Abstract
This meeting report from the XV Banff conference describes the creation of a multiorgan transplant gene panel by the Banff Molecular Diagnostics Working Group (MDWG). This Banff Human Organ Transplant (B‐HOT) panel is the culmination of previous work by the MDWG to identify a broadly useful gene panel based on whole transcriptome technology. A data‐driven process distilled a gene list from peer‐reviewed comprehensive microarray studies that discovered and validated their use in kidney, liver, heart, and lung transplant biopsies. These were supplemented by genes that define relevant cellular pathways and cell types plus 12 reference genes used for normalization. The 770 gene B‐HOT panel includes the most pertinent genes related to rejection, tolerance, viral infections, and innate and adaptive immune responses. This commercially available panel uses the NanoString platform, which can quantitate transcripts from formalin‐fixed paraffin‐embedded samples. The B‐HOT panel will facilitate multicenter collaborative clinical research using archival samples and permit the development of an open source large database of standardized analyses, thereby expediting clinical validation studies. The MDWG believes that a pathogenesis and pathway based molecular approach will be valuable for investigators and promote therapeutic decision‐making and clinical trials.
Keywords: biomarker, biopsy, classification systems: Banff classification, clinical research/practice, diagnostic techniques and imaging, pathology/histopathology
Short abstract
This Banff meeting report summarizes the progress of the Banff Molecular Diagnostics Working Group, which generated consensus on a transplant‐specific discovery gene panel and a potential roadmap for its validation for diagnostic application.
Abbreviations
- ABMR
antibody‐mediated rejection
- B‐HOT
Banff Human Organ Transplant
- CLIA
Clinical Laboratory Improvement Amendments
- DIP
data integration platform
- DSA
donor specific antibody
- FFPE
formalin fixed, paraffin embedded
- MDWG
Molecular Diagnostics Working Group
- TCMR
T cell–mediated rejection
1. INTRODUCTION
The XV Banff Conference for Allograft Pathology was held on September 23‐27, 2019, in Pittsburgh,Pennsylvania. One main topic, continuing a theme from two previous Banff meetings, was to include applications of molecular techniques for transplant biopsies and to articulate a roadmap for the clinical adoption of molecular transplant diagnostics for allograft biopsies. 1 This meeting report summarizes the progress made by the Banff Molecular Diagnostics Working Group (MDWG) and the resulting next steps from the 2019 conference.
2. CHALLENGES IN MOLECULAR TRANSPLANT DIAGNOSTICS
The MDWG identified several challenges in the clinical application of molecular diagnostics. Different assays that measure different sets of genes validated for slightly different clinical contexts create a major analytical challenge. Enrolling patients into multicenter molecular diagnostic trials becomes problematic if local molecular diagnostic tests and risk stratification are done by noncomparable assays. The lack of a diagnostic gold standard for clinical validation of new molecular diagnostics requires multicenter standardization and independent validation in prospective randomized trials. Clinical and pathologic indications for molecular testing need to be defined and validated. Molecular tests must be cost effective to increase diagnostic utility beyond histopathology. For useful molecular diagnostics turnaround time needs to match immediate clinical needs. The integration of molecular tests with other diagnostic and clinical information requires standardization to make diagnosis and risk stratification comparable between centers. Industry partnerships are needed to advance the field, but transparency and appropriate disclosure of potential conflicts of interest are paramount. The MDWG believes that the present report shows a pathway that can address many of these issues.
3. EVOLUTION OF MOLECULAR TRANSPLANT DIAGNOSTICS
Over the past 20 years, we estimate that more than 4000 organ transplant biopsies have been studied by whole transcriptome microarrays. 2 These have been conducted independently by several research groups, covering transplant biopsies of kidneys 3 , 4 , 5 , 6 , 7 and, to a lesser extent, other organs. 8 , 9 , 10 , 11 , 12 , 13 Different analytical approaches addressing relevant research questions from these data have been made available and reproduced by several research groups and transplant centers, covering a broad spectrum of phenotypes and patient demographics. 14 These studies led to potential diagnostic applications as well as major novel mechanistic insights with changes to the Banff classification, for example, the adoption of C4d‐negative antibody‐mediated rejection (ABMR) and chronic‐active T cell–mediated rejection (TCMR) as new diagnostic categories. 3 , 14 , 15 Using transcriptome arrays the molecular phenotype in renal allografts correlates well with relevant rejection clinical entities and phenotypes. 2 , 16 In liver transplantation, microarray studies confirmed that liver biopsies with TCMR share very similar transcriptional phenotypes with those in renal allograft biopsies. 12 , 13 Transcriptional similarities are also present in heart and lung allograft biopsies. 8 , 9 , 10 , 11 These publications show that groups of genes within certain molecular pathways are statistically significantly associated with specific Banff histological lesions, rejection phenotypes, and Banff diagnostic categories. Transcript analysis also reveals potentially important underlying heterogeneities not perceived by pathology alone within diagnostic groups. 17
In 2013 molecular diagnostics were added as an aspirational goal to the Banff classification. 15 The molecular quantification of endothelial cell associated transcripts and classifier‐based prediction of donor specific antibody‐mediated tissue injury were adopted as diagnostic features/lesions equivalent to C4d for the diagnosis of ABMR. This was noted to be a forward‐looking proposal at the time, because there was no consensus around which endothelial genes should be quantified and no independent multi‐institutional validation for any diagnostic classifier or gene set. The main impetus in 2013 to adopt a molecular diagnostic option into the classification, despite these limitations, was to set the future direction for the Banff classification and to promote collaborative and multi‐institutional, open source efforts to advance the field by validating, standardizing, and making molecular transplant diagnostics accessible to the broad transplant community. This is a foundational value of the Banff consortium. 18
At the 2015 meeting, the Banff MDWG recommended the creation of molecular consensus gene sets as classifiers derived from the overlap between published and reproduced gene lists that associate with the main clinical phenotypes of TCMR and ABMR. 1 Similar roadmaps and processes for clinical adoption have been reviewed extensively and proposed by other key opinion leaders in the field. 19 , 20 , 21 , 22 Collaborative multicenter studies were proposed to close identified knowledge gaps and enable practical molecular diagnostic incorporation into diagnostic classifications. 22 The 2017 Banff meeting identified an initial validated, consensus gene list with potential specific indications for molecular testing. 23 Importantly presented at this meeting was a new technology, Nanostring, which uses robust multiplex transcript quantitation from formalin‐fixed, paraffin‐embedded (FFPE) biopsies. The compelling advantage of NanoString is that it performs transcriptional analysis on routine histological samples allowing correlation of both histologic with molecular phenotypes on the same tissue. 1
4. CURRENT STATE OF MOLECULAR TRANSPLANT DIAGNOSTICS
Most of the published research studies for molecular testing on biopsies has been performed using microarrays on an extra biopsy core stored in RNAlater Stabilization Solution. The pioneering work by Halloran and colleagues was the basis of a commercial test (Molecular Microscope MMDx) now offered by One Lambda Inc. 17 , 24 , 25 , 26 These insightful, prospective studies showed strong associations of transcript patterns with the histological Banff lesions and diagnosis but also identified discrepancies. 17 These discrepancies require further investigation to reveal the optimal integration of histology and molecular biopsy features that are informative of outcome and response to therapy. No prospective randomized outcome trial using microarray assays as the end point has been conducted, in part because of the technical challenges and the long follow‐up required. Although microarray analysis is the most established method for biopsies, alternative approaches, less invasive than a biopsy, are attractive and under investigation, such as urine and blood transcript analysis.
