Abstract
Tumor progression is governed by various growth factors and cytokines in the tumor microenvironment (TME). Among these, transforming growth factor-β (TGF-β) is secreted by various cell types residing in the TME and promotes tumor progression by inducing the epithelial-to-mesenchymal transition (EMT) of cancer cells and tumor angiogenesis. TGF-β comprises three isoforms, TGF-β1, -β2, and -β3, and transduces intracellular signals via TGF-β type I receptor (TβRI) and TGF-β type II receptor (TβRII). For the purpose of designing ligand traps that reduce oncogenic signaling in the TME, chimeric proteins comprising the ligand-interacting ectodomains of receptors fused with the Fc portion of immunoglobulin are often used. For example, chimeric soluble TβRII (TβRII-Fc) has been developed as an effective therapeutic strategy for targeting TGF-β ligands, but several lines of evidence indicate that TβRII-Fc more effectively traps TGF-β1 and TGF-β3 than TGF-β2, whose expression is elevated in multiple cancer types. In the present study, we developed a chimeric TGF-β receptor containing both TβRI and TβRII (TβRI-TβRII-Fc) and found that TβRI-TβRII-Fc trapped all TGF-β isoforms, leading to inhibition of both the TGF-β signal and TGF-β–induced EMT of oral cancer cells, whereas TβRII-Fc failed to trap TGF-β2. Furthermore, we found that TβRI-TβRII-Fc suppresses tumor growth and angiogenesis more effectively than TβRII-Fc in a subcutaneous xenograft model of oral cancer cells with high TGF-β expression. These results suggest that TβRI-TβRII-Fc may be a promising tool for targeting all TGF-β isoforms in the TME.
Keywords: angiogenesis, cancer therapy, epithelial-to-mesenchymal transition (EMT), Fc chimeric receptor, oral squamous cell cancer, transforming growth factor-β (TGF-β), tumor microenvironment, cancer therapy, Fc receptor, angiogenesis, ligand trap
The tumor microenvironment (TME) is a complex system comprising tumor cells, stromal cells, endothelial cells, immune cells, and the extracellular matrix. The TME plays important roles during tumor progression via growth regulation and metastasis of cancer cells (1). Recent lines of evidence have revealed that the TME composition changes during tumor progression in response to oncogenic signals from tumor cells; therefore, effective anti-tumor therapies should target not only cancer cells but also other TME components.
Cells in the TME secrete various growth factors and cytokines to create a microenvironment favoring tumorigenesis. Among these factors, transforming growth factor-β (TGF-β) is secreted by various cell types residing in the TME. The pleiotropic functions of TGF-β, including not only the regulation of cell growth, differentiation, and apoptosis but also cell motility and invasion, have been widely studied (2). Although TGF-β can function as an early-stage tumor suppressor, it may also act as a tumor promoter (3). TGF-β appears to create a favorable TME for cancer cells, stimulating cancer metastasis (3). These dual roles in tumorigenesis likely depend on an increased level of active TGF-β within the TME as well as dysregulation of TGF-β signaling pathways that leads to the reprogramming of the cellular signals that drive tumorigenesis.
TGF-β comprises three isoforms, TGF-β1, -β2, and -β3. All three isoforms share high sequence similarity and essentially signal through the same receptors; however, their effects on tumorigenesis may depend on the cellular context and differences in their temporal and spatial expressions (4). The canonical TGF-β signaling pathway relies on the activation of Smad transcription factors upon binding of TGF-βs to their transmembrane receptors: TGF-β type I receptor (TβRI) and TGF-β type II receptor (TβRII). Binding of TGF-β to TβRII stimulates phosphorylation of TβRI, leading to the activation of Smad transcription factors and expression of direct target genes (5). TGF-β1 and TGF-β3 bind with the high affinity to TβRII dimers in the absence of TβRI; however, TGF-β2 exhibits lower affinity for dimers formed by either TβRI or TβRII (6, 7), suggesting that binding of TGF-β2 requires the formation of a functional complex between the two types of these receptors.
Although increased expression of all three biologically active TGF-β isoforms has been reported in various cancers, including colorectal cancer (8), breast cancer (9), and glioblastoma (10), substantial TGF-β2 expression seems to be particularly associated with the progression of multiple types of cancer (11). This increased TGF-β2 level may drive tumorigenesis by stimulating the epithelial-to-mesenchymal transition (EMT) of cancer cells and facilitating metastasis. Moreover, analysis of a public database, the Cancer Cell Line Encyclopedia (RRID:SCR_013836) revealed elevated TGF-β2 expression in multiple cancer cell lines, including glioma and osteosarcoma, concordant with previous reports showing elevated TGF-β2 expression in glioma patients (12).
High TGF-β2 expression is associated with poor prognosis of advanced gliomas (12), skin squamous carcinoma (13), and lung cancer (14). Furthermore, a Kaplan–Meier analysis (KM plotter) (15) of multiple cancer types, done by our group, showed that higher TGF-β2 but not TGF-β1 or TGF-β3 (Fig. S1, A–C) levels were correlated with poor survival of patients with head and neck squamous cell carcinoma.
Oral squamous cell carcinoma (OSCC) is a common malignant tumor of the head and neck region. It represents the sixth most common cancer in the world, with over 300,000 cases diagnosed and 145,000 deaths annually (16). Despite treatment advances, the survival of patients with OSCC has not significantly improved in recent decades. OSCC has a poor clinical outcome due to primary metastases to the lymph nodes, lungs, liver, or other distant organs. High TGF-β expression has been detected in specimens obtained from patients with oral cancer cells metastasizing to bones (17). Moreover, we previously reported that TGF-β signals could stimulate oral cancer cell migration (18), suggesting the important role of TGF-β in oral cancer progression.
TGF-β increases the migratory and invasive properties of cancer cells via inducing EMT, a process during which epithelial cells lose their apical-basal polarity, becoming more motile and invasive by gaining properties attributed to mesenchymal cells (19). TGF-β signals induce the expression of various EMT-related transcription factors, resulting in decreased expression of proteins participating in cell junction complexes (e.g. E-cadherin and claudin-1). Down-regulation of the epithelial cell markers is accompanied by increased expression of multiple mesenchymal marker genes (e.g. smooth muscle protein 22α (SM22α), vimentin, or matrix metalloproteinases) that results in reorganization of cell cytoskeletal architecture and a mesenchymal and invasive phenotype in cancer cells (20).
Various strategies targeting the TGF-β signals have been used to develop effective anti-tumor therapies, with several clinical studies under way (21). Many trials were based on the development of antibodies targeting TGF-β to prevent its binding to its receptors. Fresolimumab (GC1008), a mAb neutralizing all TGF-β isoforms, has been tested in the treatment of renal cell carcinoma (22) and metastatic breast cancer (23). Small molecules have also been developed to inhibit TGF-β signaling by targeting the kinase activity of TGF-β receptors. Among them, galunisertib (LY2157299) is in the advanced phases of clinical trials media for the treatment of hepatocellular and pancreatic adenocarcinomas (NCT02423343) (24). Other approaches, involving belagenpumatucel-L, a vaccine including cancer cell lines modified with antisense TGF-β2 for the treatment of non-small-cell lung cancer, are also in progress (25). However, despite promising results, some of those therapeutic strategies show limited efficacy or show skin and heart toxicity (26).
Other attempts have been made to develop ligand trap-based strategies. Ligand traps are chimeric proteins comprising the ligand-interacting ectodomains of receptors fused with human IgG1 Fc domains. Ligand traps are promising tools that can be used to reduce the concentrations of ligands in the extracellular environment. Several chimeric proteins capable of inhibiting TGF-β have been designed and successfully tested, demonstrating their efficacy for selective blocking of TGF-β family ligands in pathological conditions (27, 28); however, their therapeutic potential is isoform-selective, as they trap TGF-β1 and TGF-β3 but not TGF-β2 (27). Because high TGF-β2 expression is associated with the progression or resistance of multiple tumors (8, 11), there is a need to develop an effective strategy to eliminate all three TGF-β isoforms. TGF-β2 signaling requires the formation of a functional complex between TβRI and TβRII (7). del Re et al. (29) reported that the combined exposure of mink lung epithelial cells, Mv1Lu, to a mixture of the soluble forms of TβRI-Fc and TβRII-Fc could inhibit TGF-β2 signaling, suggesting efficient ligand trapping via a combination of exogenously added receptors. However, due to difficulties in precisely controlling the effective concentration of both ligand traps and differences in the half-lives of TGF-β receptor types, this method may not be useful for in vivo applications. To overcome these problems, in the present study, we developed a molecular ligand trap, TβRI-TβRII-Fc, a chimeric protein that interacted with all TGF-β isoforms. We also showed its effective application in vitro to inhibit TGF-β signals and in vivo to inhibit tumor growth of oral cancer cells, thus demonstrating the chimeric Fc molecular trap as a promising tool for the development of effective TME-targeted therapies.
