Abstract
Targeting extracellular vesicle (EV) secretion can have potential clinical implications for cancer therapy, however the precise regulatory mechanisms of EV secretion are not fully understood. Recently, we have shown a novel pathway of EV biogenesis in PCa cell lines, PC3 and PC3M. However, as the characteristics of EVs are divergent even among PCa cell lines, we hypothesized that other pathways or common regulatory pathways of EV biogenesis still exist. Here, we performed quantitative high‐throughput screening to determine the key regulatory genes involved in EV biogenesis in 22Rv1 cells, which secrete a different type of EVs. In total, 1728 miRNAs were screened and miR‐1908 was selected as the potential miRNA regulating EV biogenesis in 22Rv1 cells. Subsequently, we investigated target genes of miR‐1908 using siRNA screening and identified that spermidine synthase (SRM) was the key regulator of EV secretion in 22Rv1 cells. Attenuation of SRM expression significantly inhibited secretion of EVs in 22Rv1 cells, and overexpression of SRM was confirmed in PCa tissues. Furthermore, we found that the number of endosome compartments was increased in cellular cytoplasm after knockdown of the SRM gene. In conclusion, our results showed that miR‐1908‐mediated regulation of SRM can control secretion of EVs in PCa. In addition, these data suggested that the EV secretion pathway was dependent on cellular characteristics.
Keywords: biogenesis of extracellular vesicles, extracellular vesicles, high‐throughput screening, prostate cancer
Our study reports for the first time that miR‐1908 negatively regulates extracellular vesicle (EV) secretion in prostate cancer (PCa). In addition, we identified spermidine synthase (SRM) as the target gene of miR‐1908, which regulates EV secretion in 22Rv1 cells. Our study suggested that the targeting of SRM in PCa could be a novel therapeutic target for PCa.
1. INTRODUCTION
‘Extracellular vesicle’ (EV) is a comprehensive term for different types of small‐sized membrane vesicles secreted by various cell types including exosomes, microvesicles, apoptotic bodies, and prostasomes, which have been reported to play important roles in intercellular communication. 1 , 2 Accumulating evidence has shown that these EVs play important roles in tumor progression, 3 especially in the field of urology, in which prostasomes derived from normal prostate epithelial cells or prostate cancer (PCa) cells are a well known type of EV, and are reported not only to function in normal physiological processes but are also associated with PCa pathogenesis. 4 , 5 Potential targeting of cell‐cell communication via EVs has been proposed as a novel approach for cancer treatment, 6 highlighting that an understanding of the molecular mechanisms of EV secretion in cancer cells may lead to the development of novel therapeutic strategies against EVs from cancer cells.
miRNAs through their dysregulated profiles function as regulators of both tumor progression and suppression in almost all types of cancer. 7 We have shown previously by comprehensive miRNA screening that miRNAs can regulate EV secretion in cancer cells. 8 In this study, 2 metastatic PCa cell lines, PC3 and PC3M, were employed; we found that miR‐26a regulates EV secretion in these cell lines through suppression of SHC4, PFDN4, and CHORDC1. 8 In addition, we have reported previously that the characteristics of EVs from PC3/PC3M and 22Rv1 cells differ even though PC3/PC3M and 22Rv1 are both castration‐resistant prostate cancer (CRPC) cell lines. 9 For instance, the expression levels of EV marker proteins such as Flottilin‐1, Annexin 2, HSP70, and Integrin beta 1 were hardly detected in EVs from 22Rv1. Conversely, CD81 and TSG101 expression levels in EVs in the 22Rv1 cell line were higher than expression levels in EVs from PC3/PC3M cells, 9 suggesting that different types of EVs are secreted from these 2 different types of PCa cell lines. However, there has been no report determining if these PCa cell lines have common EV secretion pathways, or not. In the present study, using comprehensive miRNA screening we found that miR‐1908 negatively regulates EV secretion in 22Rv1 cells, and identified spermidine synthase (SRM) as the target gene of miR‐1908.
2. MATERIALS AND METHODS
2.1. Cell culture
The human PCa cell line 22Rv1 was purchased from the American Type Culture Collection (ATCC CRL‐2505). 22Rv1 cells were cultured in DMEM (Thermo Fisher Scientific) supplemented with 10% heat‐inactivated fetal bovine serum (FBS) and 1% antibiotic‐antimycotic solution (AA, Invitrogen) at 37°C. Human metastatic PCa cell lines PC3 (ATCC CRL‐1435) and PC‐3M‐luc‐C6 (PC3M) (Xenogen) were cultured in RPMI 1640 medium (Thermo Fisher Scientific) supplemented with 10% heat‐inactivated FBS and 1% AA. For routine maintenance, each cell line was grown as a monolayer at 37°C in 5% carbon dioxide in air and 95% relative humidity.
2.2. Preparation of conditioned medium and extracellular vesicle purification
PCa cells were washed with phosphate‐buffered saline (PBS) and conditioned medium (CM) was replaced with advanced DMEM or RPMI medium (Thermo Fisher Scientific) containing AA and 2 mmol/L l‐glutamine. EVs from CM were isolated using a differential ultracentrifugation protocol, as previously reported. 10 Briefly, after cell incubation for 48 h, CM was centrifuged at 2000 g for 10 min at 4°C. To thoroughly remove cellular debris, the resulting supernatants were filtered through a 0.22 µm filter (Millipore). The filtered CM was centrifuged for 70 min at 110 000 g to pellet enriched EVs. Pellets washed with 11 mL PBS were subjected to ultracentrifugation at 110 000 g for another 70 min. EV pellets were stored in a refrigerator at 4°C in PROKEEP low protein adsorption tubes (WATSON) until use.
2.3. Immunoblotting
The following antibodies used for immunoblotting were purchased from COSMO BIO: mouse monoclonal anti‐human CD9 (12A12, dilution 1:1000) and mouse monoclonal anti‐human CD63 (clone 8A12, dilution 1:1000). Mouse monoclonal anti‐human TSG101 (51/TSG101, dilution 1:200) was purchased from BD Biosciences. Rabbit polyclonal anti‐calnexin (#2433, dilution 1:1000) was purchased from Cell Signaling Technology. Rabbit polyclonal anti‐SRM (#19858‐1‐AP, dilution 1:1000) was purchased from Proteintech. Mouse monoclonal anti‐Actin (clone C4, dilution 1:1000) was purchased from Merck Millipore.