Recently, more practical technologies based on FFPE biopsy analysis are now available, in particular the NanoString nCounter system (NanoString Technologies, Seattle, WA). Several NanoString publications using FFPE transplant specimens identify similar transcript associations with the molecular and histologic phenotypes as those reported in microarray studies. 3 , 4 , 13 , 14 , 15 , 16 , 17 , 18 , 27 , 28 , 29 , 30 , 31 , 32 , 33 Among the advantages of NanoString are (1) a separate core processed at the time of biopsy is not required; (2) transcripts are assessed in the same sample analyzed by light microscopy; and (3) large retrospective and longitudinal analyses of archived samples can be readily performed in the setting of multicenter studies, which will enable retrospective randomization with long‐term survival end points available (Table 1). 27 Over 1000 publications have reported its application and value. The NanoString system yields comparable results between FFPE and fresh frozen samples, with a higher sensitivity than that of microarrays and about equal to reverse transcription polymerase chain reaction (RT‐PCR). 34 , 35 , 36 This technology in one assay uses color‐coded molecular barcodes that can hybridize directly up to 800 different targets with highly reproducibility. NanoString thereby closes a gap between genome‐wide expression (ie, microarrays and RNA sequencing as whole transcriptome discovery platforms) and mRNA expression profiling of a single target (ie, RT‐PCR). But unlike quantitative RT‐PCR, the NanoString system does not require enzymes and uses a single reaction per sample regardless of the level of multiplexing. Thus, it is simpler for the user and requires less sample per experiment for multiplex experiments, for example, pathway analysis, assessment of biomarker panels, or assessment of custom‐made gene sets. The NanoString system is approved for clinical diagnostics and paired with user‐friendly analytical software, thus representing a simple, relatively fast (24‐hour turnaround time), automated platform that is well poised for integration into the routine diagnostic workflows in existing pathology laboratories. 37 Synthetic DNA standard oligonucleotides, corresponding to each target probe in the panel, allow normalization of expression results between different reagent batches, platforms, and users, This permits standardization of diagnostic thresholds across multiple laboratories, a major challenge using microarrays and RNA sequencing. 27 A major disadvantage of the NanoString approach is the need to predefine the gene panel and the restriction to 800 probes, making it better for follow‐up studies once the discovery phase with microarrays has winnowed the possibilities to the most informative transcripts. The other disadvantages, shared with microarrays and RNASeq, is the loss of anatomic localization and the need for a biopsy.
TABLE 1.
Technical comparison of gene expression analysis using formalin‐fixed paraffin‐embedded (FFPE) tissue with NanoString nCounter vs fresh tissue with DNA microarrays
| Feature | FFPE tissue with NanoString nCounter | Fresh tissue with cDNA microarrays |
|---|---|---|
| Maximum number of transcript targets | 800 | >47 000 a |
| Off‐the‐shelf panels available | Yes | Yes |
| Custom panels available | Yes | Yes |
| Recommended RNA input quantity | 100 ng | 50‐500 ng |
| Requires reverse transcription/amplification | No | Yes |
| Approximate assay turnaround time b | 24‐40 h | 25.5‐37.5 h |
| Analysis software provided by manufacturer | Yes c | Yes d |
| Ability to use same sample for histology and gene expression analysis, that is, ability for histomolecular integration | Yes | No |
| Immediate access to long‐term clinical follow‐up data on archival clinical samples (FFPE) | Yes | No |
| Food and Drug Administration approved |
Yes for platform Yes for specific clinical assays e |
No for platform Yes for specific clinical assay f |
| Approximate assay cost per sample g | $275 | $1000‐3000 |
| Integration with local (decentralized) clinical workflow | Simple due to local testing (no shipment of samples) on regulatory approved platform using simple open source analytics | Complex (shipment of sample to referral lab, no regulatory approval of platform, complex analytics) |
Affymetrix GeneChip Human Genome U133 Plus 2.0 Array.
Dependent on multiple variables: instrument settings, RNA input quantity, technician experience, etc. Time excludes RNA extraction time and sample shipment time if applicable.
NanoString nSolver Analysis Software.
Affymetrix Transcriptome Analysis Console Software.
NanoString Prosigna Breast Cancer Prognostic Gene Signature Assay.
Roche AmpliChip CYP450 Test, a pharmacogenetics assay to determine the genotype of two cytochrome P450 enzymes: 2D6 and 2C19.
Including RNA isolation but excluding instrument expenses and labor for RNA extraction. Reagent cost varies with number of transcript targets and samples. Microarrays costs vary on scale of economy by provider.
5. GENERATION OF A BANFF HUMAN ORGAN TRANSPLANT (B‐HOT) PANEL
The B‐HOT panel includes the validated genes found informative from major peer reviewed microarray and NanoString studies on kidney, heart, lung, and liver allograft biopsies, identified by the MDWG through literature review. A list of the genes with corresponding key publications is given in the Data S1. In detail, candidate genes were identified using the key words “transplantation,” “kidney, “heart, ” “lung, ” ‘liver, ” “gene expression, ” “molecule, ” and “transcripts. ” Mining these publications for genes listed as significantly associated with any study variable revealed 2521 publications indexed in PubMed concerning more than 4000 genes. After redundant and duplicate genes were removed, the list contained 1749 genes. Then the MDWG members identified overlap between these genes and genes described in the peer‐reviewed literature 2 , 8 , 9 , 10 , 11 , 12 , 29 , 32 , 33 , 38 , 39 , 40 , 41 , 42 , 43 , 44 , 45 , 46 , 47 , 48 , 49 , 50 , 51 , 52 , 53 , 54 , 55 , 56 , 57 , 58 , 59 , 60 , 61 , 62 , 63 , 64 , 65 as being strongly associated with relevant clinical phenotypes and identified 1050 genes to be considered for inclusion. In the next step, a list including all genes with consensus expert opinion were selected and for which all Hugo duplicates were then combined, leaving 670 unique genes.
We initiated discussions with NanoString and learned they would be willing to make our panel widely available. However, their commercial panels typically have 770 genes, so they provided suggestions for addition genes to delineate relevant cellular pathways and cell types that have been used in other panels. Using an independent data‐driven process, NanoString Technologies Inc recommended additional genes within relevant molecular pathways related to the 670 genes that were most informative by their Ingenuity Pathways. The final B‐HOT panel included 758 genes covering the most pertinent genes from the core pathways and processes related to host responses to rejection of transplanted tissue, tolerance, drug‐induced toxicity, transplantation‐associated viral infections (BK polyomavirus, cytomegalovirus, Epstein‐Barr virus) plus 12 internal reference genes for quality control and normalization (Figures 1 and 2, Table 2).Through that approach the B‐HOT gene panel was defined, further engineered, and made commercially available (https://www.NanoString.com/products/gene‐expression‐panels/gene‐expression‐panels‐overview/human‐organ‐transplant‐panel). The pathways added to the list are given in Figure 2 and in more detail in the Table S1.
FIGURE 1.