Results
An Fc chimeric protein containing extracellular domains of both TGF-β type I and II receptors effectively inhibits Smad2/3/4 signals mediated by all TGF-β isoforms
A previous report demonstrated that the soluble extracellular domain of the TGF-β type II receptor fused to the Fc portion of human IgG (TβRII-Fc) inhibited the biological activities of TGF-β1 and TGF-β3 but not TGF-β2 (29). Moreover, the inhibitory effects of TβRII-Fc were strengthened by the addition of the soluble TGF-β type I receptor (TβRI-Fc) (29). To examine these effects, we constructed expression plasmids encoding TβRI-Fc, TβRII-Fc, and Fc alone (Control-Fc), as a negative control (Fig. 1A). The chimeric proteins were efficiently expressed in HEK293T cells and secreted into culture media, as shown in immunoblots with anti-human IgG-Fc antibodies (Fig. 1B). We first examined whether constructed chimeric proteins are able to trap TGF-β ligands using ELISA (Fig. 1C). As expected, TβRI-Fc chimeric protein did not exhibit significant interaction with any of TGF-β isoforms as compared with Control-Fc (negative control). On the other hand, recombinant TβRII-Fc chimeric protein could interact with immobilized TGF-β1 and TGF-β3 proteins, but not with TGF-β2. Next, we examined the ability of various Fc chimeric proteins to inhibit TGF-β signals using HEK-Blue TGF-β reporter cells. The HEK-Blue TGF-β reporter system allows direct monitoring of TGF-β signaling by quantification of secreted alkaline phosphatase (SEAP), which is expressed under the control of Smad2/3/4-inducible elements. HEK-Blue TGF-β reporter cells treated with any of the three TGF-β isoforms, TGF-β1, -β2, or -β3, could activate the TGF-β signal transduction pathway, as revealed by increased SEAP activity (Fig. 1D). Preincubation of TGF-βs with conditioned media of HEK293T cells expressing Control-Fc or TβRI-Fc did not decrease the level of the TGF-β signal (Fig. 1D). Although TβRII-Fc chimeric protein could significantly inhibit the signal activated by TGF-β1 and TGF-β3, it did not suppress signals generated in response to the TGF-β2 isoform. Similar results were also obtained for the combination of both chimeric proteins, TβRI-Fc and TβRII-Fc. This mixture was sufficient to inhibit TGF-β1 and TGF-β3 but not TGF-β2 signals (Fig. 1D).
Figure 1.
TβRI-TβRII-Fc protein specifically traps all three TGF-β isoforms, resulting in Smad2/3/4 signal inhibition. A, schematic representation of the Fc chimeric receptors used in this study. hIgG-Fc, Fc portion of human IgG; TβRI, extracellular domain of TGF-β type I receptor; TβRII, extracellular domain of TGF-β type II receptor. B, expression of Fc chimeric proteins in HEK293T cells. Conditioned media from HEK293T cells were collected 48 h post-transfection with plasmids expressing various Fc chimeric proteins, and the concentration of each Fc chimeric protein was determined by ELISA. Culture media containing 119 ng of each Fc chimeric protein, respectively, were analyzed by immunoblotting with anti-IgG-Fc antibodies. The bands (black arrowheads) correspond to the expressed ligand traps secreted into the culture media. C, affinity of soluble chimeric receptors toward TGF-β ligands as measured by ELISA. Binding affinity of recombinant Fc chimeric proteins was assayed toward recombinant TGF-β1, TGF-β2, or TGF-β3 ligands. Values represent colorimetric changes of alkaline phosphatase substrate conjugated to anti-human-Fc antibodies and measured at 450–620 nm. D, effect of Fc chimeric proteins on Smad2/3/4 signal activation by various TGF-β isoforms in HEK-Blue TGF-β reporter cells cultured in the absence or presence of medium containing TGF-β1 (Tβ1), TGF-β2 (Tβ2), or TGF-β3 (Tβ3) (1 ng/ml) and conditioned media from HEK293T cells expressing soluble chimeric receptor, Control-Fc, TβRI-Fc, TβRII-Fc, TβRI-TβRII-Fc, or a combination of TβRI-Fc and TβRII-Fc, as described under “Experimental procedures.” The values represent Smad2/3/4 signal activation corresponding to the colorimetric changes of the Quanti-Blue substrate by SEAP at 640 nm. E, inhibitory potential of Fc chimeric proteins toward various TGF-β family members. TGF-β isoforms (1 ng/ml) and activin A (1 ng/ml) were incubated with conditioned media from HEK293T cells containing Control-Fc, TβRI-Fc, TβRII-Fc, TβRI-TβRII-Fc, or SB431542 (10 μm), followed by the measurement of Smad2/3/4 signals activated in HEK-Blue TGF-β reporter cells for 24 h. Samples incubated without soluble Fc chimeric proteins (Control) or containing the TGF-β signal inhibitor, SB431542 (10 μm) were considered as negative and positive controls, respectively. The values are represented as the -fold changes of control samples treated with the indicated ligands and inhibitor. Error bars, S.D. *, p < 0.05; **, p < 0.01; ***, p < 0.001; N.S., not significant.
To target all TGF-β isoforms, we generated Fc chimeric receptors containing both TGF-β type I and II receptors (TβRI-TβRII-Fc: Fig. 1, A and B) and found that the TβRI-TβRII-Fc chimeric protein could interact not only with TGF-β1 and TGF-β3 proteins, but also with TGF-β2 (Fig. 1C). Consistent with these results, TβRI-TβRII-Fc effectively inhibited the signals mediated by all three isoforms (Fig. 1D). Whereas we observed that 50 ng of TβRI-TβRII-Fc chimeric protein inhibited the signals induced by all TGF-β isoforms, smaller amounts of TβRI-TβRII-Fc (5 and 10 ng) were not capable of effectively inhibiting TGF-β2 but significantly inhibited both TGF-β1 and TGF-β3 (Fig. S2), suggesting the dose-dependent effect of this chimeric protein. The dose-dependent effect was also observed for TβRII-Fc chimeric protein, toward TGF-β1 and -β3 ligands, with no effect on TGF-β2 (Fig. S2), strengthening the conclusion that only TβRI-TβRII-Fc chimeric protein traps all isotypes of TGF-β.
We next studied the specificity of the inhibitory capacity of TβRI-TβRII-Fc. Whereas activin A, a member of the TGF-β family, binds to activin type I and II receptors that are distinct from TGF-β receptors, it also activates Smad2/3/4 signals in HEK-Blue TGF-β reporter cells. This effect could be abolished by SB431542, an inhibitor of TGF-β and activin type I receptors (Fig. 1E). In contrast, none of the Fc chimeras trapped activin A (Fig. 1E), suggesting that TβRI-TβRII-Fc can trap all TGF-β isoforms but not activin A.
TβRI-TβRII-Fc protein suppresses the expression of target genes of TGF-β signals in oral cancer cells
High TGF-β2 expression is correlated with poor head and neck cancer patient survival, as suggested by Kaplan–Meier analysis (Fig. S1B). Thus, we used SAS cells, an OSCC cell line, to examine whether TGF-β receptor-Fc chimeras could be used to change the extracellular milieu of cancer cells and influence their response to TGF-β present in the extracellular environment. We first tested the effect of chimeric proteins on TGF-β signaling in SAS cells. SAS cells were incubated in conditioned media derived from HEK293T cells expressing Fc chimeric proteins in the absence or presence of TGF-β1, -β2, or -β3, respectively, followed by an analysis of the expression of transmembrane prostate androgen–induced protein (TMEPAI; PMEPA1), a direct downstream target of TGF-β, by quantitative RT-PCR (qRT-PCR). As shown in Fig. 2A, increased TMEPAI expression was detected in control SAS cells treated with either TGF-β isoform. As expected, the presence of SB431542 decreased TMEPAI expression to the control level (Fig. 2A). Consistent with the results obtained in the HEK-Blue TGF-β reporter system, TMEPAI was not inhibited in cells treated with TGF-β in the presence of either Control-Fc or the TβRI-Fc. The TβRII-Fc chimeric protein inhibited TMEPAI expression induced by either TGF-β1 or TGF-β3 and had no effect on cells incubated with the TGF-β2 isoform. The inhibition of signals generated upon incubation with any of the three isoforms was observed only in the presence of the TβRI-TβRII-Fc chimeric protein (Fig. 2A), suggesting that only TβRI-TβRII-Fc protein is a potent inhibitor of all three TGF-β isoforms. The inhibitory effect of TβRI-TβRII-Fc on all three TGF-β isoforms was confirmed by the analysis of plasminogen activator inhibitor-1 (PAI-1; SERPINE1) expression, another gene directly regulated by TGF-β. Up-regulation of PAI-1 induced by any of the three TGF-β isoforms was inhibited only in the presence of TβRI-TβRII-Fc chimeric protein and not the other chimeric proteins, TβRI-Fc or TβRII-Fc (Fig. 2B).
Figure 2.
TβRI-TβRII-Fc protein suppresses the expression of genes activated by TGF-β signals in SAS oral cancer cells. SAS cells were cultured for 24 h in a mixture of conditioned media from HEK293T cells expressing various chimeric receptors (Control-Fc, TβRI-Fc, TβRII-Fc, or TβRI-TβRII-Fc) and TGF-β isoforms (TGF-β1 (Tβ1), TGF-β2 (Tβ2), or TGF-β3 (Tβ3)) (2 ng/ml) and subjected to qRT-PCR analysis for the expression of TMEPAI (PMEPA1) (A) and PAI-1 (SERPINE1) (B). The negative and positive controls were incubated without soluble Fc chimeric proteins (Control) or with 10 μm SB431542, respectively. All data are normalized to the expression of β-actin (ACTB). Error bars, S.D. *, p < 0.05; **, p < 0.01; ***, p < 0.001; N.S., not significant.
