Abstract
Background
Mesenchymal stromal cells (MSCs) are rapidly advancing as commercial therapeutics. However, there are still no adequate tools to validate the identity of MSCs and support standardization of MSC-based products. Currently accepted metrics include cell surface marker profiling and tri-lineage differentiation assays, neither of which is definitive. Transcript profiling represents a cost- and time-effective approach amenable to MSC manufacturing processes. Two independent labs recently reported non-overlapping MSC-specific transcriptomic signatures of 489 and 16 genes.
Methods
Here, we interrogated our repository of transcriptome data to determine whether routine culture manipulations including cell expansion and immune activation affect expression of the reported MSC lineage genes. These data sets comprise 4 donor populations of human umbilical cord (UC) MSCs serially cultured from cryopreservation thaw through pre-senescence, and 3 donor populations each of naïve UC and bone marrow (BM) MSCs and licensed by 3 different cytokines.
Results
Overall, 437 of 456 proposed signature genes assessed in these data sets were reliably expressed, representing an enduring lineage profile in 96% agreement with the previous studies. Serial passaging resulted in the downregulation of 3 signature genes, and one was silenced. Cytokine stimulation downregulated expression of 16 signature genes, and 3 were uniformly silenced in one or the other MSC type. Fifteen additional genes were unreliably detected, independent of culture manipulation.
Conclusion
These results validate and refine the proposed transcriptomic tools for reliable identification of MSCs after isolation through cell expansion and after inflammatory activation. We propose a 24-gene signature to support standardized and accessible MSC characterization.
Keywords: Mesenchymal stromal cell, Transcriptomics, Cell characterization, Molecular profiling, Identity assays, Umbilical cord, Bone marrow, Licensing
Background
Mesenchymal stromal cells (MSC) exhibit phenotypic and functional heterogeneity related to tissue origin, donor demographics, and processing protocols. Current metrics to establish MSC identity include plastic adherence, cell surface phenotyping, and tri-lineage differentiation [1], which do not clearly distinguish MSCs from other stromal resident cells such as fibroblasts [2] or hepatic stellate cells [3]. Two research groups recently performed deep integrative analysis of publicly available transcriptomics data for multiple MSC types compared to other stem and stromal cells to generate MSC-specific signatures [4, 5]. Using combinatorial analysis of 285 samples from public data and in-house microarray and RNA-Seq data, Roson-Burgo et al. derived an MSC lineage signature of 489 genes based primarily on genes upregulated in the bone marrow (BM), adipose, and placental MSCs compared with hematopoietic stem and progenitor cells [4]. The “Rohart MSC test”, an in silico classifier based on 16 MSC signature genes, was created and validated using over 100 transcriptome studies employing 15 different quantification platforms [5]. This test reportedly distinguishes MSCs from non-MSCs with > 95% accuracy [5].
Manufacturing of MSC therapies often includes extended cell expansion or inflammatory licensing. Although these candidate genes have been validated for cell isolation protocols, their usefulness for in-process identity testing under the pressures of cell expansion and immune activation has not been formally investigated. We recently generated two transcriptome data sets by microarray analysis of 14,500 genes. The first set of 57 arrays comprises longitudinal culture of 4 umbilical cord-derived (UC) MSC populations analyzed at every passage (P) from P1 or P2 through pre-senescence [6]. The second set of 24 arrays comprises a matrix evaluation of 3 populations each of UC and BM-MSCs at rest and after priming with TNF-α, IFN-γ, or IL-1β (submitted manuscript). All MSC populations were expanded to comparable population doublings in xeno- and serum-free media, with donor sex balanced between groups. Here, to address the need for tools with validated utility across dynamic MSC biology and tissue and donor-influenced heterogeneity, we interrogated expression of the 2 proposed MSC signatures in these 2 substantial data sets that span culture medium formulation, tissue source, donor source, cell aging, and 3 canonical cell licensing conditions.