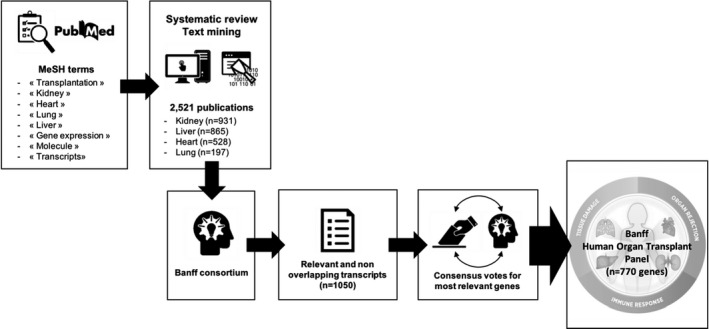
Banff Human Organ Transplant (B‐HOT) panel design process and main pathways investigated by this panel. Banff Human Organ Transplant (B‐HOT) panel design process involved 12 transplant expertsfrom 5 universities (Harvard University, Université de Paris, University of Alberta, Imperial College of London, and Erasmus MC Rotterdam). Banff consortium was composed of B. Colvin, R.N. Smith, I. Rosales, M. Mengel, B. Adam, C. Roufosse, M.C. Clahsen‐van Groningen, J.H. von der Thüsen, B. Robin, J. Dagobert, J.‐P. Duong‐van‐Huyen, and A. Loupy. The Banff Human Organ Transplant Panel logo in Figure 1 has been reproduced with permission from NanoString
FIGURE 2.
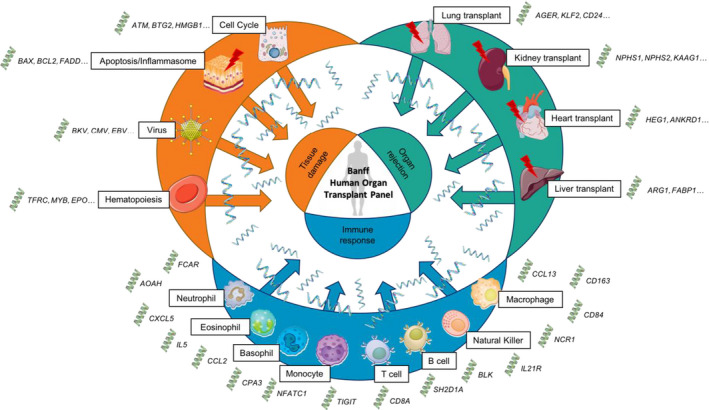
Examples of cells, pathways, and genes studied by the B‐HOT panel. Three main pathways can be identified: tissue damage, organ rejection, and immune response. The B‐HOT panel profiles a total of 758 genes across 37 pathways. Green double‐stranded DNA represents gene expression, blue single‐stranded RNA represents RNA expressed by cells or tissue. Cartoons of organs, cells, and other illustrations used in Figure 2 have been retrieved from http://smart.servier.com/, a free medical images bank of Servier
TABLE 2.
List of the 770 genes integrated in the HOT panel and their related pathways. Four groups (Tissue and cellular process, Immune system, Organ specific, Viral infection) and 17 subgroups define the genes. Twelve genes are used for internal reference. Genes can possibly be related to other pathway or involved in several processes
| Tissue and cellular process | Immune system | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Angiogenesis | CDH13 | JAK1 | PTGER4 | TIMP1 | Adaptive Immune System | Chemokine Signaling | CD209 | HFE | NFKB1 |
| ADAMTS1 | CDH5 | JAK2 | PTGS2 | TIPARP | AIRE | ACKR1 | CD83 | ICAM1 | NLRC5 |
| ADGRL4 | CDKN1A | KDR | PTPN2 | TM4SF1 | BLNK | CCL4 | CSF1 | ICAM2 | NOD2 |
| ENG | CGAS | KIT | PTPN22 | TM4SF18 | BST2 | CCL5 | CSF3R | IFI44 | NOS2 |
| ERG | CHCHD10 | KITLG | PTPN6 | TMEM178A | BTK | CCR2 | FCER1A | IFNG | OASL |
| MMRN2 | CITED4 | KLF2 | PTPRO | TNC | CCR7 | CCR4 | FCGR2A | IFNGR1 | OSMR |
| VEGFA | CLEC4C | KLF4 | RAB40C | TNFAIP6 | CD19 | CCR5 | FCGR3A/B | IFNGR2 | PAX5 |
| VEGFC | COL13A1 | KLHL13 | RAF1 | TNFRSF1A | CD22 | CMKLR1 | GNLY | IKBKB | PDCD1 |
| VWF | COL1A1 | LAMP1 | RAMP3 | TP53 | CD247 | CX3CL1 | GZMH | IKBKG | PDPN |
| Apoptosis | COL3A1 | LAYN | RAPGEF5 | TPMT | CD274 | CX3CR1 | GZMK | IKZF1 | PECAM1 |
| BAX | COL4A1 | LCN2 | RARRES1 | TPSAB1/B2 | CD276 | CXCL1/2 | IFI27 | IL10 | PIK3CD |
| BCL2 | COL4A3 | LEF1 | RASIP1 | TRAF6 | CD28 | CXCL10 | IFNA1 | IL10RB | PIK3CG |
| BCL2A1 | COL4A4 | LHX6 | RASSF9 | TRIM22 | CD3D | CXCL11 | IL1B | IL12A | POU2AF1 |
| BCL2L1 | COL4A5 | LIF | RELA | VCAN | CD3E | CXCL12 | IL33 | IL12B | PPBP |
| BCL2L11 | CRIP2 | LOX | RGN | VMP1 | CD3G | CXCL13 | KLRB1 | IL12RB2 | PRF1 |
| BIRC3 | CSF2RB | LRP2 | RHOJ | WARS | CD4 | CXCL2 | KLRC1 | IL13 | PTPN7 |
| CASP1 | CTNNB1 | LRRC32 | RHOU | WNT9A | CD40LG | CXCL5 | KLRD1 | IL15 | PTPRC |
| CASP3 | CTSL | LTBR | RNF149 | ZEB1 | CD45R0 | CXCL8 | KLRG1 | IL16 | PVR |
| CASP4 | DCAF12 | LYVE1 | ROBO4 | Hematopoiesis | CD45RA | CXCL9 | KLRK1 | IL17F | SELL |
| CASP8 | DDX50 | MAF | RORA | CD34 | CD45RB | CXCR3 | NKG7 | IL17RC | SELPLG |
| CFLAR | DNMT1 | MALL | RORC | CSF2 | CD7 | CXCR4 | NOD1 | IL1A | SERINC5 |
| FADD | DNMT3A | MAP3K1 | RPL19 | EPO | CD72 | CXCR6 | PSTPIP1 | IL1R1 | SIGIRR |
| FAS | DUSP2 | MAPK11 | RPS6 | FLT3 | CD79A | PF4 | SAMHD1 | IL1R2 | SIGLEC5 |
| FASLG | ECSCR | MAPK12 | RPS6KB1 | GATA3 | CD86 | Complement System | TAPBP | IL1RAP | SLAMF6 |
| GIMAP5 | EDA | MAPK13 | RTN4 | IKZF2 | CD8A | C1QA | TLR2 | IL1RN | SLAMF7 |
| IFI6 | EEF1A1 | MAPK14 | RXRA | IL12RB1 | CD8B | C1QB | TLR3 | IL21 | SLAMF8 |
| NLRP3 | EGFR | MAPK3 | S100A12 | IL5 | CTLA4 | C1S | TLR4 | IL21R | SLPI |
| RGS5 | EGR1 | MAPK8 | S100A8 | IL6 | CXCR5 | C3 | TLR5 | IL23A | SMAD5 |
| TNFRSF1B | EHD3 | MARCH8 | S100A9 | IL7 | FAM30A | C3AR1 | TLR7 | IL23R | SOCS1 |
| TNFRSF4 | EMP3 | MCM6 | S100B | LCK | FCAR | C5 | TLR8 | IL27 | SOCS3 |
| TNFSF10 | EPAS1 | MEF2C | S1PR1 | MYB | GZMB | C5AR1 | TLR9 | IL27RA | STAT4 |
| XAF1 | ERRFI1 | MEGF11 | SCGB1A1 | RUNX1 | HLA‐A | C9 | TREM1 | IL2RA | STAT6 |
| CellProcess | EVA1C | MEOX1 | SDC1 | TFRC | HLA‐B | CD46 | Other Immune Genes | IL2RG | TBX21 |
| ABCB1 | EZH2 | MERTK | SELP | Metabolism | HLA‐C | CD55 | ACVRL1 | IL4R | TCF7 |
| ABCC2 | F3 | MET | SEMA7A | ABCA1 | HLA‐DMA | CD59 | ADAMDEC1 | IL6R | TCL1A |
| ABCE1 | FGD2 | MIR155HG | SERPINA3 | ALDH3A2 | HLA‐DMB | CFB | AGER | IL6ST | TIGIT |
