Abstract
Melatonin is a natural hormone from the pineal gland that regulates the sleep-wake cycle. We examined the structure and physico-chemical properties of melatonin using electronic structure methods and molecular-mechanics tools. Density functional theory (DFT) was used to optimise the ground-state geometry of the molecule from frontier molecular orbitals, which were analysed using the B3LYP functional. As its electrons interacted with electromagnetic radiation, electronic excitations between different energy levels were analysed in detail using time-dependent DFT with CAM-B3LYP orbitals. The results provide a wealth of information about melatonin's electronic properties, which will enable the prediction of its bioactivity. Molecular docking studies predict the biological activity of the molecules against the coronavirus2 protein. Excellent docking scores of −7.28, −7.20, and −7.06 kcal/mol indicate that melatonin can help to defend against the viral load in vulnerable populations. Hence it can be investigated as a candidate drug for the management of COVID.
Keywords: DFT, Melatonin, Coronavirus, NLO, NBO
Highlights
-
•
Detailed quantum mechanical studies of the sleep regulating hormone melatonin
-
•
Analysed the intramolecular stabilisation and nonlinear properties
-
•
Excited state properties using TD-DFT formalism
-
•
Compound active binds to three known novel coronavirus 2019 proteins.
1. Introduction
Melatonin is a sleep-regulating hormone created by the pineal gland and is released at night [1]. It has been found to have biological activity in almost all living organisms, including plants, animals, and microbes. It can quickly enter cells through the bilipid bilayer and exhibit scavenging activity towards oxygen free radicals as well as antioxidant properties due to its low molecular weight and amphiphilic nature [2].
Peripheral tissues have been found to show a high affinity towards this hormone (melatonin), and hence act on receptors and binding sites [3,4]. Studies reported by Tan et al. show that melatonin can also be produced in the mitochondria and hence tissue melatonin levels are more than that of serum levels, hence can be used as a molecule that targets mitochondria [5]. The main physico-chemical and biological properties of melatonin are sleep-inducing effects [6,7], antioxidant behavior [[8], [9], [10], [11]], anti-inflammatory activity [[10], [11], [12], [13]], antiapoptotic effects [14], and neuroprotective [8,9,15] effects. It also regulates various physiological functions of the brain. As melatonin can diffuse quickly through the blood-brain barrier, it is effectively used in the treatment of brain injuries [7,10,16]. Rapid eye-movement sleep-behavior disorder patients are managed by a combination treatment of melatonin and clonazepam [[17], [18], [19]]. The sensing of bacteria through Toll-like receptor-4, and regulation of bacteria through altered goblet cells and antimicrobial peptides, are all involved in the anticolitic effects of melatonin in inflammatory bowel disease [20]. Melatonin is involved in the aging process, growth towards puberty, and modulation of blood pressure [21]. This versatile compound blocks proangiogenic and antiangiogenic effects caused by docetaxel and vinorelbine, which are anti-tumor drugs, and it enhances their tumor-fighting behavior [22]. This molecule can modify the redox state of the rat pancreatic stellate cell.
Melatonin is an endogenous hormone that is involved in circadian rhythm control. It is inexpensive and safe as it has a significant effect as an antioxidant and anti-inflammatory. Melatonin, chemically N-acetyl-5-methoxytryptamine, is a tryptophan derivative, has multiple physiological effects, and can be used to treat many diseases related to virus infections, especially respiratory diseases. In COVID-19 patients with digestive complications, melatonin has positive effects [23]. Melatonin can thus be used as an adjunctive or even as a regular therapy as no antiviral treatment is currently available. Electrochemical measurements of melatonin overflow demonstrate that melatonin secretion decreases with age [24]. Melatonin treatments result in the enhancement of essential oil production in Salvia species [25].
In the context of the recent COVID pandemic, melatonin can be researched as a potential molecule to control the dangerous effect of this disease. Rising patients' tolerance and decreasing the mortality in fatal virus infections would control the innate immune response and reduce inflammation during this period. Melatonin is a molecule with respective properties as it decreases the overreaction of the innate immune response and over-shoots inflammation, but also facilitates adaptive immune function [26].
Even though melatonin is a critical biomolecule, few works have been reported on the electronic structure and reactivity of this molecule except for a preliminary work reported by Turjanski and co-workers in 1999 using semi-empirical methods [27]. In this manuscript, we describe a detailed investigation into the quantum mechanical properties of melatonin, its spectral features, reactivity preferences, and the results of docking studies with three known structural protein receptors of the novel coronavirus-2. We found that melatonin docks strongly with the three proteins. We hypothesise that this compound can be used as an adjuvant medicine for the treatment of COVID-19. Also, significant rest by a person peacefully sleeping in dark surroundings will enhance the production of this hormone, which could help in the management of current patients or as a preventive measure in the vulnerable population.
2. Material and methods
The melatonin molecule was optimised using the Gaussian-09 [28] software package with the DFT-B3LYP functional and the 6-311G+ (2d, p) basis set. B3LYP is a commonly used functional and 6-311+(2d,p) basis set is medium-sized basis set with diffused functions over heavy atoms and polarization functions to bring accuracy. We performed frequency calculations to ensure that no imaginary frequency exists such that the geometry determined would correspond to a global minimum for reaching the optimised geometry. We used the same geometry for calculating frontier molecular analysis, natural bonding orbitals, and non-linear optical studies. For UV–visible spectrum simulation, we used time-dependent density functional theory (TD-DFT) with long-range corrected CAM-B3LYP functionals with 6-311G+ (2d,p) as the basis set because electronic transitions are time-dependent phenomena. TD-DFT calculations are done using the optimised geometry obtained from B3LYP/6-311G+(2d,p) simulations. The frontier molecular orbitals were viewed from the checkpoint file generated during the optimisation calculations. A wavefunction file was generated during a single point ground state calculation job, using which the subsequent analysis performed. The melatonin molecule has more than two reaction sites, for example methoxy, carbonyl–amide, and purine ring. Reaction sites of melatonin calculated using the Multiwavefunction [29,30] software, for calculating total electrostatic potential [31], average localised ionization energies [31] and non-covalent interactions [31]. As it is reported that melatonin can be used as an adjuvant therapeutic material to fight COVID-19 [32], we decided to dock the molecule with three n-CoV-2019 protein's RCSB [33] site. Melatonin is effective in critical care patients by decreasing vessel permeability, anxiety, use of sedation, and increasing the quality of sleep, which may also be beneficial to COVID-19 patients for improved clinical outcome. Melatonin especially has a high health profile. Significant data indicate that melatonin reduces virus-related diseases and will possibly also be effective in patients with COVID-19. The target proteins were downloaded, cleaned, removed alien atoms and molecules and then used for docking. The energy received from the SwissDock software [34] and the score values received from PatchDock [35], as well as the docked results collected from Bio-discovery Studio software [36] are presented.
3. Results and discussion
3.1. Geometry of melatonin
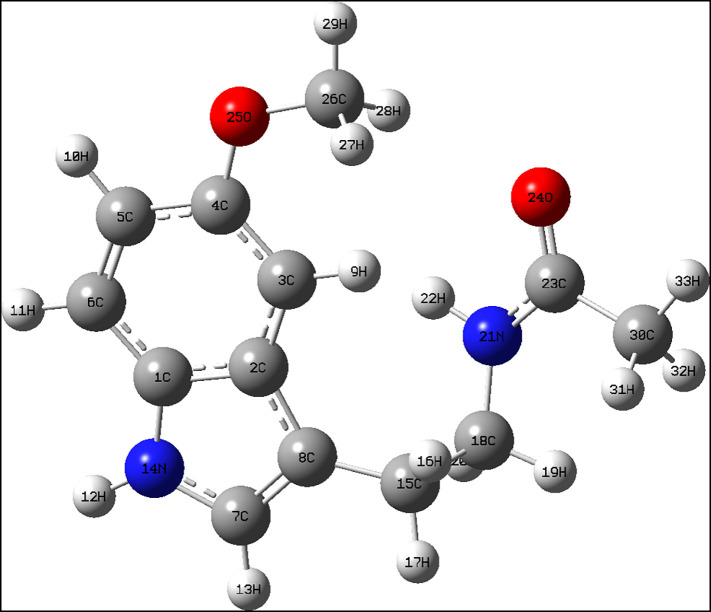
The geometry of the molecule explains its rigid structure [37]. The structure of melatonin can be explained based on its physical parameters of bond lengths and bond angles between important atoms or groups, as shown in Fig. 1 . The bond lengths of 1.3828, 1.3782 and 1.0076 Å are bonds of 1C-14N, 7C-14N, and 12H-14N, respectively, and the corresponding bond angles are109.0343°, 125.5437° and 125.4047° for 1C-14N-7C, 1C-14N-12H, and 7C-14N-12H, respectively. The bond angle of 117.8326° for 4C-25O-26C, having bond lengths of 1.3669 and 1.4227 Å for 4C—25O and 25O—26C, respectively. The bond lengths of 18C-21N, 21N-22H, 21N-23C, 23C—24O, and 23C—30C are 1.4564, 1.10147, 1.3716, 1.2284, and 1.5227 Å respectively, with the corresponding bond angles of 116.2826°, 127.4068°, 112.6290°, 121.1235°, 117.4685°, and 121.3891° for 18C-21C-22H, 18C-21C-23C, 22H-22N-23C, 21N-23C-24O, 21N-23C-30C, and 24O-23C-30C, respectively.