Methods
MSCs used to generate the data sets were validated by surface marker profiling and tri-lineage differentiation and were cultured and manipulated as previously described ([6] and submitted manuscript). Briefly, cells were maintained in TheraPEAK™ MSC growth medium chemically defined (Lonza, MD, USA) and serially propagated until senescence [6] or cultivated in xeno-free human platelet lysate (hPL)-supplemented media (RoosterBio Inc., MD, USA) and then activated by a 24-h co-incubation with TNF-α (50 ng/ml), IFN-γ (50 ng/ml), or IL-1β (80 pg/ml) in protein-free media (RoosterBio Inc.; submitted manuscript).
Microarray analysis was conducted using R Bioconductor packages [7, 8]. Gene Expression Omnibus [9, 10] data sets GSE119987 and GSE129165 were previously pre-processed using simpleaffy [11], GCRMA [12] and queried for batch effects using BatchQC [13]. For transcripts mapped by multiple probes, the probe set with highest average expression intensity across all samples was used. Pairwise comparisons of P2 versus each subsequent passage within each UC-MSC population (GSE119987) and comparisons of resting versus primed UC and BM-MSCs separately (GSE129165) were performed using eBayes in limma [14, 15]. Significantly differentially expressed probe sets had a false discovery rate-adjusted p value (p) of < 0.05 with sequential filtering for > 2-fold change.
Forty-four of the 489 Roson-Burgo MSC lineage genes [4] and 5 of 16 Rohart MSC test genes [5] are not represented on the Affymetrix GeneChip Human Genome U133A 2.0 microarray and were not analyzed.
Results
Most MSC signature genes are stable during cell expansion
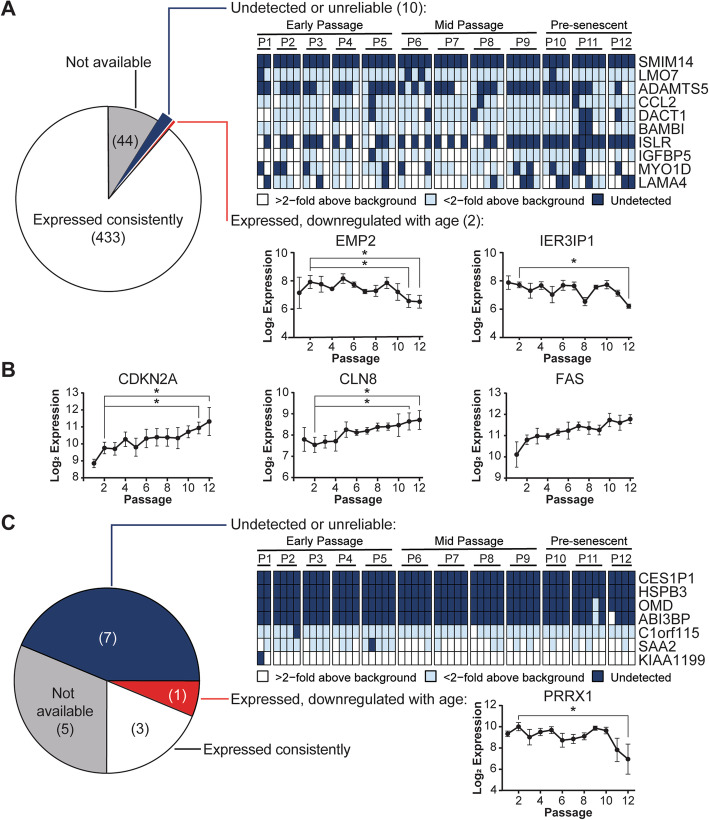
We recently reported that the transcriptomes of serially expanded UC-MSCs diverge from early passage counterparts in tiers of magnitude corresponding to early passage (P1–5), mid-passage (P6–9), and pre-senescence (P10-senescence) [6]. Intriguingly, UC-MSCs at any passage constitutively express 435 (98%) of 445 tested Roson-Burgo MSC signature genes [4], despite accumulating age-related markers during pre-senescence (Fig. 1a, b, Table 1, Additional file 1: Fig. S1A). These identity genes were consistently expressed during cell expansion, except for EMP2 and IER3IP1 which declined during pre-senescence (Fig. 1a). At P11, expression of EMP2 decreased 2.89-fold compared to P2 (p = 0.046), and 3.25-fold at P12 (p = 0.018). IER3IP1 expression also significantly decreased at P12 (− 2.91-fold, p = 0.025). However, UC-MSCs maintained moderate expression of these genes through pre-senescence. By contrast, 10 genes were unreliably expressed, with no detectable signal above microarray background in at least 1 sample (Fig. 1a). SMIM14 was never detected, while 9 others were sporadically expressed. ISLR was expressed by some early to mid-passage MSC populations but was undetected after P8.