| ACVR1 | FKBP1A | MMP12 | SERPINE1 | ALOX15 | HLA‐DPA1 | CFH | BCL6 | IL7R | TNFRSF14 |
| ADAM8 | FN1 | MMP14 | SERTAD1 | APOE | HLA‐DPB1 | CFI | BTLA | INPP5D | TNFRSF9 |
| ADORA2A | FOS | MMP9 | SHROOM3 | APOL1 | HLA‐DQA1 | CR1 | CALHM6 | IRF1 | TNFSF14 |
| AGR2 | FOSL1 | MT1A | SIRPG | APOL2 | HLA‐DQB1 | MASP1 | CCL2 | IRF4 | TNFSF18 |
| AGR3 | FOXO1 | MT2A | SKI | ARG2 | HLA‐DRA | MASP2 | CCL21 | IRF6 | TNFSF9 |
| AGT | FOXP3 | MTOR | SLA | B3GAT1 | HLA‐DRB1 | MBP | CCR3 | IRF8 | TOX2 |
| AHR | FPR1 | MUC1 | SLC11A1 | CAV1 | HLA‐DRB3 | SERPING1 | CD160 | ITGAM | TRIB1 |
| AICDA | FYN | MX2 | SLC19A3 | CETP | HLA‐E | Inflammatory Response | CD163 | ITGAX | TYK2 |
| AIM2 | GBP1 | MYBL1 | SLC22A2 | CH25H | HLA‐F | ALOX5 | CD1D | JAK3 | VCAM1 |
| AKR1C3 | GBP2 | MYC | SLC25A15 | CRHBP | HLA‐G | ANXA1 | CD2 | KIR_Activating_Subgroup_1 | VSIR |
| ALAS1 | GBP4 | NFIL3 | SLC4A1 | GAPDH | ICOS | AOAH | CD24 | KIR_Activating_Subgroup_2 | XCL1/2 |
| ANKRD1 | GDF15 | NOS3 | SMAD2 | HSD11B1 | ICOSLG | CARD16 | CD244 | KIR_Inhibiting_Subgroup_1 | |
| ANKRD22 | GEMIN7 | NOTCH1 | SMAD3 | IDO1 | IFI30 | CARD8 | CD27 | KIR_Inhibiting_Subgroup_2 | |
| APOLD1 | GNG11 | NOTCH2 | SMAD4 | IGF1 | IGHA1 | CCL13 | CD40 | KIR3DL1 | VIRAL INFECTION |
| AQP1 | HAVCR1 | NOX4 | SMARCA4 | LDLR | IGHG1 | CCL15 | CD48 | KIR3DL2 | |
| AREG | HDAC3 | NPDC1 | SOD2 | NNMT | IGHG2 | CCL18 | CD5 | KLRF1 | Virus |
| ARG1 | HDAC6 | NPPA | SOST | PLA1A | IGHG3 | CCL19 | CD58 | LAG3 | BK large T Ag |
| ARHGDIB | HDC | NPPB | SOX7 | IGHG4 | CCL20 | CD6 | LAIR1 | BK VP1 | |
| ARRB2 | HEG1 | NR4A1 | SP100 | IGHM | CCL22 | CD68 | LAP3 | CMV UL83 | |
| ASB15 | HIF1A | OR2I1P | SP140 | ORGAN SPECIFIC | IGKC | CCL3/L1 | CD69 | LGALS3 | EBV LMP2 |
| ATF3 | HK2 | P2RX4 | SPIB | IGLC1 | CCR10 | CD70 | LILRB1 | Viral Detection Genes | |
| ATM | HMGB1 | PADI4 | SPRY4 | Heart | IL17RA | CRP | CD74 | LILRB2 | EBI3 |
| ATXN3 | HPRT1 | PALMD | SRC | ACTA2 | IL2 | GBP5 | CD80 | LILRB4 | IFITM3 |
| AXL | HSP90AA1 | PDCD1LG2 | ST5 | MYL9 | IL2RB | IL10RA | CD84 | LST1 | IRF7 |
| BASP1 | HSPA12B | PDGFA | ST8SIA4 | TRDN | IL4 | IL17A | CD96 | LTA | ISG20 |
| BATF | HYAL1 | PDGFRB | STAT1 | Kidney | LCP2 | IL17RB | CEACAM3 | LTB | JUN |
| BATF3 | HYAL2 | PHEX | STAT3 | AQP2 | NFATC1 | IL18 | CHUK | LTF | MX1 |
| BDNF | IER5 | PIN1 | STAT5A | KAAG1 | NFATC2 | IL18BP | CIITA | LY96 | |
| BLK | IFIT1 | PLAAT4 | STAT5B | NPHS1 | RAG2 | IL18RAP | CPA3 | MCAM | |
| BMP2 | IFITM1 | PLAT | SYK | NPHS2 | REL | IL1RL1 | CSF3 | MICA | INTERNAL REFERENCE GENES |
| BMP4 | IFITM2 | PLAU | TANK | SLC12A3 | RELB | IL22 | CTSS | MICB | |
| BMP6 | IFNAR1 | PLAUR | TAP1 | UMOD | SELE | NFKB2 | CTSW | MIF | ABCF1 |
| BMP7 | IFNAR2 | PLK2 | TAP2 | Liver | SH2D1A | NFKBIA | CXCL14 | MME | G6PD |
| BMPER | IGF1R | PNOC | TBK1 | FABP1 | SH2D1B | NFKBIZ | CXCL16 | MPIG6B | GUSB |
| BMPR1A | IGF2R | PPM1F | TEK | HNF1A | THEMIS | PTX3 | DEFB1 | MRC1 | NRDE2 |
| BMPR1B | IGFL1 | PPP3CA | TFF3 | IGFBP1 | TNFRSF17 | TNF | EOMES | MS4A1 | OAZ1 |
| BRWD1 | IMPDH1 | PRDM1 | TGFB1 | KRT19 | TNFRSF18 | TNFAIP3 | FCER1G | MS4A2 | POLR2A |
| BTG2 | IMPDH2 | PROX1 | TGFB2 | KRT8 | TNFSF4 | TRAF4 | FCGR1A | MS4A4A | PPIA |
| CD207 | INHBC | PSEN1 | TGFBI | Lung | TNFSF8 | Innate Immune System | FCGR2B | MS4A6A | SDHA |
| CD38 | IRS1 | PSMB10 | TGFBR1 | MYOM2 | TRAT1 | B2M | FCRL2 | MS4A7 | STK11IP |
| CD44 | ISG15 | PSMB8 | TGFBR2 | SFTPA2 | TRDC | BCL3 | FGFBP2 | MYD88 | TBC1D10B |
| CD47 | ITGA4 | PSMB9 | TGIF1 | SFTPB | TRDV3 | CCR1 | FJX1 | NCAM1 | TBP |
| CD81 | ITGB2 | PSME1 | THBD | SFTPC | XBP1 | CCR6 | GZMA | NCR1 | UBB |
| CD82 | TGB6 | PSME2 | THBS1 | SFTPD | ZAP70 | CD14 | HAVCR2 | NFAM1 | |
The panel probes were also designed to cover different organ types for transplantation and for sequence homology with nonhuman primates to facilitate preclinical research applications. The panel's broad coverage of inflammatory, adaptive, and innate immune systems; signaling; and endothelial transcripts will likely be largely applicable across organ types but with some expected organ specific variation. Furthermore, parenchymal transcripts will often be organ specific and many have been included (see Table S1). We anticipate that continued discovery of other informative transcripts not included in the B‐HOT panel will occur. To provide flexibility, up to 30 custom genes can be added to the B‐HOT panel by an investigator. Although the panel has been commercialized for the nCounter platform, the gene list is not proprietary and probes based on the gene list can be designed to run on any transcript analytical platform.