To generalize the findings obtained in SAS oral cancer cells, we utilized another OSCC cell line, HSC-4. Expression of TMEPAI (Fig. S3A) and PAI-1 (Fig. S3B) induced in the presence of TGF-β2 was suppressed only upon incubation in conditioned medium derived from HEK293T cells expressing TβRI-TβRII-Fc but not the other chimeric receptors. These results suggest that TβRI-TβRII-Fc chimeric protein effectively inhibits the signals mediated by all TGF-β isoforms in multiple types of OSCC cells, whereas TβRII-Fc protein fails to inhibit TGF-β2–mediated signals.
Oral cancer cells undergo EMT in response to all TGF-β isoforms
As TGF-β is a potent inducer of EMT in various types of cancer (20, 30), we next examined whether treatment with any of the TGF-β isoforms would induce EMT in oral cancer cells. SAS cells were treated with each TGF-β isoform or SB431542 for 72 h, and the expression of various epithelial and mesenchymal markers was assessed by qRT-PCR. As shown in Fig. 3, SAS cells incubated with any of the three TGF-β isoforms showed decreased expression of genes encoding the epithelial markers, E-cadherin and claudin-1 (Fig. 3, A and B), accompanied by increased expression of the mesenchymal cell markers, SM22α, vimentin, and fibronectin (Fig. 3, C–E), suggesting that SAS cells undergo EMT in response to each TGF-β isoform. In contrast, inhibition of TGF-β signals with SB431542 increased the expression of the E-cadherin (Fig. 3A) and significantly decreased the expression of SM22α and fibronectin (Fig. 3, C and E). The effects of TGF-β isoforms on the expression of epithelial and mesenchymal markers were essentially confirmed at the protein level by immunoblotting and immunocytochemical analyses (Fig. 3, F and G). To generalize our findings, we performed the same set of experiments in HSC-4 cells. All TGF-β isoforms decreased the expression of E-cadherin (Fig. S4A). The expression of claudin-1 was decreased by TGF-β1 and -β2, but not by TGF-β3 (Fig. S4B). All TGF-β isoforms up-regulated the expression of mesenchymal markers SM22α, vimentin, and fibronectin (Fig. S4, C–E), as revealed by qRT-PCR analyses and essentially confirmed by immunoblotting (Fig. S4F) and immunocytochemical analyses (Fig. S4G). We also observed that TGF-β–induced EMT increased the size of HSC-4 cells, which was consistent with a previous report (31). Whereas the effects of TGF-β isoforms and SB431542 on the expression of epithelial and mesenchymal markers differ between SAS and HSC-4 cells, possibly due to the differences in cellular contexts, the results obtained suggest that all TGF-β isoforms are capable of inducing EMT in multiple types of OSCC cells.
Figure 3.
EMT program is activated in SAS cells in response to all TGF-β isoforms. SAS cells were cultured in the absence (Ctrl) or presence of TGF-β1 (Tβ1), TGF-β2 (Tβ2), or TGF-β3 (Tβ3) (2 ng/ml) and the TGF-β signal inhibitor, SB431542 (SB; 10 μm) for 72 h, followed by evaluation of EMT by qRT-PCR analyses (A–E), immunoblotting (F), and immunocytochemistry (G). Shown is qRT-PCR analysis for the expression of epithelial cell markers E-cadherin (A) and claudin-1 (B) and mesenchymal markers SM22α (C), vimentin (D), and fibronectin (E) in response to the ligands or inhibitor. All data are normalized to the β-actin expression. Error bars, S.D. *, p < 0.05; **, p < 0.01; ***, p < 0.001; N.S., not significant. F, immunoblot analysis with antibodies specific to epithelial and mesenchymal markers and α-tubulin. G, immunofluorescent images of SAS cells cultured in the presence or absence of ligands or inhibitor, followed by staining for E-cadherin (green), vimentin (red), and nuclei (blue). Representative images of cells cultured under each experimental condition are shown. Scale bar, 50 μm.
TβRI-TβRII-Fc protein inhibits EMT of oral cancer cells
After demonstrating that all three TGF-β isoforms induce EMT of SAS and HSC-4 oral cancer cells (Fig. 3 and Fig. S4), we next investigated how ligand traps affected the EMT of oral carcinoma cells induced by TGF-β isoforms. SAS cells were treated without or with TGF-β1, -β2, or -β3 in the presence of conditioned medium obtained from HEK293T cells expressing Control-Fc, TβRI-Fc, TβRII-Fc, or TβRI-TβRII-Fc protein and analyzed for the expression of epithelial (E-cadherin (Fig. 4A) and claudin-1 (Fig. 4B)) and mesenchymal (SM22α (Fig. 4C), vimentin (Fig. 4D), and fibronectin (Fig. 4E)) markers. SAS cells incubated with any TGF-β isoform in conditioned medium containing either Control-Fc or TβRI-Fc proteins showed decreased E-cadherin and claudin-1 expression (Fig. 4, A and B) accompanied by up-regulation of all mesenchymal markers (SM22α, vimentin, and fibronectin) (Fig. 4, C–E), suggesting that neither Control-Fc nor TβRI-Fc chimeric protein inhibited the EMT of SAS cells. Although TβRII-Fc chimeric protein could suppress TGF-β1– or TGF-β3–dependent EMT accompanied by changes in the expression of epithelial (Fig. 4, A and B) and mesenchymal markers (Fig. 4, C–E) in SAS cells, it did not prevent the EMT of SAS cells induced by TGF-β2 (Fig. 4, A–E). In contrast, SAS cells stimulated with any of the three TGF-β isoforms in conditioned medium obtained from HEK293T cells expressing TβRI-TβRII-Fc chimeric protein did not undergo EMT, with levels of the epithelial (Fig. 4, A and B) and mesenchymal markers (Fig. 4, C–E) comparable with those in the untreated control cells. Successful inhibition of EMT by TβRI-TβRII-Fc chimeric protein was also confirmed by immunocytochemical evaluation of the changes in E-cadherin and vimentin expression in response to various TGF-βs in the absence or presence of chimeric proteins (Fig. 4F). Consistent with the qRT-PCR results, neither Control-Fc nor TβRI-Fc prevented the up-regulation of vimentin expression accompanied by the down-regulation of E-cadherin, as revealed in antibody staining (Fig. 4F, top two rows). The presence of TβRII-Fc affected only the changes induced by TGF-β1 and TGF-β3 but not TGF-β2 (Fig. 4F, second row from the bottom). However, TβRI-TβRII-Fc chimeric protein led to almost complete inhibition of EMT induced by all TGF-β isoforms (Fig. 4F, bottom row). The efficacy of the TβRI-TβRII-Fc chimeric protein in suppressing EMT-associated changes in epithelial and mesenchymal marker expression was also examined at both RNA and protein levels in HSC-4 cells. As expected, the EMT of HSC-4 cells induced by any of the three TGF-β isoforms was inhibited only in the presence of the TβRI-TβRII-Fc protein (Fig. S5), suggesting that the TβRI-TβRII-Fc protein inhibited the EMT of oral cancer cells induced by all TGF-β isoforms.
Figure 4.

TβRI-TβRII-Fc protein inhibits the EMT of SAS cells. SAS cells were cultured for 72 h in conditioned media from HEK293T cell expressing various Fc chimeric proteins (Control-Fc, TβRI-Fc, TβRII-Fc, or TβRI-TβRII-Fc) in the presence of TGF-β1 (Tβ1), TGF-β2 (Tβ2), or TGF-β3 (Tβ3) (2 ng/ml). The stimulation of EMT was evaluated by qRT-PCR (A–E) and immunocytochemistry (F). Shown is qRT-PCR analysis for the expression of epithelial cell markers E-cadherin (A) and claudin-1 (B) and mesenchymal markers SM22α (C), vimentin (D), and fibronectin (E). All data are normalized to the β-actin expression. Error bars, S.D. *, p < 0.05; **, p < 0.01; ***, p < 0.001; N.S., not significant. F, immunofluorescent images of SAS cells cultured in conditioned media from HEK293T cells expressing various Fc chimeric proteins (Control-Fc, TβRI-Fc, TβRII-Fc, or TβRI-TβRII-Fc) for 72 h in the presence or absence of the ligands. The cells were stained for E-cadherin (green), vimentin (red), and nuclei (blue). Representative images of cells cultured under each experimental condition are shown. Scale bar, 50 μm.
TβRI-TβRII-Fc protein inhibits tumor growth in vivo
TGF-β signals promote tumor progression in multiple types of cancer, including oral cancer. Thus, to test the effect of TβRI-TβRII-Fc protein on tumor growth in vivo, we utilized a subcutaneous xenograft model of SAS oral cancer cells expressing a high level of endogenous TGF-βs. We established SAS cells expressing Control-Fc, TβRI-Fc, TβRII-Fc, and TβRI-TβRII-Fc chimeric proteins. These ligand traps were efficiently expressed in SAS cells (Fig. 5A, right) and secreted into the medium (Fig. 5A, left), although the efficiency of TβRI-TβRII-Fc secretion was lower than that of TβRII-Fc. The size of ligand traps accumulated in conditioned medium was slightly higher compared with the one detected in cell lysates, suggesting that the soluble receptor traps may have undergone posttranslational modification, glycosylation, via passing through the similar secretory conventional secretory pathway as endogenous TGF-β receptors (32).