Fig. 1.
Stability of MSC lineage profiles during prolonged UC-MSC expansion. a Most interrogated transcripts in the Roson-Burgo MSC lineage gene set (433/445) are expressed and unchanged by extended passaging. SMIM14 is never detected, while 9 other genes are expressed intermittently during cell propagation. EMP2 and IER3IP1 are downregulated with age but remain detectable. b Genes previously linked to senescence (CDKN2A) or aging (CLN8, FAS) as defined by Gene Ontology functional classification are upregulated at P10, 11, and 12 versus P2 (p < 0.05, > 1.5-fold change) [6]. c Four of the 11 tested Rohart MSC genes are reliably expressed by serially expanded UC-MSCs, although PRRX1 expression decreases in pre-senescent cells. Four genes (CES1P1, HSPB3, OMD, and ABI3BP) are essentially undetectable. C1orf115, SAA2, and KIAA1199 are expressed throughout cell expansion but are deemed unreliable since expression intensity occasionally falls below the detection limit. Graphs depict the mean ± SEM of biological replicates. Abbreviations: P, passage. *p < 0.05 and > 2-fold change
Table 1.
Summary of MSC transcriptomic lineage marker expression by MSCs during prolonged culture or licensing
| Roson-Burgo core signature | Rohart MSC test | ||
|---|---|---|---|
| Assessed | 445/489 (91%) | 11/16 (69%) | |
| Passage 2–12 UC-MSCs | Reliable | 435/445 (98%) | 4/11 (36%) |
| Resting and primed UC and BM-MSCs | Reliable | 436/445 (98%) | 5/11 (45%) |
| Overlap | Reliable | 433 | 4 (CXCL2, GDF5, MBD2, PRRX1) |
| Unreliable | 7 (ADAMTS5, DACT1, IGFBP5, ISLR, LAMA4, LMO7, SMIM14) | 6 (CES1P1, ABI3BP, C1orf115, HSPB3, OMD, SAA2) |
Transcripts in bold are uniformly silenced under specific culture conditions
Four of the 11 available Rohart MSC test genes [5], CXCL2, GDF5, MBD2, and PRRX1, were expressed in all 57 samples during UC-MSC cell expansion (Fig. 1c, Table 1, Additional file 1: Fig. S1B). Only PRRX1 expression dropped significantly, at P12 (− 9.38-fold versus P2, p = 0.040) (Fig. 1c). C1orf115, SAA2, and KIAA1199 (CEMIP) were also expressed throughout cell expansion but were each undetected at least once (Fig. 1c). CES1P1 and HSPB3 were never detected, while OMD and ABI3BP were only measurable in some pre-senescent populations (Fig. 1c).