6. NEXT STEPS: MULTICENTER ANALYTICAL AND CLINICAL VALIDATION
The Banff MDWG formed a voluntary, growing, and open international consortium, independent of commercial sponsorship, to develop future steps for validation, analyses, and database sharing. The focus of the next 2 years will be validation of the panel and discovery of the optimal algorithms and gene sets. This will be enabled by (1) the B‐HOT panel and its comprehensive probe standards for comparison between laboratories, batches, and runs; (2) a shared database containing clinical, laboratory, pathological and transcript data; and (3) access to comprehensive sophisticated bioinformatics. The next steps will be to document the analytical validity across laboratories and then determine the clinical validity. The clinical validity will be assessed by analyzing B‐HOT transcripts in 1000 or more clinical biopsies (as of this report the consortium has run the B‐HOT panel on over 600 samples). These results along with standardized clinical and pathologic information will be entered in a shared database, which will be interrogated to discover the most useful algorithms for clinical applications.
Analytical validation for regulatory approval must document accuracy, precision, analytical sensitivity (reproducibility, coefficient of variance), reportable ranges, reference interval values, and analytical specificity. Calibration and control procedures must be determined, and the laboratory must be enrolled in external proficiency testing programs. Clinical validation is the next step. Even an assay with perfect analytical validity does not automatically imply association between the test result and a relevant clinical outcome or action. This requires access to relevant patient populations’ material of adequately powered sample size to evaluate assay performance in a real‐world clinical setting. Accordingly, clinical utility of an assay needs to be established by providing evidence of improved, measurable clinical outcome or benefit that is directly related to the use of the test, that is, proof that the test adds significant value to patient care. This also needs to take into consideration how the assay is interpreted, reported, and applied in the context of clinical patient management. Ideally, proper evaluation of an assay's clinical utility requires prospective randomized control trials. 66
The B‐HOT panel will undergo all of these validation steps. In the next 2 years retrospective, well‐annotated cohorts will be analyzed for analytical and clinical validation. The MDWG is aligning joint efforts using available NanoString systems at participating centers for studying a broad spectrum of archived and well‐annotated transplant biopsies. To centralize the resulting multicenter molecular data from archived transplant biopsies together with the related clinical and outcome data, algorithms, and tools for analysis (including explorative analytics, machine learning‐based diagnostic approaches/classifiers, and risk prediction tools) with remote access by users across the world, a data integration platform (DIP) will be built 67 (Figure 3). Participating centers will be able to upload routinely collected transplant‐related patient data in an anonymized and uniform fashion. A participating investigator will then be able to use all data in the DIP. Currently underway is the development of a consensus data template representing the variables and units to be included in the DIP. The NanoString data files also include important analytical parameters (quality control measures, background subtractions, normalization values) in addition to the individual gene expression values, which will also be part of the DIP to allow for standardization across laboratories and thus multicenter analytical validation of any diagnostic assays. The output of this effort is expected to be a robust well‐characterized gene set (presumably a subset of the B‐HOT panel or additional genes) and analytic methodology for interpretation, which will be presented at a subsequent Banff meeting and published. We expect to see correlations with histologic diagnosis (including interpretations not revealed by routine pathology analysis), ongoing immunosuppressive therapy, prediction of outcome, and response to treatment. We (and others, we hope) will follow this by prospective, controlled clinical trials to fully define clinical utility.
FIGURE 3.