Secondary antibodies [horseradish peroxidase‐labeled sheep anti‐mouse IgG (NA931) and horseradish peroxidase‐linked anti‐rabbit IgG (NA934)] were purchased from GE HealthCare. The EV fraction was measured for its protein content using a Micro BCA Protein Assay kit (Thermo Fisher Scientific). Equal amounts of EVs were loaded onto 4%‐15% Mini‐PROTEAN TGX gels (Bio‐Rad). Following electrophoresis (100 V, 30 mA), proteins were transferred onto a polyvinylidene difluoride membrane. Membranes were blocked with Blocking One solution (Nacalai Tesque) and then incubated with primary antibodies. After washing, membranes were incubated with horseradish peroxidase‐conjugated sheep anti‐mouse IgG or donkey anti‐rabbit IgG and then subjected to enhanced chemiluminescence using ImmunoStar LD (Wako).
2.4. ExoScreen
AlphaLISA reagents (Perkin Elmer, Inc) consisted of AlphaScreen streptavidin‐coated donor beads (6760002), AlphaLISA unconjugated‐acceptor beads (6062011), and AlphaLISA Universal buffer (AL001F). Mouse monoclonal anti‐human CD9 antibody (clone 12A12) was used to modify either the acceptor beads or biotin, following the manufacturer's protocol. AlphaLISA assay was performed in 96‐well half‐area white plates (6005560) and read on an EnSpire Alpha 2300 Multilabel Plate reader (Perkin Elmer, Inc). A 96‐well half‐area white plate was filled with 5 µL EV or 10 µL CM, 5 nmol/L biotinylated antibodies, and 50 µg/mL AlphaLISA acceptor beads conjugated to antibodies in universal buffer. The volume of each reagent was 10 µL. The plate was then incubated for 1 h at 37°C. Without going to the washing step, 25 µL of 80 µg/mL AlphaScreen streptavidin‐coated donor beads were added. The reaction mixture was incubated in the dark for another 30 min at 37°C. The, the plate was then read on an EnSpire Alpha 2300 Multilabel Plate reader at an excitation wavelength of 680 nm and emission wavelength of 615 nm. Background signals obtained from filtrated Advanced RPMI or PBS were subtracted from the measured signals.
2.5. Cell proliferation and migration assays
A colorimetric cell proliferation assay was performed using the Cell Counting Kit‐8 (Dojindo Molecular Technologies Inc) following the manufacturer's instructions. Cell migration was characterized with wound healing and migration assays. A cell migration assay was performed in 24 well plates using BD Falcon Cell Culture Inserts (BD Biosciences) that contained uncoated transwell polycarbonate membrane filters with 8 µm pores and following the manufacturer's instructions.
2.6. Quantitative high‐throughput screening for extracellular vesicle regulators
Experiments were performed as described previously. 8 Briefly, high‐throughput miRNA screening (total 1728 miRNAs) was performed using the AccuTarget™ Human miRNA mimic library constructed based on the miRBase database v.21 (CosmoBio). A 100 µL 22Rv1 cell suspension containing 15 000 cells/well (in DMEM containing 10% serum without antibiotics) was seeded into 96‐well plates and incubated for 24 h. Then, transfections with 10 nmol/L miRNA mimics or siRNAs were performed using DharmaFECT Transfection Reagent 1 (Dharmacon) according to the manufacturer's protocol. After 24 h, medium was replaced with advanced DMEM medium containing 2 mmol/L l‐glutamine without AA. After 48 h, the medium was changed, and ExoScreen and cell proliferation assays were performed, as described above. Briefly, after 10 µL of CM was collected for ExoScreen assay, cellular viability was examined using a cell proliferation assay. Relative value of EV secretion/cell viability in each well was calculated as compared to values obtained from the negative control well.
2.7. Analysis of extracellular vesicle particles by nanoparticle tracking analysis
Nanoparticle tracking analysis (NTA) was carried out using the NanoSight LM10 with NTA2.3 Analytical software (NanoSight). For particle tracking, at least five 60 s videos were acquired of each sample with a camera gain of 7. Analysis settings were optimized and kept constant between samples. EV concentrations were calculated as particle/cell of culture to obtain the net vesicle secretion rates.
2.8. Transient transfection assay
22Rv1 cells were transfected with miRNA mimics or siRNAs using DharmaFECT Transfection Reagent 1 according to the manufacturer's protocol. The negative control miRNA (miRNA mimic negative control #1, Ambion) was used to investigate the effect of miR‐518, miR‐1253, miR‐1908, and miR‐4460 on EV secretion. siRNAs against NSD1 and SRM, and ALL STAR negative control siRNA were used to investigate the effect of these genes on EV secretion. After 24 h, CM was changed to advanced DMEM medium containing an AA and 2 mmol/L l‐glutamine. After 48 h, the medium was changed and total RNA was extracted using a miRNeasy Mini Kit (Qiagen) according to the manufacturer's instructions; then, SRM expression was determined using real‐time PCR. CM was collected and purified for EVs using ultracentrifugation.
2.9. Microarray analysis
Gene expression for mRNA was analyzed using the Agilent Array platform (8x60K, Agilent Technologies), following the manufacturer's protocols. The Gene Expression Omnibus accession number is GSE 143350.
2.10. RNA extraction and real‐time PCR analysis
Total RNA was extracted from cultured cells using QIAzol and a miRNeasy Mini Kit (Qiagen) according to the manufacturer's protocols. Purity and concentration of all RNA samples were quantified using a NanoDrop ND‐1000 spectrophotometer (Thermo Fisher Scientific). A reverse transcription reaction was performed using a High‐Capacity cDNA Reverse Transcription Kit (Applied Biosystems) and a random hexamer primer. Real‐time PCR analyses were performed using StepOne Plus and TaqMan Universal PCR Master Mix (Thermo Fisher Scientific). mRNA expression was normalized to β‐actin. TaqMan probes for NSD1, SRM, and β‐actin were purchased from Applied Biosystems.
2.11. Plasmid constructs and luciferase reporter assay
The following annealed oligos (Thermo Fisher Scientific) were used to construct 3′‐UTR‐reporter vectors.
