
| Rhus aromatica | Anacardiaceae | HSV-1 and HSV-2 | (34) | |
| Rhus aromatica | Anacardiaceae | HSV-1 and HSV-2 | (34) | |
| Rhus parviflora | Anacardiaceae | HIV-1 | (35) | |
| Schinus terebinthifolia | Anacardiaceae | HSV-1 | (36) | |
| Arisaema Tortuosum | Araceae | Acyclovir-resistant HSV-2 and HSV-1 | (40) | |
| Jasonia montana | Asteraceae | Poliomyelitis-1 virus | (47) | |
| Baccharis gaudichaudiana DC | Asteraceae | Bovine viral diarrhea virus, HSV-1, Poliovirus type 2 (PV-2), and vesicular stomatitis virus (VSV) | (49) | |
| Pluchea sagittalis (Lam.) Cabrera | Asteraceae | Bovine viral diarrhea virus (BVDV) (HSV-1), poliovirus type 2 (PV-2), and vesicular stomatitis virus (VSV) | (49) | |
| Tagetes minuta L | Asteraceae | Bovine viral diarrhea virus, HSV-1, poliovirus type 2 (PV-2), and vesicular stomatitis virus | (49) | |
| Eupatorium perfoliatum | Asteraceae | Influenza A virus (IAV) H1N1 | (50) | |
| Silybum marianum | Asteraceae | Chikungunya virus (CHIKV), hepatitis C virus (HCV) | (51) | |
| Tanacetum parthenium | Asteraceae | HSV-1 | (52) | |
| Taraxacum officinale | Asteraceae | HCV | (53) | |
| Senna angustifolia | Fabaceae | Dengue virus serotype-2 (DENV-2) | (55) | |
| Tridax procumbers | Asteraceae | Dengue virus serotype-2 (DENV-2) | (55) | |
| Vernonia cinerea | Asteraceae | Dengue virus serotype-2 (DENV-2) | (55) | |
| Epimedium koreanum Nakai | Berberidaceae | Porcine epidermic diarrhea virus (PEDV) | (58) | |
| Canarium album (Lour.) | Burseraceae | Influenza A virus (IAV) | (62) | |
| Polyphenols | Cistus incanus | Cistaceae | HIV (clinical HIV-1 and HIV-2) and Filoviruses, Ebola, and Marburg virus | (69) |
| Combretum adenogonium | Combretaceae | HIV-1 | (72) | |
| Cornus canadensis | Cornaceae | HSV-1 | (74) | |
| Taxodium distichum | Cupressaceae | Influenza A and B viruses | (75) | |
| Cyperus rotundus | Cyperaceae | HSV-1, HBV | (76) | |
| Equisetum giganteum | Equisetaceae | HSV-2 | (78) | |
| Euphorbia hirta | Euphorbiaceae | HIV-1, HIV-2, SIV mac 251 | (80) | |
| Euphorbia sikkimensis | Euphorbiaceae | HIV-1 | (81) | |
| Acacia arabica | Fabaceae | Influenza A virus H9N2 | (83) | |
| Aspalathus linearis | Fabaceae | Rhesus rotavirus (RRV), simian rotavirus (SA-11) infection | (85) | |
| Vachellia nilotica | Fabaceae | HSV-2 | (87) | |
| Acacia catechu | Fabaceae | HIV-1 | (88) | |
| Acacia catechu | Fabaceae | HIV-1 | (88) | |
| Quercus persica | Fagaceae | HSV-I | (90) | |
| Geranium thunbergii | Geraniaceae | Influenza virus, H1N1, H3N2, influenza type B | (91) | |
| Pelargonium sidoides | Geraniaceae | HIV-1 | (92) | |
| Ribes nigrum | Grossulariaceae | Influenza A virus | (94) | |
| Hamamelis virginiana | Hamamelidaceae | Influenza A virus and human papillomavirus | (95) | |
| Prunella vulgaris | Lamiaceae | Lentivirus | (101) | |
| Scutellaria baicalensis | Lamiaceae | RSV, HIV, Influenza, and Dengue viruses | (104) | |
| Strychnos pseudoquina | Loganiaceae | HSV-1 (KOS strain) and HSV-2 (333 strain) | (108) | |
| Punica granatum | Lythraceae | HSV-2 | (109) | |
| Magnolia officinalis | Magnoliaceae | Dengue virus type 2 | (111) | |
| Cissampelos pareira Linn | Menispermaceae | Dengue virus types 1-4 (DENV-1-4) | (113) | |
| Ficus benjamina | Moraceae | HSV-1 and HSV-2), varicella zoster virus (VZV | (114) | |
| Ficus carica | Moraceae | HSV-1, HSV-1, ECV-11, and ADV, influenza virus | (115) | |
| Ficus religiosa | Moraceae | HSV-2 | (116) | |
| Syzygium aromaticum L. | Myrtaceae | HSV and HCV | (119) | |
| Paulownia tomentosa | Paulowniaceae | SARS-CoV papain-like protease (PLpro) | (123) | |
| Phyllanthus amarus | Phyllanthaceae | Acyclovir-resistant HSV strains, hepatitis B virus (HBV), HCV, and HIV | (126) | |
| Polyphenols | Phyllanthus niruri | Phyllanthaceae | Acyclovir-resistant HSV strains, hepatitis B virus (HBV), HCV, HIV | (126) |
| Phyllanthus urinaria | Phyllanthaceae | Acyclovir-resistant HSV strains, hepatitis B virus (HBV), HCV and HIV | (126) | |
| Phyllanthus watsonii | Phyllanthaceae | Acyclovir-resistant HSV strains, hepatitis B virus (HBV), HCV, and HIV | (126) | |
| Limonium sinense | Plumbaginaceae | HCV | (128) | |
| Plumbago indica | Plumbaginaceae | Influenza A (H1N1) | (129) | |
| Agrimonia pilosa | Rosaceae | Influenza viruses (H1N1 and H3N2) | (135) | |
| Prunus dulcis | Rosaceae | HSV-1 | (136) | |
| Pavetta tomentosa | Rubiaceae | Dengue virus (DENV) | (138) | |
| Aegle marmelos | Rutaceae | Human coxsackieviruses B1-B6, rotavirus SA-11 | (139) | |
| Dimocarpus longan | Sapindaceae | HCV (genotype 2a strain JFH1) | (140) | |
| Torreya nucifera | Taxaceae | SARS-CoV 3CLpro | (144) | |
| Viola diffusa | Violaceae | Hepatitis B virus | (147) | |
| Alpinia katsumadai | Zingiberaceae | influenza virus type A | (148) | |
| Illicium verum Hook. f. | Schisandraceae | Grouper iridovirus infection (GIV) | (190) | |
| Camellia sinensis | Theaceae | HIV, HTLV-1, HCV, influenza, and HBV | (145, 146) | |
| Ocimum sanctum | Lamiaceae | Dengue virus serotype-1 (DENV-1) | (26, 99) | |
| Achillea fragrantissima | Asteraceae | Poliomyelitis-1 virus | (47, 48) | |
| Ephedra alata | Ephedraceae | HSV | (47, 77) | |
| Tamarix nilotica | Tamaricaceae | HSV | (47, 143) | |
| Moringa peregrina | Moringaceae | HSV | (47, 189) | |
| Capparis sinaica | Capparaceae | Avian influenza strain H5N1 | (47, 64) | |
| Ficus sycomorus | Moraceae | HSV-1 | (56, 118) | |
| Balanites aegyptiaca | Balanitaceae | VSV | (56, 57) | |
| Terminalia mollis | Combretaceae | HSV-0 | (56, 73) | |
| Tuberaria lignosa | Cistaceae | HIV | (70, 71) | |
| Anthemis hyaline | Asreraceae | SARS-CoV | (152) | |
| Alnus japonica | Betulaceae | SARS-CoV | (59) | |
| Cassia tora | Fabaceae | SARS-CoV | (156) | |
| Psoralea corylifolia | Fabaceae | SARS-CoV | (150) | |
| Taxillus chinensis | Loranthaceae | SARS-CoV | (268) | |
| Polyphenols | Citrus sinensis | Rutaceae | SARS-CoV | (152) |
| Polygonum multiflorum | Polygonaceae | SARS-CoV | (158) | |
| Rheum officinale | Polygonaceae | SARS-CoV | (158) | |
| Rheum palmatum | Polygonaceae | SARS-CoV | (159) | |
| Citrus sinensis | Rutaceae | SARS-CoV | (152) | |
| Alkaloids | Sambucus nigra | Adoxaceae | Dengue virus serotype-2 (DENV-2) | (29) |
| Iresine Herbstii | Amaranthaceae | Newcastle disease virus (NDV) | (31) | |
| Combretum adenogonium | Combretaceae | HIV-1 | (72) | |
| Cyperus rotundus | Cyperaceae | HSV-1, HBV | (76) | |
| Equisetum giganteum | Equisetaceae | HSV-2 | (78) | |
| Cissampelos pareira Linn | Menispermaceae | Dengue virus types 1-4 (DENV-1-4) | (113) | |
| Ficus religiosa | Moraceae | HSV-2 | (116) | |
| Phyllanthus amarus | Phyllanthaceae | HCV | (125) | |
| Plumbago indica | Plumbaginaceae | Influenza A (H1N1) | (129) | |
| Pavetta tomentosa | Rubiaceae | Dengue virus (DENV) | (138) | |
| Tarenna asiatica | Rubiaceae | Dengue virus (DENV) | (138) | |
| Moringa peregrina | Moringaceae | HSV | (47, 189) | |
| Capparis sinaica | Capparaceae | HSV | (47, 65) | |
| Balanites aegyptiaca | Balanitaceae | VSV | (56, 57) | |
| Lycoris radiata | Amaryllis | SARS-CoV | (151) | |
| Acanthopanacis cortex | Araliaceae | SARS-CoV | (134) | |
| Saponins | Iresine Herbstii | Amaranthaceae | Newcastle disease virus (NDV) | (31) |
| Anacardium occidentale | Anacardiaceae | Simian (SA-11) virus | (33) | |
| Panax ginseng | Araliaceae | RSV | (41) | |
| Panax ginseng | Araliaceae | Murine norovirus (MNV) and feline calicivirus (FCV) | (42) | |
| Balanites aegyptiaca | Balanitaceae | VSV | (56, 57) | |
| Capparis sinaica | Capparaceae | HSV | (47, 65) | |
| Combretum adenogonium | Combretaceae | HIV-1 | (72) | |
| Vachellia nilotica | Fabaceae | HSV-2 | (87) | |
| Lindernia crustacea | Linderniaceae | Epstein–Barr virus (EBV) | (107) | |
| Artocarpus integrifolia | Moraceae | (SA-11) and human (HCR3) rotaviruses | (33) | |
| Ficus religiosa | Moraceae | HSV-2 | (116) | |
| Saponins | Ficus sycomorus | Moraceae | HSV-1 | (56, 118) |
| Moringa peregrina | Moringaceae | HSV | (47, 189) | |
| Plumbago indica | Plumbaginaceae | Influenza A (H1N1) | (129) | |
| Pavetta tomentosa | Rubiaceae | Dengue virus (DENV) | (138) | |
| Tarenna asiatica | Rubiaceae | Dengue virus (DENV) | (138) | |
| Terpenoids | Andrographis paniculata | Acanthaceae | Dengue virus serotype-1 (DENV-1) | (26) |
| Baccharis gaudichaudiana DC | Asteraceae | Bovine viral diarrhea virus, HSV-1, Poliovirus type 2 (PV-2), and vesicular stomatitis virus (VSV) | (49) | |
| Baccharis spicata (Lam.) Baill | Asteraceae | Bovine viral diarrhea virus (BVD), HSV-1, poliovirus type 2 (PV-2), and vesicular stomatitis virus (VSV) | (49) | |
| Taxodium distichum | Cupressaceae | Influenza A and B viruses | (75) | |
| Euphorbia hirta | Euphorbiaceae | HIV-1, HIV-2, SIV mac 251 | (80) | |
| Jatropha multifida | Euphorbiaceae | Influenza A H1N1 virus | (82) | |
| Torreya nucifera | Taxaceae | SARS-CoV 3CLpro | (144) | |
| Agrimonia pilosa | Rosaceae | Influenza viruses (H1N1 and H3N2) | (135) | |
| Tripterygium regelii | Celastraceae | SARS-CoV | (144) | |
| Gentiana scabra | Gentianaceae | SARS-CoV | (156) | |
| Carbohydra-tes | Panax ginseng | Araliaceae | Human rotavirus | (33) |
| Panax notoginseng | Araliaceae | Influenza A H1N1 virus | (43) | |
| Equisetum giganteum | Equisetaceae | HSV-2 | (78) | |
| Prunella vulgaris | Lamiaceae | HSV-1 and HSV-2 antigens virus antigen in Vero cells | (100) | |
| Prunellae Spica | Lamiaceae | Herpes simplex virus (HSV) | (102) | |
| Laminaria japonica | Laminariaceae | RSV | (105) | |
| Plumbago indica | Plumbaginaceae | Influenza A (H1N1) | (129) | |
| Ardisia chinensis Benth | Primulaceae | Coxsackie B3 Virus | (131) | |
| Capparis sinaica | Capparaceae | HSV | (47, 65) | |
| Balanites aegyptiaca | Balanitaceae | VSV | (56, 57) | |
| Carissa edulis | Apocynaceae | herpes simplex virus, chickenpox, and shingles | (38) |
Alkaloids are another class of natural organic compounds which are classified into several groups based on their heterocyclic ring, such as tropanes, pyrrolidines, isoquinoline purines, imidazoles, quinolizidines, indoles, piperidines, and pyrrolizidines (278). Alkaloids are very promising against HIV-1, HSV-1, HSV-2, DNV, VSV, Influenza virus, and Newcastle disease virus (NDV) (Table 4). Different kinds of alkaloids showed anti-SARS activity including emetine, Ipecac, Macetaxime, tylophorine, and 7-methoxy cryptopleurine, through the inhibition of protease enzyme, RNA synthesis, and protein synthesis (244, 279). In addition, some alkaloids act against SARS CoV as a nucleic acid intercalating agent such as tetrandrine, fangchinoline, cepharanthine, and lycorine through degrading nucleic acids and inhibiting spike and nucleocapsid proteins (280). Virtual screening analysis revealed that 10-Hydroxyusambarensine and Cryptoquindoline—two alkaloid compound isolated from African medicinal plants showed anti-SARS CoV and anti-SARS CoV-2 activity through inhibition of their Mpro (256). Chloroquine, a derivative of alkaloid, is found to be active against anti-SARS CoV-2 (281). So, some PSMs as alkaloids can be alternative drug targets for COVID-19 (280).