Fig. 1.
Geometry of melatonin.
3.2. Frontier molecular orbital (FMO) properties of melatonin
The frontier molecular orbitals are highly reactive orbitals of other molecules, and some chemical descriptors [38] are shown in Table 1 . The higher occupied molecular orbital (HOMO), lower unoccupied molecular orbital (LUMO), and energy gap of melatonin are −127.9554, −15.7755, and 112.1798 kcal/mol, respectively [39,40]. The energy gap is significant, indicating that the molecule is inherently stable. The ionization energy, electron affinity, hardness, softness, chemical potential, electronegativity, electrophilicity index, and nucleophilicity index of melatonin are 127.9554, 15.7755, 55.9989, 7031.7032, 71.8624, −71.8624, 46.1094, and 8539.8661 kcal/mol, respectively. Interaction of the melatonin with the biological target can be explained by the softness value. The softness value is high (7031.7032 kcal/mol), indicating that the molecule can positively interact with biological systems and show the desired effect [41].
Table 1.
Frontier molecular orbital properties for melatonin.
| Chemical descriptors | Energy in kcal/mol |
|---|---|
| HOMO | −127.9554 |
| LUMO | −15.7755 |
| Ionization energy (I = ɛHOMO = −HOMO) | 127.9554 |
| Electron affinity (A = ɛLUMO = −LUMO) | 15.7755 |
| Energy gap = HOMO − LUMO | 112.1798 |
| Global hardness (η = (I − A) / 2) | 55.9989 |
| Global softness (S = 1 / η) | 7031.7032 |
| Chemical potential (μ = (I + A) / 2) | 71.8624 |
| Electronegativity (χ = −μ) | −71.8624 |
| Electrophilicity index (ω = μ2 / 2η) | 46.1094 |
| Nucleophilicity index (N = 1 / ω) | 8539.8611 |
3.3. Electron transition study and excited-state properties of melatonin in solution
The electron transition study explains electron-transfer excited states. We used the TD–DFT formalism using CAM-B3LYP functionals and 6-311G+ (2d,p) basis sets in an implicit solvation atmosphere of methanol using the IEFPCM model. As transmission occurs, some energy is also emitted. Melatonin electron transitions to HOMO having a pyrrole ring and oxygen (in methoxy), HOMO-1 over the pyrrole ring and ethyl carbons, and HOMO-2 over acetamide oxygen, nitrogen, and ethyl carbons with energies of −7.04, −7.23, and −8.79 eV, respectively. Melatonin electron transitions to LUMO which is over the pyrrole ring, LUMO+, which is over the pyrrole ring, oxygen (in methoxy), ethyl carbons, and acetamide nitrogen and carbon, and LUMO+2 having acetamide carbons, acetamide nitrogen, and carbons with energies of 0.37, 0.74, and 0.97 eV, respectively. The electronic spectral data using TD–DFT simulations indicate a significant λmax of 256.07 nm in a methanol solvent. The transitions are due to the movement of electrons from HOMO-1 to LUMO (69%) and HOMO to LUMO (18%) with an oscillator strength of 0.1639. The electronic transitions are due to charge transfer transitions from one region of the molecule to another, which indicates its inherent stability due to electronic excitations.
3.4. Non-linear optical behavior of melatonin
Scientists and technologists working in the molecular electronics field are continuously searching for compounds with substantial non-linear optical (NLO) activity. Such compounds find immense application in electronic displays, surveillance equipment, and consumer electronic gadgets. Computationally, the ability of a molecule to act as an NLO material can be determined from the polarizability and hyperpolarizability data [[42], [43], [44], [45]]. The NLO properties of melatonin are shown in Table 2 . This is an essential behavior of melatonin that has a light absorption nature, movement of electrons or protons, as compared with a standard NLO material such as urea [46]. The dipole moment of melatonin is 4.4961 D, which is 1.4750 times greater than urea. Hyperpolarizability [47], mean polarizability, and anisotropy of the polarizability of melatonin are 544.5474, 180.1312, and 349.4465 esu, and which are 9.6495, 5.0524, and 5.0588 times greater than urea, respectively. The compound is not centrosymmetric, hence generates second-order spherical harmonics and beta hyperpolarizability functions. This compound can hence be used as an organic non-linear optically active substance in organic electronic appliances.
Table 2.
Non-linear optics property for melatonin.
| Non-linear property | Melatonin | Urea | Comparison of melatonin with urea |
|---|---|---|---|
| Dipole moment (μ) | 4.4961 D | 3.0482 D | 1.4750 times greater than urea |
| Hyperpolarizability (β) | 544.5474 esu | 56.4324 esu | 9.6495 times greater than urea |
| Mean polarizability (α0) | 180.1312 esu | 35.6521 esu | 5.0524 times greater than urea |
| Anisotropy of the polarizability (Δα) | 349.4465 esu | 69.0764 esu | 5.0588 times greater than urea |
3.5. Nature of NBO study of melatonin
A molecule, especially one with profound biological activity, may have many intramolecular electron delocalisation and hyperconjugative stabilisation regions. Natural bond orbital analysis, which is a quantum mechanical method, is useful for this type of study. The molecular orbital properties of melatonin for the occupancy of the natural orbitals were performed by the NBO suite [43] embedded in the Gaussian software.
From donor-bonding orbital σ (C1-C2) with occupancy is 1.9578 to acceptor anti-bonding orbitals σ* (C3-C4), σ* (C5-C6), and σ* (C7-C8) exhibiting the transition, the energies are 17.15, 15.53, and 18.83 kcal/mol, respectively. From σ (C3-C4) with occupancy is 1.9684 to σ* (C5-C6), and the Rydberg orbital R* (C30) with the energies are 18.09 and 21.48 kcal/mol, respectively. From σ (C5-C6) having an occupancy is 1.9735 to σ* (C3-C4), R* (30), and R* (H33) with the energies are 18.04, 10.49, and 13.46 kcal/mol respectively, from σ (N21-C23) has occupancy 1.9915 to Rydberg orbital R* (C30) and R* (H33) with the energies are 11.05 and 30.53 kcal/mol, respectively, from σ (C23-O24) having the occupancy is 1.9932 to σ* (C1-C2), σ* (C2-C3), R* (C23) and R* (H33) with the energies are 41.93, 23.39, 78.00, and 193.98 kcal/mol, respectively, from σ (C23-C30) has occupancy is 1.9858 to R* (H17), R* (C18), R* (H19), R* (C23), R* (O24), R* (C30), R* (H33), σ* (C1-C2), σ* (C1-C6), σ* (C2-C3), σ* (C26-H27), σ* (C30-H31), σ* (C30-H32), and σ* (C30-H33) having the energies are 28.16, 21.27, 32.72, 87.90, 38.44, 425.33, 813.65, 497.63, 11.50, 136.64, 11.32, 41.85, 17.14, and 22.64 kcal/mol, respectively, from σ (O25-C26) having occupancy, is 1.9922 to R* (C30) and σ* (C1-C2) with the energies are 23.21 and 17.18 kcal/mol, respectively, from σ (C26-H27) to σ* (C1-C2) having the energy 12.48 kcal/mol with the occupancy is 1.9940, from σ (C30-H31) with the occupancy is 1.9763 to R* (H17), R* (C18), R* (H19), R* (C23), R* (O24), R* (C30), R* (H33), σ* (C1-C2), σ* (C2-C3), σ* (C23-O24), σ* (C30-H31), and σ* (C30-H32) having the energies are 19.45, 15.35, 21.77, 59.71, 25.54, 343.96, 489.68, 774.25, 102.18, 15.87, 27.07, and 24.50 kcal/mol respectively, from σ (C30-H32) with occupancy is 1.9781 to R* (H17), R* (C18), R* (H19), R* (C23), R* (O24), R* (C30), R* (H33), σ* (C1-C2), σ* (C2-C3), σ* (C30-H32), and σ* (C30-H33) having the energies are 17.26, 13.74, 21.60, 54.83, 20.96, 333.66, 470.61, 779.22, 99.22, 27.01, and 17.77 kcal/mol, respectively, and from σ (C30-H33) with the occupancy is 1.9877 to R* (H17), R* (C18), R* (C19), R* (C23), R* (C24), R* (O25), R* (C26), R* (H28), R* (C30), R* (H33), σ* (C1-C2), σ* (C1-C6), σ* (C2-C3), σ* (C2-C4), σ* (C18-H19), σ* (C23-O24), σ* (C26-H27), σ* (C30-C31), σ* (C30-H32), and σ* (C30-H33) having the energies are 53.21, 44.19, 62.43, 168.90, 69.68, 10.22, 20.79, 25.37, 1071.78, 1521.31, 28.56, 19.70, 311.41, 10.87, 10.40, 10.54, 24.34, 59.42, 64.53, and 31.23 kcal/mol, respectively.