NSD1_3′UTR_S: 5′‐TCGAGTGCTTTTTTGCCGCGCCCCCCCCCCCCCGCCCCCATAGATTGTCAGCTGTAAGTGAAACTCCTAGTGAAAGC
NSD1_3′UTR_AS: 5′‐GGCCGCTTTCACTAGGAGTTTCACTTACAGCTGACAATCTATGGGGGCGGGGGGGGGGGGGCGCGGCAAAAAAGCAC
NSD1_3′UTR_mut_S: 5′‐TCGAGTGCTTTTTTGCCGCGCCCCCCCCCGGGGCGGCCCATAGATTGTCAGCTGTAAGTGAAACTCCTAGTGAAAGC
NSD1_3′UTR_mut_AS: 5′‐GGCCGCTTTCACTAGGAGTTTCACTTACAGCTGACAATCTATGGGCCGCCCCGGGGGGGGGCGCGGCAAAAAAGCAC
SRM_3′UTR_1_S: 5′‐TCGAGCGGCATTCAGACTTGGGTTCAAATTCCCACCATGCCCCGCCCCCTATGTGGACAAATTGAGAAAGCAAGTGTGGGCACCGC
SRM_3′UTR_1_AS: 5′‐GGCCGCGGTGCCCACACTTGCTTTCTCAATTTGTCCACATAGGGGGCGGGGCATGGTGGGAATTTGAACCCAAGTCTGAATGCCGC
SRM_3′UTR_mut_1_S: 5′‐TCGAGCGGCATTCAGACTTGGGTTCAAATTCCCACCATGGGGGCGGCCCTATGTGGACAAATTGAGAAAGCAAGTGTGGGCACCGC
SRM_3′UTR_mut_1_AS: 5′‐GGCCGCGGTGCCCACACTTGCTTTCTCAATTTGTCCACATAGGGCCGCCCCCATGGTGGGAATTTGAACCCAAGTCTGAATGCCGC
SRM_3′UTR_2_S: 5′‐TCGAGTGGGCCAGGCACTTTCATTTAACCTTCACAACCACCTGCCCGCCAGGCATTGTTCCTGCTGCACAGAGGCAAGGCGATGTGC
SRM_3′UTR_2_AS: 5′‐GGCCGCACATCGCCTTGCCTCTGTGCAGCAGGAACAATGCCTGGCGGGCAGGTGGTTGTGAAGGTTAAATGAAAGTGCCTGGCCCAC
SRM_3′UTR_mut_2_S: 5′‐TCGAGTGGGCCAGGCACTTTCATTTAACCTTCACAACCACCTGGGGCGGTGGCATTGTTCCTGCTGCACAGAGGCAAGGCGATGTGC
SRM_3′UTR_mut_2_AS: 5′‐GGCCGCACATCGCCTTGCCTCTGTGCAGCAGGAACAATGCCACCGCCCCAGGTGGTTGTGAAGGTTAAATGAAAGTGCCTGGCCCAC
The annealed oligos for 3′UTR of NSD1 or SRM were subcloned into thepsiCHECK2 vector that had been digested with XhoI 35 and NotI. To mutate miR‐1908 recognition sites in NSD1 or SRM, annealed oligos were used.
Luciferase reporter assays were performed by co‐transfecting HEK293 cells with 8 µL of 5 µmol/L miR‐1908 mimic or negative control miRNA, and with 500 ng of psiCHECK2 reporter plasmids using Lipofectamine 3000 (Thermo Fisher Scientific); a dual‐luciferase reporter assay (Promega) was performed after 48 h.
2.12. Immunofluorescence
Cells cultured on glass dishes were washed once with PBS and fixed in 4% paraformaldehyde (Wako). After fixation, cells were washed with PBS and permeabilized with 0.1% Triton X‐100 for 15 min. Then, cells were washed with PBS and incubated with Blocking One buffer to block nonspecific binding. The glass dishes were incubated with mouse monoclonal anti‐human CD63 (clone 8A12), rinsed with 0.05% Tween 20/PBS, incubated with the appropriate fluorescently labeled secondary antibodies, and then examined using confocal microscopy.
2.13. Live‐cell staining with LysoTracker
Lysotracker Red (50 nmol/L, Molecular Probes, Invitrogen) was added to the cells in serum‐free advanced DMEM, followed by incubation at 37°C for 2 h. After incubation, cells were washed thoroughly with advanced DMEM and observed (FluoView 1000 Olympus Confocal Microscope) with excitation filters 510‐560 nm and barrier filter 590 nm.
2.14. Statistical analysis
Unless otherwise described, data are presented as the mean ± SE, and statistical significance was determined using Student t test. In the dot plot, bars indicate the median and interquartile range, and statistical significance was determined using Student t test. A P‐value < .05 was considered statistically significant.
3. RESULTS
3.1. Difference in characteristics of EVs secreted from prostate cancer cells
To confirm the difference in EVs between PC3/PC3M and 22Rv1, EVs derived from these PCa cell lines were isolated by ultracentrifugation and characterized using phase‐contrast electron microscopy and NTA. The number of secreted EV particles from 22Rv1 cells was higher than that from PC3M cells (Figure 1A). In addition, we observed bilayer‐membrane vesicles whose size was heterogeneous, ranging in diameter from 50 to 300 nm in PC3, however size of EVs from 22Rv1 cells was almost homogeneous (Figure 1B). Compared with NTA data in a previous report, 9 these data suggested that 22Rv1 cells possessed a different EV secretion pathway from PC3 or PC3M cells, and promoted us to examine 22Rv1 cells to elucidate the novel EV secretion pathway in PCa.
Figure 1.