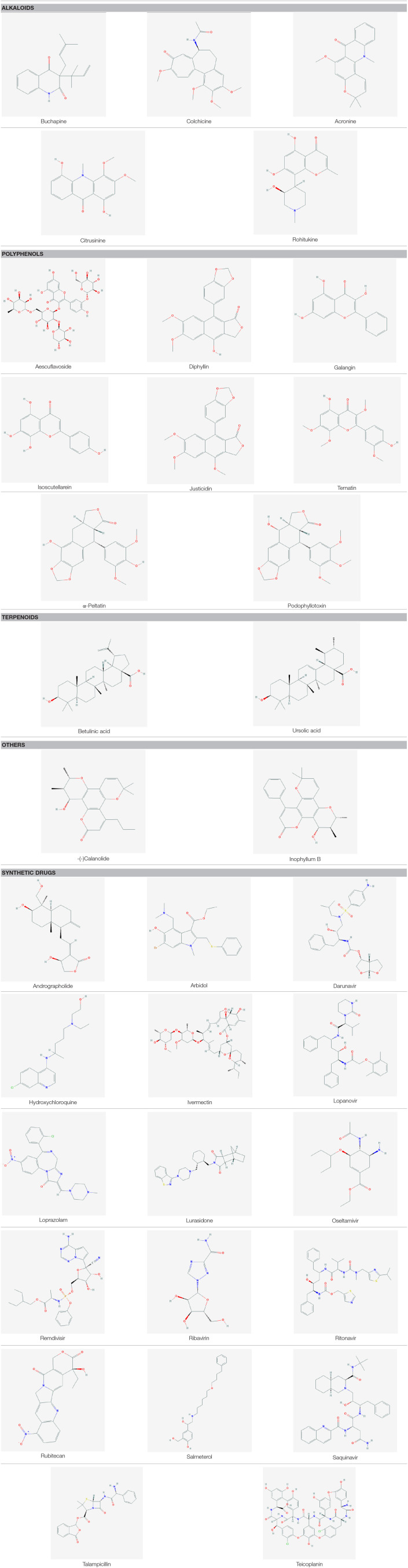
Another class of PSMs, saponins (amphipathic glycosides), are found ubiquitously in plants which showed antiviral activities against Newcastle disease virus (NDV), Simian (SA-11) virus, Murine norovirus (MNV) and Feline calicivirus (FCV), RSV, VSV, HSV-1,HSv-2, HIV-1, Epstein–Barr virus (EBV), (SA-11) and human (HCR3) rotaviruses, Influenza virus, and Dengue virus (Table 4). Plants produce five carbon isoprene derived terpenes which are the largest and most diverse group of PSM. They are classified by monoterpenes, diterpenes, triterpenes, sesterterpenes, hemi terpenes, and sesquiterpenes (282). They exhibited antiviral activity against Bovine viral diarrhea virus, HSV-1, Poliovirus type 2 (PV-2) and vesicular stomatitis virus (VSV), Dengue virus serotype-1 (DENV-1), Influenza A and B viruses, HIV-1, HIV-2, SIV mac 251, and SARS-CoV (Table 4). Ten diterpenes, two sesquiterpenes, and two triterpenes showed anti-SARS activity with IC50 of 3–10 μM (283). In silico analysis also revealed that terpene Ginkgolide A can strongly inhibit SARS CoV-2 protease enzyme (284). Carbohydrates, mainly classified as monosaccharides, disaccharides, polysaccharides, and oligosaccharides (282), are found as antiviral agent against Human rotavirus, Influenza A virus, HSV-1, HSV-2, Herpes simplex virus (HSV), RSV, Coxsackie B3 Virus, and VSV [(285); Table 4]. Acyclovir is an FDA (Food and Drug Administration) approved antiviral drug which is obtained from Carissa edulis (Supplementary Table 1). It is mainly used for herpes simplex virus, chickenpox, and shingles. The group basis structure of some major compounds can be found in Table 5.
Table 5.
Structures of some major PSMs and Drugs used against SARS CoV-2.
 |
Drug Discovery From PSMs: Addressing the Major Challenges Toward Future Insights
Drug discovery from plant metabolites refers to the extraction and purification of active ingredients from conventional cures. Natural plant products comprise complicated chemical structures which differ according to their numerous species. There are several classes of PSMs which are responsible for the biological activities of herbal medicines. PSMs exert their actions on molecular targets that differ from one case to the other. These targets may be enzymes, mediators, transcription factors, or even nucleic acids (286). Good knowledge of the chemical composition of plants leads to a better understanding of their possible and specific medicinal value. Drug discovery and development have become a wide interdisciplinary field over recent decades and many factors are involved in the successful evolution from a bioactive compound into a potential drug [(287, 288); Figure 2]. When existing methods with advanced technologies are applied, it can lead to a modern revelation of drugs, benefitting medicinal purposes (223, 289). The development of modern technologies has streamlined the screening of natural products in discovering new drugs. Research for drug discovery must create robust and prudent lead molecules, which is progressed from a screening hit to a drug candidate through structural elucidation and structure recognizable proof available from high throughput technology like GC–MS, NMR, IR, HPLC, and HPTLC. Utilizing these advanced technologies gives us an opportunity to perform research in screening novel molecules employing a computer program and database to set up common items as a major source for drug discovery. It finally leads to lead structure discovery. Powerful new technologies are revolutionizing natural herbal drug discovery (223). Steps associated with the drug discovery process from natural resources is illustrated (Figure 3).
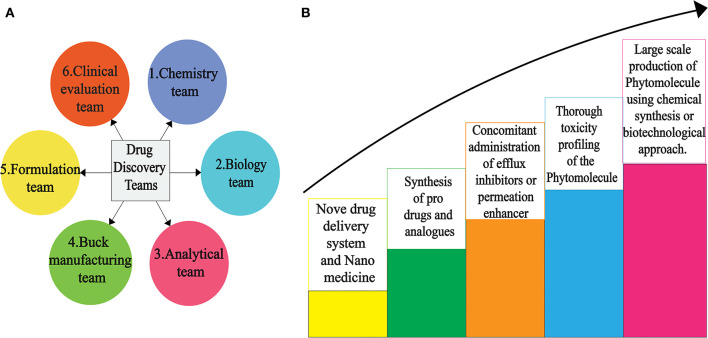
Figure 2.
Scientific teams (A) to overcome various hurdles for successful novel drug discovery (B) from PSMs.
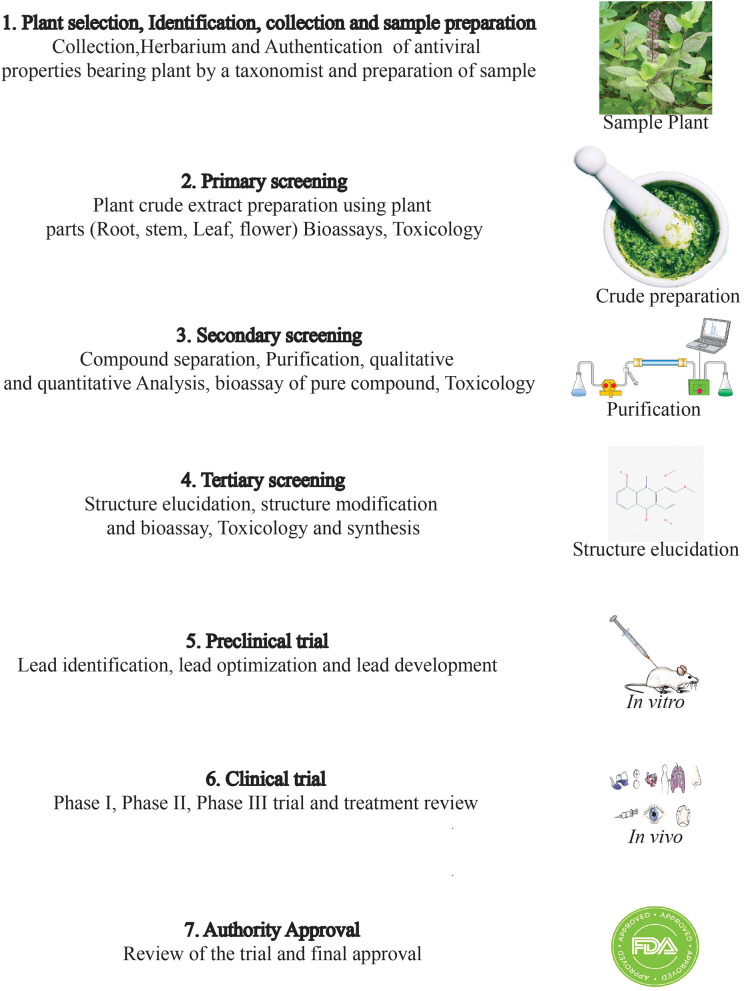
Figure 3.
Various steps involved in the tedious drug discovery process from plant sources.
However, several factors involving the conversion of a desirable compound into a valuable drug candidate include availability, bioavailability, intellectual property, and the strong pharmacokinetic profile of the compound (268, 290). Sometimes researchers find great bioactivity of a plant-derived compound in in vitro analysis but unfortunately, the desired compound becomes ineffectual under in vivo conditions (291). In vivo is a very crucial step to move to animal trials or subsequent clinical trials. Even if the compound shows promising activity in in vivo assay but it can still become ineffective in animal model trials due to a poor pharmacokinetic profile (292). Under in vivo condition, the target compound remains in direct contact with cells, while in animal models the compound moves to various stages where it might lose its bioactivity (292). For example, despite curcumin having promising antioxidant, anticancer, anti-inflammatory, and antimicrobial activities, it has not been released as a drug yet due to its poor bioavailability (292). Another propitious drug candidate, epigallocatechin gallate (EGCG), showed antioxidant, antihypertensive, anticancer, antimicrobial, and anti-inflammatory activity (293, 294) but unfortunately, it has also failed to obtain drug designation due to the same reason mentioned for curcumin (292).
To remedy these problems, researchers around the world are working to develop new approaches. Changing the administration route might increase the bioavailability of a compound. For example, the bioavailability of an anti-inflammatory compound, andrographolide, is increased when it is administered intravenously instead of through oral administration (295). Other methods to enhance the bioavailability of target compound include using drug delivery systems, the nano-formulation of a drug, using adjuvant systems, or altering structural analogs (220, 296). Furthermore, the modification of pharmacokinetic profiles of compounds like absorption, distribution, metabolism, and excretion can escalate its probability as drug candidate (268). Indeed, there is an urgent need for specific protocols for invention of novel bioactive compounds and for this purpose it is very crucial for related organizations, companies, and agencies, including the World Health Organization (WHO), Food and Drug Administration (FDA), European Medicines Agency (EMA), World Trade Organization (WTO), International Conference on Harmonization (ICH), World Intellectual Property Organization (WIPO), biotech companies, pharmaceutical pharmaceuticals companies, and several other companies and agencies, to work together. However, plant-originated therapeutics need to be taken under consideration against SARS-CoV-2 as they have already shown promising hopes for different critical conditions caused by deadly pathogens.
The seven major drug targets of SARS CoV-2 were described before (176). Similarly, screening of PSMs for drug establishment by molecular docking is efficient in terms of time and cost. Even the development of vaccines through computational biology was found to be effective for previous severe viruses like MERS using animal models, target antigens, and probable vaccine candidates (181). But still, there exists a lack of a complete review for PSMs as alternative drug therapeutics. Our review aims at establishing PSMs as a strong and safe candidate for the treatment of SARS CoV-2. Through suggesting probable antiviral plant metabolites or screening, druggability analysis of plant metabolites against SARS-CoV-2 has become a time-saving practice (280, 297). Without establishing a drug development pipeline that includes clinical trials, these suggested candidate PSMs will end up only in journal publications or be shelved as herbal formulations on a supermarket store as a traditional medicine and will never be a modern drug. Undoubtedly, the plant an underutilized source of novel bioactive compounds and is one of the hotspots to fight against this microbial resistance war. The decrypting of PSMs is not increasing so much in comparison to the number of metabolites produced from plants. A biotechnological approach can offer a desired amount of secondary metabolites in a rapid and eco-friendly way against SARS-CoV-2 (298). In addition, plant metabolomics are now used as a tool for discovery of novel drugs from plant resources (299). Characterization of genes and proteins involved in secondary metabolic pathways are also very crucial to understand. Therefore, omics approaches (transcriptomics, proteomics, and metabolomics) have paramount importance in food research and drug discovery (300, 301) for human welfare. Genetic modifications for engineering plant metabolites can be helpful for reaching a specific drug. Quality control of natural products is also very important. So, laboratory support, skilled manpower, and funding is also very important for drug discovery from natural resources.
Conclusions
Scientists all around the world are trying to discover the most effective antiviral drug to combat SARS CoV-2. In this situation, our study accentuated some plant secondary metabolites that showed prominent antiviral activity against coronaviruses through impeding the main machinery used in their pathogenesis and replication cycle. The in vitro, in vivo, and in silico investigations revealed numerous plant-derived compounds with promising anti- SARS CoV and anti- SARS CoV-2 activity [Table 2; (179, 219, 222, 233–261, 297)]. Plants are a dramatically underutilized source of bioactive compounds with a broad spectrum of antiviral activities. Some Chinese traditional plant formulations have been reported as being anti- SARS CoV-2 and this formulation is also provided in COVID patients (302, 303). We reported here on 219 plants which act against a wide range of DNA/RNA viruses, but the plant PSMs that showed promising activity against SARS CoV and MERS might be a desired drug candidate against SARS CoV-2. So, this review gathered all antiviral plants in a single platform to facilitate laboratory-based research for the development of novel drug/molecular therapeutics to overcome this and future pandemic situations. The world is facing a serious health crisis, and it needs an effective solution to combat the burning flame of COVID-19. Researchers are trying to find an effective way to overcome this situation, and the present study could help them to think with a new dimension by using the knowledge from the databases based on the plant metabolites (304, 305). Finally, advanced and rapid acting extraction, purification, and characterization techniques used for plant metabolites as well as multidisciplinary expertise and funding are very essential for novel drug discovery.
Literature Study
Articles were selected and identified by searching specific keywords and journal citations for each section of a manuscript. Related peer reviewed scientific journal articles were screened from different journal depositories after reviewing abstracts and original data.
Author's Note
The authors initiated this project to facilitate the research on molecular therapeutics from plant sources as an immediate action in response to the COVID-19 pandemic situation.