Cytokine activation has little impact on expression of MSC lineage genes
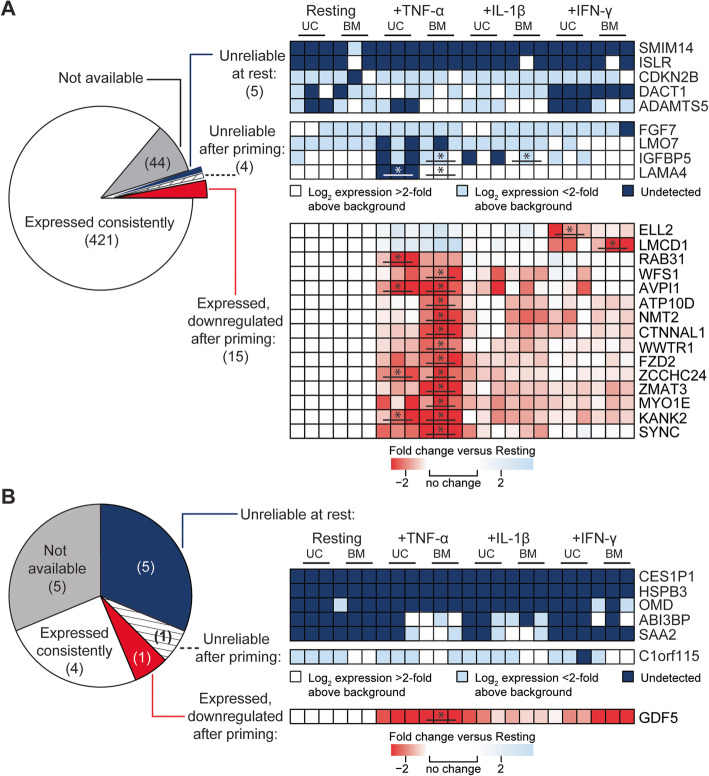
Three populations each of resting UC and BM-MSCs cultured in hPL-supplemented media (submitted manuscript) expressed 440 (99%) of 445 signature genes proposed by Roson-Burgo et al. [4] (Fig. 2a, Additional file 2: Fig. S2A). Fifteen of these 440 genes were expressed at significantly lower levels after cytokine licensing with TNF-α or IFN-γ, but were still detected (p < 0.05, > 2-fold change; Fig. 2a). Four genes were undetected in at least 1 population of licensed MSCs (Fig. 2a); LAMA4 was specifically silenced in all TNF-α primed UC-MSCs (p = 0.024).
Fig. 2.
Impact of cytokine activation on MSC lineage genes in UC and BM-MSCs. a A small cohort of Roson-Burgo MSC lineage genes is undetected before (5/445) or after (4/445) cytokine stimulation. Of these, DACT1, ADAMTS5, and LAMA4 are silenced in a cytokine-specific manner. Fifteen genes are significantly downregulated after conditioning by TNF-α or IFN-γ but remain detectable. b Four Rohart MSC test genes are consistently expressed in naïve and activated UC and BM-MSCs. In addition, GDF5 is downregulated but still measurable in all primed MSC populations. C1orf115 was also consistently expressed, except in one IFN-γ primed UC-MSC population. CES1P1 and HSPB3 were never detected, while OMD, ABI3BP, and SAA2 are only detected in some licensed populations, predominantly BM-MSCs. *p < 0.05 and > 2-fold change
ADAMTS5 and DACT1 were detected in 4 of 6 resting MSC populations and were also influenced by inflammatory activation (Fig. 2a). DACT1 expression was abrogated in all IFN-γ primed MSC populations, while TNF-α or IL-1β stimulated DACT1 expression in all MSC populations. ADAMTS5 was specifically silenced in all IFN-γ primed UC-MSCs and remained off in 2 of 3 TNF-α primed UC-MSC populations. In BM-MSCs, however, ADAMTS5 was not affected by IFN-γ priming but increased in response to TNF-α or IL-1β activation. Notably, 7 of 9 Roson-Burgo genes unreliably detected in naïve or licensed MSCs are consistent with those found unreliable during prolonged UC-MSC expansion (Table 1).
Six of the 11 available Rohart MSC test genes [5] were expressed by all 6 resting UC and BM-MSC populations cultivated in hPL (Fig. 2b, Table 1, Additional file 2: Fig. S2B), although GDF5 was significantly downregulated to moderate levels in TNF-α activated BM-MSCs (− 20.44-fold, p = 0.012). C1orf115 became undetectable in one IFN-γ conditioned UC-MSC population (Fig. 2b). Consistent with serially expanded UC-MSCs, CES1P1 and HSPB3 were undetected in all MSC samples (Fig. 2b). ABI3BP and SAA2 were also not detected in resting UC and-BM-MSCs but could be induced in a subset of donor populations responsive to TNF-α or IL-1β (Fig. 2b). These 5 genes, together with OMD, were also unreliable in serially passaged UC-MSCs (Table 1).