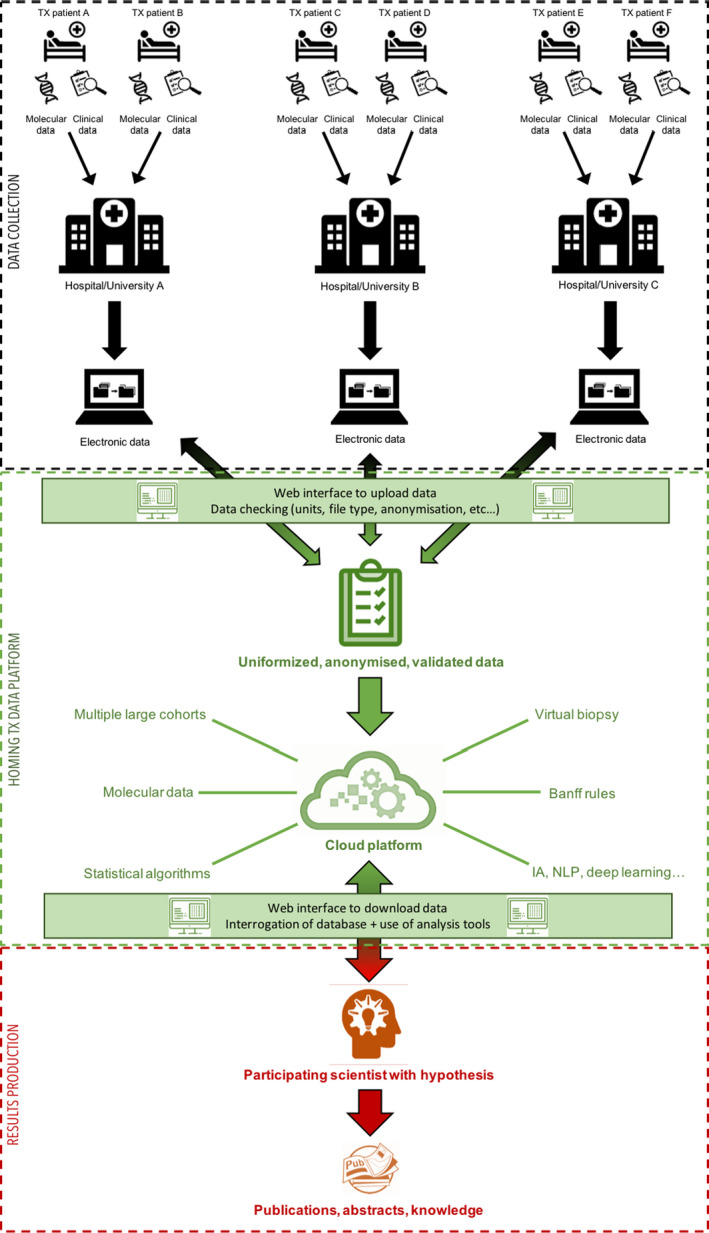
Data integration platform (DIP) design. Three elements are identified: (1) data production (histology, molecular, and clinical) by participating hospital; (2) DIP (web interface, cloud computing) to centralize, check, and validate all data; and (3) results production by any participating physician/scientist using built in analytical tools
As a first evaluation, after the Banff meeting, a member of the MDWG, Neal Smith, performed an in silico assessment of the B‐HOT panel genes using the archived Genomic Spatial Event databases from Halloran's group 5 , 46 , 68 that contains 764 kidney biopsy samples with microarray data and diagnostic classification as TCMR, chronic‐active ABMR, mixed, acute kidney injury, no rejection, and normal. Briefly, 3 bioinformatics methods were used to see if they could identify the 6 diagnostic groups from the transcripts: (1) supervised, using diagnostic and pathogenesis based transcripts sets of Halloran; 16 (2) semisupervised, using Nanostring pathways (Data S1) plus CIBERSORT cells types; and (3) unsupervised principal component analysis. Results confirmed the correlation of expected gene sets in each analysis with the 6 diagnostic categories (Smith, manuscript in preparation). A description of the initial B‐HOT results in kidney transplants to be presented at the 2020 American Transplant Conference reveals both expected and novel correlations with pathologic categories. 69
The B‐HOT panel will be commercially available for research use only. Whether B‐HOT leads to a clinically indicated laboratory developed test remains to be seen. If it does, it will probably be a simplified panel. In the future, the international, open source, multicenter Banff DIP can serve as a reference point for generating a molecular diagnostic “gold‐standard” in transplantation, similar to the Banff histology lesions and diagnoses agreed upon in 1991. 70 As the Banff consensus rules for histology underwent refinement over the last 28 years as new knowledge emerged, any molecular “consensus” will also need to undergo constant refinement and, no doubt further, technological innovation. Only through integration with clinical decision‐making and end points in clinical trials can the true clinical utility of molecular diagnostics be demonstrated. 67
DISCLOSURE
The authors of this manuscript have conflicts of interest to disclose as described by the American Journal of Transplantation. Michael Mengel received honoraria from Novartis, CSL Behring, Vitaeris. Mark Haas received consulting fees from Shire ViroPharma, AstraZeneca, Novartis, and CareDx, and honoraria from CareDx. Robert Colvin is a consultant for Shire ViroPharma, CSL Behring, Alexion and eGenesis. Candice Roufosse has received consulting fees from Achillion and UCB. Ivy Rosales is a consultant for eGenesis. Enver Akalin received honorarium and research grant support from CareDx. Marian Clahsen‐van Groningen received grant support from Astellas Pharma (paid to the Erasmus MC). A. Jake Demetris receives research support from Q2 Solutions and is a member of an Adjudication Committee for Novartis. None of these conflicts are relevant to this article. The other authors have no conflicts of interest to disclose. None of the authors has a financial interest in NanoString.
Supporting information
Supplementary Material
Supplementary Material
ACKNOWLEDGMENTS
The 2019 Banff meeting received sponsorship from CareDx, CSL Behring, Elsevier, Eppendorf, GenDx, Hansa Biopharma, Histogenetics, Immucor, Omion, OneLambda, NanoString, Novartis, Takeda, Veloxis, and Vitaeris.
Mengel M, Loupy A, Haas M, et al. Banff 2019 Meeting Report: Molecular diagnostics in solid organ transplantation–Consensus for the Banff Human Organ Transplant (B‐HOT) gene panel and open source multicenter validation. Am J Transplant. 2020;20:2305–2317. 10.1111/ajt.16059
M. Mengel, A. Loupy, B. Adam, and R.B. Colvin contributed equally to this report.
Contributor Information
Michael Mengel, Email: mmengel@ualberta.ca.
Alexandre Loupy, Email: alexandre.loupy@inserm.fr.
DATA AVAILABILITY STATEMENT
Data sharing is not applicable to this article as no new data were created or analyzed in this study.
REFERENCES
- 1. Loupy A, Haas M, Solez K, et al. The Banff 2015 Kidney Meeting Report: current challenges in rejection classification and prospects for adopting molecular pathology. Am J Transplant. 2017;17:28‐41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Halloran PF, Venner JM, Madill‐Thomsen KS, et al. Review: the transcripts associated with organ allograft rejection. Am J Transplant. 2018;18:785‐795. [DOI] [PubMed] [Google Scholar]
- 3. Halloran PF, Matas A, Kasiske BL, et al. Molecular phenotype of kidney transplant indication biopsies with inflammation in scarred areas. Am J Transplant. 2019;19:1356‐1370. [DOI] [PubMed] [Google Scholar]
- 4. Halloran PF, Reeve J, Akalin E, et al. Real time central assessment of kidney transplant indication biopsies by microarrays: the INTERCOMEX Study. Am J Transplant. 2017;17:2851‐2862. [DOI] [PubMed] [Google Scholar]
- 5. Reeve J, Sellarés J, Mengel M, et al. Molecular diagnosis of T cell‐mediated rejection in human kidney transplant biopsies. Am J Transplant. 2013;13:645‐655. [DOI] [PubMed] [Google Scholar]
- 6. Sirota M, Sarwal MM. Transplantomics: toward precision medicine in transplantation research. Transplantation. 2017;101:1777‐1782. [DOI] [PubMed] [Google Scholar]
- 7. Vitalone MJ, Sigdel TK, Salomonis N, et al. Transcriptional perturbations in graft rejection. Transplantation. 2015;99:1882‐1893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Mengel M, Sis B, Kim D, et al. The molecular phenotype of heart transplant biopsies: relationship to histopathological and clinical variables. Am J Transplant. 2010;10:2105‐2115. [DOI] [PubMed] [Google Scholar]
- 9. Loupy A, Duong Van Huyen JP, Hidalgo L, et al. Expression profiling for the identification and classification of antibody‐mediated heart rejection. Circulation. 2017;135:917‐935. [DOI] [PubMed] [Google Scholar]
- 10. Dromparis P, Aboelnazar NS, Wagner S, et al. Ex vivo perfusion induces a time‐ and perfusate‐dependent molecular repair response in explanted porcine lungs. Am J Transplant. 2019;19:1024‐1036. [DOI] [PubMed] [Google Scholar]
- 11. Halloran KM, Parkes MD, Chang J, et al. Molecular assessment of rejection and injury in lung transplant biopsies. J Heart Lung Transplant. 2019;38:504‐513. [DOI] [PubMed] [Google Scholar]
- 12. Bonaccorsi‐Riani E, Pennycuick A, Londoño M‐C, et al. Molecular characterization of acute cellular rejection occurring during intentional immunosuppression withdrawal in liver transplantation. Am J Transplant. 2016;16:484‐496. [DOI] [PubMed] [Google Scholar]