Author Contributions
FB and MH designed the project. FB prepared the first draft. FB, SH, TR, and MH have investigated the data and completed the manuscript. All authors have read through the manuscript and approved it for submission and publication.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We are thankful to Mr. Md. Fayezur Rahman (MSS in Economics, Comilla University) for drawing the figures as authors expected. Our sincere gratitude to Mr. Mohammad Saifullah (MA in English, University of Chittagong) for proofreading of the manuscript. We are grateful to the editors and reviewers for their valuable comments during the review process to improve the manuscript.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmed.2020.00444/full#supplementary-material
Different plant families showing antiviral properties. (Each portion of the pie chart describes a specific Family alongside its total number of plants that have antiviral properties).
List of secondary metabolites found from medicinal plants.
References
- 1.Brian DA, Baric RS. Coronavirus genome structure and replication. Curr Top Microbiol Immunol. (2005) 287:1–30. 10.1007/3-540-26765-4_1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. (2020) 395:565–74. 10.1016/S0140-6736(20)30251-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Cavanagh D. Coronavirus avian infectious bronchitis virus. Vet Res. (2007) 38:281–97. 10.1051/vetres:2006055 [DOI] [PubMed] [Google Scholar]
- 4.Ismail MM, Tang Y, Saif YM. Pathogenicity of turkey coronavirus in turkeys and chickens. Avian Dis. (2003) 47:515–22. 10.1637/5917 [DOI] [PubMed] [Google Scholar]
- 5.Fehr AR, Perlman S. Coronaviruses: an overview of their replication and pathogenesis. Methods Mol Biol. (2015) 1282:1–23. 10.1007/978-1-4939-2438-7_1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Lan Jun, Jiwan Ge, Jinfang Yu, Sisi Shan, Huan Zhou, et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature. (2020) 581:215–20 10.1101/2020.02.19.956235 [DOI] [PubMed] [Google Scholar]
- 7.Chen Y, Peng H, Wang L, Zhao Y, Zeng L, Gao H, et al. Infants born to mothers with a new coronavirus (COVID-19). Front Pediatr. (2020) 8:104. 10.3389/fped.2020.00104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.WHO Director-General's Opening Remarks at the Media Briefing on COVID-19. (2020). Available online at: https://www.who.int/dg/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19 (accessed May 15, 2020).
- 9.WHO Director-General's Opening Remarks at the Media Briefing on COVID-19. (2020). Available online at: https://www.who.int/dg/speeches/detail/who-director-general-sopening-remarks-at-the-media-briefing-on-covid-19 (accessed April 5, 2020).
- 10.Kanne JP. Chest CT findings in 2019 novel coronavirus (2019-nCoV) infections from Wuhan, China: Key points for the radiologist. Radiology. (2020) 295:16–7. 10.1148/radiol.2020200241 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zumla A, Chan JFW, Azhar EI, Hui DSC, Yuen KY. Coronaviruses-drug discovery and therapeutic options. Nat Rev Drug Discov. (2016) 15:327–47. 10.1038/nrd.2015.37 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Heymann DL, Rodier G. Global surveillance, national surveillance, and SARS. Emerg Infect Dis. (2004) 10:173–5. 10.3201/eid1002.031038 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Zaki AM, Van Boheemen S, Bestebroer TM, Osterhaus ADME, Fouchier RAM. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N Engl J Med. (2012) 367:1814–20. 10.1056/NEJMoa1211721 [DOI] [PubMed] [Google Scholar]
- 14.Shetty R, Ghosh A, Honavar SG, Khamar P, Sethu S. Therapeutic opportunities to manage COVID-19/SARS-CoV-2 infection: present and future. Indian J Ophthalmol. (2020) 68:693–702. 10.4103/ijo.IJO_639_20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hirano T, Murakami M. COVID-19: a new virus, but a familiar receptor and cytokine release syndrome. Immunity. (2020) 52:731–3. 10.1016/j.immuni.2020.04.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Pedersen SF, Ho YC. SARS-CoV-2: a storm is raging. J Clin Invest. (2020) 130:2202–5. 10.1172/JCI137647 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Coperchini F, Chiovato L, Croce L, Magri F, Rotondi M. The cytokine storm in COVID-19: an overview of the involvement of the chemokine/chemokine-receptor system. Cytokine Growth Factor Rev. (2020) 53:25–32. 10.1016/j.cytogfr.2020.05.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Moses T, Goossens A. Plants for human health: greening biotechnology and synthetic biology. J Exp Bot. (2017) 68:4009–11. 10.1093/jxb/erx268 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Schaal B. Plants and people: our shared history and future. Plants People Planet. (2019) 1:14–9. 10.1002/ppp3.12 [DOI] [Google Scholar]
- 20.World Health Organization WHO Global Report on Traditional and Complementary Medicine 2019. World Health Organization; (2019). [Google Scholar]
- 21.Hoque Haque MI, Chowdhury ABMA, Shahjahan M, Harun MGD. Traditional healing practices in rural Bangladesh: a qualitative investigation. BMC Complement Altern Med. (2018) 18:62. 10.1186/s12906-018-2129-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ozioma E, Okaka A. Herbal medicines in African traditional medicine. In: Ozioma EJ, Chinwe OAN. editors. Herbal Medicine. Intratech open; (2019). p. 1–25. [Google Scholar]
- 23.Jassim SAA, Naji MA. Novel antiviral agents: a medicinal plant perspective. J Appl Microbiol. (2003) 95:412–27. 10.1046/j.1365-2672.2003.02026.x [DOI] [PubMed] [Google Scholar]
- 24.Hussain W, Haleem KS, Khan I, Tauseef I, Qayyum S, Ahmed B, et al. Medicinal plants: a repository of antiviral metabolites. Future Virol. (2017) 12:299–308. 10.2217/fvl-2016-0110 [DOI] [Google Scholar]
- 25.Özçelik B, Kartal M, Orhan I. Cytotoxicity, antiviral and antimicrobial activities of alkaloids, flavonoids, and phenolic acids. Pharm Biol. (2011) 49:396–402. 10.3109/13880209.2010.519390 [DOI] [PubMed] [Google Scholar]
- 26.Pick A, Ling K, Khoo BF, Seah CH, Foo KY, Cheah RK. Inhibitory activities of methanol extracts of Andrographis Paniculata and Ocimum Sanctum against Dengue-1 virus. In: International Conference on Biological, Environment & Food Engineering. Bali: (2014). p. 6. [Google Scholar]
- 27.Namazi R, Zabihollahi R, Behbahani M, Rezae A. Inhibitory activity of Avicennia marina, a medicinal plant in persian folk medicine, against HIV and HSV. Iran J Pharm Res.. (2013) 12:435–43. [PMC free article] [PubMed] [Google Scholar]
- 28.Zhou B, Yang Z, Feng Q, Liang X, Li J, Zanin M, et al. Aurantiamide acetate from baphicacanthus cusia root exhibits anti-inflammatory and anti-viral effects via inhibition of the NF-κB signaling pathway in influenza A virus-infected cells. J Ethnopharmacol. (2017) 199:60–7. 10.1016/j.jep.2017.01.038 [DOI] [PubMed] [Google Scholar]
- 29.Castillo-Maldonado I, Moreno-Altamirano MMB, Serrano-Gallardo LB. Anti-dengue serotype-2 activity effect of Sambucus nigra leaves-and flowers-derived compounds. Virol Res Rev. (2017) 1:1–5. 10.15761/VRR.1000117 [DOI] [Google Scholar]
- 30.Chen C, Zuckerman DM, Brantley S, Sharpe M, Childress K, Hoiczyk E, et al. Sambucus nigra extracts inhibit infectious bronchitis virus at an early point during replication. BMC Vet Res. (2014) 10:24. 10.1186/1746-6148-10-24 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Andleeb R, Ashraf A, Muzammil S, Naz S, Asad F, Ali T, et al. Analysis of bioactive composites and antiviral activity of Iresine herbstii extracts against Newcastle disease virus in ovo. Saudi J Biol Sci. (2020) 27:335–40. 10.1016/j.sjbs.2019.10.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Szlμvik L, Minμrovits J, Forgo P. Alkaloids from Leucojum vernum and antiretroviral activity of Amaryllidaceae alkaloids. Planta Medica. (2004) 70:871–3. 10.1055/s-2004-827239 [DOI] [PubMed] [Google Scholar]
- 33.Lopes RC, Oliveira DB, Costa SS, Miranda MMFS, Romanos MT V, Santos NSO, et al. In vitro anti-rotavirus activity of some medicinal plants used in Brazil against diarrhea. J Ethnopharmacol. (2005) 99:403–7. 10.1016/j.jep.2005.01.032 [DOI] [PubMed] [Google Scholar]
- 34.Reichling J, Neuner A, Sharaf M, Harkenthal M, Schnitzler P. Antiviral activity of Rhus aromatica (fragrant sumac) extract against two types of herpes simplex viruses in cell culture. Pharmazie. (2009) 64:538–41. [PubMed] [Google Scholar]
- 35.Modi M, Nutan, Pancholi B, Kulshrestha S, Rawat AKS, Malhotra S, et al. Anti-HIV-1 activity, protease inhibition and safety profile of extracts prepared from Rhus parviflora. BMC Complement Altern Med. (2013) 13:158. 10.1186/1472-6882-13-158 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Nocchi SR, Companhoni MVP, De Mello JCP, Dias Filho BP, Nakamura CV, Carollo CA, et al. Antiviral activity of crude hydroethanolic extract from Schinus terebinthifolia against herpes simplex virus type 1. Planta Med. (2017) 83:509–18. 10.1055/s-0042-117774 [DOI] [PubMed] [Google Scholar]
- 37.Park JY, Ko JA, Kim DW, Kim YM, Kwon HJ, Jeong HJ, et al. Chalcones isolated from Angelica keiskei inhibit cysteine proteases of SARS-CoV. J Enzyme Inhib Med Chem. (2016) 31:23–30. 10.3109/14756366.2014.1003215 [DOI] [PubMed] [Google Scholar]
- 38.Tolo FM, Rukunga GM, Muli FW, Njagi ENM, Njue W, Kumon K, et al. Anti-viral activity of the extracts of a Kenyan medicinal plant Carissa edulis against Herpes simplex virus. J Ethnopharmacol. (2006) 104:92–9. 10.1016/j.jep.2005.08.053 [DOI] [PubMed] [Google Scholar]
- 39.Bonvicini F, Lianza M, Mandrone M, Poli F, Gentilomi GA, Antognoni F. Hemidesmus indicus (L.) R. Br. Extract inhibits the early step of herpes simplex type 1 and type 2 replication. New Microbiol. (2018) 41:187–94. [PubMed] [Google Scholar]
- 40.Rittà M, Marengo A, Civra A, Lembo D, Cagliero C, Kant K, et al. Antiviral activity of a Arisaema tortuosum leaf extract and some of its constituents against herpes simplex virus type 2. Planta Med. (2020) 86:267–75. 10.1055/a-1087-8303 [DOI] [PubMed] [Google Scholar]
- 41.Lee JS, Ko E, Hwang HYESUK, Lee Y, Kwon Y, Kim M, et al. Antiviral activity of ginseng extract against respiratory syncytial virus infection. Int J Mol Med. (2014) 34:183–90. 10.3892/ijmm.2014.1750 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Lee MH, Lee B, Jung J, Cheon D, Kim K, Choi C. Antiviral effect of Korean red ginseng extract and ginsenosides on murine norovirus and feline calicivirus as surrogates for human norovirus. J Ginseng Res. (2011) 35:429–35. 10.5142/jgr.2011.35.4.429 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Choi J, Jin Y, Lee H, Oh TW, Yim N, Cho W. Protective effect of Panax notoginseng root water extract against influenza a virus infection by enhancing antiviral interferon-mediated immune responses and natural killer cell activity. Front Immunol. (2017) 8:1542. 10.3389/fimmu.2017.01542 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Glatthaar-Saalmüller B, Fal AM, Schönknecht K, Conrad F, Sievers H, Saalmüller A. Antiviral activity of an aqueous extract derived from Aloe arborescens Mill. against a broad panel of viruses causing infections of the upper respiratory tract. Phytomedicine. (2015) 22:911–20. 10.1016/j.phymed.2015.06.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Zandi K, Zadeh MA, Sartavi K, Rastian Z. Antiviral activity of Aloe vera against herpes simplex virus type 2 : an in vitro study. Afr J Biotechnol. (2007) 6:1770203 10.5897/AJB2007.000-2276 [DOI] [Google Scholar]
- 46.Moradi M, Rafieian-kopaei M, Karimi A. A review study on the effect of Iranian herbal medicines against in vitro replication of herpes simplex virus. Avicenna J Phytomed. (2016) 6:506–15. [PMC free article] [PubMed] [Google Scholar]
- 47.Soltan MM, Zaki AK. Antiviral screening of forty-two Egyptian medicinal plants. J Ethnopharmacol. (2009) 126:102–7. 10.1016/j.jep.2009.08.001 [DOI] [PubMed] [Google Scholar]
- 48.Patocka J, Navratilova Z. Achillea fragrantissima : pharmacology review. Clin Oncol. (2019) 4:1601. [Google Scholar]
- 49.Visintini M, Redko F, Muschietti L, Campos R, Martino V, Cavallaro LV. in vitro antiviral activity of plant extracts from Asteraceae medicinal plants. Virol J. (2013) 10:245. 10.1186/1743-422X-10-245 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Derksen A, Kühn J, Hafezi W, Sendker J, Ehrhardt C, Ludwig S, et al. Antiviral activity of hydroalcoholic extract from Eupatorium perfoliatum L. against the attachment of influenza A virus. J Ethnopharmacol. (2016) 188:144–52. 10.1016/j.jep.2016.05.016 [DOI] [PubMed] [Google Scholar]
- 51.Lani R, Hassandarvish P, Chiam CW, Moghaddam E, Chu JJH, Rausalu K, et al. Antiviral activity of silymarin against chikungunya virus. Sci Rep. (2015) 5:11421. 10.1038/srep11421 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Benassi-Zanqueta É, Marques CF, Valone LM, Pellegrini BL, Bauermeister A, Ferreira ICP, et al. Evaluation of anti-HSV-1 activity and toxicity of hydroethanolic extract of Tanacetum parthenium (L.) Sch.Bip. (Asteraceae). Phytomedicine. (2019) 55:249–54. 10.1016/j.phymed.2018.06.040 [DOI] [PubMed] [Google Scholar]