From core bonding orbital C (C23), which has the occupancy 1.9994 electrons move to anti-bonding R* (C23), R* (C30), and R* (H33) with the transition energies are 111.01, 25.20, and 254.09 kcal/mol, respectively, from C (C23) with the occupancy is 19,994 to R* (H33), σ* (C2-C8), and σ* (C26-H28) having the energies are 165.30, 100.33, and 100.05 kcal/mol, respectively, from C (O24) to σ* (C26-H29) has the energy is 104.46 kcal/mol with occupancy is 1.9997, and from C (C30) having the occupancy is 1.9992 to R* (H17), R* (C23), R* (C30), R* (H33), σ* (C2-C8), σ* (C26-H28), σ* (C30-H31), and σ* (C30-H32) with the energies are 69.05, 1429.01, 485.11, 2218.86, 1038.03, 1094.80, 255.02, and 368.22 kcal/mol, respectively. From lone pair orbital n (N14) with the occupancy is 1.6360 to σ* (C7-C8) with the energy 34.96 kcal/mol, from n (N21) has the occupancy is 1.7134 to σ* (C23-O24) having the energy 48.68 kcal/mol, and from n (O24) having the occupancy is 1.9771 to R* (C23), R* (C30), R* (H33), σ* (C1-C2), and σ* (C2-C3) with the energies are 45.23, 251.23, 124.44, 180.36, and 80.00 kcal/mol, respectively. From anti-bonding orbital σ* (C1-C2) having the occupancy 0.4953 to R* (C30) and σ* (C2-C3) with the energies are 349.61 and 107.48 kcal/mol, respectively, from σ* (C5-C6) has occupancy is 0.3175 to R* (C30) with the energy is 108.84 kcal/mol, and from σ* (C23-O24) has occupancy is 0.2792 to R* (C30) with the energy is 78.70 kcal/mol. The inherent stabilisation of the molecule is evident from the series of hyperconjugative interactions presented above. These interactions can also be between the melatonin and the surrounding solvent molecules, which reveals its stabilisation in biological medium and also between the molecule and the target proteins used in the docking.
3.6. Total electrostatic potentials (ESP) and average localised ionization energy (ALIE) of melatonin
The electrostatic potential [48,49] explains how reactive sites can undergo nucleophilic or electrophilic addition or substitution reactions of melatonin, shown in Fig. 2 within 11.00 Bohr [3], and a color change from blue to red indicates charges on elements from −0.100 to 0.100. The blue color appears in both methoxy-oxygen and acetamide-oxygen; these are electron-rich sites, and electrophiles can quickly attack them. The red color appears on all of the hydrogens; these are electron-poor sites, and nucleophiles can quickly attack them.
Fig. 2.
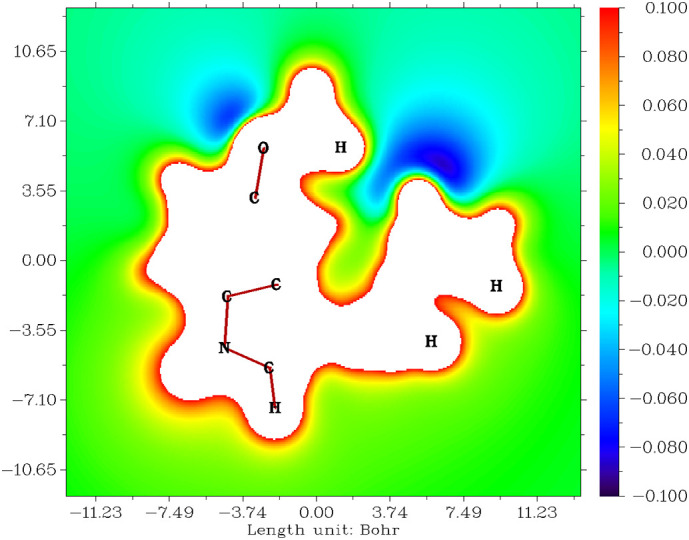
Electrostatic potentials of melatonin.
The ALIE clarifies the stability of any molecule based on saturated and unsaturated bond electron movements which are localised or delocalised [48]. The number of the resonance structure is proportional to the stability of the molecule. The ALIE of melatonin shown in Fig. 3 is within the range of ±11.00 Bohr, color is from indigo to red, and the numerical value is from 0.000 to 2.000. The blue color of protons in the methoxy group, three protons in the six-membered ring in the indole group, methyl protons, and both adjacent carbons in the acetamide group are all sites that act as electrophiles. The red color of acetamide‑carbon, conjugated carbon with oxygen atoms, and both acetamide-amide and methoxy groups are all sites that act as nucleophiles. These blue and red regions represent saturated bonds. The bluish-green regions are on indole rings to methoxy carbons via oxygen and acetamide chains. This indicates that delocalised electrons and unsaturated bonds lead to several resonance structures and explains the stability of melatonin. The electrophilic and nucleophilic reactive centres identified above interact with the COVID virus proteins and provide various electrostatic and non-covalent interactions and increases drug affinity.
Fig. 3.
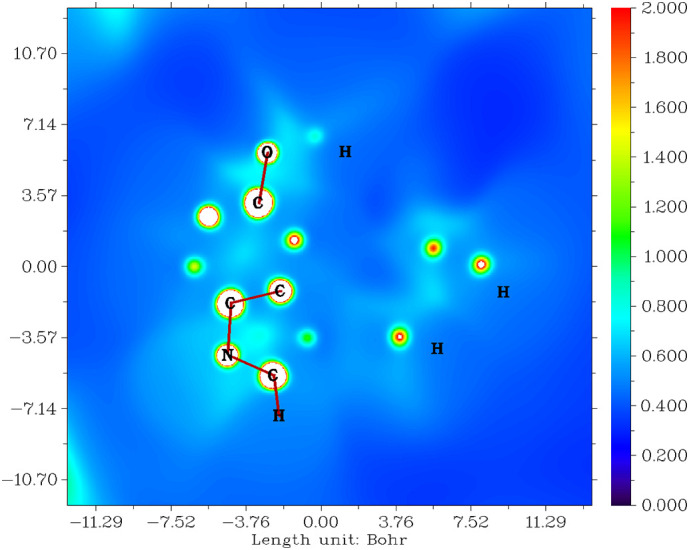
Average localised ionization energy for melatonin.
3.7. Non-covalent interaction (NCI) properties of melatonin
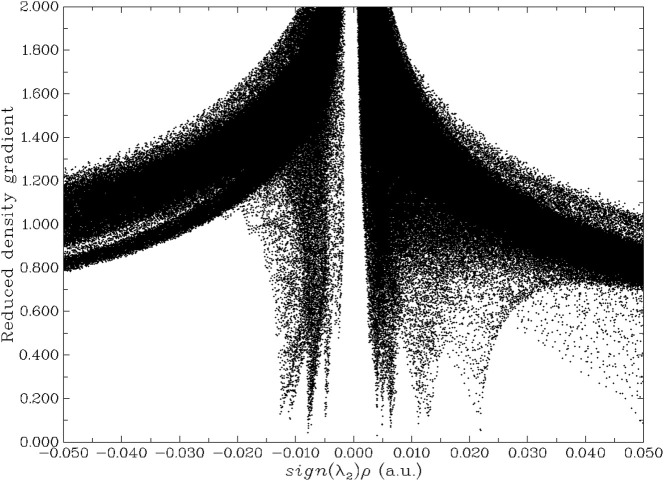
Non-covalent interactions are a valuable biological property of molecules and are non-bonded directly, but are bound by some forces such as hydrogen bonding, van der Waals bonding, and/or steric constraints. Non-covalent interactions of melatonin are shown in Fig. 4 plotted as a graph with energy plotted versus a reduced density gradient [50]. Hydrogen bonds appear from −0.005 to −0.015 a.u. between oxygen and protons from the acetamide group. The Van der Waals force ranges from −0.005 to 0.004 a.u. between acetamide-oxygen and its adjacent protons in both methyl and methoxy groups. The steric force ranges from 0.004 to 0.023 a.u. between the indole ring, the methyl group, and the carbonyl-amide group. Non-covalent interactions are a group of interactions like hydrogen bond, pi-stacking, hydrophobic interactions, van der Waal's forces, ion-dipole interactions and dipole-dipole interactions responsible for the stabilisation of the molecule and the docking between melatonin and the COVID proteins.
Fig. 4.
Non-covalent interactions of melatonin.
3.8. Molecular docking
Molecular docking is one of the essential functions of biologically active molecules. This is the theoretical evidence to design the structure and reactivity relationship of a molecule. At present, the COVID-19 pandemic caused by a new strain of the coronavirus is creating havoc throughout the world. We made efforts to dock the melatonin with the three proteins isolated from the virus, represented through the PDB ID: 6LU7, 6M03, and 6W63 were deposited in the database as mentioned in the methodology section.
With the rapid spread of the novel coronavirus globally, the design of vaccines is of great importance. SARS-CoV-2 is an enveloped, non-segmented and single stranded, positive sense RNA virus. The best drug target among coronaviruses is the main protease, Mpro, also called 3CL protease [51,52]. This is a key coronavirus enzyme, and plays a vital role in mediating viral replication and transcription. It is identified as having a mechanism-based inhibitor [53,54].