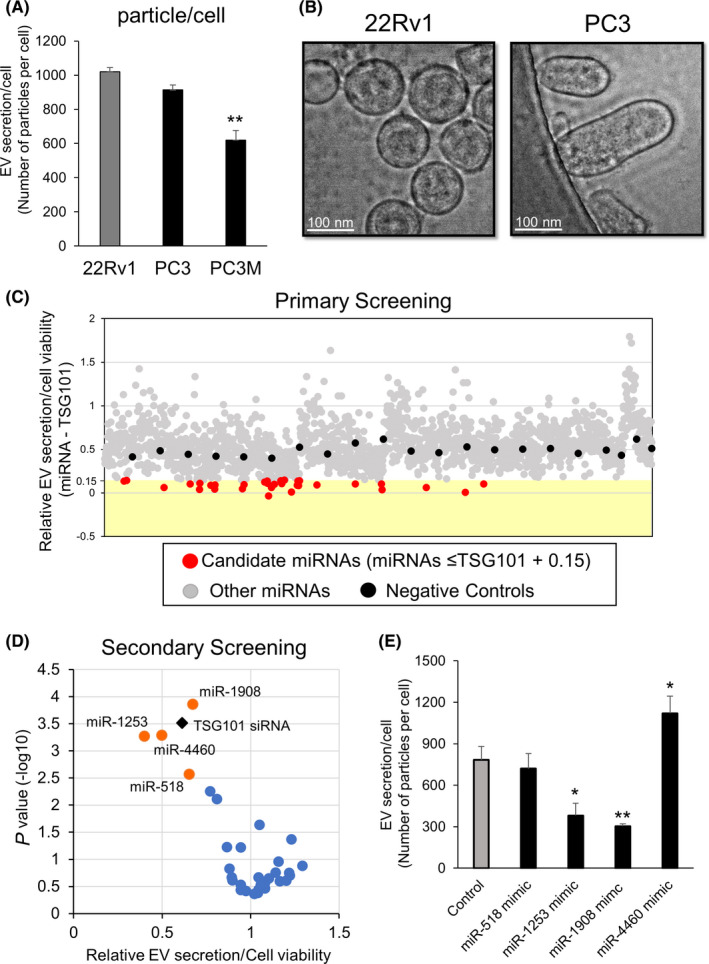
Screening of miRNAs regulating extracellular vesicle secretion. A, Amount of extracellular vesicle (EV) secretion in 22Rv1, PC3, and PC3M cell. The number of EVs was examined using a nanoparticle tracking system. Values represent mean ± SE (n = 3). The statistical significance was calculated using Dunnett's t test (**P < .01). B, Representative image of EVs isolated from 22Rv1 and PC3 cells under transmission electron microscopy. Scale bars, 100 nm. C, Result of primary screening. Distribution of the score for each sample calculated from the amount of EVs and cell viability; 1728 miRNAs are plotted as red dots or gray dots according to cut‐off value (score < 0.15). D, Result of the secondary screening. Distribution of the scores for 33 selected samples. Thirty‐three miRNAs are plotted according to cut‐off value (score < 0.15). E, Effect of miR‐518, miR‐1253, miR‐1908, and miR‐4460 on the EV secretion per 22Rv1 cell. Number of EV secreted from each 22Rv1 cell was examined using a nanoparticle tracking system. Values represent mean ± SE (n = 3). The statistical significance was calculated using Dunnett's t test (*P < .05, **P < .01)
We collected EVs from 22Rv1 cells (Figure S1A) and confirmed that EVs were positive for CD9, CD63, and TSG101 markers, and were negative for calnexin and actin (Figure S1B). Considering these data, we decided to perform the first screening using CD9 antibodies to determine any change in EV secretion after transfection with each miRNA mimic. In addition, to determine any effects of miRNA transfections on cell proliferation, we also performed a Cell Counting Kit‐8 assay. To date, several genes have been reported to regulate EV biogenesis 11 , 12 , 13 , 14 ; we decided to use one of these genes as a positive control gene for this assay. We transfected siRNA targeting these genes and evaluated the effect on EV secretion, and selected TSG101 as the positive control gene (Figure S2A). Then, we performed the first high‐throughput screening (Figure S2B). We selected candidate miRNAs according to the criteria described in Figure S2C. We performed this high‐throughput screening twice (primary screening and secondary screening) and selected 4 candidate miRNAs: miR‐518, miR‐1253, miR‐1908, and miR‐4460 (Figure 1C,D). For validation, we evaluated the number of EV particles secreted by each 22Rv1 cell that was transfected with candidate miRNA mimics, using NTA and cell counting (Figure 1E). Finally, we selected miR‐1908 as the potential regulator of EV secretion in 22Rv1 cells because 22Rv1 cells transfected with miR‐1908 showed the maximum reduction in number of secreted EV particles. Furthermore, we searched some of the relevant public databases (GSE32448, 46602, and 55945) and found that miR‐1908 was downregulated in PCa tissues (Figure S2D). Intriguingly, based on this miRNA screening, we could not see any suppressive effect of EV secretion in 22Rv1 cells by miR‐26a, which was the most effective miRNA for suppression of EV secretion in PC3M. 8 Likewise, miR‐1908 was not an EV suppressive miRNA in PC3M cells (Figure S2E). These results suggested that the EV secretion pathway in 22Rv1 cells was different from that in PC3M cells.
3.2. Selection of candidate genes potentially regulating extracellular vesicle secretion in prostate cancer cells
To further elucidate the mechanism of EV secretion in 22Rv1 cells, we conducted further screening using siRNAs. We performed an mRNA microarray analysis in 22Rv1 cells following transfections with miR‐1908 mimic or negative control miRNA. To identify genes that could potentially be targeted by miR‐1908, we used TargetScan and found that miR‐1908 overexpression in 22Rv1 cells resulted in downregulation of 104 genes compared with control cells (Figure 2A). Therefore, in this screening, we screened for genes regulating EV secretion by siRNA‐mediated targeting of each candidate 104 genes (Figure 2B). We selected genes that showed a relative value of EV secretion/cell viability lower than that of miR‐1908 treated cells, as evaluated using ExoScreen and cell proliferation assay. Results of each screening are shown in Figure 2C. Consequently, we identified 3 genes, NSD1, SRM, and ZF385A as candidate genes that could potentially be regulating EV secretion in 22Rv1 (Figure 2C). Then, we validated our data by evaluating the effect of knocking down these 3 genes (following transfections with their respective targeted siRNAs) on the number of secreted EVs in 22Rv1 cells, using NTA and cell counting. The number of secreted EV particles was decreased after treatment with siRNAs of NSD1 and SRM (Figure 2D). In addition, we found that expression levels of NSD1 and SRM were significantly upregulated in PCa tissue compared with normal tissues (Figure 2E).
Figure 2.