- 53.Rehman S, Ijaz B, Fatima N, Aun S. Sciencedirect therapeutic potential of Taraxacum officinale against HCV NS5B polymerase : in-vitro and in silico study. Biomed Pharmacother. (2016) 83:881–91. 10.1016/j.biopha.2016.08.002 [DOI] [PubMed] [Google Scholar]
- 54.He W, Han H, Wang W, Gao B. Anti-influenza virus effect of aqueous extracts from dandelion. Virol J. (2011) 8:538. 10.1186/1743-422X-8-538 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Rothan HA, Zulqarnain M, Ammar YA, Tan EC, Rahman NA, Yusof R. Screening of antiviral activities in medicinal plants extracts against dengue virus using dengue NS2B-NS3 protease assay. Trop Biomed. (2014) 31:286–96. [PubMed] [Google Scholar]
- 56.Mgole S, Pieters L, David O, Apers S, Vingerhoets R, Cos P, et al. Screening of some Tanzanian medicinal plants from Bunda District for antibacterial, antifungal and antiviral activities. J Ethnopharmacol. (2008) 119:58–66. 10.1016/j.jep.2008.05.033 [DOI] [PubMed] [Google Scholar]
- 57.Al-thobaiti SA, Zeid IMA. Phytochemistry and pharmaceutical evaluation of Balanites aegyptiaca : an overview. J Exp Biol Agric Sci. (2018) 6:453–65 10.18006/2018.6(3).453.465 [DOI] [Google Scholar]
- 58.Cho WK, Kim H, Choi YJ, Yim NH, Yang HJ, Ma JY. Epimedium koreanum Nakai water extract exhibits antiviral activity against porcine epidermic diarrhea virus in vitro and in vivo. Evid Based Complement Altern Med. (2012) 2012:985150. 10.1155/2012/985151 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Park JY, Jeong HJ, Kim JH, Kim YM, Park SJ, Kim D, et al. Diarylheptanoids from Alnus japonica inhibit papain-like protease of severe acute respiratory syndrome coronavirus. Biol Pharm Bull. (2012) 35:2036–42. 10.1248/bpb.b12-00623 [DOI] [PubMed] [Google Scholar]
- 60.Tung NH, Kwon HJ, Kim JH, Ra JC, Ding Y, Kim JA, et al. Anti-influenza diarylheptanoids from the bark of Alnus japonica. Bioorganic Med Chem Lett. (2010) 20:1000–3. 10.1016/j.bmcl.2009.12.057 [DOI] [PubMed] [Google Scholar]
- 61.Lin CW, Tsai FJ, Tsai CH, Lai CC, Wan L, Ho TY, et al. Anti-SARS coronavirus 3C-like protease effects of Isatis indigotica root and plant-derived phenolic compounds. Antiviral Res. (2005) 68:36–42. 10.1016/j.antiviral.2005.07.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Chen F, Yang L, Huang Y, Chen Y, Sang H, Duan W, et al. Isocorilagin, isolated from Canarium album (Lour.) Raeusch, as a potent neuraminidase inhibitor against influenza A virus. Biochem Biophys Res Commun. (2020) 523:183–9. 10.1016/j.bbrc.2019.12.043 [DOI] [PubMed] [Google Scholar]
- 63.César GZJ, Alfonso MGG, Marius MM, Elizabeth EM, Ángel CBM, Maira HR, Guadalupe CLM, et al. Inhibition of HIV-1 reverse transcriptase, toxicological and chemical profile of Calophyllum brasiliense extracts from Chiapas, Mexico. Fitoterapia. (2011) 82:1027–34. 10.1016/j.fitote.2011.06.006 [DOI] [PubMed] [Google Scholar]
- 64.Ibrahim AK, Youssef AI, Arafa AS, Ahmed SA. Anti-H5N1 virus flavonoids from Capparis sinaica Veill. Nat Prod Res. (2013) 27:2149–53. 10.1080/14786419.2013.790027 [DOI] [PubMed] [Google Scholar]
- 65.Ghazal EA, Khamis IMA, Elhaw MHM. Chemical constituents of Capparis sinaica Veill. plant and its antimicrobial effects. Middle East J Appl Sci. (2015) 5:411–22. [Google Scholar]
- 66.Lam S-Z, Ng T-B. A protein with antiproliferative, antifungal and HIV-1 reverse transcriptase inhibitory activities from caper (Capparis spinosa) seeds. Phytomedicine. (2009) 16:444–50. 10.1016/j.phymed.2008.09.006 [DOI] [PubMed] [Google Scholar]
- 67.Callies O, Bedoya LM, Beltra M, Mun A, Obrego P, Osorio AA, et al. Isolation, structural modification, and HIV inhibition of pentacyclic lupane-type triterpenoids from Cassine xylocarpa and Maytenus cuzcoina. J Nat Prod. (2015) 78:1045–55. 10.1021/np501025r [DOI] [PubMed] [Google Scholar]
- 68.Inoue R. Orally administered Salacia reticulata extract reduces h1n1 influenza clinical symptoms in murine lung tissues putatively due to enhanced natural killer cell activity. Front Immunol. (2016) 7:115. 10.3389/fimmu.2016.00115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Rebensburg S, Helfer M, Schneider M, Koppensteiner H, Eberle J, Schindler M, et al. Potent in vitro antiviral activity of Cistus incanus extract against HIV and Filoviruses targets viral envelope proteins. Sci Rep. (2016) 6:20394. 10.1038/srep20394 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Javed T, Ashfaq UA, Riaz S, Rehman S, Riazuddin S. In-vitro antiviral activity of Solanum nigrum against Hepatitis C Virus. Virol J. (2011) 8:26. 10.1186/1743-422X-8-26 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Alcami J, Bermejo P. Phytomedicine Ellagitannins from Tuberaria lignosa as entry inhibitors of HIV. Phytomedicine. (2010) 17:69–74. 10.1016/j.phymed.2009.08.008 [DOI] [PubMed] [Google Scholar]
- 72.Mushi NF, Mbwambo ZH, Innocent E, Tewtrakul S. Activities of aqueous ethanolic extracts from Combretum adenogonium Steud. Ex A. Rich (Combretaceae). BMC Complement Altern Med. (2012) 12:163. 10.1186/1472-6882-12-163 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Liu M, Katerere DR, Gray AI, Seidel V. Phytochemical and antifungal studies on Terminalia mollis and Terminalia brachystemma. Fitoterapia. (2009) 80:369–73. 10.1016/j.fitote.2009.05.006 [DOI] [PubMed] [Google Scholar]
- 74.Lavoie S, Côté I, Pichette A, Gauthier C, Ouellet M, Nagau-Lavoie F, et al. Chemical composition and anti-herpes simplex virus type 1 (HSV-1) activity of extracts from Cornus canadensis. BMC Complement Altern Med. (2017) 17:2. 10.1186/s12906-017-1618-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Hsieh CF, Chen YL, Lin CF, Ho JY, Huang CH, Chiu CH, et al. An extract from Taxodium distichum targets hemagglutinin- and neuraminidase-related activities of influenza virus in vitro. Sci Rep. (2016) 6:36015. 10.1038/srep36015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Xu H, Ma Y, Huang X, Geng C, Wang H. Bioactivity-guided isolation of anti-hepatitis B virus active sesquiterpenoids from the traditional Chinese medicine : Rhizomes of Cyperus rotundus. J Ethnopharmacol. (2015) 171:131–40. 10.1016/j.jep.2015.05.040 [DOI] [PubMed] [Google Scholar]
- 77.Danciu C, Muntean D, Alexa E, Farcas C, Oprean C, Zupko I, et al. Phytochemical characterization and evaluation of the antimicrobial, antiproliferative and pro-apoptotic potential of Ephedra alata Decne. hydroalcoholic extract against the MCF-7 breast cancer cell line. Molecules. (2018) 24:13. 10.3390/molecules24010013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Churqui MP, Lind L, Thörn K, Svensson A, Savolainen O, Aranda KT, et al. Extracts of Equisetum giganteum L and Copaifera reticulate Ducke show strong antiviral activity against the sexually transmitted pathogen herpes simplex virus type 2. J Ethnopharmacol. (2018) 210:192–7. 10.1016/j.jep.2017.08.010 [DOI] [PubMed] [Google Scholar]
- 79.Shamsabadipour S, Ghanadian M, Saeedi H, Reza Rahimnejad M, Mohammadi-Kamalabadi M, Ayatollahi SM, et al. Triterpenes and steroids from Euphorbia denticulata Lam. with anti-herpes symplex virus activity. Iran J Pharm Res. (2013) 12:759–67. [PMC free article] [PubMed] [Google Scholar]
- 80.Gyuris A, Szlávik L, Minárovits J, Vasas A, Molnár J, Hohmann J. Antiviral activities of extracts of Euphorbia hirta L. against HIV-1, HIV-2 and SIVmac251. In Vivo. (2008) 23:429–32. [PubMed] [Google Scholar]
- 81.Jiang C, Luo P, Zhao Y, Hong J, Morris-Natschke SL, Xu J, et al. Carolignans from the aerial parts of Euphorbia sikkimensis and their anti-HIV activity. J Nat Prod. (2016) 79:578–83. 10.1021/acs.jnatprod.5b01012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Shoji M, Woo SY, Masuda A, Win NN, Ngwe H, Takahashi E, et al. Anti-influenza virus activity of extracts from the stems of Jatropha multifida Linn. Collected in Myanmar. BMC Complement Altern Med. (2017) 17:76. 10.1186/s12906-017-1612-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Ghoke SS, Sood R, Kumar N, Pateriya AK, Bhatia S, Mishra A, et al. Evaluation of antiviral activity of Ocimum sanctum and Acacia arabica leaves extracts against H9N2 virus using embryonated chicken egg model. BMC Complement Altern Med. (2018) 18:174. 10.1186/s12906-018-2238-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Makau JN, Watanabe K, Mohammed MMD, Nishida N. Antiviral activity of peanut (Arachis hypogaea L.) skin extract against human influenza viruses. J Med Food. (2018) 21:777–84. 10.1089/jmf.2017.4121 [DOI] [PubMed] [Google Scholar]
- 85.Knipping K, Garssen J, van't Land B. An evaluation of the inhibitory effects against rotavirus infection of edible plant extracts. Virol J. (2012) 9:137. 10.1186/1743-422X-9-137 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Fahmy NM, Al-Sayed E, Moghannem S, Azam F, El-Shazly M, Singab AN. Breaking down the barriers to a natural antiviral agent: antiviral activity and molecular docking of Erythrina speciosa extract, fractions, and the major compound. Chem Biodivers. (2020) 17:2. 10.1002/cbdv.201900511 [DOI] [PubMed] [Google Scholar]
- 87.Donalisio M, Cagno V, Civra A, Gibellini D, Musumeci G, Rittà M, et al. The traditional use of Vachellia nilotica for sexually transmitted diseases is substantiated by the antiviral activity of its bark extract against sexually transmitted viruses. J Ethnopharmacol. (2018) 213:403–8. 10.1016/j.jep.2017.11.039 [DOI] [PubMed] [Google Scholar]
- 88.Nutan SK, Modi M, Dezzutti CS, Kulshreshtha S, Rawat AKS, Srivastava SK, et al. Extracts from Acacia catechu suppress HIV-1 replication by inhibiting the activities of the viral protease and Tat. Virol J. (2013) 10:309. 10.1186/1743-422X-10-309 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Karimi A, Rafieian-kopaei M, Moradi M. Anti -herpes simplex virus type-1 activity and phenolic content of crude ethanol extract and four corresponding fractions of Quercus brantii L Acorn. J Evid Based Complement Altern Med. (2016) 22:455–61. 10.1177/2156587216676421 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Rafieian-kopaei M, Saeedi M, Asgari S, Karimi A, Moradi M-T. Antiviral activity of Quercus persica L.: high efficacy and low toxicity. Adv Biomed Res. (2013) 2:36. 10.4103/2277-9175.109722 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Choi JG, Kim YS, Kim JH, Chung HS. Antiviral activity of ethanol extract of Geranii Herba and its components against influenza viruses via neuraminidase inhibition. Sci Rep. (2019) 9:12132. 10.1038/s41598-019-48430-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Helfer M, Koppensteiner H, Schneider M, Rebensburg S, Forcisi S, Müller C, et al. The root extract of the medicinal plant Pelargonium sidoides is a potent HIV-1 attachment inhibitor. PLoS ONE. (2014) 9:e87487. 10.1371/journal.pone.0087487 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Michaelis M, Doerr HW, Cinatl J. Investigation of the influence of EPs® 7630, a herbal drug preparation from Pelargonium sidoides, on replication of a broad panel of respiratory viruses. Phytomedicine. (2011) 18:384–6. 10.1016/j.phymed.2010.09.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Haasbach E, Hartmayer C, Hettler A, Sarnecka A, Wulle U, Ehrhardt C, et al. Antiviral activity of Ladania067, an extract from wild black currant leaves against influenza a virus in vitro and in vivo. Front Microbiol. (2014) 5:171. 10.3389/fmicb.2014.00171 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Theisen LL, Erdelmeier CAJ, Spoden GA, Boukhallouk F, Sausy A, Florin L, et al. Tannins from Hamamelis virginiana bark extract: characterization and improvement of the antiviral efficacy against influenza, a virus and human papillomavirus. PLoS ONE. (2014) 9:e88062. 10.1371/journal.pone.0088062 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Erdelmeier CAJ, Cinatl J, Rabenau H, Doerr HW, Biber A, Koch E. Antiviral and antiphlogistic activities of Hamamelis virginiana bark. Planta Med. (1996) 62:241–5. 10.1055/s-2006-957868 [DOI] [PubMed] [Google Scholar]
- 97.Chen SG, Leu YL, Cheng ML, Ting SC, Liu CC, Wang S Der, et al. Anti-enterovirus 71 activities of Melissa officinalis extract and its biologically active constituent rosmarinic acid. Sci Rep. (2017) 7:12264. 10.1038/s41598-017-12388-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Schnitzler P, Schuhmacher A, Astani A, Reichling J. Melissa officinalis oil affects infectivity of enveloped herpesviruses. Phytomedicine. (2008) 15:734–40. 10.1016/j.phymed.2008.04.018 [DOI] [PubMed] [Google Scholar]
- 99.Tang LIC, Ling APK, Koh RY, Chye SM, Voon KGL. Screening of anti-dengue activity in methanolic extracts of medicinal plants. BMC Complement Altern Med. (2012) 12:3. 10.1186/1472-6882-12-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Chiu LCM, Zhu W, Ooi VEC. A polysaccharide fraction from medicinal herb Prunella vulgaris downregulates the expression of herpes simplex virus antigen in Vero cells. J Ethnopharmacol. (2004) 93:63–8. 10.1016/j.jep.2004.03.024 [DOI] [PubMed] [Google Scholar]
- 101.Brindley MA, Widrlechner MP, McCoy JA, Murphy P, Hauck C, Rizshsky L, et al. Inhibition of lentivirus replication by aqueous extracts of Prunella vulgaris. Virol J. (2009) 6:8. 10.1186/1743-422X-6-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Ma F-W, Kong S-Y, Tan H-S, Wu R, Xia B, Zhou Y, et al. Structural characterization and antiviral effect of a novel polysaccharide PSP-2B from Prunellae Spica. Carbohydr Polym. (2016) 152:699–709. 10.1016/j.carbpol.2016.07.062 [DOI] [PubMed] [Google Scholar]