The main targeting protease protein (PDB 6LU7) is widely studied. A series of frontier molecular orbital based interaction analyses were performed on the complex between the main protease of COVID-19 and the peptide-like inhibitor whose fundamental structure was obtained from the protein (PDB 6LU7) [55]. Another targeted protease protein, in an apo form (PDB 6M03), shows the most stable form after binding with the selected drug, Threonine 111 residue with the help of several covalent bond interaction with a −6 kcal/mol docking affinity [56]. Lasinavir, Brecanavir, Telinavir, Rotigaptide, 1,3-Bis-(2-ethoxycarbonylchromon-5-yloxy)-2-(lysyloxy)propane, and Pimelautide can be considered as the main protease inhibitors of COVID-19 by docking them to the binding cavities of apo (PDB 6M03) and holo (PDB 6LU7). Another protease protein (PDB 6W63) [57] is a reversible inhibitor. The flavonoid narcissoside is reported to have a high affinity towards the protease protein (PDB 6W63) according to molecular docking studies. Thus these three protease proteins (PDB 6LU7, PDB 6M03, and PDB 6W63) can be included in the category of non-structural proteins in the structure of SARS-CoV-2 [58,59].
From Table 3 , the result from SwissDock explains the biological activity of melatonin with coronavirus proteins (PDB ID: 6LU7, 6M03, and 6W63). In general, the total ΔG is more than −5.00 kcal/mol is right active. Luckily melatonin has a total ΔG of −7.28, −7.20, and −7.06 kcal/mol with coronavirus2 proteins PDB ID: 6LU7, 6M03, and 6W63, respectively, and the total ΔG is directly proportional to the full fitness energy values which are −1219.44, −1194.87, and 1179.13 kcal/mol, respectively. It can also be seen from this table the: inter-full fitness, intra-full fitness, full solvent fitness, full surface fitness, ΔG complex polar solvent, ΔG complex nonpolar solvent, ΔG protein polar solvent, ΔG protein nonpolar solvent, ΔG ligand polar solvent, ΔG ligand nonpolar solvent, ΔG Van der Waals force, and ΔG electric force relationships between melatonin and coronavirus2 proteins, as referred.
Table 3.
| 6LU7 | 6M03 | 6W63 | |
|---|---|---|---|
| Energy | −13.8144 kcal/mol | −12.942 kcal/mol | −13.0971 kcal/mol |
| Simple fitness | −13.8144 kcal/mol | −12.942 kcal/mol | −13.0971 kcal/mol |
| Full fitness | −1219.4406 kcal/mol | −1194.8722 kcal/mol | −1179.1298 kcal/mol |
| Inter full fitness | −39.8859 kcal/mol | −36.4921 kcal/mol | −35.9845 kcal/mol |
| Intra full fitness | 2.02143 kcal/mol | 2.2279 kcal/mol | 3.07976 kcal/mol |
| Solvent full fitness | −1401.02 kcal/mol | −1380.43 kcal/mol | −1366.94 kcal/mol |
| Surface full fitness | 219.444 kcal/mol | 219.822 kcal/mol | 220.715 kcal/mol |
| Extra full fitness | 0 kcal/mol | 0 kcal/mol | 0 kcal/mol |
| ΔG complex solvent polar | −1401.02 kcal/mol | −1380.43 kcal/mol | −1366.94 kcal/mol |
| ΔG complex solvent nonpolar | 219.444 kcal/mol | 219.822 kcal/mol | 220.715 kcal/mol |
| ΔG protein solvent polar | −1411.41 kcal/mol | −1385.67 kcal/mol | −1372.14 kcal/mol |
| ΔG protein solvent nonpolar | 221.095 kcal/mol | 221.3 kcal/mol | 222.123 kcal/mol |
| ΔG ligand solvent polar | −9.24543 kcal/mol | −10.2561 kcal/mol | −11.0251 kcal/mol |
| ΔG ligand solvent nonpolar | 5.98505 kcal/mol | 6.02087 kcal/mol | 5.53225 kcal/mol |
| ΔG van der Waals force | −39.8859 kcal/mol | −36.4921 kcal/mol | −35.9845 kcal/mol |
| ΔG electric force | 0 kcal/mol | 0 kcal/mol | 0 kcal/mol |
| Total ΔG | −7.283887 kcal/mol 4 | −7.2031846 kcal/mol | −7.057701 kcal/mol |
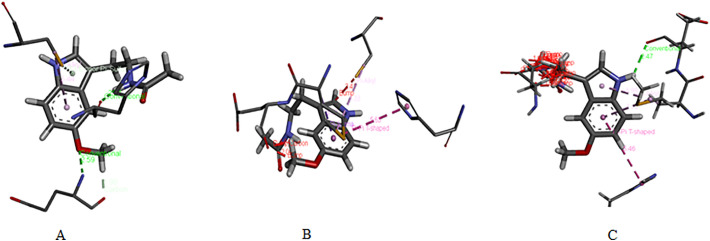
The result from PatchDock are as follows: the score values are 3772, 3730, and 3588, total surface interacting area; 402.10, 416.60, and 427.80 Å [2]; and the minimum atomic contact energies are −150.61, −157.56, and −194.63 kcal/mol for melatonin with coronavirus2 proteins PDB ID: 6LU7, 6M03, and 6W63, respectively. Figs. 5 and S1 show the skeletal structure and protein residue interactions between melatonin and coronavirus2 protein PDB ID: 6LU7, 6M03, and 6W63. Table 4 explains what protein residues are interacting with melatonin, and details the residue names, labels, hydrophobic values, pKa values, average isotropic displacements, secondary structures, residue solvent accessibility, sidechain solvent accessibility, percent solvent accessibility, and percent sidechain solvent accessibility values of coronavirus2 proteins.
Fig. 5.
Skeletal structure of interactions between melatonin and 6LU7 (A), 6M03 (B), and 6W63 (C) coronavirus2 protein residues.
Table 4.
Interactions between melatonin and coronavirus2 protein residues.
| PDB IDs | Name | Label | Hydrophobicity | pKa | Avg. isotropic displacement (Å)2 | Secondary structure | Residue solvent accessibility (Å)2 | Sidechain solvent accessibility (Å)2 | Percent solvent accessibility (Å)2 | Percent sidechain solvent accessibility (Å)2 |
|---|---|---|---|---|---|---|---|---|---|---|
| 6LU7 | Histidine | A:His41 | −3.2 | 6 | 38.866 | Helix | 34.075 | 21.85 | 19.464 | 17.602 |
| Methionine | A:Met49 | 1.9 | – | 61.459 | Turn | 69.279 | 50.785 | 38.073 | 39.363 | |
| Tyrosine | A:Tyr54 | −1.3 | 10 | 37.275 | Helix | 177.112 | 58.584 | 80.665 | 34.767 | |
| Leucine | A:Leu141 | 3.8 | – | 41.296 | Coil | 146.696 | 129.317 | 96.704 | 130.127 | |
| Asparagine | A:Asn142 | −3.5 | – | 45.276 | Turn | 167.056 | 128.991 | 113.179 | 133.44 | |
| Glycine | A:Gly143 | −0.4 | – | 38.475 | Turn | 60.921 | 15.095 | 85.049 | 62.827 | |
| Serine | A:Ser144 | −0.8 | – | 31.77 | Coil | 35.79 | 16.07 | 31.539 | 26.549 | |
| Cysteine | A:Cys145 | 2.5 | 9 | 32.06 | Turn | 26.583 | 21.399 | 20.472 | 27.646 | |
| Histidine | A:His164 | −3.2 | 6 | 30.766 | Sheet | 57.755 | 46.916 | 32.991 | 37.796 | |
| Methionine | A:Met165 | 1.9 | – | 37.842 | Sheet | 34.047 | 29.015 | 18.711 | 22.49 | |
| Glutamic acid | A:Glu166 | −3.5 | 4.3 | 36.048 | Sheet | 122.751 | 89.056 | 69.599 | 72.155 | |