Figure 5.
TβRI-TβRII-Fc protein produced by SAS cells and secreted into the extracellular space can trap endogenous TGF-βs. SAS cells were infected with lentiviruses expressing various ligand traps (Control-Fc, TβRI-Fc, TβRII-Fc, or TβRI-TβRII-Fc). A, expression of each Fc chimeric protein in SAS cells and their secretion into culture media examined by immunoblotting with anti-human IgG-Fc antibody. The same number of cells expressing each Fc chimeric protein was seeded into a 6-well plate and cultured for 48 h. The 84 μg of total proteins (cell lysate) or 50 μl of conditioned medium per lane were loaded onto the gel. The bands (black arrowheads) correspond to the expressed ligand traps secreted into the culture media (left; conditioned medium) or present in the cell lysates (right; cell lysate). α-Tubulin was used as an internal loading control. B, TβRI-TβRII-Fc protein secreted by SAS cells into culture media interacts with endogenous TGF-βs released into the extracellular space. Conditioned media from SAS cells expressing each ligand trap cultured for 48 h were used in a HEK-Blue TGF-β reporter assay as described under “Experimental procedures,” with 10 μm SB431542 used as a positive control for total inhibition of TGF-β signals. The values represent the colorimetric changes of Quanti-Blue substrate measured at 640 nm. C, effects of ligand trap expression on SAS cell proliferation. SAS cells expressing Fc chimeric proteins were grown in 12-well plates for 72 h. Cell proliferation was determined by WST-1 assay. Error bars, S.D. **, p < 0.01; ***, p < 0.001; N.S., not significant.
Next, we examined whether the ligand traps would interact with endogenous TGF-βs released from the established SAS cells. We cultured HEK-Blue TGF-β reporter cells in the conditioned medium of the SAS cells expressing Control-Fc in the absence or presence of SB431542 and confirmed that SAS cells secrete a significant level of TGF-βs whose signals are inhibited by SB431542 (Fig. 5B). Whereas TβRI-Fc or TβRII-Fc released from the established SAS cells was not able to inhibit the TGF-β signals, TGF-β signals were significantly decreased in the presence of conditioned medium derived from SAS cells expressing the TβRI-TβRII-Fc chimeric protein (Fig. 5B), suggesting that only the TβRI-TβRII-Fc chimeric protein effectively trapped endogenous TGF-β proteins. We also studied the endogenous expression of TGF-β isoforms in the SAS cells expressing various ligand traps and found that their expression was not altered (Fig. S6).
Previous reports have shown that TGF-β has an inhibitory effect on epithelial cell growth (33). Consistent with these reports, we also showed that SAS cells responded to all TGF-β isoforms and proliferated less compared with untreated cells (Fig. S7), revealing TGF-β–dependent cell growth inhibition. Thus, the overexpression of Fc chimeric proteins by SAS cells could potentially stimulate their cell growth by trapping the ligands present in the extracellular milieu. However, none of the established SAS cell lines expressing Fc chimeric proteins showed significant changes in proliferation compared with the cells expressing Control-Fc (Fig. 5C).
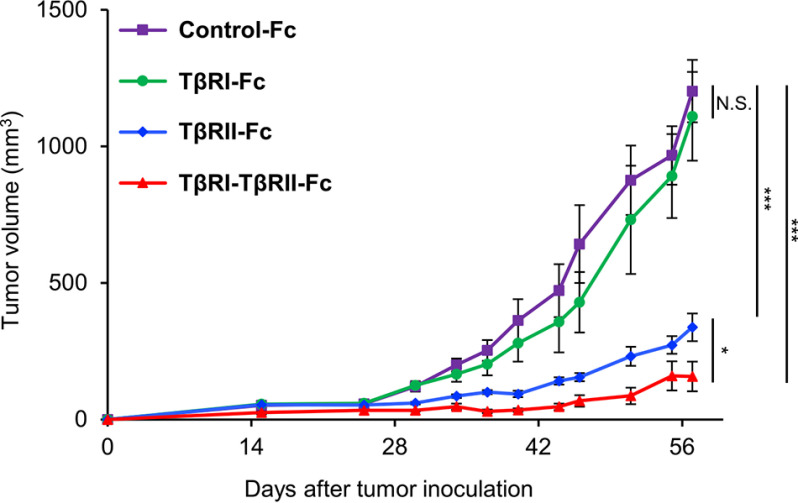
Using the SAS cells expressing the Fc chimeric proteins, we investigated their effect on the growth of oral tumors in vivo. SAS cells expressing Control-Fc, TβRI-Fc, TβRII-Fc, and TβRI-TβRII-Fc chimeric proteins were injected into the left abdominal walls of BALB/c nude mice, and the resulting tumors were allowed to grow for 8 weeks. There was no difference in tumor size between those formed from SAS cells expressing Control-Fc or TβRI-Fc. However, SAS cells expressing TβRII-Fc or TβRI-TβRII-Fc chimeric proteins formed much smaller primary tumors compared with those for other chimeric proteins (Fig. 6). Furthermore, the primary tumors that developed from SAS cells expressing TβRI-TβRII-Fc were statistically smaller compared with those formed by SAS cells expressing TβRII-Fc protein. Considering that the level of secreted TβRI-TβRII-Fc proteins was lower than that of TβRII-Fc (Fig. 5A), these results suggest that TβRI-TβRII-Fc inhibited the tumor growth most effectively via trapping TGF-βs present in the TME.
Figure 6.
TβRI-TβRII-Fc protein inhibits tumor growth in vivo. SAS cells expressing Control-Fc, TβRI-Fc, TβRII-Fc, and TβRI-TβRII-Fc were injected into the left abdominal walls of BALB/c nude mice. Tumor growth was monitored for 8 weeks. Error bars, S.E. *, p < 0.05; ***, p < 0.001; N.S., not significant.
TβRI-TβRII-Fc protein inhibits tumor angiogenesis in vivo
We next attempted to elucidate the mechanisms by which the TβRII-Fc and TβRI-TβRII-Fc chimeric proteins inhibited tumor growth. Because in vitro proliferation of SAS cells was not altered by TβRII-Fc or TβRI-TβRII-Fc (Fig. 5C), this inhibition of in vivo tumor growth did not result from the proliferative ability of cancer cells. Tumors comprise not only cancer cells but also other stromal components including tumor vessels that supply cancer cells with oxygen and nutrients and, thus, contribute to tumor growth. Because TGF-β signals have been implicated in angiogenesis (3), we examined blood vessel formation in tumors derived from SAS cells expressing each chimeric protein using antibodies against vascular endothelial markers, platelet and endothelial cell adhesion molecule 1 (PECAM-1) and CD34. As shown in Fig. 7A, we observed the formation of tumor vessels that were stained for both PECAM-1 and CD34 in the tumors formed by SAS cells expressing either Control-Fc or TβRI-Fc. Quantitative analysis of dissected tumors with antibodies recognizing PECAM-1 revealed that PECAM-1–positive areas in tumors formed by SAS cells expressing TβRI-Fc were not significantly different from that of Control-Fc (Fig. 7B). On the other hand, we observed a significant decrease in the density of tumor vessels formed by cells expressing TβRII-Fc as compared with Control-Fc (Fig. 7). Furthermore, the extent of tumor vessel formation in tumors expressing TβRI-TβRII-Fc was significantly lower than that of TβRII-Fc (Fig. 7), which is consistent with the results of tumor formation (Fig. 6). These results suggest that the ligand traps in the extracellular space may affect the angiogenic response of endothelial cells within the TME, leading to the reduction in tumor size.
Figure 7.
TβRI-TβRII-Fc protein inhibits blood vessel formation in vivo. SAS cells expressing Control-Fc, TβRI-Fc, TβRII-Fc, and TβRI-TβRII-Fc chimeric proteins were injected into the left abdominal walls of BALB/c nude mice. The tumors were excised 63 days (9 weeks) after the inoculations and examined for vascular density. A, immunostaining for PECAM-1 (red), CD34 (green), and nuclei (blue) of sections obtained from tumors derived from cells expressing Control-Fc, TβRI-Fc, TβRII-Fc, and TβRI-TβRII-Fc proteins, respectively. Scale bar, 100 μm. Negative control represents staining with secondary antibody only. B, level of angiogenesis was quantified. Values represent the PECAM-1–positive areas (%) in the observed fields. Error bars, S.E. *, p < 0.05; **, p < 0.01; N.S., not significant.
Discussion
The crucial roles of TGF-β in controlling various steps of oncogenic processes have increased the urgency to develop effective therapeutic strategies targeting TGF-β signaling. In the present study, we developed a TβRI-TβRII-Fc chimeric protein that effectively trapped all TGF-β isoforms to inhibit the TGF-β signals. We also showed that the chimeric protein inhibited the TGF-β-dependent EMT of oral cancer cells. Additionally, our in vivo approach in SAS oral cancer cells demonstrated its anti-tumor-forming ability and effects on the TME composition, suggesting that TβRI-TβRII-Fc may be a promising tool for the development of effective TME-targeted therapies.
Reports of the role of TGF-β signaling in numerous pathophysiological processes have led to increased efforts to develop effective inhibitors (21). Although several small-molecule inhibitors, including SB431542, galunisertib (LY2157299), and LY2109761, have been successfully used in in vitro and in vivo studies, their fast turnover and the potential to induce the resistance of cancer cells with prolonged drug administration (34, 35) has prompted the search for different approaches.