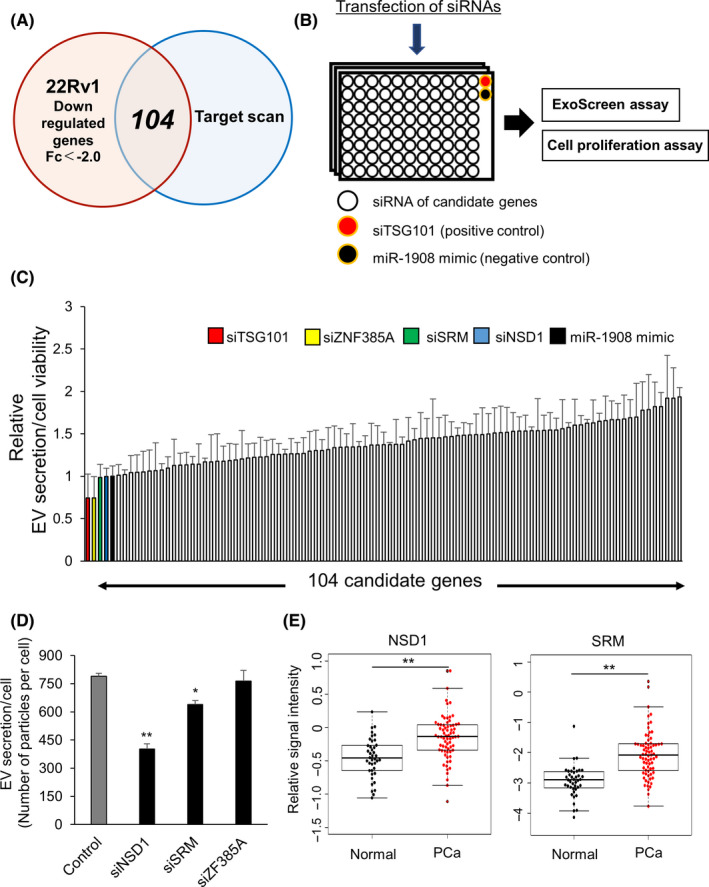
NSD1 and SRM are candidate genes for regulating extracellular vesicle secretion. A, Venn diagram of predicted miR‐1908 targets (TargetScan) and transcripts that were experimentally repressed >2‐fold by miR‐1908 overexpression in 22Rv1 cells relative to the control condition. B, Schematic diagram of the high‐throughput screening to detect genes regulating extracellular vesicle (EV) secretion. C, Results of the screening from candidate 104 genes. The effect of 104 genes and miR‐1908 (control) on the secretion of EVs and cell viability. Secretion of EVs was evaluated by ExoScreen, and cell viability was examined by Cell Counting Kit‐8 assay. Relative values represent mean ± SE (n = 3). D, Effect of transfections with NSD1 siRNA, SRM siRNA, and ZNF385A siRNA on EV secretion per 22Rv1 cell. The amount of EV secreted per cell was examined using a nanoparticle tracking system. Values represent mean ± SE (n = 3). **P < .01. E, Expression levels of NSD1 and SRM in prostate cancer and normal prostate tissue clinical specimens (GSE6099). **P < .01
3.3. miR‐1908 regulates extracellular vesicle secretion by targeting SRM
To examine whether miR‐1908 could directly regulate these genes, we constructed luciferase reporters of SRM or NSD1, which are target candidates of miR‐1908, at the 3′‐UTR of the reporter gene (Figures 3A and S3A), and performed luciferase reporter assays. Ectopic expression of miR‐1908 significantly suppressed the luciferase activity of wild‐type SRM 3′‐UTRs but not their mutant 3′‐UTRs (Figure 3B). In addition, we confirmed that miR‐1908 regulated expression levels of SRM, as depicted by PCR and immunoblot analysis (Figure 3C,D). These results provided experimental evidence that miR‐1908 could directly repress translation of SRM. As for NSD1, we found suppression of luciferase activity not only in wild‐type 3′‐UTR of NSD1, but also in the mutant 3′‐UTR of NSD1, suggesting that miR‐1908 is involved in suppression of NSD1 through a regulatory mechanism that is different from conventional miRNA regulation (Figure S3B).
Figure 3.
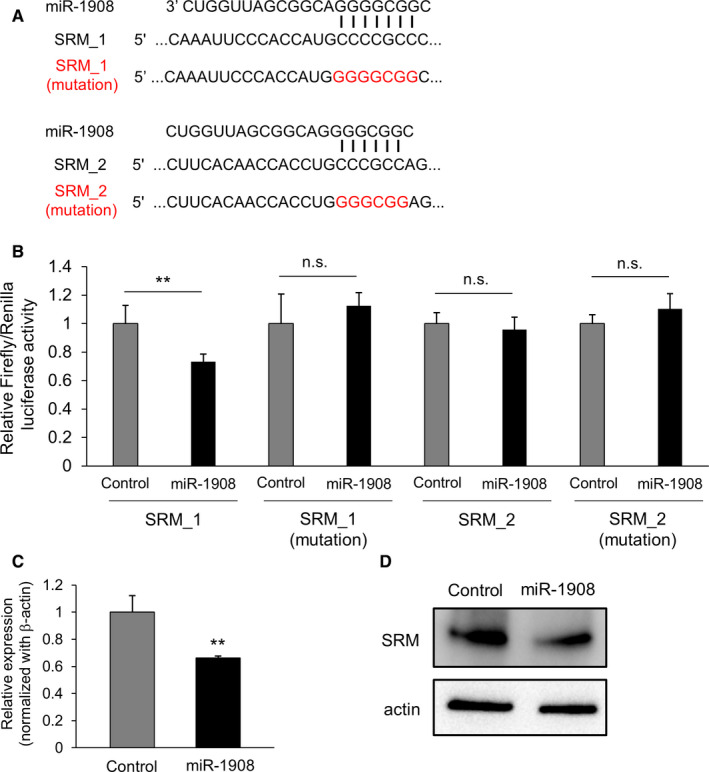
miR‐1908 directly regulates the expression levels of spermidine synthase (SRM). A, Summary of miR‐1908 target sites and mutated sites (shown in red) in the 3′‐UTR of SRM. B, Target validation of SRM was performed using a luciferase reporter assay. Values are depicted as fold change relative to the negative control miRNA (control). The values represent mean ± SE (n = 3). **P < .01; and n.s. indicates not significant. C, Effect of miR‐1908 on the expression levels of the target genes in 22Rv1 cells. The values are depicted as fold change relative to the negative control miRNA (control). The values represent mean ± SE (n = 3). **P < .01. D, Immunoblot analysis of 22Rv1 cells transfected with miR‐1908 mimic or negative control miRNA (control). 10 µg of the cell lysates derived from the transfected PCa cells was loaded for detection of SRM and actin
3.4. SRM did not affect cell viability in prostate cancer cells
We confirmed downregulation of SRM by its target siRNA in 22Rv1 cells (Figure S4A,B). Then, to evaluate the property changes in 22Rv1 cells after repression of SRM by siRNA, we observed cellular morphology, proliferation, and motility. As a result, attenuation of SRM in 22Rv1 did not affect these phenotypes (Figure S5A‐D). Therefore, these results suggested that SRM regulates EV secretion but not cell‐autonomous phenotypes such as cellular morphology, proliferation, and motility. In addition, we searched the expression levels of SRM in CRCP tissue using public databases (GSE70770). Although there was no significant difference between hormone‐sensitive prostate cancer (HSPC) and CRPC, SRM was significantly upregulated in HSPC and CRPC tissue compared with normal prostate tissue (Figure S6A). As we could not find any significant difference in SRM expression in HSPC and CRPC tissues (Figure S6A), we concluded that the relationship between the machinery of miR‐1908 and SRM was general in any type of PCa, not only in CRPC. Furthermore, we examined whether expression levels of SRM predicted progression‐free survival, based on mRNA expression in PCa (TCGA, PanCancer Atlas) through the cBioPortal (http://www.cbioportal.org/). Our analysis revealed that PCa patients with higher expression of SRM tended to have shorter progression‐free intervals (P = .181; Figure S6B).