- 13. Feng S, Bucuvalas JC, Demetris AJ, et al. Evidence of chronic allograft injury in liver biopsies from long‐term pediatric recipients of liver transplants. Gastroenterology. 2018;155(6):1838‐1851.e7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Naesens M, Sarwal MM. Molecular diagnostics in transplantation. Nat Rev Nephrol. 2010;6:614‐628. [DOI] [PubMed] [Google Scholar]
- 15. Haas M, Sis B, Racusen LC, et al. Banff 2013 meeting report: inclusion of C4d‐negative antibody‐mediated rejection and antibody‐associated arterial lesions. Am J Transplant. 2014;14:272‐283. [DOI] [PubMed] [Google Scholar]
- 16. Halloran PF, De Freitas DG, Einecke G, et al. The molecular phenotype of kidney transplants. Am J Transplant. 2010;10:2215‐2222. [DOI] [PubMed] [Google Scholar]
- 17. Madill‐Thomsen K, Perkowska‐Ptasińska A, Böhmig GA, et al. Discrepancy analysis comparing molecular and histology diagnoses in kidney transplant biopsies. Am J Transplant. 2020;20:1341–1350. [DOI] [PubMed] [Google Scholar]
- 18. Mengel M, Sis B, Halloran PF. SWOT analysis of Banff: strengths, weaknesses, opportunities and threats of the international Banff consensus process and classification system for renal allograft pathology. Am J Transplant. 2007;7:2221‐2226. [DOI] [PubMed] [Google Scholar]
- 19. Volk HD, Sawitzki B, Reinke P. Molecular analysis of renal allograft biopsies–more than a nice toy for researchers? Am J Transplant. 2013;13:539‐540. [DOI] [PubMed] [Google Scholar]
- 20. Jamshaid F, Froghi S, Di Cocco P, et al. Novel non‐invasive biomarkers diagnostic of acute rejection in renal transplant recipients: a systematic review. Int J Clin Pract. 2018;72(8):e13220. [DOI] [PubMed] [Google Scholar]
- 21. Abecassis M, Kaplan B. Transplantation: biomarkers in transplantation‐the devil is in the detail. Nat Rev Nephrol. 2015;11:204‐205. [DOI] [PubMed] [Google Scholar]
- 22. Naesens M, Friedewald J, Mas V, et al. A practical guide to the clinical implementation of biomarkers for subclinical rejection following kidney transplantation. Transplantation. 2020;104:700‐707. [DOI] [PubMed] [Google Scholar]
- 23. Haas M, Loupy A, Lefaucheur C, et al. The Banff 2017 Kidney Meeting Report: revised diagnostic criteria for chronic active T cell‐mediated rejection, antibody‐mediated rejection, and prospects for integrative endpoints for next‐generation clinical trials. Am J Transplant. 2018;18:293‐307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Halloran PF, Pereira AB, Chang J, et al. Microarray diagnosis of antibody‐mediated rejection in kidney transplant biopsies: an international prospective study (INTERCOM). Am J Transplant. 2013;13:2865‐2874. [DOI] [PubMed] [Google Scholar]
- 25. Halloran PF, Pereira AB, Chang J, et al. Potential impact of microarray diagnosis of T cell‐mediated rejection in kidney transplants: the INTERCOM study. Am J Transplant. 2013;13:2352‐2363. [DOI] [PubMed] [Google Scholar]
- 26. Reeve J, Böhmig GA, Eskandary F, et al. Generating automated kidney transplant biopsy reports combining molecular measurements with ensembles of machine learning classifiers. Am J Transplant. 2019;19:2719‐2731. [DOI] [PubMed] [Google Scholar]
- 27. Adam B, Afzali B, Dominy KM, et al. Multiplexed color‐coded probe‐based gene expression assessment for clinical molecular diagnostics in formalin‐fixed paraffin‐embedded human renal allograft tissue. Clin Transplant. 2016;30:295‐305. [DOI] [PubMed] [Google Scholar]
- 28. Dominy KM, Willicombe M, Al Johani T, et al. Molecular assessment of C4d‐positive renal transplant biopsies without evidence of rejection. Kidney Int Rep. 2019;4:148‐158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Adam BA, Smith RN, Rosales IA, et al. Chronic antibody‐mediated rejection in nonhuman primate renal allografts: validation of human histological and molecular phenotypes. Am J Transplant. 2017;17:2841‐2850. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Matsunami M, Rosales IA, Adam BA, et al. Long‐term kinetics of intragraft gene signatures in renal allograft tolerance induced by transient mixed chimerism. Transplantation. 2019;103:e334‐e344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. van der Zwan M, Baan CC, Colvin RB, et al. Immunomics of renal allograft acute T cell‐mediated rejection biopsies of tacrolimus‐ and belatacept‐treated patients. Transplant Direct. 2019;5:e418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Smith RN, Adam BA, Rosales IA, et al. RNA expression profiling of renal allografts in a nonhuman primate identifies variation in NK and endothelial gene expression. Am J Transplant. 2018;18:1340‐1350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Smith RN, Matsunami M, Adam BA, et al. RNA expression profiling of nonhuman primate renal allograft rejection identifies tolerance. Am J Transplant. 2018;18:1328‐1339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Geiss GK, Bumgarner RE, Birditt B, et al. Direct multiplexed measurement of gene expression with color‐coded probe pairs. Nat Biotechnol. 2008;26:317‐325. [DOI] [PubMed] [Google Scholar]
- 35. Payton JE, Grieselhuber NR, Chang L‐W, et al. High throughput digital quantification of mRNA abundance in primary human acute myeloid leukemia samples. J Clin Invest. 2009;119:1714‐1726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Goytain A, Ng T. NanoString nCounter technology: high‐throughput RNA validation. Methods Mol Biol. 2020;2079:125‐139. [DOI] [PubMed] [Google Scholar]
- 37. Nielsen T, Wallden B, Schaper C, et al. Analytical validation of the PAM50‐based Prosigna Breast Cancer Prognostic Gene Signature Assay and nCounter Analysis System using formalin‐fixed paraffin‐embedded breast tumor specimens. BMC Cancer. 2014;14:177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Venner JM, Famulski KS, Badr D, et al. Molecular landscape of T cell‐mediated rejection in human kidney transplants: prominence of CTLA4 and PD ligands. Am J Transplant. 2014;14:2565‐2576. [DOI] [PubMed] [Google Scholar]
- 39. Vitalone MJ, Ganguly B, Hsieh S, et al. Transcriptional profiling of belatacept and calcineurin inhibitor therapy in renal allograft recipients. Am J Transplant. 2014;14:1912‐1921. [DOI] [PubMed] [Google Scholar]
- 40. Sellarés J, Reeve J, Loupy A, et al. Molecular diagnosis of antibody‐mediated rejection in human kidney transplants. Am J Transplant. 2013;13:971‐983. [DOI] [PubMed] [Google Scholar]
- 41. Wherry EJ, Ha S‐J, Kaech SM, et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity. 2007;27:670‐684. [DOI] [PubMed] [Google Scholar]