Engineered molecular ligand traps comprising biologically active molecules and the IgG-Fc domain have become valuable reagents for in vivo and in vitro research. Many Fc-conjugated ligands have been generated and tested in various pathological conditions. Among such Fc-chimeric drugs, etanercept, a soluble TNF-α type II receptor fused to the Fc portion of IgG1, traps TNF-α to attenuate rheumatoid arthritis (36). One of the most valuable characteristics conferred by the Fc domain in vivo is an increased plasma half-life of the protein of interest, which can improve therapeutic efficacy, an attribute that has made Fc chimeric proteins attractive biotherapeutic agents.
Previous reports in various types of preclinical models including lung and breast cancer have shown that, in a similar manner to the neutralizing antibody, soluble TGF-β type II receptor fused with Fc protein of IgG (TβRII-Fc) targets TGF-β and reduces tumor growth and metastasis (37). Of note, Qin et al. (38) constructed a trivalent TGF-β receptor trap, consisting of a single extracellular domain of betaglycan, TGF-β type III receptor (BGE), flanked by two TβRII extracellular domains (RII) and showed that the trivalent TGF-β receptor trap RII-BGE-RII (RER) effectively blocked the interaction between TGF-β and TβRII, resulting in TGF-β signaling inhibition in multiple types of prostate cancer cells as well as inhibition of early-stage tumorigenesis and tumor cell invasion in murine models of prostate cancer (38) and gynecologic cancers (39). These reports suggest that ligand traps containing the TβRII could be useful tools to inhibit TGF-β signaling in various types of cancer.
Among the three TGF-β isoforms, TGF-β2 has a 100–1,000-fold lower binding affinity to TβRII than TGF-β1 and TGF-β3 (40). This reduced affinity underlies its reduced biological activity in cells lacking betaglycan, which promotes the assembly of the TβRI-TβRII signaling complex with TGF-β2 (41). del Re and colleagues (29) reported that the biological activity of TGF-β2 was not affected by TβRII-Fc but was attenuated by the combination of two Fc-chimeras, TβRI-Fc and TβRII-Fc. However, our data showed that the combination of TβRI-Fc and TβRII-Fc did not trap TGF-β2 (Fig. 1C), which may reflect the difference in affinity of TGF-β2 to both receptors. Moreover, if TβRI-Fc and TβRII-Fc are added separately, the two chimeric proteins might have different half-lives, which is a critical factor for effective therapy. Furthermore, by having type I and II receptors in one molecule, TβRI-TβRII-Fc can trap TGF-β ligands more effectively than two independent Fc chimeric proteins. Thus, our approach to combine two receptors in a single Fc chimeric protein assures the same distributions of both receptors and increases the probability of trapping, leading to more effective therapy.
TGF-β2 is overexpressed in various cancer types, including glioma (12) and pancreatic cancer (42), and its expression is also correlated with tumor progression. Several studies have reported attempts to target TGF-β2 in tumor tissues.
TGF-β2 has been implicated not only in cancer but also in other pathological conditions. Recent reports showed that increased TGF-β2 secretion was correlated with liver- and diabetes-associated kidney fibrosis (43, 44). Therefore, TβRI-TβRII-Fc may be a promising tool for the prevention and treatment of pathological conditions caused by TGF-βs, especially TGF-β2.
Experimental procedures
Cell culture and reagents
The SAS human oral squamous carcinoma cell line was obtained from the RIKEN BioResource Center Cell Bank. HEK293T and 293FT cells were obtained from Invitrogen. HEK-Blue TGF-β reporter cells (HEK293 cell–derived TGF-β–responsive reporter cells) were purchased from InvivoGen. The cells were maintained in Dulbecco's modified Eagle's medium (DMEM; Nacalai Tesque) supplemented with 10% fetal bovine serum (Sigma), 100 units/ml penicillin, and 100 μg/ml streptomycin (Nacalai Tesque). The culture medium for 293FT cells was also supplemented with 1% minimal essential medium nonessential amino acid solution (Nacalai Tesque). TGF-β1, -β2, and -β3 were purchased from Peprotech and used at concentrations of 1, 2, or 3 ng/ml, depending on the experiments. Activin A was purchased from R&D Systems and used at a concentration of 1 ng/ml. SB431542 was obtained from WAKO Chemicals and used at concentrations of 5 or 10 μm, depending on the experiment.
Construction of chimeric proteins
The experimental procedures were approved by the Genetically Modified Organisms Safety Committee of Tokyo Medical and Dental University (registration number G2019-026C). To generate the Control-Fc construct, the Fc region of human IgG was fused to the interleukin-2 signal peptide amplified from the pFUSE-IgG1-Fc vector and inserted into the BglII and ApaI restriction sites of the pENTR201 vector (Invitrogen). For construction of the other ligand traps, TβRI-Fc and TβRII-Fc, the cDNA encoding the extracellular domain (ECD) of human TβRI and TβRII were amplified from human TβRI and TβRII cDNAs using KOD Plus Neo DNA polymerase (TOYOBO) with gene-specific primer sets. The bands corresponding to the ECDs of human TβRI ECD1-128 (384 bp) and TβRII ECD1-184 (486 bp) were digested and ligated in-frame into pENTR201-IgG1 Fc to produce pENTR201-TβRI-IgG1Fc and pENTR201-TβRII-IgG1Fc, respectively. The primers used for amplification were as follows: hTβRIFc-BamHI-F 5′-GGATCCGGGACCATGGAGGCGGCGGTCGCT-3′ (forward), hTβRIFc-EcoRV-R 5′-GATATCAGCTGCCAGTTCCACAGGACCAAG-3′ (reverse), hTβRIIFc-BamHI-F 5′-GGATCCTCTGCCATGGGTCGGGGGCTGCTCA-3′ (forward), and hTβRIIFc-EcoRV-R 5′-GATATCGTCAGGATTGCTGGTGTTATATTC-3′ (reverse). To generate the TβRI-TβRII-Fc protein, the ECD of TβRII lacking a signal peptide sequence (ECD23-184) was amplified with KOD Plus Neo DNA polymerase using the following set of primers: hTβRIIFc-EcoRV-F 5′-GATATCACGATCCCACCGCACGTTCAGAAG-3′ (forward) and hTβRIIFc-BglII-R 5′-AGATCTGTCAGGATTGCTGGTGTTATATTC-3′ (reverse). The amplified product (552 bp) was digested and inserted in-frame between the EcoRV and BglII restriction sites of pENTR201-TβRI-IgG1Fc to produce the pENTR201-TβRI-TβRII-IgG1Fc vector. Each construct was then sequenced on both strands to confirm the fidelity of the subcloned cDNAs. Gateway Technology (Thermo Fisher Scientific) was used to generate lentiviral vectors encoding Control-Fc, TβRI-Fc, TβRII-Fc, and TβRI-TβRII-Fc, respectively. Control-Fc, TβRI-Fc, TβRII-Fc and TβRI-TβRII-Fc cDNAs subcloned into pENTR201 vectors were then transferred into the pCSII-EF-RfA lentiviral expression vector (a gift from Dr. Hiroyuki Miyoshi, Keio University) by the LR recombination reaction (Thermo Fisher Scientific) and used for either HEK293T cell transfection to produce recombinant Fc proteins or lentiviral particle preparation, as described below.
Production of Fc chimeric proteins
HEK293T cells (9.0 × 106) were seeded onto 10-cm tissue culture dishes and transfected with pCSII-EF-RfA vectors expressing each Fc chimeric protein using Lipofectamine 2000 (Invitrogen) according to the manufacturer's protocol. Four hours post-transfection, the medium was replaced with serum-free Opti-MEM (Gibco), and the cells were incubated for 48 h to allow sufficient accumulation of Fc chimeric proteins. The concentrations of each Fc chimeric protein in the conditioned medium were determined by ELISA with the Human IgG ELISA Quantitation Set (Bethyl Laboratories) according to the manufacturer's protocol. Two batches of conditioned media were prepared to use for downstream applications; batch-1 (Control-Fc, 211.17 μg/ml; TβRI-Fc, 5.13 μg/ml; TβRII-Fc, 11.87 μg/ml; and TβRI-TβRII-Fc, 1.39 μg/ml) and batch-2 (Control-Fc, 101.75 μg/ml; TβRI-Fc, 9.25 μg/ml; TβRII-Fc, 15.37 μg/ml; and TβRI-TβRII-Fc, 2.97 μg/ml). The collected conditioned media were aliquoted and stored at −80 °C until use.
Determination of Fc chimeric protein-trapping activity
The 96-well plates were coated for 4 h at 4 °C with a 10 nm concentration of each TGF-β ligand (TGF-β1, -β2, or -β3) solution in 10 mm NaHCO3. The wells were extensively washed with PBS, 0.01% Tween 20 and masked with Bullet Blocking One Reagent (Nacalai Tesque) for 10 min. The 1 μm recombinant Control-Fc, TβRI-Fc, TβRII-Fc, or TβRI-TβRII-Fc solutions were added to each immobilized ligand and incubated overnight at 4 °C. The wells were washed with PBS, 0.01% Tween 20, followed by the addition of horseradish peroxidase–conjugated polyclonal rabbit anti-goat antibody (Dako, P0160, 1:2000 dilution), which is cross-reactive to human IgG, and incubation for 4 h at 4 °C. Effective trapping of Fc chimeric proteins was visualized with one-step Ultra TMB-ELISA Reagent (Thermo Fisher Scientific; 50 μl/well). The reaction was stopped by the addition of an equal volume of 2 m H2SO4. The amount of Fc chimeric proteins trapped by each immobilized ligand was quantified at 450-620 nm using a microplate reader (Bio-Rad, model 680).