A panel of 24 validated MSC lineage genes may be suited for transcript-based MSC lineage assays
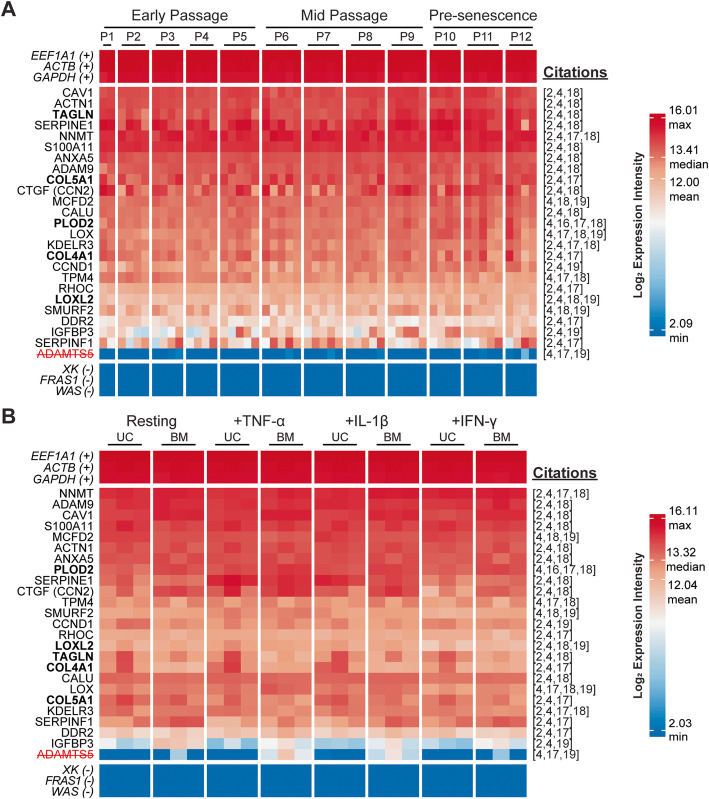
Four hundred and thirty-three Roson-Burgo signature genes and 4 Rohart MSC test genes were expressed across our 2 data sets, and together represent an enduring lineage profile (Table 1, Additional file 1: Fig. S1, Additional file 2: Fig. S2). MSC gene signatures tabulated by Roson-Burgo et al. [4] reveal that 25 of their 433 genes were previously validated in 2 or more studies [2, 16–19] (Fig. 3). Of the 4 Rohart MSC test genes reliably expressed in our data, only PRRX1 was independently cited as an MSC identity gene [17]. The panel of 25 genes was detected at moderate to high levels across our data sets, except for ADAMTS5 (Fig. 3). Of these, COL4A1, COL5A1, LOXL2, TAGLN, and PLOD2 are known to be upregulated in MSCs versus fibroblasts [2, 16]. Stability of this refined panel of 24 MSC lineage genes confirmed here and by others [2, 4, 16–19] supports its utility for standardized assays.
Fig. 3.
Candidate genes for a refined MSC lineage gene profile. Twenty-four genes previously determined to be specific to the MSC lineage are reliably expressed in both a serially expanded UC-MSCs and b cytokine-licensed UC and BM-MSCs. ADAMTS5 is undetected in multiple samples in each data set and is not suitable as an enduring marker gene. Genes in bold are upregulated in MSCs versus fibroblasts [2, 16]. Italicized genes are positive (+) and negative (−) controls, the latter of which have involvement in genetic disease
Discussion
Recent advances in MSC therapies have revealed a need for reliable and accessible MSC identity tests to support cell production [20]. Two independent groups have proposed MSC transcriptomic signature profiles that purportedly distinguish MSCs from other stromal and stem cells [4, 5], potential tools that could address this gap. Despite deep and rigorous data mining to elucidate these signatures, no genes are common to both profiles. To our knowledge, there has been no reported use of the signature proposed by Roson-Burgo et al. [4]. Literature citing Rohart et al. [5] report successful Rohart MSC test scores for either comparative qualification of cell samples [21, 22] or for examining effects of chondrogenic differentiation [23]. Thus, transcriptomic profiling is a nascent MSC classification tool. Here, we queried whether MSC gene signatures are applicable under the pressures of routine MSC manipulation including cell expansion and immune licensing by interrogating two recent transcriptome data sets encompassing these parameters.