3.5. SRM regulates extracellular vesicle secretion in prostate cancer cells
For further analysis of SRM in EV secretion in 22Rv1 cells, we performed immunofluorescence and LysoTracker staining. To examine the effect of SRM on EV secretion, we stained cells with LysoTracker, which labels acidic organelles such as late endosome or lysosome, after treatment with siRNA targeting SRM. As a result, the number of EVs stained with LysoTracker increased following downregulation of SRM in 22Rv1 cells (Figure 4A), suggesting that knockdown of SRM could lead to accumulation of either late endosomes or lysosomes. To clarify this further, we analyzed the effects of SRM knockdown by staining for the CD63 marker. We found that attenuation of SRM significantly increased the number of CD63‐positive structures in cellular cytoplasm (Figure 4B). These results suggested that SRM regulates the release of EVs into extracellular space in 22Rv1 cells. It is already known that SRM is an enzyme that catalyzes production of spermidine from putrescine and decarboxylated S‐adenosylmethionine. 15 Therefore, we examined the effects of spermidine on EV secretion in 22Rv1 cells. However, we did not observe any effects on EV secretion after treatment with spermidine in SRM under attenuated or steady‐state condition (Figure S7).
Figure 4.
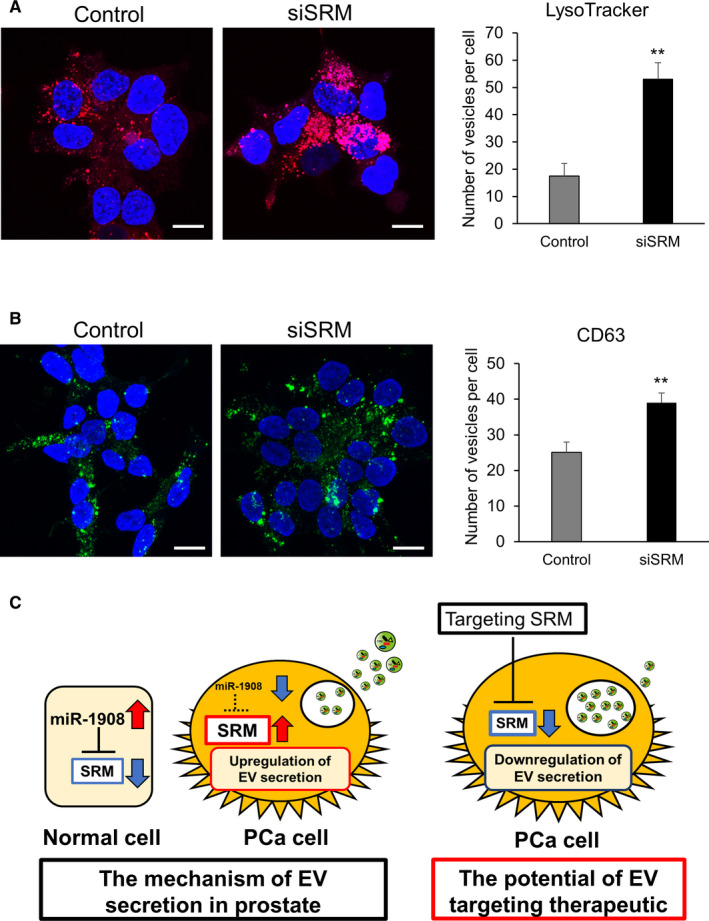
Sspermidine synthase (SRM) promotes the secretion of extracellular vesicles in prostate cancer cells. A, Representative images of LysoTracker staining in control and SRM knockdown 22Rv1 cells. Scale bars, 10 μm. The number of vesicles in 22Rv1 cells was counted. Values represent mean ± SE (n = 3). B, Confocal immunofluorescence micrographs of 22Rv1 cells transfected with SRM siRNA. Cells were stained with anti‐CD63 antibody. Scale bars, 10 µm. The number of vesicles in PCa cells was counted. The values represent mean ± SE (n = 5). C, Schematic model of the regulation of extracellular vesicle (EV) secretion in PCa. The novel signaling network of miR‐1908 and its target SRM regulating EV secretion in PCa cells. In the normal cells, expression of miR‐1908 was high, therefore expression of SRM was suppressed. However, because of downregulation of miR‐1908 in PCa cells, expression of SRM was upregulated and led to the acceleration of EV secretion from PCa cells, resulting in the promotion of cancer malignancy. Therefore, targeting SRM in PCa could be a novel therapeutic target for PCa
Collectively, we performed screening of 1728 species of miRNAs using our established EV detection method and elucidated a novel signaling network involving miR‐1908 and its target SRM that could regulate EV secretion in 22Rv1 cells (Figure 4C). In addition, these data also suggested that the EV secretion pathway is different among PCa cell types.