| Histidine | A:His172 | −3.2 | 6 | 30.017 | Sheet | 116.35 | 52.725 | 66.461 | 42.475 | |
| Aspartic acid | A:Asp187 | −3.5 | 3.9 | 36.639 | Coil | 47.507 | 41.597 | 32.569 | 43.935 | |
| Arginine | A:Arg188 | −4.5 | 12 | 50.478 | Coil | 85.675 | 77.361 | 37.35 | 43.846 | |
| Glutamine | A:Gln189 | −3.5 | – | 43.289 | Coil | 134.002 | 110.464 | 75.154 | 88.119 | |
| 6M03 | Histidine | A:His41 | −3.2 | 6 | 44.353 | Helix | 90.166 | 20.405 | 51.505 | 16.438 |
| Methionine | A:Met49 | 1.9 | – | 58.974 | Helix | 123.264 | 51.423 | 67.741 | 39.858 | |
| Phenylalanine | A:Phe140 | 2.8 | – | 37.895 | Coil | 117.687 | 68.935 | 58.843 | 46.284 | |
| Leucine | A:Leu141 | 3.8 | – | 44.94 | Coil | 122.991 | 116.737 | 81.078 | 117.47 | |
| Asparagine | A:Asn142 | −3.5 | – | 54.727 | Turn | 157.144 | 128.895 | 106.464 | 133.34 | |
| Glycine | A:Gly143 | −0.4 | – | 42.9 | Turn | 45.267 | 23.146 | 63.196 | 96.335 | |
| Serine | A:Ser144 | −0.8 | – | 36.803 | Coil | 21.118 | 6.917 | 18.61 | 11.427 | |
| Cysteine | A:Cys145 | 2.5 | 9 | 38.353 | Turn | 31.983 | 26.799 | 24.631 | 34.622 | |
| Histidine | A:His163 | −3.2 | 6 | 33.44 | Sheet | 20.538 | 6.9 | 11.732 | 5.558 | |
| Histidine | A:His164 | −3.2 | 6 | 33.951 | Sheet | 61.269 | 57.028 | 34.998 | 45.942 | |
| Methionine | A:Met165 | 1.9 | – | 57.177 | Sheet | 20.059 | 18.046 | 11.024 | 13.988 | |
| Glutamic acid | A:Glu166 | −3.5 | 4.3 | 50.616 | Sheet | 99.525 | 74.28 | 56.43 | 60.184 | |
| Leucine | A:Leu167 | 3.8 | – | 52.523 | Coil | 45.031 | 40.757 | 29.685 | 41.013 | |
| Proline | A:Pro168 | −1.6 | – | 65.724 | Turn | 122.6 | 53.337 | 96.378 | 56.913 | |
| Histidine | A:His172 | −3.2 | 6 | 38.068 | Sheet | 77.66 | 41.72 | 44.361 | 33.609 | |
| Glutamine | A:Gln189 | −3.5 | – | 61.694 | Coil | 130.781 | 102.85 | 73.347 | 82.045 | |
| 6W63 | Histidine | A:His41 | −3.2 | 6 | 24.23 | Helix | 109.161 | 50.29 | 62.354 | 40.513 |
| Cysteine | A:Cys44 | 2.5 | 9 | 45.568 | Coil | 60.751 | 12.814 | 46.786 | 16.554 | |
| Methionine | A:Met49 | 1.9 | – | 66.152 | Coil | 82.928 | 78.088 | 45.574 | 60.526 | |
| Leucine | A:Leu50 | 3.8 | – | 65.366 | Coil | 113.773 | 105.164 | 75.001 | 105.82 | |
| Proline | A:Pro52 | −1.6 | – | 45.346 | Coil | 39.512 | 5.032 | 31.061 | 5.369 | |
| Tyrosine | A:Tyr54 | −1.3 | 10 | 30.317 | Helix | 185.906 | 70.381 | 84.67 | 41.768 | |
| Histidine | A:His164 | −3.2 | 6 | 20.287 | Sheet | 125.035 | 91.897 | 71.422 | 74.032 | |
| Methionine | A:Met165 | 1.9 | – | 32.264 | Sheet | 45.811 | 24.487 | 25.176 | 18.979 | |
| Glutamic acid | A:Glu166 | −3.5 | 4.3 | 28.999 | Sheet | 171.193 | 136.411 | 97.065 | 110.52 | |
| Leucine | A:Leu167 | 3.8 | – | 34.562 | Coil | 66.89 | 48.305 | 44.095 | 48.608 | |
| Proline | A:Pro168 | −1.6 | – | 66.174 | Turn | 165.818 | 82.018 | 130.351 | 87.517 | |
| Aspartic acid | A:Asp187 | −3.5 | 3.9 | 23.642 | Coil | 45.934 | 39.584 | 31.491 | 41.809 | |
| Arginine | A:Arg188 | −4.5 | 12 | 45.901 | Coil | 103.831 | 91.196 | 45.265 | 51.687 | |
| Glutamine | A:Gln189 | −3.5 | – | 41.43 | Coil | 115.264 | 100.696 | 64.644 | 80.327 | |
| Threonine | A:Thr190 | −0.7 | – | 46.264 | Coil | 63.345 | 43.578 | 47.773 | 50.505 | |
| Glutamine | A:Gln192 | −3.5 | – | 40.842 | Coil | 75.091 | 18.832 | 42.114 | 15.023 |
Table 3 and Fig. S2 show the residue structure of the favorable non-bond interactions between melatonin and coronavirus2 proteins. Table 5 lists favorable non-bond interactions of 6LU7 having conventional hydrogen bonds, carbon-hydrogen bonds, pi-donor hydrogen bonds, pi-sulfur, and pi-alkyl with melatonin. 6M03 has pi-sigma, pi-pi T-shaped, and pi-alkyl with melatonin, while 6W63 has pi-pi T-shaped, pi-alkyl, and pi-alkyl with melatonin along with the bond distance from chemistry. Fig. S2 and Table 6 show the residue structure of the unfavorable non-bond interactions between melatonin and coronavirus2 proteins. Protein 6LU7 does not have any unfavorable/steric interactions, protein 6M03 having three unfavorable non-bond interactions, and protein 6W63 having 33 unfavorable bump/non-bond interactions. Fig. S2 and Table 7 show unsatisfied bonds within melatonin interacting with coronavirus proteins. When interacting, protein 6LU7 has one hydrogen donor and one oxygen acceptor, protein 6M03 has two hydrogen donors and two oxygen acceptors, and protein 6W63 has one hydrogen donor and two oxygen acceptors with melatonin [60,61].
Table 5.
Favorable non-bond interactions between melatonin and coronavirus2 proteins.
| PDB IDs | Distance (Å) | Category | Type | From | From chemistry | To | To chemistry |
|---|---|---|---|---|---|---|---|
| 6LU7 | 2.59208 | Hydrogen bond | Conventional hydrogen bond | A:GLU166:N | H-donor | :UNK0:O | H-acceptor |
| 2.90848 | Hydrogen bond | Conventional hydrogen bond | :UNK0:H | H-donor | A:HIS164:O | H-acceptor | |
| 1.89804 | Hydrogen bond | Carbon hydrogen bond | :UNK0:H | H-donor | A:GLU166:O | H-acceptor | |
| 4.14288 | Hydrogen bond; other | Pi-donor hydrogen bond; Pi-sulfur | A:CYS145:SG | H-donor; sulfur | :UNK0 | Pi-orbitals; Pi-orbitals | |
| 5.43636 | Hydrophobic | Pi-alkyl | :UNK0 | Pi-orbitals | A:CYS145 | Alkyl | |
| 6M03 | 3.92902 | Hydrophobic | Pi-sigma | A:MET165:CA | C-H | :UNK0 | Pi-orbitals |
| 5.63023 | Hydrophobic | Pi-Pi T-shaped | A:HIS41 | Pi-orbitals | :UNK0 | Pi-orbitals | |
| 5.32694 | Hydrophobic | Pi-alkyl | :UNK0 | Pi-orbitals | A:CYS145 | Alkyl | |
| 6W63 | 2.47167 | Hydrogen bond | Conventional hydrogen bond | :UNK0:H | H-donor | A:GLU166:O | H-acceptor |
| 5.46262 | Hydrophobic | Pi-Pi T-shaped | A:HIS41 | Pi-orbitals | :UNK0 | Pi-orbitals | |
| 4.3243 | Hydrophobic | Pi-alkyl | :UNK0 | Pi-orbitals | A:MET165 | Alkyl | |
| 3.98009 | Hydrophobic | Pi-alkyl | :UNK0 | Pi-orbitals | A:MET165 | Alkyl |
Table 6.
Unfavorable non-bond between melatonin and coronavirus2 proteins.