The human UC and BM-MSCs evaluated here are commercially available as starting material for cell therapy products and meet the International Society of Cell and Gene Therapy minimal criteria for MSCs [1]. All resting MSC populations had near-perfect compliance with the comprehensive Roson-Burgo panel, whether they were cultured in chemically defined or hPL-supplemented media. This signature gene profile proved largely stable, as few genes were affected by cell aging or licensing. Seven genes, ADAMTS5, DACT1, IGFBP5, ISLR, LAMA4, LMO7, and SMIM14, proved unreliable in both data sets, and 5 others were unreliable either during cell expansion or following activation. Reliable detection of 433 signature genes across the test matrix validates their utility as MSC identity genes. The Rohart MSC test is available for datasets submitted to Stemformatics (www.stemformatics.org) [24, 25], but current wait times for curation and analysis limit its accessibility, particularly for industrialized processes. Eleven of the 16 proposed signature genes were represented in the array data. Interestingly, only 41% of these markers were reliably detected in our test matrix. Whether these 4 Rohart MSC genes can still distinguish MSCs from similar cell types, for example using the “bootsPLS” R package also developed by Rohart et al. [26], is unknown.
Our results support transcriptomics as a potential classification tool for MSCs, using lineage genes that are stably detected through pre-senescence. Cell surface markers [1] are not ideal MSC classifiers since they can be downregulated with passaging [27], and their relative abundance can vary by MSC origin and between donors [27, 28]. Tri-lineage differentiation reportedly has substantially more impact on the proposed MSC transcriptome signatures than our cytokine licensing experiments [2, 5, 23]. In this study, we identified a core panel of MSC signature genes with minimal donor and tissue origin influence that remain stable in multiple licensing conditions.
A defined transcriptomic signature would support existing minimal criteria to improve characterization of MSC master cell banks, manufactured products, and intermediates. A refined gene profile amenable to small-scale quantitative polymerase chain reactions (qPCR) is more accessible than microarray and next-generation sequencing technologies for product screening. Although the size and proprietary test framework of the Rohart MSC test set [5] is attractive for this purpose, many genes were not represented or were unexpressed in our data set. By contrast, 91% of the Roson-Burgo signature genes were available in our test matrix, and 97% of which were reliably detected. Overall, 437 (96%) of the 456 analyzed lineage genes were expressed during culture manipulation. Curation of these findings using supporting data from previously reported MSC lineage studies reveals 24 genes with potential utility as an MSC identity test for rapid, standardized, and cost-effective MSC product characterization by qPCR. This gene panel represents a substantial refinement to commercially available MSC identity tests, one of which uses markers purportedly identified in a single study of BM-MSCs and CD34+ hematopoietic precursors [29, 30]. By contrast, the panel of 24 candidate genes validated here includes 5 markers shown to be upregulated in MSCs versus fibroblasts in multiple studies and may be particularly useful to estimate fibroblast contamination.
Supplementary information
Acknowledgements
The authors are grateful to Catherine Wood for technical support and critical review of the manuscript and Tissue Regeneration Therapeutics Inc. (Toronto, Canada) for provision of human umbilical cord perivascular cells.
Abbreviations
- BM
Bone marrow
- hPL
Human platelet lysate
- MSC
Mesenchymal stromal cell
- P
Passage
- p
p value
- qPCR
Quantitative polymerase chain reaction
- UC
Umbilical cord
Authors’ contributions
DMW performed the work, analyzed the data, and wrote the manuscript. LRB designed the research and wrote the manuscript. Both authors read and approved the final manuscript.
Funding
This work was funded in part by the National Research Council of Canada Industrial Research Assistance Program Project 914919.
Availability of data and materials
The microarray data used to generate these results are available at the NCBI Gene Expression Omnibus [9, 10] through accession numbers GSE119987 and GSE129165.
Ethics approval and consent to participate
The human-sourced cells used in the study were obtained from commercial suppliers under human ethics approved protocols and provided to Aurora BioSolutions as anonymized samples for research purposes.
Consent for publication
Not applicable
Competing interests
LRB is an officer and a shareholder of the Aurora BioSolutions Inc. DMW declares no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information accompanies this paper at 10.1186/s13287-020-01873-7.