4. DISCUSSION
Extracellular vesicles have been known to modulate cancer progression for about 10 years 16 ; several recent reports have shown the potential therapeutic value of reducing cancer‐derived EVs in inhibiting cancer proliferation and dissemination. 6 We recently showed that administration of human‐specific antibodies against CD9 and CD63 markers, which are enriched on the surface of EVs, could significantly decrease metastasis in a human breast cancer xenograft mouse model. 17 This study provided promising evidence that inhibition of circulating EVs could be a novel strategy for cancer treatment. Despite significant advances in the understanding of the role of EVs in cancer progression, the mechanism involved in EV biogenesis, which has a great potential in therapeutic targeting in cancers, remains obscure.
Recently, we elucidated a novel pathway of EV biogenesis in PCa cell lines, PC3 and PC3M, using quantitative high‐throughput screening. 8 However, characteristics of EVs were different among PCa cell lines. 9 These data prompted us to examine the other pathways of EV biogenesis and secretion in a different type of PCa cell line, such as 22Rv1. For the first screen, we utilized an miRNA mimic library, and selected miR‐1908 as the miRNA potentially regulating EV secretion in PCa cells. Several studies have reported the role of miR‐1908 in cancers. In liver cancer and osteosarcomas, miR‐1908 was aberrantly expressed, 18 , 19 , 20 especially in osteosarcoma, where overexpression of miR‐1908 promoted cell proliferation, migration, and invasion, 19 and was strongly associated with poor prognosis. 20 However, expression of miR‐1908 was significantly downregulated in non–small‐cell lung cancer (NSCLC), and transfection of miR‐1908 mimic reduced cell proliferation. 21 Wang et al 22 reported decreased expression levels of miR‐1908 in PCa. However, the role of miR‐1908 in PCa has not been elucidated. Our study is the first to report the novel role of miR‐1908 as an inhibitor of secretion of EVs in PCa cells. Intriguingly, as shown in Figure 1D, we could not see any suppressive effect of EV secretion by miR‐26a in 22Rv1 cells. Conversely, miR‐1908 did not suppress secretion of EVs from PC3M cells (Figure S2E). These results suggested that the EV secretion pathway in 22Rv1 cells was different from that in PC3M cells.
In addition, we hypothesized that by screening for target genes of miR‐1908, we could identify specific genes involved in EV secretion. Therefore, we performed screening using a cherry‐picked siRNA library and identified SRM as the gene regulating EV secretion in PCa cells. Expression levels of SRM were upregulated in PCa tissue. In addition, although attenuation of SRM did not affect cell proliferation or migration, patients with higher expression of SRM tended to have shorter progression‐free survival. These results suggested that the contribution of SRM in cancer progression might be through non–cell‐autonomous mechanisms such as promotion of EV secretion in PCa cells, resulting in the modulation of the cancer microenvironment and leading to cancer progression.
SRM is a synthase that catalyzes the conversion of putrescine to spermidine, a polyamine that is particularly abundant in sperm. 23 Previous reports have shown the role of spermidine in tumor biology, however these roles remain controversial. Several studies have reported the oncogenic properties of spermidine. 24 , 25 , 26 , 27 Pless et al 26 reported that SAM486A, an inhibitor of S‐adenosylmethionine‐decarboxylase that leads to low spermidine concentrations, had moderate success in non‐Hodgkin's lymphoma. Soda et al 27 reported that spermidine may inhibit colon carcinogenesis in mice, however it promotes tumor growth once cancer has developed. Contrary to these oncogenic properties of spermidine, several reports have shown that spermidine administration can reduce the incidence of hepatocellular carcinoma and tumor progression of colorectal cancer in mouse models. 28 , 29 In our study, exogenous treatment with spermidine did not affect cell proliferation or EV secretion of PCa cells. Therefore, the precise mechanism by which SRM in EV secretion should be investigated in a future study; our data suggested that SRM regulates EV secretion.
In conclusion, in this study, we noticed divergence of characteristics for EVs between PC3/PC3M and 22Rv1 cells and screened EV regulator miRNAs and genes using 22Rv1 cells. We identified a novel mechanism of miR‐1908 regulated EV secretion in PCa. This difference in EV regulating pathways reflected the difference in EV characteristics. This novel signaling network for miR‐1908 and its target SRM in regulating EV secretion could be a novel therapeutic target for PCa.
DISCLOSURE
The authors declare no conflict of interest.
Supporting information
Figs S1‐S7
ACKNOWLEDGMENTS
We thank Ms. Ayako Inoue for supporting this study. This study was supported by the Practical Research for Innovative Cancer Control (19ck0106366h0003) from the Japan Agency for Medical Research and Development; the ‘Development of Diagnostic Technology for Detection of miRNA in Body Fluids’ grant from AMED; Kobayashi Foundation for Cancer Research; Princess Takamatsu Cancer Research Fund; Yasuda Medical Foundation; Yamaguchi Endocrine Research Foundation; and the Foundation for Promotion of Cancer Research in Japan.
Urabe F, Kosaka N, Sawa Y, et al. The miR-1908/SRM regulatory axis contributes to extracellular vesicle secretion in prostate cancer. Cancer Sci. 2020;111:3258–3267. 10.1111/cas.14535
Contributor Information
Nobuyoshi Kosaka, Email: nkosaka@tokyo-med.ac.jp.
Yusuke Yamamoto, Email: yuyamamo@ncc.go.jp.