Smad2/3/4-responsive reporter assay (HEK-Blue TGF-β reporter assay)
The effect of Fc chimeric proteins on TGF-β/Smad signal activation by TGF-β family members (TGF-β1, -β2, and -β3 and activin A) was determined using HEK-Blue TGF-β reporter cells seeded into 96-well plates (1.0 × 105 cells/well) and incubated overnight at 37 °C in 5% CO2. The next day, the medium was replaced with 200 μl of serum-free DMEM, and the cells were incubated for 3 h. Equal volumes of preformed Fc chimeric protein/ligand complexes were then added to the cells, followed by incubation for 24 h. The Fc chimeric protein/ligand complexes were formed as follows. HEK293T-conditioned medium corresponding to 50 ng of Control-Fc, TβRI-Fc, TβRII-Fc, or TβRI-TβRII-Fc protein (calculated from total concentration assayed with ELISA) was mixed with Opti-MEM to make a total volume of 100 μl, followed by the addition of TGF-β family ligand (1 ng/ml TGF-β1, -β2, or -β3 or 1 ng/ml activin A) and incubation for 2 h at 37 °C. A mixture of SB431542 and ligands in serum-free DMEM was used as a positive control for inhibition of TGF-β signals. The activation of TGF-β/Smad signals was detected using the QUANTI-Blue substrate (InvivoGen) following incubation for 30 min at 37 °C. The colorimetric change of the substrate by the released SEAP was quantified at 640 nm using a microplate reader.
To analyze the trapping efficiency of the culture media derived from SAS cells expressing Fc chimeric proteins, SAS cells were incubated in serum-free DMEM for 48 h. Next, 100 μl of conditioned medium was added to HEK-Blue TGF-β reporter cells seeded into 96-well plates, and the cells were starved in serum-free DMEM for 3 h. The cells were then incubated for an additional 24 h, and activated TGF-β/Smad signals were detected as described above.
Treatment of OSCC cell lines with Fc chimeric proteins
SAS or HSC-4 cells were seeded into 12-well plates and cultured overnight at 37 °C in 5% CO2. The next day, the medium was replaced with 1 ml of serum-free Opti-MEM. The Fc chimeric protein/ligand complexes were prepared by mixing the conditioned medium from HEK293T cells expressing Fc chimeric proteins (500 ng of Fc chimeric proteins in 500 μl of Opti-MEM) with TGF-β1, -β2, or -β3, respectively, and incubated for 2 h at 37 °C to allow efficient formation of the ligand/Fc chimeric protein complexes. A mixture of SB431542 and ligands in serum-free DMEM was used as a positive control for TGF-β signal inhibition. The preformed ligand/Fc chimeric protein complexes were then added to SAS or HSC-4 cells. The cells were then incubated with ligand/Fc chimeric protein complexes for 4 or 72 h (depending on the experiment) and subjected to gene expression analysis by qRT-PCR or immunocytochemistry.
RNA isolation and quantitative RT-PCR
Total RNA was isolated using Sepasol RNA I Super G Reagent (Nacalai Tesque) and reverse-transcribed using ReverTraAce qPCR RT Master Mix (TOYOBO). The qRT-PCR analysis was performed using SYBR Green (Roche Applied Science) on a Step One Plus real-time PCR system (Applied Biosystems). All expression data were normalized to the expression of β-actin. The genes and corresponding primer sequences are listed in Table S1.
Immunoblotting analysis
Immunoblotting analysis was performed as described previously (45). Antibodies to E-cadherin (Cell Signaling Technology, 14472, 1:1000 dilution), claudin-1 (Abcam, ab15098, 1:2000 dilution), fibronectin (BD Biosciences, 610077, 1:5000 dilution), vimentin (Abcam, ab92547, 1:1000 dilution), SM22α (Abcam, ab14106, 1:2000 dilution), human IgG-Fc fragment (Bethyl Laboratories, A80-105, 1:5000 dilution), and α-tubulin (Abcam, ab4074, 1:10,000 dilution) were used for the detection of separated proteins. Binding of specific antibodies was detected using ECL detection reagent (GE Healthcare) and visualized with a Fusion Solo S imaging system (Vilber).
Immunocytochemistry
SAS or HSC-4 cells were seeded on coverglasses placed into 12-well tissue culture plates and treated with TGF-β1, -β2, or -β3 in the presence or absence of Fc chimeric proteins for 72 h. The cells were then fixed with an ice-cold mixture of methanol/acetone (1:1) for 20 s on ice and blocked with 1% BSA, followed by incubation with anti-E-cadherin (Cell Signaling Technology, 3195, 1:200 dilution) and anti-vimentin (Abcam, ab92547, 1:400 dilution) antibodies. The proteins were visualized with anti-mouse IgG (H + L) Alexa Fluor 488–conjugated (Thermo Fisher Scientific, A-11001) and anti-rabbit IgG (H + L) Alexa Fluor 594–conjugated (Thermo Fisher Scientific, A-21207) secondary antibodies. Hoechst33342 (Cell Signaling Technology) was used for nuclear staining. Images were obtained with a TCS SP8 confocal microscope (Leica Microsystems GmbH).
Lentivirus production and SAS cell infection
Lentivirus particles were produced by transfecting 293FT cells with the expression plasmids (pCSII-EF-RfA-Control-Fc, pCSII-EF-RfA-TβRI-Fc, pCSII-EF-RfA-TβRII-Fc, or pCSII-EF-RfA-TβRI-TβRII-Fc) and packaging plasmids (pCMV-VSV-G-RSV-Rev and pCAG-HIVgp) using Lipofectamine 2000 (Thermo Fisher Scientific). The viral supernatants were collected 48 h after transfection and concentrated for 7 days at 4 °C using a Lenti-X Concentrator (Takara Bio). The concentrated lentiviral particles were used to infect 1.2 × 105 SAS cells/well in 12-well tissue culture plates.
Cell proliferation assay
Cells were seeded into 12-well culture plates and grown for the indicated period followed by WST-1 cell proliferation assay (Dojindo) according to the manufacturer's protocol. The experiments were performed in duplicates.
Subcutaneous xenograft model
The animal experimental procedures were approved by the Institutional Animal Care and Use Committee of Tokyo Medical and Dental University (registration number A2019-251C2) and performed according to the guidelines of the Animal Care Standards of Tokyo Medical and Dental University. SAS cells (1 × 103 cells in 100 μl of Matrigel; Corning) expressing the Fc chimeric proteins ; SAS-Control-Fc, SAS-TβRI-Fc, SAS-TβRII-Fc, and SAS-TβRI-TβRII-Fc were inoculated subcutaneously into the left abdominal walls of BALB/c 6-week-old male immunodeficient nude mice. Tumor growth was assessed at the indicated times by caliper measurements and calculated from the minor axis and major radius using the formula, V = L × W2 × 0.5, with V corresponding to tumor volume (mm3), L to the length of the tumor (mm), and W to the width of the tumor (mm).
Immunohistochemistry
Immunofluorescent staining of human oral cancer xenografts was performed as described previously (46). Tumors formed in BALB/c nude mice grafted with SAS-Control-Fc cells, SAS-TβRI-Fc cells, SAS-TβRII-Fc cells, or SAS-TβRI-TβRII-Fc cells were harvested on day 63. The tumors were excised, embedded in OCT compound, and snap-frozen in a dry ice/acetone-hexane bath. To evaluate the formation of blood vessels in the xenografted tumors, the frozen tumor xenograft samples were sectioned at a thickness of 5 μm using a cryostat CMV 1950 (Leica Microsystems GmbH), fixed with 4% paraformaldehyde, blocked with 1% BSA, and incubated with anti-PECAM-1 antibody (BioLegend, 102502, 1:400) and anti-CD34 antibody (EP373Y) (Abcam, ab81289, 1: 400) overnight at 4 °C. The samples were then incubated with secondary antibodies (anti-rat IgG (H + L) Alexa Fluor 594–conjugated secondary antibodies (Thermo Fisher Scientific A21209) to visualize PECAM-1 and anti-rabbit IgG (H + L) Alexa Fluor 488–conjugated secondary antibodies (Thermo Fisher Scientific A21206) to detect CD34) and Hoechst 33342 (Cell Signaling Technology) for 1 h before visualizing the stained molecules. Images (94 fields of view for tumor xenografts from mice inoculated with SAS cells expressing Control-Fc, 60 fields of view for TβRI-Fc xenografts, 64 fields of view for TβRII-Fc xenografts, and 43 fields of view for TβRI-TβRII-Fc xenografts) were obtained using an All-in-One fluorescent microscope, BZ-X710 (Keyence) with a ×20 objective lens. ImageJ software (National Institutes of Health) was used for the quantification of PECAM-1–positive vessels. All images were converted into binary (black/white) images, and black areas (corresponding to vessels positive for PECAM-1) were quantified. The values represented the percentage of PECAM-1–positive area in each observed field.