- 103.Mancini DAP, Torres RP, Pinto JR, Mancini-Filho J. Inhibition of DNA virus: Herpes-1 (HSV-1) in cellular culture replication, through an antioxidant treatment extracted from rosemary spice. Brazilian J Pharm Sci. (2009) 45:127–33. 10.1590/S1984-82502009000100016 [DOI] [Google Scholar]
- 104.Shi H, Ren K, Lv B, Zhang W, Zhao Y, Tan RX, et al. Baicalin from Scutellaria baicalensis blocks respiratory syncytial virus (RSV) infection and reduces inflammatory cell infiltration and lung injury in mice. Sci Rep. (2016) 6:35851. 10.1038/srep35851 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Cao YG, Hao Y, Li ZH, Liu ST, Wang LX. Antiviral activity of polysaccharide extract from Laminaria japonica against respiratory syncytial virus. Biomed Pharmacother. (2016) 84:1705–10. 10.1016/j.biopha.2016.10.082 [DOI] [PubMed] [Google Scholar]
- 106.Yarmolinsky L, Zaccai M, Ben-shabat S, Mills D, Huleihel M. Antiviral activity of ethanol extracts of Ficus binjamina and Lilium candidum in vitro. (2009) 26:307–13. 10.1016/j.nbt.2009.08.005 [DOI] [PubMed] [Google Scholar]
- 107.Tsai YC, Hohmann J, El-Shazly M, Chang LK, Dankó B, Kúsz N, et al. Bioactive constituents of Lindernia crustacea and its anti-EBV effect via Rta expression inhibition in the viral lytic cycle. J Ethnopharmacol. (2020) 250:112493. 10.1016/j.jep.2019.112493 [DOI] [PubMed] [Google Scholar]
- 108.Boff L, Silva IT, Argenta DF, Farias LM, Alvarenga LF, Pádua RM. Strychnos pseudoquina A. St. Hil.: a Brazilian medicinal plant with promising in vitro antiherpes activity. J Appl Microbiol. (2016) 121:1519–29. 10.1111/jam.13279 [DOI] [PubMed] [Google Scholar]
- 109.Arunkumar J, Rajarajan S. Study on antiviral activities, drug-likeness and molecular docking of bioactive compounds of Punica granatum L. to Herpes simplex virus - 2 (HSV-2). Microb Pathog. (2018) 118:301–9. 10.1016/j.micpath.2018.03.052 [DOI] [PubMed] [Google Scholar]
- 110.Haidari M, Ali M, Ward Casscells S, Madjid M. Pomegranate (Punica granatum) purified polyphenol extract inhibits influenza virus and has a synergistic effect with oseltamivir. Phytomedicine. (2009) 16:1127–36. 10.1016/j.phymed.2009.06.002 [DOI] [PubMed] [Google Scholar]
- 111.Fang C, Chen S, Wu H, Ping Y, Lin C. Honokiol, a Lignan biphenol derived from the magnolia tree, inhibits Dengue virus type 2 infection. Viruses. (2015) 7:4894–910. 10.3390/v7092852 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Baatartsogt T, Bui VN, Trinh DQ, Yamaguchi E, Gronsang D, Thampaisarn R, et al. High antiviral effects of hibiscus tea extract on the H5 subtypes of low and highly pathogenic avian influenza viruses. J Vet Med Sci. (2016) 78:1405–11. 10.1292/jvms.16-0124 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Sood R, Raut R, Tyagi P, Pareek PK, Barman TK, Singhal S, et al. Cissampelos pareira Linn: Natural source of potent antiviral activity against all four Dengue virus serotypes. PLoS Negl Trop Dis. (2015) 9:e0004255. 10.1371/journal.pntd.0004255 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Yarmolinsky L, Huleihel M, Zaccai M, Ben-shabat S. Potent antiviral flavone glycosides from Ficus benjamina leaves. Fitoterapia. (2012) 83:362–7. 10.1016/j.fitote.2011.11.014 [DOI] [PubMed] [Google Scholar]
- 115.Aref HL, Gaaliche B, Fekih A. in vitro cytotoxic and antiviral activities of Ficus carica latex extracts. Nat Prod Res. (2011) 25:310–9. 10.1080/14786419.2010.528758 [DOI] [PubMed] [Google Scholar]
- 116.Ghosh M, Civra A, Rittà M, Cagno V, Mavuduru SG, Awasthi P, et al. Ficus religiosa L. bark extracts inhibit infection by herpes simplex virus type 2 in vitro. Arch Virol. (2016) 161:3509–14. 10.1007/s00705-016-3032-3 [DOI] [PubMed] [Google Scholar]
- 117.Huang NC, Hung WT, Tsai WL, Lai FY, Lin YS, Huang MS, et al. Ficus septica plant extracts for treating Dengue virus in vitro. Peer J. (2017) 5:e3448. 10.7717/peerj.3448 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Sa S, Abdullahi H. Antifungal activity and phytochemical analysis of Ficus sycomorus leaf extract on malassezia glubosa. Adv Plants Agric Res. (2018) 8:432–6. 10.15406/apar.2018.08.00362 [DOI] [Google Scholar]
- 119.Batiha GES, Alkazmi LM, Wasef LG, Beshbishy AM, Nadwa EH, Rashwan EK. Syzygium aromaticum L. (myrtaceae): traditional uses, bioactive chemical constituents, pharmacological and toxicological activities. Biomolecules. (2020) 10:202. 10.3390/biom10020202 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Benzekri R, Bouslama L, Papetti A, Hammami M, Smaoui A, Limam F. Anti HSV-2 activity of Peganum harmala (L.) and isolation of the active compound. Microb Pathog. (2018) 114:291–8. 10.1016/j.micpath.2017.12.017 [DOI] [PubMed] [Google Scholar]
- 121.Li J, Yang X, Huang L. Anti-Influenza virus activity and constituents characterization of Paeonia delavayi extracts. Molecules. (2016) 21:1133. 10.3390/molecules21091133 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Ho JY, Chang HW, Lin CF, Liu CJ, Hsieh CF, Horng JT. Characterization of the anti-influenza activity of the Chinese herbal plant Paeonia lactiflora. Viruses. (2014) 6:1861–75. 10.3390/v6041861 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Cho JK, Curtis-Long MJ, Lee KH, Kim DW, Ryu HW, Yuk HJ, et al. Geranylated flavonoids displaying SARS-CoV papain-like protease inhibition from the fruits of Paulownia tomentosa. Bioorganic Med Chem. (2013) 21:3051–7. 10.1016/j.bmc.2013.03.027 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Lv J, Yu S, Wang Y, Wang D, Zhu H, Cheng R, et al. Anti-hepatitis B virus norbisabolane sesquiterpenoids from Phyllanthus acidus and the establishment of their absolute configurations using theoretical calculations. J Org Chem. (2014) 79:5432–47. 10.1021/jo5004604 [DOI] [PubMed] [Google Scholar]
- 125.Ravikumar YS, Ray U, Nandhitha M, Perween A, Raja Naika H, Khanna N, et al. Inhibition of hepatitis C virus replication by herbal extract: Phyllanthus amarus as potent natural source. Virus Res. (2011) 158:89–97. 10.1016/j.virusres.2011.03.014 [DOI] [PubMed] [Google Scholar]
- 126.Tan WC, Jaganath IB, Manikam R, Sekaran SD. Evaluation of antiviral activities of four local Malaysian phyllanthus species against herpes simplex viruses and possible antiviral target. Int J Med Sci. (2013) 10:1817–29. 10.7150/ijms.6902 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Zhang X, Yang LM, Liu GM, Liu YJ, Zheng CB, Lv YJ, et al. Potent anti-HIV activities and mechanisms of action of a pine cone extract from Pinus yunnanensis. Molecules. (2012) 17:6916–29. 10.3390/molecules17066916 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Hsu WC, Chang SP, Lin LC, Li CL, Richardson CD, Lin CC, et al. Limonium sinense and gallic acid suppress hepatitis C virus infection by blocking early viral entry. Antiviral Res. (2015) 118:139–47. 10.1016/j.antiviral.2015.04.003 [DOI] [PubMed] [Google Scholar]
- 129.Chavan RD, Shinde P, Girkar K, Madage R, Chowdhary A. Assessment of Anti-Influenza activity and hemagglutination inhibition of Plumbago indica and Allium sativum extracts. Pharmacognosy Res. (2016) 8:105–11. 10.4103/0974-8490.172562 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Xiong HR, Luo J, Hou W, Xiao H, Yang ZQ. The effect of Emodin, an anthraquinone derivative extracted from the roots of Rheum tanguticum, against herpes simplex virus in vitro and in vivo. J Ethnopharmacol. (2011) 133:718–23. 10.1016/j.jep.2010.10.059 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Su M, Li Y, Leung KT, Cen Y, Li T, Chen R, et al. Antiviral activity and constituent of Ardisia Chinensis Benth against coxsackie B3 virus. Phytother Res. (2006) 20:634–9. 10.1002/ptr.1912 [DOI] [PubMed] [Google Scholar]
- 132.Hossan MS, Fatima A, Rahmatullah M, Khoo TJ, Nissapatorn V, Galochkina AV, et al. Antiviral activity of Embelia ribes Burm. f. against influenza virus in vitro. Arch Virol. (2018) 163:2121–31. 10.1007/s00705-018-3842-6 [DOI] [PubMed] [Google Scholar]
- 133.Hung TC, Jassey A, Lin CJ, Liu CH, Lin CC, Yen MH, et al. Methanolic extract of rhizoma coptidis inhibits the early viral entry steps of hepatitis C virus infection. Viruses. (2018) 10:669. 10.3390/v10120669 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Kim HB, Lee CY, Kim SJ, Han JH, Choi KH. Medicinal herb extracts ameliorate impaired growth performance and intestinal lesion of newborn piglets challenged with the virulent porcine epidemic diarrhea virus. J Anim Sci Technol. (2015) 57:33. 10.1186/s40781-015-0065-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Shin WJ, Lee KH, Park MH, Seong BL. Broad-spectrum antiviral effect of Agrimonia pilosa extract on influenza viruses. Microbiol Immunol. (2010) 54:11–9. 10.1111/j.1348-0421.2009.00173.x [DOI] [PubMed] [Google Scholar]
- 136.Bisignano C, Mandalari G, Smeriglio A, Trombetta D, Pizzo MM, Pennisi R, et al. Almond skin extracts abrogate HSV-1 replication by blocking virus binding to the cell. Viruses. (2017) 9:178. 10.3390/v9070178 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Ratnoglik SL, Aoki C, Sudarmono P, Komoto M, Deng L, Shoji I, et al. Antiviral activity of extracts from Morinda citrifolia leaves and chlorophyll catabolites, pheophorbide a and pyropheophorbide a, against hepatitis C virus. Microbiol Immunol. (2014) 58:188–94. 10.1111/1348-0421.12133 [DOI] [PubMed] [Google Scholar]
- 138.Pratheeba T, Taranath V, Sai Gopal DVR, Natarajan D. Antidengue potential of leaf extracts of Pavetta tomentosa and Tarenna asiatica (Rubiaceae) against dengue virus and its vector Aedes aegypti (Diptera: Culicidae). Heliyon. (2019) 5:e02732. 10.1016/j.heliyon.2019.e02732 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Brijesh S, Daswani P, Tetali P, Antia N, Birdi T. Studies on the antidiarrhoeal activity of Aegle marmelos unripe fruit : validating its traditional usage. BMC Complement Altern Med. (2009) 12:47. 10.1186/1472-6882-9-47 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Apriyanto DR, Aoki C, Hartati S, Hanafi M, Kardono LBS, Arsianti A, et al. Anti-hepatitis C virus activity of a crude extract from longan (Dimocarpus longan Lour.) leaves. Jpn J Infect Dis. (2016) 69:213–20. 10.7883/yoken.JJID.2015.107 [DOI] [PubMed] [Google Scholar]
- 141.Song JH, Ahn JH, Kim SR, Cho S, Hong EH, Kwon BE, et al. Manassantin B shows antiviral activity against coxsackievirus B3 infection by activation of the STING/TBK-1/IRF3 signalling pathway. Sci Rep. (2019) 9:9413. 10.1038/s41598-019-45868-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Wang GW, Hu WT, Huang BK, Qin LP. Illicium verum: a review on its botany, traditional use, chemistry and pharmacology. J Ethnopharmacol. (2011) 136:10–20. 10.1016/j.jep.2011.04.051 [DOI] [PubMed] [Google Scholar]
- 143.Abdelgawad AAM. Tamarix nilotica (Ehrenb) bunge: a review of phytochemistry and pharmacology. J Microb Biochem Technol. (2017) 9:544–53. 10.4172/1948-5948.1000340 [DOI] [Google Scholar]
- 144.Ryu YB, Jeong HJ, Kim JH, Kim YM, Park JY, Kim D, et al. Biflavonoids from Torreya nucifera displaying SARS-CoV 3CLpro inhibition. Bioorganic Med Chem. (2010) 18:7940–7. 10.1016/j.bmc.2010.09.035 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Karamese M, Aydogdu S, Karamese SA, Altoparlak U. Preventive effects of a major component of green tea, Epigallocathechin-3-Gallate, on Hepatitis-B virus DNA replication. Asian Pac J Cancer Prev. (2015) 16:40199–202. 10.7314/APJCP.2015.16.10.4199 [DOI] [PubMed] [Google Scholar]
- 146.Xu J, Wang J, Deng F, Hu Z, Wang H. Green tea extract and its major component epigallocatechin gallate inhibits hepatitis B virus in vitro. Antiviral Res. (2008) 78:242–9. 10.1016/j.antiviral.2007.11.011 [DOI] [PubMed] [Google Scholar]
- 147.Dai J, Tao H, Min Q, Zhu Q. Anti-hepatitis B virus activities of friedelolactones from Viola diffusa Ging. Phytomedicine. (2015) 22:724–9. 10.1016/j.phymed.2015.05.001 [DOI] [PubMed] [Google Scholar]
- 148.Kwon HJ, Kim HH, Yoon SY, Ryu YB, Chang JS, Cho KO, et al. in vitro inhibitory activity of Alpinia katsumadai extracts against influenza virus infection and hemagglutination. Virol J. (2010) 7:307. 10.1186/1743-422X-7-307 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Thuy BTP, My TTA, Hai NTT, Hieu LT, Hoa TT, Thi Phuong Loan H, et al. Investigation into SARS-CoV-2 resistance of compounds in garlic essential oil. ACS Omega. (2020) 5:8312–8320. 10.1021/acsomega.0c00772 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Kim DW, Seo KH, Curtis-Long MJ, Oh KY, Oh JW, Cho JK, et al. Phenolic phytochemical displaying SARS-CoV papain-like protease inhibition from the seeds of Psoralea corylifolia. J Enzyme Inhib Med Chem. (2014) 29:59–63. 10.3109/14756366.2012.753591 [DOI] [PubMed] [Google Scholar]