References
- 1.Dominici M, Le Blanc K, Mueller I, Slaper-Cortenbach I, Fc M, Ds K, et al. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy. 2006;8:315–317. doi: 10.1080/14653240600855905. [DOI] [PubMed] [Google Scholar]
- 2.Jääger K, Islam S, Zajac P, Linnarsson S, Neuman T. RNA-seq analysis reveals different dynamics of differentiation of human dermis- and adipose-derived stromal stem cells. PLoS One. 2012;7:e38833. doi: 10.1371/journal.pone.0038833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Chinnadurai R, Sands J, Rajan D, Liu X, Arafat D, Das R, et al. Molecular genetic and immune functional responses distinguish bone marrow mesenchymal stromal cells from hepatic stellate cells. Stem Cells. 2019;37:1075–1082. doi: 10.1002/stem.3028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Roson-Burgo B, Sanchez-Guijo F, Del Cañizo C, De Las Rivas J. Insights into the human mesenchymal stromal/stem cell identity through integrative transcriptomic profiling. BMC Genomics. 2016;17:944. doi: 10.1186/s12864-016-3230-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Rohart F, Mason EA, Matigian N, Mosbergen R, Korn O, Chen T, et al. A molecular classification of human mesenchymal stromal cells. PeerJ. 2016;4:e1845. doi: 10.7717/peerj.1845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Wiese DM, Ruttan CC, Wood CA, Ford BN, Braid LR. Accumulating transcriptome drift precedes cell aging in human umbilical cord-derived mesenchymal stromal cells serially cultured to replicative senescence. Stem Cells Transl Med. 2019;8:945–958. doi: 10.1002/sctm.18-0246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.R Core Team . R: a language and environment for statistical computing. Vienna: R Foundation for Statistical Computing; 2017. [Google Scholar]
- 8.Huber W, Carey VJ, Gentleman R, Anders S, Carlson M, Carvalho BS, et al. Orchestrating high-throughput genomic analysis with bioconductor. Nat Methods. 2015;12:115–121. doi: 10.1038/nmeth.3252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Edgar R, Domrachev M, Lash AE. Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002;30:207–210. doi: 10.1093/nar/30.1.207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, et al. NCBI GEO: archive for functional genomics data sets - update. Nucleic Acids Res. 2013;41:D991–D995. doi: 10.1093/nar/gks1193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Miller CJ. Simpleaffy: very simple high level analysis of Affymetrix data. 2019. [Google Scholar]
- 12.Wu J, Gentry RI. Wcf JMJ. Gcrma: background adjustment using sequence information. R version 2.58.0. 2019. [Google Scholar]
- 13.Manimaran S, Selby HM, Okrah K, Ruberman C, Leek JT, Quackenbush J, et al. BatchQC: interactive software for evaluating sample and batch effects in genomic data. Bioinformatics. 2016;32:3836–3838. doi: 10.1093/bioinformatics/btw538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Phipson B, Lee S, Majewski IJ, Alexander WS, Smyth GK. Robust hyperparameter estimation protects against hypervariable genes and improves power to detect differential expression. Ann Appl Stat. 2016;10:946–963. doi: 10.1214/16-AOAS920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47–e59. doi: 10.1093/nar/gkv007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Wagner W, Wein F, Seckinger A, Frankhauser M, Wirkner U, Krause U, et al. Comparative characteristics of mesenchymal stem cells from human bone marrow, adipose tissue, and umbilical cord blood. Exp Hematol. 2005;33:1402–1416. doi: 10.1016/j.exphem.2005.07.003. [DOI] [PubMed] [Google Scholar]