Statistical analyses
Values are presented as means ± S.D. or S.E. Significant differences between means were determined using one-tailed unpaired Student's t test. Differences between means were considered statistically significant for p values <0.05.
Data availability
All data described are contained within the article.
Supplementary Material
Acknowledgments
We thank Dr. Hiroyuki Miyoshi for the lentiviral vectors, Maki Saito for technical assistance, and the members of the Department of Molecular Pathology of the University of Tokyo and the Department of Biochemistry of Tokyo Medical and Dental University (TMDU) for technical assistance and critical discussion.
This article contains supporting information.
Author contributions—K. T., Y. A., T. M., H. T., Y. Y., D. K., and M. S. data curation; K. T., Y. A., and T. W. formal analysis; K. T., Y. A., K. A. P.-I., T. M., H. T., D. K., M. S., and T. W. validation; K. T., Y. A., K. A. P.-I., T. M., H. T., D. K., M. S., K. M., and T. W. investigation; K. T., H. T., and T. W. visualization; K. T., K. M., and T. W. methodology; K. T., K. A. P.-I., and T. W. writing-original draft; K. T., Y. A., K. A. P.-I., T. M., H. T., Y. Y., D. K., M. S., K. M., and T. W. writing-review and editing; K. A. P.-I., Y. Y., K. M., and T. W. supervision; K. A. P.-I., Y. Y., and T. W. funding acquisition; K. T., K. A. P.-I., Y. Y., and T. W. project administration; K. M. and T. W. conceptualization; T. W. resources.
Funding and additional information—This work was supported by Project for Cancer Research and Therapeutic Evolution (P-CREATE), Japan Agency for Medical Research and Development (AMED) Grant 17cm0106613h0001 (to T. W.); Grant-in-Aid for Young Scientists (B) 15K21394 (to Y. Y.) and Grants-in-Aid for Scientific Research (C) 17K11828 (to K. A. P.-I.) and 17K07157 (to Y. Y.) from the Japan Society for the Promotion of Science (JSPS); and the Project for Promoting Leading-edge Research in Oral Science at Tokyo Medical and Dental University (TMDU) (to K. T., K. A. P. -I., Y. Y., and T. W.).
Conflict of interest—Y. A. is an employee of Nippon Kayaku, Co., Ltd.
- TME
- tumor microenvironment
- TGF-β
- transforming growth factor-β
- TβRI
- TGF-β type I receptor
- TβRII
- TGF-β type II receptor
- EMT
- epithelial-to-mesenchymal transition
- OSCC
- oral squamous cell carcinoma
- SEAP
- secreted alkaline phosphatase
- TMEPAI
- transmembrane prostate androgen–induced protein
- qRT-PCR
- quantitative RT-PCR
- PAI-1
- plasminogen activator inhibitor-1
- PECAM-1
- platelet and endothelial cell adhesion molecule 1
- DMEM
- Dulbecco's modified Eagle's medium
- ECD
- extracellular domain.
References
- 1. Hanahan D., and Weinberg R. A. (2011) Hallmarks of cancer: the next generation. Cell 144, 646–674 10.1016/j.cell.2011.02.013 [DOI] [PubMed] [Google Scholar]
- 2. Morikawa M., Derynck R., and Miyazono K. (2016) TGF-β and the TGF-β family: context-dependent roles in cell and tissue physiology. Cold Spring Harb. Perspect. Biol. 8, a021873 10.1101/cshperspect.a021873 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Miyazono K., Katsuno Y., Koinuma D., Ehata S., and Morikawa M. (2018) Intracellular and extracellular TGF-β signaling in cancer: some recent topics. Front. Med. 12, 387–411 10.1007/s11684-018-0646-8 [DOI] [PubMed] [Google Scholar]
- 4. Heldin C.-H., and Moustakas A. (2016) Signaling receptors for TGF-β family members. Cold Spring Harb. Perspect. Biol. 8, a022053 10.1101/cshperspect.a022053 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Derynck R., and Budi E. H. (2019) Specificity, versatility, and control of TGF-β family signaling. Sci. Signal. 12, eaav5183 10.1126/scisignal.aav5183 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Baardsnes J., Hinck C. S., Hinck A. P., and O'Connor-McCourt M. D. (2009) TβR-II discriminates the high- and low-affinity TGF-β isoforms via two hydrogen-bonded ion pairs. Biochemistry 48, 2146–2155 10.1021/bi8019004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Ehrlich M., Gutman O., Knaus P., and Henis Y. I. (2012) Oligomeric interactions of TGF-β and BMP receptors. FEBS Lett. 586, 1885–1896 10.1016/j.febslet.2012.01.040 [DOI] [PubMed] [Google Scholar]
- 8. Tang Y. A., Chen Y. F., Bao Y., Mahara S., Yatim S. M. J. M., Oguz G., Lee P. L., Feng M., Cai Y., Tan E. Y., Fong S. S., Yang Z. H., Lan P., Wu X. J., and Yu Q. (2018) Hypoxic tumor microenvironment activates GLI2 via HIF-1α and TGF-β2 to promote chemoresistance in colorectal cancer. Proc. Natl. Acad. Sci. U. S. A. 115, E5990–E5999 10.1073/pnas.1801348115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Hachim M. Y., Hachim I. Y., Dai M., Ali S., and Lebrun J. J. (2018) Differential expression of TGFβ isoforms in breast cancer highlights different roles during breast cancer progression. Tumour Biol. 40, 1010428317748254 10.1177/1010428317748254 [DOI] [PubMed] [Google Scholar]
- 10. Seystahl K., Tritschler I., Szabo E., Tabatabai G., and Weller M. (2015) Differential regulation of TGF-β-induced, ALK-5-mediated VEGF release by SMAD2/3 versus SMAD1/5/8 signaling in glioblastoma. Neuro Oncol. 17, 254–265 10.1093/neuonc/nou218 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Riemenschneider M. J., Hirblinger M., Vollmann-Zwerenz A., Hau P., Proescholdt M. A., Jaschinski F., Rothhammer-Hampl T., Wosikowski K., Janicot M., and Leo E. (2015) TGF-β isoforms in cancer: immunohistochemical expression and Smad-pathway-activity-analysis in thirteen major tumor types with a critical appraisal of antibody specificity and immunohistochemistry assay validity. Oncotarget 6, 26770–26781 10.18632/oncotarget.5780 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Kjellman C., Olofsson S. P., Hansson O., Von Schantz T., Lindvall M., Nilsson I., Salford L. G., Sjögren H. O., and Widegren B. (2000) Expression of TGF-β isoforms, TGF-β receptors, and SMAD molecules at different stages of human glioma. Int. J. Cancer 89, 251–258 10.1002/1097-0215(20000520)89:3<251::AID-IJC7>3.0.CO;2-5 [DOI] [PubMed] [Google Scholar]
- 13. Kim K. H., Back J. H., Zhu Y., Arbesman J., Athar M., Kopelovich L., Kim A. L., and Bickers D. R. (2011) Resveratrol targets transforming growth factor-β2 signaling to block UV-induced tumor progression. J. Invest. Dermatol. 131, 195–202 10.1038/jid.2010.250 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Serizawa M., Murakami H., Watanabe M., Takahashi T., Yamamoto N., and Koh Y. (2014) Peroxisome proliferator-activated receptor γ agonist efatutazone impairs transforming growth factor-β2-induced motility of epidermal growth factor receptor tyrosine kinase inhibitor-resistant lung cancer cells. Cancer Sci. 105, 683–689 10.1111/cas.12411 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Nagy A., Lánczky A., Menyhárt O., and Győrffy B. (2018) Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci. Rep. 8, 9227 10.1038/s41598-018-27521-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Riviera C. (2015) Essentials of oral cancer. Int. J. Clin. Exp. Pathol. 8, 11884–11894 10.5281/zenodo.192487 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Nakamura R., Kayamori K., Oue E., Sakamoto K., Harada K., and Yamaguchi A. (2015) Transforming growth factor-β synthesized by stromal cells and cancer cells participates in bone resorption induced by oral squamous cell carcinoma. Biochem. Biophys. Res. Commun. 458, 777–782 10.1016/j.bbrc.2015.02.013 [DOI] [PubMed] [Google Scholar]
- 18. Takahashi K., Podyma-Inoue K. A., Takao C., Yoshimatsu Y., Muramatsu T., Inazawa J., and Watabe T. (2018) Regulatory role of transforming growth factor-β signals in the migration and tumor formation of HOC313-LM cells, an oral squamous cell carcinoma. J. Stomatological Soc. Jpn. 85, 52–61 [Google Scholar]
- 19. Nieto M. A., Huang R. Y., Jackson R. A., and Thiery J. P. (2016) EMT: 2016. Cell 166, 21–45 10.1016/j.cell.2016.06.028 [DOI] [PubMed] [Google Scholar]