- 151.Li SY, Chen C, Zhang HQ, Guo HY, Wang H, Wang L, et al. Identification of natural compounds with antiviral activities against SARS-associated coronavirus. Antiviral Res. (2005) 67:18–23. 10.1016/j.antiviral.2005.02.007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152.Ulasli M, Gurses SA, Bayraktar R, Yumrutas O, Oztuzcu S, Igci M, et al. The effects of Nigella sativa (Ns), Anthemis hyalina (Ah) and Citrus sinensis (Cs) extracts on the replication of coronavirus and the expression of TRP genes family. Mol Biol Rep. (2014) 41:1703–11. 10.1007/s11033-014-3019-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Ryu YB, Park SJ, Kim YM, Lee JY, Seo WD, Chang JS, et al. SARS-CoV 3CLpro inhibitory effects of quinone-methide triterpenes from Tripterygium regelii. Bioorganic Med Chem Lett. (2010) 20:1873–76. 10.1016/j.bmcl.2010.01.152 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Loizzo MR, Saab AM, Tundis R, Statti GA, Menichimi F, Lampronti D, et al. Phytochemical analysis and in vitro antiviral activities of the essential oils of seven Lebanon species. Chem Biodivers. (2008) 5:461–70. 10.1002/cbdv.200890045 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Wu C, Liu Y, Yang Y, Zhang P, Zhong W, Wang Y, et al. Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharm Sin B. (2020) 10:766–88. 10.1016/j.apsb.2020.02.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Wen CC, Shyur LF, Jan JT, Liang PH, Kuo CJ, Arulselvan P, et al. Traditional Chinese medicine herbal extracts of Cibotium barometz, Gentiana scabra, Dioscorea batatas, Cassia tora, and Taxillus chinensis inhibit SARS-CoV replication. J Tradit Complement Med. (2011) 1:41–50. 10.1016/S2225-4110(16)30055-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.Zhuang M, Jiang H, Suzuki Y, Li X, Xiao P, Tanaka T, et al. Procyanidins and butanol extract of Cinnamomi Cortex inhibit SARS-CoV infection. Antiviral Res. (2009) 82:73–81. 10.1016/j.antiviral.2009.02.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Ho TY, Wu SL, Chen JC, Li CC, Hsiang CY. Emodin blocks the SARS coronavirus spike protein and angiotensin-converting enzyme 2 interaction. Antiviral Res. (2007) 74:92–101. 10.1016/j.antiviral.2006.04.014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Luo W, Su X, Gong S, Qin Y, Liu W, Li J, et al. Anti-SARS coronavirus 3C-like protease effects of Rheum palmatum L. extracts. Biosci Trends. (2009) 3:124–6. [PubMed] [Google Scholar]
- 160.Yang C, Lee Y, Hsu H, Shih C, Chao Y, Lee S. Targeting coronaviral replication and cellular JAK2 mediated dominant NF- κB activation for comprehensive and ultimate inhibition of coronaviral activity. Sci Rep. (2017) 7:4105. 10.1038/s41598-017-04203-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Adhikari SP, Meng S, Wu Y, Mao Y, Ye R, Wang Q, et al. Novel Coronavirus during the early outbreak period: epidemiology, causes, clinical manifestation and diagnosis, prevention and control. Infect Dis Poverty. (2020) 9:29. 10.1186/s40249-020-00646-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Nickbakhsh S, Ho A, Marques DFP, McMenamin J, Gunson RN, Murcia PR. Epidemiology of seasonal coronaviruses: establishing the context for the emergence of Coronavirus disease 2019. J Infect Dis. (2020) 222:17–25. 10.1093/infdis/jiaa185 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Wu D, Wu T, Liu Q, Yang Z. The SARS-CoV-2 outbreak: what we know. Int J Infect Dis. (2020) 94:44–8. 10.1016/j.ijid.2020.03.004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Li Q, Guan X, Wu P, Wang X, Zhou L, Tong Y, et al. Early transmission dynamics in Wuhan, China, of novel coronavirus-infected pneumonia. N Engl J Med. (2020) 382:1199–207. 10.1056/NEJMoa2001316 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165.Phan LT, Nguyen TV, Luong QC, Nguyen TV, Nguyen TV, Nguyen HT, et al. Importation and human-to-human transmission of a novel Coronavirus in Vietnam. N Engl J Med. (2020) 382:872–4. 10.1056/NEJMc2001272 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Cots JM, Alós J, Bárcena M, Boleda X. COVID-19 in Brazil: “So what?”. Lancet. (2020) 395:1461. 10.1016/S0140-6736(20)31095-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.COVID-19 Dashboard by the Center for Systems Science Engineering (CSSE) at Johns Hopkins University (JHU). Available online at: https://coronavirus.jhu.edu/map.html (accessed July 5, 2020).
- 168.Yang H, Bartlam M, Rao Z. Drug design targeting the main protease, the Achilles Heel of coronaviruses. Curr Pharm Des. (2006) 12:4573–90. 10.2174/138161206779010369 [DOI] [PubMed] [Google Scholar]
- 169.Mousavizadeh L, Ghasemi S. Genotype and phenotype of COVID-19: their roles in pathogenesis. J Microbiol Immunol Infect. (2020). 10.1016/j.jmii.2020.03.022. [Epub ahead of print]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Ou X, Liu Y, Lei X, Li P, Mi D, Ren L, et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat Commun. (2020) 11:1620. 10.1038/s41467-020-15562-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171.Coutard B, Valle C, de Lamballerie X, Canard B, Seidah NG, Decroly E. The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antiviral Res. (2020) 176:140742. 10.1016/j.antiviral.2020.104742 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172.Liang Y, Wang M, Chien C, Yarmishyn AA. Highlight of immune pathogenic response and hematopathologic effect in SARS-CoV, MERS-CoV and SARS-Cov-2 infection. Front Immunol. (2020) 11:1022. 10.3389/fimmu.2020.01022 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 173.Chen B, Tian E, He B, Tian L, Han R, Wang S, et al. Overview of lethal human coronaviruses. Sig Transduct Target Ther. (2020) 5:89. 10.1038/s41392-020-0190-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174.Zhou Y, Hou Y, Shen J, Huang Y, Martin W, Cheng F. Network-based drug repurposing for novel coronavirus 2019-nCoV/SARS-CoV-2. Cell Discov. (2020) 6:14. 10.1038/s41421-020-0153-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.Xu J, Zhao S, Teng T, Abdalla AE, Zhu W, Xie L, et al. Systematic comparison of two animal-to-human transmitted human Coronaviruses: SARS-CoV-2 and SARS-CoV. Viruses. (2020) 12:244. 10.3390/v12020244 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Prajapat M, Sarma P, Shekhar N, Avti P, Sinha S, Kaur H, et al. Drug targets for corona virus: a systematic review. Indian J Pharmacol. (2020) 52:56–65. 10.4103/ijp.IJP_115_20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.Rabaan AA, Al-Ahmed SH, Haque S, Sah R, Tiwari R, Malik YS, et al. SARS-CoV-2, SARS-CoV, and MERS-COV: a comparative overview. Le Infez Med. (2020) 28:174–84. [PubMed] [Google Scholar]
- 178.Tang X, Wu C, Li X, Song Y, Yao X, Wu X, et al. On the origin and continuing evolution of SARS-CoV-2. Natl Sci Rev. (2020) 3:nwaa036 10.1093/nsr/nwaa036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179.Zhang H, Penninger JM, Li Y, Zhong N, Slutsky AS. Angiotensin-converting enzyme 2 (ACE2) as a SARS-CoV-2 receptor: molecular mechanisms and potential therapeutic target. Intensive Care Med. (2020) 46:586–90. 10.1007/s00134-020-05985-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180.Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. (2020) 2:52 10.1016/j.cell.2020.02.052 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181.Skariayachan S, Challapilli SB, Pacckirisamy S, Kumargowda ST, Sridhar VS. Recent aspects on the pathogenesis mechanism, animal models and novel therapeutics interventions for Middle East respiratory syndrome coronavirus infections. Front Microbiol. (2019) 10:569. 10.3389/fmicb.2019.00569 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 182.Shereen MA, Khan S, Kazmi A, Bashir N, Siddique R. COVID-19 infection: origin, transmission, and characteristics of human coronaviruses. J Adv Res. (2020) 24:91–8. 10.1016/j.jare.2020.03.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 183.Rampersad S, Tennant P. Replication and expression strategies of viruses. Viruses. (2018) 2018:55–82. 10.1016/B978-0-12-811257-1.00003-6 [DOI] [Google Scholar]
- 184.Wang H, Xue S, Yang H, Chen C. Recent progress in the discovery of inhibitors targeting coronavirus proteases. Virol Sin. (2016) 31:24–30. 10.1007/s12250-015-3711-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185.Cao W, Li T. COVID-19: towards understanding of pathogenesis. Cell Res. (2020) 30:367–9. 10.1038/s41422-020-0327-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 186.Rockx B, Kuiken T, Herfst S, Bestebroer T, Lamers MM, Oude Munnink BB, et al. Comparative pathogenesis of COVID-19, MERS, and SARS in a nonhuman primate model. Science. (2020) 368:1012–5. 10.1126/science.abb7314 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 187.Chen Y, Liu Q, Guo D. Emerging coronaviruses: genome structure, replication, and pathogenesis. J Med Virol. (2020) 92:418–23. 10.1002/jmv.25681 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 188.Kim D, Lee JY, Yang JS, Kim JW, Kim VN, Chang H. The architecture of SARS-CoV-2 transcriptome. Cell. (2020) 181:914–21. 10.1016/j.cell.2020.04.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189.Shi Y, Wang Y, Shao C, Huang J, Gan J, Huang X, et al. COVID-19 infection: the perspectives on immune responses. Cell Death Differ. (2020) 27:1451–4. 10.1038/s41418-020-0530-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 190.Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. (2020) 395:497–506. 10.1016/S0140-6736(20)30183-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Rothan HA, Byrareddy SN. The epidemiology and pathogenesis of coronavirus disease (COVID-19) outbreak. J Autoimmun. (2020) 109:102433. 10.1016/j.jaut.2020.102433 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 192.Devaux CA, Rolain JM, Raoult D. ACE2 receptor polymorphism: susceptibility to SARS-CoV-2, hypertension, multi-organ failure, and COVID-19 disease outcome. J Microbiol Immunol Infect. (2020) 53:425–35. 10.1016/j.jmii.2020.04.015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193.Tahir ul Qamar M, Alqahtani SM, Alamri MA, Chen LL. Structural basis of SARS-CoV-2 3CLpro and anti-COVID-19 drug discovery from medicinal plants. J Pharm Anal. (in press) 10.20944/preprints202002.0193.v1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 194.Shang J, Ye G, Shi K, Wan Y, Luo C, Aihara H, et al. Structural basis of receptor recognition by SARS-CoV-2. Nature. (2020) 581:221–4. 10.1038/s41586-020-2179-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 195.Hilgenfeld R. From SARS to MERS: crystallographic studies on coronaviral proteases enable antiviral drug design. FEBS J. (2014) 281:4085–96. 10.1111/febs.12936 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 196.Zeng Q, Langereis MA, van Vliet ALW, Huizinga EG, de Groot RJ. Structure of coronavirus hemagglutinin-esterase offers insight into corona and influenza virus evolution. Proc Natl Acad Sci USA. (2008) 105:9065–9. 10.1073/pnas.0800502105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 197.Wang W, Xu Y, Gao R, Lu R, Han K, Wu G, et al. Detection of SARS-CoV-2 in different types of clinical specimens. JAMA. (2020) 1843–44. 10.1001/jama.2020.3786 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 198.Saxena SK, Kumar S, Maurya VK, Sharma R, Dandu HR, Bhatt MLB. Current insight into the novel coronavirus disease 2019 (COVID-19). Nat Public Health Emerg Collect. (2020) 2019:1–8. 10.1007/978-981-15-4814-7_132582323 [DOI] [Google Scholar]
- 199.Liu C, Zhou Q, Li Y, Garner LV, Watkins SP, Carter LJ, et al. Research and development on therapeutic agents and vaccines for covid-19 and related human coronavirus diseases. ACS Cent Sci. (2020) 6:315–31. 10.1021/acscentsci.0c00272 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 200.Salata C, Calistri A, Parolin C, Palù G. Coronaviruses: a paradigm of new emerging zoonotic diseases. Pathog Dis. (2020) 77:ftaa006. 10.1093/femspd/ftaa006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 201.Wilder-smith A, Chiew CJ, Lee VJ. Can we contain the COVID-19 outbreak with the same measures as for SARS? Lancet Infect Dis. (2020) 20:e102–7. 10.1016/S1473-3099(20)30129-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 202.Li G, De Clercq E. Therapeutic options for the 2019 novel coronavirus (2019-nCoV). Nat Rev Drug Discov. (2020) 19:149–50. 10.1038/d41573-020-00016-0 [DOI] [PubMed] [Google Scholar]
- 203.Andersen PI, Ianevski A, Lysvand H, Vitkauskiene A, Oksenych V, Bjørås M, et al. Discovery and development of safe-in-man broad-spectrum antiviral agents. Int J Infect Dis. (2020) 93:268–76. 10.1016/j.ijid.2020.02.018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 204.Liu M, Yu Q, Xiao H, Yi Y, Cheng H, Putra DF, et al. Antiviral activity of Illicium verum Hook. f. extracts against grouper iridovirus infection. J Fish Dis. (2020) 43:531–40. 10.1111/jfd.13146 [DOI] [PubMed] [Google Scholar]