| PDB IDs | Distance (Å) | Category | Type | From | From chemistry | To | To chemistry |
|---|---|---|---|---|---|---|---|
| 6LU7 | Nil | ||||||
| 6M03 | 2.41199 | Unfavorable | Unfavorable bump | A:CYS145:SG | Steric | :UNK0:C | Steric |
| 1.99847 | Unfavorable | Unfavorable bump | A:GLU166:O | Steric | :UNK0:C | Steric | |
| 1.63812 | Unfavorable | Unfavorable bump; carbon hydrogen bond | :UNK0:H | Steric; H-donor | A:GLU166:O | Steric; H-acceptor | |
| 6W63 | 2.13058 | Unfavorable | Unfavorable bump | A:GLN189:CA | Steric | :UNK0:C | Steric |
| 1.22635 | Unfavorable | Unfavorable bump | A:GLN189:CA | Steric | :UNK0:H | Steric | |
| 2.37385 | Unfavorable | Unfavorable bump | A:GLN189:CB | Steric | :UNK0:C | Steric | |
| 1.77146 | Unfavorable | Unfavorable bump | A:GLN189:CG | Steric | :UNK0:C | Steric | |
| 0.87813 | Unfavorable | Unfavorable bump | A:GLN189:CG | Steric | :UNK0:H | Steric | |
| 2.2544 | Unfavorable | Unfavorable bump | A:GLN189:CG | Steric | :UNK0:N | Steric | |
| 1.84974 | Unfavorable | Unfavorable bump | A:GLN189:CG | Steric | :UNK0:H | Steric | |
| 1.96838 | Unfavorable | Unfavorable bump | A:GLN189:CD | Steric | :UNK0:C | Steric | |
| 1.11711 | Unfavorable | Unfavorable bump | A:GLN189:CD | Steric | :UNK0:C | Steric | |
| 0.42714 | Unfavorable | Unfavorable bump | A:GLN189:CD | Steric | :UNK0:H | Steric | |
| 1.76113 | Unfavorable | Unfavorable bump | A:GLN189:CD | Steric | :UNK0:H | Steric | |
| 1.81942 | Unfavorable | Unfavorable bump | A:GLN189:OE1 | Steric | :UNK0:C | Steric | |
| 0.89379 | Unfavorable | Unfavorable bump | A:GLN189:OE1 | Steric | :UNK0:H | Steric | |
| 2.09198 | Unfavorable | Unfavorable bump | A:GLN189:NE2 | Steric | :UNK0:N | Steric | |
| 1.10771 | Unfavorable | Unfavorable bump | A:GLN189:NE2 | Steric | :UNK0:C | Steric | |
| 2.13841 | Unfavorable | Unfavorable bump | A:GLN189:NE2 | Steric | :UNK0:O | Steric | |
| 0.54957 | Unfavorable | Unfavorable bump | A:GLN189:NE2 | Steric | :UNK0:C | Steric | |
| 1.46472 | Unfavorable | Unfavorable bump | A:GLN189:NE2 | Steric | :UNK0:H | Steric | |
| 1.1745 | Unfavorable | Unfavorable bump | A:GLN189:NE2 | Steric | :UNK0:H | Steric | |
| 1.45993 | Unfavorable | Unfavorable bump | A:GLN189:NE2 | Steric | :UNK0:H | Steric | |
| 1.25941 | Unfavorable | Unfavorable bump | A:GLN189:HA | Steric | :UNK0:C | Steric | |
| 0.61273 | Unfavorable | Unfavorable bump | A:GLN189:HA | Steric | :UNK0:H | Steric | |
| 1.52091 | Unfavorable | Unfavorable bump | A:GLN189:HA | Steric | :UNK0:H | Steric | |
| 1.40921 | Unfavorable | Unfavorable bump | A:GLN189:HG1 | Steric | :UNK0:H | Steric | |
| 0.92704 | Unfavorable | Unfavorable bump | A:GLN189:HG2 | Steric | :UNK0:C | Steric | |
| 0.19936 | Unfavorable | Unfavorable bump | A:GLN189:HG2 | Steric | :UNK0:H | Steric | |
| 1.69586 | Unfavorable | Unfavorable bump | A:GLN189:HE21 | Steric | :UNK0:N | Steric | |
| 1.11219 | Unfavorable | Unfavorable bump | A:GLN189:HE21 | Steric | :UNK0:C | Steric | |
| 1.37181 | Unfavorable | Unfavorable bump | A:GLN189:HE21 | Steric | :UNK0:C | Steric | |
| 1.39001 | Unfavorable | Unfavorable bump | A:GLN189:HE21 | Steric | :UNK0:H | Steric | |
| 1.45408 | Unfavorable | Unfavorable bump | A:GLN189:HE22 | Steric | :UNK0:C | Steric | |
| 0.8524 | Unfavorable | Unfavorable bump | A:GLN189:HE22 | Steric | :UNK0:C | Steric | |
| 0.83783 | Unfavorable | Unfavorable bump | A:GLN189:HE22 | Steric | :UNK0:H | Steric | |
Table 7.
Unsatisfied bonds in melatonin with coronavirus2 proteins.
Table 8 shows non-covalent interactions between melatonin and coronavirus2 proteins. Hydrophobic groups of protein 6LU7 residues are A:His41, A:Leu141, A:Cys145, A:His164, A:Met165, and A:Glu166; those of protein 6M03 residues are A:His41, A:Met49, A:Phe140, A:Leu141, A:Cys145, A:Met165, A:Glu166, and A:Leu167; and those of protein residues 6W63 are A:His41, A:Cys44, A:Met49, A:Leu50, A:Met165, A:Glu166, A:Leu167, and A:Gln189 with melatonin as shown in Fig. S2 and Table 8. The hydrophilic groups of protein 6LU7 residues are A:His41, A:Asn142, A:His164, A:Glu166, A:His172, A:Asp187, A:Arg188, and A:Gln189; those of protein 6M03 residues are A:His41, A:Asn142, A:His163, A:His164, A:Glu166, A:His172, and A:Gln189; and those of protein 6W63 residues are A:His41, A:His164, A:Glu166, A:Asp187, A:Arg188, A:Gln189, and A:Gln192 with melatonin as shown in Fig. S3 and Table 8. Neutral groups of protein 6LU7 residues are A:Tyr54, A:Gly143, and A:Ser144; those of protein 6M03 residues are A:Gly143, A:Ser144, and A:Pro168; and those of protein 6W63 residues are A:Pro52, A:Tyr54, A:Arg188, and A:Thr190 with melatonin as shown in Fig. S4 and Table 8. Acidic groups of protein 6LU7 residues are A:Glu166 and A:Asp187; that of protein 6M03 residue is A:Glu166; and protein 6W63 residues are A:Glu166 and A:Asp187 with melatonin as shown in Table 8 and Fig. S5. Basic group interactions of protein 6LU7 residues are A:His41, A:His164, A:His172, and A:Arg188; those of protein 6M03 residues are A:His41, A:His163, A:His164, and A:His172; and those of protein 6W63 residues are A:His41, A:His164, and A:Arg188 as shown in Table 8 and Fig. S6. Tan et.al has shown that melatonin and derivatives has excellent biological responses like acting against oxidative stress and free radical scavenging [[62], [63], [64], [65], [66]]. Our studies show that melatonin molecule can interact with different proteins present in the n-CoV-19 virus and inhibit their proliferation. These results need further clinical follow up and could assist in the management of COVID pneumonia.
Table 8.
Non-covalent interactions between melatonin and coronavirus2 proteins.
| PDB IDs | Hydrophobicity | Hydrophilicity | Neutral group | Acidic group | Basic group |
|---|---|---|---|---|---|
| 6LU7 | A:His41, A:Leu141, A:Cys145, A:His164, A:Met165 and A:Glu166 | A:His41, A:Asn142, A:His164, A:Glu166, A:His172, A:Asp187, A:Arg188 and A:Gln189 | A:Tyr54, A:Gly143 and A:Ser144 | A:Glu166 and A:Asp187 | A:His41, A:His164, A:His172 and A:Arg188 |
| 6M03 | A:His41, A:Met49, A:Phe140, A:Leu141, A:Cys145, A:Met165, A:Glu166 and A:Leu167 | A:His41, A:Asn142, A:His163, A:His164, A:Glu166, A:His172 and A:Gln189 | A:Gly143, A:Ser144 and A:Pro168 | A:Glu166 | A:His41, A:His163, A:His164 and A:His172 |
| 6W63 | A:His41, A:Cys44, A:Met49, A:Leu50, A:Met165, A:Glu166, A:Leu167 and A:Gln189 | A:His41, A:His164, A:Glu166, A:Asp187, A:Arg188, A:Gln189 and A:Gln192 | A:Pro52, A:Tyr54, A:Arg188 and A:Thr190 | A:Glu166 and A:Asp187 | A:His41, A:His164 and A:Arg188 |
4. Conclusions
We conducted a detailed quantum-mechanical investigation of the hormone melatonin and regulation of the sleep-wake cycle. Natural bonding orbital studies revealed the intensity of several intramolecular interactions. The various frontier molecular orbital data explain the nature and physical parameters of melatonin, and the non-linear optical properties are compared with urea which is a standard material. Wavefunction studies gave information about electrostatic potentials, average localised ionization and non-covalent interactions. These data helped to predict the reactivity and identify the active site of the reactivity of the molecule. Melatonin docks with novel coronavirus proteins and shows a variety of interactions with an excellent docking score, which leads to inhibition of the virus proteins leading to its destruction. Hence, clinicians can consider incorporating melatonin also in the COVID-19 treatment regime after further studies.
CRediT authorship contribution statement
Nabil Al-Zaqri: Cenceptualization, Funding acquisition. T. Pooventhiran: Investegation, methodology, original draft. Ali Alsalme: Investegation, original draft. Ismail Warad: Resources, review, methods. Athira M. John: Methodology, writing draft. Renjith Thomas: Conceptualization; Formal analysis; Investigation; Methodology; Project administration; Resources; Software; Supervision; Validation; Visualization; Writing - review & editing.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgements
Researchers Supporting Project number (RSP-2020/78), King Saud University, Riyadh, Saudi Arabia.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.molliq.2020.114082.
Appendix A. Supplementary data
Fig. S1. Interactions between melatonin and 6LU7 (A), 6M03 (B), and 6W63 (C) and coronavirus2 protein residues.
Fig. S2. Hydrophobic interactions between melatonin and 6LU7 (A), 6M03 (B), and 6W63 (C) coronavirus2 protein residues.
Fig. S3. Hydrophilic interactions between melatonin and 6LU7 (A), 6M03 (B), and 6W63 (C) coronavirus2 protein residues.
Fig. S4. Neutral group of interactions between melatonin and 6LU7 (A), 6M03 (B), and 6W63 (C) coronavirus2 protein residues.
Fig. S5. Acidic group of interactions between melatonin and 6LU7 (A), 6M03 (B), and 6W63 (C) coronavirus2 protein residues.