REFERENCES
- 1. Urabe F, Kosaka N, Ito K, Kimura T, Egawa S, Ochiya T. Extracellular vesicles as biomarkers and therapeutic targets for cancer. Am J Physiol Cell Physiol. 2020;318(1):C29‐C39. [DOI] [PubMed] [Google Scholar]
- 2. Fujita Y, Yoshioka Y, Ochiya T. Extracellular vesicle transfer of cancer pathogenic components. Cancer Sci. 2016;107(4):385‐390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Becker A, Thakur BK, Weiss JM, Kim HS, Peinado H, Lyden D. Extracellular vesicles in cancer: cell‐to‐cell mediators of metastasis. Cancer Cell. 2016;30(6):836‐848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Urabe F, Kosaka N, Kimura T, Egawa S, Ochiya T. Extracellular vesicles: toward a clinical application in urological cancer treatment. Int J Urol. 2018;25(6):533‐543. [DOI] [PubMed] [Google Scholar]
- 5. Kosaka N, Iguchi H, Yoshioka Y, Hagiwara K, Takeshita F, Ochiya T. Competitive interactions of cancer cells and normal cells via secretory microRNAs. J Biol Chem. 2012;287(2):1397‐1405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Kosaka N, Yoshioka Y, Fujita Y, Ochiya T. Versatile roles of extracellular vesicles in cancer. J Clin Investig. 2016;126(4):1163‐1172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Lu J, Getz G, Miska EA, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435(7043):834‐838. [DOI] [PubMed] [Google Scholar]
- 8. Urabe F, Kosaka N, Sawa Y, et al. miR‐26a regulates extracellular vesicle secretion from prostate cancer cells via targeting SHC4, PFDN4, and CHORDC1. Sci Adv. 2020;6(18):eaay3051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Yoshioka Y, Konishi Y, Kosaka N, Katsuda T, Kato T, Ochiya T. Comparative marker analysis of extracellular vesicles in different human cancer types. J Extracell Vesicles. 2013;2(1):20424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Yokoi A, Yoshioka Y, Yamamoto Y, et al. Malignant extracellular vesicles carrying MMP1 mRNA facilitate peritoneal dissemination in ovarian cancer. Nat Commun. 2017;8:14470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Baietti MF, Zhang Z, Mortier E, et al. Syndecan‐syntenin‐ALIX regulates the biogenesis of exosomes. Nat Cell Biol. 2012;14(7):677‐685. [DOI] [PubMed] [Google Scholar]
- 12. Colombo M, Moita C, van Niel G, et al. Analysis of ESCRT functions in exosome biogenesis, composition and secretion highlights the heterogeneity of extracellular vesicles. J Cell Sci. 2013;126(Pt 24):5553‐5565. [DOI] [PubMed] [Google Scholar]
- 13. Trajkovic K, Hsu C, Chiantia S, et al. Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science. 2008;319(5867):1244‐1247. [DOI] [PubMed] [Google Scholar]
- 14. Ostrowski M, Carmo NB, Krumeich S, et al. Rab27a and Rab27b control different steps of the exosome secretion pathway. Nat Cell Biol. 2010;12(1):19‐30. [DOI] [PubMed] [Google Scholar]
- 15. Wu H, Min J, Ikeguchi Y, et al. Structure and mechanism of spermidine synthases. Biochemistry. 2007;46(28):8331‐8339. [DOI] [PubMed] [Google Scholar]
- 16. Kosaka N, Iguchi H, Yoshioka Y, Takeshita F, Matsuki Y, Ochiya T. Secretory mechanisms and intercellular transfer of microRNAs in living cells. J Biol Chem. 2010;285(23):17442‐17452. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Nishida‐Aoki N, Tominaga N, Takeshita F, Sonoda H, Yoshioka Y, Ochiya T. Disruption of circulating extracellular vesicles as a novel therapeutic strategy against cancer metastasis. Mol Ther. 2017;25(1):181‐191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Jin JC, Jin XL, Zhang X, Piao YS, Liu SP. Effect of OSW‐1 on microRNA expression profiles of hepatoma cells and functions of novel microRNAs. Mol Med Rep. 2013;7(6):1831‐1837. [DOI] [PubMed] [Google Scholar]
- 19. Yuan H, Gao Y. MicroRNA‐1908 is upregulated in human osteosarcoma and regulates cell proliferation and migration by repressing PTEN expression. Oncol Rep. 2015;34(5):2706‐2714. [DOI] [PubMed] [Google Scholar]
- 20. Lian D, Wang ZZ, Liu NS. MicroRNA‐1908 is a biomarker for poor prognosis in human osteosarcoma. Eur Rev Med Pharmacol Sci. 2016;20(7):1258‐1262. [PubMed] [Google Scholar]
- 21. Ma Y, Feng J, Xing X, et al. miR‐1908 overexpression inhibits proliferation, changing Akt activity and p53 expression in hypoxic NSCLC cells. Oncol Res. 2016;24(1):9‐15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Wang L‐Y, Cui J‐J, Zhu T, et al. Biomarkers identified for prostate cancer patients through genome‐scale screening. Oncotarget. 2017;8(54):92055‐92063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Allen RD, Roberts TK. Role of spermine in the cytotoxic effects of seminal plasma. Am J Reprod Immunol Microbiol. 1987;13(1):4‐8. [DOI] [PubMed] [Google Scholar]
- 24. Nowotarski SL, Woster PM, Casero RA Jr. Polyamines and cancer: implications for chemotherapy and chemoprevention. Expert Rev Mol Med. 2013;15:e3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Gerner EW, Meyskens FL Jr. Polyamines and cancer: old molecules, new understanding. Nat Rev Cancer. 2004;4(10):781‐792. [DOI] [PubMed] [Google Scholar]
- 26. Pless M, Belhadj K, Menssen HD, et al. Clinical efficacy, tolerability, and safety of SAM486A, a novel polyamine biosynthesis inhibitor, in patients with relapsed or refractory non‐Hodgkin's lymphoma: results from a phase II multicenter study. Clin Cancer Res. 2004;10(4):1299‐1305. [DOI] [PubMed] [Google Scholar]
- 27. Soda K, Kano Y, Chiba F, Koizumi K, Miyaki Y. Increased polyamine intake inhibits age‐associated alteration in global DNA methylation and 1,2‐dimethylhydrazine‐induced tumorigenesis. PLoS One. 2013;8(5):e64357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Yue F, Li W, Zou J, et al. Spermidine prolongs lifespan and prevents liver fibrosis and hepatocellular carcinoma by activating MAP1S‐mediated autophagy. Cancer Res. 2017;77(11):2938‐2951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Miao H, Ou J, Peng Y, et al. Macrophage ABHD5 promotes colorectal cancer growth by suppressing spermidine production by SRM. Nat Commun. 2016;7:11716. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figs S1‐S7