- 205.Fuzimoto AD, Isidoro C. The antiviral and the coronavirus-host protein pathways inhibiting properties of herbs and natural compounds - additional weapons in the fight against the COVID-19 pandemic? J Tradit Complement Med. (2020) 10:405–19. 10.1016/j.jtcme.2020.05.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 206.Chen L, Xiong J, Bao L, Shi Y. Convalescent plasma as a potential therapy for COVID-19. Lancet Infect Dis. (2020) 20:398–400. 10.1016/S1473-3099(20)30141-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 207.Cao X. COVID-19: immunopathology and its implications for therapy. Nat Rev Immunol. (2020) 20:269–70. 10.1038/s41577-020-0308-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 208.Vankadari N. Arbidol: a potential antiviral drug for the treatment of SARS-CoV-2 by blocking trimerization of the spike glycoprotein. Int J Antimicrob Agents. (2020) 28:105998. 10.1016/j.ijantimicag.2020.105998 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 209.Khan SA, Zia K, Ashraf S, Uddin R, Ul-Haq Z. Identification of chymotrypsin-like protease inhibitors of SARS-CoV-2 via integrated computational approach. J Biomol Struct Dyn. (2020) 2020:1–10. 10.1080/07391102.2020.1751298 [DOI] [PubMed] [Google Scholar]
- 210.Delang L, Abdelnabi R, Neyts J. Favipiravir as a potential countermeasure against neglected and emerging RNA viruses. Antiviral Res. (2018) 153:85–94. 10.1016/j.antiviral.2018.03.003 [DOI] [PubMed] [Google Scholar]
- 211.Wang M, Cao R, Zhang L, Yang X, Liu J, Xu M. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. (2020) 30:269–71. 10.1038/s41422-020-0282-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 212.Caly L, Druce JD, Catton MG, Jans DA, Wagstaff KM. The FDA-approved drug ivermectin inhibits the replication of SARS-CoV-2 in vitro. Antiviral Res. (2020) 178:3–6. 10.1016/j.antiviral.2020.104787 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 213.Cao B, Wang Y, Wen D, Liu W, Wang J, Fan G, et al. A trial of lopinavir-ritonavir in adults hospitalized with severe covid-19. N Engl J Med. (2020) 382:1787–99. 10.1056/NEJMc2008043 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 214.Elmezayen AD, Al-Obaidi A, Sahin AT, Yelekçi K. Drug repurposing for coronavirus (COVID-19): in silico screening of known drugs against coronavirus 3CL hydrolase and protease enzymes. J Biomol Struct Dyn. (2020) 2020:1–13. 10.1080/07391102.2020.1758791 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 215.Muralidharan N, Sakthivel R, Velmurugan D, Gromiha MM. Computational studies of drug repurposing and synergism of lopinavir, oseltamivir and ritonavir binding with SARS-CoV-2 protease against COVID-19. J Biomol Struct Dyn. (2020) 2020:1–6. 10.1080/07391102.2020.1752802 [DOI] [PubMed] [Google Scholar]
- 216.Dong L, Hu S, Gao J. Discovering drugs to treat coronavirus disease 2019 (COVID-19). Drug Discov Ther. (2020) 14:58–60. 10.5582/ddt.2020.01012 [DOI] [PubMed] [Google Scholar]
- 217.Sarma P, Sekhar N, Prajapat M, Avti P, Kaur H, Kumar S, et al. In-silico homology assisted identification of inhibitors of RNA binding against 2019-nCoV N-protein (N terminal domain). J Biomol Struct Dyn. (2020) 2020:1–9. 10.1080/07391102.2020.1753580 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 218.Baron SA, Devaux C, Colson P, Raoult D, Rolain JM. Teicoplanin: an alternative drug for the treatment of COVID-19? Int J Antimicrob Agents. (2020) 55:105944. 10.1016/j.ijantimicag.2020.105944 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 219.Enmozhi SK, Raja K, Sebastine I, Joseph J. Andrographolide as a potential inhibitor of SARS-CoV-2 main protease: an in silico approach. J Biomol Struct Dyn. (2020) 2020:1–7. 10.1080/07391102.2020.1760136 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 220.Peng S, Li Z, Zou L, Liu W, Liu C, McClements DJ. Enhancement of curcumin bioavailability by encapsulation in Sophorolipid-coated nanoparticles: an in vitro and in vivo study. Agric Food Chem. (2018) 66:1488–97. 10.1021/acs.jafc.7b05478 [DOI] [PubMed] [Google Scholar]
- 221.Wu R, Wang L, Kuo HCD, Shannar A, Peter R, Chou PJ, et al. An update on current therapeutic drugs treating COVID-19. Curr Pharmacol Rep. (2020) 11:1–15. 10.1007/s40495-020-00216-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 222.Maurya VK, Kumar S, Bhatt MLB, Saxena SK. Therapeutic development and drugs for the treatment of COVID-19. Nat Public Health Emerg Collect. (2020) 2019:109–26. 10.1007/978-981-15-4814-7_10 [DOI] [Google Scholar]
- 223.Koparde AA, Doijad RC, Magdum CS, Koparde AA. Natural products in drug discovery. In: Doijad RC, editor. Pharmacognosy - Medicinal Plants. Rijeka: IntechOpen; (2019). p. 14 10.5772/intechopen.82860 [DOI] [Google Scholar]
- 224.Ekor M. The growing use of herbal medicines: issues relating to adverse reactions and challenges in monitoring safety. Front Neurol. (2014) 4:177. 10.3389/fphar.2013.00177 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 225.Ganjhu RK, Mudga PP, Maity H, Dowarha D, Devadiga S, Nag S, et al. Herbal plants and plant preparations as a remedial approach for viral diseases. Virus Dis. (2015) 26:225–36. 10.1007/s13337-015-0276-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 226.Chitemerere TA, Mukanganyama S. Evaluation of cell membrane integrity as a potential antimicrobial target for plant products. BMC Complement Altern Med. (2014) 14:278. 10.1186/1472-6882-14-278 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 227.Anandhi D, Srinivasan PT, Kumar GP, Jagatheesh S. Original research article DNA fragmentation induced by the glycosides and flavonoids from C. coriaria. Int J Curr Microbiol App Sci. (2014) 3:666–73. [Google Scholar]
- 228.Zhao X, Zhao F, Zhong N. Quorum sensing inhibition and anti-biofilm activity of traditional Chinese medicine. In: El-Samragy Y, editor. Food Safety - Some Global Trends. Itratech Open; (2018). p. 14 10.5772/intechopen.74658 [DOI] [Google Scholar]
- 229.Radulovic NS, Blagojevic PD, Stojanovic-Radic ZZ, Stojanovic NM. Antimicrobial plant metabolites: structural diversity and mechanism of action. Curr Med Chem. (2013) 20:932–52. 10.2174/0929867311320070008 [DOI] [PubMed] [Google Scholar]
- 230.Mogosanu GD, Grumezescu AM, Huang KS, Bejenaru LE, Bejenaru C. Prevention of microbial communities: novel approaches based natural products. Curr Pharm Biotechnol. (2015) 16:94–111. 10.2174/138920101602150112145916 [DOI] [PubMed] [Google Scholar]
- 231.Anand U, Jacobo-Herrera N, Altemimi A, Lakhssassi N. A comprehensive review on medicinal plants as antimicrobial therapeutics: potential avenues of biocompatible drug discovery. Metabolites. (2019) 9:258. 10.3390/metabo9110258 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 232.Zhang D, Wu K, Zhang X, Deng S, Peng B. In silico screening of Chinese herbal medicines with the potential to directly inhibit 2019 novel coronavirus. J Integr Med. (2020) 18:152–8. 10.1016/j.joim.2020.02.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 233.Kiran G, Karthik L, Shree Devi MS, Sathiyarajeswaran P, Kanakavalli K, Kumar KM, et al. In silico computational screening of Kabasura Kudineer - official Siddha Formulation and JACOM against SARS-CoV-2 spike protein. J Ayurveda Integr Med. (in press) 10.1016/j.jaim.2020.05.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 234.Subbaiyan A, Ravichandran K, Singh SV, Sankar M, Thomas P, Dhama K, et al. In silico molecular docking analysis targeting SARS-CoV-2 spike protein and selected herbal constituents. J Pure Appl Microbiol. (2020) 14:989–98. 10.22207/JPAM.14.SPL1.37 [DOI] [Google Scholar]
- 235.Nivetha R, Bhuvaragavan S, Janarthanan S. Inhibition of multiple SARS-CoV-2 proteins by an antiviral biomolecule, seselin from Aegle marmelos deciphered using molecular docking analysis. Res Sq. (2020) 2020:v1 10.21203/rs.3.rs-31134/v1 [DOI] [PubMed] [Google Scholar]
- 236.Basu A, Sarkar A, Maulik U. Computational approach for the design of potential spike protein binding natural compounds in SARS- CoV2. Pharmacodynamics. (2020). 10.21203/rs.3.rs-33181/v1. [Epub ahead of print]. [DOI] [Google Scholar]
- 237.Krishnasamy R, Anand T, Baba M, Bharath MV, Phuntsho J, Arunachalam D, et al. In silico analysis of active compounds from siddha herbal infusion of Ammaiyar Koondhal Kudineer (Akk) against SARS-CoV-2 spike protein and its ACE2 receptor complex. SSRN Online J. (2020). 10.2139/ssrn.3578294. [Epub ahead of print]. [DOI] [Google Scholar]
- 238.Pandit M, Latha N. In silico studies reveal potential antiviral activity of phytochemicals from medicinal plants for the treatment of COVID-19 infection. Res Sq. (2020). 10.21203/rs.3.rs-22687/v1. [Epub ahead of print]. [DOI] [Google Scholar]
- 239.Pandeya KB, Ganeshpurkar A, Kumar Mishra M. Natural RNA dependent RNA polymerase inhibitors: molecular docking studies of some biologically active alkaloids of Argemone mexicana. Med Hypotheses. (2020) 2020:109905 10.1016/j.mehy.2020.109905 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 240.Aanouz I, Belhassan A, El-Khatabi K, Lakhlifi T, El-ldrissi M, Bouachrine M. Moroccan medicinal plants as inhibitors against SARS-CoV-2 main protease: computational investigations. J Biomol Struct Dyn. (2020) 2020:1–9. 10.1080/07391102.2020.1758790 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 241.Bhardwaj VK, Singh R, Sharma J, Rajendran V, Purohit R, Kumar S. Identification of bioactive molecules from tea plant as SARS-CoV-2 main protease inhibitors. J Biomol Struct Dyn. (2020) 2020:1–11. 10.1080/07391102.2020.1766572 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 242.Serseg T, Benarous K, Yousfi M. Hispidin and Lepidine E: two natural compounds and folic acid as potential inhibitors of 2019-novel coronavirus main protease (2019-nCoVMpro), molecular docking and SAR study. Curr Comput Aided Drug Des. (2020). 10.2174/1573409916666200422075440. [Epub ahead of print]. [DOI] [PubMed] [Google Scholar]
- 243.Kumar A, Choudhir G, Shukla SK, Sharma M, Tyagi P, Bhushan A, et al. Identification of phytochemical inhibitors against main protease of COVID-19 using molecular modeling approaches. J Biomol Struct Dyn. (2020) 2020:1–11. 10.21203/rs.3.rs-31210/v1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 244.Islam MT, Sarkar C, El-Kersh DM, Jamaddar S, Uddin SJ, Shilpi JA, et al. Natural products and their derivatives against coronavirus: a review of the non-clinical and pre-clinical data. Phyther Res. (2020). 10.1002/ptr.6700 [DOI] [PubMed] [Google Scholar]
- 245.Sayed AM, Khattab AR, AboulMagd AM, Hassan HM, Rateb ME, Zaid H, et al. Nature as a treasure trove of potential anti-SARS-CoV drug leads: a structural/mechanistic rationale. RSC Adv. (2020) 10:19790–802. 10.1039/D0RA04199H [DOI] [PMC free article] [PubMed] [Google Scholar]
- 246.Sepay N, Sepay N, Al Hoque A, Mondal R, Halder UC, Muddassir M. In silico fight against novel coronavirus by finding chromone derivatives as inhibitor of coronavirus main proteases enzyme. Struct Chem. (2020) 2020:1–10. 10.1007/s11224-020-01537-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 247.Umesh U, Kundu D, Selvaraj C, Singh SK, Dubey VK. Identification of new anti-nCoV drug chemical compounds from Indian spices exploiting SARS-CoV-2 main protease as target. J Biomol Struct Dyn. (2020) 2020:1–9. 10.1080/07391102.2020.1763202 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 248.Singh A, Mishra A. Leucoefdin a potential inhibitor against SARS CoV-2 M. J Biomol Struct Dyn. (2020) 2020:1–6. 10.1080/07391102.2020.1777903 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 249.Gupta MK, Vemula S, Donde R, Gouda G, Behera L, Vadde R. In-silico approaches to detect inhibitors of the human severe acute respiratory syndrome coronavirus envelope protein ion channel. J Biomol Struct Dyn. (2020) 2020:1–11. 10.1080/07391102.2020.1751300 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 250.Joshi RS, Jagdale SS, Bansode SB, Shankar SS, Tellis MB, Pandya VK, et al. Discovery of potential multi-target-directed ligands by targeting host-specific SARS-CoV-2 structurally conserved main protease. J Biomol Struct Dyn. (2020) 2020:1–6. 10.20944/preprints202004.0068.v2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 251.Joshi T, Joshi T, Sharma P, Mathpal S, Pundir H, Bhatt V, et al. In silico screening of natural compounds against COVID-19 by targeting Mpro and ACE2 using molecular docking. Eur Rev Med Pharmacol Sci. (2020) 24:4529–36. 10.26355/eurrev_202004_21036 [DOI] [PubMed] [Google Scholar]
- 252.Salman S, Shah FH, Idrees J, Idrees F, Velagala S, Ali J, et al. Virtual screening of immunomodulatory medicinal compounds as promising anti-SARS-COV-2 inhibitors. Future Virol. (2020). 10.2217/fvl-2020-0079. [Epub ahead of print]. [DOI] [Google Scholar]
- 253.Wahedi HM, Ahmad S, Abbasi SW. Stilbene-based natural compounds as promising drug candidates against COVID-19. J Biomol Struct Dyn. (2020) 2020:1–10. 10.1080/07391102.2020.1762743 [DOI] [PubMed] [Google Scholar]
- 254.Abdelli I, Hassani F, Bekkel Brikci S, Ghalem S. In silico study the inhibition of angiotensin converting enzyme 2 receptor of COVID-19 by Ammoides verticillata components harvested from Western Algeria. J Biomol Struct Dyn. (2020) 2020:1–14. 10.1080/07391102.2020.1763199 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 255.Murugan NA, Pandian CJ, Jeyakanthan J. Computational investigation on Andrographis paniculata phytochemicals to evaluate their potency against SARS-CoV-2 in comparison to known antiviral compounds in drug trials. J Biomol Struct Dyn. (2020) 2020:1–12. 10.1080/07391102.2020.1777901 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 256.Gyebi GA, Ogunro OB, Adegunloye AP, Ogunyemi OM, Afolabi SO. Potential inhibitors of coronavirus 3-chymotrypsin-like protease (3CLpro): an in silico screening of alkaloids and Terpenoids from African medicinal plants. J Biomol Struct Dyn. (2020) 20201–13. 10.1080/07391102.2020.1764868 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 257.Sampangi-Ramaiah MH, Vishwakarma R, Shaanker RU. Molecular docking analysis of selected natural products from plants for inhibition of SARS-CoV-2 main protease. Curr Sci. (2020) 118:1087–92. [Google Scholar]