Fig. S6. Basic group of interactions between melatonin and 6LU7 (A), 6M03 (B), and 6W63 (C) coronavirus2 protein residues.
References
- 1.Stehle J.H., Saade A., Rawashdeh O., Ackermann K., Jilg A., Sebestény T., Maronde E. A survey of molecular details in the human pineal gland in the light of phylogeny, structure, function and chronobiological diseases. J. Pineal Res. 2011;51:17–43. doi: 10.1111/j.1600-079X.2011.00856.x. [DOI] [PubMed] [Google Scholar]
- 2.Hardeland R., Pandi-Perumal S.R. Melatonin, a potent agent in antioxidative defense: actions as a natural food constituent, gastrointestinal factor, drug and prodrug. Nutr. Metab. 2005;2:22. doi: 10.1186/1743-7075-2-22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Sugden D. Melatonin: binding site characteristics and biochemical and cellular responses. Neurochem. Int. 1994;24:147–157. doi: 10.1016/0197-0186(94)90101-5. [DOI] [PubMed] [Google Scholar]
- 4.Vanĕcek J., Pavlík A., Illnerová H. Brain Res. 1987;435:359–362. doi: 10.1016/0006-8993(87)91625-8. [DOI] [PubMed] [Google Scholar]
- 5.Tan D.-X., Manchester L.C., Qin L., Reiter R.J. Melatonin: a mitochondrial targeting molecule involving mitochondrial protection and dynamics. Int. J. Mol. Sci. 2016;17:2124. doi: 10.3390/ijms17122124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Doghramji K. J. Clin. Sleep Med. 2007;3:S17–S23. [PMC free article] [PubMed] [Google Scholar]
- 7.Prater W.T., Swamy M., Beane M.D., Lester D.B. J. Behav. Brain Sci. 2018;08:117–125. [Google Scholar]
- 8.Pappolla M.A., Chyan Y.J., Poeggeler B., Frangione B., Wilson G., Ghiso J., Reiter R.J. J. Neural Transm. 2000;107:203–231. doi: 10.1007/s007020050018. [DOI] [PubMed] [Google Scholar]
- 9.Sharma R., McMillan C.R., Niles L.P. J. Pineal Res. 2007;43:245–254. doi: 10.1111/j.1600-079X.2007.00469.x. [DOI] [PubMed] [Google Scholar]
- 10.Sharma R., McMillan C.R., Tenn C.C., Niles L.P. Brain Res. 2006;1068:230–236. doi: 10.1016/j.brainres.2005.10.084. [DOI] [PubMed] [Google Scholar]
- 11.Barlow K.M., Esser M.J., Veidt M., Boyd R. J. Neurotrauma. 2019;36:523–537. doi: 10.1089/neu.2018.5752. [DOI] [PubMed] [Google Scholar]
- 12.Golabchi A., Wu B., Li X., Carlisle D.L., Kozai T.D.Y., Friedlander R.M., Cui X.T. Biomaterials. 2018;180:225–239. doi: 10.1016/j.biomaterials.2018.07.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Esposito E., Cuzzocrea S. Curr. Neuropharmacol. 2010;8:228–242. doi: 10.2174/157015910792246155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Liu S., Madu C.O., Lu Y. Oncomedicine. 2018;3:37–47. [Google Scholar]
- 15.Tan D.X. Curr. Neuropharmacol. 2010;8:161. doi: 10.2174/157015910792246263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Srinivasan V., Pandi-Perumal S.R., Maestroni G.J.M., Esquifino A.I., Hardeland R., Cardinali D.P. Neurotox. Res. 2005;7:293–318. doi: 10.1007/BF03033887. [DOI] [PubMed] [Google Scholar]
- 17.Mccarter S.J., Boswell C.L., Louis E.K.S., Dueffert G., Slocumb N., Boeve B.F., Silber M.H., Ch M.B.B., Olson E.J., Tippmann-Peikert M. NIH Public Access. Sleep Med. 2014;14:237–242. doi: 10.1016/j.sleep.2012.09.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jinny C., Suzanne F., Meaghan C., Sean W.C., Fi J., Anderson C. Differential impact of sleep deprivation and circadian timing on reflexive versus inhibitory control of attention. Sci. Rep. 2020;10:1–10. doi: 10.1038/s41598-020-63144-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Gendy M.N.S., Lagzdins D., Schaman J., Le Foll B. Melatonin for Treatment-Seeking alcohol Use Disorder patients with sleeping problems: a randomised clinical pilot trial. Sci. Rep. 2020;10:1–10. doi: 10.1038/s41598-020-65166-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kim S.W., Kim S., Son M., Cheon J.H., Park Y.S. Melatonin controls microbiota in colitis by goblet cell differentiation and antimicrobial peptide production through toll-like receptor 4 signalling. Sci. Rep. 2020;10 doi: 10.1038/s41598-020-59314-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Bagheri H., Afkhami A., Hashemi P., Ghanei M. RSC Adv. 2015;5:21659–21669. [Google Scholar]
- 22.González-gonzález A., González A., Rueda N., Alonso-gonzález C., Menéndez J.M., Martínez-campa C., Mitola S., Cos S. Usefulness of melatonin as complementary to chemotherapeutic agents at different stages of the angiogenic process. Sci. Rep. 2020;10 doi: 10.1038/s41598-020-61622-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Reiter R.J., Ma Q., Sharma R. Treatment of ebola and other infectious diseases: melatonin ‘goes viral’. Melatonin Res. 2020;3(1):43–57. [Google Scholar]
- 24.Diss L.B., Robinson S.D., Wu Y., Fidalgo S., Yeoman M.S., Patel B.A. Age-related changes in melatonin release in the murine distal colon. Chem. Neurosci. 2013;4:879–887. doi: 10.1021/cn4000617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Bidabadi S.S., Vanderweide J., Sabbatini P. Exogenous melatonin improves glutathione content, redox state and increases essential oil production in two Salvia species under drought stress. Sci. Rep. 2020;10 doi: 10.1038/s41598-020-63986-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Tan D.X., Hardeland R. Potential utility of melatonin in deadly infectious diseases related to the overreaction of innate immune response and destructive inflammation: focus on COVID-19. Melatonin Research. 2020;3(1):120–143. doi: 10.32794/mr11250052. (Mar. 2020) [DOI] [Google Scholar]
- 27.Turjanski A.G., Rosenstein R.E., Estrin D.A. Solvation and conformational properties of melatonin: a computational study. J. Mol. Model. 1999;5:271–280. doi: 10.1007/s0089490050271. [DOI] [Google Scholar]
- 28.M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, B. Mennucci, G. A. Petersson, H. Nakatsuji, M. Caricato, X. Li, H. P. Hratchian, A. F. Izmaylov, J. Bloino, G. Zheng, J. L. Sonnenberg, M. Hada, … D. J. Fox, Gaussian09 Revision D.1 9.5 (n.d.).