- 20. Moustakas A., and Heldin C.-H. (2016) Mechanisms of TGFβ-induced epithelial-mesenchymal transition. J. Clin. Med. 5, 63 10.3390/jcm5070063 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Huynh L. K., Hipolito C. J., and ten Dijke P. (2019) A perspective on the development of TGF-β inhibitors for cancer treatment. Biomolecules 9, E743 10.3390/biom9110743 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Morris J. C., Tan A. R., Olencki T. E., Shapiro G. I., Dezube B. J., Reiss M., Hsu F. J., Berzofsky J. A., and Lawrence D. P. (2014) Phase I study of GC1008 (Fresolimumab): a human anti-transforming growth factor-β (TGFβ) monoclonal antibody in patients with advanced malignant melanoma or renal cell carcinoma. PLoS ONE 9, e90353 10.1371/journal.pone.0090353 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Formenti S. C., Lee P., Adams S., Goldberg J. D., Li X., Xie M. W., Ratikan J. A., Felix C., Hwang L., Faull K. F., Sayre J. W., Hurvitz S., Glaspy J. A., Comin-Anduix B., Demaria S., et al. (2018) Focal irradiation and systemic TGFβ blockade in metastatic breast cancer. Clin. Cancer Res. 24, 2493–2504 10.1158/1078-0432.CCR-17-3322 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Yingling J. M., McMillen W. T., Yan L., Huang H., Sawyer J., Graff J., Clawson D. K., Britt K. S., Anderson B. D., Beight D. W., Desaiah D., Lahn M. M., Benhadji K. A., Lallena M. J., Holmgaard R. B., et al. (2018) Preclinical assessment of galunisertib (LY2157299 monohydrate), a first-in-class transforming growth factor-β receptor type I inhibitor. Oncotarget 9, 6659–6677 10.18632/oncotarget.23795 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Giaccone G., Bazhenova L. A., Nemunaitis J., Tan M., Juhász E., Ramlau R., van den Heuvel M. M., Lal R., Kloecker G. H., Eaton K. D., Chu Q., Dunlop D. J., Jain M., Garon E. B., Davis C. S., et al. (2015) A phase III study of belagenpumatucel-L, an allogeneic tumour cell vaccine, as maintenance therapy for non-small cell lung cancer. Eur. J. Cancer 51, 2321–2329 10.1016/j.ejca.2015.07.035 [DOI] [PubMed] [Google Scholar]
- 26. Colak S., and ten Dijke P. (2017) Targeting TGF-β signaling in cancer. Trends Cancer 3, 56–71 10.1016/j.trecan.2016.11.008 [DOI] [PubMed] [Google Scholar]
- 27. Yung L. M., Nikolic I., Paskin-Flerlage S. D., Pearsall R. S., Kumar R., and Yu P. B. (2016) A selective transforming growth factor-β ligand trap attenuates pulmonary hypertension. Am. J. Respir. Crit. Care Med. 194, 1140–1151 10.1164/rccm.201510-1955OC [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Zhou X., Wang J. L., Lu J., Song Y., Kwak K. S., Jiao Q., Rosenfeld R., Chen Q., Boone T., Simonet W. S., Lacey D. L., Goldberg A. L., and Han H. Q. (2010) Reversal of cancer cachexia and muscle wasting by ActRIIB antagonism leads to prolonged survival. Cell 142, 531–543 10.1016/j.cell.2010.07.011 [DOI] [PubMed] [Google Scholar]
- 29. del Re E., Babitt J. L., Pirani A., Schneyer A. L., and Lin H. Y. (2004) In the absence of type III receptor, the transforming growth factor (TGF)-β type II-B receptor requires the type I receptor to bind TGF-β2. J. Biol. Chem. 279, 22765–22772 10.1074/jbc.M401350200 [DOI] [PubMed] [Google Scholar]
- 30. Katsuno Y., Lamouille S., and Derynck R. (2013) TGF-β signaling and epithelial-mesenchymal transition in cancer progression. Curr. Opin. Oncol. 25, 76–84 10.1097/CCO.0b013e32835b6371 [DOI] [PubMed] [Google Scholar]
- 31. Lamouille S., and Derynck R. (2007) Cell size and invasion in TGF-β-induced epithelial to mesenchymal transition is regulated by activation of the mTOR pathway. J. Cell Biol. 178, 437–451 10.1083/jcb.200611146 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Miyazono K., Olofsson A., Colosetti P., and Heldin C.-H. (1991) A role of the latent TGF-β1-binding protein in the assembly and secretion of TGF-β1. EMBO J. 10, 1091–1101 10.1002/j.1460-2075.1991.tb08049.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Zhang Y., Alexander P. B., and Wang X. F. (2017) TGF-β family signaling in the control of cell proliferation and survival. Cold Spring Harb. Perspect. Biol. 9, a022145 10.1101/cshperspect.a022145 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Serova M., Tijeras-Raballand A., Dos Santos C., Albuquerque M., Paradis V., Neuzillet C., Benhadji K. A., Raymond E., Faivre S., and de Gramont A. (2015) Effects of TGF-β signalling inhibition with galunisertib (LY2157299) in hepatocellular carcinoma models and in ex vivo whole tumor tissue samples from patients. Oncotarget 6, 21614–21627 10.18632/oncotarget.4308 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Connolly E. C., Saunier E. F., Quigley D., Luu M. T., De Sapio A., Hann B., Yingling J. M., and Akhurst R. J. (2011) Outgrowth of drug-resistant carcinomas expressing markers of tumor aggression after long-term TβRI/II kinase inhibition with LY2109761. Cancer Res. 71, 2339–2349 10.1158/0008-5472.CAN-10-2941 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Weinblatt M. E., Kremer J. M., Bankhurst A. D., Bulpitt K. J., Fleischmann R. M., Fox R. I., Jackson C. G., Lange M., and Burge D. J. (1999) A trial of etanercept, a recombinant tumor necrosis factor receptor:Fc fusion protein, in patients with rheumatoid arthritis receiving methotrexate. N. Engl. J. Med. 340, 253–259 10.1056/NEJM199901283400401 [DOI] [PubMed] [Google Scholar]
- 37. Muraoka R. S., Dumont N., Ritter C. A., Dugger T. C., Brantley D. M., Chen J., Easterly E., Roebuck L. R., Ryan S., Gotwals P. J., Koteliansky V., and Arteaga C. L. (2002) Blockade of TGF-β inhibits mammary tumor cell viability, migration, and metastases. J. Clin. Invest. 109, 1551–1559 10.1172/JCI0215234 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Qin T., Barron L., Xia L., Huang H., Villarreal M. M., Zwaagstra J., Collins C., Yang J., Zwieb C., Kodali R., Hinck C. S., Kim S. K., Reddick R. L., Shu C., O'Connor-McCourt M. D., et al. (2016) A novel highly potent trivalent TGF-β receptor trap inhibits early-stage tumorigenesis and tumor cell invasion in murine Pten-deficient prostate glands. Oncotarget 7, 86087–86102 10.18632/oncotarget.13343 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Zhu H., Gu X., Xia L., Zhou Y., Bouamar H., Yang J., Ding X., Zwieb C., Zhang J., Hinck A. P., Sun L. Z., and Zhu X. (2018) A novel TGFβ trap blocks chemotherapeutics-induced TGFβ1 signaling and enhances their anticancer activity in gynecologic cancers. Clin. Cancer Res. 24, 2780–2793 10.1158/1078-0432.CCR-17-3112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Cheifetz S., Hernandez H., Laiho M., ten Dijke P., Iwata K. K., and Massagué J. (1990) Distinct transforming growth factor-β (TGF-β) receptor subsets as determinants of cellular responsiveness to three TGF-β isoforms. J. Biol. Chem. 265, 20533–20538 [PubMed] [Google Scholar]
- 41. López-Casillas F., Wrana J. L., and Massagué J. (1993) Betaglycan presents ligand to the TGFβ signaling receptor. Cell 73, 1435–1444 10.1016/0092-8674(93)90368-Z [DOI] [PubMed] [Google Scholar]
- 42. Friess H., Yamanaka Y., Büchler M., Ebert M., Beger H. G., Gold L. I., and Korc M. (1993) Enhanced expression of transforming growth factor β isoforms in pancreatic cancer correlates with decreased survival. Gastroenterology 105, 1846–1856 10.1016/0016-5085(93)91084-U [DOI] [PubMed] [Google Scholar]
- 43. Hill C., Flyvbjerg A., Rasch R., Bak M., and Logan A. (2001) Transforming growth factor-β2 antibody attenuates fibrosis in the experimental diabetic rat kidney. J. Endocrinol. 170, 647–651 10.1677/joe.0.1700647 [DOI] [PubMed] [Google Scholar]
- 44. Zepeda-Morales A. S., Del Toro-Arreola S., García-Benavides L., Bastidas-Ramírez B. E., Fafutis-Morris M., Pereira-Suárez A. L., and Bueno-Topete M. R. (2016) Liver fibrosis in bile duct-ligated rats correlates with increased hepatic IL-17 and TGF-β2 expression. Ann. Hepatol. 15, 418–426 10.5604/16652681.1198820 [DOI] [PubMed] [Google Scholar]
- 45. Goto K., Kamiya Y., Imamura T., Miyazono K., and Miyazawa K. (2007) Selective inhibitory effects of Smad6 on bone morphogenetic protein type I receptors. J. Biol. Chem. 282, 20603–20611 10.1074/jbc.M702100200 [DOI] [PubMed] [Google Scholar]
- 46. Suzuki Y., Ohga N., Morishita Y., Hida K., Miyazono K., and Watabe T. (2010) BMP-9 induces proliferation of multiple types of endothelial cells in vitro and in vivo. J. Cell Sci. 123, 1684–1692 10.1242/jcs.061556 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data described are contained within the article.