- 17.Pedemonte E, Benvenuto F, Casazza S, Mancardi G, Oksenberg JR, Uccelli A, et al. The molecular signature of therapeutic mesenchymal stem cells exposes the architecture of the hematopoietic stem cell niche synapse. BMC Genomics. 2007;8:65. doi: 10.1186/1471-2164-8-65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Tsai M-S, Hwang S-M, Chen K-D, Lee Y-S, Hsu L-W, Chang Y-J, et al. Functional network analysis of the transcriptomes of mesenchymal stem cells derived from amniotic fluid, amniotic membrane, cord blood, and bone marrow. Stem Cells. 2007;25:2511–2523. doi: 10.1634/stemcells.2007-0023. [DOI] [PubMed] [Google Scholar]
- 19.Kubo H, Shimizu M, Taya Y, Kawamoto T, Michida M, Kaneko E, et al. Identification of mesenchymal stem cell (MSC)-transcription factors by microarray and knockdown analyses, and signature molecule-marked MSC in bone marrow by immunohistochemistry. Genes Cells. 2009;14:407–424. doi: 10.1111/j.1365-2443.2009.01281.x. [DOI] [PubMed] [Google Scholar]
- 20.Kasoju N, Wang H, Zhang B, George J, Gao S, Triffitt JT, et al. Transcriptomics of human multipotent mesenchymal stromal cells: retrospective analysis and future prospects. Biotechnol Adv. 2017;35:407–418. doi: 10.1016/j.biotechadv.2017.04.005. [DOI] [PubMed] [Google Scholar]
- 21.Sinclair KA, Yerkovich ST, Chen T, McQualter JL, Hopkins PM-A, Wells CA, et al. Mesenchymal stromal cells are readily recoverable from lung tissue, but not the alveolar space, in healthy humans. Stem Cells. 2016;34:2548–2558. doi: 10.1002/stem.2419. [DOI] [PubMed] [Google Scholar]
- 22.Cruz T, López-Giraldo A, Noell G, Guirao A, Casas-Recasens S, Garcia T, et al. Smoking impairs the immunomodulatory capacity of lung resident mesenchymal stem cells in COPD. Am J Respir Cell Mol Biol. 2019;61:575–583. doi: 10.1165/rcmb.2018-0351OC. [DOI] [PubMed] [Google Scholar]
- 23.Larson BL, Yu SN, Park H, Estes BT, Moutos FT, Bloomquist CJ, et al. Chondrogenic, hypertrophic, and osteochondral differentiation of human mesenchymal stem cells on three-dimensionally woven scaffolds. J Tissue Eng Regen Med. 2019;13:1453–1465. doi: 10.1002/term.2899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wells CA, Mosbergen R, Korn O, Choi J, Seidenman N, Matigian NA, et al. Stemformatics: visualisation and sharing of stem cell gene expression. Stem Cell Res. 2013;10:387–395. doi: 10.1016/j.scr.2012.12.003. [DOI] [PubMed] [Google Scholar]
- 25.Choi J, Pacheco CM, Mosbergen R, Korn O, Chen T, Nagpal I, et al. Stemformatics: visualize and download curated stem cell data. Nucleic Acids Res. 2019;47(D1):D841–D846. doi: 10.1093/nar/gky1064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rohart F, Lê Cao K-A, Wells CA. bootsPLS: bootstrap subsamplings of sparse partial least squares - discriminant analysis for classification and signature identification. R package version 1.1.2. 2018. [Google Scholar]
- 27.Tomar GB, Srivastava RK, Gupta N, Barhanpurkar AP, Pote ST, Jhaveri HM, et al. Human gingiva-derived mesenchymal stem cells are superior to bone marrow-derived mesenchymal stem cells for cell therapy in regenerative medicine. Biochem Biophys Res Commun. 2010;393:377–383. doi: 10.1016/j.bbrc.2010.01.126. [DOI] [PubMed] [Google Scholar]
- 28.Najar M, Raicevic G, Fayyad-Kazan H, De Bruyn C, Bron D, Toungouz M, et al. Immune-related antigens, surface molecules and regulatory factors in human-derived mesenchymal stromal cells: the expression and impact of inflammatory priming. Stem Cell Rev Rep. 2012;8:1188–1198. doi: 10.1007/s12015-012-9408-1. [DOI] [PubMed] [Google Scholar]
- 29.STEMCELL™ Technologies Inc., Human mesenchymal stem cell qPCR array. https://www.stemcell.com/products/human-mesenchymal-stem-cell-qpcr-array.html. Accessed July 9 2020.
- 30.Silva WA, Jr, Covas DT, Panepucci RA, Proto-Siqueira R, Siufi JLC, Zanette DL, et al. The profile of gene expression of human marrow mesenchymal stem cells. Stem Cells. 2003;21:661–669. doi: 10.1634/stemcells.21-6-661. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The microarray data used to generate these results are available at the NCBI Gene Expression Omnibus [9, 10] through accession numbers GSE119987 and GSE129165.