- 29.http://sobereva.com/multiwfn
- 30.Lu T., Chen F. Multiwfn: a multifunctional wavefunction analyser. J. Comp. Chem. 2012;33:580–592. doi: 10.1002/jcc.22885. [DOI] [PubMed] [Google Scholar]
- 31.Johnson E.R., Keinan S., Mori-Sánchez P., Contreras-García J., Cohen A.J., Yang W. Revealing noncovalent interactions. J. Am. Chem. Soc. 2010;132:6498–6506. doi: 10.1021/ja100936w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zhang R., Wang X., Ni L., Di X., Ma B., Niu S., Liu C., Reiter R.J. COVID-19: melatonin as a potential adjuvant treatment. Life Sci. 2020;250 doi: 10.1016/j.lfs.2020.117583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.https://www.rcsb.org/
- 34.http://www.swissdock.ch/
- 35.https://bioinfo3d.cs.tau.ac.il/PatchDock/
- 36.Allouche A. Software news and updates Gabedit—a graphical user interface for computational chemistry software. J. Comp. Chem. 2012;32:174–182. doi: 10.1002/jcc. [DOI] [PubMed] [Google Scholar]
- 37.De Almeida W.B., Santos H.F.D., O’Malley P.J. A molecular mechanics and semi-empirical conformational analysis of the herbicide diuron inhibitor of photosystem II. Struct. Chem. 1995;6:383–389. [Google Scholar]
- 38.Parr R.G., Pearson R.G. Absolute hardness: companion parameter to absolute electronegativity. J. Am. Chem. Soc. 1983;105:7512–7516. [Google Scholar]
- 39.Thomas R., Mary Y.S., Resmi K.S., Narayana B., Sarojini B.K., Vijayakumar G., Van Alsenoy C. Two neoteric pyrazole compounds as potential anti-cancer agents: Synthesis, electronic structure, physico-chemical properties and docking analysis. J. Mol. Struct. 2019;1181:455–466. [Google Scholar]
- 40.Thadathil D.A., Varghese S., Akshaya K.B., Thomas R., Varghese A. An insight into photophysical investigation of (E)-2-fluoro-N′-(1-(4-nitrophenyl) ethylidene) benzohydrazide through solvatochromism approaches and computational studies. J. Fluoresc. 2019;29:1013–1027. doi: 10.1007/s10895-019-02415-y. [DOI] [PubMed] [Google Scholar]
- 41.Srikanth K.E., Veeraiah A., Pooventhiran T., Thomas R., Solomon K.A., Soma Raju C.J.S., Latha J.N.L. Detailed molecular structure (XRD), conformational search, spectroscopic characterization (IR, Raman, UV, fluorescence), quantum mechanical properties and bioactivity prediction of a pyrrole analogue. Heliyon. 2020;6 doi: 10.1016/j.heliyon.2020.e04106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Mary Y.S., Mary Y.S., Thomas R., Narayana B., Samshuddin S., Sarojini B.K., Armaković S., Armaković S.J., Pillai G.G. Theoretical studies on the structure and various physico-chemical and biological properties of a terphenyl derivative with immense anti-protozoan activity. Aromat. Compd. 2019:1–16. doi: 10.1080/10406638.2019.1624974. [DOI] [Google Scholar]
- 43.Haruna K., Kumar V.S., Sheena Mary Y.S., Popoola S.A., Thomas R., Roxy M.S., Al-Saadi A.A. Conformational profile, vibrational assignments, NLO properties and molecular docking of biologically active herbicide1,1-dimethyl-3-phenylurea. Heliyon. 2019;5 doi: 10.1016/j.heliyon.2019.e01987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Beegum S., Mary Y.S., Mary Y.S., Thomas R., Armaković S., Armaković S.J., Zitko J., Dolezal M., Van Alsenoy C. Exploring the detailed spectroscopic characteristics, chemical and biological activity of two cyanopyrazine-2-carboxamide derivatives using experimental and theoretical tools. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2020;224 doi: 10.1016/j.saa.2019.117414. [DOI] [PubMed] [Google Scholar]
- 45.Thomas R., Mary Y.S., Resmi K.S., Narayana B., Sarojini S.B.K., Armaković S., Armaković S.J., Vijayakumar G., Alsenoy C.V., Mohan B.J. Synthesis and spectroscopic study of two new pyrazole derivatives with detailed computational evaluation of their reactivity and pharmaceutical potential. J. Mol. Struct. 2019;1181:599–612. doi: 10.1016/j.molstruc.2019.01.014. [DOI] [Google Scholar]
- 46.Priya Y.S., Rao K.R., Chalapathi P.V., Veeraiah A., Srikanth K.E., Mary Y.S., Thomas R. Intricate spectroscopic profiling, light harvesting studies and other quantum mechanical properties of 3-phenyl-5-isooxazolone using experimental and computational strategies. J. Mol. Struct. 2020;1203 http://www.ncbi.nlm.nih.gov/pubmed/127461 [Google Scholar]
- 47.Glendening E.D., Reed A.E., Carpenter J.E., Weinhold F. In: NBO Version 3.1. TCI, University of Wisconsin, editor. Madison; 1998. [Google Scholar]
- 48.Al-Otaibi J.S., Almuqrin A.H., Mary Y.S., Thomas R. Modeling the conformational preference, spectroscopic properties, UV light harvesting efficiency, biological receptor inhibitory ability and other physico-chemical properties of five imidazole derivatives using quantum mechanical and molecular mechanics tools. J. Mol. Liq. 2020;310 http://www.ncbi.nlm.nih.gov/pubmed/112871 [Google Scholar]
- 49.Bulat F.A., Toro-Labbé A., Brinck T., Murray J.S., Politzer P. Quantitative analysis of molecular surfaces: areas, volumes, electrostatic potentials and average local ionisation energies. J. Mol. Model. 2010;16:1679–1691. doi: 10.1007/s00894-010-0692-x. [DOI] [PubMed] [Google Scholar]
- 50.Johnson E.R., Keinan S., Mori-Sánchez P., Contreras-García J., Cohen A.J., Yang W. Revealing non-covalent interactions. J. Am. Chem. Soc. 2010;132:6498–6506. doi: 10.1021/ja100936w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Anand K., Ziebuhr J., Wadhwani P., Mesters J.R., Hilgenfeld R. Coronavirus main proteinase (3CLpro) structure: basis for design of anti-SARS drugs. Science (80-. ). 2003;300(5626):1763–1767. doi: 10.1126/science.1085658. [DOI] [PubMed] [Google Scholar]
- 52.Jin Z. Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature. 2020;582(7811):289–293. doi: 10.1038/s41586-020-2223-y. [DOI] [PubMed] [Google Scholar]
- 53.Peele K.A. Molecular docking and dynamic simulations for antiviral compounds against SARS-CoV-2: a computational study. Informatics Med. Unlocked. 2020;19(May):100345. doi: 10.1016/j.imu.2020.100345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Hatada R. Fragment molecular orbital based interaction analyses on COVID-19 main protease - inhibitor N3 complex (PDB ID:6LU7) J. Chem. Inf. Model. 2020 doi: 10.1021/acs.jcim.0c00283. in Press. [DOI] [PubMed] [Google Scholar]
- 55.Zhang Z., Zhao B., Jin Y., Liu Z., Yang X., Rao H. RCSB; 2020. RCSB PDB - 6M03: the crystal structure of COVID-19 main protease in apo form. https://www.rcsb.org/structure/6M03
- 56.Durdagi S., Aksoydan B., Dogan B., Sahin K., Shahraki A. Screening of clinically approved and investigation drugs as potential inhibitors of COVID-19 main protease: a virtual drug repurposing study. ChemRxiv. 2020;(March):1–31. (A.D.) [Google Scholar]
- 57.Mesecar . RCSB; 2020. RCSB PDB - 6W63: Structure of COVID-19 Main Protease Bound to Potent Broad-spectrum Non-covalent Inhibitor X77. [DOI] [Google Scholar]
- 58.Dubey K., Dubey R. Computation screening of narcissoside a glycosyloxyflavone for potential novel coronavirus 2019 (COVID-19) inhibitor. Biom. J. 2020;2019(xxxx):4–8. doi: 10.1016/j.bj.2020.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Osman E.E.A., Toogood P.L., Neamati N. COVID-19: living through another pandemic. ACS Infect. Dis. 2020 doi: 10.1021/acsinfecdis.0c00224. [DOI] [PubMed] [Google Scholar]
- 60.Al-Otaibi J.S., Mary Y.S., Mary Y.S., Panicker C.Y., Thomas R. Cocrystals of pyrazinamide with p-toluenesulfonic and ferulic acids: DFT investigations and molecular docking studies. J. Mol. Struct. 2019;1175:916–926. [Google Scholar]
- 61.Al-Otaibi J.S., Mary Y.S., Mary Y.S., Thomas R. Quantum mechanical and photovoltaic studies on the cocrystals of hydrochlorothiazide with isonazid and malonamide. J. Mol. Struct. 2019;1197:719–726. [Google Scholar]
- 62.Tan D.X., Manchester L.C., Reiter R.J., Qi W.B., Karbownik M., Calvo J.R. Significance of melatonin in antioxidative defense system: reactions and products. Biol. Signals Recept. 2000;9:137–159. doi: 10.1159/000014635. [DOI] [PubMed] [Google Scholar]
- 63.Tan D.X., Chen L.D., Poeggeler B., Manchester L.C., Reiter R.J. Melatonin: a potent, endogenous hydroxyl radical scavenger. Endocr. J. 1993;1:57–60. [Google Scholar]
- 64.Tan D.X., Manchester L.C., Fuentes-Broto L., Paredes S.D., Reiter R.J. Significance and application of melatonin in the regulation of brown adipose tissue metabolism: relation to human obesity. Obes. Rev. 2011;12:167–188. doi: 10.1111/j.1467-789X.2010.00756.x. [DOI] [PubMed] [Google Scholar]
- 65.Tan D.X., Manchester L.C., Reiter R.J., Plummer B.F. Cyclic 3-hydroxymelatonin: a melatonin metabolite generated as a result of hydroxyl radical scavenging. Biol. Signals Recept. 1999;8:70–74. doi: 10.1159/000014571. [DOI] [PubMed] [Google Scholar]
- 66.Tan D.X., Hardeland R., Back K., Manchester L.C., Alatorre-Jimenez M.A., Reiter R.J. On the significance of an alternate pathway of melatonin synthesis via 5-methoxytryptamine: comparisons across species. J. Pineal Res. 2016;61:27–40. doi: 10.1111/jpi.12336. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Fig. S1. Interactions between melatonin and 6LU7 (A), 6M03 (B), and 6W63 (C) and coronavirus2 protein residues.
Fig. S2. Hydrophobic interactions between melatonin and 6LU7 (A), 6M03 (B), and 6W63 (C) coronavirus2 protein residues.
Fig. S3. Hydrophilic interactions between melatonin and 6LU7 (A), 6M03 (B), and 6W63 (C) coronavirus2 protein residues.
Fig. S4. Neutral group of interactions between melatonin and 6LU7 (A), 6M03 (B), and 6W63 (C) coronavirus2 protein residues.
Fig. S5. Acidic group of interactions between melatonin and 6LU7 (A), 6M03 (B), and 6W63 (C) coronavirus2 protein residues.
Fig. S6. Basic group of interactions between melatonin and 6LU7 (A), 6M03 (B), and 6W63 (C) coronavirus2 protein residues.