- 42. Rebollo‐Mesa I, Nova‐Lamperti E, Mobillo P, et al. Biomarkers of tolerance in kidney transplantation: are we predicting tolerance or response to immunosuppressive treatment? Am J Transplant. 2016;16:3443‐3457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. O'Connell PJ, Zhang W, Menon MC, et al. Biopsy transcriptome expression profiling to identify kidney transplants at risk of chronic injury: a multicentre, prospective study. Lancet. 2016;388:983‐993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Menon MC, Chuang PY, Li Z, et al. Intronic locus determines SHROOM3 expression and potentiates renal allograft fibrosis. J Clin Invest. 2015;125:208‐221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Mengel M, Reeve J, Bunnag S, et al. Molecular correlates of scarring in kidney transplants: the emergence of mast cell transcripts. Am J Transplant. 2009;9:169‐178. [DOI] [PubMed] [Google Scholar]
- 46. Famulski KS, de Freitas DG, Kreepala C, et al. Molecular phenotypes of acute kidney injury in kidney transplants. J Am Soc Nephrol. 2012;23:948‐958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Lefaucheur C, Viglietti D, Hidalgo LG, et al. Complement‐activating anti‐HLA antibodies in kidney transplantation: allograft gene expression profiling and response to treatment. J Am Soc Nephrol. 2018;29:620‐635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Stegall MD, Park WD, Kim DY, et al. Changes in intragraft gene expression secondary to ischemia reperfusion after cardiac transplantation. Transplantation. 2002;74:924‐930. [DOI] [PubMed] [Google Scholar]
- 49. Deng MC, Eisen HJ, Mehra MR, et al. Noninvasive discrimination of rejection in cardiac allograft recipients using gene expression profiling. Am J Transplant. 2006;6:150‐160. [DOI] [PubMed] [Google Scholar]
- 50. Streblow DN, Kreklywich CN, Andoh T, et al. The role of angiogenic and wound repair factors during CMV‐accelerated transplant vascular sclerosis in rat cardiac transplants. Am J Transplant. 2008;8:277‐287. [DOI] [PubMed] [Google Scholar]
- 51. Afzali B, Chapman E, Racapé M, et al. Molecular assessment of microcirculation injury in formalin‐fixed human cardiac allograft biopsies with antibody‐mediated rejection. Am J Transplant. 2017;17:496‐505. [DOI] [PubMed] [Google Scholar]
- 52. Kearns MJ, Miller SD, Cheung A, et al. A rodent model of cardiac donation after circulatory death and novel biomarkers of cardiac viability during ex vivo heart perfusion. Transplantation. 2017;101:e231‐e239. [DOI] [PubMed] [Google Scholar]
- 53. Adam N, Coutance G, Viailly P‐J, et al. Reverse transcriptase multiplex ligation‐dependent probe amplification in endomyocardial biopsies for the diagnosis of cardiac allograft rejection. J Heart Lung Transplant. 2020;39:115‐124. [DOI] [PubMed] [Google Scholar]
- 54. Patil J, Lande JD, Li NA, et al. Bronchoalveolar lavage cell gene expression in acute lung rejection: development of a diagnostic classifier. Transplantation. 2008;85:224‐231. [DOI] [PubMed] [Google Scholar]
- 55. Yeung JC, Zamel R, Klement W, et al. Towards donor lung recovery‐gene expression changes during ex vivo lung perfusion of human lungs. Am J Transplant. 2018;18:1518‐1526. [DOI] [PubMed] [Google Scholar]
- 56. Sacreas A, Yang JYC, Vanaudenaerde BM, et al. The common rejection module in chronic rejection post lung transplantation. PLoS One. 2018;13:e0205107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Greenland JR, Wang P, Brotman JJ, et al. signatures common to allograft rejection are associated with lymphocytic bronchitis. Clin Transplant. 2019;33:e13515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Weigt SS, Wang X, Palchevskiy V, et al. Usefulness of gene expression profiling of bronchoalveolar lavage cells in acute lung allograft rejection. J Heart Lung Transplant. 2019;38:845‐855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Halloran K, Parkes MD, Timofte IL, et al. Molecular phenotyping of rejection‐related changes in mucosal biopsies from lung transplants. Am J Transplant. 2020;20:954‐966. [DOI] [PubMed] [Google Scholar]
- 60. Sreekumar R, Rasmussen DL, Wiesner RH, et al. Differential allograft gene expression in acute cellular rejection and recurrence of hepatitis C after liver transplantation. Liver Transpl. 2002;8:814‐821. [DOI] [PubMed] [Google Scholar]
- 61. Inkinen K, Lahesmaa R, Brandt A, et al. DNA microarray‐based gene expression profiles of cytomegalovirus infection and acute rejection in liver transplants. Transplant Proc. 2005;37:1227‐1229. [DOI] [PubMed] [Google Scholar]
- 62. Martínez‐Llordella M, Lozano JJ, Puig‐Pey I, et al. Using transcriptional profiling to develop a diagnostic test of operational tolerance in liver transplant recipients. J Clin Invest. 2008;118:2845‐2857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Gehrau RC, Mas VR, Dumur CI, et al. Regulation of molecular pathways in ischemia‐reperfusion injury after liver transplantation. Transplantation. 2013;96:926‐934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Kurian SM, Fouraschen SMG, Langfelder P, et al. Genomic profiles and predictors of early allograft dysfunction after human liver transplantation. Am J Transplant. 2015;15:1605‐1614. [DOI] [PubMed] [Google Scholar]
- 65. Londoño M‐C, Souza LN, Lozano J‐J, et al. Molecular profiling of subclinical inflammatory lesions in long‐term surviving adult liver transplant recipients. J Hepatol. 2018;69:626‐634. [DOI] [PubMed] [Google Scholar]
- 66. Burckart GJ, Amur S, Goodsaid FM, et al. Qualification of biomarkers for drug development in organ transplantation. Am J Transplant. 2008;8:267‐270. [DOI] [PubMed] [Google Scholar]
- 67. Loupy A, Aubert O, Orandi BJ,, et al. Prediction system for risk of allograft loss in patients receiving kidney transplants: international derivation and validation study. BMJ 2019;366:l4923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Halloran PF, Reeve JP, Pereira AB, et al. Antibody‐mediated rejection, T cell‐mediated rejection, and the injury‐repair response: new insights from the Genome Canada studies of kidney transplant biopsies. Kidney Int. 2014;85:258‐264. [DOI] [PubMed] [Google Scholar]
- 69. Rosales I, Smith RN, Acheampong E, et al. Routine human renal allograft biopsies analyzed for mRNA content with a novel Nanostring mRNA Human Organ Transplant (HOT) panel. Am J Transplant. 2020; Abstract to be presented at 2020 ATC. [Google Scholar]
- 70. Solez K, Axelsen RA, Benediktsson H, et al. International standardization of criteria for the histologic diagnosis of renal allograft rejection: the Banff working classification of kidney transplant pathology. Kidney Int. 1993;44:411‐422. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Material
Supplementary Material
Data Availability Statement
Data sharing is not applicable to this article as no new data were created or analyzed in this study.