- 258.Kumar V, Dhanjal JK, Bhargava P, Kaul A, Wang J, Zhang H, et al. Withanone and withaferin-A are predicted to interact with transmembrane protease serine 2 (TMPRSS2) and block entry of SARS-CoV-2 into cells. J Biomol Struct Dyn. (2020) 2020:1–13. 10.1080/07391102.2020.1775704 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 259.Borkotoky S, Banerjee M. A computational prediction of SARS-CoV-2 structural protein inhibitors from Azadirachta indica (Neem). J Biomol Struct Dyn. (2020) 2020:1–7. 10.1080/07391102.2020.1774419 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 260.Elfiky AA. Natural products may interfere with SARS-CoV-2 attachment to the host cell. J Biomol Struct Dyn. (2020) 2020:1–10. 10.21203/rs.3.rs-22458/v1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 261.Ahmad S, Abbasi HW, Shahid S, Gul S, Abbasi SW. Molecular docking, simulation and MM-PBSA studies of nigella sativa compounds: a computational quest to identify potential natural antiviral for COVID-19 treatment. J Biomol Struct Dyn. (2020) 2020:1–9. 10.1080/07391102.2020.1775129 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 262.Saito K. Phytochemical genomics-a new trend. Curr Opin Plant Biol. (2013) 16:373–80. 10.1016/j.pbi.2013.04.001 [DOI] [PubMed] [Google Scholar]
- 263.Mathur S, Hoskins C. Drug development: lessons from nature. Biomed Rep. (2017) 6:612–4. 10.3892/br.2017.909 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 264.Wink M. Plant secondary metabolites modulate insect behavior-steps toward addiction? Front Physiol. (2018) 9:364. 10.3389/fphys.2018.00364 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 265.Ebenezer KS, Manivannan R, Punniyamoorthy A, Tamilselvan C. Plant secondary metabolites of antiviral properties a rich medicinal source for drug discovery: a mini review. J Drug Deliv Ther. (2019) 9:161–7. 10.22270/jddt.v9i5.3471 [DOI] [Google Scholar]
- 266.Calixto JB. The role of natural products in modern drug discovery. An Acad Bras Cienc. (2019) 91:20190105. 10.1590/0001-3765201920190105 [DOI] [PubMed] [Google Scholar]
- 267.Hardman WE. Diet components can suppress inflammation and reduce cancer risk. Nutr Res Prac.t. (2014) 8:233–40. 10.4162/nrp.2014.8.3.233 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 268.Atanasov AG, Waltenberger B, Pferschy-Wenzig EM, Linder T, Wawrosch C, Uhrin P, et al. Discovery and resupply of pharmacologically active plant-derived natural products: a review. Biotechnol Adv. (2015) 33:1582–614. 10.1016/j.biotechadv.2015.08.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 269.Annunziata G, Sanduzzi Zamparelli M, Santoro C, Ciampaglia R, Stornaiuolo M, Tenore GC, et al. May polyphenols have a role against coronavirus infection? An overview of in vitro evidence. Front Med. (2020) 7:240. 10.3389/fmed.2020.00240 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 270.Chowdhury P, Sahuc M, Rouillé C, Rivière C, Bonneau N, Vandeputte A, et al. Theaflavins, polyphenols of black tea, inhibit entry of hepatitis C virus in cell culture. PLoS ONE. (2018) 13:11. 10.1371/journal.pone.0198226 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 271.Vázquez-Calvo Á, Oya N, Martín-Acebes MA, Garcia-Moruno E, Saiz J. Antiviral properties of the natural polyphenols delphinidin and epigallocatechin gallate against the flaviviruses West Nile virus, Zika virus, and dengue virus. Front Microbiol. (2017) 8:1314. 10.3389/fmicb.2017.01314 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 272.Park JY, Yuk HJ, Ryu HW, Lim SH, Kim KS, Park KH, et al. Evaluation of polyphenols from Broussonetia papyrifera as coronavirus protease inhibitors. J Enzyme Inhib Med Chem. (2017) 32:504–12. 10.1080/14756366.2016.1265519 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 273.Abdelmageed MI, Abdelmoneim AH, Mustafa MI, Elfadol NM, Murshed NS, Shantier SW, et al. Design of a multiepitope-based peptide vaccine against the E protein of human COVID-19: an immunoinformatics approach. BioMed Res Int. (2020) 2020:2683286. 10.1101/2020.02.04.934232 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 274.Ghosh R, Chakraborty A, Biswas A, Chowdhuri S. Evaluation of green tea polyphenols as novel coronavirus (SARS CoV-2) main protease (Mpro) inhibitors – an in silico docking and molecular dynamics simulation study. J Biomol Struct Dyn. (2020) 1–11. 10.1080/07391102.2020.1779818 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 275.Jo S, Kim S, Shin DH, Kim MS. Inhibition of SARS-CoV 3CL protease by flavonoids. J Enzyme Inhib Med Chem. (2020) 35:145–51. 10.1080/14756366.2019.1690480 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 276.Peterson L. In silico molecular dynamics docking of drugs to the inhibitory active site of SARS-CoV-2 protease and their predicted toxicology and ADME. SSRN Electron J. (2020). 10.26434/chemrxiv.12155523.v1 [DOI] [Google Scholar]
- 277.Owis A, Marwa EH, Dalia EA, Omar A, Usama A, Mohamed K. Molecular docking reveals the potential of Salvadora persica flavonoids to inhibit COVID-19 virus main protease. RSC Adv. (2020) 10:19570–75. 10.1039/D0RA03582C [DOI] [PMC free article] [PubMed] [Google Scholar]
- 278.Thawabteh A, Juma S, Bader M, Karaman D, Scrano L, Bufo SA, et al. The biological activity of natural alkaloids against herbivores, cancerous cells and pathogens. Toxins. (2019) 11:656–83. 10.3390/toxins11110656 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 279.Kim DE, Min JS, Jang MS, Lee JY, Shin YS, Park CM, et al. Natural bis-benzylisoquinoline alkaloids-tetrandrine, fangchinoline, and cepharanthine, inhibit human coronavirus oc43 infection of mrc-5 human lung cells. Biomolecules. (2019) 9:696. 10.3390/biom9110696 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 280.Wink M. Potential of DNA intercalating alkaloids and other plant secondary metabolites against SARS-CoV-2 causing COVID-19. Diversity. (2020) 12:175 10.3390/d12050175 [DOI] [Google Scholar]
- 281.Colson P, Rolain J, Roult D. Chloroquine for the 2019 novel coronavirus SARS-CoV-2. Int J Antimicrob Agents. (2020) 55:105923 10.1016/j.ijantimicag.2020.105923 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 282.Hussein RA, El-Anssary AA. Plants secondary metabolites: the key drivers of the pharmacological actions of medicinal plants. Herb Med. (2019) 10.5772/intechopen.76139 [DOI] [Google Scholar]
- 283.Wen CC, Kuo YH, Jan JT, Liang PH, Wang SY, Liu HG, et al. Specific plant terpenoids and lignoids possess potent antiviral activities against severe acute respiratory syndrome coronavirus. J Med Chem. (2007) 50:4087–95. 10.1021/jm070295s [DOI] [PubMed] [Google Scholar]
- 284.Álvarez DM, Castillo E, Duarte LF, Arriagada J, Corrales N, Farías MA, et al. Current antivirals and novel botanical molecules interfering with herpes simplex virus infection. Front Microbiol. (2020) 11:139. 10.3389/fmicb.2020.00139 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 285.Sun Z, Yu C, Wang W, Yu G, Zhang T, Zhang L, et al. Aloe polysaccharides inhibit Influenza A virus infection—a promising natural anti-flu drug. Front Microbiol. (2018) 9:2338. 10.3389/fmicb.2018.02338 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 286.Mera IFG, Falconí DEG, Córdova VM. Secondary metabolites in plants: main classes, phytochemical analysis and pharmacological activities. Rev Bionatura. (2019) 4:1000–9. 10.21931/RB/2019.04.04.11 [DOI] [Google Scholar]
- 287.Strovel J, Sittampalam S, Coussens NP, Hughes M, Inglese J, Kurtz A, et al. Early Drug Discovery and Development Guidelines: for Academic Researchers, Collaborators, and Start-Up Companies. Assay Guidance Manual. Bethesda, MD: Eli Lilly & Company and the National Center for Advancing Translational Sciences; (2004). p. 1–35. [PubMed] [Google Scholar]
- 288.Krüger A, Gonçalves Maltarollo V, Wrenger C, Kronenberger T. ADME profiling in drug discovery and a new path paved on silica. In: Gaitonde V, editor. Drug Discovery and Development - New Advances. Itratech open; (2020). p. 32 10.5772/intechopen.86174 [DOI] [Google Scholar]
- 289.Seca AML, Pinto DCGA. Biological potential and medical use of secondary metabolites. Medicines. (2019) 6:66. 10.3390/medicines6020066 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 290.Dias DA, Urban S, Roessner U. A historical overview of natural products in drug discovery. Metabolites. (2012) 2:303–36. 10.3390/metabo2020303 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 291.Anand U, Nandy S, Mundhra A, Das N, Pandey DK, Dey A. A review on antimicrobial botanicals, phytochemicals and natural resistance modifying agents from Apocynaceae family: possible therapeutic approaches against multidrug resistance in pathogenic microorganisms. Drug Resist Update. (2020) 51:100695. 10.1016/j.drup.2020.100695 [DOI] [PubMed] [Google Scholar]
- 292.Anwar N, Teo Y, Joash T. The role of plant metabolites in drug discovery: current challenges and future perspectives. In: Swamy MK, Akhtar MS. editors. Natural Bio-active Compounds, Volume 2: Chemistry, Pharmacology and Health Care Practices. New York, NY: Springer Publications; (2019). p. 25–51. 10.1007/978-981-13-7205-6_2 [DOI] [Google Scholar]
- 293.Gonzalez-Alfonso JL, Peñalver P, Ballesteros AO, Morales JC, Plou FJ. Effect of α-Glycosylation on the stability, antioxidant properties, toxicity, and neuroprotective activity of (-)-Epigallocatechin gallate. Front Nutr. (2019) 6:30 10.3389/fnut.2019.00030 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 294.Sharifi-Rad M, Pezzani R, Redaelli M, Zorzan M, Imran M, Khalil AA, et al. Preclinical pharmacological activities of Epigallocatechin-3-gallate in signaling pathways: an update on cancer. Molecules. (2020) 25:467. 10.3390/molecules25030467 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 295.Banerjee A, Czinn SJ, Reiter RJ, Blanchard TG. Crosstalk between endoplasmic reticulum stress and anti-viral activities: a novel therapeutic target for COVID-19. Life Sci. (2020) 255:117842. 10.1016/j.lfs.2020.117842 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 296.Karade PG, Jadhav NR. Colon targeted curcumin microspheres laden with ascorbic acid for bioavailability enhancement. J Microencapsul. (2018) 35:372–80. 10.1080/02652048.2018.1501111 [DOI] [PubMed] [Google Scholar]
- 297.Azim KF, Ahmed SR, Banik A, Khan MMR, Deb A, Somana SR. Screening and druggability analysis of some plant metabolites against SARS-CoV-2: an integrative computational approach. Inform Med Unlocked. (2020) 20:100367. 10.1016/j.imu.2020.100367 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 298.Capell T, Twyman RM, Armario NV, Ma JKC, Schillberg S, Christou P. Potential applications of plant biotechnology against SARS-CoV-2. Trends Plant Sci. (2020) 25:635–43. 10.1016/j.tplants.2020.04.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 299.Nakabayashi R, Saito K. Metabolomics for unknown plant metabolites. Anal Bioanal Chem. (2013) 405:5005–11. 10.1007/s00216-013-6869-2 [DOI] [PubMed] [Google Scholar]
- 300.Li D Halitschke R Baldwin IT Gaquerel E . Information theory tests critical predictions of plant defense theory for specialized metabolism. Sci Adv. (2020) 6:eaaz0381. 10.1126/sciadv.aaz0381 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 301.Bhuiyan FR, Campos NA, Swennen R, Carpentier S. Characterizing fruit ripening in plantain and Cavendish bananas: a proteomics approach. J Proteomics. (2020) 214:103632. 10.1016/j.jprot.2019.103632 [DOI] [PubMed] [Google Scholar]
- 302.Runfeng L, Yunlong H, Jicheng H, Weiqi P, Qinhai M, Yongxia S, et al. Lianhuaqingwen exerts anti-viral and anti-inflammatory activity against novel coronavirus (SARS-CoV-2). Pharmacol Res. (2020) 156:104761. 10.1016/j.phrs.2020.104761 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 303.Cyranoski D. China is promoting coronavirus treatments based on unproven traditional medicines. Nature. (2020). 10.1038/d41586-020-01284-x. [Epub ahead of print]. [DOI] [PubMed] [Google Scholar]
- 304.Nguyen-Vo TH, Nguyen L, Do N, Nguyen TN, Trinh K, Cao H, et al. Plant metabolite databases: from herbal medicines to modern drug discovery. J Chem Inf Model. (2020) 60:1101–10. 10.1021/acs.jcim.9b00826 [DOI] [PubMed] [Google Scholar]
- 305.Johnson SR, Lange BM. Open-access metabolomics databases for natural product research: present capabilities and future potential. Front Bioeng Biotechnol. (2015) 3:22. 10.3389/fbioe.2015.00022 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Different plant families showing antiviral properties. (Each portion of the pie chart describes a specific Family alongside its total number of plants that have antiviral properties).
List of secondary metabolites found from medicinal plants.